Comparative Genomics of Seasonal Senescence in Forest Trees
Abstract
:1. Introduction
1.1. Genetic Regulation of Leaf Senescence
1.1.1. Regulation at the Transcriptional Level
1.1.2. Regulation at the Post-Transcriptional Level
1.1.3. Regulation at the Translational Level
1.1.4. Regulation at the Post-Translational Level
1.2. The Role of Phytohormones in the Regulation of Leaf Senescence
2. Results
3. Discussion
4. Materials and Methods
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ABA-WDS | ABA-water deficit stress domain |
| ADR1 | activated disease resistance 1 |
| AP2 | apetala 2 |
| Bet v 1 | pathogenesis-related protein Bet v 1 family |
| DRM1 | dormancy-associated protein 1 |
| EXL2 | exordium like 2 |
| JMJ16 | jumonji domain-containing protein 16 |
| LRR-RLK | leucine-rich repeat receptor-like kinase |
| NAC | NAC transcription factor |
| NB-ARC | nucleotide-binding domain shared with APAF-1, various R-proteins and CED-4 |
| Oxidored FMN | NADH:flavin oxidoreductase/NADH oxidase family |
| Phi 1 | phosphate-induced protein 1 conserved region |
| Pkinase | protein kinase domain |
| ROS | reactive oxygen species |
| RPK1 | receptor protein kinase 1 |
| SAG201 | senescence-associated gene 201 |
| WRKY | WRKY transcription factor |
References
- Kim, J.; Kim, J.H.; Lyu, J.I.; Woo, H.R.; Lim, P.O. New insights into the regulation of leaf senescence in Arabidopsis. J. Exp. Bot. 2018, 69, 787–799. [Google Scholar] [CrossRef] [Green Version]
- Cho, E.J.; Choi, S.H.; Kim, J.H.; Kim, J.E.; Lee, M.H.; Chung, B.Y.; Woo, H.R.; Kim, J.H. A mutation in plant-specific SWI2/SNF2-like chromatin-remodeling proteins, DRD1 and DDM1, delays leaf senescence in Arabidopsis thaliana. PLoS ONE 2016, 11, e0146826. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Zhang, S.; Zhou, B.; Luo, X.; Zhou, X.F.; Cai, B.; Jin, Y.H.; Niu, D.; Lin, J.; Cao, X.; et al. The histone H3K4 demethylase JMJ16 represses leaf senescence in Arabidopsis. Plant Cell 2019, 31, 430–443. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Steele, J.F.C.; Hughes, R.K.; Banfield, M.J. Structural and biochemical studies of an NB-ARC domain from a plant NLR immune receptor. PLoS ONE 2019, 14, e0221226. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, X.; Chen, J.; Xie, Z.; Gao, J.; Ren, G.; Gao, S.; Zhou, X.; Kuai, B. Jasmonic acid promotes degreening via MYC2/3/4- and ANAC019/055/072-mediated regulation of major chlorophyll catabolic genes. Plant J. 2015, 84, 597–610. [Google Scholar] [CrossRef]
- Kim, Y.S.; Sakuraba, Y.; Han, S.H.; Yoo, S.C.; Paek, N.C. Mutation of the Arabidopsis NAC016 transcription factor delays leaf senescence. Plant Cell Physiol. 2013, 54, 1660–1672. [Google Scholar] [CrossRef]
- Jin, H.K.; Hye, R.W.; Kim, J.; Pyung, O.L.; In, C.L.; Seung, H.C.; Hwang, D.; Hong, G.N. Trifurcate feed-forward regulation of age-dependent cell death involving MiR164 in Arabidopsis. Science 2009, 323, 1053–1057. [Google Scholar] [CrossRef]
- Li, Z.; Peng, J.; Wen, X.; Guo, H. ETHYLENE-INSENSITIVE3 is a senescence-associated gene that accelerates age-dependent leaf senescence by directly repressing MiR164 transcription in Arabidopsis. Plant Cell 2013, 25, 3311–3328. [Google Scholar] [CrossRef] [Green Version]
- Schommer, C.; Palatnik, J.F.; Aggarwal, P.; Chételat, A.; Cubas, P.; Farmer, E.E.; Nath, U.; Weigel, D. Control of jasmonate biosynthesis and senescence by MiR319 targets. PLoS Biol. 2008, 6, e230. [Google Scholar] [CrossRef] [Green Version]
- Marin, E.; Jouannet, V.; Herz, A.; Lokerse, A.S.; Weijers, D.; Vaucheret, H.; Nussaume, L.; Crespi, M.D.; Maizel, A. MiR390, Arabidopsis TAS3 TasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell 2012, 22, 1104–1117. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; Lu, L.; Mayer, K.S.; Scalf, M.; Qian, S.; Lomax, A.; Smith, L.M.; Zhong, X. POWERDRESS interacts with HISTONE DEACETYLASE 9 to promote aging in Arabidopsis. eLife 2016, 5, e17214. [Google Scholar] [CrossRef] [PubMed]
