Prognostic and Immunological Implications of FAM72A in Pan-Cancer and Functional Validations
Abstract
1. Introduction
2. Results
2.1. Landscape of FAM72A mRNA Expression Levels in Pan-Cancer Tissues
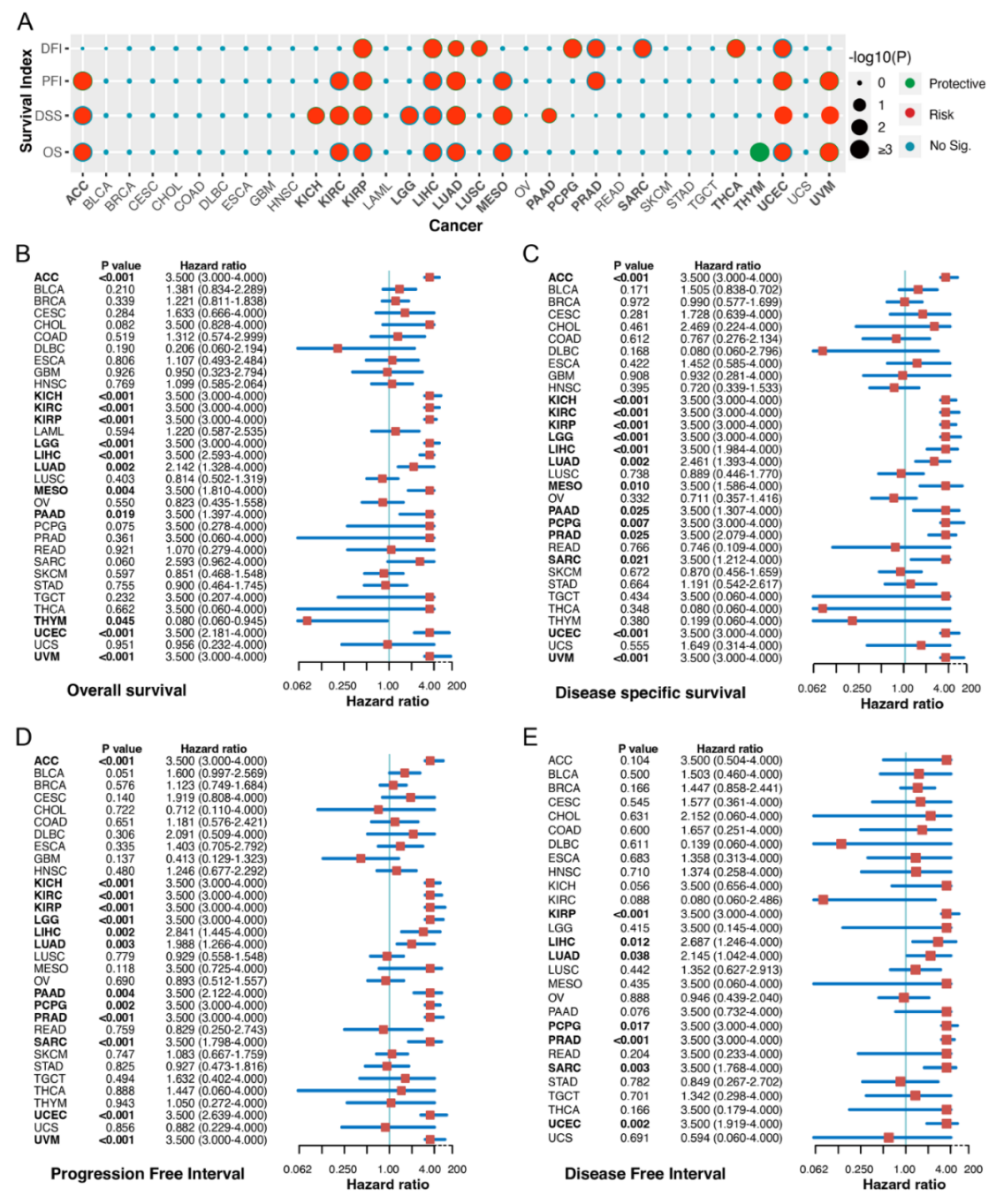
2.2. Correlation between FAM72A Expression Level and Cancer Prognosis
2.3. Diagnostic Value of FAM72A for Pan-Cancer
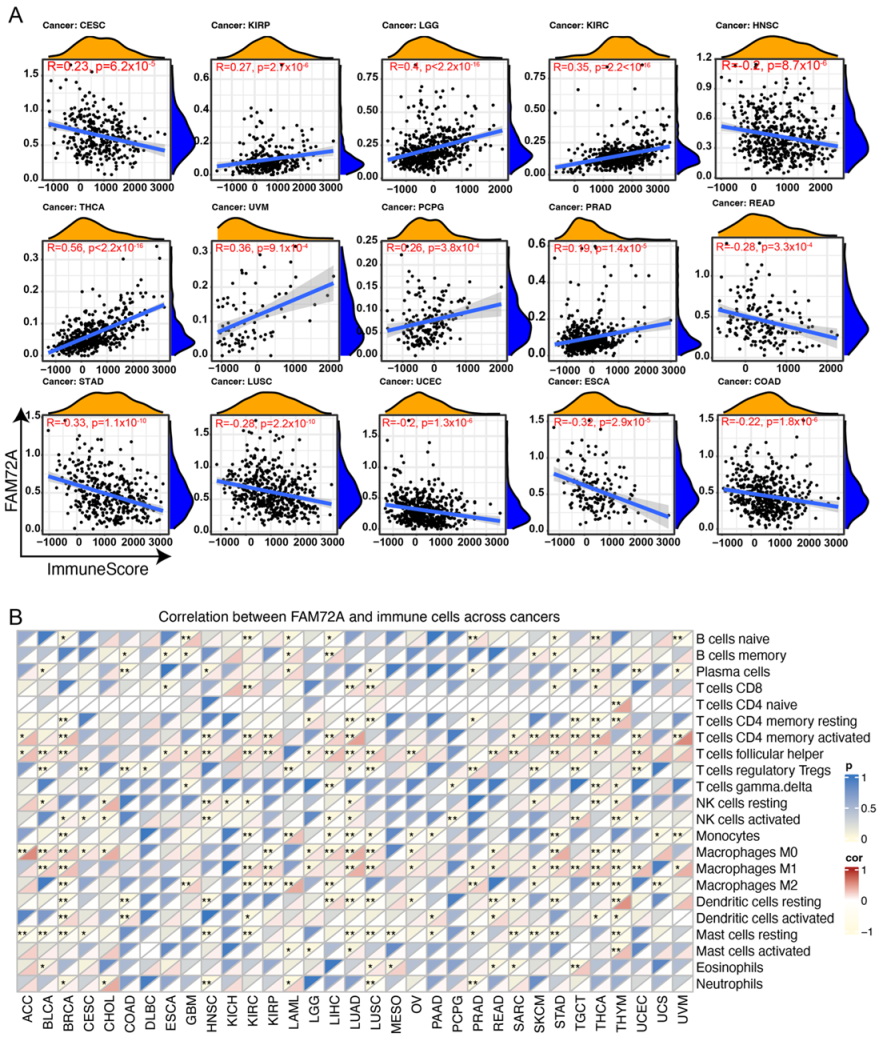
2.4. Correlation between FAM72A and Tumor Microenvironment in Different Tumors
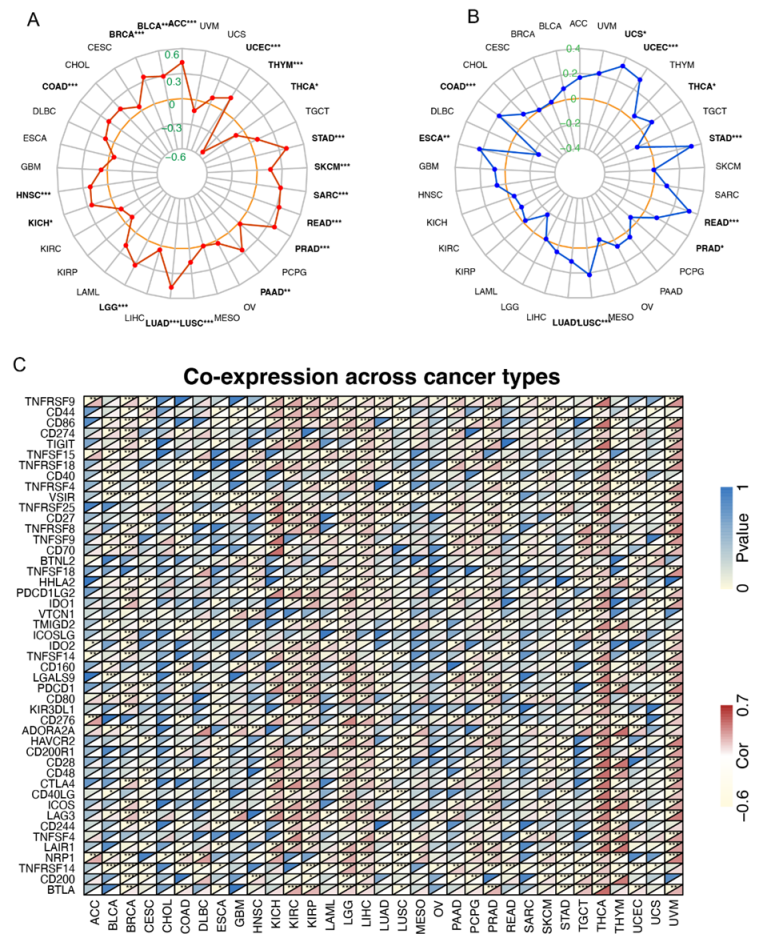
2.5. Correlations of FAM72A Expression with TMB, MSI, and Immune Checkpoint Genes in Cancers
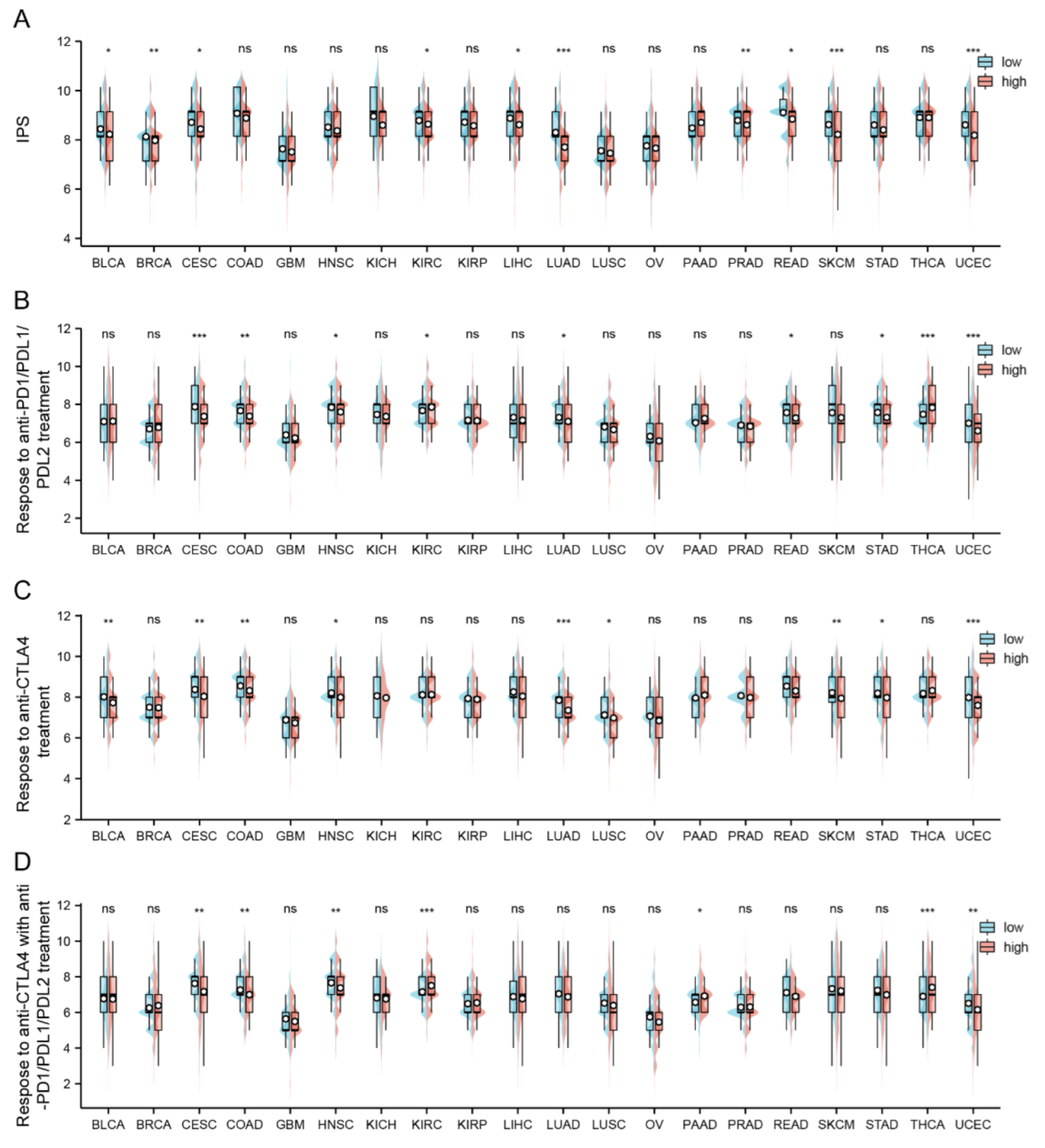
2.6. Relation between FAM72A and Immunophenoscore across Cancers
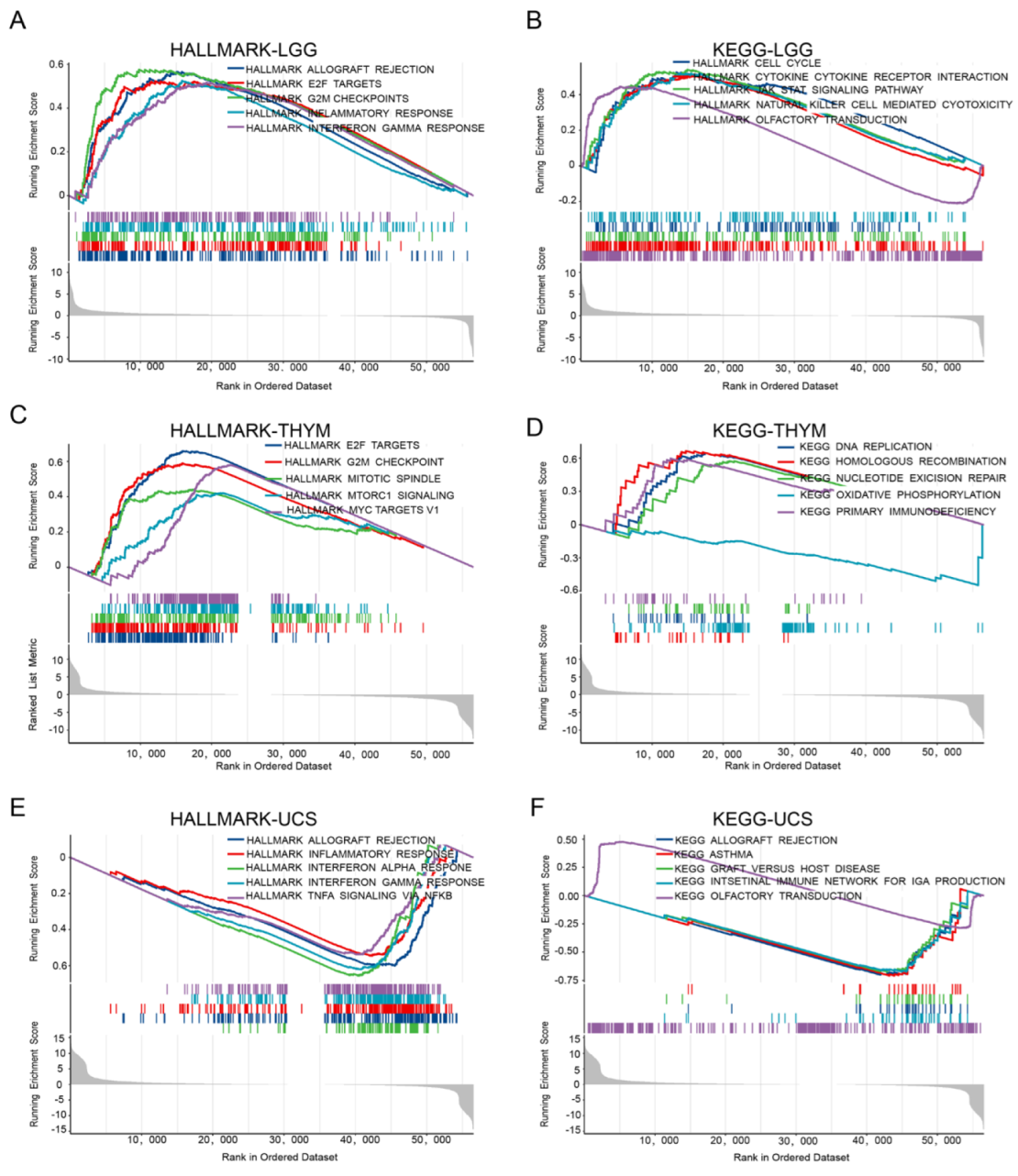
2.7. The Signaling Pathway Associated with FAM72A
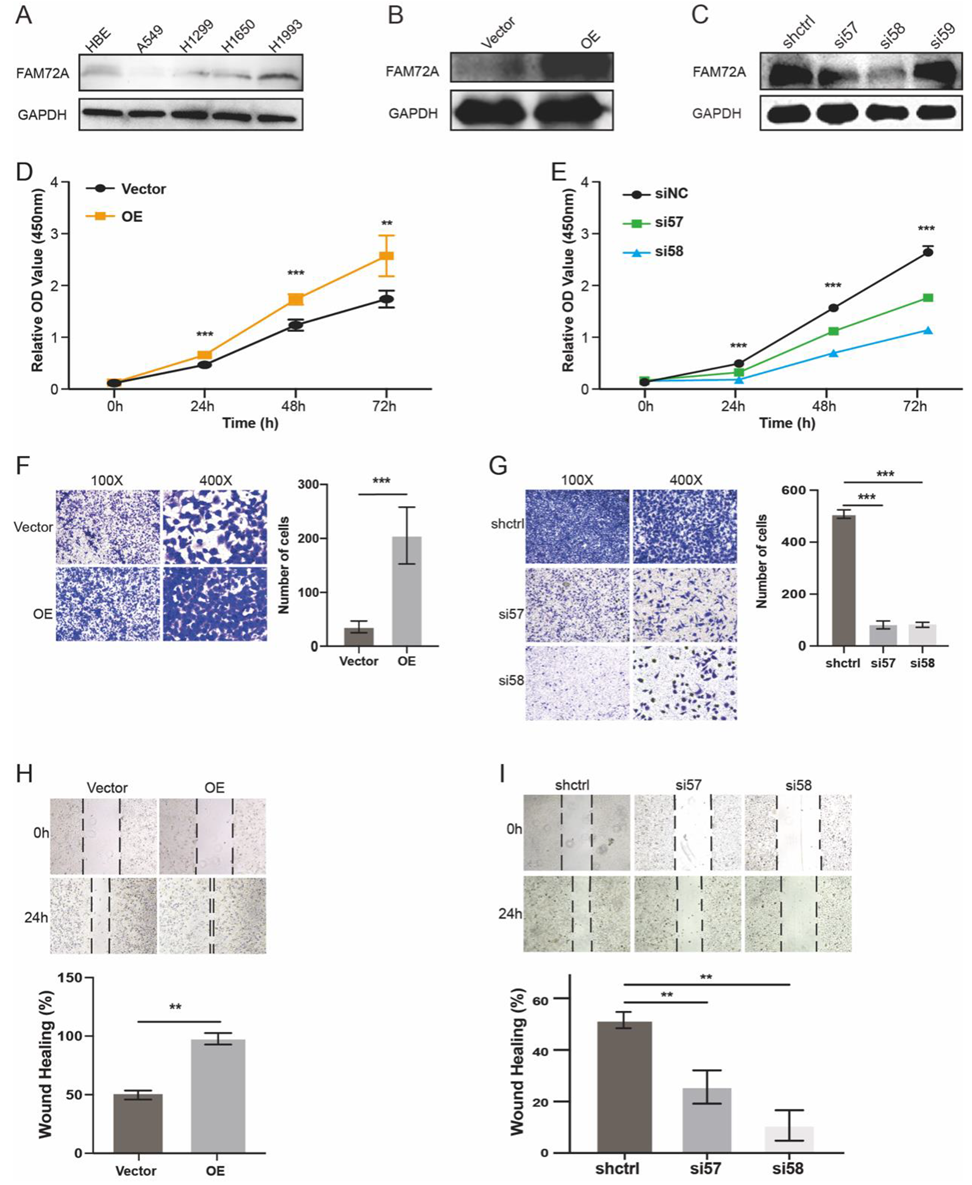
2.8. In Vitro Validation
3. Discussion
3.1. Expression and Prognostic Implication
3.2. Tumor Microenvironment
3.3. Correlation to TMB, MSI, and ICPs
3.4. Functional Analysis
4. Materials and Methods
4.1. Data Collection
4.2. Estimation of FAM72A Activity in Patient Samples
4.3. Prognostic Analysis
4.4. FAM72A’s Capacity to Distinguish Tumor from Non-Tumor Tissues
4.5. Implication of FAM72A Expression in Tumor Immune Microenvironment
4.6. Correlation Analysis of FAM72A with TMB, MSI, Checkpoint Genes, and Immunophenoscore
4.7. Gene Set Enrichment Analysis in Pan-Cancer
4.8. Cell Culture
4.9. Establishment of Stable Cell Lines
4.10. Western Blotting
4.11. Cell Proliferation
4.12. Cell Migration Assay
4.13. Wound-Healing Assay
