Profiling Microbial Communities in Idiopathic Granulomatous Mastitis
Abstract
1. Introduction
- Comparisons between IGM and LM pus microbial communities;
- Comparisons between IGM pus and skin microbial communities in the same patient.
2. Results
2.1. Study Population
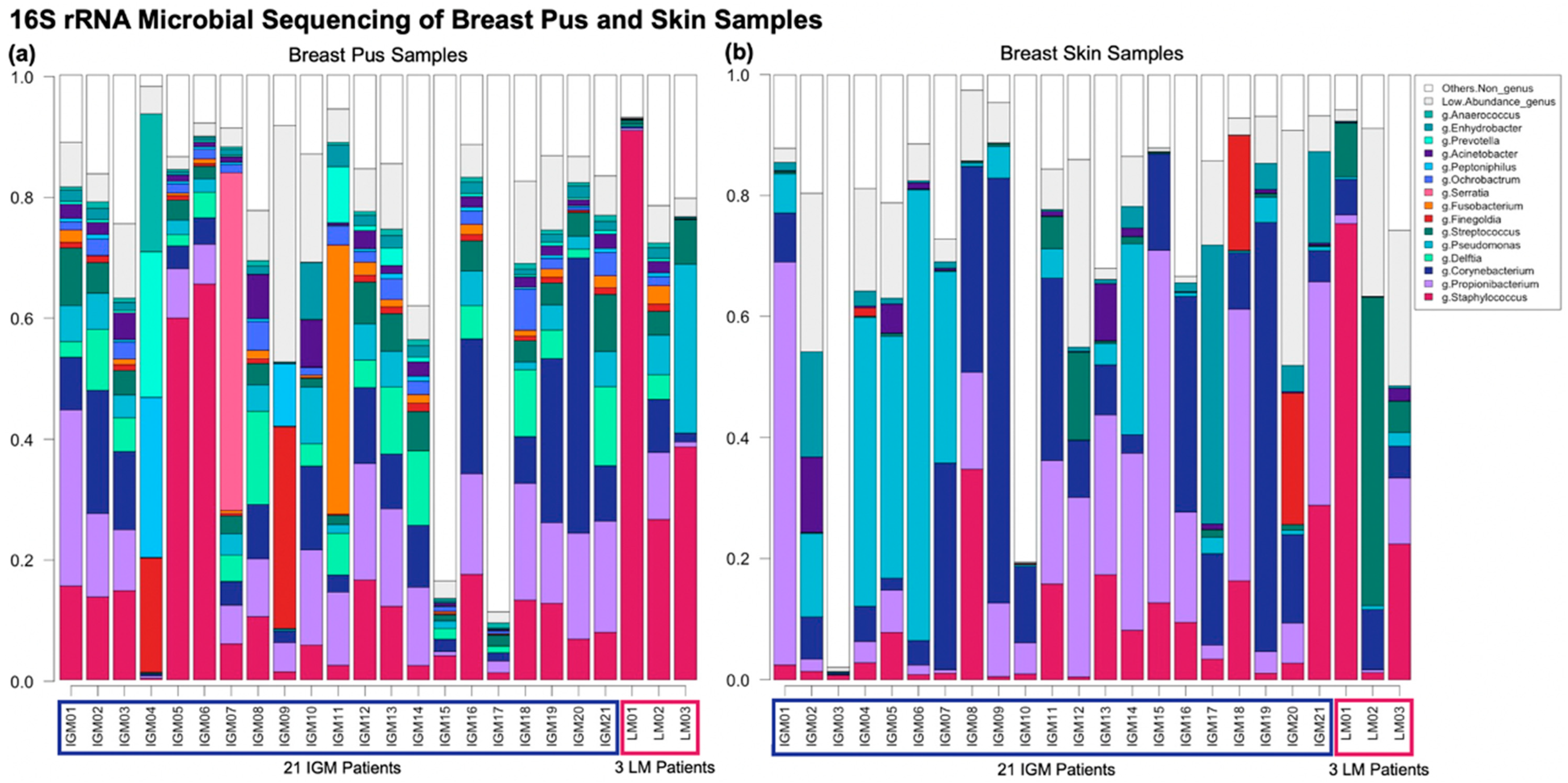
2.2. Comparing IGM vs. LM Microbial Taxonomic Profiles
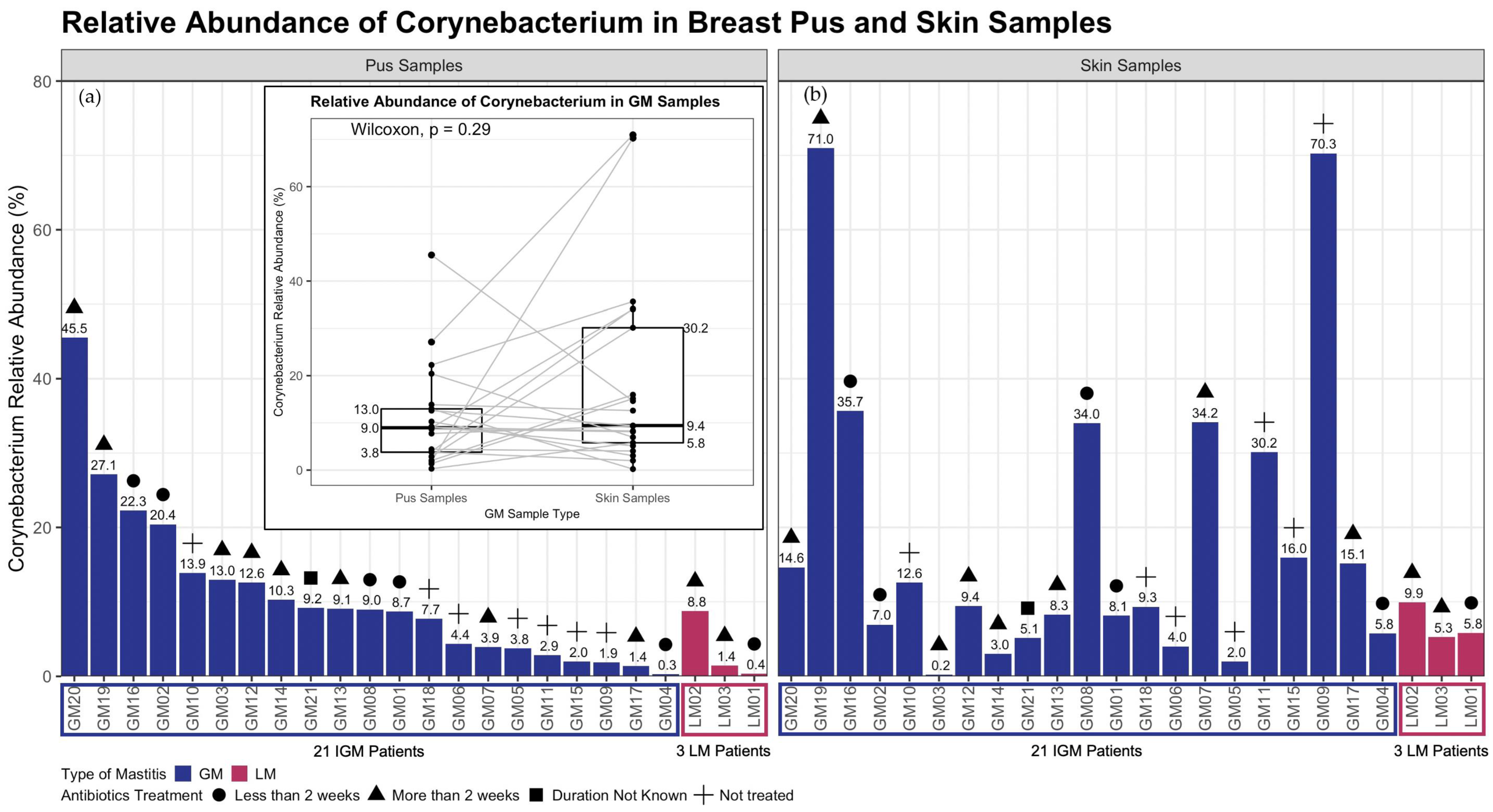
2.3. Corynebacterium
