Genetic Determinants of Fiber-Associated Traits in Flax Identified by Omics Data Integration
Abstract
:1. Introduction
2. Results and Discussion
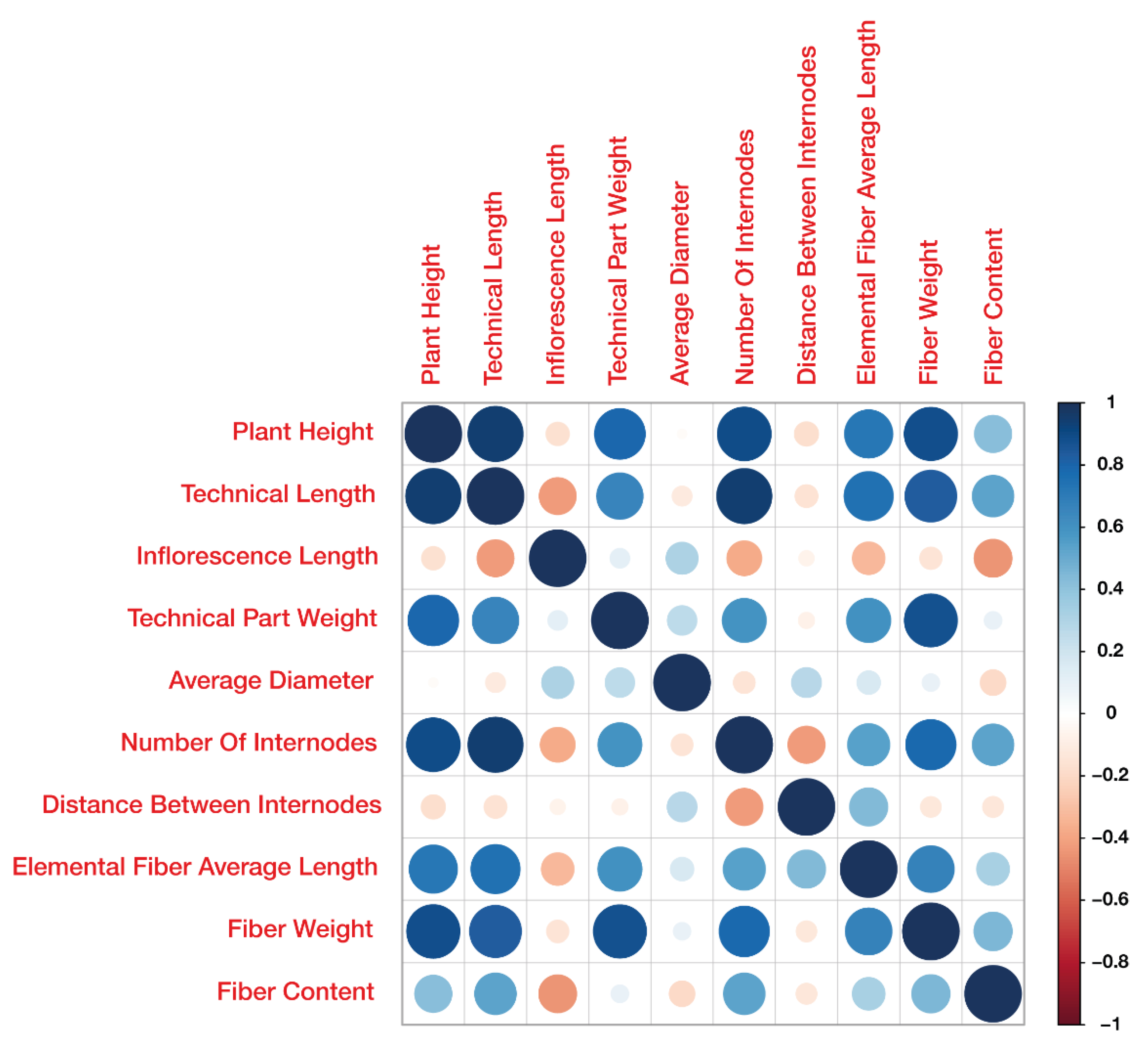
2.1. Phenotypic Variation
2.2. Association Mapping of Fiber Traits
2.3. Fiber-Related Candidate Genes
2.4. Candidate Genes Related to the Synthesis and Modification of Cell Wall Components
2.5. Candidate Genes Controlling Cell Fate Determination, Growth and Elongation
2.6. Candidate Genes Associated with Vesicular Transport, Intracellular Trafficking of the Cell Wall Components and Solute Transport across Membranes
2.7. Candidate Genes Involved in Hormonal Regulation
2.8. Candidate Genes Associated with Resistance to Plant Diseases
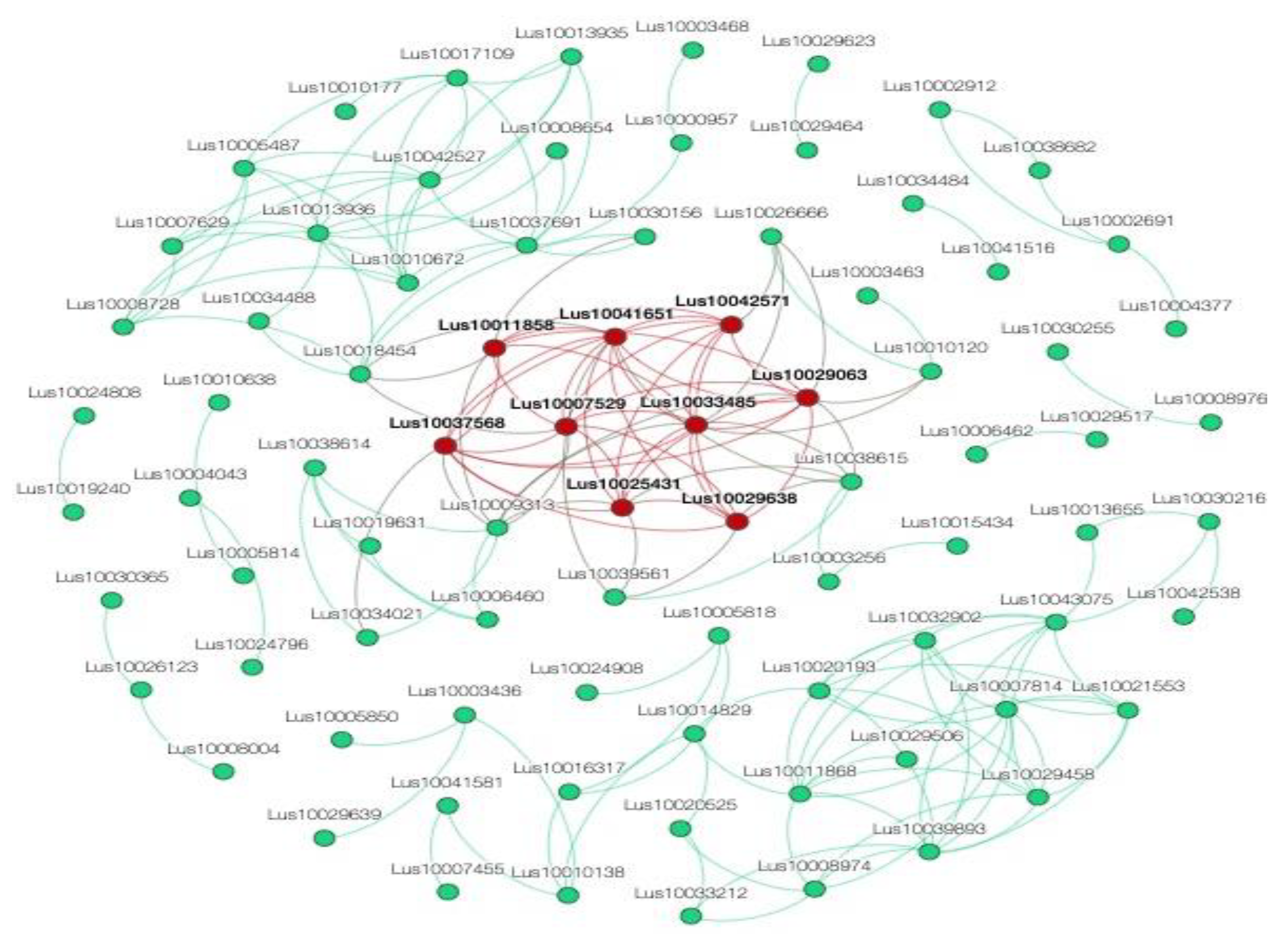
2.9. Network Analysis
3. Materials and Methods
3.1. Plant Material and Phenotyping
3.2. DNA Sequencing and Variant Calling
3.3. Genetic Data Analysis
3.4. GWAS
3.5. Candidate Genes
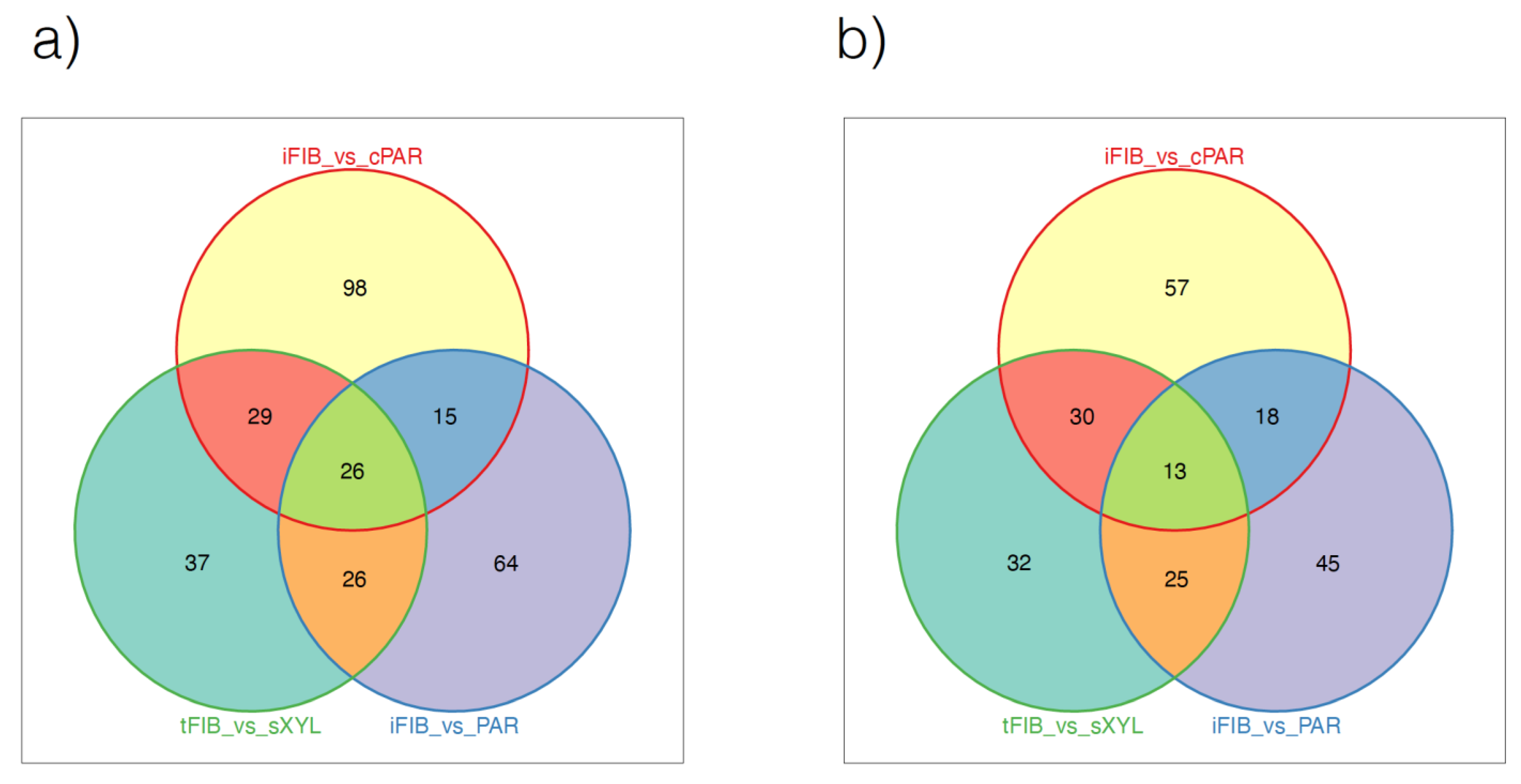
3.6. Differential Gene Expression in Fiber Cells
3.7. Co-Expression Network Construction
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| Plant height | (PH) |
| Technical stem length | (TL) |
| Technical part weight | (TW) |
| Stem diameter | (SD) |
| Number of internodes | (NI) |
| Distance between internodes | (DI) |
| Fiber content | (FC) |
| Fiber weight | (FW) |
| Elementary fiber length | (ELF) |
| Stem slenderness index | (SSI) |
| Stem tapering index | (STI) |
References
- Goudenhooft, C.; Bourmaud, A.; Baley, C. Flax (Linum usitatissimum L.) fibers for composite reinforcement: Exploring the link between plant growth, cell walls development, and fiber properties. Front. Plant Sci. 2019, 10, 411. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Allaby, R.G.; Peterson, G.W.; Merriwether, D.A.; Fu, Y.-B. Evidence of the domestication history of flax (Linum usitatissimum L.) from genetic diversity of the Sad2 locus. Theor. Appl. Genet. 2005, 112, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Goudenhooft, C.; Alméras, T.; Bourmaud, A.; Baley, C. The remarkable slenderness of flax plant and pertinent factors affecting its mechanical stability. Biosyst. Eng. 2019, 178, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Goudenhooft, C.; Bourmaud, A.; Baley, C. Varietal selection of flax over time: Evolution of plant architecture related to influence on the mechanical properties of fibers. Ind. Crop. Prod. 2017, 97, 56–64. [Google Scholar] [CrossRef]
- Fogorasi, M.; Barbu, I. The potential of natural fibres for automotive sector—Review. Iop. Conf. Ser. Mater. Sci. Eng. 2017, 252, 012044. [Google Scholar] [CrossRef] [Green Version]
- Mellerowicz, E.J.; Gorshkova, T.A. Tensional stress generation in gelatinous fibres: A review and possible mechanism based on cell-wall structure and composition. J. Exp. Bot. 2012, 63, 551–565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roach, M.J.; Mokshina, N.Y.; Badhan, A.; Snegireva, A.V.; Hobson, N.; Deyholos, M.K.; Gorshkova, T.A. Development of cellulosic secondary walls in flax fibers requires β-galactosidase. Plant Physiol. 2011, 156, 1351–1363. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gorshkova, T.; Chernova, T.; Mokshina, N.; Gorshkov, V.; Kozlova, L.; Gorshkov, O. Transcriptome analysis of intrusively growing flax fibers isolated by laser microdissection. Sci. Rep. 2018, 8, 14570. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baley, C.; Goudenhooft, C.; Gibaud, M.; Bourmaud, A. Flax stems: From a specific architecture to an instructive model for bioinspired composite structures. Bioinspir. Biomim. 2018, 13, 026007. [Google Scholar] [CrossRef] [PubMed]
- Rozhmina, T.; Bankin, M.; Samsonova, A.; Kanapin, A.; Samsonova, M. A comprehensive dataset of flax (Linum uitatissimum L.) Phenotypes. Data Brief 2021, 37, 107224. [Google Scholar] [CrossRef] [PubMed]
- Barabaschi, D.; Tondelli, A.; Desiderio, F.; Volante, A.; Vaccino, P.; Valè, G.; Cattivelli, L. Next generation breeding. Plant Sci. 2016, 242, 3–13. [Google Scholar] [CrossRef]
- Porter, H.F.; O’Reilly, P.F. Multivariate simulation framework reveals performance of multi-trait GWAS methods. Sci. Rep. 2017, 7, 38837. [Google Scholar] [CrossRef] [Green Version]
- Casale, F.P.; Rakitsch, B.; Lippert, C.; Stegle, O. Efficient set tests for the genetic analysis of correlated traits. Nat. Methods 2015, 12, 755–758. [Google Scholar] [CrossRef] [PubMed]
- Stephens, M. A unified framework for association analysis with multiple related phenotypes. PLoS ONE 2013, 8, e65245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ritchie, M.D.; Holzinger, E.R.; Li, R.; Pendergrass, S.A.; Kim, D. Methods of integrating data to uncover genotype–phenotype interactions. Nat. Rev. Genet. 2015, 16, 85–97. [Google Scholar] [CrossRef] [PubMed]
- Soto-Cerda, B.J.; Duguid, S.; Booker, H.; Rowland, G.; Diederichsen, A.; Cloutier, S. Genomic regions underlying agronomic traits in linseed (Linum usitatissimum L.) as revealed by association mapping. J. Integr. Plant Biol. 2014, 56, 75–87. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chandrawati; Singh, N.; Kumar, R.; Kumar, S.; Singh, P.K.; Yadav, V.K.; Ranade, S.A.; Yadav, H.K. Genetic diversity, population structure and association analysis in linseed (Linum usitatissimum L.). Physiol. Mol. Biol. Plants 2017, 23, 207–219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Soto-Cerda, B.J.; Cloutier, S.; Quian, R.; Gajardo, H.A.; Olivos, M.; You, F.M. Genome-wide association analysis of mucilage and hull content in flax (Linum usitatissimum L.) seeds. Int. J. Mol. Sci. 2018, 19, 2870. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- You, F.M.; Xiao, J.; Li, P.; Yao, Z.; Jia, G.; He, L.; Kumar, S.; Soto-Cerda, B.; Duguid, S.D.; Booker, H.M.; et al. Genome-wide association study and selection signatures detect genomic regions associated with seed yield and oil quality in flax. Int. J. Mol. Sci. 2018, 19, 2303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, D.; Dai, Z.; Yang, Z.; Tang, Q.; Deng, C.; Xu, Y.; Wang, J.; Chen, J.; Zhao, D.; Zhang, S.; et al. Combined genome-wide association analysis and transcriptome sequencing to identify candidate genes for flax seed fatty acid metabolism. Plant Sci. 2019, 286, 98–107. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.; Jiang, H.; Yan, W.; Yang, L.; Ye, J.; Wang, Y.; Yan, Q.; Chen, J.; Gao, Y.; Duan, L.; et al. Resequencing 200 flax cultivated accessions identifies candidate genes related to seed size and weight and reveals signatures of artificial selection. Front. Plant Sci. 2020, 10, 1682. [Google Scholar] [CrossRef] [Green Version]
