Unraveling Molecular and Genetic Insights into Neurodegenerative Diseases: Advances in Understanding Alzheimer’s, Parkinson’s, and Huntington’s Diseases and Amyotrophic Lateral Sclerosis
Abstract
1. Introduction
2. Alzheimer’s Disease (AD)
2.1. Amyloid Precursor Protein
2.2. Presenilin1
2.3. Presenilin2
2.4. Apolipoprotein E
2.5. APOE and Tau
2.6. APOE and Neuroinflammation
2.7. Important APOE Mutations Involved in AD Onset
2.7.1. c.-488C>A
2.7.2. c.-24+38G>A
2.7.3. c.-24+288G>A
2.7.4. c.-23-377A>G
2.7.5. A18T
2.8. Microtubule-Associated Protein Tau
2.9. Important MAPT Mutations Involved in AD Onset
2.9.1. MAPT IVS10+12 C>T
2.9.2. MAPT A152T
2.9.3. MAPT K257T
2.9.4. MAPT L266V
2.10. The Evolving Landscape of Alzheimer’s Disease Donanemab Treatment: Exploring Current and Future Perspectives
2.11. Brief Reflection Point
3. Parkinson’s Disease (PD)
3.1. Perspectives of Treatment
3.2. Brief Reflection Point
4. Huntington’s Disease (HD)
4.1. Treatment
4.2. Brief Reflection Point
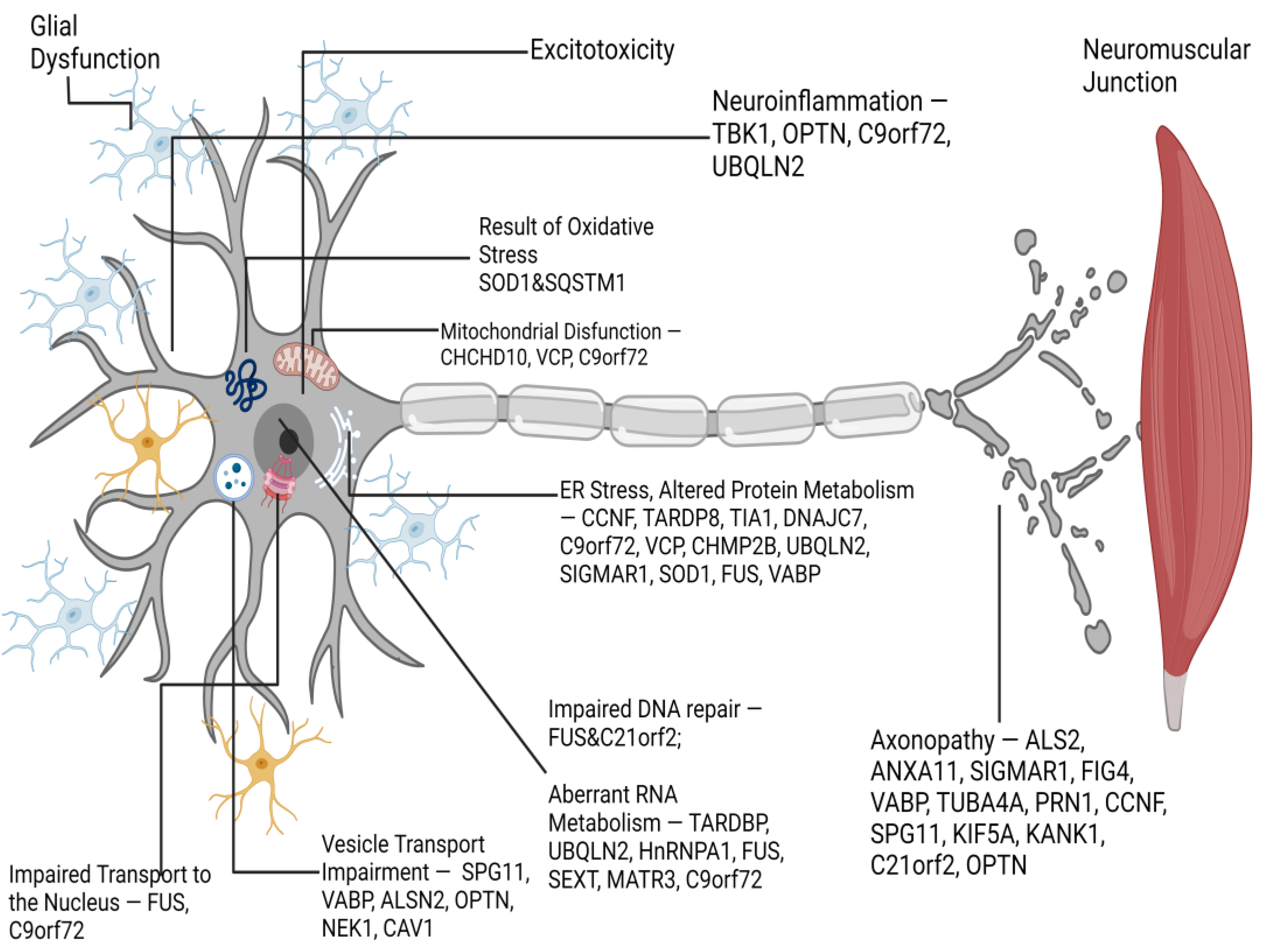
5. Amyotrophic Lateral Sclerosis (ALS)
5.1. Perspectives for Treatment
5.2. Brief Reflection Point
6. Conclusions and Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ND | Neurodegenerative disease |
| AD | Alzheimer’s disease |
| APP | Amyloid precursor protein |
| Aβ | Amyloid beta peptide |
| HCHWA-D | Hereditary cerebral hemorrhage with amyloidosis |
| PiD | Pick’s disease |
| GFAP | Glial fibrillary acidic protein |
| SPECT | Single-photon emission computed tomography |
| PET | Positron emission tomography |
| APOE | Apolipoprotein E |
| BBB | Brain Blood Barrier |
| HDL | High density lipoprotein |
| CNS | Central Nervous System |
| NFTs | Neurofibrillary Tangles |
| GWAS | Genome-wide association study |
| PP2A | Protein phosphatase 2A |
| FTD | Frontotemporal Dementia |
| CTD | Chronic traumatic encephalopathy |
| CBD | Corticobasal degeneration |
| DAM | Disease-associated microglia |
| MGnD | Microglial neurodegenerative |
| TREM2 | Triggering receptor expressed on myeloid cells 2 |
| CRP | C-reactive protein |
| MCI | Mild cognitive impairment |
| MAPT | Microtubule-associated protein tau |
| PSP | Progressive supranuclear palsy |
| PHF | Abnormally hyperphosphorylated taus |
| CL | Centiloids |
| SUVR | Standardized Uptake Ration |
| PD | Parkinson’s disease |
| SNpc | Substantia nigra pars compacta |
| DA | Dopamine |
| DMV | Dorsal motor nucleus of vagus |
| OB | Olfactory bulb |
| LC | Locus coeruleus |
| IML | Intermediolateral nucleus in spinal cord |
| ENS | Enteric nervous system |
| EOPD | Early-onset Parkinson’s disease |
| mitoQC | Mitochondrial quality control |
| DFO | Deferrioxamine |
| DFP | Deferiprone |
| HD | Huntington’s disease |
| MSNs | Medium spiny neurons |
| mHTT | Mutation of the huntingtin gene |
| HTT | Huntingtin gene |
| CBP | cAMP response element-binding protein |
| MSK1 | Mitogen-activated and stress-activated protein kinase 1 |
| PGC-1a | Proliferator-activated receptor gamma coactivator alpha |
| HDAC | Histone deacetylase |
| BDNF | Brain-derived neurotrophic factor |
| ASOs | Antisense oligonucleotides |
| RNAi | RNA interference |
| CoQ10 | Coenzyme Q10 |
| CREST-E | Creatine safety tolerability efficacy in Huntington’s disease |
| ALS | Amyotrophic lateral sclerosis |
| FALS | Familial amyotrophic lateral sclerosis |
| DPR | Dipeptide repeat proteins |
| FUS | Fused in Sarcoma |
| miRNA | Micro RNA |
| PTM | Post-translational modification |
| UMN | Upper motor neuron |
| LMN | Lower motor neuron |
| PMA | Progressive muscular atrophy |
| PBP | Pseudobulbar Palsy |
References
- Berry, R.M. The genetic revolution and the physician’s duty of confidentiality: The role of the old Hippocratic virtues in the regulation of the new genetic intimacy. BMJ 1997, 18, 401–441. [Google Scholar] [CrossRef] [PubMed]
- De Castro, M. Johann Gregor Mendel: Paragon of experimental science. Mol. Genet. Genom. Med. 2016, 4, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Saceleanu, V.M.; Mohan, A.G.; Covache-Busuioc, R.A.; Costin, H.P.; Ciurea, A.V. Wilhelm von Waldeyer: Important Steps in Neural Theory, Anatomy and Citology. Brain Sci. 2022, 12, 224. [Google Scholar] [CrossRef] [PubMed]
- Kovacs, G.G. Concepts and classification of neurodegenerative diseases. In Handbook of Clinical Neurology; Elsevier: Amsterdam, The Netherlands, 2018; pp. 301–307. [Google Scholar] [CrossRef]
- Brettschneider, J.; Del Tredici, K.; Lee, V.M.-Y.; Trojanowski, J.Q. Spreading of pathology in neurodegenerative diseases: A focus on human studies. Nat. Rev. Neurosci. 2015, 16, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.L.; Lee, V.M.Y. Cell-to-cell transmission of pathogenic proteins in neurodegenerative diseases. Nat. Med. 2014, 20, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Castellani, R.J.; Rolston, R.K.; Smith, M.A. Alzheimer Disease. Disease-a-Month 2010, 56, 484–546. [Google Scholar] [CrossRef]
- Ciurea, V.A.; Covache-Busuioc, R.-A.; Mohan, A.G.; Costin, H.P.; Voicu, V.; Academy, B.R. Alzheimer’s disease: 120 years of research and progress. J. Med. Life 2023, 16, 173–177. [Google Scholar] [CrossRef]
- Gatz, M.; Reynolds, C.A.; Fratiglioni, L.; Johansson, B.; Mortimer, J.A.; Berg, S.; Fiske, A.; Pedersen, N.L. Role of Genes and Environments for Explaining Alzheimer Disease. Arch. Gen. Psychiatry 2006, 63, 168–174. [Google Scholar] [CrossRef]
- An, F.; Zhao, R.; Xuan, X.; Xuan, T.; Zhang, G.; Wei, C. Calycosin ameliorates advanced glycation end product-induced neurodegenerative changes in cellular and rat models of diabetes-related Alzheimer’s disease. Chem. Interact. 2022, 368, 110206. [Google Scholar] [CrossRef]
- Van Cauwenberghe, C.; Van Broeckhoven, C.; Sleegers, K. The genetic landscape of Alzheimer disease: Clinical implications and perspectives. Anesth. Analg. 2016, 18, 421–430. [Google Scholar] [CrossRef]
- Roberts, S.; Ripellino, J.; Ingalls, K.; Robakis, N.; Felsenstein, K. Non-amyloidogenic cleavage of the beta-amyloid precursor protein by an integral membrane metalloendopeptidase. J. Biol. Chem. 1994, 269, 3111–3116. [Google Scholar] [CrossRef]
- Gao, C.-X.; Wu, Q.; Sun, J.-X.; Song, X.-H.; Wang, J.; Xiong, C.-Q.; Teng, F.-X. Blocking beta 2-adrenergic receptor inhibits dendrite ramification in a mouse model of Alzheimer’s disease. Neural Regen. Res. 2017, 12, 1499–1506. [Google Scholar] [CrossRef]
- Okochi, M.; Tagami, S.; Yanagida, K.; Takami, M.; Kodama, T.S.; Mori, K.; Nakayama, T.; Ihara, Y.; Takeda, M. γ-Secretase Modulators and Presenilin 1 Mutants Act Differently on Presenilin/γ-Secretase Function to Cleave Aβ42 and Aβ43. Cell Rep. 2013, 3, 42–51. [Google Scholar] [CrossRef]
- Hampel, H.; Hu, Y.; Hardy, J.; Blennow, K.; Chen, C.; Perry, G.; Kim, S.H.; Villemagne, V.L.; Aisen, P.; Vendruscolo, M.; et al. The amyloid-β pathway in Alzheimer’s disease: A plain language summary. Neurodegener. Dis. Manag. 2023, 13, 141–201. [Google Scholar] [CrossRef]
- Tosh, J.L.; Rhymes, E.R.; Mumford, P.; Whittaker, H.T.; Pulford, L.J.; Noy, S.J.; Cleverley, K.; Strydom, A.; Fisher, E.M.C.; Wiseman, F.K.; et al. Genetic dissection of down syndrome-associated alterations in APP/amyloid-β biology using mouse models. Sci. Rep. 2021, 11, 5736. [Google Scholar] [CrossRef] [PubMed]
- Jonsson, T.; Atwal, J.K.; Steinberg, S.; Snaedal, J.; Jonsson, P.V.; Bjornsson, S.; Stefansson, H.; Sulem, P.; Gudbjartsson, D.F.; Maloney, J.; et al. A mutation in APP protects against Alzheimer’s disease and age-related cognitive decline. Nature 2012, 488, 96–99. [Google Scholar] [CrossRef]
- Benilova, I.; Gallardo, R.; Ungureanu, A.-A.; Cano, V.C.; Snellinx, A.; Ramakers, M.; Bartic, C.; Rousseau, F.; Schymkowitz, J.; De Strooper, B. The Alzheimer Disease Protective Mutation A2T Modulates Kinetic and Thermodynamic Properties of Amyloid-β (Aβ) Aggregation. J. Biol. Chem. 2014, 289, 30977–30989. [Google Scholar] [CrossRef]
- Liu, Y.-W.; He, Y.-H.; Zhang, Y.-X.; Cai, W.-W.; Yang, L.-Q.; Xu, L.-Y.; Kong, Q.-P. Absence of A673T variant in APP gene indicates an alternative protective mechanism contributing to longevity in Chinese individuals. Neurobiol. Aging 2014, 35, 935.e11–935.e12. [Google Scholar] [CrossRef] [PubMed]
- Mengel-From, J.; Jeune, B.; Pentti, T.; McGue, M.; Christensen, K.; Christiansen, L. The APP A673T frequency differs between Nordic countries. Neurobiol. Aging 2015, 36, 2909.e1–2909.e4. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Giaccone, G.; Morbin, M.; Moda, F.; Botta, M.; Mazzoleni, G.; Uggetti, A.; Catania, M.; Moro, M.L.; Redaelli, V.; Spagnoli, A.; et al. Neuropathology of the recessive A673V APP mutation: Alzheimer disease with distinctive features. Acta Neuropathol. 2010, 120, 803–812. [Google Scholar] [CrossRef] [PubMed]