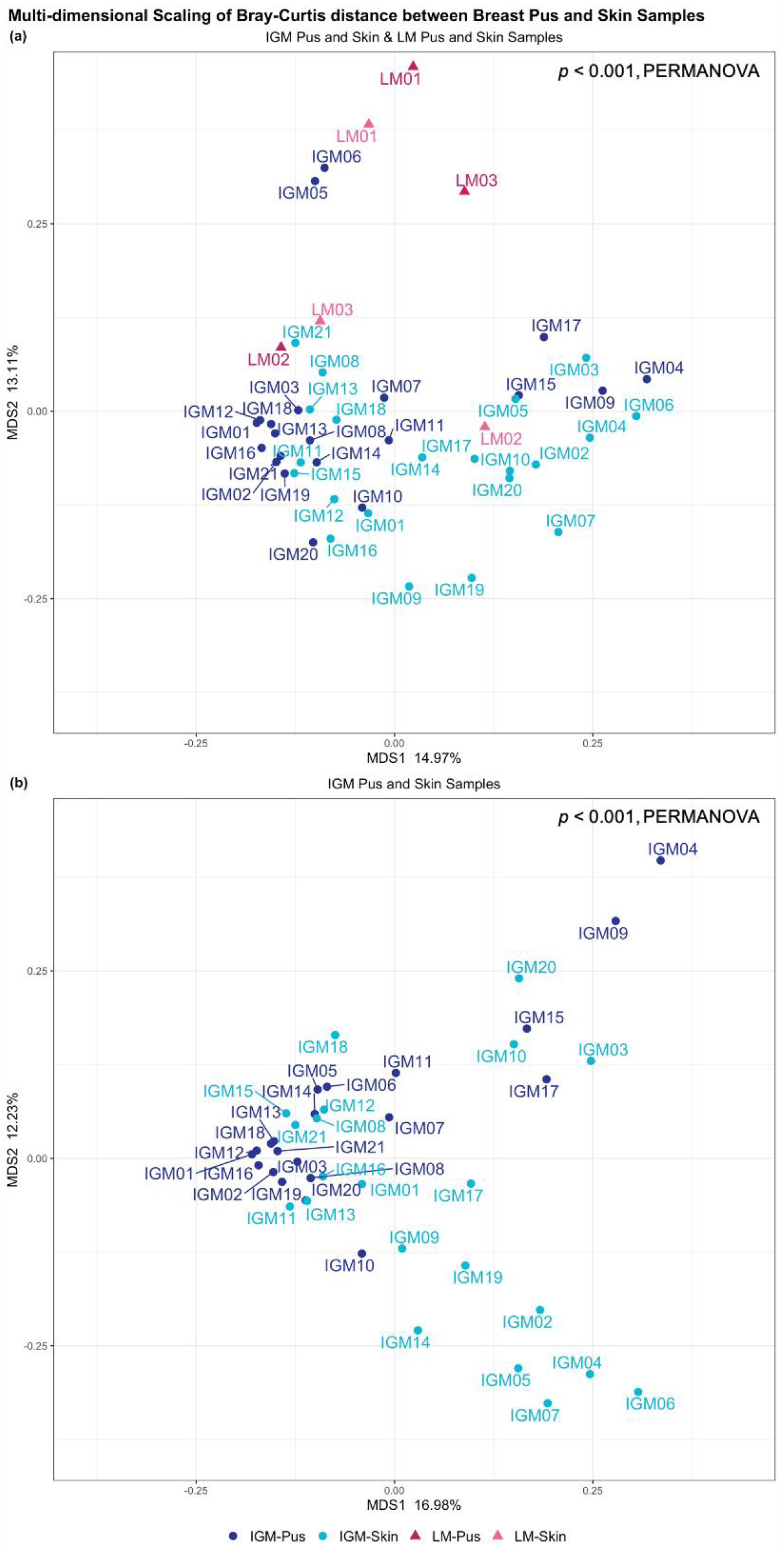
2.4. Associations between Sample Types and Metagenomic Features
3. Discussion
4. Materials and Methods
4.1. Study Population
4.2. Patient Recruitment and Sample Collection
- Pus sample collection: Swab was allowed to absorb infected fluid for 30 s while rotating the swab;
- Skin sample collection: Swab was used to rub skin area for 10 to 15 strokes with moderate pressure. Swab was rotated and sampling was repeated.