- Soto-Cerda, B.J.; Aravena, G.; Cloutier, S. Genetic dissection of flowering time in flax (Linum usitatissimum L.) through single- and multi-locus genome-wide association studies. Mol. Genet. Genom. 2021, 296, 877–891. [Google Scholar] [CrossRef] [PubMed]
- Sertse, D.; You, F.M.; Ravichandran, S.; Soto-Cerda, B.J.; Duguid, S.; Cloutier, S. Loci harboring genes with important role in drought and related abiotic stress responses in flax revealed by multiple GWAS models. Theor. Appl. Genet. 2021, 134, 191–212. [Google Scholar] [CrossRef] [PubMed]
- Xie, D.; Dai, Z.; Yang, Z.; Sun, J.; Zhao, D.; Yang, X.; Zhang, L.; Tang, Q.; Su, J. Genome-wide association study identifying candidate genes influencing important agronomic traits of flax (Linum usitatissimum L.) using SLAF-seq. Front. Plant Sci. 2018, 8, 2232. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, J.; Zhao, Q.; Zhang, L.; Li, S.; Ma, Y.; Pan, L.; Lin, H.; Wu, G.; Yuan, H.; Yu, Y.; et al. QTL mapping of fiber-related traits based on a high-density genetic map in flax (Linum usitatissimum L.). Front. Plant Sci. 2018, 9, 885. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, J.; Long, Y.; Wang, L.; Dang, Z.; Zhang, T.; Song, X.; Dang, Z.; Pei, X. Consensus genetic linkage map construction and QTL mapping for plant height-related traits in linseed flax (Linum usitatissimum L.). BMC Plant Biol. 2018, 18, 160. [Google Scholar] [CrossRef]
- Kumar, S.; You, F.M.; Duguid, S.; Booker, H.; Rowland, G.; Cloutier, S. QTL for fatty acid composition and yield in linseed (Linum usitatissimum L.). Theor. Appl. Genet. 2015, 128, 965–984. [Google Scholar] [CrossRef] [PubMed]
- Xie, D.; Dai, Z.; Yang, Z.; Tang, Q.; Sun, J.; Yang, X.; Song, X.; Lu, Y.; Zhao, D.; Zhang, L.; et al. Genomic variations and association study of agronomic traits in flax. BMC Genom. 2018, 19, 512. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gorshkov, O.; Chernova, T.; Mokshina, N.; Gogoleva, N.; Suslov, D.; Tkachenko, A.; Gorshkova, T. Intrusive growth of phloem fibers in flax stem: Integrated analysis of MiRNA and MRNA expression profiles. Plants 2019, 8, 47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mokshina, N.; Gorshkov, O.; Galinousky, D.; Gorshkova, T. Genes with bast fiber-specific expression in flax plants—Molecular keys for targeted fiber crop improvement. Ind. Crop. Prod. 2020, 152, 112549. [Google Scholar] [CrossRef]
- Duk, M.; Kanapin, A.; Rozhmina, T.; Bankin, M.; Surkova, S.; Samsonova, A.; Samsonova, M. The genetic landscape of fiber flax. Front. Plant Sci. 2021, 12, 764612. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, Z. GAPIT version 3: Boosting power and accuracy for genomic association and prediction. bioRxiv 2020. bioRxiv: 2020.11.29.403170. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Stephens, M. Efficient algorithms for multivariate linear mixed models in genome-wide association studies. Nat. Methods 2014, 11, 407–409. [Google Scholar] [CrossRef]
- Struck, A.W.; Thompson, M.L.; Wong, L.S.; Micklefield, J. S-adenosyl-methionine-dependent methyltransferases: Highly versatile enzymes in biocatalysis, biosynthesis and other biotechnological applications. Chembiochem. 2012, 13, 2642–2655. [Google Scholar] [CrossRef] [PubMed]
- Azevedo, C.; Santos-Rosa, M.J.; Shirasu, K. The U-box protein family in plants. Trends Plant Sci. 2001, 6, 354–358. [Google Scholar] [CrossRef]
- Yang, W.; Chen, S.; Cheng, Y.; Zhang, N.; Ma, Y.; Wang, W.; Tian, H.; Li, Y.; Hussain, S.; Wang, S. Cell wall/vacuolar inhibitor of fructosidase 1 regulates ABA response and salt tolerance in Arabidopsis. Plant Signal. Behav. 2020, 15, 1744293. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, X.-R.; Lian, H.; Ni, D.-A.; He, Y.; Chen, X.-Y.; Ruan, Y.-L. Evidence that high activity of vacuolar invertase is required for cotton fiber and Arabidopsis root elongation through osmotic dependent and independent pathways, respectively. Plant Physiol. 2010, 154, 744–756. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mravec, J.; Skůpa, P.; Bailly, A.; Hoyerová, K.; Křeček, P.; Bielach, A.; Petrášek, J.; Zhang, J.; Gaykova, V.; Stierhof, Y.-D.; et al. Subcellular homeostasis of phytohormone auxin is mediated by the ER-localized PIN5 transporter. Nature 2009, 459, 1136–1140. [Google Scholar] [CrossRef] [PubMed]
- Majda, M.; Robert, S. The role of auxin in cell wall expansion. Int. J. Mol. Sci. 2018, 19, 951. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fawal, N.; Li, Q.; Savelli, B.; Brette, M.; Passaia, G.; Fabre, M.; Mathé, C.; Dunand, C. PeroxiBase: A database for large-scale evolutionary analysis of peroxidases. Nucleic Acids Res. 2013, 41, D441–D444. [Google Scholar] [CrossRef] [PubMed]
- Francoz, E.; Ranocha, P.; Nguyen-Kim, H.; Jamet, E.; Burlat, V.; Dunand, C. Roles of cell wall peroxidases in plant development. Phytochemistry 2015, 112, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Ul Haq, S.; Khan, A.; Ali, M.; Khattak, A.M.; Gai, W.-X.; Zhang, H.-X.; Wei, A.-M.; Gong, Z.-H. Heat shock proteins: Dynamic biomolecules to counter plant biotic and abiotic stresses. Int. J. Mol. Sci. 2019, 20, 5321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Drew, D.; North, R.A.; Nagarathinam, K.; Tanabe, M. Structures and general transport mechanisms by the major facilitator superfamily (MFS). Chem. Rev. 2021, 121, 5289–5335. [Google Scholar] [CrossRef]