- Woo, H.R.; Goh, C.H.; Park, J.H.; De La Serve, B.T.; Kim, J.H.; Park, Y.I.; Nam, H.G. Extended leaf longevity in the Ore4-1 mutant of Arabidopsis with a reduced expression of a plastid ribosomal protein gene. Plant J. 2002, 31, 331–340. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Woo, H.R.; Kim, H.J.; Nam, H.G.; Lim, P.O. Plant leaf senescence and death-regulation by multiple layers of control and implications for aging in general. J. Cell Sci. 2013, 126, 4823–4833. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miao, Y.; Laun, T.M.; Smykowski, A.; Zentgraf, U. Arabidopsis MEKK1 can take a short cut: It can directly interact with senescence-related WRKY53 transcription factor on the protein level and can bind to its promoter. Plant Mol. Biol. 2007, 65, 63–76. [Google Scholar] [CrossRef] [PubMed]
- Miao, Y.; Zentgraf, U. A HECT E3 ubiquitin ligase negatively regulates Arabidopsis leaf senescence through degradation of the transcription factor WRKY53. Plant J. 2010, 63, 179–188. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.C.; Hong, S.W.; Whang, S.S.; Lim, P.O.; Nam, H.G.; Koo, J. Age-dependent action of an ABA-inducible receptor kinase, RPK1, as a positive regulator of senescence in Arabidopsis leaves. Plant Cell Physiol. 2011, 52, 651–662. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Wang, H.L.; Li, Z.; Guo, H. Genetic network between leaf senescence and plant immunity: Crucial regulatory nodes and new insights. Plants 2020, 9, 495. [Google Scholar] [CrossRef] [Green Version]
- Lim, P.O.; Lee, I.C.; Kim, J.; Kim, H.J.; Ryu, J.S.; Woo, H.R.; Nam, H.G. Auxin response factor 2 (ARF2) plays a major role in regulating auxin-mediated leaf longevity. J. Exp. Bot. 2010, 61, 1419–1430. [Google Scholar] [CrossRef]
- Emms, D.M.; Kelly, S. OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol. 2019, 20, 238. [Google Scholar] [CrossRef] [Green Version]
- Gernandt, D.S.; Willyard, A.; Syring, J.V.; Liston, A. The Conifers (Pinophyta). In Genetics, Genomics and Breeding of Conifers; Plomion, C., Bousquet, J., Kole, C., Eds.; CRC Press: Boca Raton, FL, USA; Science Publishers: New York, NY, USA, 2011; pp. 1–39. ISBN 9780429065934. [Google Scholar]
- Maddison, D.R.; Schulz, K.-S. (Eds.) The Tree of Life Project. Zootaxa 2007, 1668, 19–40. Available online: http://tolweb.org (accessed on 22 March 2022). [CrossRef]
