YdfD, a Lysis Protein of the Qin Prophage, Is a Specific Inhibitor of the IspG-Catalyzed Step in the MEP Pathway of Escherichia coli
Abstract
1. Introduction
2. Results
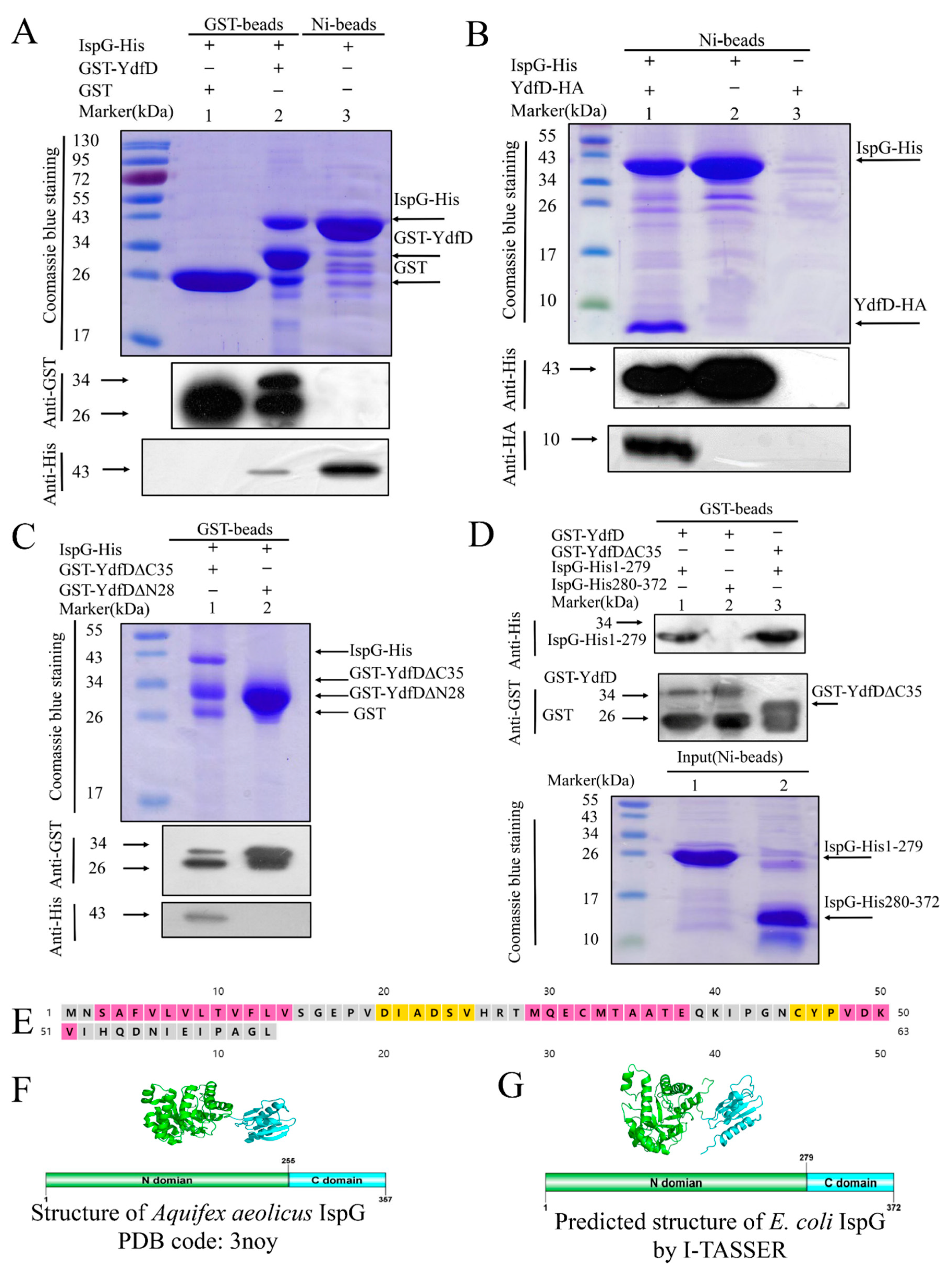
2.1. Protein-Protein Binding between YdfD and IspG
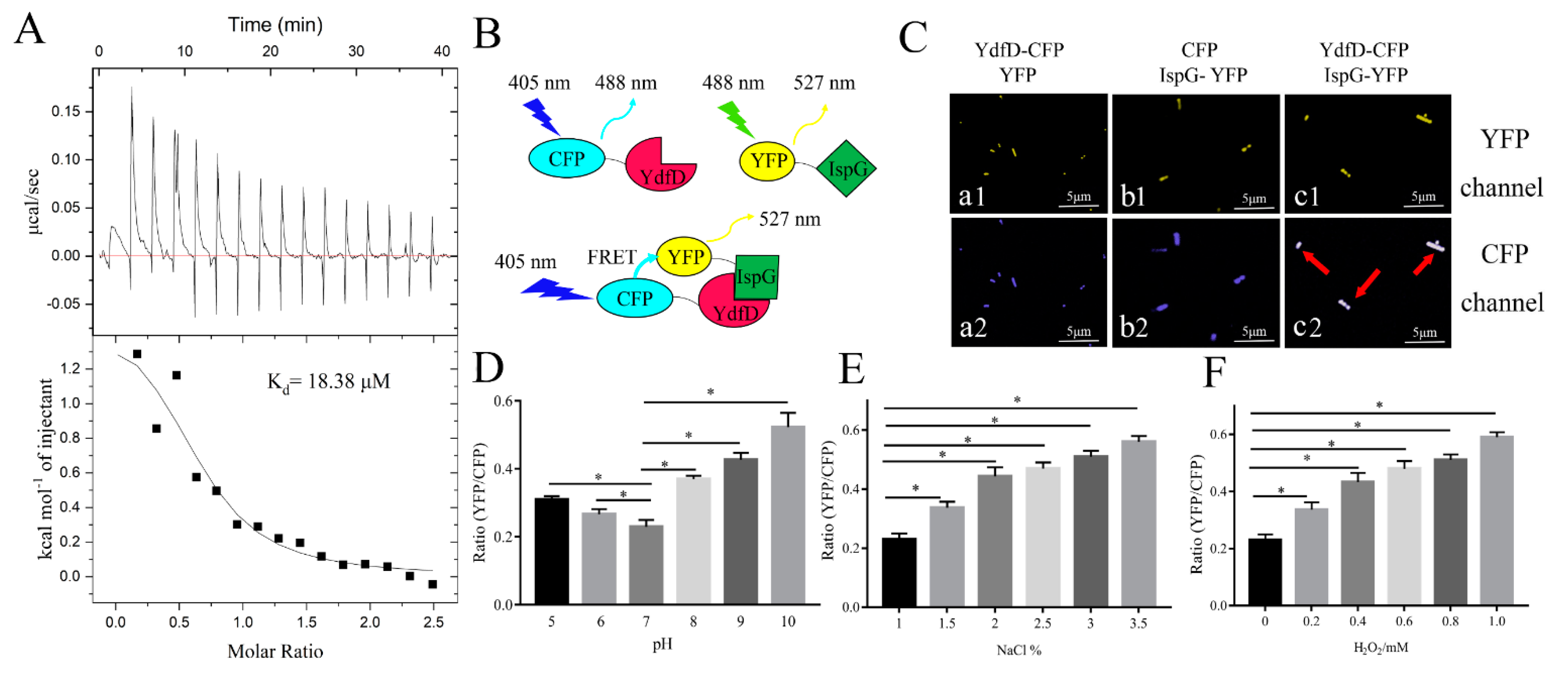
2.2. YdfD Forms Stable Complex with IspG in Solution
2.3. YdfD and IspG Interactions In Vitro and Vivo
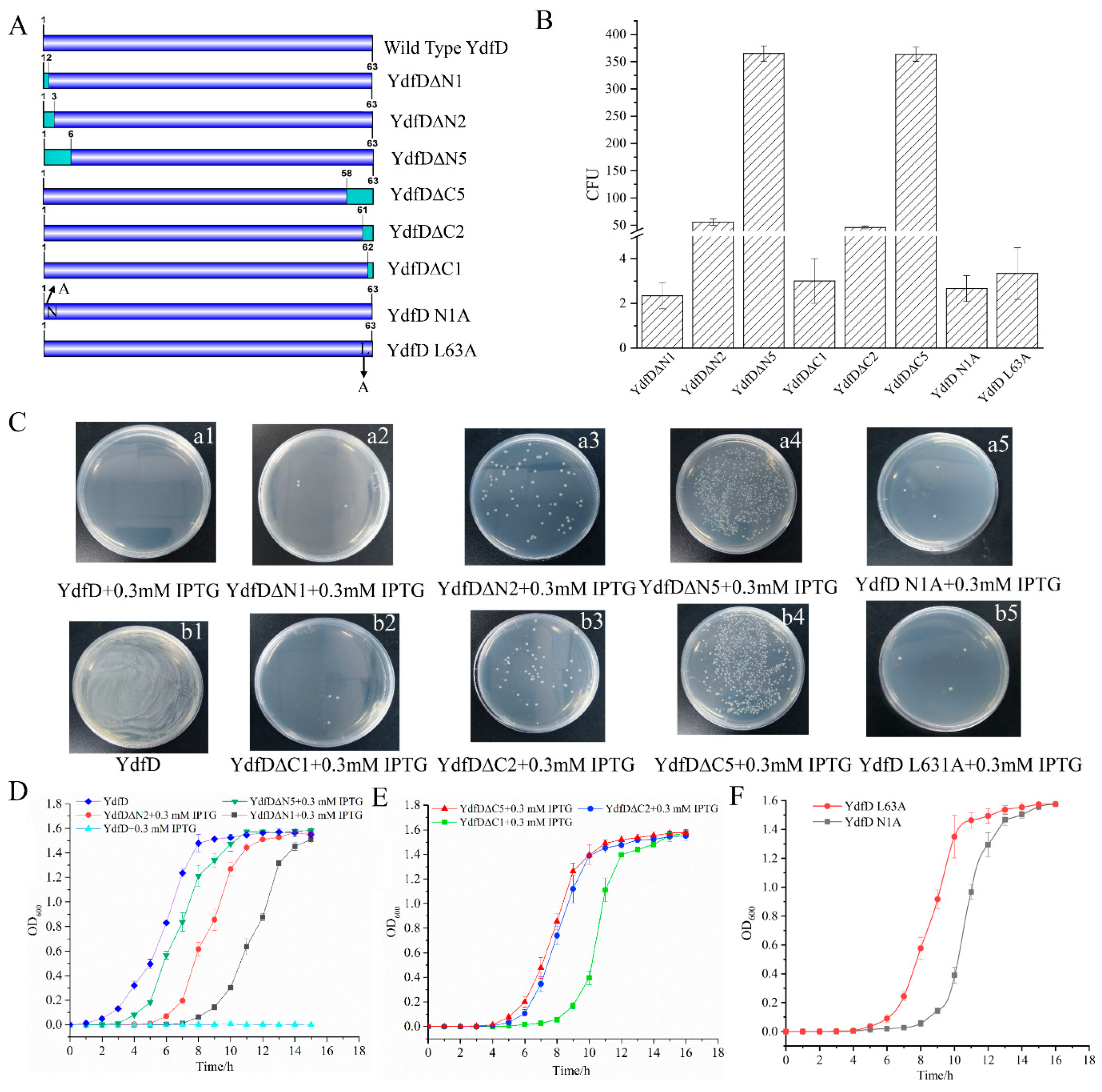
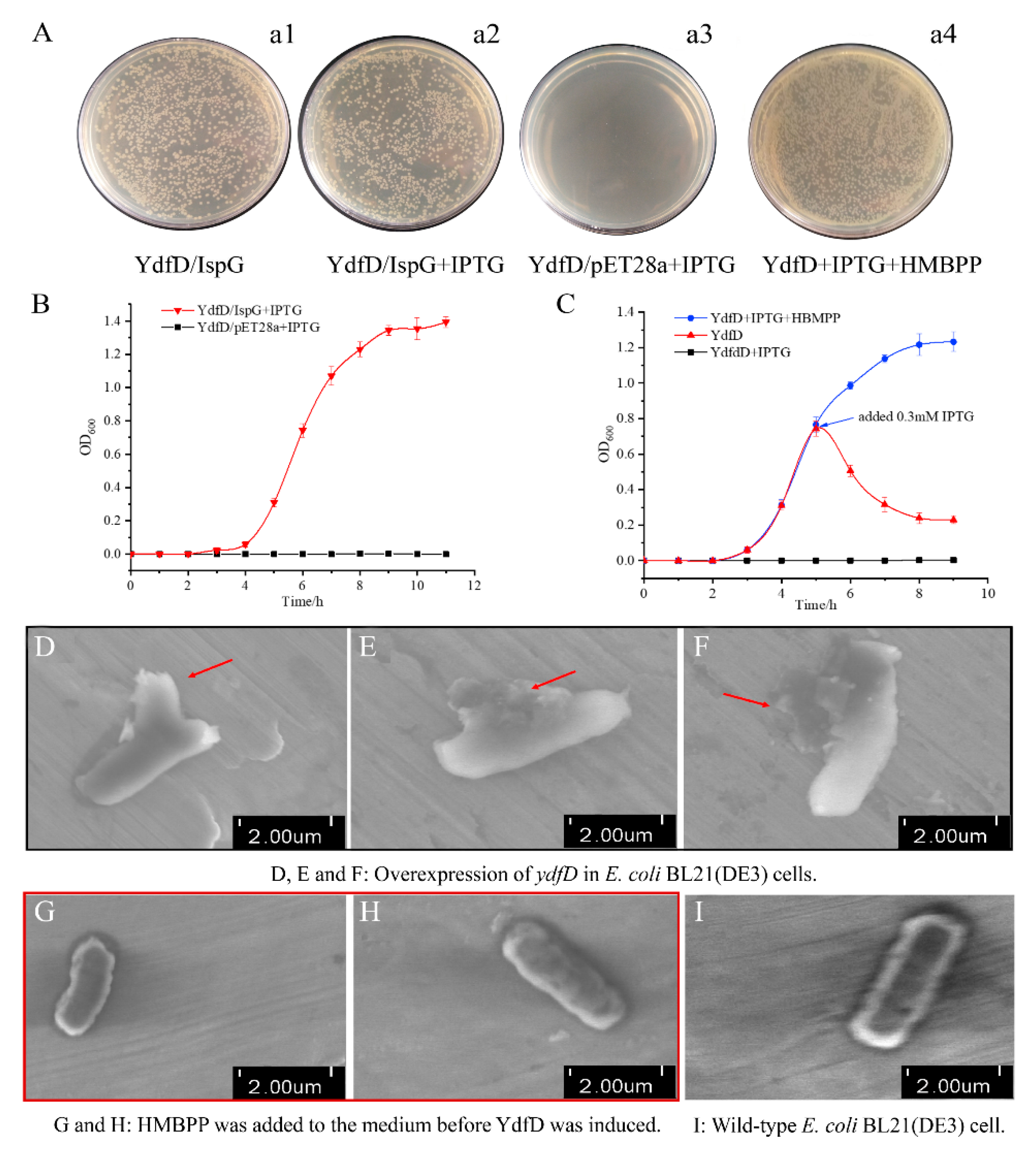
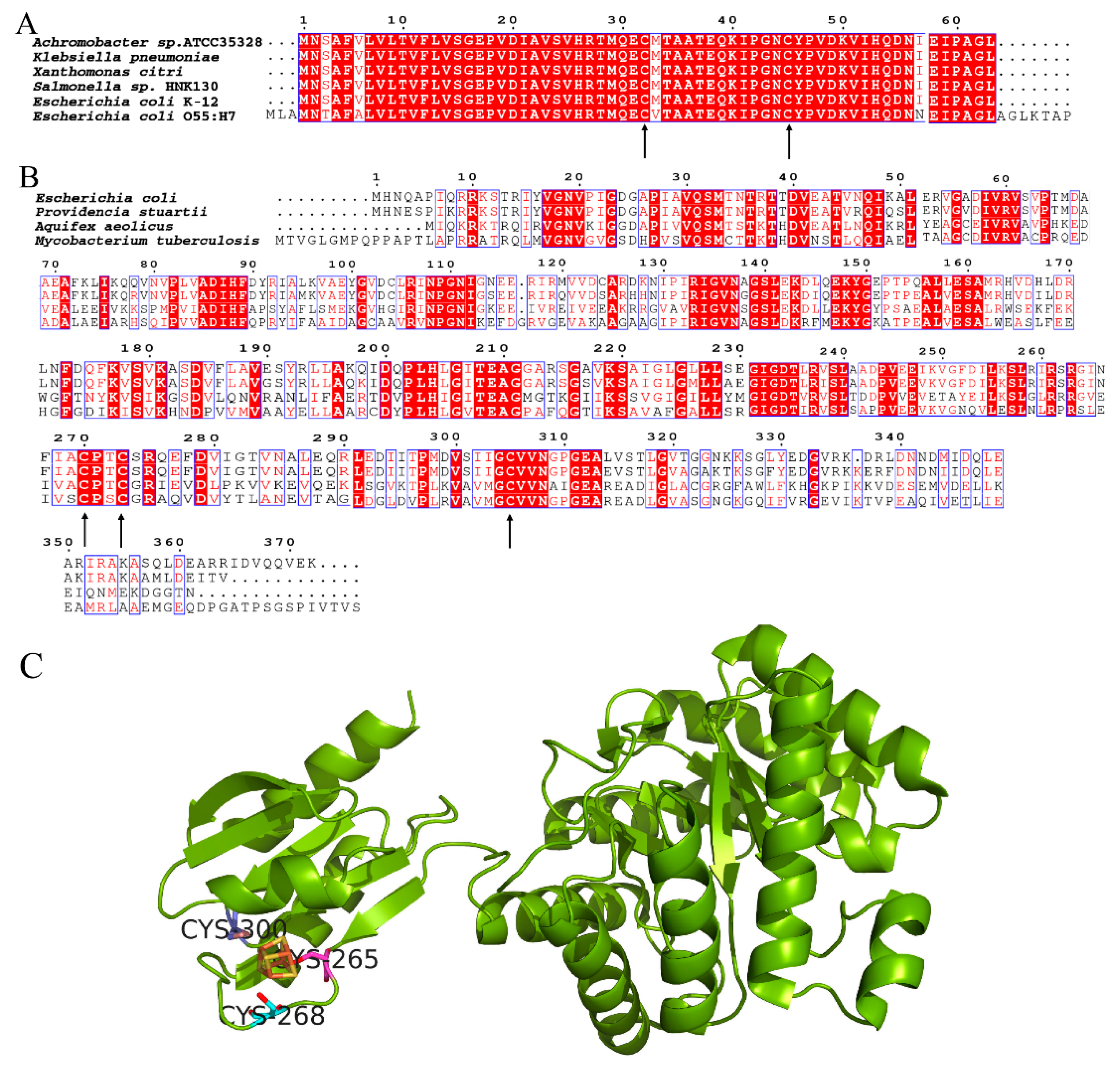
2.4. The Intact YdfD Protein Is Required for Its Cell Lysis Function
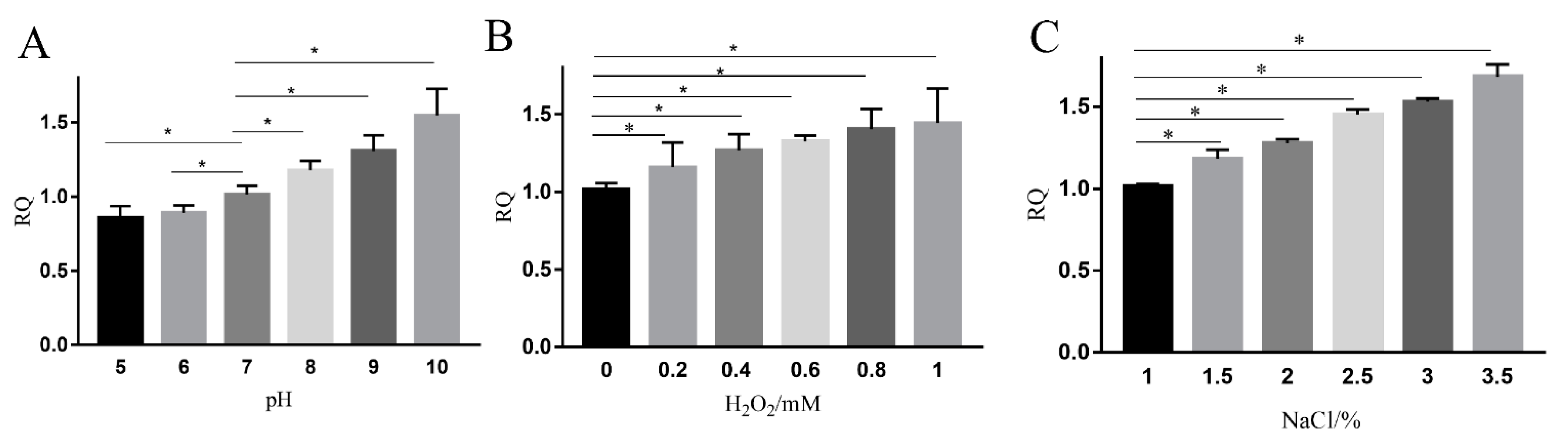
2.5. mRNA Levels of ydfD Increased under Stress
2.6. Compensation Experiments
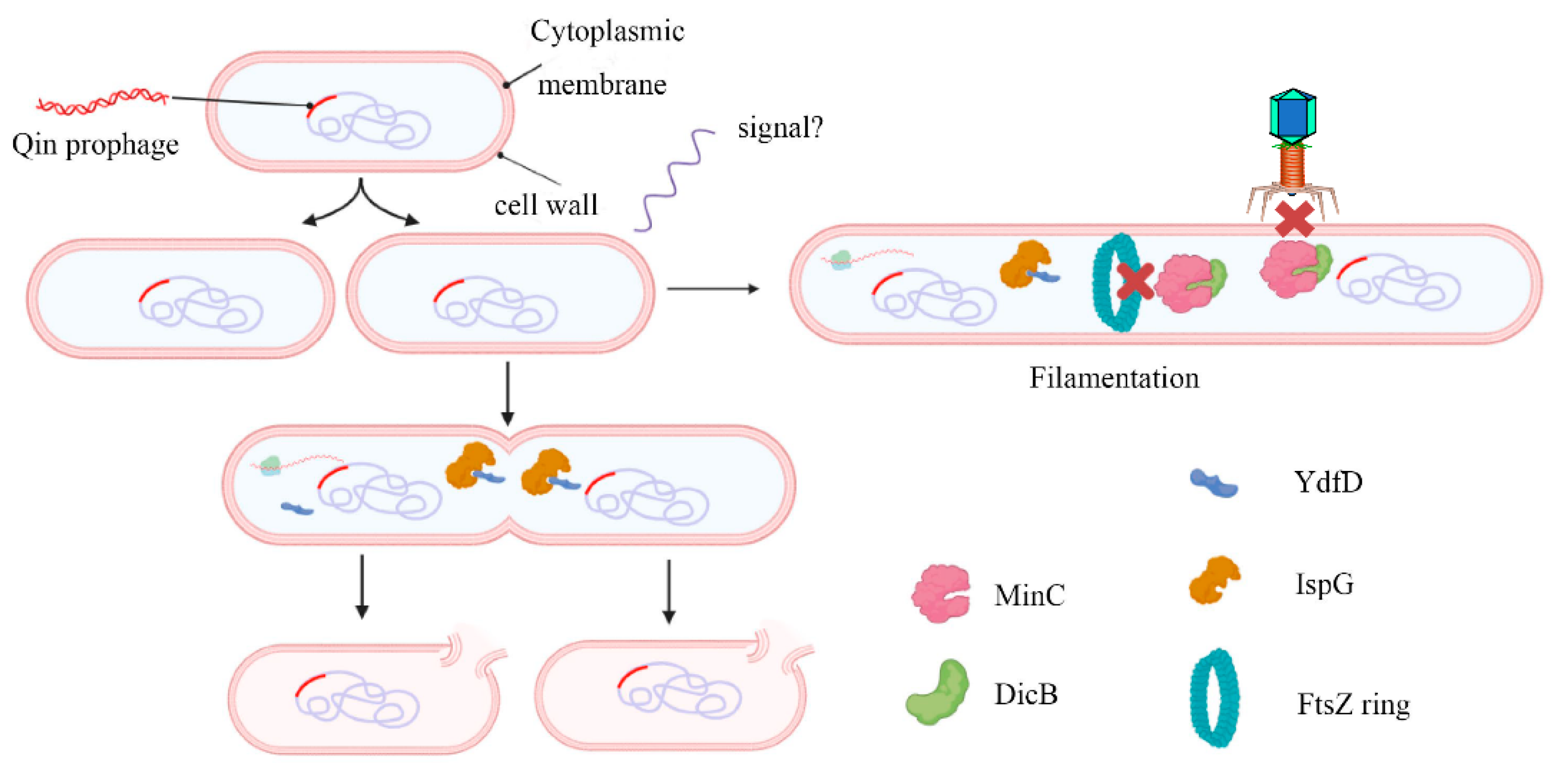
3. Discussion
4. Materials and Methods
4.1. Cell Strains and Plasmids Construction
4.2. Protein Expression and Purification
4.3. Mass Spectrometry
4.4. Pull-Down Assays
4.5. Cell Survival Assays
4.6. Stress Experiments
4.7. Fluorescence Resonance Energy Transfer
4.8. Isothermal Titration Calorimetry
4.9. Compensation Experiment
4.10. Scanning Electron Microscopy
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Arndt, D.; Marcu, A.; Liang, Y.J.; Wishart, D.S. PHAST, PHASTER and PHASTEST: Tools for finding prophage in bacterial genomes. Brief. Bioinform. 2019, 20, 1560–1567. [Google Scholar] [CrossRef] [PubMed]
- Reis-Cunha, J.L.; Bartholomeu, D.C.; Manson, A.L.; Earl, A.M.; Cerqueira, G.C. ProphET, prophage estimation tool: A stand-alone prophage sequence prediction tool with self-updating reference database. PLoS ONE 2019, 14, e0223364. [Google Scholar] [CrossRef] [PubMed]
- Song, W.C.; Sun, H.X.; Zhang, C.; Cheng, L.; Peng, Y.; Deng, Z.Q.; Wang, D.; Wang, Y.; Hu, M.; Liu, W.E.; et al. Prophage Hunter: An integrative hunting tool for active prophages. Nucleic Acids Res. 2019, 47, W74–W80. [Google Scholar] [CrossRef] [PubMed]
- Blattner, F.R.; Plunkett, G.; Bloch, C.A.; Perna, N.T.; Burland, V.; Riley, M.; Collado-Vides, J.; Glasner, J.D.; Rode, C.K.; Mayhew, G.F.; et al. The complete genome sequence of Escherichia coli K-12. Science 1997, 277, 1453–1642. [Google Scholar] [CrossRef]
- Lederberg, E.M.; Lederberg, J. Genetic Studies of Lysogenicity in Escherichia-Coli. Genetics 1953, 38, 51–64. [Google Scholar] [CrossRef]