- Liberato, M.V.; Silveira, R.L.; Prates, É.T.; de Araujo, E.A.; Pellegrini, V.O.A.; Camilo, C.M.; Kadowaki, M.A.; Neto, M.d.O.; Popov, A.; Skaf, M.S.; et al. Molecular characterization of a family 5 glycoside hydrolase suggests an induced-fit enzymatic mechanism. Sci. Rep. 2016, 6, 23473. [Google Scholar] [CrossRef] [Green Version]
- Korolev, A.V.; Chan, J.; Naldrett, M.J.; Doonan, J.H.; Lloyd, C.W. Identification of a novel family of 70° KDa microtubule-associated proteins in Arabidopsis cells. Plant J. 2005, 42, 547–555. [Google Scholar] [CrossRef]
- Lim, E.-K.; Doucet, C.J.; Li, Y.; Elias, L.; Worrall, D.; Spencer, S.P.; Ross, J.; Bowles, D.J. The activity of Arabidopsis glycosyltransferases toward salicylic acid, 4-hydroxybenzoic acid, and other benzoates. J. Biol. Chem. 2002, 277, 586–592. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaundal, A.; Sandhu, D.; Duenas, M.; Ferreira, J.F.S. Expression of the high-affinity K+ transporter 1 (PpHKT1) gene from almond rootstock ‘Nemaguard’ improved salt tolerance of transgenic Arabidopsis. PLoS ONE 2019, 14, e0214473. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rennie, E.A.; Hansen, S.F.; Baidoo, E.E.K.; Hadi, M.Z.; Keasling, J.D.; Scheller, H.V. Three members of the Arabidopsis glycosyltransferase family 8 are xylan glucuronosyltransferases. Plant Physiol. 2012, 159, 1408–1417. [Google Scholar] [CrossRef] [Green Version]
- Ochando, I.; González-Reig, S.; Ripoll, J.-J.; Vera, A.; Martínez-Laborda, A. Alteration of the shoot radial pattern in Arabidopsis thaliana by a gain-of-function allele of the class III HD-Zip gene INCURVATA. Int. J. Dev. Biol. 2004, 52, 953–961. [Google Scholar] [CrossRef]
- Shi, J.X.; Malitsky, S.; Oliveira, S.D.; Branigan, C.; Franke, R.B.; Schreiber, L.; Aharoni, A. SHINE transcription factors act redundantly to pattern the archetypal surface of Arabidopsis flower organs. PLoS Genet. 2011, 7, e1001388. [Google Scholar] [CrossRef] [PubMed]
- Nishigaki, N.; Yoshimi, Y.; Kuki, H.; Kunieda, T.; Hara-Nishimura, I.; Tsumuraya, Y.; Takahashi, D.; Dupree, P.; Kotake, T. Galactoglucomannan structure of Arabidopsis seed-coat mucilage in GDP-mannose synthesis impaired mutants. Physiol. Plant. 2021, 173, 1244–1252. [Google Scholar] [CrossRef] [PubMed]
- Vercruysse, J.; Baekelandt, A.; Gonzalez, N.; Inzé, D. Molecular networks regulating cell division during Arabidopsis leaf growth. J. Exp. Bot. 2019, 71, 2365–2378. [Google Scholar] [CrossRef]
- Ito, T.; Meyerowitz, E.M. Overexpression of a gene encoding a cytochrome P450, CYP78A9, induces large and seedless fruit in Arabidopsis. Plant Cell 2000, 12, 1541–1550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, W.; Wang, Z.; Cui, R.; Li, J.; Li, Y. Maternal control of seed size by EOD3/CYP78A6 in Arabidopsis thaliana. Plant J. 2012, 70, 929–939. [Google Scholar] [CrossRef]
- Kang, B.-H.; Busse, J.S.; Bednarek, S.Y. Members of the Arabidopsis dynamin-like gene family, ADL1, are essential for plant cytokinesis and polarized cell growth. Plant Cell 2003, 15, 899–913. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mindrebo, J.T.; Nartey, C.M.; Seto, Y.; Burkart, M.D.; Noel, J.P. Unveiling the functional diversity of the alpha/beta hydrolase superfamily in the plant kingdom. Curr. Opin. Struc. Biol. 2016, 41, 233–246. [Google Scholar] [CrossRef] [Green Version]
- Pires, I.S.; Negrão, S.; Pentony, M.M.; Abreu, I.A.; Oliveira, M.M.; Purugganan, M.D. Different evolutionary histories of two cation/proton exchanger gene families in plants. BMC Plant Biol. 2013, 13, 97. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuroda, H.; Yanagawa, Y.; Takahashi, N.; Horii, Y.; Matsui, M. A comprehensive analysis of interaction and localization of Arabidopsis SKP1-LIKE (ASK) and F-Box (FBX) proteins. PLoS ONE 2012, 7, e50009. [Google Scholar] [CrossRef] [Green Version]
- Rocha, J.; Cicéron, F.; de Sanctis, D.; Lelimousin, M.; Chazalet, V.; Lerouxel, O.; Breton, C. Structure of Arabidopsis thaliana FUT1 reveals a variant of the GT-B class fold and provides insight into Xyloglucan fucosylation. Plant Cell 2016, 28, 2352–2364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tamasloukht, B.; Lam, M.S.-J.W.Q.; Martinez, Y.; Tozo, K.; Barbier, O.; Jourda, C.; Jauneau, A.; Borderies, G.; Balzergue, S.; Renou, J.-P.; et al. Characterization of a cinnamoyl-CoA reductase 1 (CCR1) mutant in maize: Effects on lignification, fibre development, and global gene expression. J. Exp. Bot. 2011, 62, 3837–3848. [Google Scholar] [CrossRef]
- Brembu, T.; Winge, P.; Bones, A.M. The small GTPase AtRAC2/ROP7 is specifically expressed during late stages of xylem differentiation in Arabidopsis. J. Exp. Bot. 2005, 56, 2465–2476. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Waghmare, S.; Lileikyte, E.; Karnik, R.; Goodman, J.K.; Blatt, M.R.; Jones, A.M.E. SNAREs SYP121 and SYP122 mediate the secretion of distinct cargo subsets. Plant Physiol. 2018, 178, 1679–1688. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Q.; Wang, H.; Zhang, Z.; Wu, J.; Feng, Y.; Zhu, Z. Divergence in function and expression of the NOD26-like intrinsic proteins in plants. BMC Genomics 2009, 10, 313. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Irshad, M.; Canut, H.; Borderies, G.; Pont-Lezica, R.; Jamet, E. A new picture of cell wall protein dynamics in elongating cells of Arabidopsis thaliana: Confirmed actors and newcomers. BMC Plant Biol. 2008, 8, 94. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moneo-Sánchez, M.; Izquierdo, L.; Martín, I.; Labrador, E.; Dopico, B. Subcellular location of Arabidopsis thaliana subfamily A1 β-galactosidases and developmental regulation of transcript levels of their coding genes. Plant Physiol. Biochem. 2016, 109, 137–145. [Google Scholar] [CrossRef] [PubMed]