- Klepikova, A.V.; Kasianov, A.S.; Gerasimov, E.S.; Logacheva, M.D.; Penin, A.A. A high resolution map of the Arabidopsis thaliana developmental transcriptome based on RNA-Seq profiling. Plant J. 2016, 88, 1058–1070. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Chen, C.; Chen, Z. Expression profiles of the Arabidopsis WRKY gene superfamily during plant defense response. Plant Mol. Biol. 2003, 51, 21–37. [Google Scholar] [CrossRef] [PubMed]
- Gaudet, P.; Livstone, M.S.; Lewis, S.E.; Thomas, P.D. Phylogenetic-based propagation of functional annotations within the gene ontology consortium. Brief. Bioinform. 2011, 12, 449–462. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Cordewener, J.H.G.; America, A.H.P.; Shan, W.; Bouwmeester, K.; Govers, F. Arabidopsis lectin receptor kinases LecRK-IX.1 and LecRK-IX.2 are functional analogs in regulating Phytophthora resistance and plant cell death. Mol. Plant Microbe Interact. 2015, 28, 1032–1048. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, C.W.; Yang, S.H.; Shin, K.H.; Lee, S.C.; Kim, S.H. The AtLRK10L1.2, Arabidopsis ortholog of wheat LRK10, is involved in ABA-mediated signaling and drought resistance. Plant Cell Rep. 2015, 34, 447–455. [Google Scholar] [CrossRef]
- McHale, L.; Tan, X.; Koehl, P.; Michelmore, R.W. Plant NBS-LRR proteins: Adaptable guards. Genome Biol. 2006, 7, 212. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Cui, J.; Jin, B.; Zhao, J.; Xu, H.; Lu, Z.; Li, W.; Li, X.; Li, L.; Liang, E.; et al. Multifeature analyses of vascular cambial cells reveal longevity mechanisms in old Ginkgo biloba trees. Proc. Natl. Acad. Sci. USA 2020, 117, 2201–2210. [Google Scholar] [CrossRef] [Green Version]
- Chini, A.; Grant, J.J.; Seki, M.; Shinozaki, K.; Loake, G.J. Drought tolerance established by enhanced expression of the CC–NBS–LRR Gene, ADR1, requires salicylic acid, EDS1 and ABI1. Plant J. 2004, 38, 810–822. [Google Scholar] [CrossRef]
- Schröder, F.; Lisso, J.; Müssig, C. Expression pattern and putative function of EXL1 and Homologous genes in Arabidopsis. Plant Signal. Behav. 2012, 7, 22–27. [Google Scholar] [CrossRef] [Green Version]
