Deep-4mCGP: A Deep Learning Approach to Predict 4mC Sites in Geobacter pickeringii by Using Correlation-Based Feature Selection Technique
Abstract
:1. Introduction
2. Results and Discussion
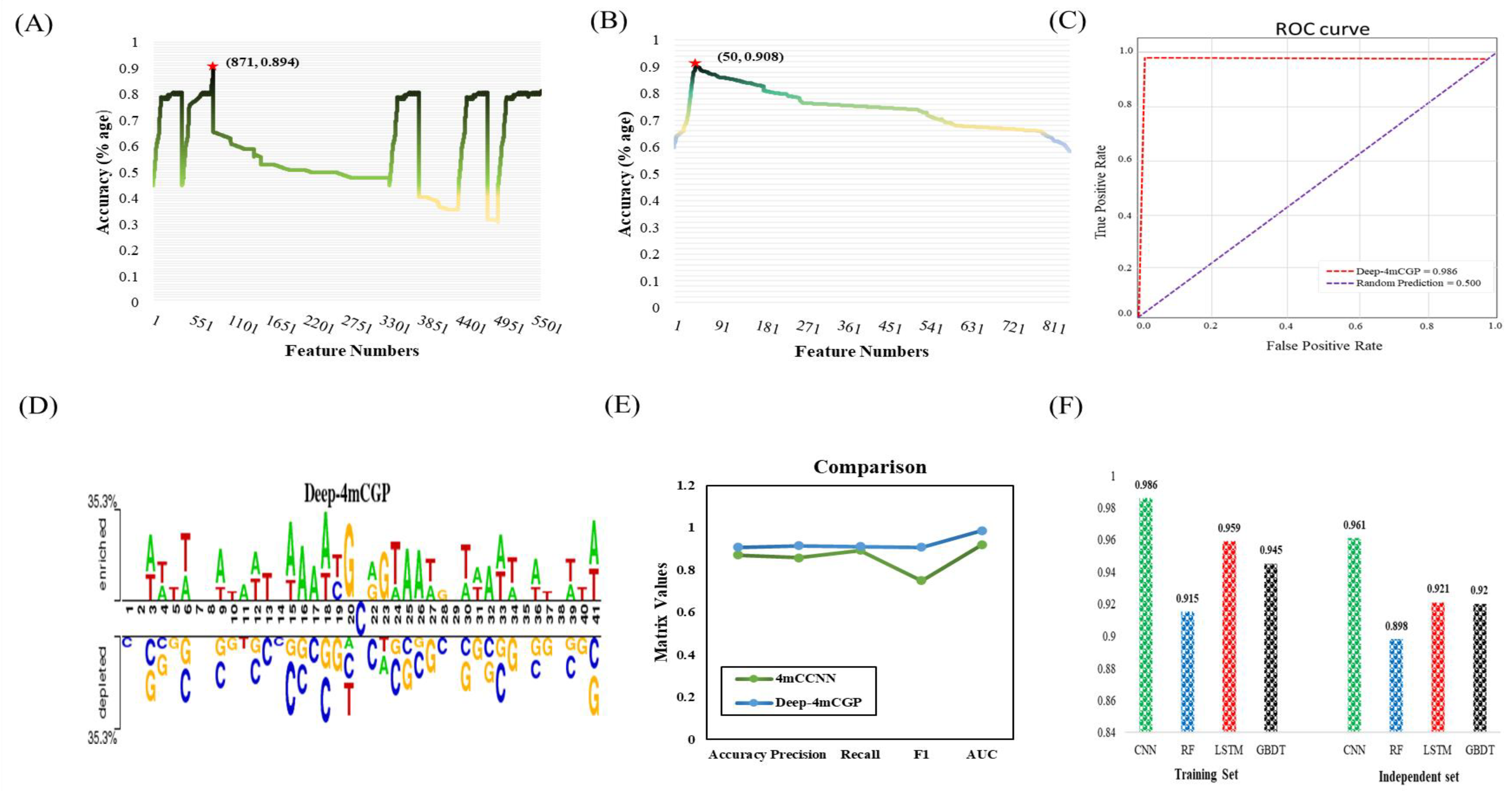
2.1. Performance Evaluation
2.2. Sequence Composition Analysis
2.3. Comparison on the Basis of Independent Data
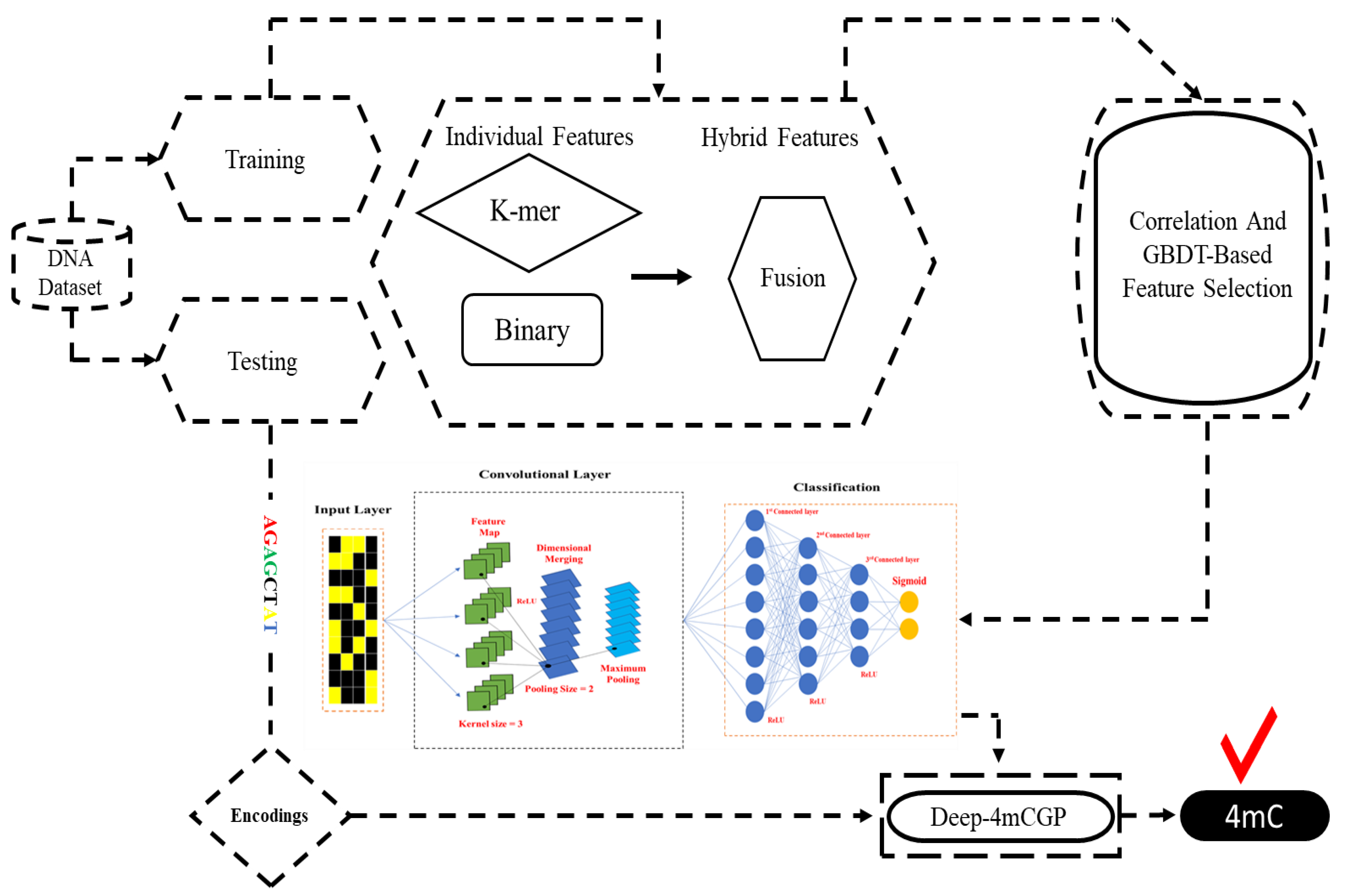
3. Materials and Methods
3.1. Feature Descriptors
3.1.1. k-mer
3.1.2. Binary
3.2. Feature Selection
3.2.1. Correlation
3.2.2. GBDT with IFS
| Algorithms 1: Correlation and GBDT-based Feature Selection Algorithm |
| Input: Training Data: = Q (L1, L2, ……, Lk, Lc) Output: Qbest 1st Round 1 Begin 2 for i = 1 to k do 3 r = calculate correlational coefficient (Li, Lc) 4 end 5 p = 0.05 6 ρ= 0 (⸫ if there is no correlation among the Fi and Fc) 7 for i = 1 to k do 8 t = to calculate the significance (r, ρ) for Li (⸫ by utilizing the t-test value from Equation (5)) 9 if t > critical value 10 Qbest = Q list 11 end 12 return Qbest 2nd Round Input: Qbest: = Where, ( = data and = label) LF: = P (, q ()) 13 By initializing the model 14 : = argument minimum 15 for I = {1, 2, 3, 4, 5…, n} do 16 for k = {1, 2, 3, 4, 5…, K} do 17 Pseudo residual error calculations: = 18 end 19 end 20 On the basis of , = {}, we built a decision tree 21 for j = {1, 2, 3, 4, 5…., J} do 22 = argument minimum 23 end 24 Updating the model = + 25 q (x) = Output: The decision tree function q (x) |
3.3. Convolutional Neural Network
3.4. Metrics Evaluation
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Schübeler, D. Function and information content of DNA methylation. Nature 2015, 517, 321–326. [Google Scholar] [CrossRef]
- Ao, C.; Yu, L.; Zou, Q. Prediction of bio-sequence modifications and the associations with diseases. Brief. Funct. Genom. 2021, 20, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Pataillot-Meakin, T.; Pillay, N.; Beck, S. 3-methylcytosine in cancer: An underappreciated methyl lesion? Epigenomics 2016, 8, 451–454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yalcin, D.; Otu, H.H. An Unbiased Predictive Model to Detect DNA Methylation Propensity of CpG Islands in the Human Genome. Curr. Bioinform. 2021, 16, 179–196. [Google Scholar] [CrossRef]