- Rueggeberg, K.G.; Toba, F.A.; Bird, J.G.; Franck, N.; Thompson, M.G.; Hay, A.G. The lysis cassette of DLP12 defective prophage is regulated by RpoE. Microbiology 2015, 161, 1683–1693. [Google Scholar] [CrossRef]
- Stern, A.; Sorek, R. The phage-host arms race: Shaping the evolution of microbes. Bioessays 2011, 33, 43–51. [Google Scholar] [CrossRef]
- Obeng, N.; Pratama, A.A.; van Elsas, J.D. The Significance of Mutualistic Phages for bacterial Ecology and Evolution. Trends Microbiol. 2016, 24, 440–449. [Google Scholar] [CrossRef]
- Labrie, S.J.; Samson, J.E.; Moineau, S. Bacteriophage resistance mechanisms. Nat. Rev. Microbiol. 2010, 8, 317–327. [Google Scholar] [CrossRef]
- Casjens, S. Prophages and bacterial genomics: What have we learned so far? Mol. Microbiol. 2003, 49, 277–300. [Google Scholar] [CrossRef]
- Chamakura, K.; Young, R. Phage single-gene lysis: Finding the weak spot in the bacterial cell wall. J. Biol. Chem. 2019, 294, 3350–3358. [Google Scholar] [CrossRef] [PubMed]
- Wang, I.N.; Smith, D.L.; Young, R. Holins: The protein clocks of bacteriophage infections. Annu. Rev. Microbiol. 2000, 54, 799–825. [Google Scholar] [CrossRef] [PubMed]
- Zampara, A.; Sørensen, M.C.H.; Grimon, D.; Antenucci, F.; Vitt, A.R.; Bortolaia, V.; Briers, Y.; Brøndsted, L. Exploiting phage receptor binding proteins to enable endolysins to kill Gram-negative bacteria. Sci. Rep. 2020, 10, 12087. [Google Scholar] [CrossRef] [PubMed]
- Reed, C.A.; Langlais, C.; Kuznetsov, V.; Young, R. Inhibitory mechanism of the Qβ lysis protein A2. Mol. Microbiol. 2012, 86, 836–844. [Google Scholar] [CrossRef] [PubMed]
- Bernhardt, T.G.; Struck, D.K.; Young, R. The lysis protein E of phi X174 is a specific inhibitor of the MraY-catalyzed step in peptidoglycan synthesis. J. Biol. Chem. 2001, 276, 6093–6097. [Google Scholar] [CrossRef]
- Chamakura, K.R.; Sham, L.T.; Davis, R.M.; Min, L.; Cho, H.; Ruiz, N.; Bernhardt, T.G.; Young, R. A viral protein antibiotic inhibits lipid II flippase activity. Nat. Microbiol. 2017, 2, 1480–1484. [Google Scholar] [CrossRef]
- Espion, D.; Kaiser, K.; Dambly-Chaudiere, C. A third defective lambdoid prophage of Escherichia coli K12 defined by the lambda derivative, lambdaqin111. J. Mol. Biol. 1983, 170, 611–633. [Google Scholar] [CrossRef]
- Béjar, S.; Cam, K.; Bouché, J.P. Control of cell division in Escherichia coli. DNA sequence of dicA and of a second gene complementing mutation dicA1, dicC. Nucleic Acids Res. 1986, 14, 6821–6833. [Google Scholar] [CrossRef]
