Plastid Transformation: New Challenges in the Circular Economy Era
Abstract
1. Introduction
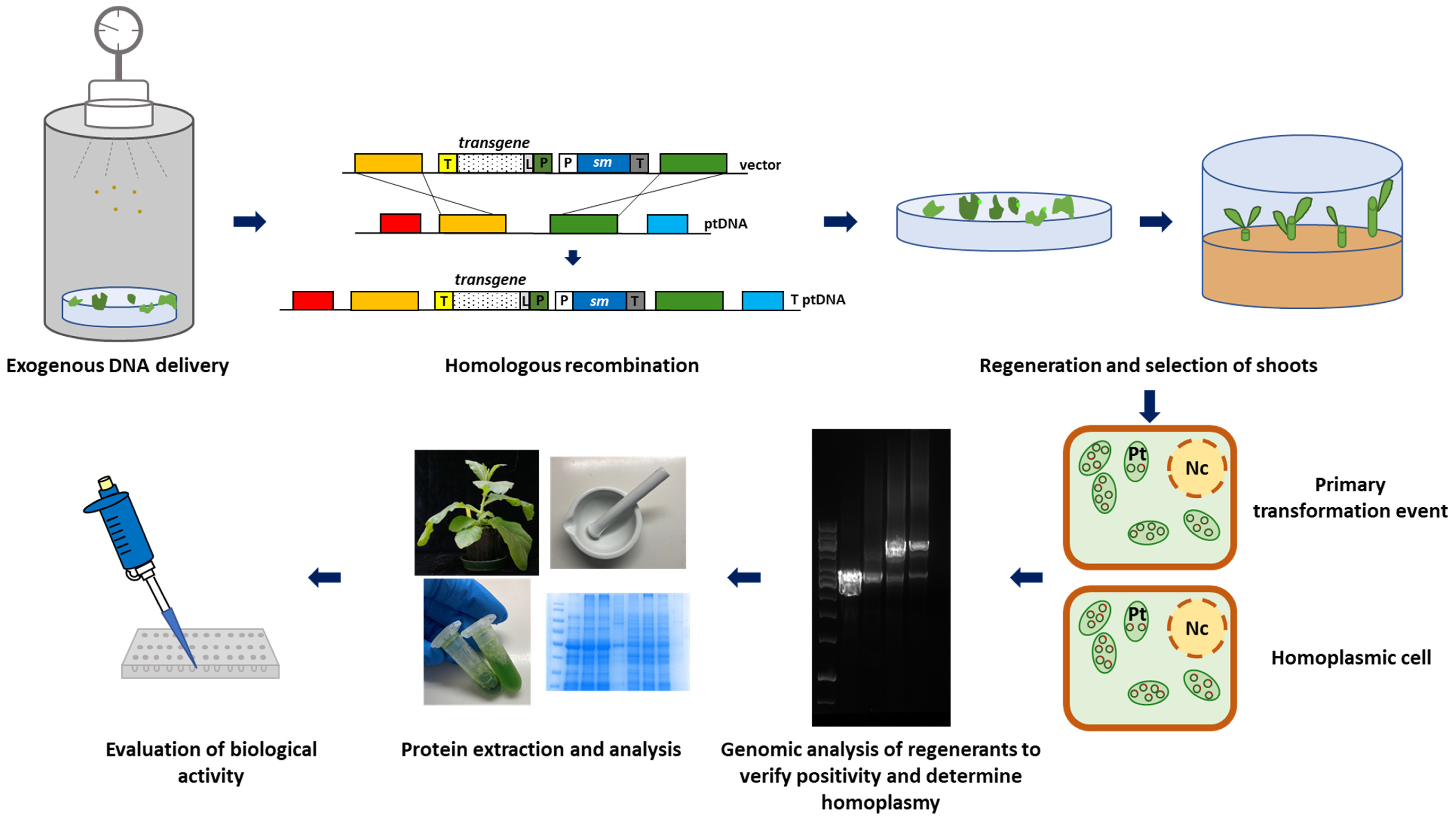
2. Plastid Transformation Technology
3. New Advances in Plastid Expression of Recombinant Proteins
4. Plastid-Based Enzymes for the Biorefinery Industry
| Heterologous Protein | Gene | Source | Expression Level | Phenotype of Transplastomic Plants | Physico-Chemical Characterization | References |
|---|---|---|---|---|---|---|
| acetyl xylan esterase | axe1 | Trichoderma reesei | color change observed | wild type | - | [30] |
| β-glucosidase | bgl1 | Trichoderma reesei | 14 U/mg CTSP a | wild type | 50 °C, pH 5.2 b | [30] |
| β-glucosidase | bglC1 | Thermobifida fusca | 5–40% TSP c | mutant | Topt 45 °C, pHopt 7.0 | [33] |
| β-glucosidase | bglC | Thermobifida fusca | up to 12% TSP | nd d | 50 °C b | [34] |
| β-glucosidase | celB | Pyrococcus furiosus | up to 75% TSP | wild type | Topt 105 °C, pH 4.0–5.0 | [12] |
| β-glucosidase | bgl1 | Aspergillus niger | 20 mg/g TSP | wild type | Topt 40 °C, pH 5.0 | [35] |
| β-mannase | man1 | Trichoderma reesei | 25 U/g FW | mutant (pale green) | 40 °C < Topt < 70 °C, 3.0 < pHopt < 7.0 | [36] |
| cellulases | celA | Thermotoga neapolitana | 23 mg/g TSP | wild type | Topt 65 °C, pH 5.0 | [35] |
| cellulases | celB | Thermotoga neapolitana | 21 mg/g TSP | wild type | Topt 65 °C, pH 5.0 | [35] |
| cutinase | cut | Fusarium solani | 15 U/mg CTSP | wild type | 30 °C, pH 8.0 b | [30] |
| cutinase | cut | Trichoderma reesei | nd | mutant | nd | [37] |
| endo-1,4-β-glucanase | cel6A | Thermobifida fusca | up to 2% TSP | nd b | 50 °C b | [38] |
| endoglucanase | cel6A | Thermobifida fusca | up to 10% TSP | wild type | nd | [39] |
| endo-β-glucanase E1 catalytic domain (E1cd) | e1cd | Acidothermus cellulolyticus | 12% TSP | mutant | 55 °C b | [40] |
| endoglucanase | celD | Clostridium thermocellum | up to 4930 U/g FW | wild type | Topt 60 °C, pHopt 6.0 | [30] |
| endoglucanase | egI | Trichoderma reesei | 339 U/mg CTSP | wild type | 50 °C, pH 5.2 b | [30] |
| endoglucanase | cel9A | Thermobifida fusca | 40% TSP | mutant | nd | [33] |
| endoglucanase | egph | Pyrococcus horikoshii | 25% TSP | mutant (pale green) | 85 °C, pH 5.5 b | [41] |
| endoglucanase | cel7, endoV, celK1 | Chaetomium globosum, Paenibacillus sp. | 8–10% TPC e | nd | 60 °C b | [42,43] |
| endoglucanase | endo | Sulfolobus solfataricus | 2% TSP | mutant (severe retarded growth) | nd | [12] |
| exo-cellobiohydrolase | cel6B | Thermobifida fusca | 3% TSP | nd b | 50 °C b | [38] |
| exo-cellobiohydrolase | cel3 | Phanerochaete chrysosporium | 0.4 U/mg protein | nd | nd | [42,43] |
| exoglucanase | celO | Clostridium thermocellum | 442 U/mg CTSP | wild type | 60 °C, pH 5.2 b | [30] |
| exoglucanase | cel6B | Thermobifida fusca | 5% TSP | mutant | nd | [33] |
| exoglucanase | cel6 | Chaetomium globosum | 0.4 U/mg protein | nd | nd | [42,43] |
| lipase | lipY | Mycobacterium tuberculosis | nd | wild type | - | [30] |
| manganese peroxidase | mnP-2 | Phanerochaete chrysosporium | nd | normal (high growth rate) | 65 °C, pH 6.0 | [44] |
| pectate lyases | pelA, pelB, pelD | Fusarium solani | up to 32 U/g FW | wild type | Topt 40 °C, 6.0 > pHopt > 8.0 | [30] |
| pectinase | pelA | Streptomyces thermocarboxydus | nd | wild type | 60 °C, pH 7.0 | [44] |
| swollenin | swo1 | Trichoderma reesei | swelling observed | wild type | - | [30] |
| swollenin | swo1 | Trichoderma reesei | nd | mutant | nd | [37] |
| xylanase | xynA | Bacillus subtilis strain NG-27 | 6% TSP | wild type | Topt 70 °C pHopt 8.4 | [29] |
| xylanase | xyn2 | Trichoderma reesei | 421 U/mg CTSP | wild type | 50 °C, pH 5.2 cb | [30] |
| xylanase | xyl10B | Thermotoga maritima | up to 15% TSP | wild type | termostable at 85 °C for 30 min | [31] |
| xylanase | xynA | Clostridium cellulovorans | 0.5% TSP | wild type | 40 °C b | [32] |
| xylanase | xyn10A | Aspergillus niger | up to 3% TSP | wild type | 40 °C b | [32] |
| xylanase | xyn11B | Aspergillus niger | up to 6% TSP | wild type | 40 °C b | [32] |
| xylanase | xyn | Alicyclobacillus acidocaldarius | 36% TSP | wild type | Topt 80 °C pH 6.0 | [12] |
| xyloglucanase | xeg74 | Thermobifida fusca | 5–40% TSP | mutant (pigment deficiency, retarded growth) | nd | [33] |
5. New Challenges for Biocatalysts and the Circular Economy
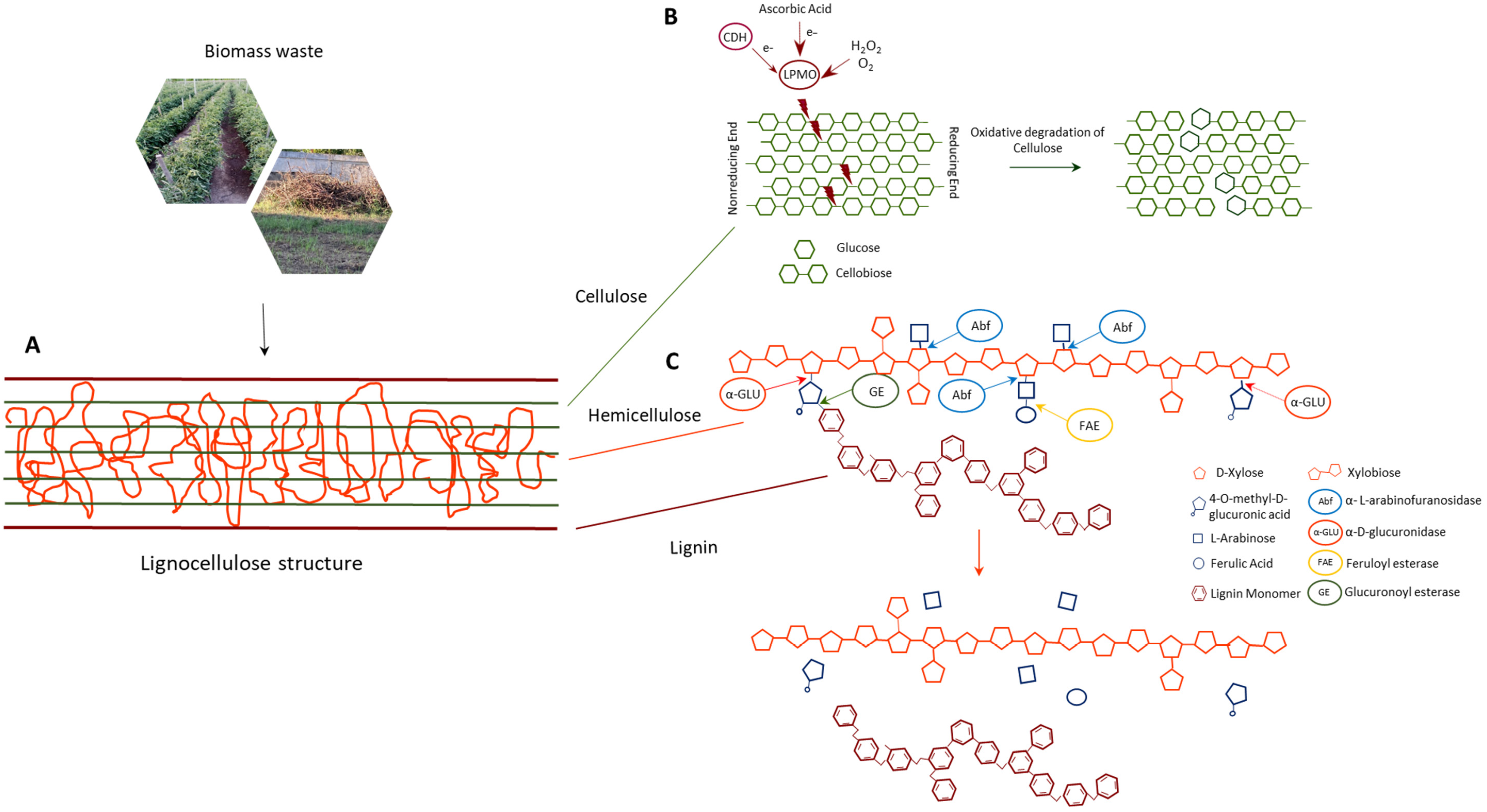
5.1. Enzymes for Lignocellulosic Waste Biorefinery
5.1.1. Lytic Polysaccharide Monooxygenases
5.1.2. Arabinofuranosidases
5.1.3. α-Glucuronidases
5.1.4. Feruloyl Esterases
5.1.5. Glucuronoyl Esterases
5.2. Other Enzymes for Waste Biorefinery
5.2.1. Chitinases
5.2.2. Chitin Deacetylase
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sherwood, J. The significance of biomass in a circular economy. Bioresour. Technol. 2020, 300, 122755. [Google Scholar] [CrossRef] [PubMed]