4.3. DNA Extraction and 16S Ribosomal rRNA Sequencing
- Denaturation at 95 °C for 30 seconds;
- Annealing at 59 °C for 30 seconds; and
- Extension at 72 °C for 1 minute.
4.4. Sequencing Data Processing
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bellavia, M.; Damiano, G.; Palumbo, V.D.; Spinelli, G.; Tomasello, G.; Marrazzo, A.; Ficarella, S.; Bruno, A.; Sammartano, A.; Fiorentini, T.; et al. Granulomatous mastitis during chronic antidepressant therapy: Is it possible a conservative therapeutic approach? J. Breast Cancer 2012, 15, 371–372. [Google Scholar] [CrossRef] [PubMed]
- Kessler, E.; Wolloch, Y. Granulomatous mastitis: A lesion clinically simulating carcinoma. Am. J. Clin. Pathol. 1972, 58, 642–646. [Google Scholar] [CrossRef] [PubMed]
- Tekgoz, E.; Colak, S.; Cinar, M.; Yilmaz, S. Treatment of idiopathic granulomatous mastitis and factors related with disease recurrence. Turk. J. Med. Sci. 2020, 50, 1380–1386. [Google Scholar] [CrossRef]
- Steuer, A.B.; Stern, M.J.; Cobos, G.; Castilla, C.; Joseph, K.A.; Pomeranz, M.K.; Femia, A.N. Clinical characteristics and medical management of idiopathic granulomatous mastitis. JAMA Dermatol. 2020, 156, 460–464. [Google Scholar] [CrossRef] [PubMed]
- Mathew Thomas, V.; Alexander, S.A.; Bindal, P.; Vredenburgh, J. Idiopathic granulomatous mastitis—A mystery yet to be unraveled: A case series and review of literature. Cureus 2020, 12, e6895. [Google Scholar] [CrossRef]
- Pluguez-Turull, C.W.; Nanyes, J.E.; Quintero, C.J.; Alizai, H.; Mais, D.D.; Kist, K.A.; Dornbluth, N.C. Idiopathic Granulomatous Mastitis: Manifestations at Multimodality Imaging and Pitfalls. Radiographics 2018, 38, 330–356. [Google Scholar] [CrossRef]
- Altintoprak, F.; Kivilcim, T.; Ozkan, O.V. Aetiology of idiopathic granulomatous mastitis. World J. Clin. Cases. 2014, 2, 852–858. [Google Scholar] [CrossRef]
- Taylor, G.B.; Paviour, S.D.; Musaad, S.; Jones, W.O.; Holland, D.J. A clinicopathological review of 34 cases of inflammatory breast disease showing an association between corynebacteria infection and granulomatous mastitis. Pathology 2003, 35, 109–119. [Google Scholar]
- Paviour, S.; Musaad, S.; Roberts, S.; Taylor, G.; Taylor, S.; Shore, K.; Lang, S.; Holland, D. Corynebacterium species isolated from patients with mastitis. Clin. Infect. Dis. 2002, 35, 1434–1440. [Google Scholar] [CrossRef]
- Ang, L.M.; Brown, H. Corynebacterium accolens isolated from breast abscess: Possible association with granulomatous mastitis. J. Clin. Microbiol. 2007, 45, 1666–1668. [Google Scholar] [CrossRef]
- Yu, H.J.; Deng, H.; Ma, J.; Huang, S.J.; Yang, J.M.; Huang, Y.F.; Mu, X.P.; Zhang, L.; Wang, Q. Clinical metagenomic analysis of bacterial communities in breast abscesses of granulomatous mastitis. Int. J. Infect. Dis. 2016, 53, 30–33. [Google Scholar] [CrossRef] [PubMed]
- Omranipour, R.; Vasigh, M. Mastitis, breast abscess, and granulomatous mastitis. In Diseases of the Breast during Pregnancy and Lactation; Alipour, S., Omranipour, R., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 53–61. [Google Scholar] [CrossRef]
- Kvist, L.J. Toward a clarification of the concept of mastitis as used in empirical studies of breast inflammation during lactation. J. Hum. Lact. 2010, 26, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Scott, D.M. Inflammatory diseases of the breast. Best Pract. Res. Clin. Obstet. Gynaecol. 2022, 83, 72–87. [Google Scholar] [CrossRef] [PubMed]
- Amir, L.H.; Academy of Breastfeeding Medicine Protocol Committee. ABM clinical protocol #4: Mastitis, revised March 2014. Breastfeed. Med. 2014, 9, 239–243. [Google Scholar] [CrossRef]
