Systematic Comparison of Plant Promoters in Nicotiana spp. Expression Systems
Abstract
1. Introduction
2. Results
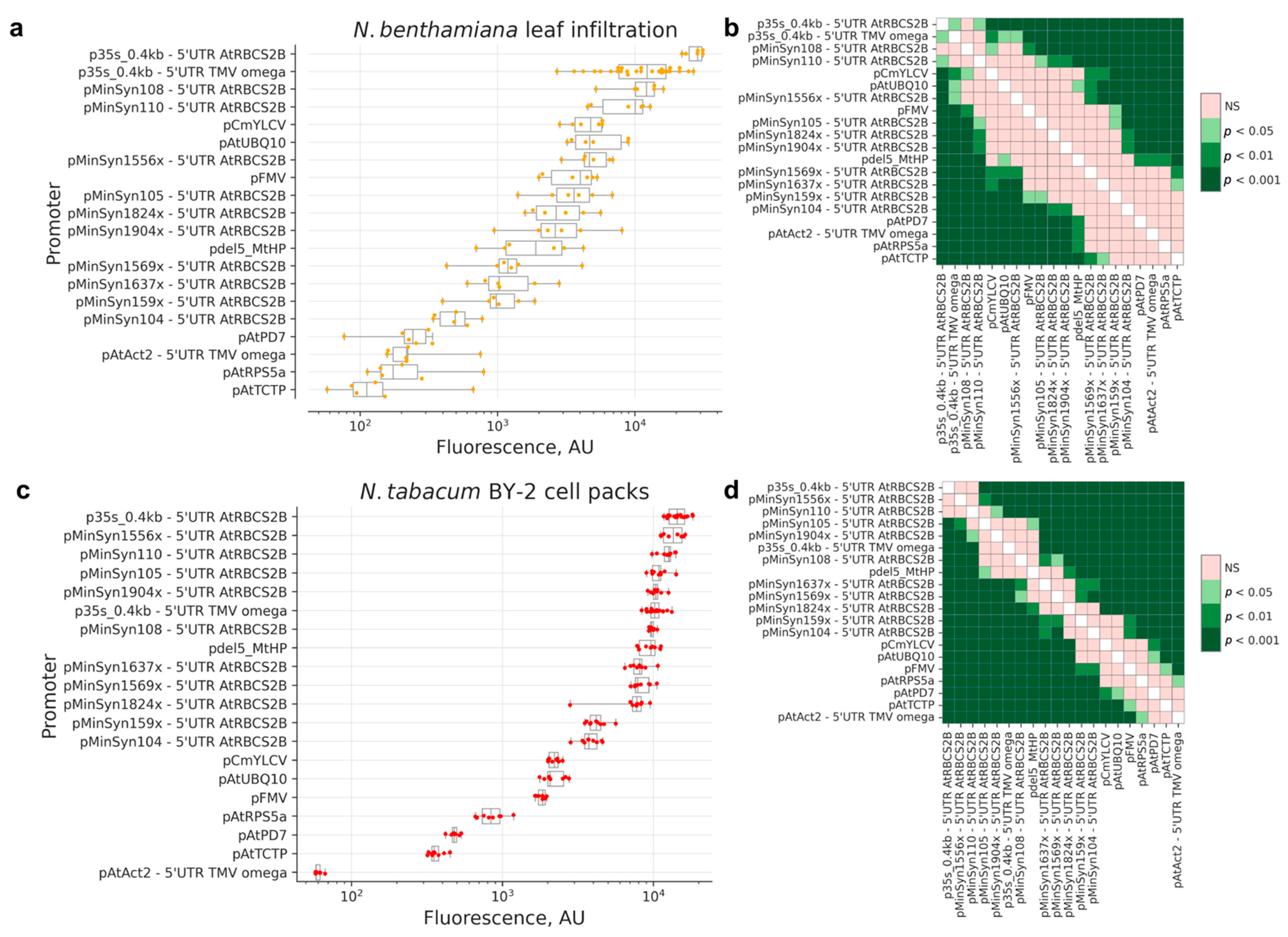
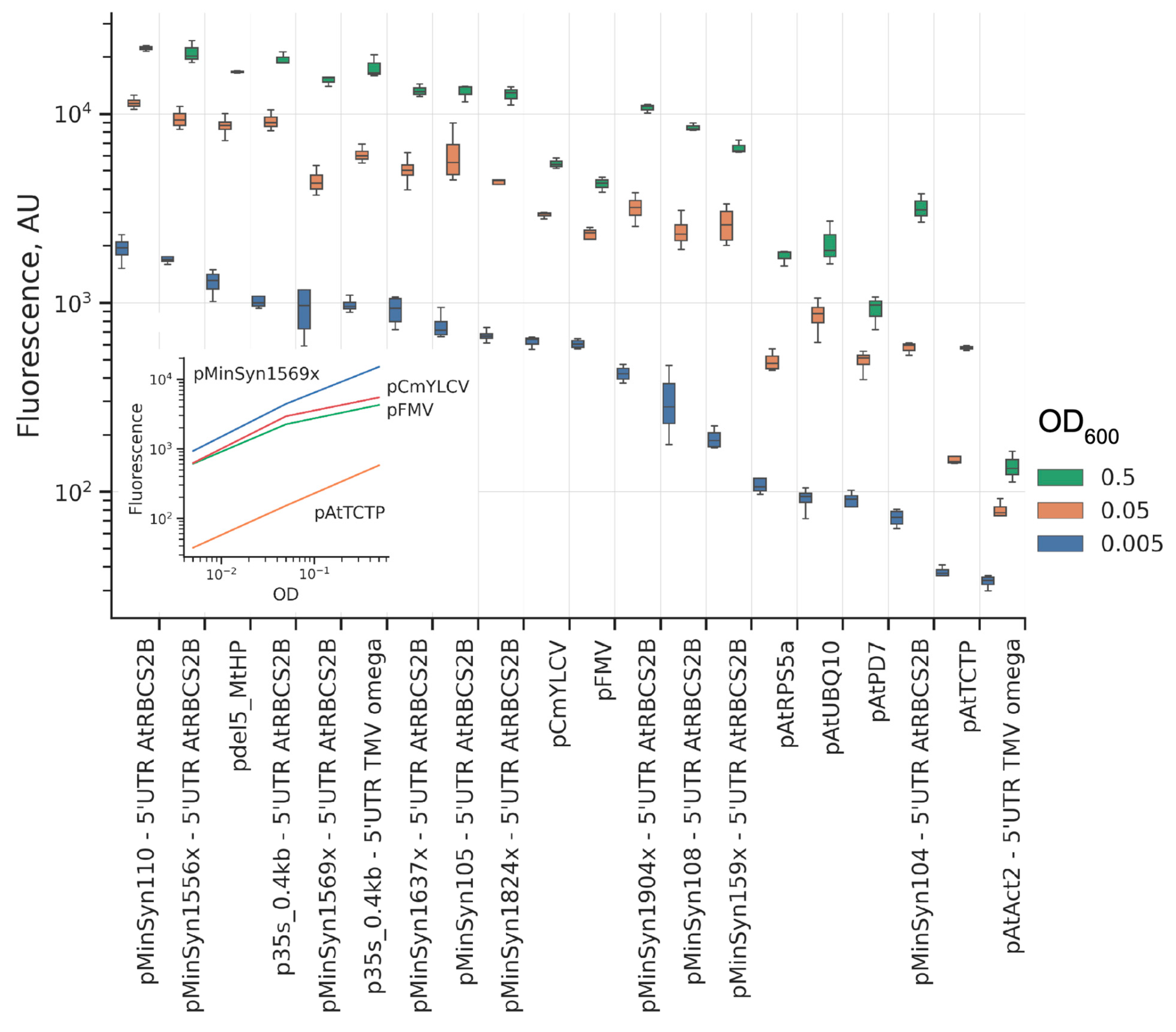
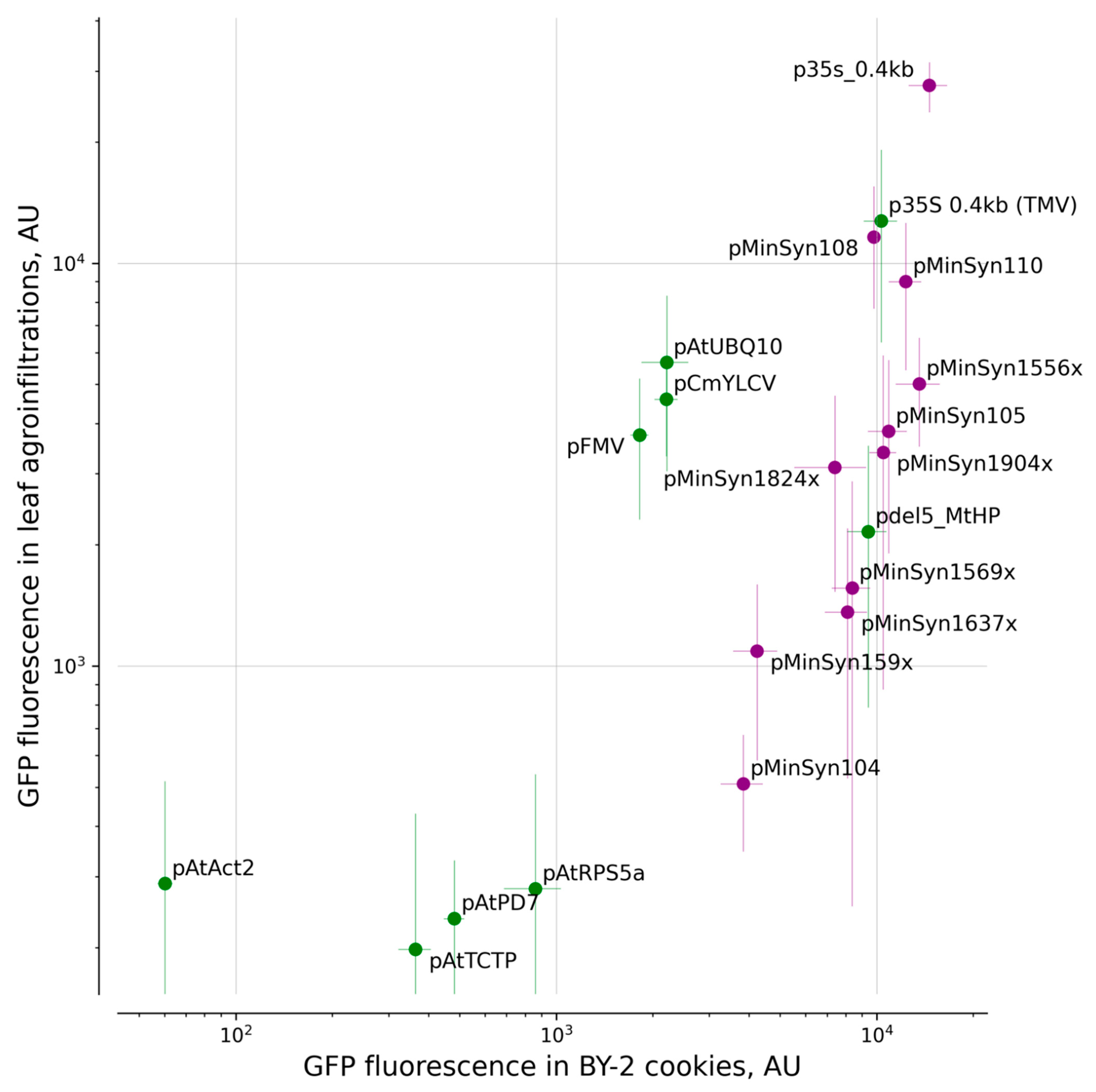
2.1. Comparison of Promoters
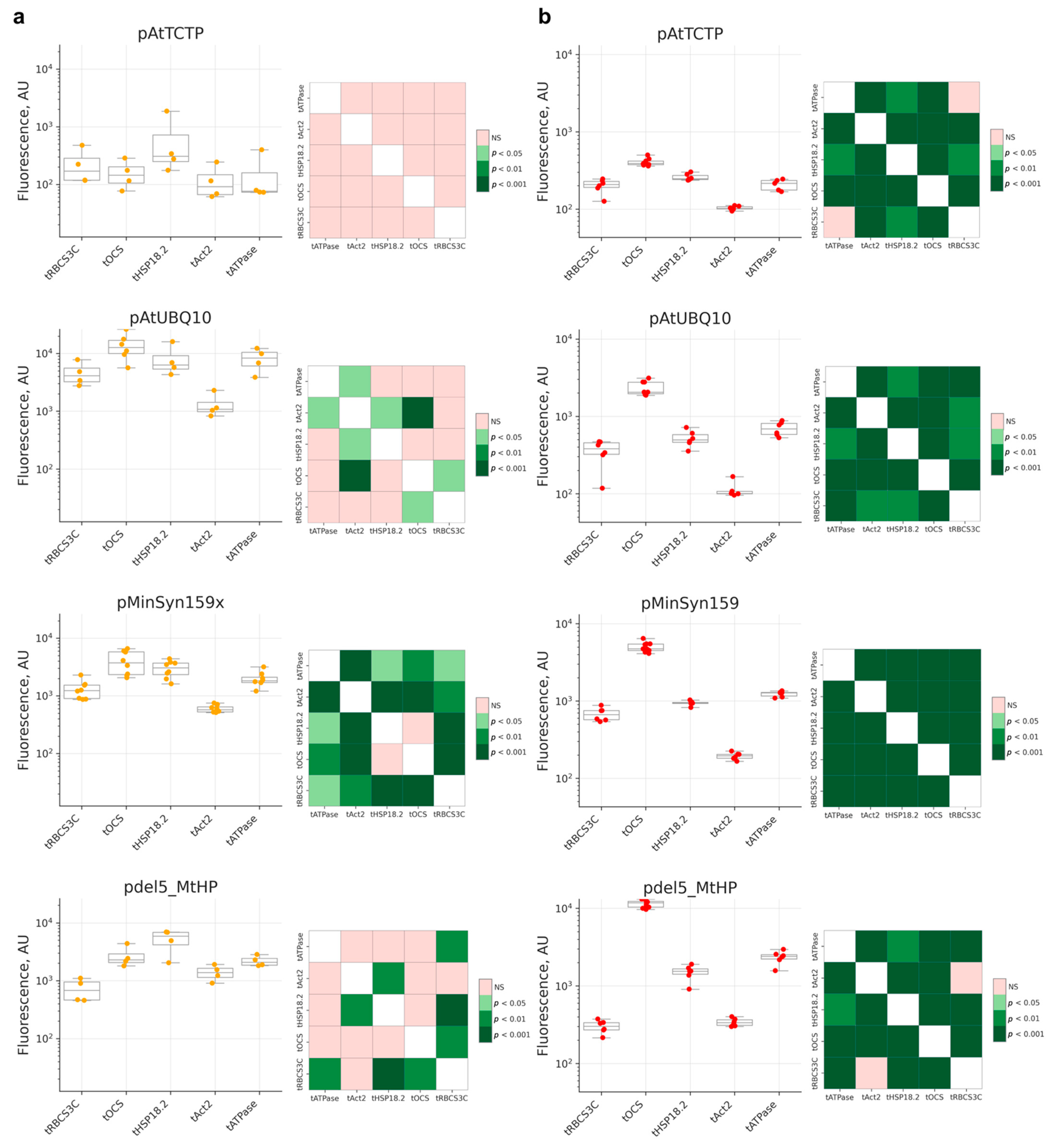
2.2. Comparison of Terminators
3. Discussion
4. Materials and Methods
4.1. Plasmids
4.2. Transformation of Agrobacterium tumefaciens
4.3. Nicotiana benthamiana Plants
4.4. Nicotiana tabacum BY-2 Cell Culture
4.5. Transient Transformation of Nicotiana benthamiana Leaves
4.6. Imaging of N. benthamiana Leaves
4.7. Transient Transformation of BY-2 Cell Culture
4.8. Measuring EGFP Fluorescence in BY-2 Cells
4.9. Data Presentation and Statistics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Amack, S.C.; Antunes, M.S. CaMV35S Promoter—A Plant Biology and Biotechnology Workhorse in the Era of Synthetic Biology. Curr. Plant Biol. 2020, 24, 100179. [Google Scholar] [CrossRef]
- Rajeevkumar, S.; Anunanthini, P.; Sathishkumar, R. Epigenetic Silencing in Transgenic Plants. Front. Plant Sci. 2015, 6, 693. [Google Scholar] [CrossRef] [PubMed]
- Porto, M.S.; Pinheiro, M.P.N.; Batista, V.G.L.; dos Santos, R.C.; de Filho, P.A.M.; de Lima, L.M. Plant Promoters: An Approach of Structure and Function. Mol. Biotechnol. 2014, 56, 38–49. [Google Scholar] [CrossRef]