- Villemagne, V.L.; Ataka, S.; Mizuno, T.; Brooks, W.S.; Wada, Y.; Kondo, M.; Jones, G.; Watanabe, Y.; Mulligan, R.S.; Nakagawa, M.; et al. High Striatal Amyloid β-Peptide Deposition Across Different Autosomal Alzheimer Disease Mutation Types. Arch. Neurol. 2009, 66, 1537–1544. [Google Scholar] [CrossRef] [PubMed]
- Readhead, B.; Haure-Mirande, J.-V.; Zhang, B.; Haroutunian, V.; Gandy, S.; Schadt, E.E.; Dudley, J.T.; Ehrlich, M.E. Molecular systems evaluation of oligomerogenic APPE693Q and fibrillogenic APPKM670/671NL/PSEN1Δexon9 mouse models identifies shared features with human Alzheimer’s brain molecular pathology. Mol. Psychiatry 2016, 21, 1099–1111. [Google Scholar] [CrossRef] [PubMed]
- Natté, R.; Maat-Schieman, M.L.C.; Haan, J.; Bornebroek, M.; Roos, R.A.; Van Duinen, S.G. Dementia in hereditary cerebral hemorrhage with amyloidosis-Dutch type is associated with cerebral amyloid angiopathy but is independent of plaques and neurofibrillary tangles. Ann. Neurol. 2001, 50, 765–772. [Google Scholar] [CrossRef] [PubMed]
- Nishitsuji, K.; Tomiyama, T.; Ishibashi, K.; Kametani, F.; Ozawa, K.; Okada, R.; Maat-Schieman, M.L.; Roos, R.A.; Iwai, K.; Mori, H. Cerebral vascular accumulation of Dutch-type Aβ42, but not wild-type Aβ42, in hereditary cerebral hemorrhage with amyloidosis, Dutch type. J. Neurosci. Res. 2007, 85, 2917–2923. [Google Scholar] [CrossRef]
- Uddin, M.S.; Tewari, D.; Sharma, G.; Kabir, M.T.; Barreto, G.E.; Bin-Jumah, M.N.; Perveen, A.; Abdel-Daim, M.M.; Ashraf, G.M. Molecular Mechanisms of ER Stress and UPR in the Pathogenesis of Alzheimer’s Disease. Mol. Neurobiol. 2020, 57, 2902–2919. [Google Scholar] [CrossRef] [PubMed]
- Alberdi, E.; Wyssenbach, A.; Alberdi, M.; Sánchez-Gómez, M.V.; Cavaliere, F.; Rodríguez, J.J.; Verkhratsky, A.; Matute, C. Ca2+-dependent endoplasmic reticulum stress correlates with astrogliosis in oligomeric amyloid β-treated astrocytes and in a model of Alzheimer’s disease. Aging Cell 2013, 12, 292–302. [Google Scholar] [CrossRef]
- Tomiyama, T.; Nagata, T.; Shimada, H.; Teraoka, R.; Fukushima, A.; Kanemitsu, H.; Takuma, H.; Kuwano, R.; Imagawa, M.; Ataka, S.; et al. A new amyloid β variant favoring oligomerization in Alzheimer’s-type dementia. Ann. Neurol. 2008, 63, 377–387. [Google Scholar] [CrossRef]
- McKnelly, K.J.; Kreutzer, A.G.; Howitz, W.J.; Haduong, K.; Yoo, S.; Hart, C.; Nowick, J.S. Effects of Familial Alzheimer’s Disease Mutations on the Assembly of a β-Hairpin Peptide Derived from Aβ16–36. Biochemistry 2022, 61, 446–454. [Google Scholar] [CrossRef]
- Paik, Y.K.; Chang, D.J.; Reardon, C.A.; Walker, M.D.; Taxman, E.; Taylor, J.M. Identification and characterization of transcriptional regulatory regions associated with expression of the human apolipoprotein E gene. J. Biol. Chem. 1988, 263, 13340–13349. [Google Scholar] [CrossRef]
- Maloney, B.; Ge, Y.-W.; Alley, G.M.; Lahiri, D.K. Important differences between human and mouse APOE gene promoters: Limitation of mouse APOE model in studying Alzheimer’s disease. J. Neurochem. 2007, 103, 1237–1257. [Google Scholar] [CrossRef]
- Yee, A.; Tsui, N.B.; Kwan, R.Y.; Leung, A.Y.; Lai, C.K.; Chung, T.; Lau, J.Y.; Fok, M.; Dai, D.L.; Lau, L.-T. Apolipoprotein E Gene Revisited: Contribution of Rare Variants to Alzheimer’s Disease Susceptibility in Southern Chinese. Curr. Alzheimer Res. 2021, 18, 67–79. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Mägi, R.; Milani, L.; Lauschke, V.M. Global genetic diversity of human apolipoproteins and effects on cardiovascular disease risk. J. Lipid Res. 2018, 59, 1987–2000. [Google Scholar] [CrossRef] [PubMed]
- Saunders, A.M. Gene identification in Alzheimer’s disease. Pharmacogenomics 2001, 2, 239–249. [Google Scholar] [CrossRef] [PubMed]
- Theuns, J.; Del-Favero, J.; Dermaut, B.; van Duijn, C.M.; Backhovens, H.; Broeck, M.V.D.; Serneels, S.; Corsmit, E.; Van Broeckhoven, C.; Cruts, M. Genetic variability in the regulatory region of presenilin 1 associated with risk for Alzheimer’s disease and variable expression. Hum. Mol. Genet. 2000, 9, 325–331. [Google Scholar] [CrossRef] [PubMed]
- Cruts, M.; Theuns, J.; Van Broeckhoven, C. Locus-specific mutation databases for neurodegenerative brain diseases. Hum. Mutat. 2012, 33, 1340–1344. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Zhou, R.; Yang, G.; Shi, Y. Analysis of 138 pathogenic mutations in presenilin-1 on the in vitro production of Aβ42 and Aβ40 peptides by γ-secretase. Proc. Natl. Acad. Sci. USA 2016, 114, E476–E485. [Google Scholar] [CrossRef]
- Wines-Samuelson, M.; Schulte, E.C.; Smith, M.J.; Aoki, C.; Liu, X.; Kelleher, R.J.; Shen, J. Characterization of Age-Dependent and Progressive Cortical Neuronal Degeneration in Presenilin Conditional Mutant Mice. PLoS ONE 2010, 5, e10195. [Google Scholar] [CrossRef]
- Kauwe, J.S.K.; Jacquart, S.; Chakraverty, S.; Wang, J.; Mayo, K.; Fagan, A.M.; Holtzman, D.M.; Morris, J.C.; Goate, A.M. Extreme cerebrospinal fluid amyloid β levels identify family with late-onset Alzheimer’s disease presenilin 1 mutation. Ann. Neurol. 2007, 61, 446–453. [Google Scholar] [CrossRef]
- Kumar-Singh, S.; Theuns, J.; Van Broeck, B.; Pirici, D.; Vennekens, K.; Corsmit, E.; Cruts, M.; Dermaut, B.; Wang, R.; Van Broeckhoven, C. Mean age-of-onset of familial alzheimer disease caused by presenilin mutations correlates with both increased Aβ42 and decreased Aβ40. Hum. Mutat. 2006, 27, 686–695. [Google Scholar] [CrossRef]
- Koriath, C.; Kenny, J.; Adamson, G.; Druyeh, R.; Taylor, W.; Beck, J.; Quinn, L.; Mok, T.H.; Dimitriadis, A.; Norsworthy, P.; et al. Predictors for a dementia gene mutation based on gene-panel next-generation sequencing of a large dementia referral series. Mol. Psychiatry 2020, 25, 3399–3412. [Google Scholar] [CrossRef]
- Gallo, M.; Frangipane, F.; Cupidi, C.; De Bartolo, M.; Turone, S.; Ferrari, C.; Nacmias, B.; Grimaldi, G.; Laganà, V.; Colao, R.; et al. The novel PSEN1 M84V mutation associated to frontal dysexecutive syndrome, spastic paraparesis, and cerebellar atrophy in a dominant Alzheimer’s disease family. Neurobiol. Aging 2017, 56, 213.e7–213.e12. [Google Scholar] [CrossRef] [PubMed]
- Hsu, S.; Gordon, B.A.; Hornbeck, R.; Norton, J.B.; Levitch, D.; Louden, A.; Ziegemeier, E.; Laforce, R., Jr.; Chhatwal, J.; Day, G.S.; et al. Discovery and validation of autosomal dominant Alzheimer’s disease mutations. Alzheimers Res. Ther. 2018, 10, 67. [Google Scholar] [CrossRef] [PubMed]
- Kakuda, N.; Takami, M.; Okochi, M.; Kasuga, K.; Ihara, Y.; Ikeuchi, T. Switched Aβ43 generation in familial Alzheimer’s disease with presenilin 1 mutation. Transl. Psychiatry 2021, 11, 558. [Google Scholar] [CrossRef]
- López-García, S.; Jiménez-Bonilla, J.; Delgado, A.L.; Balaguer, P.O.; Ceberio, J.I.; Marraco, I.B.; Rodríguez, E.R.; Sánchez-Juan, P. A Rare PSEN1 (Leu85Pro) Mutation Causing Alzheimer’s Disease in a 29-Year-Old Woman Presenting as Corticobasal Syndrome. J. Alzheimers Dis. 2019, 70, 655–658. [Google Scholar] [CrossRef] [PubMed]
- Arber, C.; Llerena, C.V.; Toombs, J.; Pocock, J.M.; Ryan, N.S.; Fox, N.C.; Zetterberg, H.; Hardy, J.; Wray, S. Amyloid precursor protein processing in human neurons with an allelic series of the PSEN1 intron 4 deletion mutation and total presenilin-1 knockout. Brain Commun. 2019, 1, fcz024. [Google Scholar] [CrossRef] [PubMed]
- De Jonghe, C.; Cruts, M.; Rogaeva, E.A.; Tysoe, C.; Singleton, A.; Vanderstichele, H.; Meschino, W.; Dermaut, B.; Vanderhoeven, I.; Backhovens, H.; et al. Aberrant Splicing in the Presenilin-1 Intron 4 Mutation Causes Presenile Alzheimer’s Disease by Increased A 42 Secretion. Hum. Mol. Genet. 1999, 8, 1529–1540. [Google Scholar] [CrossRef] [PubMed]
- Szaruga-Bracke, M.; Veugelen, S.; Benurwar, M.; Lismont, S.; Sepulveda-Falla, D.; Lleo, A.; Ryan, N.S.; Lashley, T.; Fox, N.; Murayama, S.; et al. Qualitative changes in human γ-secretase underlie familial Alzheimer’s disease. J. Exp. Med. 2015, 212, 2003–2013. [Google Scholar] [CrossRef]
- Fox, N.C.; Kennedy, A.M.; Harvey, R.J.; Lantos, P.L.; Roques, P.K.; Collinge, J.; Hardy, J.; Hutton, M.; Stevens, J.M.; Warrington, E.K.; et al. Clinicopathological features of familial Alzheimer’s disease associated with the M139V mutation in the presenilin 1 gene. Pedigree but not mutation specific age at onset provides evidence for a further genetic factor. Brain 1997, 120, 491–501. [Google Scholar] [CrossRef]