- Gonzali, S.; Loreti, E.; Solfanelli, C.; Novi, G.; Alpi, A.; Perata, P. Identification of sugar-modulated genes and evidence for in vivo sugar sensing in Arabidopsis. J. Plant Res. 2006, 119, 115–123. [Google Scholar] [CrossRef]
- Kuzmin, D.A.; Feranchuk, S.I.; Sharov, V.V.; Cybin, A.N.; Makolov, S.V.; Putintseva, Y.A.; Oreshkova, N.V.; Krutovsky, K.V. Stepwise large genome assembly approach: A case of Siberian larch (Larix sibirica Ledeb.). BMC Bioinform. 2019, 20, 35–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; Van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rice, P.; Longden, L.; Bleasby, A. EMBOSS: The European Molecular Biology Open Software Suite. Trends Genet. 2000, 16, 276–277. [Google Scholar] [CrossRef]
- Huerta-Cepas, J.; Forslund, K.; Coelho, L.P.; Szklarczyk, D.; Jensen, L.J.; Von Mering, C.; Bork, P. Fast genome-wide functional annotation through orthology assignment by EggNOG-Mapper. Mol. Biol. Evol. 2017, 34, 2115–2122. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [Green Version]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2014, 12, 59–60. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [Green Version]
- Emms, D.M.; Kelly, S. STAG: Species Tree Inference from All Genes. bioRxiv 2018, 267914. [Google Scholar] [CrossRef]
- Huson, D.H.; Scornavacca, C. Dendroscope 3: An interactive tool for rooted phylogenetic trees and networks. Syst. Biol. 2012, 61, 1061–1067. [Google Scholar] [CrossRef] [Green Version]
- Huson, D.H.; Richter, D.C.; Rausch, C.; Dezulian, T.; Franz, M.; Rupp, R. Dendroscope: An interactive viewer for large phylogenetic trees. BMC Bioinform. 2007, 8, 460. [Google Scholar] [CrossRef] [Green Version]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hammer, D.A.T.; Ryan, P.D.; Hammer, Ø.; Harper, D.A.T. Past: Paleontological statistics software package for education and data analysis. Palaeontol. Electron. 2001, 4, 178. [Google Scholar]
| Group | Type | Species | Eukaryotes | Undefined | Total |
|---|---|---|---|---|---|