- Angelopoulou, A.; Field, D.; Ryan, C.A.; Stanton, C.; Hill, C.; Ross, R.P. The microbiology and treatment of human mastitis. Med. Microbiol. Immunol. 2018, 207, 83–94. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, E.; de Andres, J.; Manrique, M.; Pareja-Tobes, P.; Tobes, R.; Martinez-Blanch, J.F.; Codoner, F.M.; Ramon, D.; Fernandez, L.; Rodriguez, J.M. Metagenomic Analysis of Milk of Healthy and Mastitis-Suffering Women. J. Hum. Lact. 2015, 31, 406–415. [Google Scholar] [CrossRef]
- Patel, S.H.; Vaidya, Y.H.; Patel, R.J.; Pandit, R.J.; Joshi, C.G.; Kunjadiya, A.P. Culture independent assessment of human milk microbial community in lactational mastitis. Sci. Rep. 2017, 7, 7804. [Google Scholar] [CrossRef]
- Marin, M.; Arroyo, R.; Espinosa-Martos, I.; Fernandez, L.; Rodriguez, J.M. Identification of emerging human mastitis pathogens by MALDI-TOF and assessment of their antibiotic resistance patterns. Front. Microbiol. 2017, 8, 1258. [Google Scholar] [CrossRef]
- Mediano, P.; Fernandez, L.; Jimenez, E.; Arroyo, R.; Espinosa-Martos, I.; Rodriguez, J.M.; Marin, M. microbial diversity in milk of women with mastitis: Potential role of coagulase-negative staphylococci, viridans group streptococci, and corynebacteria. J. Hum. Lact. 2017, 33, 309–318. [Google Scholar] [CrossRef]
- Shojaee, L.; Rahmani, N.; Moradi, S.; Motamedi, A.; Godazandeh, G. Idiopathic granulomatous mastitis: Challenges of treatment in iranian women. BMC Surg. 2021, 21, 206. [Google Scholar] [CrossRef]
- Uysal, E.; Soran, A.; Sezgin, E.; Granulomatous Mastitis Study Group. Factors related to recurrence of idiopathic granulomatous mastitis: What do we learn from a multicentre study? ANZ J. Surg. 2018, 88, 635–639. [Google Scholar] [CrossRef] [PubMed]
- Azizi, A.; Prasath, V.; Canner, J.; Gharib, M.; Sadat Fattahi, A.; Naser Forghani, M.; Sajjadi, S.; Farhadi, E.; Vasigh, M.; Kaviani, A.; et al. Idiopathic granulomatous mastitis: Management and predictors of recurrence in 474 patients. Breast J. 2020, 26, 1358–1362. [Google Scholar] [CrossRef] [PubMed]
- Basim, P.; Argun, D.; Argun, F. Risk factors for idiopathic granulomatous mastitis recurrence after patient-tailored treatment: Do we need an escalating treatment algorithm? Breast Care 2022, 17, 172–179. [Google Scholar] [CrossRef]
- Bukin, Y.S.; Galachyants, Y.P.; Morozov, I.V.; Bukin, S.V.; Zakharenko, A.S.; Zemskaya, T.I. The effect of 16S rRNA region choice on bacterial community metabarcoding results. Sci. Data 2019, 6, 190007. [Google Scholar] [CrossRef] [PubMed]
- Meisel, J.S.; Hannigan, G.D.; Tyldsley, A.S.; SanMiguel, A.J.; Hodkinson, B.P.; Zheng, Q.; Grice, E.A. Skin microbiome surveys are strongly influenced by experimental design. J. Investig. Dermatol. 2016, 136, 947–956. [Google Scholar] [CrossRef] [PubMed]
- Chakravorty, S.; Helb, D.; Burday, M.; Connell, N.; Alland, D. A detailed analysis of 16S ribosomal RNA gene segments for the diagnosis of pathogenic bacteria. J. Microbiol. Methods 2007, 69, 330–339. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Porchas, M.; Villalpando-Canchola, E.; Vargas-Albores, F. Significant loss of sensitivity and specificity in the taxonomic classification occurs when short 16S rRNA gene sequences are used. Heliyon 2016, 2, e00170. [Google Scholar] [CrossRef]
- Blackmon, M.M.; Nguyen, H.; Mukherji, P. Acute Mastitis. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Fetherston, C. Risk factors for lactation mastitis. J. Hum. Lact. 1998, 14, 101–109. [Google Scholar] [CrossRef]
- Foxman, B.; D’Arcy, H.; Gillespie, B.; Bobo, J.K.; Schwartz, K. Lactation mastitis: Occurrence and medical management among 946 breastfeeding women in the United States. Am. J. Epidemiol. 2002, 155, 103–114. [Google Scholar] [CrossRef]
- Bouton, M.E.; Winton, L.M.; Gandhi, S.G.; Jayaram, L.; Patel, P.N.; PJ, O.N.; Komenaka, I.K. Temporal resolution of idiopathic granulomatous mastitis with resumption of bromocriptine therapy for prolactinoma. Int. J. Surg. Case Rep. 2015, 10, 8–11. [Google Scholar] [CrossRef][Green Version]