- Robertson, K.D. DNA methylation and human disease. Nat. Rev. Genet. 2005, 6, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Iyer, L.M.; Abhiman, S.; Aravind, L. Natural history of eukaryotic DNA methylation systems. Prog. Mol. Biol. Transl. Sci. 2011, 101, 25–104. [Google Scholar] [PubMed]
- Flusberg, B.A.; Webster, D.R.; Lee, J.H.; Travers, K.J.; Olivares, E.C.; Clark, T.A.; Korlach, J.; Turner, S.W. Direct detection of DNA methylation during single-molecule, real-time sequencing. Nat. Methods 2010, 7, 461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Doherty, R.; Couldrey, C. Exploring genome wide bisulfite sequencing for DNA methylation analysis in livestock: A technical assessment. Front. Genet. 2014, 5, 126. [Google Scholar] [CrossRef] [Green Version]
- Boch, J.; Bonas, U. Xanthomonas AvrBs3 family-type III effectors: Discovery and function. Annu. Rev. Phytopathol. 2010, 48, 419–436. [Google Scholar] [CrossRef]
- Chen, W.; Yang, H.; Feng, P.; Ding, H.; Lin, H. iDNA4mC: Identifying DNA N4-methylcytosine sites based on nucleotide chemical properties. Bioinformatics 2017, 33, 3518–3523. [Google Scholar] [CrossRef]
- Wei, L.; Su, R.; Luan, S.; Liao, Z.; Manavalan, B.; Zou, Q.; Shi, X. Iterative feature representations improve N4-methylcytosine site prediction. Bioinformatics 2019, 35, 4930–4937. [Google Scholar] [CrossRef]
- Tang, Q.; Kang, J.; Yuan, J.; Tang, H.; Li, X.; Lin, H.; Huang, J.; Chen, W. DNA4mC-LIP: A linear integration method to identify N4-methylcytosine site in multiple species. Bioinformatics 2020, 36, 3327–3335. [Google Scholar] [CrossRef] [PubMed]
- Manavalan, B.; Basith, S.; Shin, T.H.; Wei, L.; Lee, G. Meta-4mCpred: A Sequence-Based Meta-Predictor for Accurate DNA 4mC Site Prediction Using Effective Feature Representation. Mol. Ther.-Nucleic Acids 2019, 16, 733–744. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khanal, J.; Nazari, I.; Tayara, H.; Chong, K.T. 4mCCNN: Identification of N4-methylcytosine sites in prokaryotes using convolutional neural network. IEEE Access 2019, 7, 145455–145461. [Google Scholar] [CrossRef]