- Jores, T.; Tonnies, J.; Wrightsman, T.; Buckler, E.S.; Cuperus, J.T.; Fields, S.; Queitsch, C. Synthetic Promoter Designs Enabled by a Comprehensive Analysis of Plant Core Promoters. Nat. Plants 2021, 7, 842–855. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.-M.; Kallam, K.; Tidd, H.; Gendarini, G.; Salzman, A.; Patron, N.J. Rational Design of Minimal Synthetic Promoters for Plants. Nucleic Acids Res. 2020, 48, 11845–11856. [Google Scholar] [CrossRef]
- Ali, S.; Kim, W.-C. A Fruitful Decade Using Synthetic Promoters in the Improvement of Transgenic Plants. Front. Plant Sci. 2019, 10, 1433. [Google Scholar] [CrossRef]
- Engler, C.; Youles, M.; Gruetzner, R.; Ehnert, T.-M.; Werner, S.; Jones, J.D.G.; Patron, N.J.; Marillonnet, S. A Golden Gate Modular Cloning Toolbox for Plants. ACS Synth. Biol. 2014, 3, 839–843. [Google Scholar] [CrossRef]
- Andreou, A.I.; Nirkko, J.; Ochoa-Villarreal, M.; Nakayama, N. Mobius Assembly for Plant Systems Highlights Promoter-Terminator Interaction in Gene Regulation. bioRxiv 2021. [Google Scholar] [CrossRef]
- Tian, C.; Zhang, Y.; Li, J.; Wang, Y. Benchmarking Intrinsic Promoters and Terminators for Plant Synthetic Biology Research. BioDesign Res. 2022, 2022, 9834989. [Google Scholar] [CrossRef]
- de Felippes, F.F.; McHale, M.; Doran, R.L.; Roden, S.; Eamens, A.L.; Finnegan, E.J.; Waterhouse, P.M. The Key Role of Terminators on the Expression and Post-Transcriptional Gene Silencing of Transgenes. Plant J. 2020, 104, 96–112. [Google Scholar] [CrossRef]