- Funke, G.; von Graevenitz, A.; Clarridge, J.E., 3rd; Bernard, K.A. Clinical microbiology of coryneform bacteria. Clin. Microbiol. Rev. 1997, 10, 125–159. [Google Scholar] [CrossRef] [PubMed]
- Dobinson, H.C.; Anderson, T.P.; Chambers, S.T.; Doogue, M.P.; Seaward, L.; Werno, A.M. Antimicrobial Treatment Options for Granulomatous Mastitis Caused by Corynebacterium Species. J. Clin. Microbiol. 2015, 53, 2895–2899. [Google Scholar] [CrossRef] [PubMed]
- Co, M.; Cheng, V.C.C.; Wei, J.; Wong, S.C.Y.; Chan, S.M.S.; Shek, T.; Kwong, A. Idiopathic granulomatous mastitis: A 10-year study from a multicentre clinical database. Pathology 2018, 50, 742–747. [Google Scholar] [CrossRef] [PubMed]
- Wolfrum, A.; Kummel, S.; Theuerkauf, I.; Pelz, E.; Reinisch, M. Granulomatous Mastitis: A Therapeutic and Diagnostic Challenge. Breast Care 2018, 13, 413–418. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.C.Y.; Poon, R.W.S.; Chen, J.H.K.; Tse, H.; Lo, J.Y.C.; Ng, T.K.; Au, J.C.K.; Tse, C.W.S.; Cheung, I.Y.Y.; Yuk, M.T.; et al. Corynebacterium kroppenstedtii Is an Emerging Cause of Mastitis Especially in Patients With Psychiatric Illness on Antipsychotic Medication. Open Forum Infect. Dis. 2017, 4, ofx096. [Google Scholar] [CrossRef]
- Chalmers, R.; McClellan, P.; Silva, V.; Shutt, N.; Restini, C. Red flags for the differential diagnosis of granulomatous mastitis: A case report. J. Med. Case Rep. 2020, 14, 215. [Google Scholar] [CrossRef]
- Hartmann, J.E.; Albrich, W.C.; Dmitrijeva, M.; Kahlert, C.R. The effects of corticosteroids on the respiratory microbiome: A systematic review. Front. Med. 2021, 8, 588584. [Google Scholar] [CrossRef]
- Huang, E.Y.; Inoue, T.; Leone, V.A.; Dalal, S.; Touw, K.; Wang, Y.; Musch, M.W.; Theriault, B.; Higuchi, K.; Donovan, S.; et al. Using corticosteroids to reshape the gut microbiome: Implications for inflammatory bowel diseases. Inflamm. Bowel Dis. 2015, 21, 963–972. [Google Scholar] [CrossRef]
- Vich Vila, A.; Collij, V.; Sanna, S.; Sinha, T.; Imhann, F.; Bourgonje, A.R.; Mujagic, Z.; Jonkers, D.; Masclee, A.A.M.; Fu, J.; et al. Impact of commonly used drugs on the composition and metabolic function of the gut microbiota. Nat. Commun. 2020, 11, 362. [Google Scholar] [CrossRef]
- Graham, M.E.; Herbert, W.G.; Song, S.D.; Raman, H.N.; Zhu, J.E.; Gonzalez, P.E.; Walther-Antonio, M.R.S.; Tetel, M.J. Gut and vaginal microbiomes on steroids: Implications for women’s health. Trends Endocrinol. Metab. 2021, 32, 554–565. [Google Scholar] [CrossRef]
- Laforest, S.; Pelletier, M.; Denver, N.; Poirier, B.; Nguyen, S.; Walker, B.R.; Durocher, F.; Homer, N.Z.M.; Diorio, C.; Andrew, R.; et al. Estrogens and glucocorticoids in mammary adipose tissue: Relationships with body mass index and breast cancer features. J. Clin. Endocrinol. Metab. 2020, 105, e1504–e1516. [Google Scholar] [CrossRef]
- Wilson, M.; Mello, M.J.; Gruppuso, P.A. Antibiotics and the human microbiome: A survey of prescribing clinicians’ knowledge and opinions regarding the link between antibiotic-induced dysbiosis and immune-mediated disease. R. I. Med. J. 2021, 104, 59–63. [Google Scholar] [PubMed]
- Ramirez, J.; Guarner, F.; Bustos Fernandez, L.; Maruy, A.; Sdepanian, V.L.; Cohen, H. Antibiotics as major disruptors of gut microbiota. Front. Cell Infect. Microbiol. 2020, 10, 572912. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Osto, M.; Geurts, L.; Everard, A. Involvement of gut microbiota in the development of low-grade inflammation and type 2 diabetes associated with obesity. Gut Microbes 2012, 3, 279–288. [Google Scholar] [CrossRef] [PubMed]
- Nel Van Zyl, K.; Matukane, S.R.; Hamman, B.L.; Whitelaw, A.C.; Newton-Foot, M. Effect of antibiotics on the human microbiome: A systematic review. Int. J. Antimicrob. Agents 2022, 59, 106502. [Google Scholar] [CrossRef]