- Manavalan, B.; Basith, S.; Shin, T.H.; Lee, D.Y.; Wei, L.; Lee, G. 4mCpred-EL: An ensemble learning framework for identification of DNA N4-methylcytosine sites in the mouse genome. Cells 2019, 8, 1332. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hasan, M.M.; Manavalan, B.; Shoombuatong, W.; Khatun, M.S.; Kurata, H. i4mC-Mouse: Improved identification of DNA N4-methylcytosine sites in the mouse genome using multiple encoding schemes. Comput. Struct. Biotechnol. J. 2020, 18, 906–912. [Google Scholar] [CrossRef] [PubMed]
- Zulfiqar, H.; Khan, R.S.; Hassan, F.; Hippe, K.; Hunt, C.; Ding, H.; Song, X.-M.; Cao, R. Computational identification of N4-methylcytosine sites in the mouse genome with machine-learning method. Math. Biosci. Eng. 2021, 18, 3348–3363. [Google Scholar] [CrossRef]
- Ye, P.; Luan, Y.; Chen, K.; Liu, Y.; Xiao, C.; Xie, Z. MethSMRT: An integrative database for DNA N6-methyladenine and N4-methylcytosine generated by single-molecular real-time sequencing. Nucleic Acids Res. 2016, 45, D85–D89. [Google Scholar] [CrossRef] [Green Version]
- Smith, Z.D.; Meissner, A. DNA methylation: Roles in mammalian development. Nat. Rev. Genet. 2013, 14, 204–220. [Google Scholar] [CrossRef]
- Vacic, V.; Iakoucheva, L.M.; Radivojac, P. Two Sample Logo: A graphical representation of the differences between two sets of sequence alignments. Bioinformatics 2006, 22, 1536–1537. [Google Scholar] [CrossRef] [Green Version]
- Gers, F.A.; Schmidhuber, J.; Cummins, F. Learning to forget: Continual prediction with LSTM. Neural Comput. 2000, 12, 2451–2471. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Chow, J.-H.; Chen, J.; Zheng, Z. Stochastic gradient boosted distributed decision trees. In Proceedings of the 18th ACM Conference on Information and Knowledge Management, Hong Kong, China, 2–6 November 2009; pp. 2061–2064. [Google Scholar]
- Qi, Y. Random forest for bioinformatics. In Ensemble Machine Learning; Springer: Berlin/Heidelberg, Germany, 2012; pp. 307–323. [Google Scholar]
