Arabinose Plays an Important Role in Regulating the Growth and Sporulation of Bacillus subtilis NCD-2
Abstract
1. Introduction
2. Results
2.1. Screening of Nutrients That Facilitate the Metabolism of Strain NCD-2
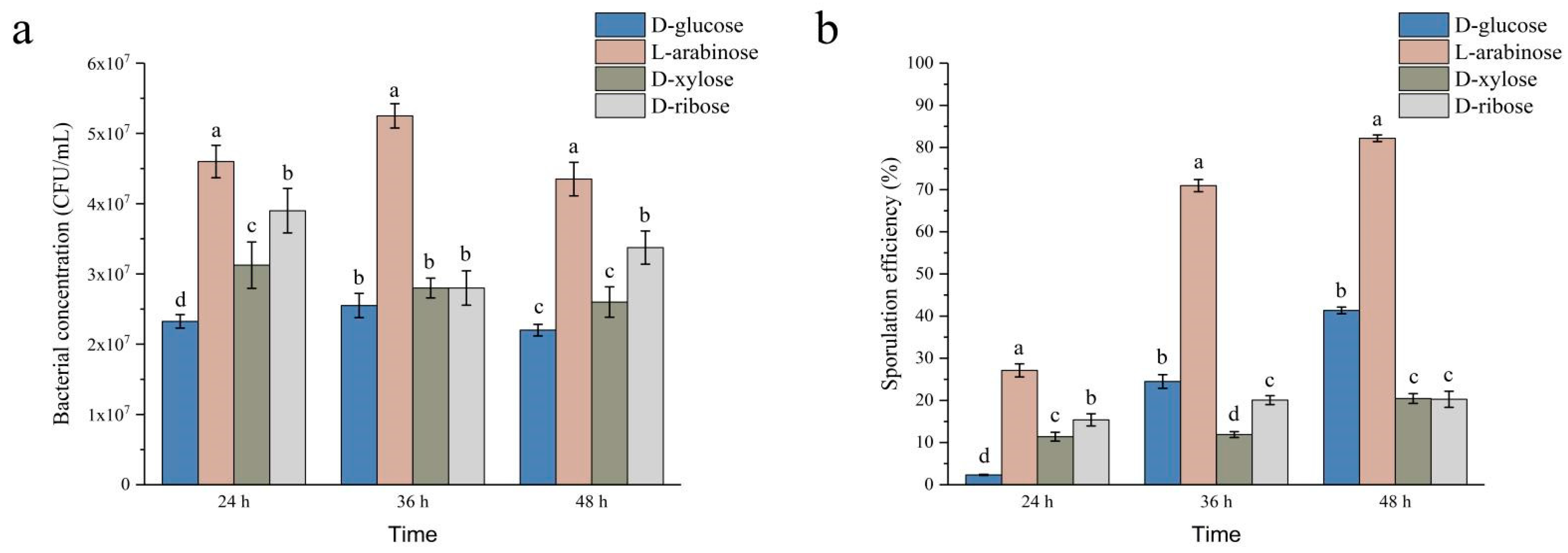
2.2. Effects of L-Arabinose, D-Ribose, and D-Xylose on Growth and Sporulation Efficiency of Strain NCD-2
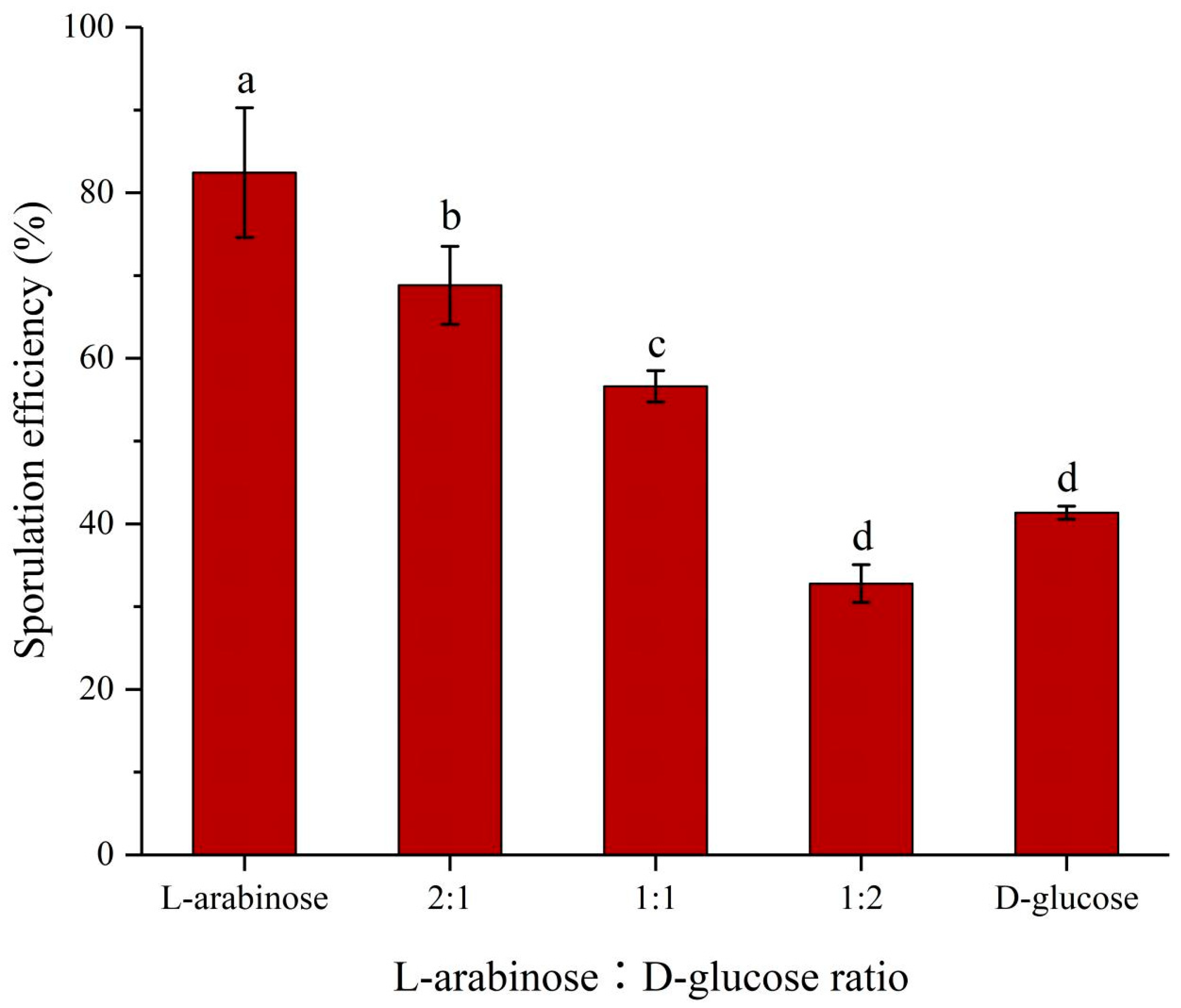
2.3. Effects of Different Proportions of L-Arabinose and D-Glucose on Sporulation of Strain NCD-2
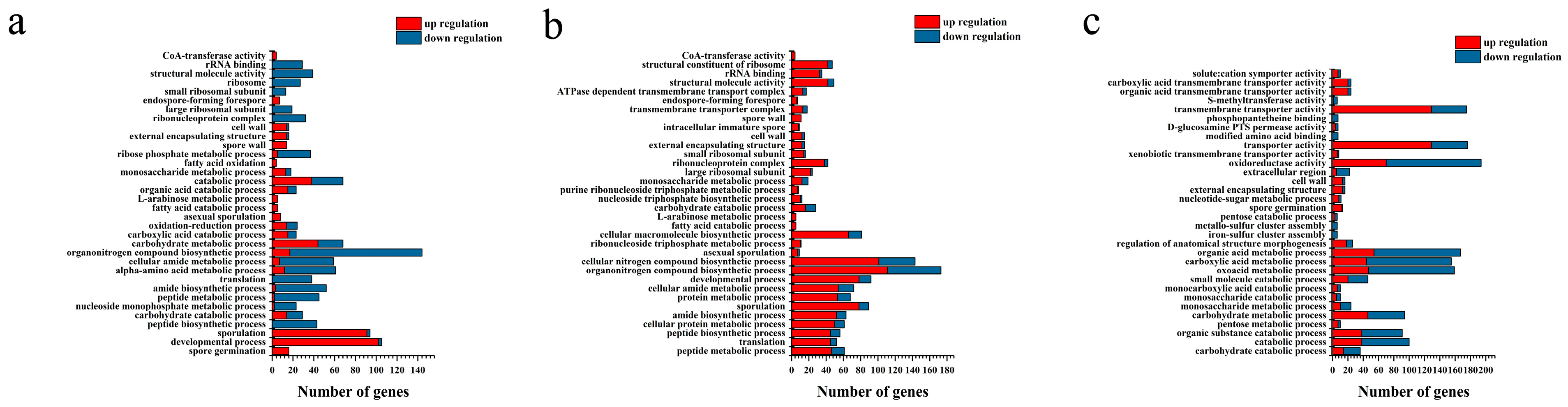
2.4. Transcriptome Analysis
2.5. Confirmation of Transcriptional Results with qRT-PCR
2.6. Analysis of Genes Associated with Sporulation in Strain NCD-2
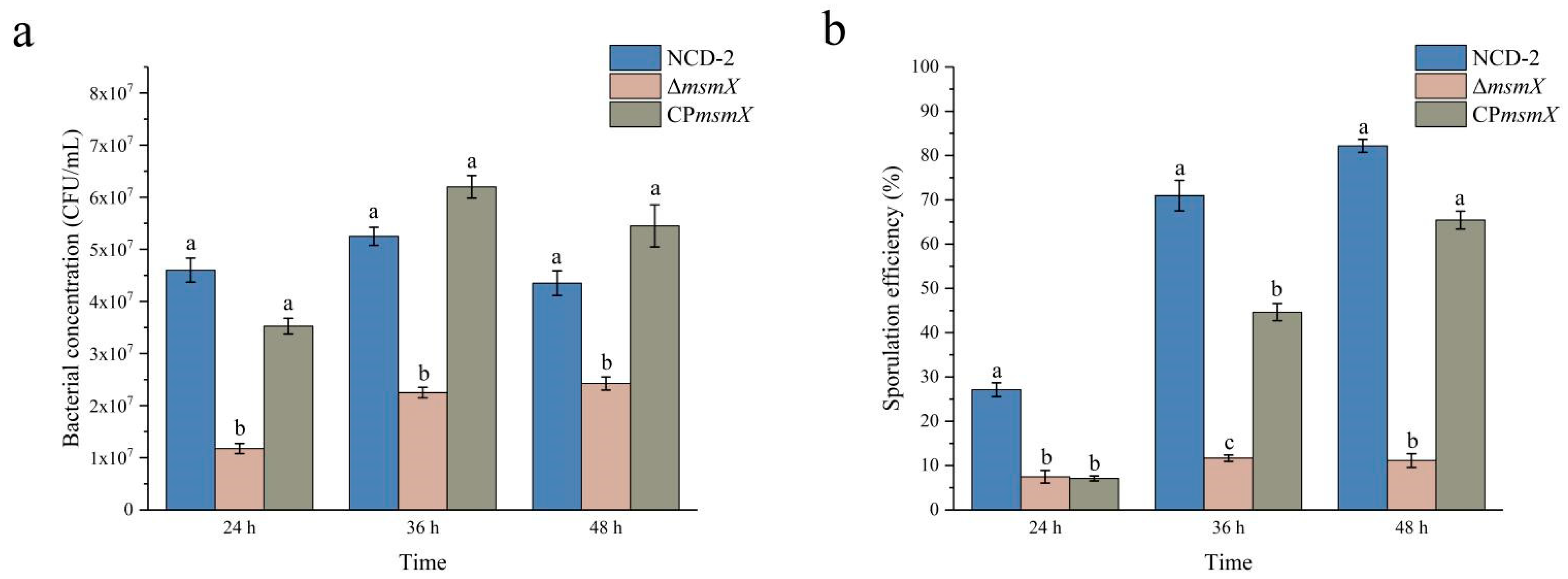
2.7. Deletion of the msmX Gene Decreased the Sporulation Efficiency in Strain NCD-2
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Growth Conditions
4.2. Biolog Phenotype MicroArray Analysis
4.3. Determination of Cell Concentration and Sporulation Efficiency
4.4. RNA Extraction and RNA Sequencing
4.5. Transcriptome Data and Differential Gene Expression Analysis
4.6. Confirmation of Transcriptome Analysis Results
4.7. Function Analysis of msmX Gene
4.8. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Fiers, M.; Edel-Hermann, V.; Chatot, C.; Hingrat, Y.L.; Steinberg, A.C. Potato soil-borne diseases. A review. Agro. Sustain. Dev. 2012, 32, 93–132. [Google Scholar] [CrossRef]
- Bisutti, I.L.; Pelz, J.; Büttner, C.; Stephan, D. Field assessment on the influence of RhizoVital® 42 fl. and Trichostar® on strawberries in the presence of soil-borne diseases. Crop Prot. 2017, 96, 195–203. [Google Scholar] [CrossRef]