- Vangoitsenhoven, R.; Cresci, G.A.M. Role of microbiome and antibiotics in autoimmune diseases. Nutr. Clin. Pract. 2020, 35, 406–416. [Google Scholar] [CrossRef]
- Wisplinghoff, H. 181—Pseudomonas spp., Acinetobacter spp. and miscellaneous gram-negative bacilli. In Infectious Diseases, 4th ed.; Cohen, J., Powderly, W.G., Opal, S.M., Eds.; Elsevier: Amsterdam, The Netherlands, 2017; pp. 1579–1599. [Google Scholar] [CrossRef]
- Chiller, K.; Selkin, B.A.; Murakawa, G.J. Skin microflora and bacterial infections of the skin. J. Investig. Dermatol. Symp. Proc. 2001, 6, 170–174. [Google Scholar] [CrossRef]
- Davis, C.P. Normal flora. In Medical Microbiology; Baron, S., Ed.; University of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996. [Google Scholar]
- Ryan, M.P.; Pembroke, J.T. The genus ochrobactrum as major opportunistic pathogens. Microorganisms 2020, 8, 1797. [Google Scholar] [CrossRef]
- Herman, T.F.; Hashmi, M.F. Cephalexin. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Smieja, M. Current indications for the use of clindamycin: A critical review. Can. J. Infect. Dis. 1998, 9, 22–28. [Google Scholar] [CrossRef]
- Kester, M.; Karpa, K.D.; Vrana, K.E. 4—Treatment of infectious diseases. In Elsevier’s Integrated Review Pharmacology, 2nd ed.; Kester, M., Karpa, K.D., Vrana, K.E., Eds.; W.B. Saunders: Philadelphia, PA, USA, 2012; pp. 41–78. [Google Scholar] [CrossRef]
- Waller, D.G.; Sampson, A.P. 51—Chemotherapy of infections. In Medical Pharmacology and Therapeutics (Fifth Edition), Waller, D.G., Sampson, A.P., Eds.; Elsevier: Amsterdam, The Netherlands, 2018; pp. 581–629. [Google Scholar] [CrossRef]
- GlaxoSmithKline Inc. Product monograph, including Patient Medication Information. In Clavulin. Amoxicillin/Clavulanate Potassium Tablets. Amoxicillin/Clavulanate Potassium for Oral Suspension; GlaxoSmithKline Inc.: Mississauga, ON, Canada, 2020. [Google Scholar]
- Bjerre, R.D.; Hugerth, L.W.; Boulund, F.; Seifert, M.; Johansen, J.D.; Engstrand, L. Effects of sampling strategy and DNA extraction on human skin microbiome investigations. Sci. Rep. 2019, 9, 17287. [Google Scholar] [CrossRef]
- Johnson, J.S.; Spakowicz, D.J.; Hong, B.Y.; Petersen, L.M.; Demkowicz, P.; Chen, L.; Leopold, S.R.; Hanson, B.M.; Agresta, H.O.; Gerstein, M.; et al. Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis. Nat. Commun. 2019, 10, 5029. [Google Scholar] [CrossRef] [PubMed]
- Quince, C.; Walker, A.W.; Simpson, J.T.; Loman, N.J.; Segata, N. Shotgun metagenomics, from sampling to analysis. Nat. Biotechnol. 2017, 35, 833–844. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.H.; Kukkillaya, V.U.; Wilm, A.; Lay, C.; Ho, E.X.; Low, L.; Hibberd, M.L.; Nagarajan, N. Species identification and profiling of complex microbial communities using shotgun Illumina sequencing of 16S rRNA amplicon sequences. PLoS ONE 2013, 8, e60811. [Google Scholar] [CrossRef] [PubMed]
- Miller, C.S.; Baker, B.J.; Thomas, B.C.; Singer, S.W.; Banfield, J.F. EMIRGE: Reconstruction of full-length ribosomal genes from microbial community short read sequencing data. Genome Biol. 2011, 12, R44. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Mallick, H.; Rahnavard, A.; McIver, L.J.; Ma, S.; Zhang, Y.; Nguyen, L.H.; Tickle, T.L.; Weingart, G.; Ren, B.; Schwager, E.H.; et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 2021, 17, e1009442. [Google Scholar] [CrossRef] [PubMed]

| Total n = 24 | IGM n = 21 | LM n = 3 | p-Value 1 | |
|---|---|---|---|---|
| Demographics | ||||
| Median age at diagnosis (years, IQR) | 34 (30.50–40.25) | 34 (31.00–41.00) | 31 (28.00–34.00) | 0.358 |
| Recruitment site (n, %) | 0.422 | |||
| KKH | 13 (54.17) | 10 (47.62) | 3 (100.00) | |
| SGH | 9 (37.50) | 9 (42.86) | 0 (0) | |