- Rademacher, T.; Sack, M.; Blessing, D.; Fischer, R.; Holland, T.; Buyel, J. Plant Cell Packs: A Scalable Platform for Recombinant Protein Production and Metabolic Engineering. Plant Biotechnol. J. 2019, 17, 1560–1566. [Google Scholar] [CrossRef]
- Sanger, M.; Daubert, S.; Goodman, R.M. Characteristics of a Strong Promoter from Figwort Mosaic Virus: Comparison with the Analogous 35S Promoter from Cauliflower Mosaic Virus and the Regulated Mannopine Synthase Promoter. Plant Mol. Biol. 1990, 14, 433–443. [Google Scholar] [CrossRef]
- Stavolone, L.; Kononova, M.; Pauli, S.; Ragozzino, A.; de Haan, P.; Milligan, S.; Lawton, K.; Hohn, T. Cestrum Yellow Leaf Curling Virus (CmYLCV) Promoter: A New Strong Constitutive Promoter for Heterologous Gene Expression in a Wide Variety of Crops. Plant Mol. Biol. 2003, 53, 663–673. [Google Scholar] [CrossRef]
- Han, Y.-J.; Kim, Y.-M.; Hwang, O.-J.; Kim, J.-I. Characterization of a Small Constitutive Promoter from Arabidopsis Translationally Controlled Tumor Protein (AtTCTP) Gene for Plant Transformation. Plant Cell Rep. 2015, 34, 265–275. [Google Scholar] [CrossRef]
- Xiao, K.; Zhang, C.; Harrison, M.; Wang, Z.-Y. Isolation and Characterization of a Novel Plant Promoter That Directs Strong Constitutive Expression of Transgenes in Plants. Mol. Breed. 2005, 15, 221–231. [Google Scholar] [CrossRef]