- He, G.; Liu, P.; Zhao, H.; Sun, J. The HD-ZIP II transcription factors regulate plant architecture through the auxin pathway. Int. J. Mol. Sci. 2020, 21, 3250. [Google Scholar] [CrossRef] [PubMed]
- Schreiber, K.J.; Bentham, A.; Williams, S.J.; Kobe, B.; Staskawicz, B.J. Multiple domain associations within the Arabidopsis immune receptor RPP1 regulate the activation of programmed cell death. PLoS Pathog. 2016, 12, e1005769. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Zhou, M. The ALMT gene family performs multiple functions in plants. Agronomy 2018, 8, 20. [Google Scholar] [CrossRef] [Green Version]
- Rauf, M.; Arif, M.; Fisahn, J.; Xue, G.-P.; Balazadeh, S.; Mueller-Roeber, B. NAC transcription factor SPEEDY HYPONASTIC GROWTH regulates flooding-induced leaf movement in Arabidopsis. Plant Cell 2013, 25, 4941–4955. [Google Scholar] [CrossRef] [Green Version]
- Rajaraman, J.; Douchkov, D.; Hensel, G.; Stefanato, F.L.; Gordon, A.; Ereful, N.; Caldararu, O.F.; Petrescu, A.-J.; Kumlehn, J.; Boyd, L.A.; et al. An LRR/malectin receptor-like kinase mediates resistance to non-adapted and adapted powdery mildew fungi in barley and wheat. Front. Plant Sci. 2016, 7, 1836. [Google Scholar] [CrossRef]
- Li, Z.; Liu, D.; Xia, Y.; Li, Z.; Jing, D.; Du, J.; Niu, N.; Ma, S.; Wang, J.; Song, Y.; et al. Identification of the WUSCHEL-related homeobox (WOX) gene family, and interaction and functional analysis of TaWOX9 and TaWUS in wheat. Int. J. Mol. Sci. 2020, 21, 1581. [Google Scholar] [CrossRef] [Green Version]
- Xu, J.; Wang, X.; Guo, W. The cytochrome P450 superfamily: Key players in plant development and defense. J. Integr. Agr. 2015, 14, 1673–1686. [Google Scholar] [CrossRef] [Green Version]
- Eggers, R.; Jammer, A.; Jha, S.; Kerschbaumer, B.; Lahham, M.; Strandback, E.; Toplak, M.; Wallner, S.; Winkler, A.; Macheroux, P. The scope of flavin-dependent reactions and processes in the model plant Arabidopsis thaliana. Phytochemistry 2021, 189, 112822. [Google Scholar] [CrossRef]
- Etchells, J.P.; Provost, C.M.; Mishra, L.; Turner, S.R. WOX4 and WOX14 act downstream of the PXY receptor kinase to regulate plant vascular proliferation independently of any role in vascular organisation. Development 2013, 140, 2224–2234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gou, M.; Yang, X.; Zhao, Y.; Ran, X.; Song, Y.; Liu, C.-J. Cytochrome b 5 is an obligate electron shuttle protein for syringyl lignin biosynthesis in Arabidopsis. Plant Cell 2019, 31, 1344–1366. [Google Scholar] [CrossRef] [Green Version]
- Davies, G.; Henrissat, B. Structures and mechanisms of glycosyl hydrolases. Structure 1995, 3, 853–859. [Google Scholar] [CrossRef] [Green Version]
- Woo, H.-H.; Jeong, B.R.; Hirsch, A.M.; Hawes, M.C. Characterization of Arabidopsis AtUGT85A and AtGUS gene families and their expression in rapidly dividing tissues. Genomics 2007, 90, 143–153. [Google Scholar] [CrossRef] [Green Version]
- Jin, S.-H.; Ma, X.-M.; Kojima, M.; Sakakibara, H.; Wang, Y.-W.; Hou, B.-K. Overexpression of glucosyltransferase UGT85A1 influences trans-zeatin homeostasis and trans-zeatin responses likely through O-glucosylation. Planta 2013, 237, 991–999. [Google Scholar] [CrossRef] [PubMed]
- Baucher, M.; Pérez-Morga, D.; Jaziri, M.E. Insight into plant annexin function: From shoot to root signaling. Plant Signal. Behav. 2012, 7, 524–528. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoffmann, L.; Besseau, S.; Geoffroy, P.; Ritzenthaler, C.; Meyer, D.; Lapierre, C.; Pollet, B.; Legrand, M. Silencing of hydroxycinnamoyl-coenzyme a shikimate/quinate hydroxycinnamoyltransferase affects phenylpropanoid biosynthesis. Plant Cell 2004, 16, 1446–1465. [Google Scholar] [CrossRef] [PubMed]
- Hebda, A.; Liszka, A.; Lewandowska, A.; Lyczakowski, J.J.; Gabryś, H.; Krzeszowiec, W. Upregulation of GLRs expression by light in Arabidopsis leaves. BMC Plant Biol. 2022, 22, 197. [Google Scholar] [CrossRef]
- Merkulova, M.; Hurtado-Lorenzo, A.; Hosokawa, H.; Zhuang, Z.; Brown, D.; Ausiello, D.A.; Marshansky, V. Aldolase directly interacts with ARNO and modulates cell morphology and acidic vesicle distribution. Am. J. Physiol.-Cell Physiol. 2011, 300, C1442–C1455. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.; Singh, K.B. Characterization of salicylic acid-responsive, Arabidopsis Dof domain proteins: Overexpression of OBP3 leads to growth defects. Plant J. 2000, 21, 329–339. [Google Scholar] [CrossRef]
- Ranocha, P.; Dima, O.; Nagy, R.; Felten, J.; Corratgé-Faillie, C.; Novák, O.; Morreel, K.; Lacombe, B.; Martinez, Y.; Pfrunder, S.; et al. Arabidopsis WAT1 is a vacuolar auxin transport facilitator required for auxin homoeostasis. Nat. Commun. 2013, 4, 2625. [Google Scholar] [CrossRef] [Green Version]
- Ranocha, P.; Denancé, N.; Vanholme, R.; Freydier, A.; Martinez, Y.; Hoffmann, L.; Köhler, L.; Pouzet, C.; Renou, J.; Sundberg, B.; et al. Walls are thin 1 (WAT1), an Arabidopsis homolog of medicago truncatula NODULIN21, is a tonoplast-localized protein required for secondary wall formation in fibers. Plant J. 2010, 63, 469–483. [Google Scholar] [CrossRef] [PubMed]
- Denancé, N.; Szurek, B.; Noël, L.D. Emerging functions of nodulin-like proteins in non-nodulating plant species. Plant Cell Physiol. 2014, 55, 469–474. [Google Scholar] [CrossRef] [PubMed]
- Geng, P.; Zhang, S.; Liu, J.; Zhao, C.; Wu, J.; Cao, Y.; Fu, C.; Han, X.; He, H.; Zhao, Q. MYB20, MYB42, MYB43, and MYB85 regulate phenylalanine and lignin biosynthesis during secondary cell wall formation. Plant Physiol. 2019, 182, 1272–1283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Minic, Z.; Jouanin, L. Plant glycoside hydrolases involved in cell wall polysaccharide degradation. Plant Physiol. Biochem. 2006, 44, 435–449. [Google Scholar] [CrossRef]