- Ahmed, F.F.; Khatun, M.S.; Mosharaf, M.P.; Mollah, M.N.H. Prediction of Protein-protein Interactions in Arabidopsis thaliana Using Partial Training Samples in a Machine Learning Framework. Curr. Bioinform. 2021, 16, 865–879. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, Y.; Wang, R.; Lu, J.; Ma, X.; Qiu, M. PSAC: Proactive Sequence-aware Content Caching via Deep Learning at the Network Edge. IEEE Trans. Netw. Sci. Eng. 2020, 7, 2145–2154. [Google Scholar] [CrossRef]
- Su, W.; Liu, M.L.; Yang, Y.H.; Wang, J.S.; Li, S.H.; Lv, H.; Dao, F.Y.; Yang, H.; Lin, H. PPD: A Manually Curated Database for Experimentally Verified Prokaryotic Promoters. J. Mol. Biol. 2021, 433, 166860. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.K.; Srivastava, R. Protein Secondary Structure Prediction Using Character bi-gram Embedding and Bi-LSTM. Curr. Bioinform. 2021, 16, 333–338. [Google Scholar] [CrossRef]
- Hasan, M.M.; Alam, M.A.; Shoombuatong, W.; Deng, H.W.; Manavalan, B.; Kurata, H. NeuroPred-FRL: An interpretable prediction model for identifying neuropeptide using feature representation learning. Brief. Bioinform. 2021, 22, bbab167. [Google Scholar] [CrossRef] [PubMed]
- Charoenkwan, P.; Chiangjong, W.; Nantasenamat, C.; Hasan, M.M.; Manavalan, B.; Shoombuatong, W. StackIL6: A stacking ensemble model for improving the prediction of IL-6 inducing peptides. Brief. Bioinform. 2021, 22, bbab172. [Google Scholar] [CrossRef] [PubMed]
- Zulfiqar, H.; Sun, Z.J.; Huang, Q.L.; Yuan, S.S.; Lv, H.; Dao, F.Y.; Lin, H.; Li, Y.W. Deep-4mCW2V: A sequence-based predictor to identify N4-methylcytosine sites in Escherichia coli. Methods 2021, in press. [Google Scholar] [CrossRef]
- Ju, Z.; Wang, S.-Y. Prediction of Neddylation Sites Using the Composition of k-spaced Amino Acid Pairs and Fuzzy SVM. Curr. Bioinform. 2020, 15, 725–731. [Google Scholar] [CrossRef]
- Zhang, D.; Chen, H.-D.; Zulfiqar, H.; Yuan, S.-S.; Huang, Q.-L.; Zhang, Z.-Y.; Deng, K.-J. iBLP: An XGBoost-based predictor for identifying bioluminescent proteins. Comput. Math. Methods Med. 2021, 2021, 6664362. [Google Scholar] [CrossRef]
- Lv, H.; Dao, F.-Y.; Zulfiqar, H.; Lin, H. DeepIPs: Comprehensive assessment and computational identification of phosphorylation sites of SARS-CoV-2 infection using a deep learning-based approach. Brief. Bioinform. 2021, 22, bbab244. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Huang, Z.; Kong, L. CSBPI Site: Multi-Information Sources of Features to RNA Binding Sites Prediction. Curr. Bioinform. 2021, 16, 691–699. [Google Scholar] [CrossRef]