- Gantner, J.; Ordon, J.; Ilse, T.; Kretschmer, C.; Gruetzner, R.; Löfke, C.; Dagdas, Y.; Bürstenbinder, K.; Marillonnet, S.; Stuttmann, J. Peripheral Infrastructure Vectors and an Extended Set of Plant Parts for the Modular Cloning System. PLoS ONE 2018, 13, e0197185. [Google Scholar] [CrossRef]
- Jiang, P.; Zhang, K.; Ding, Z.; He, Q.; Li, W.; Zhu, S.; Cheng, W.; Zhang, K.; Li, K. Characterization of a Strong and Constitutive Promoter from the Arabidopsis Serine Carboxypeptidase-like Gene AtSCPL30 as a Potential Tool for Crop Transgenic Breeding. BMC Biotechnol. 2018, 18, 59. [Google Scholar] [CrossRef]
- Nagaya, S.; Kawamura, K.; Shinmyo, A.; Kato, K. The HSP Terminator of Arabidopsis Thaliana Increases Gene Expression in Plant Cells. Plant Cell Physiol. 2010, 51, 328–332. [Google Scholar] [CrossRef] [PubMed]
- Johansen, L.K.; Carrington, J.C. Silencing on the Spot. Induction and Suppression of RNA Silencing in the Agrobacterium-Mediated Transient Expression System. Plant Physiol. 2001, 126, 930–938. [Google Scholar] [CrossRef] [PubMed]
- Iverson, S.V.; Haddock, T.L.; Beal, J.; Densmore, D.M. CIDAR MoClo: Improved MoClo Assembly Standard and New E. Coli Part Library Enable Rapid Combinatorial Design for Synthetic and Traditional Biology. ACS Synth. Biol. 2016, 5, 99–103. [Google Scholar] [CrossRef]