- Tedeschi, F.; Rizzo, P.; Rutten, T.; Altschmied, L.; Bäumlein, H. RWP-RK domain-containing transcription factors control cell differentiation during female gametophyte development in Arabidopsis. New Phytol. 2017, 213, 1909–1924. [Google Scholar] [CrossRef]
- De Araújo, A.C.; Fonseca, F.C.D.A.; Cotta, M.G.; Alves, G.S.C.; Miller, R.N.G. Plant NLR receptor proteins and their potential in the development of durable genetic resistance to biotic stresses. Biotechnol. Res. Innov. 2019, 3, 80–94. [Google Scholar] [CrossRef]
- Schaller, A.; Stintzi, A.; Rivas, S.; Serrano, I.; Chichkova, N.V.; Vartapetian, A.B.; Martínez, D.; Guiamét, J.J.; Sueldo, D.J.; Hoorn, R.A.L.; et al. From structure to function—A family portrait of plant subtilases. New Phytol. 2018, 218, 901–915. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bischoff, V.; Nita, S.; Neumetzler, L.; Schindelasch, D.; Urbain, A.; Eshed, R.; Persson, S.; Delmer, D.; Scheible, W.-R. TRICHOME BIREFRINGENCE and its homolog AT5G01360 encode plant-specific DUF231 proteins required for cellulose biosynthesis in Arabidopsis. Plant Physiol. 2010, 153, 590–602. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bris, P.L.; Wang, Y.; Barbereau, C.; Antelme, S.; Cézard, L.; Legée, F.; D’Orlando, A.; Dalmais, M.; Bendahmane, A.; Schuetz, M.; et al. Inactivation of LACCASE8 and LACCASE5 genes in brachypodium distachyon leads to severe decrease in lignin content and high increase in saccharification yield without impacting plant integrity. Biotechnol. Biofuels 2019, 12, 181. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bürstenbinder, K.; Savchenko, T.; Müller, J.; Adamson, A.W.; Stamm, G.; Kwong, R.; Zipp, B.J.; Dinesh, D.C.; Abel, S. Arabidopsis Calmodulin-binding protein IQ67-domain 1 localizes to microtubules and interacts with kinesin light chain-related protein-1. J. Biol. Chem. 2013, 288, 1871–1882. [Google Scholar] [CrossRef] [Green Version]
- Bürstenbinder, K.; Möller, B.; Plötner, R.; Stamm, G.; Hause, G.; Mitra, D.; Abel, S. The IQD family of calmodulin-binding proteins links calcium signaling to microtubules, membrane subdomains, and the nucleus. Plant Physiol. 2017, 173, 1692–1708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhong, R.; Richardson, E.A.; Ye, Z.-H. The MYB46 transcription factor is a direct target of SND1 and regulates secondary wall biosynthesis in Arabidopsis. Plant Cell 2007, 19, 2776–2792. [Google Scholar] [CrossRef] [Green Version]
- Cordoba, E.; Aceves-Zamudio, D.L.; Hernández-Bernal, A.F.; Ramos-Vega, M.; León, P. Sugar regulation of SUGAR TRANSPORTER PROTEIN 1 (STP1) expression in Arabidopsis thaliana. J. Exp. Bot. 2015, 66, 147–159. [Google Scholar] [CrossRef] [Green Version]
- Ma, R.; Yuan, H.; An, J.; Hao, X.; Li, H. A Gossypium hirsutum GDSL lipase/hydrolase gene (GhGLIP) appears to be involved in promoting seed growth in Arabidopsis. PLoS ONE 2018, 13, e0195556. [Google Scholar] [CrossRef] [Green Version]
- Lai, C.-P.; Huang, L.-M.; Chen, L.-F.O.; Chan, M.-T.; Shaw, J.-F. Genome-wide analysis of GDSL-type esterases/lipases in Arabidopsis. Plant Mol. Biol. 2017, 95, 181–197. [Google Scholar] [CrossRef] [PubMed]
- Yadav, V.K.; Yadav, V.K.; Pant, P.; Singh, S.P.; Maurya, R.; Sable, A.; Sawant, S.V. GhMYB1 regulates SCW stage-specific expression of the GhGDSL promoter in the fibres of Gossypium hirsutum L. Plant Biotechnol. J. 2017, 15, 1163–1174. [Google Scholar] [CrossRef] [PubMed]
- Li, S.-B.; Xie, Z.-Z.; Hu, C.-G.; Zhang, J.-Z. A review of auxin response factors (ARFs) in plants. Front. Plant Sci. 2016, 7, 47. [Google Scholar] [CrossRef] [Green Version]
- Yang, Y.S.; Strittmatter, S.M. The reticulons: A family of proteins with diverse functions. Genome Biol. 2007, 8, 234. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Mitsuda, N.; Yoshizumi, T.; Horii, Y.; Oshima, Y.; Ohme-Takagi, M.; Matsui, M.; Kakimoto, T. Two types of BHLH transcription factor determine the competence of the pericycle for lateral root initiation. Nat. Plants 2021, 7, 633–643. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Sun, J.; Liu, N.; Sun, X.; Liu, C.; Wu, L.; Liu, G.; Zeng, F.; Hou, C.; Han, S.; et al. A novel cysteine-rich receptor-like kinase gene, TaCRK2, contributes to leaf rust resistance in wheat. Mol. Plant Pathol. 2020, 21, 732–746. [Google Scholar] [CrossRef] [Green Version]
- Parker, J.E.; Coleman, M.J.; Szabò, V.; Frost, L.N.; Schmidt, R.; van der Biezen, E.A.; Moores, T.; Dean, C.; Daniels, M.J.; Jones, J.D. The Arabidopsis downy mildew resistance gene RPP5 shares similarity to the toll and interleukin-1 receptors with N and L6. Plant Cell 1997, 9, 879–894. [Google Scholar] [CrossRef] [Green Version]
- Piovesana, M.; Wood, A.K.M.; Smith, D.P.; Deery, M.J.; Bayliss, R.; Carrera, E.; Napier, J.A.; Kurup, S.; Matthes, M.C. A point mutation in the kinase domain of CRK10 leads to xylem vessel collapse and activates defence responses. bioRxiv 2021. bioRxiv: 2021.08.16.456532. [Google Scholar] [CrossRef]
- Cosgrove, D.J. Plant cell growth and elongation. In eLS; John Wiley & Sons, Ltd.: New York, NY, USA, 2014. [Google Scholar] [CrossRef]
- Wu, S.-W.; Kumar, R.; Iswanto, A.B.B.; Kim, J.-Y. Callose balancing at plasmodesmata. J. Exp. Bot. 2018, 69, 5325–5339. [Google Scholar] [CrossRef]
- Ageeva, M.V.; Petrovská, B.; Kieft, H.; Sal’nikov, V.V.; Snegireva, A.V.; van Dam, J.E.G.; van Veenendaal, W.L.H.; Emons, A.M.C.; Gorshkova, T.A.; van Lammeren, A.A.M. Intrusive growth of flax phloem fibers is of intercalary type. Planta 2005, 222, 565–574. [Google Scholar] [CrossRef]
- Cassab, G.I. Plant cell wall proteins. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1998, 49, 281–309. [Google Scholar] [CrossRef] [PubMed]
- Hellmann, E.; Ko, D.; Ruonala, R.; Helariutta, Y. Plant vascular tissues—Connecting tissue comes in all shapes. Plants 2018, 7, 109. [Google Scholar] [CrossRef] [Green Version]