- Chávez-Gutiérrez, L.; Bammens, L.; Benilova, I.; Vandersteen, A.; Benurwar, M.; Borgers, M.; Lismont, S.; Zhou, L.; Van Cleynenbreugel, S.; Esselmann, H.; et al. The mechanism of γ-Secretase dysfunction in familial Alzheimer disease. EMBO J. 2012, 31, 2261–2274. [Google Scholar] [CrossRef]
- Ringman, J.M.; Gylys, K.H.; Medina, L.D.; Fox, M.; Kepe, V.; Flores, D.L.; Apostolova, L.G.; Barrio, J.R.; Small, G.; Silverman, D.H.; et al. Biochemical, neuropathological, and neuroimaging characteristics of early-onset Alzheimer’s disease due to a novel PSEN1 mutation. Neurosci. Lett. 2011, 487, 287–292. [Google Scholar] [CrossRef]
- Devi, G.; Fotiou, A.; Jyrinji, D.; Tycko, B.; DeArmand, S.; Rogaeva, E.; Song, Y.-Q.; Medieros, H.; Liang, Y.; Orlacchio, A.; et al. Novel presenilin 1 mutations associated with early onset of dementia in a family with both early-onset and late-onset Alzheimer disease. Arch. Neurol. 2000, 57, 1454–1457. [Google Scholar] [CrossRef] [PubMed]
- Levy-Lahad, E.; Wasco, W.; Poorkaj, P.; Romano, D.M.; Oshima, J.; Pettingell, W.H.; Yu, C.E.; Jondro, P.D.; Schmidt, S.D.; Wang, K.; et al. Candidate gene for the chromosome 1 familial Alzheimer’s disease locus. Science 1995, 269, 973–977. [Google Scholar] [CrossRef] [PubMed]
- Perrone, F.; Cacace, R.; Van Mossevelde, S.; Bossche, T.V.D.; De Deyn, P.P.; Cras, P.; Engelborghs, S.; van der Zee, J.; Van Broeckhoven, C. Genetic screening in early-onset dementia patients with unclear phenotype: Relevance for clinical diagnosis. Neurobiol. Aging 2018, 69, 292.e7–292.e14. [Google Scholar] [CrossRef] [PubMed]
- Jia, L.; Fu, Y.; Shen, L.; Zhang, H.; Zhu, M.; Qiu, Q.; Wang, Q.; Yan, X.; Kong, C.; Hao, J.; et al. PSEN1, PSEN2, and APP mutations in 404 Chinese pedigrees with familial Alzheimer’s disease. Alzheimers Dement. 2020, 16, 178–191. [Google Scholar] [CrossRef]
- Pang, Y.; Li, T.; Wang, Q.; Qin, W.; Li, Y.; Wei, Y.; Jia, L. A Rare Variation in the 3′ Untranslated Region of the Presenilin 2 Gene Is Linked to Alzheimer’s Disease. Mol. Neurobiol. 2021, 58, 4337–4347. [Google Scholar] [CrossRef]
- Wallon, D.; Rousseau, S.; Rovelet-Lecrux, A.; Quillard-Muraine, M.; Guyant-Maréchal, L.; Martinaud, O.; Pariente, J.; Puel, M.; Rollin-Sillaire, A.; Pasquier, F.; et al. The French Series of Autosomal Dominant Early Onset Alzheimer’s Disease Cases: Mutation Spectrum and Cerebrospinal Fluid Biomarkers. J. Alzheimers Dis. 2012, 30, 847–856. [Google Scholar] [CrossRef]
- Marcon, G.; Giaccone, G.; Cupidi, C.; Balestrieri, M.; Beltrami, C.A.; Finato, N.; Bergonzi, P.; Sorbi, S.; Bugiani, O.; Tagliavini, F. Neuropathological and Clinical Phenotype of an Italian Alzheimer Family with M239V Mutation of Presenilin 2 Gene. J. Neuropathol. Exp. Neurol. 2004, 63, 199–209. [Google Scholar] [CrossRef][Green Version]
- Huang, Y.; Weisgraber, K.H.; Mucke, L.; Mahley, R.W. Apolipoprotein E: Diversity of Cellular Origins, Structural and Biophysical Properties, and Effects in Alzheimer’s Disease. J. Mol. Neurosci. 2004, 23, 189–204. [Google Scholar] [CrossRef]
- Liu, C.-C.; Kanekiyo, T.; Xu, H.; Bu, G. Apolipoprotein E and Alzheimer disease: Risk, mechanisms and therapy. Nat. Rev. Neurol. 2013, 9, 106–118. [Google Scholar] [CrossRef]
- Kara, E.; Marks, J.D.; Fan, Z.; Klickstein, J.A.; Roe, A.D.; Krogh, K.A.; Wegmann, S.; Maesako, M.; Luo, C.C.; Mylvaganam, R.; et al. Isoform- and cell type-specific structure of apolipoprotein E lipoparticles as revealed by a novel Forster resonance energy transfer assay. J. Biol. Chem. 2017, 292, 14720–14729. [Google Scholar] [CrossRef]
- Lin, Y.-T.; Seo, J.; Gao, F.; Feldman, H.M.; Wen, H.-L.; Penney, J.; Cam, H.P.; Gjoneska, E.; Raja, W.K.; Cheng, J.; et al. APOE4 Causes Widespread Molecular and Cellular Alterations Associated with Alzheimer’s Disease Phenotypes in Human iPSC-Derived Brain Cell Types. Neuron 2018, 98, 1141–1154.e7. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.S.; Ebbert, M.T.W.; Baker, K.E.; Cook, C.; Wang, X.; Sens, J.P.; Kocher, J.-P.; Petrucelli, L.; Fryer, J.D. Microglial translational profiling reveals a convergent APOE pathway from aging, amyloid, and tau. J. Exp. Med. 2018, 215, 2235–2245. [Google Scholar] [CrossRef] [PubMed]
- Mathys, H.; Davila-Velderrain, J.; Peng, Z.; Gao, F.; Mohammadi, S.; Young, J.Z.; Menon, M.; He, L.; Abdurrob, F.; Jiang, X.; et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 2019, 570, 332–337. [Google Scholar] [CrossRef]
- Therriault, J.; Benedet, A.L.; Pascoal, T.A.; Mathotaarachchi, S.; Chamoun, M.; Savard, M.; Thomas, E.; Kang, M.S.; Lussier, F.; Tissot, C.; et al. Association of Apolipoprotein E ε4 With Medial Temporal Tau Independent of Amyloid-β. JAMA Neurol. 2020, 77, 470–479. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Alzheimer’s Disease Neuroimaging Initiative; Yamada, K.; Liddelow, S.A.; Smith, S.T.; Zhao, L.; Luo, W.; Tsai, R.M.; Spina, S.; Grinberg, L.T.; et al. ApoE4 markedly exacerbates tau-mediated neurodegeneration in a mouse model of tauopathy. Nature 2017, 549, 523–527. [Google Scholar] [CrossRef] [PubMed]
- Harris, F.M.; Brecht, W.J.; Xu, Q.; Mahley, R.W.; Huang, Y. Increased tau Phosphorylation in Apolipoprotein E4 Transgenic Mice Is Associated with Activation of Extracellular Signal-regulated Kinase. J. Biol. Chem. 2004, 279, 44795–44801. [Google Scholar] [CrossRef]
- Wadhwani, A.R.; Affaneh, A.; Van Gulden, S.; Kessler, J.A. Neuronal apolipoprotein E4 increases cell death and phosphorylated tau release in alzheimer disease. Ann. Neurol. 2019, 85, 726–739. [Google Scholar] [CrossRef]
- Gibb, G.M.; Pearce, J.; Betts, J.C.; Lovestone, S.; Hoffmann, M.M.; Maerz, W.; Blackstock, W.P.; Anderton, B.H. Differential effects of apolipoprotein E isoforms on phosphorylation at specific sites on tau by glycogen synthase kinase-3β identified by nano-electrospray mass spectrometry. FEBS Lett. 2000, 485, 99–103. [Google Scholar] [CrossRef]
- Park, J.-S.; Ji-Seon, P.; Sang-Wook, K.; Kang, M.-J.; Kang, S.-W.; Kim, D.-H.; Yoon, S.-Y. Disease-Associated Mutations of TREM2 Alter the Processing of N-Linked Oligosaccharides in the Golgi Apparatus. Traffic 2015, 16, 510–518. [Google Scholar] [CrossRef]
- Saceleanu, V.M.; Covache-Busuioc, R.-A.; Costin, H.-P.; Glavan, L.-A.; Ciurea, A.V. An Important Step in Neuroscience: Camillo Golgi and His Discoveries. Cells 2022, 11, 4112. [Google Scholar] [CrossRef]
- Ulrich, J.D.; Ulland, T.K.; Mahan, T.E.; Nyström, S.; Nilsson, K.P.; Song, W.M.; Zhou, Y.; Reinartz, M.; Choi, S.; Jiang, H.; et al. ApoE facilitates the microglial response to amyloid plaque pathology. J. Exp. Med. 2018, 215, 1047–1058. [Google Scholar] [CrossRef] [PubMed]
- Keren-Shaul, H.; Spinrad, A.; Weiner, A.; Matcovitch-Natan, O.; Dvir-Szternfeld, R.; Ulland, T.K.; David, E.; Baruch, K.; Lara-Astaiso, D.; Toth, B.; et al. A Unique Microglia Type Associated with Restricting Development of Alzheimer’s Disease. Cell 2017, 169, 1276–1290.e17. [Google Scholar] [CrossRef] [PubMed]
- Hutton, M.; Lendon, C.L.; Rizzu, P.; Baker, M.; Froelich, S.; Houlden, H.; Pickering-Brown, S.; Chakraverty, S.; Isaacs, A.; Grover, A.; et al. Association of missense and 5′-splice-site mutations in tau with the inherited dementia FTDP-17. Nature 1998, 393, 702–705. [Google Scholar] [CrossRef] [PubMed]
- Spillantini, M.G.; Bird, T.D.; Ghetti, B. Frontotemporal Dementia and Parkinsonism Linked to Chromosome 17: A New Group of Tauopathies. Brain Pathol. 2006, 8, 387–402. [Google Scholar] [CrossRef] [PubMed]
- Poorkaj, P.; Bird, T.D.; Wijsman, E.; Nemens, E.; Garruto, R.M.; Anderson, L.; Andreadis, A.; Wiederholt, W.C.; Raskind, M.; Schellenberg, G.D. Tau is a candidate gene for chromosome 17 frontotemporal dementia. Ann. Neurol. 1998, 43, 815–825. [Google Scholar] [CrossRef] [PubMed]
- Goedert, M.; Spillantini, M.G.; Jakes, R.; Rutherford, D.; Crowther, R.A. Multiple isoforms of human microtubule-associated protein tau: Sequences and localization in neurofibrillary tangles of Alzheimer’s disease. Neuron 1989, 3, 519–526. [Google Scholar] [CrossRef]
- Baudier, J.; Cole, R. D Interactions between the microtubule-associated tau proteins and S100b regulate tau phosphorylation by the Ca2+/calmodulin-dependent protein kinase II. J. Biol. Chem. 1988, 263, 5876–5883. [Google Scholar] [CrossRef]