4.14. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Guo, C.; Zhang, X.; Fink, S.P.; Platzer, P.; Wilson, K.; Willson, J.K.V.; Wang, Z.; Markowitz, S.D. Ugene, a newly identified protein that is commonly overexpressed in cancer and binds uracil DNA glycosylase. Cancer Res. 2008, 68, 6118–6126. [Google Scholar] [CrossRef] [PubMed]
- Rogier, M.; Moritz, J.; Robert, I.; Lescale, C.; Heyer, V.; Abello, A.; Martin, O.; Capitani, K.; Thomas, M.; Thomas-Claudepierre, A.S.; et al. Fam72a enforces error-prone DNA repair during antibody diversification. Nature 2021, 600, 329–333. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Li, C.; Stewart, J.A.; Barbulescu, P.; Desivo, N.S.; Alvarez-Quilon, A.; Pezo, R.C.; Perera, M.L.W.; Chan, K.E.; Tong, A.H.Y.; et al. FAM72A antagonizes UNG2 to promote mutagenic repair during antibody maturation. Nature 2021, 600, 324–328. [Google Scholar] [CrossRef] [PubMed]
- Renganathan, S.; Pramanik, S.; Ekambaram, R.; Kutzner, A.; Kim, P.S.; Heese, K. Identification of a Chemotherapeutic Lead Molecule for the Potential Disruption of the FAM72A-UNG2 Interaction to Interfere with Genome Stability, Centromere Formation, and Genome Editing. Cancers 2021, 13, 5870. [Google Scholar] [CrossRef]
- Rahane, C.S.; Kutzner, A.; Heese, K. A cancer tissue-specific FAM72 expression profile defines a novel glioblastoma multiform (GBM) gene-mutation signature. J. Neurooncol. 2019, 141, 57–70. [Google Scholar] [CrossRef]
- Yu, Y.; Wang, Z.; Zheng, Q.; Li, J. FAM72 serves as a biomarker of poor prognosis in human lung adenocarcinoma. Aging 2021, 13, 8155–8176. [Google Scholar] [CrossRef]