- Du, Q.; Wang, H. The role of HD-ZIP III transcription factors and MiR165/166 in vascular development and secondary cell wall formation. Plant Signal. Behav. 2015, 10, e1078955. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turchi, L.; Carabelli, M.; Ruzza, V.; Possenti, M.; Sassi, M.; Peñalosa, A.; Sessa, G.; Salvi, S.; Forte, V.; Morelli, G.; et al. Arabidopsis HD-Zip II transcription factors control apical embryo development and meristem function. Development 2013, 140, 2118–2129. [Google Scholar] [CrossRef] [Green Version]
- Sanderfoot, A.A.; Kovaleva, V.; Bassham, D.C.; Raikhel, N.V. Interactions between syntaxins identify at least five SNARE complexes within the golgi/prevacuolar system of the Arabidopsis cell. Mol. Biol. Cell 2001, 12, 3733–3743. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Ruan, Y.-L. Unraveling mechanisms of cell expansion linking solute transport, metabolism, plasmodesmtal gating and cell wall dynamics. Plant Signal. Behav. 2010, 5, 1561–1564. [Google Scholar] [CrossRef] [Green Version]
- Malinovsky, F.G.; Fangel, J.U.; Willats, W.G.T. The role of the cell wall in plant immunity. Front. Plant Sci. 2014, 5, 178. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- You, F.M.; Cloutier, S. Mapping quantitative trait loci onto chromosome-scale pseudomolecules in flax. Methods Protoc. 2020, 3, 28. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Durbin, R. Fast and accurate short read alignment with burrows–wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tello, D.; Gil, J.; Loaiza, C.D.; Riascos, J.J.; Cardozo, N.; Duitama, J. NGSEP3: Accurate variant calling across species and sequencing protocols. Bioinformatics 2019, 35, 4716–4723. [Google Scholar] [CrossRef] [Green Version]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef]
- Zhang, C.; Dong, S.-S.; Xu, J.-Y.; He, W.-M.; Yang, T.-L. PopLDdecay: A fast and effective tool for linkage disequilibrium decay analysis based on variant call format files. Bioinformatics 2018, 35, 1786–1788. [Google Scholar] [CrossRef] [PubMed]
- Hill, W.G.; Weir, B.S. Variances and covariances of squared linkage disequilibria in finite populations. Theor. Popul. Biol. 1988, 33, 54–78. [Google Scholar] [CrossRef]
- Mann, H.B.; Whitney, D.R. On a test of whether one of two random variables is stochastically larger than the other. Ann. Math. Stat. 1947, 18, 50–60. [Google Scholar] [CrossRef]
- Dobin, A.; Gingeras, T.R. Mapping RNA-seq reads with STAR. Curr. Protoc. Bioinform. 2015, 51, 11–14. [Google Scholar] [CrossRef] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Almeida-Silva, F.; Venancio, T.M. BioNERO: An all-in-one R/bioconductor package for comprehensive and easy biological network reconstruction. bioRxiv 2021. bioRxiv: 2021.04.10.439287. [Google Scholar] [CrossRef]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 2008, 9, 559. [Google Scholar] [CrossRef] [Green Version]
- Bastian, M.; Heymann, S.; Jacomy, M. Gephi: An open source software for exploring and manipulating networks. Proc. Int. AAAI Conf. Web Soc. Media 2009, 3, 361–362. [Google Scholar] [CrossRef]

| QTN | Gene | Functional Annotation | Arabidopsis Gene Name | Reference | Predicted Function |
|---|---|---|---|---|---|
| Chr1:27748943 | Lus10010138 | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein | AT4G01240 | [34] | methylation and demethylation of cell wall proteins |
| Chr2:261056 | Lus10005814 | RING/U-box superfamily protein | AT1G49230 | [35] | signaling proteins for receptor kinases, regulation of hormone signaling and abiotic/biotic stress response |
| Lus10005815 | RING/U-box superfamily protein | AT1G49230 | |||
| Lus10005818 | RING/U-box superfamily protein | AT1G49230 | |||
| Chr2:2513654 | Lus10016317 | cell wall/vacuolar inhibitor of fructosidase 1 | AT1G47960, ATC/VIF1, C/VIF1 | [36,37] | regulation of ABA response and salt tolerance |
| Chr2:4360628 | Lus10020193 | auxin efflux carrier family protein | AT5G16530, PIN5 | [38,39] | regulates intracellular auxin homeostasis and metabolism |
| Chr2:4683418 | Lus10013618 | RING/U-box superfamily protein | AT5G17600 | [35] | signaling proteins for receptor kinases, regulation of hormone signaling and abiotic/biotic stress response |
| Lus10013631 | peroxidase superfamily protein, class III | AT5G1782, AtperoxP57 | [40,41] | cell wall localized proteins tightly associated to its loosening and stiffening | |
| Chr2:4872459 | Lus10013655 | HSP20-like chaperones superfamily protein | AT1G54400 | [42] | enhances membrane stability and detoxifies ROS by regulating the antioxidant enzymes system |
| Chr2:6885020 | Lus10038682 | major facilitator superfamily protein | AT2G39210 | [43] | facilitates movement of small solutes across cell membranes |
| Chr2:25126362 | Lus10006462 | cellulase (glycosyl hydrolase family 5) protein | AT1G13130 | [44] | hydrolyses the glycosidic bond |
| Chr3:4704531 | Lus10019171 | microtubule-associated proteins 70-2 | AT1G24764, ATMAP70-2 | [45] | outlining tracks of cellulose production during SCW formation and delivery of cell wall components |
| Chr3:5077352 | Lus10007972 | UDP-glucosyl transferase 89B1 | AT1G73880, UGT89B1 | [46] | |
| Chr3:5883916 | Lus10040598 | high-affinity K+ transporter 1 | AT4G10310, ATHKT1 | [47] | reducing Na+ toxicity through K+ uptake |
| Chr3:17134662 | Lus10033485 | plant glycogenin-like starch initiation protein 1 | AT3G18660, GUX1, PGSIP1 | [48] | glucuronyl transferase responsible for the addition of GlcA residues onto xylan and for secondary wall deposition |
| Chr3:17878746 | Lus10007051 | peroxidase superfamily protein, class III | AT2G4148, AtperoxP25 | [40,41] | cell wall localized proteins tightly associated to its loosening and stiffening |
| Chr3:25379963 | Lus10037568 | homeobox-leucine zipper family protein/lipid-binding START domain-containing protein | AT1G52150, ATHB15, CNA, ICU4 | [49] | confers positional information which is required to establish the number and pattern of vascular bundles in the stem, and is involved in repression of secondary cell wall development in Arabidopsis |