- Chen, Y.; Milam, S.L.; Erickson, H.P. SulA inhibits assembly of FtsZ by a simple sequestration mechanism. Biochemistry 2012, 51, 3100–3109. [Google Scholar] [CrossRef]
- Johnson, J.E.; Lackner, L.L.; de Boer, P.A. Targeting of (D)MinC/MinD and (D)MinC/DicB complexes to septal rings in Escherichia coli suggests a multistep mechanism for MinC-mediated destruction of nascent FtsZ rings. J. Bacteriol. 2002, 184, 2951–2962. [Google Scholar] [CrossRef]
- Masuda, H.; Awano, N.; Inouye, M. ydfD encodes a novel lytic protein in Escherichia coli. FEMS Microbiol. Lett. 2016, 363, fnw039. [Google Scholar] [CrossRef][Green Version]
- Hecht, S.; Eisenreich, W.; Adam, P.; Amslinger, S.; Kis, K.; Bacher, A.; Arigoni, D.; Rohdich, F. Studies on the nonmevalonate pathway to terpenes: The role of the GcpE (IspG) protein. Proc. Natl. Acad. Sci. USA 2001, 98, 14837–14842. [Google Scholar] [CrossRef] [PubMed]
- Rohmer, M. From Molecular Fossils of Bacterial Hopanoids to the Formation of Isoprene Units: Discovery and Elucidation of the Methylerythritol Phosphate Pathway. Lipids 2008, 43, 1095–1107. [Google Scholar] [CrossRef] [PubMed]
- Odom, A.R. Five questions about non-mevalonate isoprenoid biosynthesis. PLoS Pathog. 2011, 7, e1002323. [Google Scholar] [CrossRef] [PubMed]
- Anderson, R.G.; Hussey, H.; Baddiley, J. The mechanism of wall synthesis in bacteria. The organization of enzymes and isoprenoid phosphates in the membrane. Biochem. J. 1972, 127, 11–25. [Google Scholar] [CrossRef]
- Bouhss, A.; Trunkfield, A.E.; Bugg, T.D.; Mengin-Lecreulx, D. The biosynthesis of peptidoglycan lipid-linked intermediates. FEMS Microbiol. Rev. 2008, 32, 208–233. [Google Scholar] [CrossRef]
- Wang, X.; Dowd, C.S. The Methylerythritol Phosphate Pathway: Promising Drug Targets in the Fight against Tuberculosis. ACS Infect. Dis. 2018, 4, 278–290. [Google Scholar] [CrossRef] [PubMed]
- He, L.; He, P.; Luo, X.; Li, M.; Yu, L.; Guo, J.; Zhan, X.; Zhu, G.; Zhao, J. The MEP pathway in Babesia orientalis apicoplast, a potential target for anti-babesiosis drug development. Parasites Vectors 2018, 11, 452. [Google Scholar] [CrossRef]
- Rohmer, M.; Grosdemange-Billiard, C.; Seemann, M.; Tritsch, D. Isoprenoid biosynthesis as a novel target for antibacterial and antiparasitic drugs. Curr. Opin. Investig. Drugs 2004, 5, 154–162. [Google Scholar]