- Hubbe, M.A. Catalysts inspired by life. Biofuel Res. J. 2016, 3, 430. [Google Scholar] [CrossRef][Green Version]
- Atalah, J.; Cáceres-Moreno, P.; Espina, G.; Blamey, J.M. Thermophiles and the applications of their enzymes as new biocatalysts. Bioresour. Technol. 2019, 280, 478–488. [Google Scholar] [CrossRef] [PubMed]
- Martins, M.; Dinamarco, T.M.; Goldbeck, R. Recombinant chimeric enzymes for lignocellulosic biomass hydrolysis. Enzyme Microb. Technol. 2020, 140, 109647. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, L.F.; Amarelle, V.; Alves, L.D.; Viana de Siqueira, G.M.; Lovate, G.L.; Borelli, T.C.; Guazzaroni, M.-E. Genetically engineered proteins to improve biomass conversion: New advances and challenges for tailoring biocatalysts. Molecules 2019, 24, 2879. [Google Scholar] [CrossRef]
- Lambertz, C.; Garvey, M.; Klinger, J.; Heesel, D.; Klose, H.; Fischer, R.; Commandeur, U. Challenges and advances in the heterologous expression of cellulolytic enzymes: A review. Biotechnol. Biofuels 2014, 7, 135. [Google Scholar] [CrossRef]
- Buyel, J.F.; Stöger, E.; Bortesi, L. Targeted genome editing of plants and plant cells for biomanufacturing. Transgenic Res. 2021, 30, 401–426. [Google Scholar] [CrossRef]
- Scotti, N.; Rigano, M.M.; Cardi, T. Production of foreign proteins using plastid transformation. Biotechnol. Adv. 2012, 30, 387–397. [Google Scholar] [CrossRef]
- Scotti, N.; Bellucci, M.; Cardi, T. The chloroplasts as platform for recombinant proteins production. In Translation in Mitochondria and Other Organelles; Duchêne, A.-M., Ed.; Springer: Berlin/Heidelberg, Germany, 2013; pp. 225–262. [Google Scholar]
- Bock, R. Transplastomic approaches for metabolic engineering. Curr. Opin. Plant Biol. 2022, 66, 102185. [Google Scholar] [CrossRef]
- Oey, M.; Lohse, M.; Kreikemeyer, B.; Bock, R. Exhaustion of the chloroplast protein synthesis capacity by massive expression of a highly stable protein antibiotic. Plant J. 2009, 57, 436–445. [Google Scholar] [CrossRef]
- Castiglia, D.; Sannino, L.; Marcolongo, L.; Ionata, E.; Tamburino, R.; De Stradis, A.; Cobucci-Ponzano, B.; Moracci, M.; La Cara, F.; Scotti, N. High-level expression of thermostable cellulolytic enzymes in tobacco transplastomic plants and their use in hydrolysis of an industrially pretreated Arundo donax L. biomass. Biotechnol. Biofuels 2016, 9, 154. [Google Scholar] [CrossRef]
- Scotti, N.; Cardi, T. Plastid transformation as an expression tool for plant-derived biopharmaceuticals. In Transgenic Plants: Methods and Protocols; Dunwell, J.M., Wetten, A.C., Eds.; Humana Press: Totowa, NJ, USA, 2012; pp. 451–466. [Google Scholar]
- Bock, R. Engineering plastid genomes: Methods, tools, and applications in basic research and biotechnology. Annu. Rev. Plant Biol. 2015, 66, 211–241. [Google Scholar] [CrossRef]
- Bock, R. Engineering chloroplasts for high-level constitutive or inducible transgene expression. In Chloroplast Biotechnology: Methods and Protocols; Maliga, P., Ed.; Springer: New York, NY, USA, 2021; pp. 77–94. [Google Scholar]
- Apel, W.; Schulze, W.X.; Bock, R. Identification of protein stability determinants in chloroplasts. Plant J. 2010, 63, 636–650. [Google Scholar] [CrossRef]
- Singh, N.D.; Li, M.; Lee, S.B.; Schnell, D.; Daniell, H. Arabidopsis Tic40 expression in tobacco chloroplasts results in massive proliferation of the inner envelope membrane and upregulation of associated proteins. Plant Cell 2008, 20, 3405–3417. [Google Scholar] [CrossRef]
- Tissot, G.; Canard, H.; Nadai, M.; Martone, A.; Botterman, J.; Dubald, M. Translocation of aprotinin, a therapeutic protease inhibitor, into the thylakoid lumen of genetically engineered tobacco chloroplasts. Plant Biotechnol. J. 2008, 6, 309–320. [Google Scholar] [CrossRef]
- Shanmugabalaji, V.; Besagni, C.; Piller, L.E.; Douet, V.; Ruf, S.; Bock, R.; Kessler, F. Dual targeting of a mature plastoglobulin/fibrillin fusion protein to chloroplast plastoglobules and thylakoids in transplastomic tobacco plants. Plant Mol. Biol. 2013, 81, 13–25. [Google Scholar] [CrossRef]
- Bally, J.; Paget, E.; Droux, M.; Job, C.; Job, D.; Dubald, M. Both the stroma and thylakoid lumen of tobacco chloroplasts are competent for the formation of disulphide bonds in recombinant proteins. Plant Biotechnol. J. 2008, 6, 46–61. [Google Scholar] [CrossRef]
- Ahmad, N.; Michoux, F.; Nixon, P.J. Investigating the production of foreign membrane proteins in tobacco chloroplasts: Expression of an algal plastid terminal oxidase. PLoS ONE 2012, 7, e41722. [Google Scholar] [CrossRef]
- De Marchis, F.; Bellucci, M.; Pompa, A. Phaseolin expression in tobacco chloroplast reveals an autoregulatory mechanism in heterologous protein translation. Plant Biotechnol. J. 2016, 14, 603–614. [Google Scholar] [CrossRef]