| Chr3:26449997 | Lus10002912 | integrase-type DNA-binding superfamily protein | AT5G25390, SHN2 | [50] | TF, regulates floral organs’ epidermal cell elongation by affecting pectin metabolism |
| Chr4:184242 | Lus10030156 | glucose-1-phosphate adenylyltransferase family protein | AT2G39770, CYT1, EMB101, GMP1, SOZ1, VTC1 | [51] | provides GDP-mannose, which is used for cell wall carbohydrate biosynthesis, including lignin and protein glycosylation |
| Chr4:466800 | Lus10030216 | cytochrome P450, family 78, subfamily A, polypeptide 6 | CYP78A6, EOD3 | [52,53,54] | Cell expansion, enhancer of DA-1 3 and seed and fruit development |
| Chr4:647773 | Lus10030255 | DYNAMIN-like 1B | AT3G61760, ADL1B | [55] | membrane trafficking, essential for plant cytokinesis and polarized cell growth |
| Chr4:1229010 | Lus10030366 | alpha/beta-hydrolases superfamily protein | AT3G62860 | [56] | breaking of carbon–carbon bonds |
| Chr4:2763834 | Lus10029506 | cation/hydrogen exchanger 28 | AT3G52080, chx28 | [57] | cation/hydrogen exchange |
| Chr4:7789243 | Lus10039564 | F-box/RNI-like superfamily protein | AT5G67140 | [58] | SCF ubiquitin ligase complex |
| Chr4:14879575 Chr4:14879658 | Lus10041581 | lysine histidine transporter 1 | AT5G40780, LHT1 | [30] | solute transporter |
| Chr4:15210519 | Lus10041644 | fucosyltransferase 1 | AT2G03220, ATFT1, ATFUT1, FT1, MUR2 | [59] | xyloglucan synthesis |
| Lus10041651 | cinnamoyl CoA reductase 1 | AT1G15950, ATCCR1, IRX4 | [60] | lignin biosynthesis | |
| Chr7:14140267 | Lus10025431 | RAC-like 2 | AT5G45970, ATRAC2, ATROP7 | [61] | secondary cell wall of xylem vessels |
| Chr8:1099671 | Lus10013589 | syntaxin of plants 121 | AT3G11820, ATSYP121, ATSYR1, PEN1 | [62] | secretory traffic to the plasma membrane |
| Chr9:693002 Chr9:695025 | Lus10010153 | NOD26-like intrinsic protein 5;1 | AT4G1038, NIP5;1, NLM6, NLM8 | [63] | aquaporin |
| Lus10010190 | serine carboxypeptidase-like 34 | AT5G23210, SCPL34 | [64] | present in cell wall of elongating cells | |
| Chr9:1929944 | Lus10008974 | beta-galactosidase 12 | AT4G26140, BGAL12 | [65] | plant cell wall remodeling |
| Chr9:5255847 | Lus10004364 | alpha/beta-hydrolases superfamily protein | AT4G16820, PLA-I{beta]2 | [56] | breaking of carbon–carbon bonds |
| Lus10004377 | homeobox protein 2 | AT4G1678, ATHB2, HAT4, HB-2 | [66] | cell expansion and cell proliferation in the response to auxin | |
| Chr9:8233979 | Lus10007455 | disease resistance protein (TIR-NBS-LRR class) family | AT3G4448, cog1, RPP1 | [67] | plant immunity |
| Chr9:11359847 | Lus10019631 | alpha/beta-hydrolases superfamily protein | AT1G72620 | [56] | breaking of carbon–carbon bonds |
| Chr9:17239226 | Lus10042571 | aluminum-activated malate transporter family protein | AT5G46600 | [68] | transporter |
| Chr9:17408991 | Lus10042531 | NAC domain-containing protein 47 | AT3G0407, NAC047, SHYG | [69] | NAC TF, localized longitudinal cell expansion |
| Lus10042537 | malectin/receptor-like protein kinase family protein | AT3G04690, ANX1 | [70] | perception and relay processes at cell membranes | |
| Chr9:18936691 | Lus10024808 | WUSCHEL-related homeobox 4 | AT1G46480, WOX4 | [71] | regulates the dynamic balance of division and differentiation of plant stem cells |
| Lus10024818 | cytochrome P450, family 81, subfamily D, polypeptide 5 | AT4G3732, CYP81D5 | [72] | defense response | |
| Chr9:19858075 | Lus10000494 | flavin-binding monooxygenase family protein | AT4G2872, YUC8 | [73] | indole-3-pyruvate monooxygenase YUCCA8, responsible for auxin biosynthesis in leaves |
| Chr9:20032070- Chr9:20032149 | Lus10011868 | CLAVATA3/ESR-RELATED 41 | AT3G24770, CLE41 | [74] | regulates vascular cell division, vascular organization and xylem differentiation in vascular tissue |
| Lus10011858 | cytochrome B5 isoform B | AT2G32720, CB5-B | [75] | lignin biosynthesis | |
| Chr12:16047776 | Lus10043075 | O-glycosyl hydrolase family 17 protein | AT5G20870 | [76] | hydrolytic activity towards different types of hemicelluloses or callose |
| Chr13:1286479 | Lus10010665 | UDP-glycosyltransferase superfamily protein | AT1G2240, ATUGT85A1, UGT85A1 | [77,78] | cell cycle regulation, influences trans-zeatin (cytokinin) homeostasis likely through O-glycosylation |
| Lus10010666 | annexin 8 | AT5G12380, ANNAT8 | [79] | cell wall protein | |
| Chr13:3759775 | Lus10026123 | hydroxycinnamoyl-CoA shikimate/quinate hydroxycinnamoyl transferase | AT5G48930, HCT | [80] | lignin biosynthesis |
| Chr15:1632466 | Lus10013935 | alpha/beta-hydrolases superfamily protein | AT4G18550 | [56] | breaking of carbon–carbon bonds |
| Lus10013936 | alpha/beta-hydrolases superfamily protein | ||||
| Lus10013945 | O-methyltransferase family protein | AT4G35160 | transfer of methyl groups to various biomolecules |
| QTN | Gene | Functional Annotation | Arabidopsis Gene Name | Reference | Predicted Function |
|---|---|---|---|---|---|
| Chr2:734912 Chr2:748984 Chr2:752999 | Lus10019240 | glycosyl hydrolase family protein | AT5G10560 | [76] | hydrolysis of glycosidic bonds |
| Lus10019273 | basic leucine-zipper 6 | AT2G22850, AtbZIP6, bZIP6 | transcription factor | ||
| Chr2:3633762 Chr2:3633790 | Lus10003436 | glutamate receptor 2.7 | AT2G29120, GLR2.7 | [81] | non-selective cation channel |
| Chr2:6831421 | Lus10038682 | major facilitator superfamily protein | AT2G39210 | [43] | facilitates movement of small solutes across cell membranes |
| Chr2:17384359 Chr2:17384362 | Lus10033212 | RAD-like 6 | AT1G75250, RL6, RSM3 | MYB family TF | |
| Chr3:12697091 | Lus10005850 | aldolase superfamily protein | AT4G26530 | [82] | modulates V-ATPase-dependent vesicular trafficking events and actin cytoskeleton remodeling |