- Hunter, W.N. The non-mevalonate pathway of isoprenoid precursor biosynthesis. J. Biol. Chem. 2007, 282, 21573–21577. [Google Scholar] [CrossRef]
- Masini, T.; Hirsch, A.K. Development of inhibitors of the 2-C-methyl-D-erythritol 4-phosphate (MEP) pathway enzymes as potential anti-infective agents. J. Med. Chem. 2014, 57, 9740–9763. [Google Scholar] [CrossRef] [PubMed]
- Frank, A.; Groll, M. The Methylerythritol Phosphate Pathway to Isoprenoids. Chem. Rev. 2017, 117, 5675–5703. [Google Scholar] [CrossRef] [PubMed]
- Patel, H.; Nemeria, N.S.; Brammer, L.A.; Freel Meyers, C.L.; Jordan, F. Observation of Thiamin-Bound Intermediates and Microscopic Rate Constants for Their Interconversion on 1-Deoxy-d-xylulose 5-Phosphate Synthase: 600-Fold Rate Acceleration of Pyruvate Decarboxylation by d-Glyceraldehyde-3-phosphate. J. Am. Chem. Soc. 2012, 134, 18374–18379. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Yang, L.; DeColli, A.; Freel Meyers, C.; Nemeria, N.S.; Jordan, F. Conformational dynamics of 1-deoxy-d-xylulose 5-phosphate synthase on ligand binding revealed by H/D exchange MS. Proc. Natl. Acad. Sci. USA 2017, 114, 9355–9360. [Google Scholar] [CrossRef] [PubMed]
- Masini, T.; Kroezen, B.S.; Hirsch, A.K. Druggability of the enzymes of the non-mevalonate-pathway. Drug Discov. Today 2013, 18, 1256–1262. [Google Scholar] [CrossRef]
- Borrmann, S.; Lundgren, I.; Oyakhirome, S.; Impouma, B.; Matsiegui, P.B.; Adegnika, A.A.; Issifou, S.; Kun, J.F.; Hutchinson, D.; Wiesner, J.; et al. Fosmidomycin plus clindamycin for treatment of pediatric patients aged 1 to 14 years with Plasmodium falciparum malaria. Antimicrob. Agents Chemother. 2006, 50, 2713–2718. [Google Scholar] [CrossRef]
- Missinou, M.A.; Borrmann, S.; Schindler, A.; Issifou, S.; Adegnika, A.A.; Matsiegui, P.-B.; Binder, R.; Lell, B.; Wiesner, J.; Baranek, T.; et al. Fosmidomycin for malaria. Lancet 2002, 360, 1941–1942. [Google Scholar] [CrossRef]
- Borrmann, S.; Adegnika, A.A.; Moussavou, F.; Oyakhirome, S.; Esser, G.; Matsiegui, P.B.; Ramharter, M.; Lundgren, I.; Kombila, M.; Issifou, S.; et al. Short-course regimens of artesunate-fosmidomycin in treatment of uncomplicated Plasmodium falciparum malaria. Antimicrob. Agents Chemother. 2005, 49, 3749–3754. [Google Scholar] [CrossRef]