- Scotti, N.; Sannino, L.; Idoine, A.; Hamman, P.; De Stradis, A.; Giorio, P.; Maréchal-Drouard, L.; Bock, R.; Cardi, T. The HIV-1 Pr55gag polyprotein binds to plastidial membranes and leads to severe impairment of chloroplast biogenesis and seedling lethality in transplastomic tobacco plants. Transgenic Res. 2015, 24, 319–331. [Google Scholar] [CrossRef]
- Hennig, A.; Bonfig, K.; Roitsch, T.; Warzecha, H. Expression of the recombinant bacterial outer surface protein A in tobacco chloroplasts leads to thylakoid localization and loss of photosynthesis. FEBS J. 2007, 274, 5749–5758. [Google Scholar] [CrossRef] [PubMed]
- Ren, K.; Xu, W.; Ren, B.; Fu, J.; Jiang, C.; Zhang, J. A simple technology for plastid transformation with fragmented DNA. J. Exp. Bot. 2022, 73, 6078–6088. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; You, L.; Li, S.; Ma, M.; Wu, M.; Ma, L.; Bock, R.; Chang, L.; Zhang, J. In vivo assembly in Escherichia coli of transformation vectors for plastid genome engineering. Front. Plant Sci. 2017, 8, 1454. [Google Scholar] [CrossRef] [PubMed]
- Jakubiec, A.; Sarokina, A.; Choinard, S.; Vlad, F.; Malcuit, I.; Sorokin, A.P. Replicating minichromosomes as a new tool for plastid genome engineering. Nat. Plants 2021, 7, 932–941. [Google Scholar] [CrossRef] [PubMed]
- Odahara, M.; Horii, Y.; Itami, J.; Watanabe, K.; Numata, K. Functional peptide-mediated plastid transformation in tobacco, rice, and kenaf. Front. Plant Sci. 2022, 13, 989310. [Google Scholar] [CrossRef]
- Leelavathi, S.; Gupta, N.; Maiti, S.; Ghosh, A.; Reddy, V.S. Overproduction of an alkali- and thermo-stable xylanase in tobacco chloroplasts and efficient recovery of the enzyme. Mol. Breed 2003, 11, 59–67. [Google Scholar] [CrossRef]
- Verma, D.; Kanagaraj, A.; Jin, S.; Singh, N.D.; Kolattukudy, P.E.; Daniell, H. Chloroplast-derived enzyme cocktails hydrolyse lignocellulosic biomass and release fermentable sugars. Plant Biotechnol. J. 2010, 8, 332–350. [Google Scholar] [CrossRef]
- Kim, J.Y.; Kavas, M.; Fouad, W.M.; Nong, G.; Preston, J.F.; Altpeter, F. Production of hyperthermostable GH10 xylanase Xyl10B from Thermotoga maritima in transplastomic plants enables complete hydrolysis of methylglucuronoxylan to fermentable sugars for biofuel production. Plant Mol. Biol. 2011, 76, 357–369. [Google Scholar] [CrossRef]
- Kolotilin, I.; Kaldis, A.; Pereira, E.O.; Laberge, S.; Menassa, R. Optimization of transplastomic production of hemicellulases in tobacco: Effects of expression cassette configuration and tobacco cultivar used as production platform on recombinant protein yields. Biotechnol. Biofuels 2013, 6, 65. [Google Scholar] [CrossRef]
- Petersen, K.; Bock, R. High-level expression of a suite of thermostable cell wall-degrading enzymes from the chloroplast genome. Plant Mol. Biol. 2011, 76, 311–321. [Google Scholar] [CrossRef]
- Gray, B.N.; Yang, H.; Ahner, B.A.; Hanson, M.R. An efficient downstream box fusion allows high-level accumulation of active bacterial beta-glucosidase in tobacco chloroplasts. Plant Mol. Biol. 2011, 76, 345–355. [Google Scholar] [CrossRef]
- Espinoza-Sánchez, E.A.; Torres-Castillo, J.A.; Rascón-Cruz, Q.; Zavala-García, F.; Sinagawa-García, S.R. Production and characterization of fungal β-glucosidase and bacterial cellulases by tobacco chloroplast transformation. Plant Biotechnol. Rep. 2016, 10, 61–73. [Google Scholar] [CrossRef]
- Agrawal, P.; Verma, D.; Daniell, H. Expression of Trichoderma reesei β-mannanase in tobacco chloroplasts and its utilization in lignocellulosic woody biomass hydrolysis. PLoS ONE 2011, 6, e29302. [Google Scholar] [CrossRef]
- Verma, D.; Jin, S.; Kanagaraj, A.; Singh, N.D.; Daniel, J.; Kolattukudy, P.E.; Miller, M.; Daniell, H. Expression of fungal cutinase and swollenin in tobacco chloroplasts reveals novel enzyme functions and/or substrates. PLoS ONE 2013, 8, e57187. [Google Scholar] [CrossRef]
- Yu, L.X.; Gray, B.N.; Rutzke, C.J.; Walker, L.P.; Wilson, D.B.; Hanson, M.R. Expression of thermostable microbial cellulases in the chloroplasts of nicotine-free tobacco. J. Biotechnol. 2007, 131, 362–369. [Google Scholar] [CrossRef]
- Gray, B.N.; Ahner, B.A.; Hanson, M.R. High-level bacterial cellulase accumulation in chloroplast-transformed tobacco mediated by downstream box fusions. Biotechnol. Bioeng. 2009, 102, 1045–1054. [Google Scholar] [CrossRef]
- Ziegelhoffer, T.; Raasch, J.A.; Austin-Phillips, S. Expression of Acidothermus cellulolyticus E1 endo-beta-1,4-glucanase catalytic domain in transplastomic tobacco. Plant Biotechnol. J. 2009, 7, 527–536. [Google Scholar] [CrossRef]
- Nakahira, Y.; Ishikawa, K.; Tanaka, K.; Tozawa, Y.; Shiina, T. Overproduction of hyperthermostable β-1,4-endoglucanase from the Archaeon Pyrococcus horikoshii by tobacco chloroplast engineering. Biosci. Biotechnol. Biochem. 2013, 77, 2140–2143. [Google Scholar] [CrossRef]
