Genome-Wide Analysis of AP2/ERF Gene Superfamily in Ramie (Boehmeria nivea L.) Revealed Their Synergistic Roles in Regulating Abiotic Stress Resistance and Ramet Development
Abstract
:1. Introduction
2. Results
2.1. Identification, Characteristics, Phylogeny Relationship, and Classification of AP2/ERF Transcription Factors in Ramie
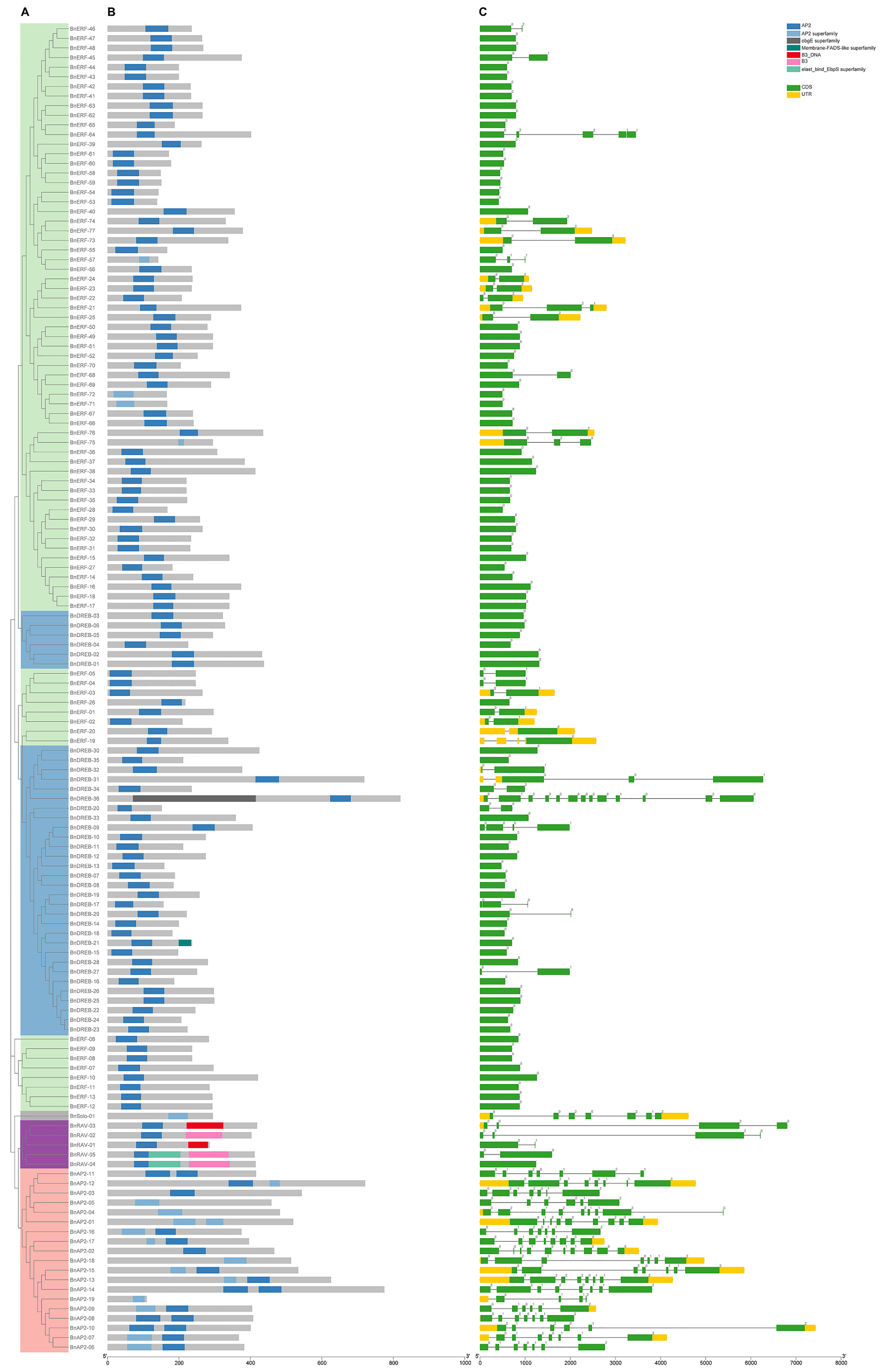
2.2. Gene Structure and Conserved Motifs Analysis of BnAP2/ERFs
2.3. Chromosome Distribution and Gene Duplication of BnAP2/ERFs
2.4. The Putative Promoter Regions Analysis of BnAP2/ERF Subfamily
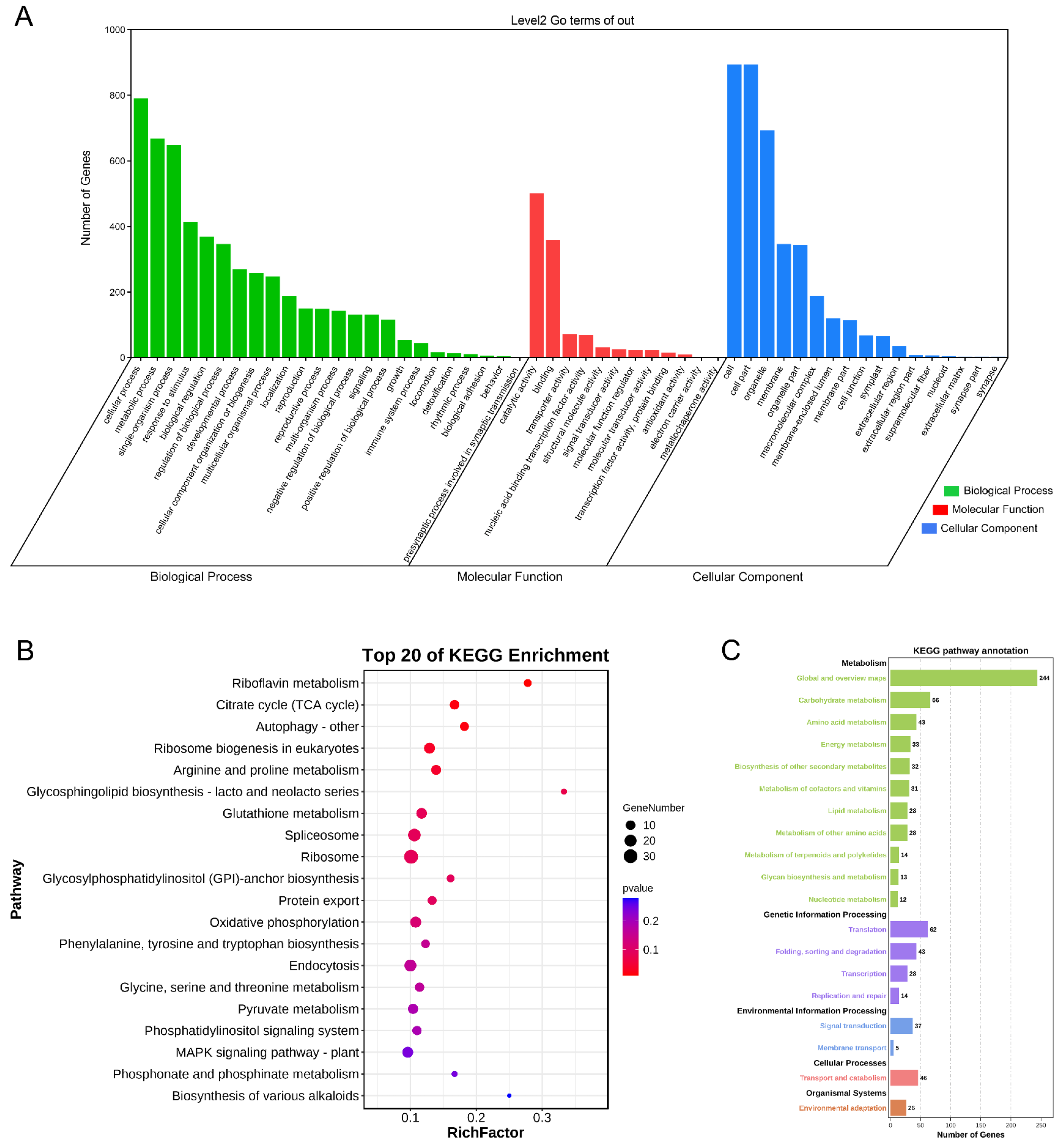
2.5. Gene Ontology Annotation and KEGG Enrichment Analysis of BnAP2/ERF Target Genes
2.6. Expression Analysis of BnAP2/ERF Genes Based on Transcriptome Data
2.6.1. Tissue Specific Expression of BnAP2/ERF Genes
2.6.2. Expression Patterns of BnAP2/ERF Genes in Response to Various Abiotic Stresses
2.6.3. Expression Patterns of BnAP2/ERF Genes in Various Ramie Varieties with Significantly Different Ramet Numbers
2.6.4. Verification of Gene Expression by qPCR
3. Discussion
3.1. Global Profile of AP2/ERF Gene Family of Ramie
3.2. The Roles of the BnAP2/ERF Gene Family in Responding to Abiotic Stresses
3.3. Candidate Genes for Improving Waterlogging Tolerance Coupling with Ramet Development
4. Materials and Methods
4.1. Identification of BnAP2/ERF Gene Superfamily
4.2. Phylogeny, Conserved Motifs, and Gene Structure Analysis
4.3. Chromosome Distribution, Gene Duplication, and Evolutionary Analysis of AP2/ERF Homologous Genes
4.4. Cis-Element Analysis
4.5. KEGG Enrichment Analysis and Gene Ontology Functional Annotation of BnAP2/ERFs Target Genes
4.6. Transcriptome Data Sources and Expression Analysis of the BnAP2/ERF Genes
4.7. BnAP2/ERF Protein–Protein Interaction Network Prediction
4.8. Plant Materials and Sampling
4.9. RNA Sample Extraction and qPCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chen, J.; Gao, G.; Yu, C.; Chen, P.; Chen, K.; Wang, X.; Bai, L.; Zhu, A. Ramie BnALDH genes and their potential role involved in adaptation to hydroponic culturing condition. Indl Crops Prod. 2020, 157, 112928. [Google Scholar] [CrossRef]
- Luan, M.B.; Jian, J.B.; Chen, P.; Chen, J.H.; Chen, J.H.; Gao, Q.; Gao, G.; Zhou, J.H.; Chen, K.M.; Guang, X.M.; et al. Draft genome sequence of ramie, Boehmeria nivea (L.) Gaudich. Mol. Ecol. Resour. 2018, 18, 639–645. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Zhu, S.; Tang, Q.; Yu, Y.; Tang, S. Identification of drought stress-responsive transcription factors in ramie (Boehmeria nivea L. Gaud). BMC Plant Biol. 2013, 13, 130. [Google Scholar] [CrossRef] [Green Version]
- Kuluev, B.; Avalbaev, A.; Nurgaleeva, E.; Knyazev, A.; Nikonorov, Y.; Chemeris, A. Role of AINTEGUMENTA-like gene NtANTL in the regulation of tobacco organ growth. J. Plant Physiol. 2015, 189, 11–23. [Google Scholar] [CrossRef] [PubMed]