- Nagata, T.; Nemoto, Y.; Hasezawa, S. Tobacco BY-2 Cell Line as the “HeLa” Cell in the Cell Biology of Higher Plants. In International Review of Cytology; Jeon, K.W., Friedlander, M., Eds.; Academic Press: Cambridge, MA, USA, 1992; Volume 132, pp. 1–30. [Google Scholar]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An Open-Source Platform for Biological-Image Analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Gengenbach, B.B.; Opdensteinen, P.; Buyel, J.F. Robot Cookies—Plant Cell Packs as an Automated High-Throughput Screening Platform Based on Transient Expression. Front. Bioeng. Biotechnol. 2020, 8, 393. [Google Scholar] [CrossRef] [PubMed]
| Element | Name | Description | Reference |
|---|---|---|---|
| Promoter | p35s_0.4kb–5′UTR TMV omega | Promoter 35s (Cauliflower Mosaic Virus), 0.4 kb in combination with 5′UTR, omega (Tobacco Mosaic Virus) or 5′UTR, RbcS2B (AT5g38420, A. thaliana) | [7] |
| p35s_0.4kb–5′UTR AtRBCS2B | |||
| pFMV | Figwort mosaic virus (FMV) 34S promoter + 5′UTR | [12] | |
| pCmYLCV | Cestrum yellow leaf curling virus (CmYLCV) promoter + 5′UTR | [13] | |
| pMinSyn104–5′UTR AtRBCS2B | Minimal synthetic promoters + 5′UTR, RbcS2B (AT5g38420, A. thaliana) | [5] | |
| pMinSyn105–5′UTR AtRBCS2B | |||
| pMinSyn108–5′UTR AtRBCS2B | |||
| pMinSyn110–5′UTR AtRBCS2B | |||
| pMinSyn159x–5′UTR AtRBCS2B | Predicted minimal synthetic promoters + 5′UTR, RbcS2B (AT5g38420, A. thaliana) | [5] * | |