- Longoni, P.; Leelavathi, S.; Doria, E.; Reddy, V.S.; Cella, R. Production by tobacco transplastomic plants of recombinant fungal and bacterial cell-wall degrading enzymes to be used for cellulosic biomass saccharification. Biomed. Res. Int. 2015, 2015, 289759. [Google Scholar] [CrossRef]
- Faè, M.; Accossato, S.; Cella, R.; Fontana, F.; Goldschmidt-Clermont, M.; Leelavathi, S.; Reddy, V.S.; Longoni, P. Comparison of transplastomic Chlamydomonas reinhardtii and Nicotiana tabacum expression system for the production of a bacterial endoglucanase. Appl. Microbiol. Biotechnol. 2017, 101, 4085–4092. [Google Scholar] [CrossRef]
- Espinoza-Sánchez, E.A.; Álvarez-Hernández, M.H.; Torres-Castillo, J.A.; Rascón-Cruz, Q.; Gutiérrez-Díez, A.; Zavala-García, F.; Sinagawa-García, S.R. Stable expression and characterization of a fungal pectinase and bacterial peroxidase genes in tobacco chloroplast. Electron. J. Biotechnol. 2015, 18, 161–168. [Google Scholar] [CrossRef]
- Jin, S.; Kanagaraj, A.; Verma, D.; Lange, T.; Daniell, H. Release of hormones from conjugates: Chloroplast expression of beta-glucosidase results in elevated phytohormone levels associated with significant increase in biomass and protection from aphids or whiteflies conferred by sucrose esters. Plant Physiol. 2011, 155, 222–235. [Google Scholar] [CrossRef] [PubMed]
- Rojas, L.F.; Zapata, P.; Ruiz-Tirado, L. Agro-industrial waste enzymes: Perspectives in circular economy. Curr. Opin. Green Sustain. Chem. 2022, 34, 100585. [Google Scholar] [CrossRef]
- Sahay, S. Deconstruction of lignocelluloses: Potential biological approaches. In Handbook of Biofuels; Sahay, S., Ed.; Academic Press: Cambridge, MA, USA, 2022; pp. 207–232. [Google Scholar]
- Levasseur, A.; Drula, E.; Lombard, V.; Coutinho, P.M.; Henrissat, B. Expansion of the enzymatic repertoire of the CAZy database to integrate auxiliary redox enzymes. Biotechnol. Biofuels 2013, 6, 41. [Google Scholar] [CrossRef]
- Laurent, C.V.F.P.; Sun, P.; Scheiblbrandner, S.; Csarman, F.; Cannazza, P.; Frommhagen, M.; van Berkel, W.J.H.; Oostenbrink, C.; Kabel, M.A.; Ludwig, R. Influence of lytic polysaccharide monooxygenase active site segments on activity and affinity. Int. J. Mol. Sci. 2019, 20, 6219. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, K.; Suzuki, M.; Taiyoji, M.; Nikaidou, N.; Watanabe, T. Chitin Binding Protein (CBP21) in the culture supernatant of Serratia marcescens 2170. Biosci. Biotechnol. Biochem. 1998, 62, 128–135. [Google Scholar] [CrossRef]
- Vaaje-Kolstad, G.; Houston, D.R.; Riemen, A.H.; Eijsink, V.G.; van Aalten, D.M. Crystal structure and binding properties of the Serratia marcescens chitin-binding protein CBP21. J. Biol. Chem. 2005, 280, 11313–11319. [Google Scholar] [CrossRef]
- Forsberg, Z.; Mackenzie, A.K.; Sorlie, M.; Rohr, A.K.; Helland, R.; Arvai, A.S.; Vaaje-Kolstad, G.; Eijsink, V.G. Structural and functional characterization of a conserved pair of bacterial cellulose-oxidizing lytic polysaccharide monooxygenases. Proc. Natl. Acad. Sci. USA 2014, 111, 8446–8451. [Google Scholar] [CrossRef]
- Bentley, S.D.; Chater, K.F.; Cerdeno-Tarraga, A.M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 2002, 417, 141–147. [Google Scholar] [CrossRef]
- Rodrigues, K.B.; Macedo, J.K.A.; Teixeira, T.; Barros, J.S.; Araujo, A.C.B.; Santos, F.P.; Quirino, B.F.; Brasil, B.; Salum, T.F.C.; Abdelnur, P.V.; et al. Recombinant expression of Thermobifida fusca E7 LPMO in Pichia pastoris and Escherichia coli and their functional characterization. Carbohydr. Res. 2017, 448, 175–181. [Google Scholar] [CrossRef]
- Russo, D.A.; Zedler, J.A.Z.; Wittmann, D.N.; Mollers, B.; Singh, R.K.; Batth, T.S.; van Oort, B.; Olsen, J.V.; Bjerrum, M.J.; Jensen, P.E. Expression and secretion of a lytic polysaccharide monooxygenase by a fast-growing cyanobacterium. Biotechnol. Biofuels 2019, 12, 74. [Google Scholar] [CrossRef]
- Li, F.; Liu, Y.; Liu, Y.; Li, Y.; Yu, H. Heterologous expression and characterization of a novel lytic polysaccharide monooxygenase from Natrialbaceae archaeon and its application for chitin biodegradation. Bioresour. Technol. 2022, 354, 127174. [Google Scholar] [CrossRef]
- Arab-Jaziri, F.; Bissaro, B.; Tellier, C.; Dion, M.; Fauré, R.; O’Donohue, M.J. Enhancing the chemoenzymatic synthesis of arabinosylated xylo-oligosaccharides by GH51 α-l-arabinofuranosidase. Carbohydr. Res. 2015, 401, 64–72. [Google Scholar] [CrossRef]
- Liu, G.; Qu, Y. Integrated engineering of enzymes and microorganisms for improving the efficiency of industrial lignocellulose deconstruction. Eng. Microbiol. 2021, 1, 100005. [Google Scholar] [CrossRef]
- Numan, M.T.; Bhosle, N.B. α-l-Arabinofuranosidases: The potential applications in biotechnology. J. Ind. Microbiol. Biotechnol. 2006, 33, 247–260. [Google Scholar] [CrossRef]