- Pan, D.L.; Wang, G.; Wang, T.; Jia, Z.H.; Guo, Z.R.; Zhang, J.Y. AdRAP2.3, a novel ethylene response factor VII from Actinidia deliciosa, enhances waterlogging resistance in transgenic tobacco through improving expression levels of PDC and ADH genes. Int. J. Mol. Sci. 2019, 20, 1189. [Google Scholar] [CrossRef] [Green Version]
- Xie, Z.; Nolan, T.; Jiang, H.; Tang, B.; Zhang, M.; Li, Z.; Yin, Y. The AP2/ERF transcription factor TINY modulates brassinosteroid-regulated plant growth and drought responses in Arabidopsis. Plant Cell. 2019, 31, 1788–1806. [Google Scholar] [CrossRef] [Green Version]
- Ritonga, F.N.; Ngatia, J.N.; Wang, Y.; Khoso, M.A.; Farooq, U.; Chen, S. AP2/ERF, an important cold stress-related transcription factor family in plants: A review. Physiol. Mol. Biol. Plants 2021, 27, 1953–1968. [Google Scholar] [CrossRef]
- Zhang, T.; Tang, Y.; Luan, Y.; Cheng, Z.; Wang, X.; Tao, J.; Zhao, D. Herbaceous peony AP2/ERF transcription factor binds the promoter of the tryptophan decarboxylase gene to enhance high-temperature stress tolerance. Plant Cell Environ. 2022, 45, 2729–2743. [Google Scholar] [CrossRef]
- Lv, K.; Li, J.; Zhao, K.; Chen, S.; Nie, J.; Zhang, W.; Liu, G.; Wei, H. Overexpression of an AP2/ERF family gene, BpERF13, in birch enhances cold tolerance through upregulating CBF genes and mitigating reactive oxygen species. Plant Sci. 2020, 292, 110375. [Google Scholar] [CrossRef] [PubMed]
- Yin, F.; Zeng, Y.; Ji, J.; Wang, P.; Zhang, Y.; Li, W. The halophyte Halostachys caspica AP2/ERF transcription factor HcTOE3 positively regulates freezing tolerance in Arabidopsis. Front. Plant Sci. 2021, 12, 638788. [Google Scholar] [CrossRef] [PubMed]
- Ye, B.B.; Shang, G.D.; Pan, Y.; Xu, Z.G.; Zhou, C.M.; Mao, Y.B.; Bao, N.; Sun, L.; Xu, T.; Wang, J.W. AP2/ERF transcription factors integrate age and wound signals for root regeneration. Plant Cell. 2020, 32, 226–241. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vahala, J.; Felten, J.; Love, J.; Gorzsás, A.; Gerber, L.; Lamminmäki, A.; Kangasjärvi, J.; Sundberg, B. A genome-wide screen for ethylene-induced ethylene response factors (ERFs) in hybrid aspen stem identifies ERF genes that modify stem growth and wood properties. New Phytol. 2013, 200, 511–522. [Google Scholar] [CrossRef]
- Marsch-Martinez, N.; Greco, R.; Becker, J.D.; Dixit, S.; Bergervoet, J.H.; Karaba, A.; de Folter, S.; Pereira, A. BOLITA, an Arabidopsis AP2/ERF-like transcription factor that affects cell expansion and proliferation/differentiation pathways. Plant Mol. Biol. 2006, 62, 825–843. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yant, L.; Mathieu, J.; Dinh, T.T.; Ott, F.; Lanz, C.; Wollmann, H.; Chen, X.; Schmid, M. Orchestration of the floral transition and floral development in Arabidopsis by the bifunctional transcription factor APETALA2. Plant Cell. 2010, 22, 2156–2170. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharma, P.; Watts, A.; Kumar, V.; Srinivasan, R.; Siwach, P. Cloning, characterization and expression analysis of APETALA2 genes of Brassica juncea (L.) Czern. Indian J. Exp. Biol. 2018, 56, 604–610. [Google Scholar]