- Wang, W.; Oldfield, E. Bioorganometallic chemistry with IspG and IspH: Structure, function, and inhibition of the [Fe4S4] proteins involved in isoprenoid biosynthesis. Angew. Chem. Int. Ed. 2014, 53, 4294–4310. [Google Scholar] [CrossRef]
- Baba, T.; Ara, T.; Hasegawa, M.; Takai, Y.; Okumura, Y.; Baba, M.; Datsenko, K.A.; Tomita, M.; Wanner, B.L.; Mori, H. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: The Keio collection. Mol. Syst. Biol. 2006, 2, 2006–2008. [Google Scholar] [CrossRef]
- Campos, N.; Rodriguez-Concepcion, M.; Seemann, M.; Rohmer, M.; Boronat, A. Identification of gcpE as a novel gene of the 2-C-methyl-D-erythritol 4-phosphate pathway for isoprenoid biosynthesis in Escherichia coli. FEBS Lett. 2001, 488, 170–173. [Google Scholar] [CrossRef]
- Sauret-Gueto, S.; Ramos-Valdivia, A.; Ibanez, E.; Boronat, A.; Rodriguez-Concepcion, M. Identification of lethal mutations in Escherichia coli genes encoding enzymes of the methylerythritol phosphate pathway. Biochem. Biophys. Res. Commun. 2003, 307, 408–415. [Google Scholar] [CrossRef]
- Buchan, D.W.A.; Jones, D.T. The PSIPRED Protein Analysis Workbench: 20 years on. Nucleic Acids Res. 2019, 47, W402–W407. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.Y.; Zhang, Y. I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Res. 2015, 43, W174–W181. [Google Scholar] [CrossRef]
- Blaskovich, M.A.T.; Hansford, K.A.; Gong, Y.; Butler, M.S.; Muldoon, C.; Huang, J.X.; Ramu, S.; Silva, A.B.; Cheng, M.; Kavanagh, A.M.; et al. Protein-inspired antibiotics active against vancomycin- and daptomycin-resistant bacteria. Nat. Commun. 2018, 9, 22. [Google Scholar] [CrossRef]
- Miura, K. An Overview of Current Methods to Confirm Protein-Protein Interactions. Protein Pept. Lett. 2018, 25, 728–733. [Google Scholar] [CrossRef]
- Kondo, S.; Ichikawa, H.; Iwo, K.; Kato, T. Base-change mutagenesis and prophage induction in strains of Escherichia coli with different DNA repair capacities. Genetics 1970, 66, 187–217. [Google Scholar] [CrossRef]
- Nanda Arun, M.; Thormann, K.; Frunzke, J.; Margolin, W. Impact of Spontaneous Prophage Induction on the Fitness of Bacterial Populations and Host-Microbe Interactions. J. Bacteriol. 2015, 197, 410–419. [Google Scholar] [CrossRef]
- Barnhart, B.J.; Cox, S.H.; Jett, J.H. Prophage induction and inactivation by UV light. J. Virol. 1976, 18, 950–955. [Google Scholar] [CrossRef]