- Lv, H.; Shi, L.; Berkenpas, J.W.; Dao, F.-Y.; Zulfiqar, H.; Ding, H.; Zhang, Y.; Yang, L.; Cao, R. Application of artificial intelligence and machine learning for COVID-19 drug discovery and vaccine design. Brief. Bioinform. 2021, 22, bbab320. [Google Scholar] [CrossRef]
- Zulfiqar, H.; Masoud, M.S.; Yang, H.; Han, S.G.; Wu, C.Y.; Lin, H. Screening of prospective plant compounds as H1R and CL1R inhibitors and its antiallergic efficacy through molecular docking approach. Comput. Math. Methods Med. 2021, 2021, 6683407. [Google Scholar] [CrossRef]
- Hasan, M.M.; Schaduangrat, N.; Basith, S.; Lee, G.; Shoombuatong, W.; Manavalan, B. HLPpred-Fuse: Improved and robust prediction of hemolytic peptide and its activity by fusing multiple feature representation. Bioinformatics 2020, 36, 3350–3356. [Google Scholar] [CrossRef]
- Govindaraj, R.G.; Subramaniyam, S.; Manavalan, B. Extremely-randomized-tree-based Prediction of N(6)-Methyladenosine Sites in Saccharomyces cerevisiae. Curr. Genom. 2020, 21, 26–33. [Google Scholar] [CrossRef]
- Li, Q.; Yu, J.; Yan, Y.; Chen, Y.; Tan, S. PsePSSM-based Prediction for the Protein-ATP Binding Sites. Curr. Bioinform. 2021, 16, 576–582. [Google Scholar]
- Dao, F.-Y.; Lv, H.; Zulfiqar, H.; Yang, H.; Su, W.; Gao, H.; Ding, H.; Lin, H. A computational platform to identify origins of replication sites in eukaryotes. Brief. Bioinform. 2021, 22, 1940–1950. [Google Scholar] [CrossRef]
- Lv, H.; Dao, F.-Y.; Zulfiqar, H.; Su, W.; Ding, H.; Liu, L.; Lin, H. A sequence-based deep learning approach to predict CTCF-mediated chromatin loop. Brief. Bioinform. 2021, 22, 1–13. [Google Scholar] [CrossRef]
- Zulfiqar, H.; Yuan, S.-S.; Huang, Q.-L.; Sun, Z.-J.; Dao, F.-Y.; Yu, X.-L.; Lin, H. Identification of cyclin protein using gradient boost decision tree algorithm. Comput. Struct. Biotechnol. J. 2021, 19, 4123–4131. [Google Scholar] [CrossRef]
- Ke, G.; Meng, Q.; Finley, T.; Wang, T.; Chen, W.; Ma, W.; Ye, Q.; Liu, T.-Y. Lightgbm: A highly efficient gradient boosting decision tree. Adv. Neural Inf. Process. Syst. 2017, 30, 3146–3154. [Google Scholar]