- Bagaev, A.; Kotlov, N.; Nomie, K.; Svekolkin, V.; Gafurov, A.; Isaeva, O.; Osokin, N.; Kozlov, I.; Frenkel, F.; Gancharova, O.; et al. Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell 2021, 39, 845–865.e7. [Google Scholar] [CrossRef]
- Charoentong, P.; Finotello, F.; Angelova, M.; Mayer, C.; Efremova, M.; Rieder, D.; Hackl, H.; Trajanoski, Z. Pan-cancer Immunogenomic Analyses Reveal Genotype-Immunophenotype Relationships and Predictors of Response to Checkpoint Blockade. Cell Rep. 2017, 18, 248–262. [Google Scholar] [CrossRef]
- Sha, D.; Jin, Z.; Budczies, J.; Kluck, K.; Stenzinger, A.; Sinicrope, F.A. Tumor Mutational Burden as a Predictive Biomarker in Solid Tumors. Cancer Discov. 2020, 10, 1808–1825. [Google Scholar] [CrossRef]
- Luchini, C.; Bibeau, F.; Ligtenberg, M.J.L.; Singh, N.; Nottegar, A.; Bosse, T.; Miller, R.; Riaz, N.; Douillard, J.-Y.; Andre, F.; et al. ESMO recommendations on microsatellite instability testing for immunotherapy in cancer, and its relationship with PD-1/PD-L1 expression and tumour mutational burden: A systematic review-based approach. Ann. Oncol. 2019, 30, 1232–1243. [Google Scholar] [CrossRef]
- Thorsson, V.; Gibbs, D.L.; Brown, S.D.; Wolf, D.; Bortone, D.S.; Yang, T.H.O.; Porta-Pardo, E.; Gao, G.F.; Plaisier, C.L.; Eddy, J.A.; et al. The Immune Landscape of Cancer. Immunity 2018, 48, 812–830.e14. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Zhang, G.; Tang, T.; Liang, T. Identification of tumor antigens and immune subtypes of pancreatic adenocarcinoma for mRNA vaccine development. Mol. Cancer 2021, 20, 44. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Nie, Y.; Gu, J.; Cai, K.; Chen, X.; Li, H.; Wang, J. Identification of Mitochondrial-Related Prognostic Biomarkers Associated With Primary Bile Acid Biosynthesis and Tumor Microenvironment of Hepatocellular Carcinoma. Front. Oncol. 2021, 11, 587479. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Liu, J.; Zhao, D.; Diao, G. A Novel Prognostic Model for Identifying the Risk of Hepatocellular Carcinoma Based on Angiogenesis Factors. Front. Genet. 2022, 13, 857215. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Chen, L.; Yang, L.; Zhou, H.; Chen, Y.; Guo, Y. Integrated systemic analysis of FAM72A to identify its clinical relevance, biological function, and relationship to drug sensitivity in hepatocellular carcinoma. Front. Oncol. 2022, 12, 1046473. [Google Scholar] [CrossRef]
- Hinshaw, D.C.; Shevde, L.A. The Tumor Microenvironment Innately Modulates Cancer Progression. Cancer Res. 2019, 79, 4557–4566. [Google Scholar] [CrossRef]
- Goliwas, K.F.; Deshane, J.S.; Elmets, C.A.; Athar, M. Moving Immune Therapy Forward Targeting TME. Physiol. Rev. 2021, 101, 417–425. [Google Scholar] [CrossRef]
- Yoshihara, K.; Shahmoradgoli, M.; Martinez, E.; Vegesna, R.; Kim, H.; Torres-Garcia, W.; Trevino, V.; Shen, H.; Laird, P.W.; Levine, D.A.; et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 2013, 4, 2612. [Google Scholar] [CrossRef]
- Chan, T.A.; Yarchoan, M.; Jaffee, E.; Swanton, C.; Quezada, S.A.; Stenzinger, A.; Peters, S. Development of tumor mutation burden as an immunotherapy biomarker: Utility for the oncology clinic. Ann. Oncol. 2019, 30, 44–56. [Google Scholar] [CrossRef]