- Kobayashi, M.; Kumagai, Y.; Yamamoto, Y.; Yasui, H.; Kishimura, H. Identification of a key enzyme for the hydrolysis of β-(1→3)-xylosyl linkage in red alga dulse xylooligosaccharide from Bifidobacterium adolescentis. Mar. Drugs 2020, 18, 174. [Google Scholar] [CrossRef]
- Thakur, A.; Sharma, K.; Goyal, A. α-l-Arabinofuranosidase: A potential enzyme for the food industry. In Green Bio-processes: Enzymes in Industrial Food Processing; Parameswaran, B., Varjani, S., Raveendran, S., Eds.; Springer: Singapore, 2019; pp. 229–244. [Google Scholar]
- Wilkens, C.; Andersen, S.; Dumon, C.; Berrin, J.G.; Svensson, B. GH62 arabinofuranosidases: Structure, function and applications. Biotechnol. Adv. 2017, 35, 792–804. [Google Scholar] [CrossRef]
- Limsakul, P.; Phitsuwan, P.; Waeonukul, R.; Pason, P.; Tachaapaikoon, C.; Poomputsa, K.; Kosugi, A.; Ratanakhanokchai, K. A novel multifunctional arabinofuranosidase/endoxylanase/beta-xylosidase GH43 enzyme from Paenibacillus curdlanolyticus B-6 and its synergistic action to produce arabinose and xylose from cereal arabinoxylan. Appl. Environ. Microbiol. 2021, 87, e0173021. [Google Scholar] [CrossRef]
- Wang, W.; Mai-Gisondi, G.; Stogios, P.J.; Kaur, A.; Xu, X.; Cui, H.; Turunen, O.; Savchenko, A.; Master, E.R. Elucidation of the molecular basis for arabinoxylan-debranching activity of a thermostable family GH62 α-l-arabinofuranosidase from Streptomyces thermoviolaceus. Appl. Environ. Microbiol. 2014, 80, 5317–5329. [Google Scholar] [CrossRef]
- Olajide, A.O.; Adesina, F.C.; Onilude, A.A. A thermostable and alkalitolerant arabinofuranosidase by Streptomyces lividus. Biotechnol. J. Int. 2020, 24, 35–47. [Google Scholar] [CrossRef]
- Sevim, E.; Inan Bektas, K.; Sevim, A.; Canakci, S.; Sahin, I.; Belduz, A.O. Purification and characterization of α-L-arabinofuranosidases from Geobacillus stearothermophilus strain 12. Biologia 2017, 72, 831–839. [Google Scholar] [CrossRef]
- İlgü, H.; Sürmeli, Y.; Şanlı-Mohamed, G. A thermophilic α-l-Arabinofuranosidase from Geobacillus vulcani GS90: Heterologous expression, biochemical characterization, and its synergistic action in fruit juice enrichment. Eur. Food Res. Technol. 2018, 244, 1627–1636. [Google Scholar] [CrossRef]
- Xie, J.; Zhao, D.; Zhao, L.; Pei, J.; Xiao, W.; Ding, G.; Wang, Z.; Xu, J. Characterization of a novel arabinose-tolerant alpha-L-arabinofuranosidase with high ginsenoside Rc to ginsenoside Rd bioconversion productivity. J. Appl. Microbiol. 2016, 120, 647–660. [Google Scholar] [CrossRef]
- Morana, A.; Paris, O.; Maurelli, L.; Rossi, M.; Cannio, R. Gene cloning and expression in Escherichia coli of a bi-functional β-D-xylosidase/α-l-arabinosidase from Sulfolobus solfataricus involved in xylan degradation. Extremophiles 2007, 11, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Saleh, M.A.; Mahmud, S.; Albogami, S.; El-Shehawi, A.M.; Paul, G.K.; Islam, S.; Dutta, A.K.; Uddin, M.S.; Zaman, S. Biochemical and molecular dynamics study of a novel GH43 alpha-l-arabinofuranosidase/beta-xylosidase from Caldicellulosiruptor saccharolyticus DSM8903. Front. Bioeng. Biotechnol. 2022, 10, 810542. [Google Scholar] [CrossRef]
- Zhang, X.J.; Wang, L.; Wang, S.; Chen, Z.L.; Li, Y.H. Contributions and characteristics of two bifunctional GH43 β-xylosidase /α-L-arabinofuranosidases with different structures on the xylan degradation of Paenibacillus physcomitrellae strain XB. Microbiol. Res. 2021, 253, 126886. [Google Scholar] [CrossRef] [PubMed]
- Underlin, E.N.; Frommhagen, M.; Dilokpimol, A.; van Erven, G.; de Vries, R.P.; Kabel, M.A. Feruloyl esterases for biorefineries: Subfamily classified specificity for natural substrates. Front. Bioeng. Biotechnol. 2020, 8, 332. [Google Scholar] [CrossRef]
- Biely, P.; Singh, S.; Puchart, V. Towards enzymatic breakdown of complex plant xylan structures: State of the art. Biotechnol. Adv. 2016, 34, 1260–1274. [Google Scholar] [CrossRef]
- Olson, D.G.; McBride, J.E.; Joe Shaw, A.; Lynd, L.R. Recent progress in consolidated bioprocessing. Curr. Opin. Biotechnol. 2012, 23, 396–405. [Google Scholar] [CrossRef]
- Bronnenmeier, K.; Meissner, H.; Stocker, S.; Staudenbauer, W.L. α-D-glucuronidases from the xylanolytic thermophiles Clostridium stercorarium and Thermoanaerobacterium saccharolyticum. Microbiology 1995, 141, 2033–2040. [Google Scholar] [CrossRef][Green Version]
- Choi, I.-D.; Kim, H.-Y.; Choi, Y.-J. Gene cloning and characterization of α-glucuronidase of Bacillus stearothermophilus No. 236. Biosci. Biotechnol. Biochem. 2000, 64, 2530–2537. [Google Scholar] [CrossRef]
- Zaide, G.; Shallom, D.; Shulami, S.; Zolotnitsky, G.; Golan, G.; Baasov, T.; Shoham, G.; Shoham, Y. Biochemical characterization and identification of catalytic residues in α-glucuronidase from Bacillus stearothermophilus T-6. Eur. J. Biochem. 2001, 268, 3006–3016. [Google Scholar] [CrossRef]