- Pérez-García, A.; Romero, D.; de Vicente, A. Plant protection and growth stimulation by microorganisms: Biotechnological applications of Bacilli in agriculture. Curr. Opin. Biotechnol. 2011, 22, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Rao, Y.K.; Tsay, K.J.; Wu, W.S.; Tzeng, Y.M. Medium optimization of carbon and nitrogen sources for the production of spores from Bacillus amyloliquefaciens B128 using response surface methodology. Process Biochem. 2007, 42, 535–541. [Google Scholar] [CrossRef]
- Posada-Uribe, L.F.; Romero-Tabarez, M.; Villegas-Escobar, V. Effect of medium components and culture conditions in Bacillus subtilis EA-CB0575 spore production. Bioproc. Biosyst. Eng. 2015, 38, 1879–1888. [Google Scholar] [CrossRef] [PubMed]
- de Vries, Y.; Atmadja, R.; Hornstra, L.; de Vos, W.; Abee, T. Influence of glutamate on growth, sporulation, and spore properties of Bacillus cereus ATCC 14579 in defined medium. App. Environ. Microb. 2005, 71, 3248–3254. [Google Scholar] [CrossRef] [PubMed]
- Donohue, T.; Bernlohr, R. Effect of cultural conditions on the concentrations of metabolic intermediates during growth and sporulation of Bacillus licheniformis. J. Bacteriol. 1978, 135, 363–372. [Google Scholar] [CrossRef]
- Singh, R.M. Role of carbon and nitrogen sources in bacterial growth and sporulation. Appl. Microbiol. 1971, 22, 131–132. [Google Scholar] [CrossRef]
- Lopez, J.M.; Marks, C.L.; Freese, E. The decrease of guanine nucleotides initiates sporulation of Bacillus subtilis. Biochim. Biophys. Acta 1979, 587, 238–252. [Google Scholar] [CrossRef]
- Monteiro, S.M.S.; Clemente, J.J.; Carrondo, M.J.T.; Cunha, A.E. Enhanced spore production of Bacillus subtilis grown in a Chemically Defined Medium. Adv. Microbiol. 2014, 4, 444–454. [Google Scholar] [CrossRef]
- Warriner, K.; Waites, W.M. Enhanced sporulation in Bacillus subtilis grown on medium containing glucose:ribose. Lett. Appl. Microbiol. 1999, 29, 97–102. [Google Scholar] [CrossRef]
- Nguyen Thi Minh, H.; Durand, A.; Loison, P.; Perrier-Cornet, J.; Gervais, P. Effect of sporulation conditions on the resistance of Bacillus subtilis spores to heat and high pressure. Appl. Microbiol. Biotechnol. 2011, 90, 1409–1417. [Google Scholar] [CrossRef] [PubMed]