- Drewes, G.; Ebneth, A.; Preuss, U.; Mandelkow, E.-M.; Mandelkow, E. MARK, a Novel Family of Protein Kinases That Phosphorylate Microtubule-Associated Proteins and Trigger Microtubule Disruption. Cell 1997, 89, 297–308. [Google Scholar] [CrossRef]
- Drewes, G.; Lichtenberg-Kraag, B.; Döring, F.; Mandelkow, E.; Biernat, J.; Goris, J.; Dorée, M.; Mandelkow, E. Mitogen activated protein (MAP) kinase transforms tau protein into an Alzheimer-like state. EMBO J. 1992, 11, 2131–2138. [Google Scholar] [CrossRef]
- Bancher, C.; Brunner, C.; Lassmann, H.; Budka, H.; Jellinger, K.; Wiche, G.; Seitelberger, F.; Grundke-Iqbal, I.; Iqbal, K.; Wisniewski, H. Accumulation of abnormally phosphorylated τ precedes the formation of neurofibrillary tangles in Alzheimer’s disease. Brain Res. 1989, 477, 90–99. [Google Scholar] [CrossRef]
- Davies, P.; Vincent, I.J. On the role of tau phosphorylation in the formation of PHF. Neurobiol. Aging 1995, 16, 420–422. [Google Scholar] [CrossRef]
- Wang, J.-Z.; Wu, Q.; Smith, A.; Grundke-Iqbal, I.; Iqbal, K. τ is phosphorylated by GSK-3 at several sites found in Alzheimer disease and its biological activity markedly inhibited only after it is prephosphorylated by A-kinase. FEBS Lett. 1998, 436, 28–34. [Google Scholar] [CrossRef]
- Yasuda, M.; Takamatsu, J.; D’Souza, I.; Crowther, R.A.; Kawamata, T.; Hasegawa, M.; Hasegawa, H.; Spillantini, M.G.; Tanimukai, S.; Poorkaj, P.; et al. A novel mutation at position +12 in the intron following exon 10 of the tau gene in familial frontotemporal dementia (FTD-Kumamoto). Ann. Neurol. 2000, 47, 422–429. [Google Scholar] [CrossRef] [PubMed]
- Takamatsu, J.; Kondo, A.; Ikegami, K.; Kimura, T.; Fujii, H.; Mitsuyama, Y.; Hashizume, Y. Selective Expression of Ser 199/202 Phosphorylated Tau in a Case of Frontotemporal Dementia. Dement. Geriatr. Cogn. Disord. 1998, 9, 82–89. [Google Scholar] [CrossRef]
- Labbé, C.; Ogaki, K.; Lorenzo-Betancor, O.; Soto-Ortolaza, A.I.; Walton, R.L.; Rayaprolu, S.; Fujioka, S.; Murray, M.E.; Heckman, M.G.; Puschmann, A.; et al. Role for the microtubule-associated protein tau variant p.A152T in risk of α-synucleinopathies. Neurology 2015, 85, 1680–1686. [Google Scholar] [CrossRef]
- Jin, S.C.; Pastor, P.; Cooper, B.; Cervantes, S.; Benitez, B.A.; Razquin, C.; Goate, A.; Cruchaga, C.; Ibero-American Alzheimer Disease Genetics Group Researchers. Pooled-DNA sequencing identifies novel causative variants in PSEN1, GRN and MAPT in a clinical early-onset and familial Alzheimer’s disease Ibero-American cohort. Alzheimers Res. Ther. 2012, 4, 34. [Google Scholar] [CrossRef] [PubMed]
- Coppola, G.; Chinnathambi, S.; Lee, J.J.; Dombroski, B.A.; Baker, M.C.; Soto-Ortolaza, A.I.; Lee, S.E.; Klein, E.; Huang, A.Y.; Sears, R.; et al. Evidence for a role of the rare p.A152T variant in MAPT in increasing the risk for FTD-spectrum and Alzheimer’s diseases. Hum. Mol. Genet. 2012, 21, 3500–3512. [Google Scholar] [CrossRef]
- Rizzini, C.; Goedert, M.; Hodges, J.R.; Smith, M.J.; Jakes, R.; Hills, R.; Xuereb, J.H.; Crowther, R.A.; Spillantini, M.G. Tau Gene Mutation K257T Causes a Tauopathy Similar to Pick’s Disease. J. Neuropathol. Exp. Neurol. 2000, 59, 990–1001. [Google Scholar] [CrossRef]
- Kobayashi, T.; Ota, S.; Tanaka, K.; Ito, Y.; Hasegawa, M.; Umeda, Y.; Motoi, Y.; Takanashi, M.; Yasuhara, M.; Anno, M.; et al. A novel L266V mutation of the tau gene causes frontotemporal dementia with a unique tau pathology. Ann. Neurol. 2003, 53, 133–137. [Google Scholar] [CrossRef]
- Hogg, M.; Grujic, Z.M.; Baker, M.; Demirci, S.; Guillozet, A.L.; Sweet, A.P.; Herzog, L.L.; Weintraub, S.; Mesulam, M.-M.; LaPointe, N.E.; et al. The L266V tau mutation is associated with frontotemporal dementia and Pick-like 3R and 4R tauopathy. Acta Neuropathol. 2003, 106, 323–336. [Google Scholar] [CrossRef]
- Schilling, S.; Zeitschel, U.; Hoffmann, T.; Heiser, U.; Francke, M.; Kehlen, A.; Holzer, M.; Hutter-Paier, B.; Prokesch, M.; Windisch, M.; et al. Glutaminyl cyclase inhibition attenuates pyroglutamate Aβ and Alzheimer’s disease–like pathology. Nat. Med. 2008, 14, 1106–1111. [Google Scholar] [CrossRef] [PubMed]
- Frost, J.L.; Liu, B.; Kleinschmidt, M.; Schilling, S.; Demuth, H.-U.; Lemere, C.A. Passive Immunization against Pyroglutamate-3 Amyloid-β Reduces Plaque Burden in Alzheimer-Like Transgenic Mice: A Pilot Study. Neurodegener. Dis. 2012, 10, 265–270. [Google Scholar] [CrossRef] [PubMed]
- Frost, J.L.; Liu, B.; Rahfeld, J.-U.; Kleinschmidt, M.; O’Nuallain, B.; Le, K.X.; Lues, I.; Caldarone, B.J.; Schilling, S.; Demuth, H.-U.; et al. An anti-pyroglutamate-3 Aβ vaccine reduces plaques and improves cognition in APPswe/PS1ΔE9 mice. Neurobiol. Aging 2015, 36, 3187–3199. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, T.; Meyer, A.; Heiser, U.; Kurat, S.; Böhme, L.; Kleinschmidt, M.; Bühring, K.-U.; Hutter-Paier, B.; Farcher, M.; DeMuth, H.-U.; et al. Glutaminyl Cyclase Inhibitor PQ912 Improves Cognition in Mouse Models of Alzheimer’s Disease—Studies on Relation to Effective Target Occupancy. Experiment 2017, 362, 119–130. [Google Scholar] [CrossRef] [PubMed]
- DeMattos, R.B.; Lu, J.; Tang, Y.; Racke, M.M.; DeLong, C.A.; Tzaferis, J.A.; Hole, J.T.; Forster, B.M.; McDonnell, P.C.; Liu, F.; et al. A Plaque-Specific Antibody Clears Existing β-amyloid Plaques in Alzheimer’s Disease Mice. Neuron 2012, 76, 908–920. [Google Scholar] [CrossRef]
- Lowe, S.L.; Willis, B.A.; Hawdon, A.; Natanegara, F.; Chua, L.; Foster, J.; Shcherbinin, S.; Ardayfio, P.; Sims, J.R. Donanemab (LY3002813) dose-escalation study in Alzheimer’s disease. Alzheimers Dement. Transl. Res. Clin. Interv. 2021, 7, e12112. [Google Scholar] [CrossRef]
- Lowe, S.L.; Evans, C.D.; Shcherbinin, S.; Cheng, Y.-J.; Willis, B.A.; Gueorguieva, I.; Lo, A.C.; Fleisher, A.S.; Dage, J.L.; Ardayfio, P.; et al. Donanemab (LY3002813) Phase 1b Study in Alzheimer’s Disease: Rapid and Sustained Reduction of Brain Amyloid Measured by Florbetapir F18 Imaging. J. Prev. Alzheimers Dis. 2021, 8, 414–424. [Google Scholar] [CrossRef]
- Shcherbinin, S.; Evans, C.D.; Lu, M.; Andersen, S.W.; Pontecorvo, M.J.; Willis, B.A.; Gueorguieva, I.; Hauck, P.M.; Brooks, D.A.; Mintun, M.A.; et al. Association of Amyloid Reduction After Donanemab Treatment With Tau Pathology and Clinical Outcomes: Association of amyloid reduction after donanemab treatment with tau pathology and clinical outcomes: The TRAILBLAZER-ALZ randomized clinical trial. JAMA Neurol. 2022, 79, 1015. [Google Scholar] [CrossRef]
- Raza, C.; Anjum, R.; Shakeel, N.U.A. Parkinson’s disease: Mechanisms, translational models and management strategies. Life Sci. 2019, 226, 77–90. [Google Scholar] [CrossRef]
- Beitz, J.M. Parkinson s disease: A review. Front. Biosci. 2014, S6, 65–74. [Google Scholar] [CrossRef]
- Day, J.O.; Mullin, S. The Genetics of Parkinson’s Disease and Implications for Clinical Practice. Genes 2021, 12, 1006. [Google Scholar] [CrossRef] [PubMed]
- Ye, H.; Robak, L.A.; Yu, M.; Cykowski, M.; Shulman, J.M. Genetics and Pathogenesis of Parkinson’s Syndrome. Annu. Rev. Pathol. Mech. Dis. 2023, 18, 95–121. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, I.J.; Pervaiz, N.; Abbasi, A.A. The Parkinson Disease gene SNCA: Evolutionary and structural insights with pathological implication. Sci. Rep. 2016, 6, 24475. [Google Scholar] [CrossRef] [PubMed]
- Rui, Q.; Ni, H.; Li, D.; Gao, R.; Chen, G. The Role of LRRK2 in Neurodegeneration of Parkinson Disease. Curr. Neuropharmacol. 2018, 16, 1348–1357. [Google Scholar] [CrossRef] [PubMed]
- Vizziello, M.; Borellini, L.; Franco, G.; Ardolino, G. Disruption of Mitochondrial Homeostasis: The Role of PINK1 in Parkinson’s Disease. Cells 2021, 10, 3022. [Google Scholar] [CrossRef]
- Crichton, R.R.; Dexter, D.T.; Ward, R.J. Brain iron metabolism and its perturbation in neurological diseases. In Metal Ions in Neurological Systems; Linert, W., Kozlowski, H., Eds.; Springer Vienna: Vienna, Austria, 2012; pp. 1–15. [Google Scholar] [CrossRef]
- Chandra, G.; Shenoi, R.; Anand, R.; Rajamma, U.; Mohanakumar, K. Reinforcing mitochondrial functions in aging brain: An insight into Parkinson’s disease therapeutics. J. Chem. Neuroanat. 2019, 95, 29–42. [Google Scholar] [CrossRef]
- Borgna-Pignatti, C.; Rugolotto, S.; De Stefano, P.; Zhao, H.; Cappellini, M.D.; Del Vecchio, G.C.; Romeo, M.A.; Forni, G.L.; Gamberini, M.R.; Ghilardi, R.; et al. Survival and complications in patients with thalassemia major treated with transfusion and deferoxamine. Haematologica 2004, 89, 1187–1193. [Google Scholar]