| Gymnosperms (Pinaceae) | deciduous | Larix sibirica | 34,323 | 4391 | 38,714 |
| evergreen | Pseudotsuga menziesii | 46,066 | 478 | 46,544 | |
| Pinus taeda | 51,113 | 527 | 51,640 | ||
| Pinus lambertiana | 38,024 | 436 | 38,460 | ||
| Picea abies | 43,626 | - | 43,626 | ||
| Angiosperms (Pentapetalae) | deciduous | Populus euphratica | 30,272 | 414 | 30,686 |
| Populus trichocarpa | 31,209 | 424 | 31,633 | ||
| Prunus persica | 22,706 | 425 | 23,131 | ||
| Prunus dulcis | 22,765 | - | 22,765 | ||
| evergreen | Coffea eugenoides | 27,514 | 1584 | 29,098 | |
| Coffea canephora | 23,204 | 2344 | 25,548 | ||
| Hevea brasiliensis | 34,297 | 867 | 35,164 | ||
| Citrus sinensis | 24,119 | 412 | 24,531 | ||
| Citrus clementina | 22,523 | 296 | 22,819 |
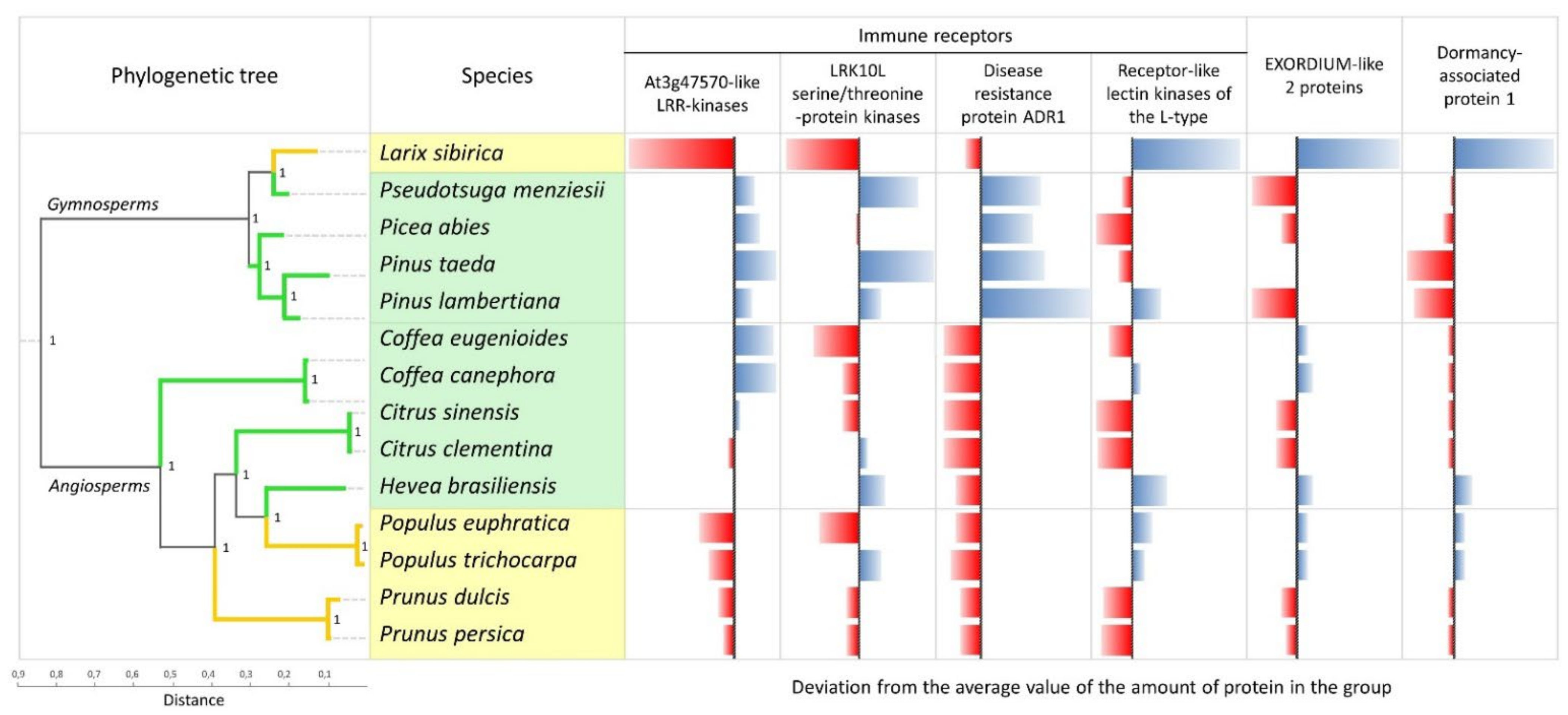
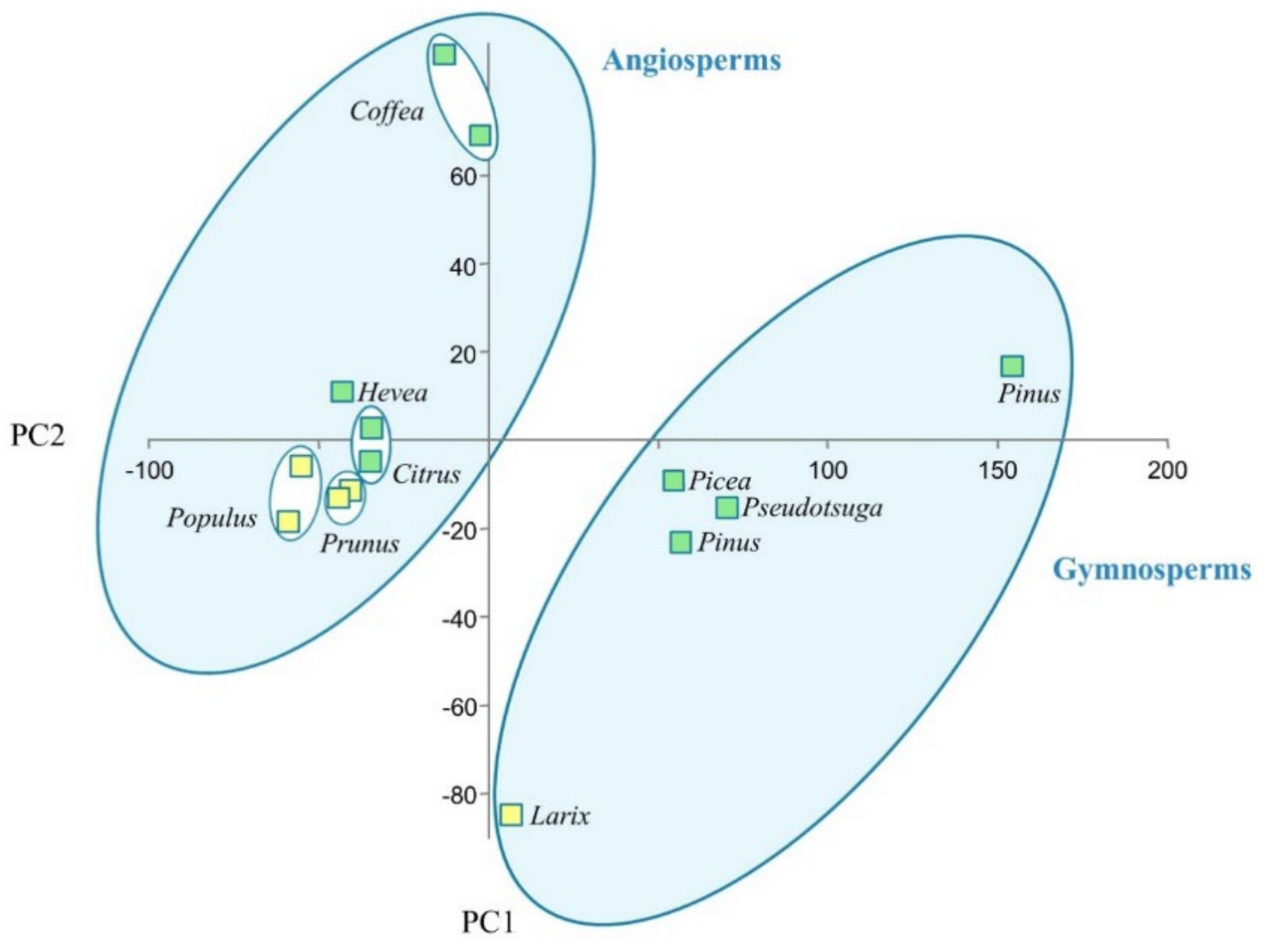
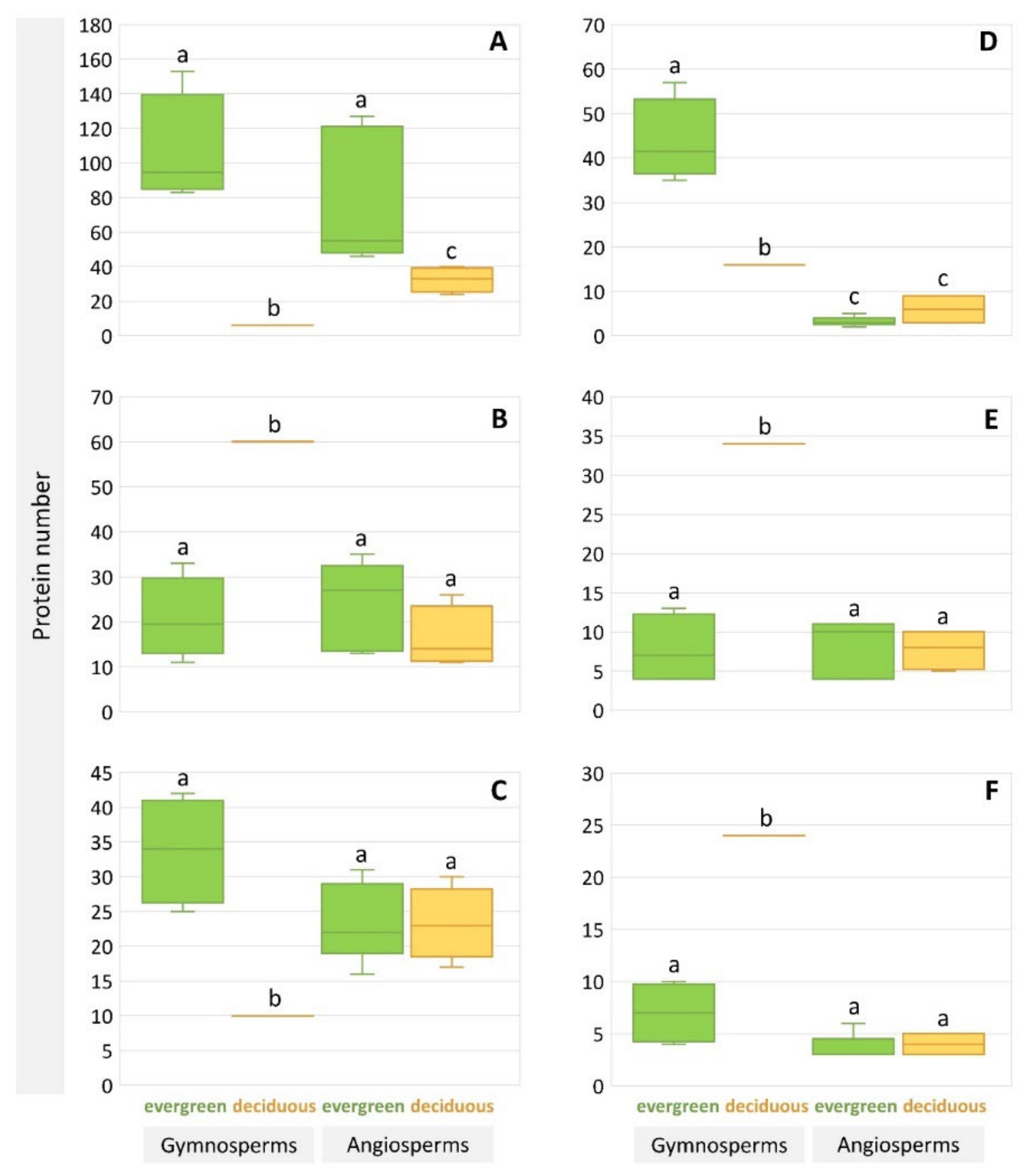
| Type | Species | At3g47570-Like LRR-Kinases | LRK10L Serine/Threonine-Protein Kinases | Disease Resistance Protein ADR1 | Receptor-Like Lectin Kinases of the L-Type | EXORDIUM-Like 2 Proteins | Dormancy-Associated Protein 1 | Total |
|---|---|---|---|---|---|---|---|---|
| deciduous | Larix sibirica | 6 | 10 | 9 | 60 | 34 | 24 | 38,714 |
| evergreen | Pseudotsuga menziensii | 90 | 38 | 31 | 20 | 4 | 10 | 46,544 |
| Pinus taeda | 153 | 42 | 32 | 11 | 13 | 4 | 51,640 | |
| Pinus lambertiana | 83 | 30 | 46 | 19 | 4 | 5 | 38,460 | |
| Picea abies | 99 | 25 | 29 | 33 | 10 | 9 | 43,626 | |
| deciduous | Populus euphratica | 24 | 17 | 6 | 16 | 10 | 5 | 30,686 |
| Populus trichocarpa | 29 | 30 | 5 | 26 | 10 | 5 | 31,633 | |
| Prunus persica | 40 | 23 | 8 | 11 | 6 | 3 | 23,131 | |
| Prunus dulcis | 37 | 23 | 8 | 12 | 5 | 3 | 22,765 | |
| evergreen | Coffea eugenoides | 115 | 16 | 2 | 35 | 10 | 3 | 29,098 |
| Coffea canephora | 127 | 22 | 2 | 30 | 11 | 3 | 25,548 | |
| Hevea brasiliensis | 50 | 31 | 6 | 27 | 11 | 6 | 35,164 | |
| Citrus sinensis | 55 | 22 | 3 | 14 | 4 | 3 | 24,531 | |
| Citrus clementina | 46 | 27 | 3 | 13 | 4 | 3 | 22,819 |
| Orthogroup | Species | Larix sibirica | Pseudotsuga menziensii | Pinus taeda | Pinus lambertiana |