| Chr4:460524 | Lus10030216 | cytochrome P450, family 78, subfamily A, polypeptide 6 | AT2G46660, CYP78A6, EOD3 | [52,53,54] | cell expansion, enhancer of DA-1 3, seed and fruit development |
| Chr4:2376445 | Lus10029458 | OBF-binding protein 3 | AT3G55370, OBP3 | [83] | TF, pays a role in plant growth and development, induced by SA and auxin |
| Lus10029464 | nodulin MtN21/EamA-like transporter family protein | AT2G39510 | [84,85,86] | amino acid and auxin transporters | |
| Chr4:9042565 | Lus10029065 | peroxidase superfamily protein, class III | AT5G05340, AtperoxP52 | [40,41] | cell wall localized proteins tightly associated to its loosening and stiffening |
| Lus10029063 | major facilitator superfamily protein | AT2G40460 | [43] | facilitates movement of small solutes across cell membranes | |
| Chr4:17106278 | Lus10004043 | myb domain protein 20 | AT1G66230 | [87] | activates lignin and phenylalanine biosynthesis genes during secondary wall formation |
| Chr4:18493683 | Lus10030007 Lus10030008 | glycosyl hydrolase 9B8 | AT2G32990, GH9B8 | [88] | membrane endo (1→4)-β-D-glucanase (cellulase), wall assembly and cell elongation, fruit ripening and floral abscission |
| Chr8:4371945 Chr8:4833196 | Lus10034491 | major facilitator superfamily protein | AT4G34950 | [43] | facilitates movement of small solutes across cell membranes |
| Lus10023931 | plant regulator RWP-RK family protein | AT2G17150 | [89] | control of cell differentiation | |
| Chr8:18364995 | Lus10007808, Lus10007809, Lus10007810, Lus10007811, Lus10007813, Lus10007814 | disease resistance protein (TIR-NBS-LRR class) family | AT5G36930 | [90] | plant immunity |
| Chr8:19022659 | Lus10002243 | subtilase family protein | AT1G04110, SDD1 | [91] | cell wall enzyme |
| Chr9:971861 | Lus10007528, Lus10007529 | TRICHOME BIREFRINGENCE-LIKE 33 | AT2G40320, TBL33 | [92] | contribute to the synthesis and deposition of secondary wall cellulose, presumably by influencing the esterification state of pectic polymers |
| Lus10007532 | laccase 5 | AT2G40370, LAC5 | [93] | lignin degradation and detoxification of lignin-derived product | |
| Chr9:7152630 | Lus10000957 | expansin-like B1 | AT4G17030, ATEXLB1, ATEXPR1, ATHEXPBETA 3.1, EXLB1, EXPR | [79] | cell wall loosening |
| Chr9:7487990 Chr9:7488554 | Lus10024499 | IQ-domain 22 | AT4G23060, IQD22 | [94,95] | Ca2+/CaM-regulated scaffold for cellular transport of specific cargo along microtubules |
| Lus10024485 | KNOTTED-like homeobox of Arabidopsis thaliana 7 | AT1G62990, IXR11, KNAT7 | [96] | Secondary-wall-associated transcription factor | |
| Chr9:8240615 Chr9:8240618 | Lus10007455 | disease resistance protein (TIR-NBS-LRR class) family | AT3G44480, cog1, RPP1 | [90] | plant immunity |
| Chr9:9241973 | Lus10006152 | cytochrome P450, family 90, subfamily D, polypeptide 1 | AT3G13730, CYP90D1 | [72] | modification of cyclic terpenes and sterols in the brassinosteroid, abscisic acid and gibberellin pathways |
| Chr9:17172377 Chr9:17190919 | Lus10042571 | aluminum-activated malate transporter family protein | AT5G46600 | [68] | transporter |
| Chr9:17511839 Chr9:17513033 | Lus10042531 | NAC domain-containing protein 47 | AT3G04070, anac047, NAC047 | [69] | transcription factor |
| Lus10042516 | sugar transporter 1 | AT1G11260, STP1 | [97] | H+/monosaccharide cotransporter | |
| Chr9:18121668 | Lus10029638 Lus10029639 | GDSL-like lipase/acylhydrolase superfamily protein | AT1G54790 | [98,99,100] | growth, biotic stress response, in cotton the GDSL (GhGDSL) li pase/hydrolase gene (CotAD_74480) is expressed during SCW biosynthesis |
| Chr11:11843614 | Lus10021553 | auxin-responsive family protein | AT3G25290 | [101] | TF, bind to AuxRE in the promoters of auxin-regulated genes |
| Chr12:16474392 | Lus10027867 | reticulon family protein | AT3G19460 | [102] | endoplasmic reticulum-Golgi trafficking, vesicle formation and membrane morphogenesis |
| Chr13:12832925 | Lus10010638 | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | AT1G27660, PFA5 | [103] | governs the competence of pericycle cells to initiate lateral root primordium formation |
| Chr13:18496268 | Lus10030901 | cysteine-rich RLK (RECEPTOR-like protein kinase) 2 | AT1G70520, CRK2 | [104] | plant immunity |
| Chr14:3502060 | Lus10020525 | disease resistance protein (TIR-NBS-LRR class) family | AT4G16950, RPP5 | [105] | plant immunity |
| Chr14:13328874 | Lus10032902 | HSP20-like chaperones superfamily protein | AT1G54400 | [42] | enhances membrane stability and detoxifies ROS by regulating the antioxidant enzymes system |
| Chr15:124776 | Lus10007632 Lus10007633 | cysteine-rich RLK (RECEPTOR-like protein kinase) 25 | AT4G05200, CRK25 | [106] | plant immunity |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kanapin, A.; Rozhmina, T.; Bankin, M.; Surkova, S.; Duk, M.; Osyagina, E.; Samsonova, M. Genetic Determinants of Fiber-Associated Traits in Flax Identified by Omics Data Integration. Int. J. Mol. Sci. 2022, 23, 14536. https://doi.org/10.3390/ijms232314536
Kanapin A, Rozhmina T, Bankin M, Surkova S, Duk M, Osyagina E, Samsonova M. Genetic Determinants of Fiber-Associated Traits in Flax Identified by Omics Data Integration. International Journal of Molecular Sciences. 2022; 23(23):14536. https://doi.org/10.3390/ijms232314536
Chicago/Turabian StyleKanapin, Alexander, Tatyana Rozhmina, Mikhail Bankin, Svetlana Surkova, Maria Duk, Ekaterina Osyagina, and Maria Samsonova. 2022. "Genetic Determinants of Fiber-Associated Traits in Flax Identified by Omics Data Integration" International Journal of Molecular Sciences 23, no. 23: 14536. https://doi.org/10.3390/ijms232314536
APA StyleKanapin, A., Rozhmina, T., Bankin, M., Surkova, S., Duk, M., Osyagina, E., & Samsonova, M. (2022). Genetic Determinants of Fiber-Associated Traits in Flax Identified by Omics Data Integration. International Journal of Molecular Sciences, 23(23), 14536. https://doi.org/10.3390/ijms232314536