- Greillier, L.; Tomasini, P.; Barlesi, F. The clinical utility of tumor mutational burden in non-small cell lung cancer. Transl. Lung Cancer Res. 2018, 7, 639–646. [Google Scholar] [CrossRef]
- Bonneville, R.; Krook, M.A.; Kautto, E.A.; Miya, J.; Wing, M.R.; Chen, H.Z.; Reeser, J.W.; Yu, L.; Roychowdhury, S. Landscape of Microsatellite Instability Across 39 Cancer Types. JCO Precis. Oncol. 2017, 2017, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Picard, E.; Verschoor, C.P.; Ma, G.C.W.; Pawelec, G. Relationships Between Immune Landscapes, Genetic Subtypes and Responses to Immunotherapy in Colorectal Cancer. Front. Immunol. 2020, 11, 369. [Google Scholar] [CrossRef] [PubMed]
- Pawlowska, A.; Skiba, W.; Suszczyk, D.; Kuryło, W.; Jakubowicz-Gil, J.; Paduch, R.; Wertel, I. The Dual Blockade of the TIGIT and PD-1/PD-L1 Pathway as a New Hope for Ovarian Cancer Patients. Cancers 2022, 14, 5757. [Google Scholar] [CrossRef]
- Fujiwara, Y.; Kato, S.; Nesline, M.K.; Conroy, J.M.; DePietro, P.; Pabla, S.; Kurzrock, R. Indoleamine 2,3-dioxygenase (IDO) inhibitors and cancer immunotherapy. Cancer Treat. Rev. 2022, 110, 102461. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Khan, S.; Huang, D.; Li, L. V-Set and immunoglobulin domain containing (VSIG) proteins as emerging immune checkpoint targets for cancer immunotherapy. Front. Immunol. 2022, 13, 938470. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.T.; Lin, C.S.; Chai, C.Y.; Liu, K.Y.; Chen, J.Y.; Hsu, S.H. Functional interaction of Ugene and EBV infection mediates tumorigenic effects. Oncogene 2011, 30, 2921–2932. [Google Scholar] [CrossRef]