- Bressuire-Isoard, C.; Broussolle, V.; Carlin, F. Sporulation environment influences spore properties in Bacillus: Evidence and insights on underlying molecular and physiological mechanisms. FEMS Microbiol. Rev. 2018, 42, 614–626. [Google Scholar] [CrossRef] [PubMed]
- Widderich, N.; Rodrigues, C.D.; Commichau, F.M.; Fischer, K.E.; Ramirez-Guadiana, F.H.; Rudner, D.Z.; Bremer, E. Salt-sensitivity of σ(H) and Spo0A prevents sporulation of Bacillus subtilis at high osmolarity avoiding death during cellular differentiation. Mol. Microbiol. 2016, 100, 108–124. [Google Scholar] [CrossRef] [PubMed]
- Li, S.Z.; Lu, X.Y.; Ma, P.; Gao, S.G.; Liu, G. Evaluation of biocontrol potential of a bacterial strain NCD-2 against cotton verticillium wilt in field trials. Acta Phytopathol. Sin. 2005, 35, 451–455. [Google Scholar]
- Guo, Q.; Dong, W.; Li, S.; Lu, X.; Wang, P.; Zhang, X.; Wang, Y.; Ma, P. Fengycin produced by Bacillus subtilis NCD-2 plays a major role in biocontrol of cotton seedling damping-off disease. Microbio. Res. 2014, 169, 533–540. [Google Scholar] [CrossRef]
- Zhao, W.S.; Ban, Y.Y.; Su, Z.H.; Li, S.Z.; Liu, X.Y.; Guo, Q.G.; Ma, P. Colonization Ability of Bacillus subtilis NCD-2 in Different Crops and Its Effect on Rhizosphere Microorganisms. Microorganisms 2023, 11, 776. [Google Scholar] [CrossRef]
- Camp, A.H.; Losick, R. A novel pathway of intercellular signalling in Bacillus subtilis involves a protein with similarity to a component of type III secretion channels. Mol. Microbiol. 2008, 69, 402–417. [Google Scholar] [CrossRef]
- Yu, D.; Fang, Y.; Tang, C.; Klosterman, S.J.; Tian, C.; Wang, Y. Genomewide transcriptome profiles reveal how Bacillus subtilis lipopeptides inhibit microsclerotia formation in Verticillium dahliae. Mol. Plant. Microbe Interact. 2019, 32, 622–634. [Google Scholar] [CrossRef]