- Wang, C.; Wang, H.; Zhang, J.; Chen, S. A seed-specific AP2-domain transcription factor from soybean plays a certain role in regulation of seed germination. Sci. China C Life Sci. 2008, 51, 336–345. [Google Scholar] [CrossRef]
- Deng, H.; Chen, Y.; Liu, Z.; Liu, Z.; Shu, P.; Wang, R.; Hao, Y.; Su, D.; Pirrello, J.; Liu, Y.; et al. SlERF.F12 modulates the transition to ripening in tomato fruit by recruiting the co-repressor TOPLESS and histone deacetylases to repress key ripening genes. Plant Cell 2022, 34, 1250–1272. [Google Scholar] [CrossRef]
- Wu, K.; Wang, S.; Song, W.; Zhang, J.; Wang, Y.; Liu, Q.; Yu, J.; Ye, Y.; Li, S.; Chen, J.; et al. Enhanced sustainable green revolution yield via nitrogen-responsive chromatin modulation in rice. Science 2020, 367, eaaz2046. [Google Scholar] [CrossRef]
- Lyu, J.; Guo, Y.; Du, C.; Yu, H.; Guo, L.; Liu, L.; Zhao, H.; Wang, X.; Hu, S. BnERF114.A1, a rapeseed gene encoding APETALA2/ETHYLENE RESPONSE FACTOR, regulates plant architecture through auxin accumulation in the apex in Arabidopsis. Int. J. Mol. Sci. 2022, 23, 2210. [Google Scholar] [CrossRef]
- Cui, L.; Feng, K.; Wang, M.; Wang, M.; Deng, P.; Song, W.; Nie, X. Genome-wide identification, phylogeny and expression analysis of AP2/ERF transcription factors family in Brachypodium distachyon. BMC Genom. 2016, 17, 636. [Google Scholar] [CrossRef] [Green Version]
- Lynch, M.; Conery, J.S. The evolutionary fate and consequences of duplicate genes. Science 2000, 290, 1151–1155. [Google Scholar] [CrossRef]
- Kabir, S.M.T.; Hossain, M.S.; Bashar, K.K.; Honi, U.; Ahmed, B.; Emdad, E.M.; Alam, M.M.; Haque, M.S.; Islam, M.S. Genome-wide identification and expression profiling of AP2/ERF superfamily genes under stress conditions in dark jute (Corchorus olitorius L.). Ind. Crops Prod. 2021, 166, 113469. [Google Scholar] [CrossRef]
- Abubakar, A.S.; Feng, X.; Gao, G.; Yu, C.; Chen, J.; Chen, K.; Wang, X.; Mou, P.; Shao, D.; Chen, P.; et al. Genome wide characterization of R2R3 MYB transcription factor from Apocynum venetum revealed potential stress tolerance and flavonoid biosynthesis genes. Genomics 2022, 114, 110275. [Google Scholar] [CrossRef] [PubMed]
- Chandler, J.W. Class VIIIb APETALA2 ethylene response factors in plant development. Trends Plant Sci. 2018, 23, 151–162. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.S.; Chen, M.; Li, L.C.; Ma, Y.Z. Functions and application of the AP2/ERF transcription factor family in crop improvement. J. Integr. Plant Biol. 2011, 53, 570–585. [Google Scholar] [CrossRef]