- Boddaert, N.; Sang, K.H.L.Q.; Rötig, A.; Leroy-Willig, A.; Gallet, S.; Brunelle, F.; Sidi, D.; Thalabard, J.-C.; Munnich, A.; Cabantchik, Z.I. Selective iron chelation in Friedreich ataxia: Biologic and clinical implications. Blood 2007, 110, 401–408. [Google Scholar] [CrossRef]
- Devos, D.; Moreau, C.; Devedjian, J.C.; Kluza, J.; Petrault, M.; Laloux, C.; Jonneaux, A.; Ryckewaert, G.; Garçon, G.; Rouaix, N.; et al. Targeting Chelatable Iron as a Therapeutic Modality in Parkinson’s Disease. Antioxid. Redox Signal. 2014, 21, 195–210. [Google Scholar] [CrossRef]
- McColgan, P.; Tabrizi, S.J. Huntington’s disease: A clinical review. Eur. J. Neurol. 2017, 25, 24–34. [Google Scholar] [CrossRef]
- Wexler, A.; Wild, E.; Tabrizi, S.J. George Huntington: A legacy of inquiry, empathy and hope. Brain 2016, 139, 2326–2333. [Google Scholar] [CrossRef]
- Bates, G.P.; Dorsey, R.; Gusella, J.F.; Hayden, M.R.; Kay, C.; Leavitt, B.R.; Nance, M.; Ross, C.A.; Scahill, R.I.; Wetzel, R.; et al. Huntington Disease. Nat. Rev. Dis. Prim. 2015, 1, 15005. [Google Scholar] [CrossRef]
- MacDonald, M.E.; Ambrose, C.M.; Duyao, M.P.; Myers, R.H.; Lin, C.; Srinidhi, L.; Barnes, G.; Taylor, S.A.; James, M.; Groot, N.; et al. The Huntington’s Disease Collaborative Research Group: A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 1993, 72, 971–983. [Google Scholar] [CrossRef]
- Squitieri, F.; Andrew, S.; Goldberg, Y.; Kremer, B.; Spence, N.; Zelsler, J.; Nichol, K.; Theilmann, J.; Greenberg, J.; Goto, J.; et al. DNA haplotype analysis of Huntington disease reveals clues to the origins and mechanisms of CAG expansion and reasons for geographic variations of prevalence. Hum. Mol. Genet. 1994, 3, 2103–2114. [Google Scholar] [CrossRef] [PubMed]
- Persichetti, F.; Srinidhi, J.; Kanaley, L.; Ge, P.; Myers, R.H.; D’Arrigo, K.; Barnes, G.T.; MacDonald, M.E.; Vonsattel, J.-P.; Gusella, J.F.; et al. Huntington’s disease CAG trinucleotide repeats in pathologically confirmed post-mortem brains. Neurobiol. Dis. 1994, 1, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Gusella, J.F.; MacDonald, M.E.; Lee, J.-M. Genetic modifiers of Huntington’s disease. Mov. Disord. 2014, 29, 1359–1365. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-M.; Gillis, T.; Mysore, J.S.; Ramos, E.M.; Myers, R.H.; Hayden, M.R.; Morrison, P.J.; Nance, M.; Ross, C.A.; Margolis, R.L.; et al. Common SNP-Based Haplotype Analysis of the 4p16.3 Huntington Disease Gene Region. Am. J. Hum. Genet. 2012, 90, 434–444. [Google Scholar] [CrossRef]
- Landles, C.; Sathasivam, K.; Weiss, A.; Woodman, B.; Moffitt, H.; Finkbeiner, S.; Sun, B.; Gafni, J.; Ellerby, L.M.; Trottier, Y.; et al. Proteolysis of Mutant Huntingtin Produces an Exon 1 Fragment That Accumulates as an Aggregated Protein in Neuronal Nuclei in Huntington Disease. J. Biol. Chem. 2010, 285, 8808–8823. [Google Scholar] [CrossRef]
- Ha, A.D.; Fung, V.S. Huntington’s disease. Curr. Opin. Neurol. 2012, 25, 491–498. [Google Scholar] [CrossRef]
- Ross, C.A.; Tabrizi, S.J. Huntington’s disease: From molecular pathogenesis to clinical treatment. Lancet Neurol. 2011, 10, 83–98. [Google Scholar] [CrossRef]
- Vonsattel, J.P.G.; DiFiglia, M. Huntington Disease. J. Neuropathol. Exp. Neurol. 1998, 57, 369–384. [Google Scholar] [CrossRef]
- Moss, D.J.H.; Pardiñas, A.F.; Langbehn, D.; Lo, K.; Leavitt, B.R.; Roos, R.; Durr, A.; Mead, S.; Holmans, P.; Jones, L.; et al. Identification of genetic variants associated with Huntington’s disease progression: A genome-wide association study. Lancet Neurol. 2017, 16, 701–711. [Google Scholar] [CrossRef]
- Travessa, A.M.; Rodrigues, F.B.; Mestre, T.A.; Ferreira, J.J. Fifteen Years of Clinical Trials in Huntington’s Disease: A Very Low Clinical Drug Development Success Rate. J. Huntingtons Dis. 2017, 6, 157–163. [Google Scholar] [CrossRef]
- Kordasiewicz, H.B.; Stanek, L.M.; Wancewicz, E.V.; Mazur, C.; McAlonis, M.M.; Pytel, K.A.; Artates, J.W.; Weiss, A.; Cheng, S.H.; Shihabuddin, L.S.; et al. Sustained Therapeutic Reversal of Huntington’s Disease by Transient Repression of Huntingtin Synthesis. Neuron 2012, 74, 1031–1044. [Google Scholar] [CrossRef]
- Agustín-Pavón, C.; Mielcarek, M.; Garriga-Canut, M.; Isalan, M. Deimmunization for gene therapy: Host matching of synthetic zinc finger constructs enables long-term mutant Huntingtin repression in mice. Mol. Neurodegener. 2016, 11, 64. [Google Scholar] [CrossRef]
- Yang, L.; Calingasan, N.Y.; Wille, E.J.; Cormier, K.; Smith, K.; Ferrante, R.J.; Beal, M.F. Combination therapy with Coenzyme Q10 and creatine produces additive neuroprotective effects in models of Parkinson’s and Huntington’s Diseases. J. Neurochem. 2009, 109, 1427–1439. [Google Scholar] [CrossRef]
- McGarry, A.; Auinger, P.; Kieburtz, K.D.; Bredlau, A.-L.; Hersch, S.M.; Rosas, H.D. Suicidality Risk Factors Across the CARE-HD, 2CARE, and CREST-E Clinical Trials in Huntington Disease. Neurol. Clin. Pract. 2022, 12, 131–138. [Google Scholar] [CrossRef]
- Basavarajappa, B.S.; Subbanna, S. Histone Methylation Regulation in Neurodegenerative Disorders. Int. J. Mol. Sci. 2021, 22, 4654. [Google Scholar] [CrossRef] [PubMed]
- Masrori, P.; Van Damme, P. Amyotrophic lateral sclerosis: A clinical review. Eur. J. Neurol. 2020, 27, 1918–1929. [Google Scholar] [CrossRef] [PubMed]
- Renton, A.E.; Majounie, E.; Waite, A.; Simon-Saánchez, J.; Rollinson, S.; Gibbs, J.R.; Schymick, J.C.; Laaksovirta, H.; van Swieten, J.C.; Myllykangas, L.; et al. A Hexanucleotide Repeat Expansion in C9ORF72 Is the Cause of Chromosome 9p21-Linked ALS-FTD. Neuron 2011, 72, 257–268. [Google Scholar] [CrossRef] [PubMed]
- Woollacott, I.O.C.; Mead, S. The C9ORF72 expansion mutation: Gene structure, phenotypic and diagnostic issues. Acta Neuropathol. 2014, 127, 319–332. [Google Scholar] [CrossRef] [PubMed]
- Balendra, R.; Isaacs, A.M. C9orf72-mediated ALS and FTD: Multiple pathways to disease. Nat. Rev. Neurol. 2018, 14, 544–558. [Google Scholar] [CrossRef] [PubMed]
- Herdewyn, S.; Zhao, H.; Moisse, M.; Race, V.; Matthijs, G.; Reumers, J.; Kusters, B.; Schelhaas, H.J.; Berg, L.H.v.D.; Goris, A.; et al. Whole-genome sequencing reveals a coding non-pathogenic variant tagging a non-coding pathogenic hexanucleotide repeat expansion in C9orf72 as cause of amyotrophic lateral sclerosis. Hum. Mol. Genet. 2012, 21, 2412–2419. [Google Scholar] [CrossRef]
- Majounie, E.; Renton, A.E.; Mok, K.; Dopper, E.G.; Waite, A.; Rollinson, S.; Chiò, A.; Restagno, G.; Nicolaou, N.; Simon-Sanchez, J.; et al. Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: A cross-sectional study. Lancet Neurol. 2012, 11, 323–330. [Google Scholar] [CrossRef]
- Xi, Z.; Zinman, L.; Grinberg, Y.; Moreno, D.; Sato, C.; Bilbao, J.M.; Ghani, M.; Hernández, I.; Ruiz, A.; Boada, M.; et al. Investigation of C9orf72 in 4 Neurodegenerative Disorders. Arch. Neurol. 2012, 69, 1583–1590. [Google Scholar] [CrossRef] [PubMed]
- Česnik, A.B.; Darovic, S.; Mihevc, S.P.; Štalekar, M.; Malnar, M.; Motaln, H.; Lee, Y.-B.; Mazej, J.; Pohleven, J.; Grosch, M.; et al. Nuclear RNA foci from C9ORF72 expansion mutation form paraspeckle-like bodies. J. Cell Sci. 2019, 132, jcs.224303. [Google Scholar] [CrossRef]
- Saccon, R.A.; Bunton-Stasyshyn, R.K.A.; Fisher, E.M.; Fratta, P. Is SOD1 loss of function involved in amyotrophic lateral sclerosis? Brain 2013, 136, 2342–2358. [Google Scholar] [CrossRef]
- Bosco, D.A.; Morfini, G.; Karabacak, N.M.; Song, Y.; Gros-Louis, F.; Pasinelli, P.; Goolsby, H.; Fontaine, B.A.; Lemay, N.; McKenna-Yasek, D.; et al. Wild-type and mutant SOD1 share an aberrant conformation and a common pathogenic pathway in ALS. Nat. Neurosci. 2010, 13, 1396–1403. [Google Scholar] [CrossRef]
- Bruijn, L.I.; Houseweart, M.K.; Kato, S.; Anderson, K.L.; Anderson, S.D.; Ohama, E.; Reaume, A.G.; Scott, R.W.; Cleveland, D.W. Aggregation and Motor Neuron Toxicity of an ALS-Linked SOD1 Mutant Independent from Wild-Type SOD1. Science 1998, 281, 1851–1854. [Google Scholar] [CrossRef]
- Saeed, M.; Yang, Y.; Deng, H.-X.; Hung, W.-Y.; Siddique, N.; Dellefave, L.; Gellera, C.; Andersen, P.M.; Siddique, T. Age and founder effect of SOD1 A4V mutation causing ALS. Neurology 2009, 72, 1634–1639. [Google Scholar] [CrossRef]
- Hensley, K.; Mhatre, M.; Mou, S.; Pye, Q.N.; Stewart, C.; West, M.; Williamson, K.S. On the Relation of Oxidative Stress to Neuroinflammation: Lessons Learned from the G93A-SOD1 Mouse Model of Amyotrophic Lateral Sclerosis. Antioxid. Redox Signal. 2006, 8, 2075–2087. [Google Scholar] [CrossRef] [PubMed]