- Ayaz, M.; Li, C.H.; Ali, Q.; Zhao, W.; Chi, Y.K.; Shafiq, M.; Ali, F.; Yu, X.Y.; Yu, Q.; Zhao, J.T.; et al. Bacterial and fungal biocontrol agents for plant disease rotection: Journey from lab to field, current Status, challenges, and global perspectives. Molecules 2023, 28, 6735. [Google Scholar] [CrossRef] [PubMed]
- Dong, L.H.; Wang, P.; Zhao, W.S.; Su, Z.H.; Zhang, X.Y.; Lu, X.Y.; Guo, Q.G. Surfactin and fengycin contribute differentially to the biological activity of Bacillus subtilis NCD-2 against cotton verticillium wilt. Biol. Control 2022, 174, 104999. [Google Scholar] [CrossRef]
- Schaeffer, P.; Millet, J.; Aubert, J.P. Catabolic repression of bacterial sporulation. Proc. Natl. Acad. Sci. USA 1965, 54, 704–711. [Google Scholar] [CrossRef] [PubMed]
- Verma, N.; Singh, N.A.; Kumar, N.; Raghu, H.V. Screening of different media for sporulation of Bacillus megaterium. Int. J. Microbiol. Res. Rev 2013, 1, 68–73. [Google Scholar]
- Monteiro, S.M.; Clemente, J.J.; Henriques, A.O.; Gomes, R.J.; Carrondo, M.J.; Cunha, A.E. A procedure for high-yield spore production by Bacillus subtilis. Biotechnol. Prog. 2005, 21, 1026–1031. [Google Scholar] [CrossRef]
- Bochner, B.R.; Gadzinski, P.; Panomitros, E. Phenotype microarrays for high-throughput phenotypic testing and assay of gene function. Genome Res. 2001, 11, 1246–1255. [Google Scholar] [CrossRef]
- Bochner, B.R. New technologies to assess genotype-phenotype relationships. Nat. Rev. Genet. 2003, 4, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Mackie, A.; Hassan, K.; Paulsen, I.; Tetu, S. Biolog Phenotype Microarrays for phenotypic characterization of microbial cells. Methods Mol. Biol. 2014, 1096, 123–130. [Google Scholar]