- Xie, Z.; Nolan, T.M.; Jiang, H.; Yin, Y. AP2/ERF transcription factor regulatory networks in hormone and abiotic stress responses in Arabidopsis. Front. Plant Sci. 2019, 10, 228. [Google Scholar] [CrossRef] [Green Version]
- Sakamoto, S.; Somssich, M.; Nakata, M.T.; Unda, F.; Atsuzawa, K.; Kaneko, Y.; Wang, T.; Bågman, A.M.; Gaudinier, A.; Yoshida, K.; et al. Complete substitution of a secondary cell wall with a primary cell wall in Arabidopsis. Nat. Plants 2018, 4, 777–783. [Google Scholar] [CrossRef]
- Li, Z.; Sheerin, D.J.; von Roepenack-Lahaye, E.; Stahl, M.; Hiltbrunner, A. The phytochrome interacting proteins ERF55 and ERF58 repress light-induced seed germination in Arabidopsis thaliana. Nat. Commun. 2022, 13, 1656. [Google Scholar] [CrossRef]
- Agarwal, G.; Garg, V.; Kudapa, H.; Doddamani, D.; Pazhamala, L.T.; Khan, A.W.; Thudi, M.; Lee, S.H.; Varshney, R.K. Genome-wide dissection of AP2/ERF and HSP90 gene families in five legumes and expression profiles in chickpea and pigeonpea. Plant Biotechnol. J. 2016, 14, 1563–1577. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Wang, Y.; Wu, M.; Li, L.; Li, C.; Han, Z.; Yuan, J.; Chen, C.; Song, W.; Wang, C. Genome-wide identification of AP2/ERF transcription factors in Cauliflower and expression profiling of the ERF family under salt and drought stresses. Front. Plant Sci. 2017, 8, 946. [Google Scholar] [CrossRef]
- Li, X.; Gao, B.; Zhang, D.; Liang, Y.; Liu, X.; Zhao, J.; Zhang, J.; Wood, A.J. Identification, classification, and functional analysis of AP2/ERF family genes in the desert moss Bryum argenteum. Int. J. Mol. Sci. 2018, 19, 3637. [Google Scholar] [CrossRef] [PubMed]
- Xing, H.; Jiang, Y.; Zou, Y.; Long, X.; Wu, X.; Ren, Y.; Li, Y.; Li, H.L. Genome-wide investigation of the AP2/ERF gene family in ginger: Evolution and expression profiling during development and abiotic stresses. BMC Plant Biol. 2021, 21, 561. [Google Scholar] [CrossRef] [PubMed]
- Rogozin, I.B.; Carmel, L.; Csuros, M.; Koonin, E.V. Origin and evolution of spliceosomal introns. Biol. Direct. 2012, 7, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, B.; Yao, X.; Zeng, Y.; Zhang, C. Genome-wide identification, characterization, and expression profiling of AP2/ERF superfamily genes under different development and abiotic stress conditions in pecan (Carya illinoinensis). Int. J. Mol. Sci. 2022, 23, 2920. [Google Scholar] [CrossRef] [PubMed]
- Parenteau, J.; Maignon, L.; Berthoumieux, M.; Catala, M.; Gagnon, V.; Abou Elela, S. Introns are mediators of cell response to starvation. Nature 2019, 565, 612–617. [Google Scholar] [CrossRef] [PubMed]
- Riechmann, J.L.; Meyerowitz, E.M. The AP2/EREBP family of plant transcription factors. Biol. Chem. 1998, 379, 633–646. [Google Scholar] [CrossRef] [PubMed]