| pMinSyn1556x–5′UTR AtRBCS2B | |||
| pMinSyn1569x–5′UTR AtRBCS2B | |||
| pMinSyn1637x–5′UTR AtRBCS2B | |||
| pMinSyn1824x–5′UTR AtRBCS2B | |||
| pMinSyn1904x–5′UTR AtRBCS2B | |||
| pAtTCTP | Small constitutive promoter from Arabidopsis translationally controlled tumor protein (AtTCTP) gene (0.3 kb) + 5′UTR | [14] | |
| pdel5_MtHP | Constitutive promoter upstream of MtHP gene isolated from Medicago truncatula + 5′UTR ** | [15] | |
| pAtUBQ10 | 800 bp fragment upstream of Arabidopsis UBQ10 (At4g05320)—strong, constitutive promoter + 5′UTR | [16] | |
| pAtRPS5a | 1700 bp upstream of Arabidopsis RPS5a (At3g11940)—strong, constitutive promoter + 5′UTR | [16] | |
| pAtPD7 | Strong and constitutive promoter from the Arabidopsis serine carboxypeptidase-like gene AtSCPL30 (456 bp upstream of the translation initiation codon ATG of AtSCPL30) + 5′UTR | [17] | |
| pAtAct2–5′UTR TMV omega | Promoter AtAct2 (AT3G18780, A. thaliana) + 5′UTR, omega (Tobacco Mosaic Virus) | [7] | |
| Terminator | tOCS | 3′UTR + terminator OCS (A. tumefaciens) | [7] |
| tAtAct2 | 3′UTR + terminator Act2 (A. thaliana) | ||
| tRBCS3C | 3′UTR + terminator RBCS3C (S. lycopersicum) | ||
| tATPase | 3′UTR + terminator ATPase (S. lycopersicum) | ||