- Sá-Nogueira, I.; Nogueira, T.V.; Soares, S.; de Lencastre, H. The Bacillus subtilis L-arabinose (ara) operon: Nucleotide sequence, genetic organization and expression. Microbiology 1997, 143 Pt 3, 957–969. [Google Scholar] [CrossRef]
- Lepesant, J.A.; Dedonder, R. Metabolism of L-arabinose in Bacillus subtilis Marburg Ind-168. C. R. Acad. Hebd. Seances Acad. Sci. D 1967, 264, 2683–2686. [Google Scholar]
- Sá-Nogueira, I.; de Lencastre, H. Cloning and characterization of araA, araB, and araD, the structural genes for L-arabinose utilization in Bacillus subtilis. J. Bacteriol. 1989, 171, 4088–4091. [Google Scholar] [CrossRef]
- Sá-Nogueira, I.; Mota, L.J. Negative regulation of L-arabinose metabolism in Bacillus subtilis: Characterization of the araR (araC) gene. J. Bacteriol. 1997, 179, 1598–1608. [Google Scholar] [CrossRef][Green Version]
- Sá-Nogueira, I.; Ramos, S.S. Cloning, functional analysis, and transcriptional regulation of the Bacillus subtilis araE gene involved in L-arabinose utilization. J. Bacteriol. 1997, 179, 7705–7711. [Google Scholar] [CrossRef]
- Ferreira, M.; Sá-Nogueira, I. A multitask ATPase serving different ABC-type sugar importers in Bacillus subtilis. J. Bacteriol. 2010, 192, 5312–5318. [Google Scholar] [CrossRef]
- Iyer, J.L.; Shetty, P.; Pai, J. Evaluation of whole cells of Bacillus subtilis as substrate for measurement of autolysin activity. Process Biochem. 2005, 40, 1593–1597. [Google Scholar] [CrossRef]
- Krogh, S.; Jørgensen, S.T.; Devine, K.M. Lysis genes of the Bacillus subtilis defective prophage PBSX. J. Bacteriol. 1998, 180, 2110–2117. [Google Scholar] [CrossRef]
- Nandy, S.K.; Venkatesh, K.V. Effect of carbon and nitrogen on the cannibalistic behavior of Bacillus subtilis. Appl. Biochem. Biotechnol. 2008, 151, 424–432. [Google Scholar] [CrossRef]
- Sahoo, S.; Rao, K.K.; Suraishkumar, G.K. Reactive oxygen species induced by shear stress mediate cell death in Bacillus subtilis. Biotechnol. Bioeng. 2006, 94, 118–127. [Google Scholar] [CrossRef] [PubMed]