- Allen, M.D.; Yamasaki, K.; Ohme-Takagi, M.; Tateno, M.; Suzuki, M. A novel mode of DNA recognition by a beta-sheet revealed by the solution structure of the GCC-box binding domain in complex with DNA. Embo. J. 1998, 17, 5484–5496. [Google Scholar] [CrossRef] [Green Version]
- De Lucas, M.; Pu, L.; Turco, G.; Gaudinier, A.; Morao, A.K.; Harashima, H.; Kim, D.; Ron, M.; Sugimoto, K.; Roudier, F.; et al. Transcriptional regulation of Arabidopsis polycomb repressive complex 2 coordinates cell-type proliferation and differentiation. Plant Cell 2016, 28, 2616–2631. [Google Scholar] [CrossRef] [Green Version]
- Riester, L.; Köster-Hofmann, S.; Doll, J.; Berendzen, K.W.; Zentgraf, U. Impact of alternatively polyadenylated isoforms of ETHYLENE RESPONSE FACTOR4 with activator and repressor function on senescence in Arabidopsis thaliana L. Genes 2019, 10, 91. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z.; Tian, L.; Latoszek-Green, M.; Brown, D.; Wu, K. Arabidopsis ERF4 is a transcriptional repressor capable of modulating ethylene and abscisic acid responses. Plant Mol. Biol. 2005, 58, 585–596. [Google Scholar] [CrossRef]
- Zhou, X.; Zhang, Z.L.; Park, J.; Tyler, L.; Yusuke, J.; Qiu, K.; Nam, E.A.; Lumba, S.; Desveaux, D.; McCourt, P.; et al. The ERF11 transcription factor promotes internode elongation by activating gibberellin biosynthesis and signaling. Plant Physiol. 2016, 171, 2760–2770. [Google Scholar] [CrossRef]
- Wei, X.; Xu, H.; Rong, W.; Ye, X.; Zhang, Z. Constitutive expression of a stabilized transcription factor group VII ethylene response factor enhances waterlogging tolerance in wheat without penalizing grain yield. Plant Cell Environ. 2019, 42, 1471–1485. [Google Scholar] [CrossRef]
- Yu, F.; Liang, K.; Fang, T.; Zhao, H.; Han, X.; Cai, M.; Qiu, F. A group VII ethylene response factor gene, ZmEREB180, coordinates waterlogging tolerance in maize seedlings. Plant Biotechnol. J. 2019, 17, 2286–2298. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; Tan, W.J.; Xie, L.J.; Qi, H.; Yang, Y.C.; Huang, L.P.; Lai, Y.X.; Tan, Y.F.; Zhou, D.M.; Yu, L.J.; et al. Polyunsaturated linolenoyl-CoA modulates ERF-VII-mediated hypoxia signaling in Arabidopsis. J. Integr. Plant Biol. 2020, 62, 330–348. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Dossa, K.; You, J.; Zhang, Y.; Li, D.; Zhou, R.; Yu, J.; Wei, X.; Zhu, X.; Jiang, S.; et al. High-resolution temporal transcriptome sequencing unravels ERF and WRKY as the master players in the regulatory networks underlying sesame responses to waterlogging and recovery. Genomics 2021, 113 Pt 1, 276–290. [Google Scholar] [CrossRef] [PubMed]
- Kondrashov, F.A.; Rogozin, I.B.; Wolf, Y.I.; Koonin, E.V. Selection in the evolution of gene duplications. Genome Biol. 2002, 3, research0008. [Google Scholar] [CrossRef] [Green Version]