- Wang, X.X.; Kim, Y.; Ma, Q.; Hong, S.H.; Pokusaeva, K.; Sturino, J.M.; Wood, T.K. Cryptic prophages help bacteria cope with adverse environments. Nat. Commun. 2010, 1, 147. [Google Scholar] [CrossRef]
- Cam, K.; Béjar, S.; Gil, D.; Bouché, J.P. Identification and sequence of gene dicB: Translation of the division inhibitor from an in-phase internal start. Nucleic Acids Res. 1988, 16, 6327–6338. [Google Scholar] [CrossRef] [PubMed]
- Ragunathan, P.T.; Vanderpool, C.K. Cryptic-Prophage-Encoded Small Protein DicB Protects Escherichia coli from Phage Infection by Inhibiting Inner Membrane Receptor Proteins. J. Bacteriol. 2019, 201, e00475-19. [Google Scholar] [CrossRef] [PubMed]
- Guerra, F.; Wang, K.; Li, J.K.; Wang, W.X.; Liu, Y.L.; Amin, S.; Oldfield, E. Inhibition of the 4Fe-4S proteins IspG and IspH: An EPR, ENDOR and HYSCORE investigation. Chem. Sci. 2014, 5, 1642–1649. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Li, J.; Wang, K.; Huang, C.; Zhang, Y.; Oldfield, E. Organometallic mechanism of action and inhibition of the 4Fe-4S isoprenoid biosynthesis protein GcpE (IspG). Proc. Natl. Acad. Sci. USA 2010, 107, 11189–11193. [Google Scholar] [CrossRef] [PubMed]
- Zepeck, F.; Gräwert, T.; Kaiser, J.; Schramek, N.; Eisenreich, W.; Bacher, A.; Rohdich, F. Biosynthesis of Isoprenoids. Purification and Properties of IspG Protein from Escherichia coli. J. Org. Chem. 2005, 70, 9168–9174. [Google Scholar] [CrossRef]
- Quitterer, F.; Frank, A.; Wang, K.; Rao, G.; O’Dowd, B.; Li, J.; Guerra, F.; Abdel-Azeim, S.; Bacher, A.; Eppinger, J.; et al. Atomic-Resolution Structures of Discrete Stages on the Reaction Coordinate of the [Fe4S4] Enzyme IspG (GcpE). J. Mol. Biol. 2015, 427, 2220–2228. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.; Gräwert, T.; Quitterer, F.; Rohdich, F.; Eppinger, J.; Eisenreich, W.; Bacher, A.; Groll, M. Biosynthesis of isoprenoids: Crystal structure of the [4Fe-4S] cluster protein IspG. J. Mol. Biol. 2010, 404, 600–610. [Google Scholar] [CrossRef] [PubMed]
- Gasteiger, E.H.C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Walker, J.M., Ed.; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. [Google Scholar]
- Oppenheim, A.B.; Kobiler, O.; Stavans, J.; Court, D.L.; Adhya, S. Switches in bacteriophage lambda development. Annu. Rev. Genet. 2005, 39, 409–429. [Google Scholar] [CrossRef]