- Jolliffe, L.K.; Langemeier, S.O.; Doyle, R.J. Hydrogen ion control of autolysin-dependent functions in Bacillus subtilis. Microbios 1983, 38, 187–194. [Google Scholar] [PubMed]
- González-Pastor, J.E.; Hobbs, E.C.; Losick, R. Cannibalism by sporulating bacteria. Science 2003, 301, 510–513. [Google Scholar] [CrossRef] [PubMed]
- González-Pastor, J.E. Cannibalism: A social behavior in sporulating Bacillus subtilis. FEMS Microbiol. Rev. 2011, 35, 415–424. [Google Scholar] [CrossRef] [PubMed]
- Eswaramoorthy, P.; Guo, T.; Fujita, M. In vivo domain-based functional analysis of the major sporulation sensor kinase, KinA, in Bacillus subtilis. J. Bacteriol. 2009, 191, 5358–5368. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Xia, L.Q.; Hu, S.B.; Yan, L.; Ding, X.Z.; Zhang, Y.M.; Yu, Z. Site-specific integration of heterologous gene into Bacillus thuringiensis chromosome and its expression. Acta Microbiol. Sin. 2008, 48, 661–666. [Google Scholar]
- Arnaud, M.; Chastanet, A.; Débarbouillé, M. New vector for efficient allelic replacement in naturally nontransformable, low-GC-content, gram-positive bacteria. Appl. Environ. Microbiol. 2004, 70, 6887–6891. [Google Scholar] [CrossRef]
- Guo, Q.; Li, S.; Lu, X.Y.; Li, B.Q.; Ma, P. PhoR/PhoP two component regulatory system affects biocontrol capability of Bacillus subtilis NCD-2. Genet. Mol. Biol. 2010, 33, 333–340. [Google Scholar] [CrossRef]


| Accession ID | Gene Name | Log2 (Ara/Glc) | Production | ||
|---|---|---|---|---|---|
| 8 h | 12 h | 16 h | |||
| WP_003231833.1 | cotE | 7.65 | 5.81 | 2.56 | Outer spore coat protein CotE |
| WP_003243364.1 | cotF | 6.59 | 9.47 | 5.54 | Spore coat protein CotF |
| WP_080344234.1 | cotG | 3.55 | 5.59 | 3.82 | Spore coat protein CotG |
| ADV92699.1 | cotM | 5.01 | 1.53 | 4.21 | Spore coat protein (outer) |
| WP_047183078.1 | cotS | 3.98 | 6.01 | 4.53 | Spore coat protein CotS |