- Li, M.-Y.; Wang, F.; Jiang, Q.; Li, R.; Ma, J.; Xiong, A.-S. Genome-wide analysis of the distribution of AP2/ERF transcription factors reveals duplication and elucidates their potential function in chinese cabbage (Brassica rapa ssp pekinensis). Plant Mol. Biol. Rep. 2013, 31, 1002–1011. [Google Scholar] [CrossRef]
- Lata, C.; Mishra, A.K.; Muthamilarasan, M.; Bonthala, V.S.; Khan, Y.; Prasad, M. Genome-wide investigation and expression profiling of AP2/ERF transcription factor superfamily in foxtail millet (Setaria italica L.). PLoS ONE 2014, 9, e113092. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fukao, T.; Barrera-Figueroa, B.E.; Juntawong, P.; Peña-Castro, J.M. Submergence and waterlogging stress in plants: A review highlighting research opportunities and understudied aspects. Front. Plant Sci. 2019, 10, 340. [Google Scholar] [CrossRef] [PubMed]
- Roeber, V.M.; Bajaj, I.; Rohde, M.; Schmülling, T.; Cortleven, A. Light acts as a stressor and influences abiotic and biotic stress responses in plants. Plant. Cell Environ. 2021, 44, 645–664. [Google Scholar] [CrossRef]
- Zhang, H.; Zhao, Y.; Zhu, J.K. Thriving under stress: How plants balance growth and the stress response. Dev. Cell 2020, 55, 529–543. [Google Scholar] [CrossRef]
- Rasheed, A.; Jie, Y.; Nawaz, M.; Jie, H.; Ma, Y.; Shah, A.N.; Hassan, M.U.; Gillani, S.F.A.; Batool, M.; Aslam, M.T.; et al. Improving drought stress tolerance in ramie (Boehmeria nivea L.) using molecular techniques. Front. Plant Sci. 2022, 13, 911610. [Google Scholar] [CrossRef]
- Xu, Y.; Tang, Q.; Dai, Z.; Yang, Z.; Cheng, C.; Deng, C.; Liu, C.; Chen, J.; Su, J. Yield components of forage ramie (Boehmeria nivea L.) and their effects on yield. Genet. Resour. Crop Evol. 2019, 66, 1601–1613. [Google Scholar] [CrossRef]
- Lopes, F.L.; Galvan-Ampudia, C.; Landrein, B. WUSCHEL in the shoot apical meristem: Old player, new tricks. J. Exp. Bot. 2021, 72, 1527–1535. [Google Scholar] [CrossRef]
- Wei, T.; Zhang, L.; Zhu, R.; Jiang, X.; Yue, C.; Su, Y.; Ren, H.; Wang, M. A Gain-of-function mutant of IAA7 inhibits stem elongation by transcriptional repression of EXPA5 genes in Brassica napus. Int. J. Mol. Sci. 2021, 22, 9018. [Google Scholar] [CrossRef]
- Zhang, M.M.; Zhang, H.K.; Zhai, J.F.; Zhang, X.S.; Sang, Y.L.; Cheng, Z.J. ARF4 regulates shoot regeneration through coordination with ARF5 and IAA12. Plant Cell Rep. 2021, 40, 315–325. [Google Scholar] [CrossRef]
- Simonini, S.; Deb, J.; Moubayidin, L.; Stephenson, P.; Valluru, M.; Freire-Rios, A.; Sorefan, K.; Weijers, D.; Friml, J.; Østergaard, L. A noncanonical auxin-sensing mechanism is required for organ morphogenesis in Arabidopsis. Genes Dev. 2016, 30, 2286–2296. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Hu, Q.; Luo, S.; Li, Q.; Yang, X.; Wang, X.; Wang, S. Expression of wild-type PtrIAA14.1, a poplar Aux/IAA gene causes morphological changes in Arabidopsis. Front. Plant Sci. 2015, 6, 388. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shukla, V.; Lombardi, L.; Pencik, A.; Novak, O.; Weits, D.A.; Loreti, E.; Perata, P.; Giuntoli, B.; Licausi, F. Jasmonate signalling contributes to primary root inhibition upon oxygen deficiency in Arabidopsis thaliana. Plants 2020, 9, 1046. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, N.; Zhan, J.; Wang, X.; Zhou, X.R.; Shi, J.; Wang, H. Genome-wide analysis of the JAZ subfamily of transcription factors and functional verification of BnC08.JAZ1-1 in Brassica napus. Biotechnol. Biofuels Bioprod. 2022, 15, 93. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.B.; Dai, Y.S.; Xie, L.J.; Yu, L.J.; Zhou, Y.; Lai, Y.X.; Yang, Y.C.; Xu, L.; Chen, Q.F.; Xiao, S. Jasmonate regulates plant responses to postsubmergence reoxygenation through transcriptional activation of antioxidant Synthesis. Plant Physiol. 2017, 173, 1864–1880. [Google Scholar] [CrossRef]
- Luo, F.L.; Nagel, K.A.; Zeng, B.; Schurr, U.; Matsubara, S. Photosynthetic acclimation is important for post-submergence recovery of photosynthesis and growth in two riparian species. Ann. Bot. 2009, 104, 1435–1444. [Google Scholar] [CrossRef] [PubMed]
- Arduini, I.; Baldanzi, M.; Pampana, S. Reduced growth and nitrogen uptake during waterlogging at tillering permanently affect yield components in late sown oats. Front. Plant Sci. 2019, 10, 1087. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El-Gebali, S.; Mistry, J.; Bateman, A.; Eddy, S.R.; Luciani, A.; Potter, S.C.; Qureshi, M.; Richardson, L.J.; Salazar, G.A.; Smart, A.; et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019, 47, D427–D432. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: A toolkit incorporating gamma-series methods and sliding window strategies. Genom. Proteom. Bioinf. 2010, 8, 77–80. [Google Scholar] [CrossRef] [Green Version]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, Y.; Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, D353–D361. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Muruganujan, A.; Ebert, D.; Huang, X.; Thomas, P.D. PANTHER version 14: More genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 2019, 47, D419–D426. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Gao, G.; Chen, P.; Chen, K.; Wang, X.; Bai, L.; Yu, C.; Zhu, A. Integrative transcriptome and proteome analysis identifies major molecular regulation pathways involved in ramie (Boehmeria nivea (L.) Gaudich) under nitrogen and water co-limitation. Plants 2020, 9, 1267. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Dong, Z.; Fang, L.; Luo, Y.; Wei, Z.; Guo, H.; Zhang, G.; Gu, Y.Q.; Coleman-Derr, D.; Xia, Q.; et al. OrthoVenn2: A web server for whole-genome comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res. 2019, 47, W52–W58. [Google Scholar] [CrossRef] [Green Version]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef]