| tHSP18.2 | 3′UTR + the heat shock protein 18.2 (HSP) terminator of Arabidopsis thaliana | [18] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shakhova, E.S.; Markina, N.M.; Mitiouchkina, T.; Bugaeva, E.N.; Karataeva, T.A.; Palkina, K.A.; Fakhranurova, L.I.; Yampolsky, I.V.; Sarkisyan, K.S.; Mishin, A.S. Systematic Comparison of Plant Promoters in Nicotiana spp. Expression Systems. Int. J. Mol. Sci. 2022, 23, 15441. https://doi.org/10.3390/ijms232315441
Shakhova ES, Markina NM, Mitiouchkina T, Bugaeva EN, Karataeva TA, Palkina KA, Fakhranurova LI, Yampolsky IV, Sarkisyan KS, Mishin AS. Systematic Comparison of Plant Promoters in Nicotiana spp. Expression Systems. International Journal of Molecular Sciences. 2022; 23(23):15441. https://doi.org/10.3390/ijms232315441
Chicago/Turabian StyleShakhova, Ekaterina S., Nadezhda M. Markina, Tatiana Mitiouchkina, Evgenia N. Bugaeva, Tatiana A. Karataeva, Kseniia A. Palkina, Liliia I. Fakhranurova, Ilia V. Yampolsky, Karen S. Sarkisyan, and Alexander S. Mishin. 2022. "Systematic Comparison of Plant Promoters in Nicotiana spp. Expression Systems" International Journal of Molecular Sciences 23, no. 23: 15441. https://doi.org/10.3390/ijms232315441
APA StyleShakhova, E. S., Markina, N. M., Mitiouchkina, T., Bugaeva, E. N., Karataeva, T. A., Palkina, K. A., Fakhranurova, L. I., Yampolsky, I. V., Sarkisyan, K. S., & Mishin, A. S. (2022). Systematic Comparison of Plant Promoters in Nicotiana spp. Expression Systems. International Journal of Molecular Sciences, 23(23), 15441. https://doi.org/10.3390/ijms232315441