- Ruile, P.; Winterhalter, C.; Liebl, W. Isolation and analysis of a gene encoding α-glucuronidase, an enzyme with a novel primary structure involved in the breakdown of xylan. Mol. Microbiol. 1997, 23, 267–279. [Google Scholar] [CrossRef]
- Suresh, C.; Kitaoka, M.; Hayashi, K. A thermostable non-xylanolytic α-glucuronidase of Thermotoga maritima MSB8. Biosci. Biotechnol. Biochem. 2003, 67, 2359–2364. [Google Scholar] [CrossRef]
- Malgas, S.; Mafa, M.S.; Mkabayi, L.; Pletschke, B.I. A mini review of xylanolytic enzymes with regards to their synergistic interactions during hetero-xylan degradation. World J. Microbiol. Biotechnol. 2019, 35, 187. [Google Scholar] [CrossRef]
- Wong, D.W.S. Feruloyl esterase. Appl. Biochem. Biotechnol. 2006, 133, 87–112. [Google Scholar] [CrossRef]
- Oliveira, D.M.; Mota, T.R.; Oliva, B.; Segato, F.; Marchiosi, R.; Ferrarese-Filho, O.; Faulds, C.B.; dos Santos, W.D. Feruloyl esterases: Biocatalysts to overcome biomass recalcitrance and for the production of bioactive compounds. Bioresour. Technol. 2019, 278, 408–423. [Google Scholar] [CrossRef]
- Zduńska, K.; Dana, A.; Kolodziejczak, A.; Rotsztejn, H. Antioxidant properties of ferulic acid and its possible application. Skin Pharmacol. Physiol. 2018, 31, 332–336. [Google Scholar] [CrossRef]
- Antonopoulou, I.; Leonov, L.; Jütten, P.; Cerullo, G.; Faraco, V.; Papadopoulou, A.; Kletsas, D.; Ralli, M.; Rova, U.; Christakopoulos, P. Optimized synthesis of novel prenyl ferulate performed by feruloyl esterases from Myceliophthora thermophila in microemulsions. Appl. Microbiol. Biotechnol. 2017, 101, 3213–3226. [Google Scholar] [CrossRef]
- Esteban-Torres, M.; Landete José, M.; Reverón, I.; Santamaría, L.; de las Rivas, B.; Muñoz, R. A Lactobacillus plantarum esterase active on a broad range of phenolic esters. Appl. Environ. Microbiol. 2015, 81, 3235–3242. [Google Scholar] [CrossRef]
- Deng, H.; Jia, P.; Jiang, J.; Bai, Y.; Fan, T.-P.; Zheng, X.; Cai, Y. Expression and characterisation of feruloyl esterases from Lactobacillus fermentum JN248 and release of ferulic acid from wheat bran. Int. J. Biol. Macromol. 2019, 138, 272–277. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.R.; Baik, S.H. Molecular cloning, purification, and characterization of a novel thermostable cinnamoyl esterase from Lactobacillus helveticus KCCM 11223. Prep. Biochem. Biotechnol. 2017, 47, 496–504. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.-H.; Baik, S.-H. Properties of recombinant novel cinnamoyl esterase from Lactobacillus acidophilus F46 isolated from human intestinal bacterium. J. Mol. Catal. B Enzym. 2015, 116, 9–15. [Google Scholar] [CrossRef]
- Ay Sal, F.; Colak, D.N.; Guler, H.I.; Canakci, S.; Belduz, A.O. Biochemical characterization of a novel thermostable feruloyl esterase from Geobacillus thermoglucosidasius DSM 2542T. Mol. Biol. Rep. 2019, 46, 4385–4395. [Google Scholar] [CrossRef] [PubMed]
- Rakotoarivonina, H.; Hermant, B.; Chabbert, B.; Touzel, J.-P.; Remond, C. A thermostable feruloyl-esterase from the hemicellulolytic bacterium Thermobacillus xylanilyticus releases phenolic acids from non-pretreated plant cell walls. Appl. Microbiol. Biotechnol. 2011, 90, 541–552. [Google Scholar] [CrossRef]
- Kmezik, C.; Bonzom, C.; Olsson, L.; Mazurkewich, S.; Larsbrink, J. Multimodular fused acetyl–feruloyl esterases from soil and gut Bacteroidetes improve xylanase depolymerization of recalcitrant biomass. Biotechnol. Biofuels 2020, 13, 60. [Google Scholar] [CrossRef]
- Blum, D.L.; Kataeva, I.A.; Li, X.-L.; Ljungdahl, L.G. Feruloyl esterase activity of the Clostridium thermocellum cellulosome can be attributed to previously unknown domains of XynY and XynZ. J. Bacteriol. 2000, 182, 1346–1351. [Google Scholar] [CrossRef]
- Prates, J.A.M.; Tarbouriech, N.; Charnock, S.J.; Fontes, C.M.G.A.; Ferreira, L.s.M.A.; Davies, G.J. The structure of the feruloyl esterase module of xylanase 10B from Clostridium thermocellum provides insights into substrate recognition. Structure 2001, 9, 1183–1190. [Google Scholar] [CrossRef]
- Arnling Bååth, J.; Mazurkewich, S.; Knudsen, R.M.; Poulsen, J.-C.N.; Olsson, L.; Lo Leggio, L.; Larsbrink, J. Biochemical and structural features of diverse bacterial glucuronoyl esterases facilitating recalcitrant biomass conversion. Biotechnol. Biofuels 2018, 11, 213. [Google Scholar] [CrossRef]
- Hämäläinen, V.; Grönroos, T.; Suonpää, A.; Heikkilä, M.W.; Romein, B.; Ihalainen, P.; Malandra, S.; Birikh, K.R. Enzymatic processes to unlock the lignin value. Front. Bioeng. Biotechnol. 2018, 6, 20. [Google Scholar] [CrossRef]
- Mosbech, C.; Holck, J.; Meyer, A.S.; Agger, J.W. The natural catalytic function of CuGE glucuronoyl esterase in hydrolysis of genuine lignin–carbohydrate complexes from birch. Biotechnol. Biofuels 2018, 11, 71. [Google Scholar] [CrossRef]