- Erez, Z.; Steinberger-Levy, I.; Shamir, M.; Doron, S.; Stokar-Avihail, A.; Peleg, Y.; Melamed, S.; Leavitt, A.; Savidor, A.; Albeck, S.; et al. Communication between viruses guides lysis-lysogeny decisions. Nature 2017, 541, 488–493. [Google Scholar] [CrossRef]
- Zong, C.H.; So, L.H.; Sepulveda, L.A.; Skinner, S.O.; Golding, I. Lysogen stability is determined by the frequency of activity bursts from the fate-determining gene. Mol. Syst. Biol. 2010, 6, 440. [Google Scholar] [CrossRef]
- Hickey, E.W.; Hirshfield, I.N. Low-pH-induced effects on patterns of protein synthesis and on internal pH in Escherichia coli and Salmonella typhimurium. Appl. Environ. Microbiol. 1990, 56, 1038–1045. [Google Scholar] [CrossRef] [PubMed]
- Booth, I.R. Regulation of cytoplasmic pH in bacteria. Microbiol. Rev. 1985, 49, 359–378. [Google Scholar] [CrossRef] [PubMed]
- Sandoval-Calderón, M.; Guan, Z.; Sohlenkamp, C. Knowns and unknowns of membrane lipid synthesis in streptomycetes. Biochimie 2017, 141, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, A.; Fontes, F.L.; Gonzalez-Juarrero, M.; McNeil, M.R.; Crans, D.C.; Jackson, M.; Crick, D.C. Partial Saturation of Menaquinone in Mycobacterium tuberculosis: Function and Essentiality of a Novel Reductase, MenJ. ACS Cent. Sci. 2015, 1, 292–302. [Google Scholar] [CrossRef]
- Crick, D.C.; Schulbach, M.C.; Zink, E.E.; Macchia, M.; Barontini, S.; Besra, G.S.; Brennan, P.J. Polyprenyl phosphate biosynthesis in Mycobacterium tuberculosis and Mycobacterium smegmatis. J. Bacteriol. 2000, 182, 5771–5778. [Google Scholar] [CrossRef]
- Bongers, M.; Chrysanthopoulos, P.K.; Behrendorff, J.B.Y.H.; Hodson, M.P.; Vickers, C.E.; Nielsen, L.K. Systems analysis of methylerythritol-phosphate pathway flux in E. coli: Insights into the role of oxidative stress and the validity of lycopene as an isoprenoid reporter metabolite. Microb. Cell Factories 2015, 14, 193. [Google Scholar] [CrossRef]
- Chang, S.-Y.; Ko, T.-P.; Liang, P.-H.; Wang, A.H.J. Catalytic Mechanism Revealed by the Crystal Structure of Undecaprenyl Pyrophosphate Synthase in Complex with Sulfate, Magnesium, and Triton. J. Biol. Chem. 2003, 278, 29298–29307. [Google Scholar] [CrossRef]