- Chen, F.; Fan, Y.; Cao, P.; Liu, B.; Hou, J.; Zhang, B.; Tan, K. Pan-Cancer Analysis of the Prognostic and Immunological Role of HSF1: A Potential Target for Survival and Immunotherapy. Oxid. Med. Cell. Longev. 2021, 2021, 5551036. [Google Scholar] [CrossRef]
- Goldman, M.J.; Craft, B.; Hastie, M.; Repecka, K.; McDade, F.; Kamath, A.; Banerjee, A.; Luo, Y.; Rogers, D.; Brooks, A.N.; et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 2020, 38, 675–678. [Google Scholar] [CrossRef]
- Tang, Z.; Li, C.; Kang, B.; Gao, G.; Li, C.; Zhang, Z. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017, 45, W98–W102. [Google Scholar] [CrossRef]
- Jin, Y.; Wang, Z.; He, D.; Zhu, Y.; Chen, X.; Cao, K. Identification of novel subtypes based on ssGSEA in immune-related prognostic signature for tongue squamous cell carcinoma. Cancer Med. 2021, 10, 8693–8707. [Google Scholar] [CrossRef]
- Jiang, J.; Ding, Y.; Chen, Y.; Lu, J.; Chen, Y.; Wu, G.; Xu, N.; Wang, H.; Teng, L. Pan-cancer analyses reveal that increased Hedgehog activity correlates with tumor immunosuppression and resistance to immune checkpoint inhibitors. Cancer Med. 2022, 11, 847–863. [Google Scholar] [CrossRef]
- Jiang, Z.; Liu, Z.; Li, M.; Chen, C.; Wang, X. Increased glycolysis correlates with elevated immune activity in tumor immune microenvironment. EBioMedicine 2019, 42, 431–442. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Feng, M.; Bai, L.; Liao, W.; Zhou, K.; Zhang, M.; Wu, Q.; Wen, F.; Lei, W.; Zhang, P.; et al. Comprehensive analysis of EMT-related genes and lncRNAs in the prognosis, immunity, and drug treatment of colorectal cancer. J. Transl. Med. 2021, 19, 391. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.; Ma, Z.; Zhang, W. Comprehensive Analysis to Identify SPP1 as a Prognostic Biomarker in Cervical Cancer. Front. Genet. 2021, 12, 732822. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Chen, H.; Yao, H.; Zhao, K.; Zhang, Y.; He, D.; Zhu, Y.; Cheng, Y.; Liu, R.; Xu, R.; et al. Turning up the heat on non-immunoreactive tumors: Pyroptosis influences the tumor immune microenvironment in bladder cancer. Oncogene 2021, 40, 6381–6393. [Google Scholar] [CrossRef]
- Chen, B.; Khodadoust, M.S.; Liu, C.L.; Newman, A.M.; Alizadeh, A.A. Profiling Tumor Infiltrating Immune Cells with CIBERSORT. Methods Mol. Biol. 2018, 1711, 243–259. [Google Scholar]
- Newman, A.M.; Liu, C.L.; Green, M.R.; Gentles, A.J.; Feng, W.; Xu, Y.; Hoang, C.D.; Diehn, M.; Alizadeh, A.A. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 2015, 12, 453–457. [Google Scholar] [CrossRef]
- Vega, D.M.; Yee, L.M.; McShane, L.M.; Williams, P.M.; Chen, L.; Vilimas, T.; Fabrizio, D.; Funari, V.; Newberg, J.; Bruce, L.K.; et al. Aligning tumor mutational burden (TMB) quantification across diagnostic platforms: Phase II of the Friends of Cancer Research TMB Harmonization Project. Ann. Oncol. 2021, 32, 1626–1636. [Google Scholar] [CrossRef]
- Wu, W.; Liu, Y.; Zeng, S.; Han, Y.; Shen, H. Intratumor heterogeneity: The hidden barrier to immunotherapy against MSI tumors from the perspective of IFN-gamma signaling and tumor-infiltrating lymphocytes. J. Hematol. Oncol. 2021, 14, 160. [Google Scholar] [CrossRef]