- Ohbuchi, T.; Takahashi, T.; Azumi, N.; Sakaino, M. Structual analysis of neutral and acidic xylooligosaccharides from hardwood kraft pulp, and their utilization by intestinal bacteria in vitro. Biosci. Biotechnol. Biochem. 2009, 73, 2070–2076. [Google Scholar] [CrossRef]
- De Graaf, M.; Nevalainen, T.J.; Scheeren, H.W.; Pinedo, H.M.; Haisma, H.J.; Boven, E. A methylester of the glucuronide prodrug DOX-GA3 for improvement of tumor-selective chemotherapy. Biochem. Pharmacol. 2004, 68, 2273–2281. [Google Scholar] [CrossRef]
- Li, X.-L.; Špániková, S.; de Vries, R.P.; Biely, P. Identification of genes encoding microbial glucuronoyl esterases. FEBS Lett. 2007, 581, 4029–4035. [Google Scholar] [CrossRef]
- De Santi, C.; Willassen, N.P.; Williamson, A. Biochemical characterization of a family 15 carbohydrate esterase from a bacterial marine arctic metagenome. PLoS ONE 2016, 11, e0159345. [Google Scholar] [CrossRef]
- Aurilia, V.; Martin, J.C.; McCrae, S.I.; Scott, K.P.; Rincon, M.T.; Flint, H.J. Three multidomain esterases from the cellulolytic rumen anaerobe Ruminococcus flavefaciens 17 that carry divergent dockerin sequences. Microbiology 2000, 146, 1391–1397. [Google Scholar] [CrossRef]
- Biely, P.; Malovíková, A.; Uhliariková, I.; Li, X.-L.; Wong, D.W.S. Glucuronoyl esterases are active on the polymeric substrate methyl esterified glucuronoxylan. FEBS Lett. 2015, 589, 2334–2339. [Google Scholar] [CrossRef]
- Krska, D.; Mazurkewich, S.; Brown, H.A.; Theibich, Y.; Poulsen, J.-C.N.; Morris, A.L.; Koropatkin, N.M.; Lo Leggio, L.; Larsbrink, J. Structural and functional analysis of a multimodular hyperthermostable xylanase-glucuronoyl esterase from Caldicellulosiruptor kristjansonii. Biochemistry 2021, 60, 2206–2220. [Google Scholar] [CrossRef]
- Arnling Bååth, J.; Mazurkewich, S.; Poulsen, J.-C.N.; Olsson, L.; Lo Leggio, L.; Larsbrink, J. Structure and function analyses reveal that a glucuronoyl esterase from Teredinibacter turnerae interacts with carbohydrates and aromatic compounds. J. Biol. Chem. 2019, 294, 6635–6644. [Google Scholar] [CrossRef]
- Souza, C.P.; Almeida, B.C.; Colwell, R.R.; Rivera, I.N.G. The Importance of chitin in the marine environment. Mar. Biotechnol. 2011, 13, 823–830. [Google Scholar] [CrossRef]
- Coutinho, P.M.; Deleury, E.; Henrissat, B. The families of carbohydrate-active enzymes in the genomic era. J. Appl. Glycosci. 2003, 50, 241–244. [Google Scholar] [CrossRef]
- Naveed, M.; Phil, L.; Sohail, M.; Hasnat, M.; Baig, M.M.F.A.; Ihsan, A.U.; Shumzaid, M.; Kakar, M.U.; Mehmood Khan, T.; Akabar, M.D.; et al. Chitosan oligosaccharide (COS): An overview. Int. J. Biol. Macromol. 2019, 129, 827–843. [Google Scholar] [CrossRef] [PubMed]
- Vaaje-Kolstad, G.; Horn, S.J.; Sørlie, M.; Eijsink, V.G.H. The chitinolytic machinery of Serratia marcescens—A model system for enzymatic degradation of recalcitrant polysaccharides. FEBS J. 2013, 280, 3028–3049. [Google Scholar] [CrossRef] [PubMed]
- Tuveng, T.R.; Hagen, L.H.; Mekasha, S.; Frank, J.; Arntzen, M.Ø.; Vaaje-Kolstad, G.; Eijsink, V.G.H. Genomic, proteomic and biochemical analysis of the chitinolytic machinery of Serratia marcescens BJL200. Biochim. Biophys. Acta Proteins Proteom. 2017, 1865, 414–421. [Google Scholar] [CrossRef] [PubMed]
- Doan, C.T.; Tran, T.N.; Wang, S.-L. Production of thermophilic chitinase by Paenibacillus sp. TKU052 by bioprocessing of chitinous fishery wastes and its application in N-acetyl-D-glucosamine production. Polymers 2021, 13, 3048. [Google Scholar] [CrossRef] [PubMed]
- Sousa, A.J.S.; Silva, C.F.B.; Sousa, J.S.; Monteiro, J.E.; Freire, J.E.C.; Sousa, B.L.; Lobo, M.D.P.; Monteiro-Moreira, A.C.O.; Grangeiro, T.B. A thermostable chitinase from the antagonistic Chromobacterium violaceum that inhibits the development of phytopathogenic fungi. Enzyme Microb. Technol. 2019, 126, 50–61. [Google Scholar] [CrossRef]
- Chen, L.; Wei, Y.; Shi, M.; Li, Z.; Zhang, S.-H. An archaeal chitinase with a secondary capacity for catalyzing cellulose and its biotechnological applications in shell and straw degradation. Front. Microbiol. 2019, 10, 1253. [Google Scholar] [CrossRef]
- Sorokin, D.Y.; Toshchakov, S.V.; Kolganova, T.V.; Kublanov, I.V. Halo(natrono)archaea isolated from hypersaline lakes utilize cellulose and chitin as growth substrates. Front. Microbiol. 2015, 6, 942. [Google Scholar] [CrossRef]
- Nishitani, Y.; Horiuchi, A.; Aslam, M.; Kanai, T.; Atomi, H.; Miki, K. Crystal structures of an archaeal chitinase ChiD and its ligand complexes. Glycobiology 2018, 28, 418–426. [Google Scholar] [CrossRef]
- Grifoll-Romero, L.; Pascual, S.; Aragunde, H.; Biarnés, X.; Planas, A. Chitin deacetylases: Structures, specificities, and biotech applications. Polymers 2018, 10, 352. [Google Scholar] [CrossRef]