| PSM02245.1 | cotT | 5.88 | 5.95 | 8.39 | Spore coat protein |
| AGE63031.1 | cotV | 4.37 | 6.32 | 5.46 | Spore coat protein (insoluble fraction) |
| WP_069486390.1 | cotW | 4.53 | 6.81 | 5.73 | Spore coat protein |
| WP_014476454.1 | cotX | 4.28 | 6.83 | 6.13 | Spore coat protein |
| WP_003231888.1 | dpaA | 6.29 | 6.84 | 4.81 | Dipicolinic acid synthetase subunit A |
| WP_003231884.1 | dpaB | 5.76 | 6.12 | 4.21 | Dipicolinate synthase subunit B |
| WP_015383228.1 | yheD | 5.17 | 3.35 | 2.27 | Spore coat associated protein YheD |
| WP_063336053.1 | gerBA | 4.55 | 2.20 | 4.73 | Spore germination protein GerKA |
| WP_003184172.1 | gerE | 3.94 | 5.01 | 3.91 | Spore germination protein GerE |
| WP_014478336.1 | gerQ | 7.95 | 4.77 | 1.32 | Spore coat protein GerQ |
| WP_047182746.1 | gerT | 5.10 | 6.02 | 6.03 | Spore germination protein GerT |
| AKE24397.1 | sigK | 6.45 | 4.08 | 3.26 | RNA polymerase sporulation-specific sigma factor |
| WP_047182864.1 | spoIIIAE | 3.84 | 2.72 | 1.41 | Stage III sporulation protein AE |
| WP_003221804.1 | spoIIID | 10.25 | 6.81 | 4.40 | Sporulation transcriptional regulator SpoIIID |
| WP_004398593.1 | spoIIM | 1.78 | 1.84 | 2.35 | Stage II sporulation protein M |
| WP_047183325.1 | spoIIQ | 5.69 | 1.78 | −1.82 | Stage II sporulation protein SpoIIQ |
| WP_004398697.1 | spoIVB | 6.19 | 1.77 | 2.12 | SpoIVB peptidase |
| WP_015483522.1 | spoIVFB | 1.25 | 1.39 | 1.84 | Stage IV sporulation protein SpoIVFB |
| WP_003230465.1 | spoVAD | 6.29 | 1.70 | 4.08 | Stage V sporulation protein AD |
| AGE63365.1 | spoVD | 2.67 | 3.20 | 2.32 | Penicillin-binding protein |
| WP_047182441.1 | yjcA | 6.69 | 5.32 | 5.67 | Sporulation protein YjcA |
| WP_015383520.1 | ykvU | 3.60 | 3.01 | 4.18 | Sporulation protein YkvU |
| WP_003223491.1 | sspA | 7.22 | 3.14 | 3.04 | Alpha/beta-type small acid-soluble spore protein |