- Foroutan, M.; Molania, R.; Pfefferle, A.; Behrenbruch, C.; Scheer, S.; Kallies, A.; Speed, T.P.; Cursons, J.; Huntington, N.D. The Ratio of Exhausted to Resident Infiltrating Lymphocytes Is Prognostic for Colorectal Cancer Patient Outcome. Cancer Immunol. Res. 2021, 9, 1125–1140. [Google Scholar] [CrossRef]
- Liu, J.N.; Kong, X.S.; Sun, P.; Wang, R.; Li, W.; Chen, Q.F. An integrated pan-cancer analysis of TFAP4 aberrations and the potential clinical implications for cancer immunity. J. Cell. Mol. Med. 2021, 25, 2082–2097. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [PubMed]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bai, Y.; Cao, K.; Zhang, P.; Ma, J.; Zhu, J. Prognostic and Immunological Implications of FAM72A in Pan-Cancer and Functional Validations. Int. J. Mol. Sci. 2023, 24, 375. https://doi.org/10.3390/ijms24010375
Bai Y, Cao K, Zhang P, Ma J, Zhu J. Prognostic and Immunological Implications of FAM72A in Pan-Cancer and Functional Validations. International Journal of Molecular Sciences. 2023; 24(1):375. https://doi.org/10.3390/ijms24010375
Chicago/Turabian StyleBai, Yuwen, Kui Cao, Ping Zhang, Jianqun Ma, and Jinhong Zhu. 2023. "Prognostic and Immunological Implications of FAM72A in Pan-Cancer and Functional Validations" International Journal of Molecular Sciences 24, no. 1: 375. https://doi.org/10.3390/ijms24010375
APA StyleBai, Y., Cao, K., Zhang, P., Ma, J., & Zhu, J. (2023). Prognostic and Immunological Implications of FAM72A in Pan-Cancer and Functional Validations. International Journal of Molecular Sciences, 24(1), 375. https://doi.org/10.3390/ijms24010375