- Yang, W.; Zhu, X.-J.; Huang, J.; Ding, H.; Lin, H. A Brief Survey of Machine Learning Methods in Protein Sub-Golgi Localization. Curr. Bioinform. 2019, 14, 234–240. [Google Scholar] [CrossRef]
- Tan, J.-X.; Li, S.-H.; Zhang, Z.-M.; Chen, C.-X.; Chen, W.; Tang, H.; Lin, H. Identification of hormone binding proteins based on machine learning methods. Math. Biosci. Eng. 2019, 16, 2466–2480. [Google Scholar] [CrossRef]
- Alim, A.; Rafay, A.; Naseem, I. PoGB-pred: Prediction of Antifreeze Proteins Sequences Using Amino Acid Composition with Feature Selection Followed by a Sequential-based Ensemble Approach. Curr. Bioinform. 2021, 16, 446–456. [Google Scholar] [CrossRef]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef] [Green Version]
- Niu, M.; Lin, Y.; Zou, Q. sgRNACNN: Identifying sgRNA on-target activity in four crops using ensembles of convolutional neural networks. Plant Mol. Biol. 2021, 105, 483–495. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Yan, J.; Chen, S.; Gong, M.; Gao, D.; Zhu, M.; Gan, W. Review of the Applications of Deep Learning in Bioinformatics. Curr. Bioinform. 2020, 15, 898–911. [Google Scholar] [CrossRef]
- Bukhari, S.A.S.; Razzaq, A.; Jabeen, J.; Khan, S.; Khan, Z. Deep-BSC: Predicting Raw DNA Binding Pattern in Arabidopsis thaliana. Curr. Bioinform. 2021, 16, 457–465. [Google Scholar] [CrossRef]
- Kwon, Y.-H.; Shin, S.-B.; Kim, S.-D. Electroencephalography based fusion two-dimensional (2D)-convolution neural networks (CNN) model for emotion recognition system. Sensors 2018, 18, 1383. [Google Scholar] [CrossRef] [Green Version]
- Mo, F.; Luo, Y.; Fan, D.A.; Zeng, H.; Zhao, Y.N.; Luo, M.; Liu, X.B.; Ma, X.L. Integrated Analysis of mRNA-seq and miRNA-seq to identify c-MYC, YAP1 and miR-3960 as Major Players in the Anticancer Effects of Caffeic Acid Phenethyl Ester in Human Small Cell Lung Cancer Cell Line. Curr. Gene Ther. 2020, 20, 15–24. [Google Scholar] [CrossRef]
- Chollet, F. Keras: Deep learning library for theano and tensorflow. Keras 2015, 7, T1. Available online: https://Keras.Io/ (accessed on 19 January 2022).