- Curti, D.; Rognoni, F.; Alimonti, D.; Malaspina, A.; Feletti, F.; Tessera, S.; Finotti, N.; Rehak, L.; Mazzini, L.; Zerbi, F.; et al. SOD1 activity and protective factors in familial ALS patients with L84F SOD1 mutation. Lateral Scler. Other Motor Neuron Disord. 2002, 3, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Ceroni, M.; Malaspina, A.; Poloni, T.E.; Alimonti, D.; Rognoni, F.; Habgood, J.; Imbesi, F.; Antonelli, P.; Alfonsi, E.; Curti, D.; et al. Clustering of ALS patients in central Italy due to the occurrence of the L84F SOD1 gene mutation. Neurology 1999, 53, 1064. [Google Scholar] [CrossRef] [PubMed]
- Juneja, T.; Pericak-Vance, M.A.; Laing, N.G.; Dave, S.; Siddique, T. Prognosis in Familial Amyotrophic Lateral Sclerosis: Progression and Survival in Patients with Glu100gly and Ala4val Mutations in Cu,Zn Superoxide Dismutase. Neurology 1997, 48, 55–57. [Google Scholar] [CrossRef]
- Zhou, Y.; Tang, J.; Lan, J.; Zhang, Y.; Wang, H.; Chen, Q.; Kang, Y.; Sun, Y.; Feng, X.; Wu, L.; et al. Honokiol alleviated neurodegeneration by reducing oxidative stress and improving mitochondrial function in mutant SOD1 cellular and mouse models of amyotrophic lateral sclerosis. Acta Pharm. Sin. B 2023, 13, 577–597. [Google Scholar] [CrossRef]
- Cassina, P.; Cassina, A.; Pehar, M.; Castellanos, R.; Gandelman, M.; de León, A.; Robinson, K.M.; Mason, R.P.; Beckman, J.S.; Barbeito, L.; et al. Mitochondrial Dysfunction in SOD1G93A-Bearing Astrocytes Promotes Motor Neuron Degeneration: Prevention by Mitochondrial-Targeted Antioxidants. J. Neurosci. 2008, 28, 4115–4122. [Google Scholar] [CrossRef]
- Zhang, X.; Li, L.; Chen, S.; Yang, D.; Wang, Y.; Zhang, X.; Wang, Z.; Le, W. Rapamycin treatment augments motor neuron degeneration in SOD1G93Amouse model of amyotrophic lateral sclerosis. Autophagy 2011, 7, 412–425. [Google Scholar] [CrossRef]
- Daoud, H.; Valdmanis, P.N.; Kabashi, E.; Dion, P.; Dupre, N.; Camu, W.; Meininger, V.; Rouleau, G.A. Contribution of TARDBP mutations to sporadic amyotrophic lateral sclerosis. J. Med. Genet. 2008, 46, 112–114. [Google Scholar] [CrossRef]
- Lattante, S.; Rouleau, G.A.; Kabashi, E. TARDBP and FUS Mutations Associated with Amyotrophic Lateral Sclerosis: Summary and Update. Hum. Mutat. 2013, 34, 812–826. [Google Scholar] [CrossRef]
- Kabashi, E.; Valdmanis, P.; Dion, P.; Spiegelman, D.; McConkey, B.J.; Velde, C.V.; Bouchard, J.-P.; Lacomblez, L.; Pochigaeva, K.; Salachas, F.; et al. TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat. Genet. 2008, 40, 572–574. [Google Scholar] [CrossRef]
- Corcia, P.; Valdmanis, P.; Millecamps, S.; Lionnet, C.; Blasco, H.; Mouzat, K.; Daoud, H.; Belzil, V.; Morales, R.; Pageot, N.; et al. Phenotype and genotype analysis in amyotrophic lateral sclerosis with TARDBP gene mutations. Neurology 2012, 78, 1519–1526. [Google Scholar] [CrossRef] [PubMed]
- Tamaoka, A.; Arai, M.; Itokawa, M.; Arai, T.; Hasegawa, M.; Tsuchiya, K.; Takuma, H.; Tsuji, H.; Ishii, A.; Watanabe, M.; et al. TDP-43 M337V Mutation in Familial Amyotrophic Lateral Sclerosis in Japan. Intern. Med. 2010, 49, 331–334. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.; Oiwa, K.; Murata, Y.; Komine, O.; Sobue, A.; Endo, F.; Takahashi, E.; Yamanaka, K. ALS-linked TDP-43M337V knock-in mice exhibit splicing deregulation without neurodegeneration. Mol. Brain 2020, 13, 8. [Google Scholar] [CrossRef] [PubMed]
- Esmaeili, M.A.; Panahi, M.; Yadav, S.; Hennings, L.; Kiaei, M. Premature death of TDP-43 (A315T) transgenic mice due to gastrointestinal complications prior to development of full neurological symptoms of amyotrophic lateral sclerosis. Int. J. Exp. Pathol. 2013, 94, 56–64. [Google Scholar] [CrossRef] [PubMed]
- Orrù, S.; Coni, P.; Floris, A.; Littera, R.; Carcassi, C.; Sogos, V.; Brancia, C. Reduced stress granule formation and cell death in fibroblasts with the A382T mutation of TARDBP gene: Evidence for loss of TDP-43 nuclear function. Hum. Mol. Genet. 2016, 25, 4473–4483. [Google Scholar] [CrossRef]
- Zanini, G.; Selleri, V.; Nasi, M.; De Gaetano, A.; Martinelli, I.; Gianferrari, G.; Lofaro, F.D.; Boraldi, F.; Mandrioli, J.; Pinti, M. Mitochondrial and Endoplasmic Reticulum Alterations in a Case of Amyotrophic Lateral Sclerosis Caused by TDP-43 A382T Mutation. Int. J. Mol. Sci. 2022, 23, 11881. [Google Scholar] [CrossRef]
- Deng, H.; Gao, K.; Jankovic, J. The role of FUS gene variants in neurodegenerative diseases. Nat. Rev. Neurol. 2014, 10, 337–348. [Google Scholar] [CrossRef]
- Corrado, L.; Del Bo, R.; Castellotti, B.; Ratti, A.; Cereda, C.; Penco, S.; Sorarù, G.; Carlomagno, Y.; Ghezzi, S.; Pensato, V.; et al. Mutations of FUS gene in sporadic amyotrophic lateral sclerosis. J. Med. Genet. 2010, 47, 190–194. [Google Scholar] [CrossRef]
- Rademakers, R.; Stewart, H.; Dejesus-Hernandez, M.; Krieger, C.; Graff-Radford, N.; Fabros, M.; Briemberg, H.; Cashman, N.; Eisen, A.; Mackenzie, I.R. Fus gene mutations in familial and sporadic amyotrophic lateral sclerosis. Muscle Nerve 2010, 42, 170–176. [Google Scholar] [CrossRef]
- Ticozzi, N.; Silani, V.; LeClerc, A.L.; Keagle, P.; Gellera, C.; Ratti, A.; Taroni, F.; Kwiatkowski, T.J.; McKenna-Yasek, D.M.; Sapp, P.C.; et al. Analysis of FUS gene mutation in familial amyotrophic lateral sclerosis within an Italian cohort. Neurology 2009, 73, 1180–1185. [Google Scholar] [CrossRef]
- Weinreich, M.; Shepheard, S.R.; Verber, N.; Wyles, M.; Heath, P.R.; Highley, J.R.; Kirby, J.; Shaw, P.J. Neuropathological characterization of a novel TANK binding kinase (TBK1) gene loss of function mutation associated with amyotrophic lateral sclerosis. Neuropathol. Appl. Neurobiol. 2019, 46, 279–291. [Google Scholar] [CrossRef]
- Oakes, J.A.; Davies, M.C.; Collins, M.O. TBK1: A new player in ALS linking autophagy and neuroinflammation. Mol. Brain 2017, 10, 5. [Google Scholar] [CrossRef]
- Gurfinkel, Y.; Polain, N.; Sonar, K.; Nice, P.; Mancera, R.L.; Rea, S.L. Functional and structural consequences of TBK1 missense variants in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Neurobiol. Dis. 2022, 174, 105859. [Google Scholar] [CrossRef]
- Zarei, S.; Carr, K.; Reiley, L.; Diaz, K.; Guerra, O.; Altamirano, P.; Pagani, W.; Lodin, D.; Orozco, G.; Chinea, A. A comprehensive review of amyotrophic lateral sclerosis. Surg. Neurol. Int. 2015, 6, 171. [Google Scholar] [CrossRef]
- Deans, C.; Maggert, K.A. What Do You Mean, “Epigenetic”? Genetics 2015, 199, 887–896. [Google Scholar] [CrossRef] [PubMed]
- Burggren, W. Epigenetic Inheritance and Its Role in Evolutionary Biology: Re-Evaluation and New Perspectives. Biology 2016, 5, 24. [Google Scholar] [CrossRef]
- Neumann, M.; Sampathu, D.M.; Kwong, L.K.; Truax, A.C.; Micsenyi, M.C.; Chou, T.T.; Bruce, J.; Schuck, T.; Grossman, M.; Clark, C.M.; et al. Ubiquitinated TDP-43 in Frontotemporal Lobar Degeneration and Amyotrophic Lateral Sclerosis. Science 2006, 314, 130–133. [Google Scholar] [CrossRef] [PubMed]
- Fecto, F.; Yan, J.; Vemula, S.P.; Liu, E.; Yang, Y.; Chen, W.; Zheng, J.G.; Shi, Y.; Siddique, N.; Arrat, H.; et al. SQSTM1 Mutations in Familial and Sporadic Amyotrophic Lateral Sclerosis. Arch. Neurol. 2011, 68, 1440–1446. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, H.; Morino, H.; Ito, H.; Izumi, Y.; Kato, H.; Watanabe, Y.; Kinoshita, Y.; Kamada, M.; Nodera, H.; Suzuki, H.; et al. Mutations of optineurin in amyotrophic lateral sclerosis. Nature 2010, 465, 223–226. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.O.; Mandrioli, J.; Benatar, M.; Abramzon, Y.; Van Deerlin, V.M.; Trojanowski, J.Q.; Gibbs, J.R.; Brunetti, M.; Gronka, S.; Wuu, J.; et al. Exome Sequencing Reveals VCP Mutations as a Cause of Familial ALS. Neuron 2010, 68, 857–864. [Google Scholar] [CrossRef]
- Boeynaems, S.; Bogaert, E.; Van Damme, P.; Bosch, L.V.D. Inside out: The role of nucleocytoplasmic transport in ALS and FTLD. Acta Neuropathol. 2016, 132, 159–173. [Google Scholar] [CrossRef]
- Vilarino-Guell, C.; Wider, C.; Soto-Ortolaza, A.I.; Cobb, S.A.; Kachergus, J.M.; Keeling, B.H.; Dachsel, J.C.; Hulihan, M.M.; Dickson, D.W.; Wszolek, Z.K.; et al. Characterization of DCTN1 genetic variability in neurodegeneration. Neurology 2009, 72, 2024–2028. [Google Scholar] [CrossRef] [PubMed]
- Araki, E.; Tsuboi, Y.; Daechsel, J.; Milnerwood, A.; Vilarino-Guell, C.; Fujii, N.; Mishima, T.; Oka, T.; Hara, H.; Fukae, J.; et al. A Novel DCTN1 mutation with late-onset parkinsonism and frontotemporal atrophy. Mov. Disord. 2014, 29, 1201–1204. [Google Scholar] [CrossRef] [PubMed]