|---|---|---|---|---|---|
| At3g47570-like LRR-kinases | Pseudotsuga menziesii | 2.2 × 10−16↓ | |||
| Pinus taeda | 2.2 × 10−16↓ | 0.001↓ | |||
| Pinus lambertiana | 2.2 × 10−16↓ | 0.492 | 0.021↑ | ||
| Picea abies | 2.2 × 10−16↓ | 0.276 | 0.043↑ | 0.766 | |
| LRK10L serine/threonine-protein kinases | Pseudotsuga menziesii | 6.8 × 10−4↓ | |||
| Pinus taeda | 4.0 × 10−4↓ | 1.000 | |||
| Pinus lambertiana | 1.4 × 10−3↓ | 0.903 | 0.906 | ||
| Picea abies | 0.041↓ | 0.207 | 0.178 | 0.281 | |
| Disease resistance protein ADR1 | Pseudotsuga menziesii | 3,8 × 10−3↓ | |||
| Pinus taeda | 6,8 × 10−3↓ | 0.802 | |||
| Pinus lambertiana | 2,4 × 10−7↓ | 0.012↓ | 0.004↓ | ||
| Picea abies | 5,0 × 10−3↓ | 1.000 | 0.798 | 0.015↑ | |
| Receptor-like lectin kinases of the L-type | Pseudotsuga menziesii | 1.1 × 10−7↑ | |||
| Pinus taeda | 6.6 × 10−13↑ | 0.071 | |||
| Pinus lambertiana | 4.2 × 10−6↑ | 0.748 | 0.026↓ | ||
| Picea abies | 8.1 × 10−4↑ | 0.053↓ | 0.000↓ | 0.164 | |
| EXORDIUM-like 2 proteins | Pseudotsuga menziesii | 1.9 × 10−8↑ | |||
| Pinus taeda | 5.0 × 10−5↑ | 0.054↓ | |||
| Pinus lambertiana | 6.0 × 10−7↑ | 1.000 | 0.142 | ||
| Picea abies | 5.6 × 10−5↑ | 0.109 | 1.000 | 0.192 | |
| DRM1-like proteins | Pseudotsuga menziesii | 5.0 × 10−3↑ | |||
| Pinus taeda | 3.8 × 10−6↑ | 0.106 | |||
| Pinus lambertiana | 5.5 × 10−4↑ | 0.442 | 0.510 | ||
| Picea abies | 4.5 × 10−3↑ | 1.000 | 0.102 | 0.437 |
| Species | NB-ARC (PF00931) | Phi 1 (PF04674) | Oxidored FMN (PF00724) | Pkinase (PF00069) | AP2 (PF00847) | ABA-WDS (PF02496) | 14-3-3 (PF00244) | Bet v 1 (PF00407) |
|---|---|---|---|---|---|---|---|---|
| Larix sibirica | 136 | 66 | 48 | 1867 | 374 | 46 | 26 | 63 |
| Pseudotsuga menziensii | 508 | 22 | 34 | 1920 | 139 | 5 | 13 | 35 |
| Pinus taeda | 849 | 27 | 18 | 2482 | 190 | 1 | 3 | 48 |
| Pinus lambertiana | 780 | 16 | 28 | 1750 | 125 | 8 | 4 | 52 |
| Picea abies | 696 | 41 | 28 | 1930 | 269 | 16 | 22 | 49 |
| Populus euphratica | 327 | 15 | 4 | 1501 | 216 | 1 | 16 | 47 |
| Populus trichocarpa | 566 | 16 | 9 | 1228 | 205 | 3 | 18 | 50 |
| Prunus persica | 388 | 10 | 10 | 1050 | 126 | 3 | 14 | 44 |