- Bonin, M.; Sreekumar, S.; Cord-Landwehr, S.; Moerschbacher, B.M. Preparation of defined chitosan oligosaccharides using chitin deacetylases. Int. J. Mol. Sci. 2020, 21, 7835. [Google Scholar] [CrossRef]
- Cheung, R.C.; Ng, T.B.; Wong, J.H.; Chan, W.Y. Chitosan: An update on potential biomedical and pharmaceutical applications. Mar. Drugs 2015, 13, 5156–5186. [Google Scholar] [CrossRef]
- Zhang, J.; Yan, N. Production of glucosamine from chitin by co-solvent promoted hydrolysis and deacetylation. ChemCatChem 2017, 9, 2790–2796. [Google Scholar] [CrossRef]
- Mine, S.; Niiyama, M.; Hashimoto, W.; Ikegami, T.; Koma, D.; Ohmoto, T.; Fukuda, Y.; Inoue, T.; Abe, Y.; Ueda, T.; et al. Expression from engineered Escherichia coli chromosome and crystallographic study of archaeal N,N′-diacetylchitobiose deacetylase. FEBS J. 2014, 281, 2584–2596. [Google Scholar] [CrossRef]
- Huang, Z.; Lv, X.; Sun, G.; Mao, X.; Lu, W.; Liu, Y.; Li, J.; Du, G.; Liu, L. Chitin deacetylase: From molecular structure to practical applications. Syst. Microbiol. Biomanufact. 2022, 2, 271–284. [Google Scholar] [CrossRef]
- Ding, Z.; Ahmed, S.; Hang, J.; Mi, H.; Hou, X.; Yang, G.; Huang, Z.; Lu, X.; Zhang, W.; Liu, S.; et al. Rationally engineered chitin deacetylase from Arthrobacter sp. AW19M34-1 with improved catalytic activity toward crystalline chitin. Carbohyd. Polym. 2021, 274, 118637. [Google Scholar] [CrossRef]
- Pawaskar, G.M.; Raval, K.; Rohit, P.; Shenoy, R.P.; Raval, R. Cloning, expression, purification and characterization of chitin deacetylase extremozyme from halophilic Bacillus aryabhattai B8W22. 3 Biotech 2021, 11, 515. [Google Scholar] [CrossRef]
- Bhat, P.; Pawaskar, G.-M.; Raval, R.; Cord-Landwehr, S.; Moerschbacher, B.; Raval, K. Expression of Bacillus licheniformis chitin deacetylase in E. coli pLysS: Sustainable production, purification and characterisation. Int. J. Biol. Macromol. 2019, 131, 1008–1013. [Google Scholar] [CrossRef]
- Tanaka, T.; Fukui, T.; Fujiwara, S.; Atomi, H.; Imanaka, T. Concerted action of diacetylchitobiose deacetylase and exo-beta-D-glucosaminidase in a novel chitinolytic pathway in the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J. Biol. Chem. 2004, 279, 30021–30027. [Google Scholar] [CrossRef]
- Mine, S.; Ikegami, T.; Kawasaki, K.; Nakamura, T.; Uegaki, K. Expression, refolding, and purification of active diacetylchitobiose deacetylase from Pyrococcus horikoshii. Protein Expr. Purif. 2012, 84, 265–269. [Google Scholar] [CrossRef]
- Huang, Z.; Mao, X.; Lv, X.; Sun, G.; Zhang, H.; Lu, W.; Liu, Y.; Li, J.; Du, G.; Liu, L. Engineering diacetylchitobiose deacetylase from Pyrococcus horikoshii towards an efficient glucosamine production. Bioresour. Technol. 2021, 334, 125241. [Google Scholar] [CrossRef] [PubMed]
- Rani Singhania, R.; Dixit, P.; Kumar Patel, A.; Shekher Giri, B.; Kuo, C.-H.; Chen, C.-W.; Di Dong, C. Role and significance of lytic polysaccharide monooxygenases (LPMOs) in lignocellulose deconstruction. Bioresour. Technol. 2021, 335, 125261. [Google Scholar] [CrossRef] [PubMed]
| Plastid Transformation Method | Strengths | Weaknesses | Reference |
|---|---|---|---|
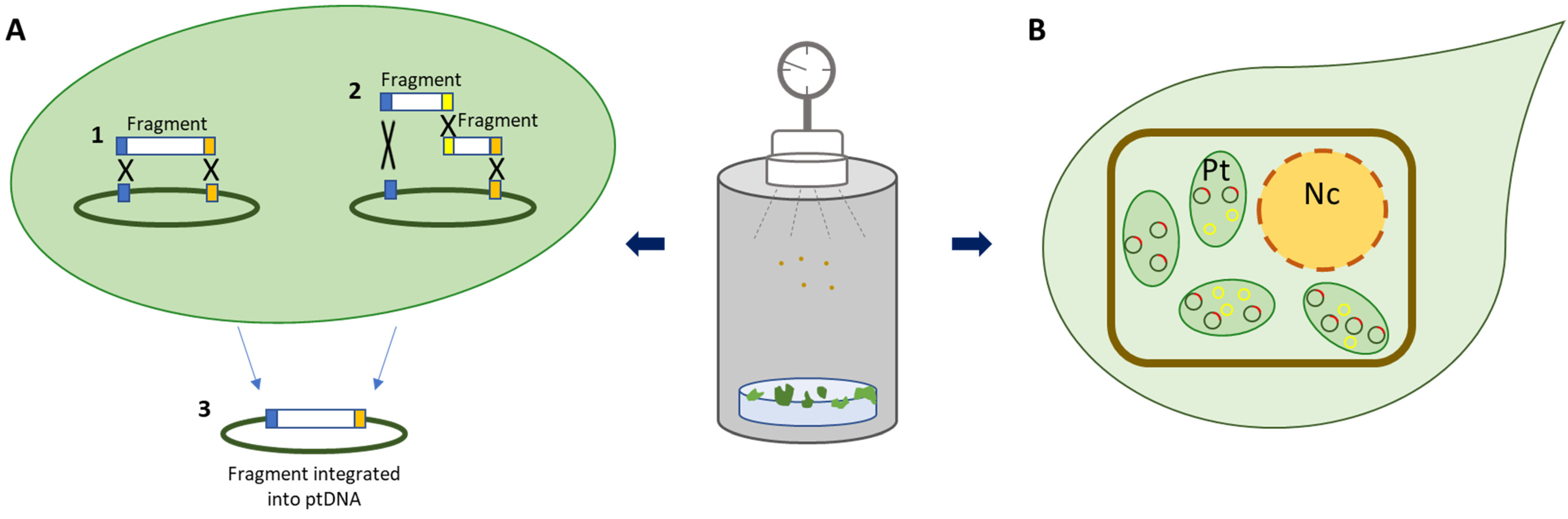
| Linear DNA fragments | No vector construction required | Sequencing of linear fragments required | [25] |
| Minichromosome | Stable and efficient amplification of foreign DNA | Need to use transplastomic plants expressing the Rep protein as plant host | [27] |
| Fusion peptide KH-AtOEP34 | Reduction of time-consuming steps | No homoplasmic plants obtained | |
| Applicable to recalcitrant crops (rice, kenaf) | Low transformation efficiency | [28] | |
| Possible integration of exogenous DNA onto nuclear DNA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tamburino, R.; Marcolongo, L.; Sannino, L.; Ionata, E.; Scotti, N. Plastid Transformation: New Challenges in the Circular Economy Era. Int. J. Mol. Sci. 2022, 23, 15254. https://doi.org/10.3390/ijms232315254
Tamburino R, Marcolongo L, Sannino L, Ionata E, Scotti N. Plastid Transformation: New Challenges in the Circular Economy Era. International Journal of Molecular Sciences. 2022; 23(23):15254. https://doi.org/10.3390/ijms232315254
Chicago/Turabian StyleTamburino, Rachele, Loredana Marcolongo, Lorenza Sannino, Elena Ionata, and Nunzia Scotti. 2022. "Plastid Transformation: New Challenges in the Circular Economy Era" International Journal of Molecular Sciences 23, no. 23: 15254. https://doi.org/10.3390/ijms232315254
APA StyleTamburino, R., Marcolongo, L., Sannino, L., Ionata, E., & Scotti, N. (2022). Plastid Transformation: New Challenges in the Circular Economy Era. International Journal of Molecular Sciences, 23(23), 15254. https://doi.org/10.3390/ijms232315254