- Al-Chalabi, A.; Hardiman, O.; Kiernan, M.C.; Chiò, A.; Rix-Brooks, B.; van den Berg, L.H. Amyotrophic lateral sclerosis: Moving towards a new classification system. Lancet Neurol. 2016, 15, 1182–1194. [Google Scholar] [CrossRef] [PubMed]
- Chiò, A.; Calvo, A.; Moglia, C.; Mazzini, L.; Mora, G.; PARALS Study Group. Phenotypic heterogeneity of amyotrophic lateral sclerosis: A population based study. J. Neurol. Neurosurg. Psychiatry 2011, 82, 740–746. [Google Scholar] [CrossRef] [PubMed]
- Finegan, E.; Chipika, R.H.; Shing, S.L.H.; Hardiman, O.; Bede, P. Pathological Crying and Laughing in Motor Neuron Disease: Pathobiology, Screening, Intervention. Front. Neurol. 2019, 10, 260. [Google Scholar] [CrossRef]
- Wijesekera, L.C.; Mathers, S.; Talman, P.; Galtrey, C.; Parkinson, M.H.; Ganesalingam, J.; Willey, E.; Ampong, M.A.; Ellis, C.M.; Shaw, C.E.; et al. Natural history and clinical features of the flail arm and flail leg ALS variants. Neurology 2009, 72, 1087–1094. [Google Scholar] [CrossRef]
- Doble, A. The pharmacology and mechanism of action of riluzole. Neurology 1996, 47 (Suppl. S4), 233S–241S. [Google Scholar] [CrossRef]
- Lacomblez, L.; Amyotrophic Lateral Sclerosis/Riluzole Study Group II; Bensimon, G.; Meininger, V.; Leigh, P.; Guillet, P. Dose-ranging study of riluzole in amyotrophic lateral sclerosis. Amyotrophic Lateral Sclerosis/Riluzole Study Group II. Lancet 1996, 347, 1425–1431. [Google Scholar] [CrossRef]
- Hinchcliffe, M.; Smith, A. Riluzole: Real-world evidence supports significant extension of median survival times in patients with amyotrophic lateral sclerosis. Degener. Neurol. Neuromuscul. Dis. 2017, 7, 61–70. [Google Scholar] [CrossRef]
- Mora, J.S.; Genge, A.; Chio, A.; Estol, C.J.; Chaverri, D.; Hernández, M.; Marín, S.; Mascias, J.; Rodriguez, G.E.; Povedano, M.; et al. Masitinib as an add-on therapy to riluzole in patients with amyotrophic lateral sclerosis: A randomized clinical trial. Amyotroph. Lateral Scler. Front. Degener. 2019, 21, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Scheltens, P.; De Strooper, B.; Kivipelto, M.; Holstege, H.; Chételat, G.; Teunissen, C.E.; Cummings, J.; van der Flier, W.M. Alzheimer’s disease. Lancet 2021, 397, 1577–1590. [Google Scholar] [CrossRef] [PubMed]
- Bloem, B.R.; Henderson, E.J.; Dorsey, E.R.; Okun, M.S.; Okubadejo, N.; Chan, P.; Andrejack, J.; Darweesh, S.K.L.; Munneke, M. Integrated and patient-centred management of Parkinson’s disease: A network model for reshaping chronic neurological care. Lancet Neurol. 2020, 19, 623–634. [Google Scholar] [CrossRef]
- Tabrizi, S.J.; Flower, M.; Ross, C.A.; Wild, E.J. Huntington disease: New insights into molecular pathogenesis and therapeutic opportunities. Nat. Rev. Neurol. 2020, 16, 529–546. [Google Scholar] [CrossRef] [PubMed]
- Deda, H.; Inci, M.; Kürekçi, A.; Sav, A.; Kayıhan, K.; Özgün, E.; Üstünsoy, G.; Kocabay, S. Treatment of amyotrophic lateral sclerosis patients by autologous bone marrow-derived hematopoietic stem cell transplantation: A 1-year follow-up. Cytotherapy 2009, 11, 18–25. [Google Scholar] [CrossRef]
- Moviglia, G.A.; Moviglia-Brandolino, M.T.; Varela, G.S.; Albanese, G.; Piccone, S.; Echegaray, G.; Martinez, G.; Blasseti, N.; Farias, J.; Farina, P.; et al. Feasibility, Safety, and Preliminary Proof of Principles of Autologous Neural Stem Cell Treatment Combined with T-Cell Vaccination for ALS Patients. Cell Transplant. 2012, 21, 57–63. [Google Scholar] [CrossRef]


| Mutation | Pathogenicity | Type of Mutation | Biological Effect | Citation |
|---|---|---|---|---|
| A673T (Icelandic) | Alzheimer’s Disease—Protective | Substitution | This particular type is linked to limited build-up of amyloid and is believed to guard against amyloid-related issues. It results in a decrease of approximately 40 percent in the production of amyloidogenic Aβ peptides, and the Aβ that is produced has a reduced tendency to form clumps. | [17,18,19,20] |
| A673V | Not Classified | Substitution | According to the CERAD criteria, a clear diagnosis of AD was made, as evidenced by substantial Aβ and tau pathology deposits (Braak stage VI) along with cerebral amyloid angiopathy. The deposits found contained elevated levels of Aβ40 and were notably larger, with fewer preamyloid deposits. Perivascular localization was frequently observed. In laboratory studies, it was discovered that A673V caused a shift in β-secretase processing of APP toward the amyloidogenic pathway and amplified Aβ aggregation. | [21,22] |
| E693Q (Dutch) | Hereditary Cerebral Hemorrhage with Amyloidosis—Pathogenic | Substitution | There is a substantial accumulation of amyloid in the cerebral blood vessels, accompanied by hemorrhages and some diffuse plaques in the brain tissue. In laboratory experiments, it was observed that this condition speeds up Aβ aggregation in vitro, leading to greater fibril formation, and may also modify APP processing. | [24,25] |
| E693del (Osaka, E693∆, E693delta | Alzheimer’s Disease—Pathogenic | Deletion | This variant led to an increased oligomerization and nucleation of Aβ aggregates in vitro. Furthermore, it was found that there was no alteration in the Aβ42/Aβ40 ratio, but there was a decrease in both Aβ42 and Aβ40. This variant was also discovered to be more resistant to degradation by neprilysin and insulin-degrading enzyme. Additionally, this variant had a greater inhibitory effect on long-term potentiation (LTP) compared to wild-type Aβ, which suggests a potential negative impact on synaptic plasticity. | [26,27,28,29] |
| E693K (Italian) | Hereditary Cerebral Hemorrhage with Amyloidosis—Pathogenic | Substitution | The observed symptoms include small to large hematomas, subarachnoid bleeding, scars with hemosiderin deposits, small infarcts, and cortical calcifications. Aβ immunoreactivity was observed in vessel walls and neuropil, but there was an absence of neurofibrillary changes and neuritic plaques. Despite a reduction in the Aβ42/Aβ40 ratio and a decrease in Aβ42 levels, the mutant peptide was found to be toxic in cells and aggregates at a faster rate. | [23] |
| E693G (Arctic, E22G) | Alzheimer’s Disease—Pathogenic | Substitution | Several carriers displayed neuropathology that was indicative of AD. Plaques were observed to have a “targetoid” shape, containing heterogeneous truncated Aβ peptides in the center and surrounded by Aβ42. Cell-based assays revealed a reduction in the production of both Aβ40 and Aβ42. Additionally, there was a decrease in proteolytic degradation of Aβ by neprilysin, a type of enzyme that breaks down proteins. | [23,25] |
| c.-488C>A (rs532314089) | Alzheimer’s Disease | Substitution | Predicted to disrupt binding of transcription factor EGR1. PHRED-scaled CADD = 0.26. Negative regulator in multiple cell types including PC12 neuronal-like rat chromaffin cells, SK-N-SH neuroblastoma cells, C6 glial cells and U373 astroctyoma cells among others | [30,31,32] |
| c.24+38G>A (rs373985746) | Alzheimer’s Disease | Substitution | Predicted benign in silico (PHRED-scaled CADD = 10). | [32] |
| c.24+288G>A (rs192348494) | Alzheimer’s Disease | Substitution | Predicted benign in silico (PHRED-scaled CADD = 12). | [32] |
| c.-23-377A>G (rs150375400) | Alzheimer’s Disease | Substitution | Predicted benign in silico (PHRED-scaled CADD = 10). | [32] |
| A18T | Alzheimer’s Disease, Cardiovascular Disease | Substitution | Predicted to disrupt signal peptide cleavage and affect APOE secretion. PHRED-scaled CADD = 22. | [32,33] |
| Mutation | Pathogenicity | Type of Mutation | Biological Effect | Citation |
|---|---|---|---|---|
| A79V | Alzheimer’s Disease —Pathogenic | Substitution | The observed neuropathology was in line with that of AD. It was observed that this variant led to an increase in the Aβ42/Aβ40 ratio and a decrease in the Aβ37/Aβ42 ratios in cells. | [39,40,41] |
| M84V | Alzheimer’s Disease —Pathogenic | Substitution | In two cases, the observed neuropathology was consistent with AD. Additionally, MRI scans revealed cortical and cerebellar atrophy in these two cases. In the third case, frontal and temporal lobe atrophy was observed. Cell studies revealed an increase in both Aβ42 and the Aβ42/Aβ40 ratio. | [42,43] |
| L85P | Alzheimer’s Disease —Pathogenic | Substitution | SPECT and PET scans showed bilateral hypoperfusion and hypometabolism in the occipital and temporal lobes. Cell studies revealed an increase in the Aβ42/Aβ40 ratio as well as increased Aβ42 levels in transfected cells. In vitro studies indicated a decrease in Aβ42 production and the complete absence of Aβ40 production. | [44,45] |