- Cao, R.; Freitas, C.; Chan, L.; Sun, M.; Jiang, H.; Chen, Z. ProLanGO: Protein function prediction using neural machine translation based on a recurrent neural network. Molecules 2017, 22, 1732. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gai, D.; Shen, X.; Chen, H. Effective Classification of Melting Curve in Real-time PCR Based on Dynamic Filter-based Convolutional Neural Network. Curr. Bioinform. 2021, 16, 820–828. [Google Scholar] [CrossRef]
- Ao, C.; Zou, Q.; Yu, L. RFhy-m2G: Identification of RNA N2-methylguanosine modification sites based on random forest and hybrid features. Methods 2021, in press. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Jia, C.; Zou, Q. 4mCPred: Machine learning methods for DNA N4-methylcytosine sites prediction. Bioinformatics 2019, 35, 593–601. [Google Scholar] [CrossRef]
- Lv, H.; Dao, F.-Y.; Zhang, D.; Guan, Z.-X.; Yang, H.; Su, W.; Liu, M.-L.; Ding, H.; Chen, W.; Lin, H. iDNA-MS: An integrated computational tool for detecting DNA modification sites in multiple genomes. Iscience 2020, 23, 100991. [Google Scholar] [CrossRef]
- Zulfiqar, H.; Dao, F.Y.; Lv, H.; Yang, H.; Zhou, P.; Chen, W.; Lin, H. Identification of Potential Inhibitors Against SARS-CoV-2 Using Computational Drug Repurposing Study. Curr. Bioinform. 2021, 16, 1320–1327. [Google Scholar] [CrossRef]
- Liu, Q.; Chen, J.; Wang, Y.; Li, S.; Jia, C.; Song, J.; Li, F. DeepTorrent: A deep learning-based approach for predicting DNA N4-methylcytosine sites. Brief. Bioinform. 2021, 22, bbaa124. [Google Scholar] [CrossRef]
| Training Data | Independent Data | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Algorithm | FS | Method | Accuracy | Precision | Recall | F1 | AUROC | Accuracy | Precision | Recall | F1 | AUROC |
| LSTM | 5460 | k-mer | 0.861 | 0.872 | 0.861 | 0.811 | 0.943 | 0.825 | 0.820 | 0.812 | 0.819 | 0.882 |
| 164 | Binary | 0.834 | 0.828 | 0.837 | 0.838 | 0.875 | 0.801 | 0.804 | 0.798 | 0.801 | 0.872 | |
| 5624 | Fusion | 0.868 | 0.865 | 0.859 | 0.862 | 0.937 | 0.810 | 0.814 | 0.808 | 0.813 | 0.902 | |
| 871 | Fusion | 0.859 | 0.857 | 0.847 | 0.857 | 0.925 | 0.808 | 0.801 | 0.807 | 0.800 | 0.876 | |
| 50 | Fusion | 0.884 | 0.878 | 0.881 | 0.879 | 0.959 | 0.841 | 0.842 | 0.839 | 0.842 | 0.921 | |
| RF | 5460 | k-mer | 0.831 | 0.862 | 0.758 | 0.664 | 0.936 | 0.809 | 0.838 | 0.761 | 0.648 | 0.909 |
| 164 | Binary | 0.772 | 0.763 | 0.755 | 0.770 | 0.863 | 0.753 | 0.748 | 0.753 | 0.756 | 0.832 | |
| 5624 | Fusion | 0.844 | 0.847 | 0.839 | 0.845 | 0.891 | 0.795 | 0.788 | 0.783 | 0.794 | 0.887 | |
| 871 | Fusion | 0.847 | 0.849 | 0.851 | 0.846 | 0.897 | 0.801 | 0.800 | 0.800 | 0.798 | 0.878 | |
| 50 | Fusion | 0.866 | 0.858 | 0.861 | 0.854 | 0.915 | 0.812 | 0.808 | 0.814 | 0.812 | 0.898 | |
| GBDT | 5460 | k-mer | 0.848 | 0.881 | 0.776 | 0.676 | 0.962 | 0.828 | 0.861 | 0.770 | 0.669 | 0.931 |
| 164 | Binary | 0.827 | 0.821 | 0.823 | 0.827 | 0.895 | 0.782 | 0.778 | 0.779 | 0.781 | 0.862 | |
| 5624 | Fusion | 0.835 | 0.832 | 0.830 | 0.832 | 0.893 | 0.786 | 0.780 | 0.786 | 0.786 | 0.882 | |