| WP_003233287.1 | sspB | 7.39 | 3.43 | 2.66 | Alpha/beta-type small acid-soluble spore protein |
| WP_003218568.1 | sspD | 7.57 | 3.93 | 4.00 | Alpha/beta-type small acid-soluble spore protein |
| BAI84385.2 | sspE | 6.59 | 4.34 | 3.30 | Gamma-type small acid-soluble spore protein |
| WP_003244950.1 | sdpC | −1.40 | −5.57 | −6.81 | Sporulation-delaying protein family toxin |
| WP_003228357.1 | sdpI | −4.53 | −3.43 | −4.90 | Immunity protein SdpI |
| WP_003243541.1 | sdpR | −4.51 | −3.34 | −2.64 | Sporulation-delaying system autorepressor SdpR |
| Strain | Genotype | Source |
|---|---|---|
| WT | Bacillus subtilis NCD-2 wild type | Lab stock |
| ΔmsmX | NCD-2 mutant, msmX-deletion mutant | This study |
| CPmsmX | Complementary of ΔmsmX with intact msmX, chloramphenicol-resistant | This study |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fu, Y.; Liu, X.; Su, Z.; Wang, P.; Guo, Q.; Ma, P. Arabinose Plays an Important Role in Regulating the Growth and Sporulation of Bacillus subtilis NCD-2. Int. J. Mol. Sci. 2023, 24, 17472. https://doi.org/10.3390/ijms242417472
Fu Y, Liu X, Su Z, Wang P, Guo Q, Ma P. Arabinose Plays an Important Role in Regulating the Growth and Sporulation of Bacillus subtilis NCD-2. International Journal of Molecular Sciences. 2023; 24(24):17472. https://doi.org/10.3390/ijms242417472
Chicago/Turabian StyleFu, Yifan, Xiaomeng Liu, Zhenhe Su, Peipei Wang, Qinggang Guo, and Ping Ma. 2023. "Arabinose Plays an Important Role in Regulating the Growth and Sporulation of Bacillus subtilis NCD-2" International Journal of Molecular Sciences 24, no. 24: 17472. https://doi.org/10.3390/ijms242417472
APA StyleFu, Y., Liu, X., Su, Z., Wang, P., Guo, Q., & Ma, P. (2023). Arabinose Plays an Important Role in Regulating the Growth and Sporulation of Bacillus subtilis NCD-2. International Journal of Molecular Sciences, 24(24), 17472. https://doi.org/10.3390/ijms242417472