| L113_I114insT (int4del) | Alzheimer’s Disease —Pathogenic | Substitution | The observed neuropathology was consistent with AD, and included neuron loss in the hippocampus and entorhinal cortex, the presence of neuritic plaques and neurofibrillary tangles in the hippocampus, and amyloid angiopathy, which was particularly evident in the cerebellum. The identified mutation involved a deletion of a G in the splice donor site of intron 4, resulting in the production of three aberrant transcripts. Further investigations indicated an increase in both Aβ42 and the Aβ42/Aβ40 ratio, as well as a reduction in Aβ40 and Aβ38 production in patient brain membranes. | [46,47,48] |
| M139V | Alzheimer’s Disease —Pathogenic | Substitution | Decrease in the levels of Aβ40, Aβ38, and Aβ37, and an increase in the levels of Aβ42 and Aβ43. In iPSC-derived neurons, the levels of mutant protein were found to be variable, suggesting protein instability. | [49,50,51] |
| Mutation | Pathogenicity | Type of Mutation | Biological Effect | Citation |
|---|---|---|---|---|
| K82fs | Tauopathy and Pick’s Disease | Deletion | The neuropathological findings were consistent with Pick’s disease. A frameshift was identified to start at K82, and the mutant protein was found to be reduced in the frontal cortex and hippocampus. | [54] |
| c.*71C>A | Alzheimer’s Disease —Pathogenic | Substitution | In one case, an MRI scan revealed widening of the sulcus, fissure, and temporal horn, along with a decrease in hippocampal volume. Additionally, FDG-PET showed hypometabolism in the bilateral frontal, parietal, and temporal lobes. Among the five affected carriers, CSF analysis showed Aβ42, total tau, and phospho-tau levels consistent with AD. The study suggests a possible reduction in the binding of PSEN2 expression suppressor miR-183-5p, which may lead to an increased Aβ42/Aβ40 ratio. | [55,56] |
| M239V | Alzheimer’s Disease —Pathogenic | Substitution | The brain pathology showed diffuse cerebral atrophy, senile plaques, neurofibrillary tangles (Braak and Braak stage VI), ectopic neurons in subcortical white matter, and extracellular “ghost” neurofibrillary tangles. In cell-based assays, there was an increase in the Aβ42/Aβ40 ratio and an increase in Aβ42 levels. However, there was no change in the proteolytic products PSEN2-CTF and PSEN2-NTF. | [57,58] |
| Mutation | Pathogenicity | Type of Mutation | Biological Effect | Citation |
|---|---|---|---|---|
| IVS10+12 C>T | Familial Danish Dementia—Pathogenic | Substitution | The mutant protein leads to the formation of tau aggregates in both neurons and glia, and isolated tau filaments exhibit a twisted, ribbon-like morphology and consist of hyperphosphorylated 4-repeat (4R) tau isoforms. The mutation also causes a destabilization of a stem-loop structure that regulates the alternative splicing of exon 10, resulting in a higher frequency of inclusion of exon 10 and an increased proportion of 4R tau isoforms. | [84,85] |
| A152T | Alzheimer’s Disease—Risk | Substitution | The presence of tau pathology is a common feature, often accompanied by Lewy bodies, amyloid plaques, or TDP-43 pathology. The mutant tau has a decreased ability to bind to microtubules, leading to less efficient microtubule assembly and impaired microtubule stability. Additionally, it has an increased propensity to form tau oligomers and is more susceptible to proteolysis by caspases. | [86,87,88] |
| K257T | Tauopathy and Frontotemporal —Pathogenic | Substitution | The patient exhibited frontotemporal atrophy with significant temporal lobe involvement. Tau-positive Pick bodies were found in the neocortex, hippocampus, and subcortical regions similar to those seen in sporadic Pick’s disease. Some cell bodies showed diffuse hyperphosphorylated tau. In vitro analysis showed that recombinant tau protein with the K257T mutation had a decreased ability to promote microtubule assembly. | [89] |
| L266V | Frontotemporal —Pathogenic | Substitution | The patient had severe atrophy of the frontal and temporal lobes, with extensive neuronal loss and gliosis. Tau-positive inclusions, including Pick bodies, and tau-positive argyrophilic astrocytes with stout filaments and round or irregular argyrophilic inclusions were also observed. In molecular studies, there were increased levels of exon 10+ tau mRNA and soluble four-repeat (4R) tau. The patient showed a decreased rate and extent of tau-induced microtubule assembly, as well as a specific increase in tau self-assembly for the 3R isoform. | [90,91] |
| Mutation | Pathogenicity | Type of Mutation | Biological Effect | Citation |
|---|---|---|---|---|
| A4V | ALS Pathogenic | Substitution | This mutation is responsible for a rapidly progressive dominant form of amyotrophic lateral sclerosis (ALS) that exclusively affects lower motor neurons, and it accounts for 50% of SOD1 mutations associated with familial ALS in North America. However, it is a rare mutation in Europe. | [142,146] |
| G93A | ALS Pathogenic | Substitution | Patients showing different oxidative markers, such as glutamate excitotoxicity, and dysfunctions at several levels, such as mitochondria, due to calcium influx, and axon as well as protein oxidation; modifications were observed at SOD1-G93A in mice. This mutation is relatively rare in the general population but it is very common in familial ALS, and multiple studies on animal models have also shown that having the SOD1-G93A mutation is enough to cause motor-neuron degeneration. | [147,148,149] |
| L84F | ALS Pathogenic | Substitution | Protein instability and misfolding that can lead to forming protein accumulations. | [144,145] |
| Mutation | Pathogenicity | Type of Mutation | Biological Effect | Citation |
|---|---|---|---|---|
| M337V | ALS Pathogenic | Substitution | Production of an abnormal TDP-43 protein that aggregates abnormally and accumulates in motor neurons, ultimately leading to their degeneration and death, and thus contributing to ALS development. | [150,154,155] |
| A315T | ALS Pathogenic | Substitution | Transgenic mice carrying the A315T mutation of TDP-43 may succumb to early death due to digestive complications before fully manifesting neurological signs associated with ALS, suggesting it also influences their digestive systems and may contribute to their early demise. Although the exact mechanisms underlying gastrointestinal complications remain poorly understood, experts speculate that abnormal TDP-43 protein build-up in intestinal cells may lead to dysfunction and damage. | [156] |
| A382T | Possible ALS Pathogenic | Substitution | Unknown mechanism. | [157,158] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ciurea, A.V.; Mohan, A.G.; Covache-Busuioc, R.-A.; Costin, H.-P.; Glavan, L.-A.; Corlatescu, A.-D.; Saceleanu, V.M. Unraveling Molecular and Genetic Insights into Neurodegenerative Diseases: Advances in Understanding Alzheimer’s, Parkinson’s, and Huntington’s Diseases and Amyotrophic Lateral Sclerosis. Int. J. Mol. Sci. 2023, 24, 10809. https://doi.org/10.3390/ijms241310809
Ciurea AV, Mohan AG, Covache-Busuioc R-A, Costin H-P, Glavan L-A, Corlatescu A-D, Saceleanu VM. Unraveling Molecular and Genetic Insights into Neurodegenerative Diseases: Advances in Understanding Alzheimer’s, Parkinson’s, and Huntington’s Diseases and Amyotrophic Lateral Sclerosis. International Journal of Molecular Sciences. 2023; 24(13):10809. https://doi.org/10.3390/ijms241310809
Chicago/Turabian StyleCiurea, Alexandru Vlad, Aurel George Mohan, Razvan-Adrian Covache-Busuioc, Horia-Petre Costin, Luca-Andrei Glavan, Antonio-Daniel Corlatescu, and Vicentiu Mircea Saceleanu. 2023. "Unraveling Molecular and Genetic Insights into Neurodegenerative Diseases: Advances in Understanding Alzheimer’s, Parkinson’s, and Huntington’s Diseases and Amyotrophic Lateral Sclerosis" International Journal of Molecular Sciences 24, no. 13: 10809. https://doi.org/10.3390/ijms241310809
APA StyleCiurea, A. V., Mohan, A. G., Covache-Busuioc, R.-A., Costin, H.-P., Glavan, L.-A., Corlatescu, A.-D., & Saceleanu, V. M. (2023). Unraveling Molecular and Genetic Insights into Neurodegenerative Diseases: Advances in Understanding Alzheimer’s, Parkinson’s, and Huntington’s Diseases and Amyotrophic Lateral Sclerosis. International Journal of Molecular Sciences, 24(13), 10809. https://doi.org/10.3390/ijms241310809