| 871 | Fusion | 0.851 | 0.853 | 0.848 | 0.854 | 0.901 | 0.814 | 0.810 | 0.815 | 0.810 | 0.893 | |
| 50 | Fusion | 0.875 | 0.874 | 0.868 | 0.860 | 0.945 | 0.836 | 0.835 | 0.830 | 0.841 | 0.920 | |
| CNN | 5460 | k-mer | 0.880 | 0.879 | 0.887 | 0.880 | 0.949 | 0.848 | 0.844 | 0.841 | 0.845 | 0.927 |
| 164 | Binary | 0.868 | 0.836 | 0.834 | 0.832 | 0.928 | 0.798 | 0.802 | 0.807 | 0.790 | 0.881 | |
| 5624 | Fusion | 0.868 | 0.865 | 0.859 | 0.862 | 0.937 | 0.810 | 0.814 | 0.808 | 0.813 | 0.903 | |
| 871 | Fusion | 0.894 | 0.877 | 0.897 | 0.889 | 0.955 | 0.846 | 0.845 | 0.841 | 0.838 | 0.920 | |
| 50 | Fusion | 0.908 | 0.914 | 0.910 | 0.908 | 0.986 | 0.868 | 0.876 | 0.773 | 0.859 | 0.961 | |
| Predictor | CV | Accuracy | Precision | Recall | F1 | AUROC | Reference |
|---|---|---|---|---|---|---|---|
| 4mcCNN | 10 (folds) | 0.871 | 0.857 | 0.893 | 0.750 | 0.921 | [14] |
| Deep-4mCGP | 10 (folds) | 0.908 | 0.914 | 0.910 | 0.908 | 0.986 | Deep-4mCGP |
| 4mcCNN | Test (Ind) | 0.826 | 0.818 | 0.823 | 0.825 | 0.920 | [14] |
| Deep-4mCGP | Test (Ind) | 0.868 | 0.876 | 0.773 | 0.859 | 0.961 | Deep-4mCGP |
| Classifier | Parameters |
|---|---|
| RF | N-estimators = 100, Learning-rate = 0.001, Mean absolute error = 0.143, Mean square error = 0.220 |
| GBDT | N-estimators = 120, Learning-rate = 0.01, Mean absolute error = 0.117, Mean square error = 0.212 |
| LSTM | nn.LSTM(input_size = feature_size, hidden_size = 128) nn.Linear(int_features = 128, out_features = 1) nn.Sigmoid() learning-rate = 0.001, Epoch = 100, Batch-size = 32 |
| CNN | nn. Conv1d (in_channels = feature size, out_channels = 32, padding = valid, strides = 1, kernel_size = 2) nn.ReLU() nn.MaxPool 1d (padding = valid, strides = 2, pool_size = 2) nn. Dropout (p = 0.5) nn.Sigmoid() Learning-rate = 0.01, epoch = 80, batch-size = 32 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zulfiqar, H.; Huang, Q.-L.; Lv, H.; Sun, Z.-J.; Dao, F.-Y.; Lin, H. Deep-4mCGP: A Deep Learning Approach to Predict 4mC Sites in Geobacter pickeringii by Using Correlation-Based Feature Selection Technique. Int. J. Mol. Sci. 2022, 23, 1251. https://doi.org/10.3390/ijms23031251
Zulfiqar H, Huang Q-L, Lv H, Sun Z-J, Dao F-Y, Lin H. Deep-4mCGP: A Deep Learning Approach to Predict 4mC Sites in Geobacter pickeringii by Using Correlation-Based Feature Selection Technique. International Journal of Molecular Sciences. 2022; 23(3):1251. https://doi.org/10.3390/ijms23031251
Chicago/Turabian StyleZulfiqar, Hasan, Qin-Lai Huang, Hao Lv, Zi-Jie Sun, Fu-Ying Dao, and Hao Lin. 2022. "Deep-4mCGP: A Deep Learning Approach to Predict 4mC Sites in Geobacter pickeringii by Using Correlation-Based Feature Selection Technique" International Journal of Molecular Sciences 23, no. 3: 1251. https://doi.org/10.3390/ijms23031251
APA StyleZulfiqar, H., Huang, Q.-L., Lv, H., Sun, Z.-J., Dao, F.-Y., & Lin, H. (2022). Deep-4mCGP: A Deep Learning Approach to Predict 4mC Sites in Geobacter pickeringii by Using Correlation-Based Feature Selection Technique. International Journal of Molecular Sciences, 23(3), 1251. https://doi.org/10.3390/ijms23031251