| NUH | 2 (8.33) | 2 (9.52) | 0 (0) | |
| Year of diagnosis (n, %) | 0.546 | |||
| 2014–2017 | 6 (25.00) | 6 (28.57) | 0 (0) | |
| 2018–2019 | 18 (75.00) | 15 (71.43) | 3 (100.00) | |
| Ethnicity (n, %) | 0.308 | |||
| Chinese | 15 (62.50) | 14 (66.67) | 1 (33.33) | |
| Malay | 4 (16.67) | 3 (14.29) | 1 (33.33) | |
| Indian | 2 (8.33) | 2 (9.52) | 0 (0) | |
| Other | 3 (12.50) | 2 (9.52) | 1 (33.33) | |
| Body mass index (kg/m2, IQR) | 26.29 (22.34–30.83) | 26.29 (24.61–31.22) | 22.19 (22.03–23.91) | 0.106 |
| Education level (n, %) | 0.727 | |||
| Primary | 2 (8.33) | 2 (9.52) | 0 (0) | |
| Secondary | 5 (20.83) | 4 (19.05) | 1 (33.33) | |
| Pre-University | 8 (33.33) | 6 (28.57) | 2 (66.67) | |
| Undergraduate | 6 (25.00) | 6 (28.57) | 0 (0) | |
| Graduate | 3 (12.50) | 3 (14.29) | 0 (0) | |
| Patient characteristics | ||||
| Pregnant, at time of sample collection (n, %) | ||||
| No | 24 (100.00) | 21 (100.00) | 3 (100.00) | |
| Lactating, at time of sample collection (n, %) | <0.001 | |||
| Yes | 3 (12.50) | 0 (0) | 3 (100.00) | |
| No | 21 (87.50) | 21 (100.00) | 0 (0) | |
| Previously Pregnant (n, %) | 1 | |||
| Yes | 21 (87.50) | 18 (85.71) | 3 (100.00) | |
| No | 3 (12.50) | 3 (14.29) | 0 (0) | |
| Number of children (n, %) | 0.769 | |||
| 0 | 3 (12.50) | 3 (14.29) | 0 (0) | |
| 1 | 13 (54.17) | 10 (47.62) | 3 (100.00) | |
| 2 | 6 (25.00) | 6 (28.57) | 0 (0) | |
| 3 | 1 (4.17) | 1 (4.76) | 0 (0) | |
| 4 | 1 (4.17) | 1 (4.76) | 0 (0) | |
| Time since last childbirth (n, %) | 0.091 | |||
| No children | 3 (12.50) | 3 (14.29) | 0 (0) | |
| Less than 2 years | 8 (33.33) | 5 (23.81) | 3 (100.00) | |
| Between 3 and 5 years | 10 (41.67) | 10 (47.62) | 0 (0) | |
| More than 5 years | 3 (12.50) | 3 (14.29) | 0 (0) | |
| Smoking (n, %) | 1 | |||
| Yes | 5 (20.83) | 5 (23.81) | 0 (0) | |
| No | 19 (79.17) | 16 (76.19) | 3 (100.00) | |
| Chronic Illness 2 Diagnosis (n, %) | 1 | |||
| Yes | 4 (16.67) | 4 (19.05) | 0 (0) | |
| No | 20 (83.33) | 17 (80.95) | 3 (100.00) | |
| Autoimmune Conditions 3 (n, %) | ||||
| No | 24 (100.00) | 21 (100.00) | 3 (100.00) | |
| Previous Infectious Disease 4 Diagnosis (n, %) | ||||
| No | 24 (100.00) | 21 (100.00) | 3 (100.00) | |
| Previous Cancer Diagnosis (n, %) | ||||
| No | 24 (100.00) | 21 (100.00) | 3 (100.00) | |
| Treatment for mastitis | ||||
| Any treatment (n, %) | 1 | |||
| Yes | 21 (87.50) | 18 (85.71) | 3 (100.00) | |
| No | 3 (12.50) | 3 (14.29) | 0 (0) | |
| Type of treatment (n, %) | 0.185 | |||
| Antibiotic treatment 5 only | 9 (42.86) | 6 (33.33) | 3 (100.00) | |
| Antibiotic and steroid treatment 6 only | 8 (38.10) | 8 (44.44) | 0 (0) | |
| Other type of treatment | 4 (19.05) | 4 (22.22) | 0 (0) | |
| Duration between antibiotic treatment and sample collection (n, %) | 0.782 | |||
| Less than 2 weeks | 7 (29.17) | 6 (28.57) | 1 (33.33) | |
| More than 2 weeks | 9 (37.50) | 7 (33.33) | 2 (66.67) | |
| Duration missing | 1 (4.17) | 1 (4.76) | 0 (0) | |
| Did not receive antibiotic treatment | 7 (29.17) | 7 (33.33) | 0 (0) |
| n (n, not 0) | Crude | Adjusted 1 | |||||||
|---|---|---|---|---|---|---|---|---|---|
| β 2 | SD 2 | p-Value 3 | q-Value 4 | β 2 | SD 2 | p-Value 3 | q-Value 4 | ||
| Genus | |||||||||
| Ochrobactrum | 42 (40) | −2.770 | 0.154 | <0.001 | <0.001 | −2.770 | 0.156 | <0.001 | <0.001 *** |
| Delftia | 42 (42) | −2.469 | 0.196 | <0.001 | <0.001 | −2.469 | 0.196 | <0.001 | <0.001 *** |
| Anaerobacillus | 42 (28) | −2.470 | 0.247 | <0.001 | <0.001 | −2.470 | 0.247 | <0.001 | <0.001 *** |
| Gordonia | 42 (42) | −1.431 | 0.159 | <0.001 | <0.001 | −1.431 | 0.159 | <0.001 | <0.001 *** |
| Methylobacterium | 42 (16) | 1.361 | 0.206 | <0.001 | <0.001 | 1.361 | 0.206 | <0.001 | <0.001 *** |
| Fusobacterium | 42 (39) | −1.577 | 0.249 | <0.001 | <0.001 | −1.577 | 0.249 | <0.001 | <0.001 *** |