| Prunus dulcis | 377 | 10 | 7 | 1046 | 129 | 4 | 13 | 46 |
| Coffea eugenoides | 948 | 13 | 13 | 1398 | 133 | 4 | 15 | 57 |
| Coffea canephora | 798 | 15 | 18 | 1281 | 104 | 3 | 15 | 47 |
| Hevea brasiliensis | 575 | 17 | 11 | 1931 | 215 | 5 | 26 | 62 |
| Citrus sinensis | 615 | 7 | 9 | 1180 | 136 | 2 | 18 | 35 |
| Citrus clementina | 578 | 7 | 10 | 1083 | 125 | 3 | 15 | 32 |
| Type | Species | NCBI GenBank # | |
|---|---|---|---|
| Gymnosperms | deciduous | Larix sibirica | GCA_004151065.1 |
| evergreen | Pseudotsuga menziensii | GCA_001517045.1 | |
| Pinus taeda | GCA_000404065.3 | ||
| Pinus lambertiana | GCA_001447015.2 | ||
| Picea abies | GCA_900491625.1 | ||
| Angiosperms | deciduous | Populus euphratica | GCA_000495115.1 |
| Populus trichocarpa | GCA_000002775.3 | ||
| Prunus persica | GCA_000346465.2 | ||
| Prunus dulcis | GCA_902201215.1 | ||
| evergreen | Coffea eugenoides | GCA_003713205.1 | |
| Coffea canephora | GCA_900059795.1 | ||
| Hevea brasiliensis | GCA_001654055.1 | ||
| Citrus sinensis | GCA_000317415.1 | ||
| Citrus clementina | GCA_000493195.1 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Batalova, A.Y.; Putintseva, Y.A.; Sadovsky, M.G.; Krutovsky, K.V. Comparative Genomics of Seasonal Senescence in Forest Trees. Int. J. Mol. Sci. 2022, 23, 3761. https://doi.org/10.3390/ijms23073761
Batalova AY, Putintseva YA, Sadovsky MG, Krutovsky KV. Comparative Genomics of Seasonal Senescence in Forest Trees. International Journal of Molecular Sciences. 2022; 23(7):3761. https://doi.org/10.3390/ijms23073761
Chicago/Turabian StyleBatalova, Anastasia Y., Yuliya A. Putintseva, Michael G. Sadovsky, and Konstantin V. Krutovsky. 2022. "Comparative Genomics of Seasonal Senescence in Forest Trees" International Journal of Molecular Sciences 23, no. 7: 3761. https://doi.org/10.3390/ijms23073761
APA StyleBatalova, A. Y., Putintseva, Y. A., Sadovsky, M. G., & Krutovsky, K. V. (2022). Comparative Genomics of Seasonal Senescence in Forest Trees. International Journal of Molecular Sciences, 23(7), 3761. https://doi.org/10.3390/ijms23073761