- Ren, B.; Hu, J.; Liu, P.; Zhao, B.; Zhang, J. Responses of nitrogen efficiency and antioxidant system of summer maize to waterlogging stress under different tillage. PeerJ 2021, 9, e11834. [Google Scholar] [CrossRef]
- Tan, L.; Gao, G.; Yu, C.; Zhu, A.; Chen, P.; Chen, K.; Chen, J.; Xiong, H. Transcriptome analysis of high-NUE (T29) and low-NUE (T13) genotypes identified different responsive patterns involved in nitrogen stress in ramie (Boehmeria nivea (L.) Gaudich). Plants 2020, 9, 767. [Google Scholar] [CrossRef]
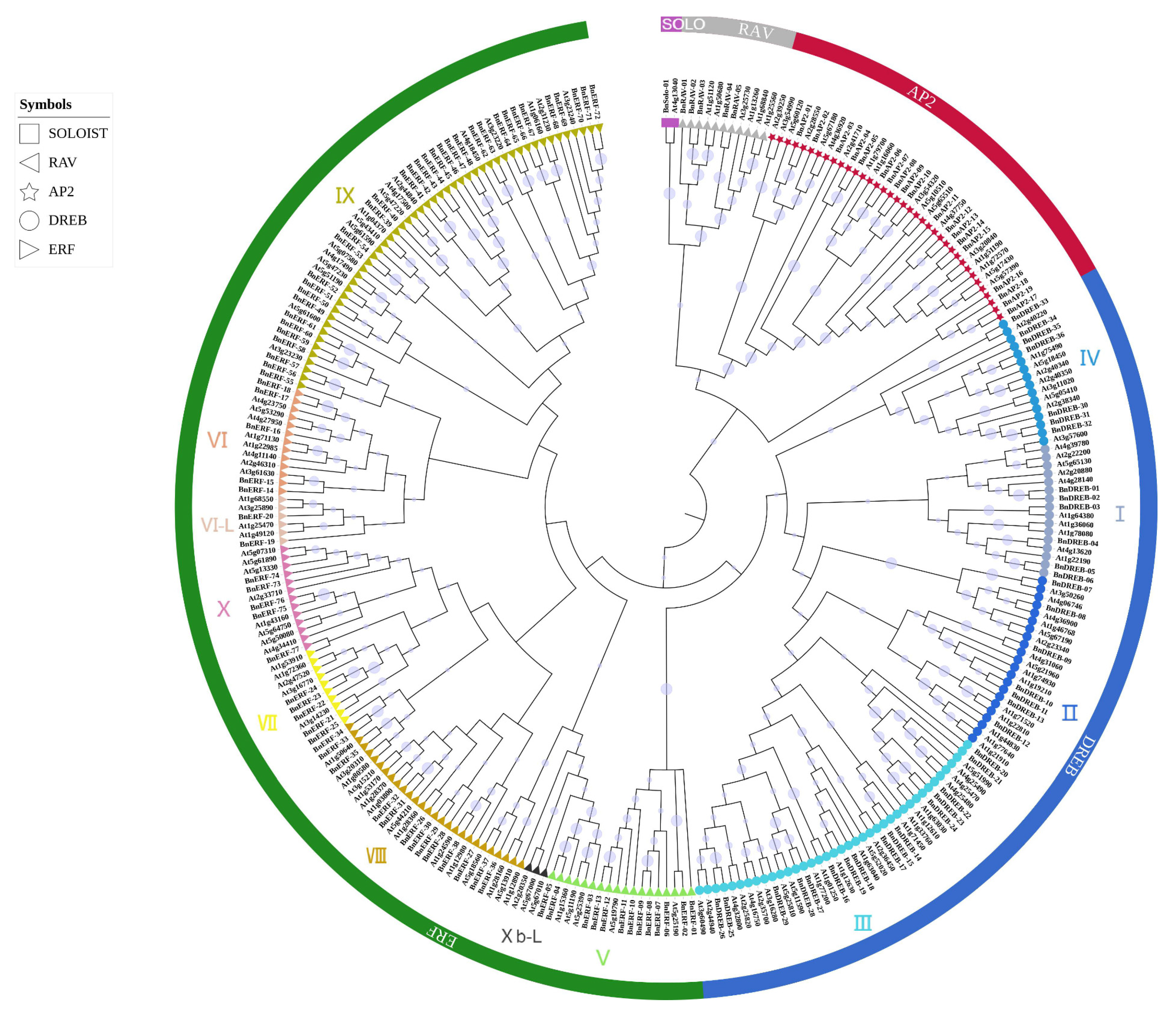








| Arabidopsis | Ramie | |||
|---|---|---|---|---|
| Classification | Group | No. | Group | No. |
| AP2 family | Total | 18 | Total | 19 |
| Double AP2/ERF domain | 14 | Double AP2/ERF domain | 13 | |
| Single AP2/ERF domain | 4 | Single AP2/ERF domain | 6 | |
| ERF family | Total | 122 | Total | 113 |
| DREB subfamily | Groups I to IV | 57 | Groups I to IV | 36 |
| ERF subfamily | Groups V to X | 58 | Groups V to X | 75 |
| ERF subfamily | Groups VI-L and Xb-L | 7 | Groups VI to L | 2 |
| Soloist | At4g13040 | 1 | BnSolo-01 | 1 |
| RAV family | 6 | 5 | ||
| Total | 147 | Total | 138 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qiu, X.; Zhao, H.; Abubakar, A.S.; Shao, D.; Chen, J.; Chen, P.; Yu, C.; Wang, X.; Chen, K.; Zhu, A. Genome-Wide Analysis of AP2/ERF Gene Superfamily in Ramie (Boehmeria nivea L.) Revealed Their Synergistic Roles in Regulating Abiotic Stress Resistance and Ramet Development. Int. J. Mol. Sci. 2022, 23, 15117. https://doi.org/10.3390/ijms232315117
Qiu X, Zhao H, Abubakar AS, Shao D, Chen J, Chen P, Yu C, Wang X, Chen K, Zhu A. Genome-Wide Analysis of AP2/ERF Gene Superfamily in Ramie (Boehmeria nivea L.) Revealed Their Synergistic Roles in Regulating Abiotic Stress Resistance and Ramet Development. International Journal of Molecular Sciences. 2022; 23(23):15117. https://doi.org/10.3390/ijms232315117
Chicago/Turabian StyleQiu, Xiaojun, Haohan Zhao, Aminu Shehu Abubakar, Deyi Shao, Jikang Chen, Ping Chen, Chunming Yu, Xiaofei Wang, Kunmei Chen, and Aiguo Zhu. 2022. "Genome-Wide Analysis of AP2/ERF Gene Superfamily in Ramie (Boehmeria nivea L.) Revealed Their Synergistic Roles in Regulating Abiotic Stress Resistance and Ramet Development" International Journal of Molecular Sciences 23, no. 23: 15117. https://doi.org/10.3390/ijms232315117
APA StyleQiu, X., Zhao, H., Abubakar, A. S., Shao, D., Chen, J., Chen, P., Yu, C., Wang, X., Chen, K., & Zhu, A. (2022). Genome-Wide Analysis of AP2/ERF Gene Superfamily in Ramie (Boehmeria nivea L.) Revealed Their Synergistic Roles in Regulating Abiotic Stress Resistance and Ramet Development. International Journal of Molecular Sciences, 23(23), 15117. https://doi.org/10.3390/ijms232315117