- Jiang, D.; Jiang, Z.; Lu, D.; Wang, X.; Liang, H.; Zhang, J.; Meng, Y.; Li, Y.; Wu, D.; Huang, Y.; et al. Migrasomes provide regional cues for organ morphogenesis during zebrafish gastrulation. Nat. Cell Biol. 2019, 21, 966–977. [Google Scholar] [CrossRef]
- Chakravorty, S.; Helb, D.; Burday, M.; Connell, N.; Alland, D. A detailed analysis of 16S ribosomal RNA gene segments for the diagnosis of pathogenic bacteria. J. Microbiol. Methods 2007, 69, 330–339. [Google Scholar] [CrossRef]
- Yang, S.; Pei, H.; Zhang, X.; Wei, Q.; Zhu, J.; Zheng, J.; Jia, Z. Characterization of DicB by partially masking its potent inhibitory activity of cell division. Open Biol. 2016, 6, 160082. [Google Scholar] [CrossRef]

| Accession | Protein | Physiological Function or Processes | Interaction with YdfD |
|---|---|---|---|
| A0A029II96 | GlmS | Hexosamine biosynthesis | No |
| E1JEU9 | ErpA | A-type iron–sulfur carrier | No |
| A7ZNX6 | FolE | Tetrahydrofolate biosynthesis | No |
| A0A070S6E1 | IspG | MEP pathway | Yes |
| A0A017JTH0 | IscS | Cysteine desulfurase | No |
| D6I572 | LpxA | Lipid biosynthesis | No |
| M8THF9 | CydA | Ubiquinol oxidase | No |
| A0A017IFA8 | KdsA | Lipopolysaccharide biosynthesis | No |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, Z.; Wang, B.; Qiu, Z.; Zhang, R.; Zheng, J.; Jia, Z. YdfD, a Lysis Protein of the Qin Prophage, Is a Specific Inhibitor of the IspG-Catalyzed Step in the MEP Pathway of Escherichia coli. Int. J. Mol. Sci. 2022, 23, 1560. https://doi.org/10.3390/ijms23031560
Lu Z, Wang B, Qiu Z, Zhang R, Zheng J, Jia Z. YdfD, a Lysis Protein of the Qin Prophage, Is a Specific Inhibitor of the IspG-Catalyzed Step in the MEP Pathway of Escherichia coli. International Journal of Molecular Sciences. 2022; 23(3):1560. https://doi.org/10.3390/ijms23031560
Chicago/Turabian StyleLu, Zhifang, Biying Wang, Zhiyu Qiu, Ruiling Zhang, Jimin Zheng, and Zongchao Jia. 2022. "YdfD, a Lysis Protein of the Qin Prophage, Is a Specific Inhibitor of the IspG-Catalyzed Step in the MEP Pathway of Escherichia coli" International Journal of Molecular Sciences 23, no. 3: 1560. https://doi.org/10.3390/ijms23031560
APA StyleLu, Z., Wang, B., Qiu, Z., Zhang, R., Zheng, J., & Jia, Z. (2022). YdfD, a Lysis Protein of the Qin Prophage, Is a Specific Inhibitor of the IspG-Catalyzed Step in the MEP Pathway of Escherichia coli. International Journal of Molecular Sciences, 23(3), 1560. https://doi.org/10.3390/ijms23031560