| Sphingobium | 42 (15) | 1.111 | 0.232 | <0.001 | 0.003 | 1.111 | 0.225 | <0.001 | 0.003 ** |
| Alkanindiges | 42 (32) | −1.436 | 0.313 | <0.001 | 0.006 | −1.436 | 0.297 | <0.001 | 0.004 ** |
| Streptococcus | 42 (42) | −0.884 | 0.168 | <0.001 | 0.002 | −0.884 | 0.168 | <0.001 | 0.005 ** |
| Achromobacter | 42 (16) | 0.832 | 0.167 | <0.001 | 0.003 | 0.832 | 0.167 | <0.001 | 0.009 ** |
| Capnocytophaga | 42 (15) | 1.131 | 0.249 | <0.001 | 0.006 | 1.131 | 0.249 | <0.001 | 0.022* |
| Mycobacterium | 42 (22) | 1.404 | 0.350 | <0.001 | 0.007 | 1.404 | 0.344 | <0.001 | 0.022 * |
| Novosphingobium | 42 (13) | 0.627 | 0.142 | <0.001 | 0.006 | 0.627 | 0.142 | <0.001 | 0.024 * |
| Peptoniphilus | 42 (42) | −1.174 | 0.271 | <0.001 | 0.008 | −1.174 | 0.271 | <0.001 | 0.025 * |
| Rothia | 42 (37) | −1.038 | 0.239 | <0.001 | 0.008 | −1.038 | 0.239 | <0.001 | 0.025 * |
| Finegoldia | 42 (42) | −1.152 | 0.271 | <0.001 | 0.008 | −1.152 | 0.271 | <0.001 | 0.027 * |
| Burkholderia | 42 (15) | 0.699 | 0.178 | <0.001 | 0.008 | 0.699 | 0.181 | <0.001 | 0.028 * |
| Roseomonas | 42 (25) | 1.328 | 0.340 | <0.001 | 0.008 | 1.328 | 0.348 | <0.001 | 0.031 * |
| Anaerococcus | 42 (42) | −0.808 | 0.221 | 0.002 | 0.027 | −0.808 | 0.216 | <0.001 | 0.036 * |
| Agrobacterium | 42 (22) | 1.336 | 0.378 | 0.001 | 0.019 | 1.336 | 0.380 | 0.001 | 0.065 |
| Hydrogenophaga | 42 (11) | 0.914 | 0.276 | 0.002 | 0.030 | 0.914 | 0.263 | 0.001 | 0.069 |
| Peptostreptococcus | 42 (9) | 0.768 | 0.226 | 0.002 | 0.035 | 0.768 | 0.226 | 0.003 | 0.120 |
| Kocuria | 42 (18) | 0.877 | 0.280 | 0.003 | 0.043 | 0.877 | 0.284 | 0.004 | 0.144 |
| Dermabacter | 42 (17) | 0.996 | 0.318 | 0.003 | 0.043 | 0.996 | 0.324 | 0.004 | 0.145 |
| Species | |||||||||
| Acinetobacter schindleri | 42 (21) | −1.571 | 0.226 | <0.001 | <0.001 | −1.571 | 0.226 | <0.001 | <0.001 *** |
| Rothia mucilaginosa | 42 (30) | −1.472 | 0.285 | <0.001 | <0.001 | −1.472 | 0.289 | <0.001 | 0.002 ** |
| Lactobacillus iners | 42 (13) | 0.695 | 0.183 | <0.001 | 0.021 | 0.695 | 0.183 | <0.001 | 0.039 * |
| Corynebacterium kroppenstedtii | 42 (38) | −0.971 | 0.245 | <0.001 | 0.019 | −0.971 | 0.245 | <0.001 | 0.053 |
| Roseomonas mucosa | 42 (24) | 1.071 | 0.321 | 0.002 | 0.033 | 1.071 | 0.329 | 0.002 | 0.116 |
| Kocuria rhizophila | 42 (12) | 0.780 | 0.246 | 0.003 | 0.045 | 0.780 | 0.251 | 0.004 | 0.162 |
| Primer | Sequences (5′-3′) |
|---|---|
| 338F | ACTYCTACGGRAGGCWGC |
| 1061R | CRRCACGAGCTGACGAC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ong, S.S.; Xu, J.; Sim, C.K.; Khng, A.J.; Ho, P.J.; Kwan, P.K.W.; Ravikrishnan, A.; Tan, K.-T.B.; Tan, Q.T.; Tan, E.Y.; et al. Profiling Microbial Communities in Idiopathic Granulomatous Mastitis. Int. J. Mol. Sci. 2023, 24, 1042. https://doi.org/10.3390/ijms24021042
Ong SS, Xu J, Sim CK, Khng AJ, Ho PJ, Kwan PKW, Ravikrishnan A, Tan K-TB, Tan QT, Tan EY, et al. Profiling Microbial Communities in Idiopathic Granulomatous Mastitis. International Journal of Molecular Sciences. 2023; 24(2):1042. https://doi.org/10.3390/ijms24021042
Chicago/Turabian StyleOng, Seeu Si, Jia Xu, Choon Kiat Sim, Alexis Jiaying Khng, Peh Joo Ho, Philip Kam Weng Kwan, Aarthi Ravikrishnan, Kiat-Tee Benita Tan, Qing Ting Tan, Ern Yu Tan, and et al. 2023. "Profiling Microbial Communities in Idiopathic Granulomatous Mastitis" International Journal of Molecular Sciences 24, no. 2: 1042. https://doi.org/10.3390/ijms24021042
APA StyleOng, S. S., Xu, J., Sim, C. K., Khng, A. J., Ho, P. J., Kwan, P. K. W., Ravikrishnan, A., Tan, K.-T. B., Tan, Q. T., Tan, E. Y., Tan, S.-M., Putti, T. C., Lim, S. H., Tang, E. L. S., Nagarajan, N., Karnani, N., Li, J., & Hartman, M. (2023). Profiling Microbial Communities in Idiopathic Granulomatous Mastitis. International Journal of Molecular Sciences, 24(2), 1042. https://doi.org/10.3390/ijms24021042







