Evolutionary Divergence of Phosphorylation to Regulate Interactive Protein Networks in Lower and Higher Species
Abstract
1. Introduction
2. Results
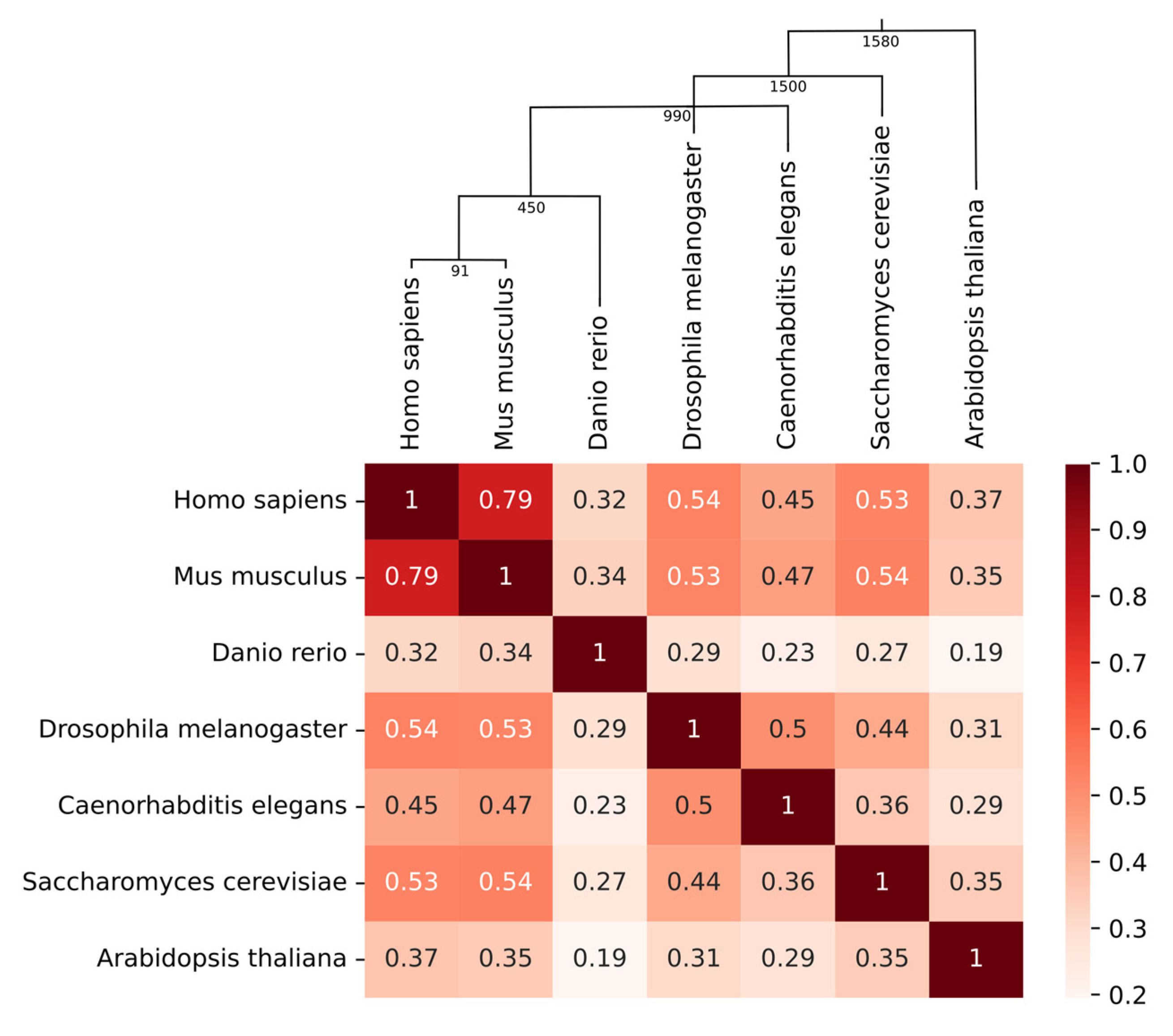
2.1. Conservation of Phosphorylation in Homolog Proteins of Phylogenetically Distant Species
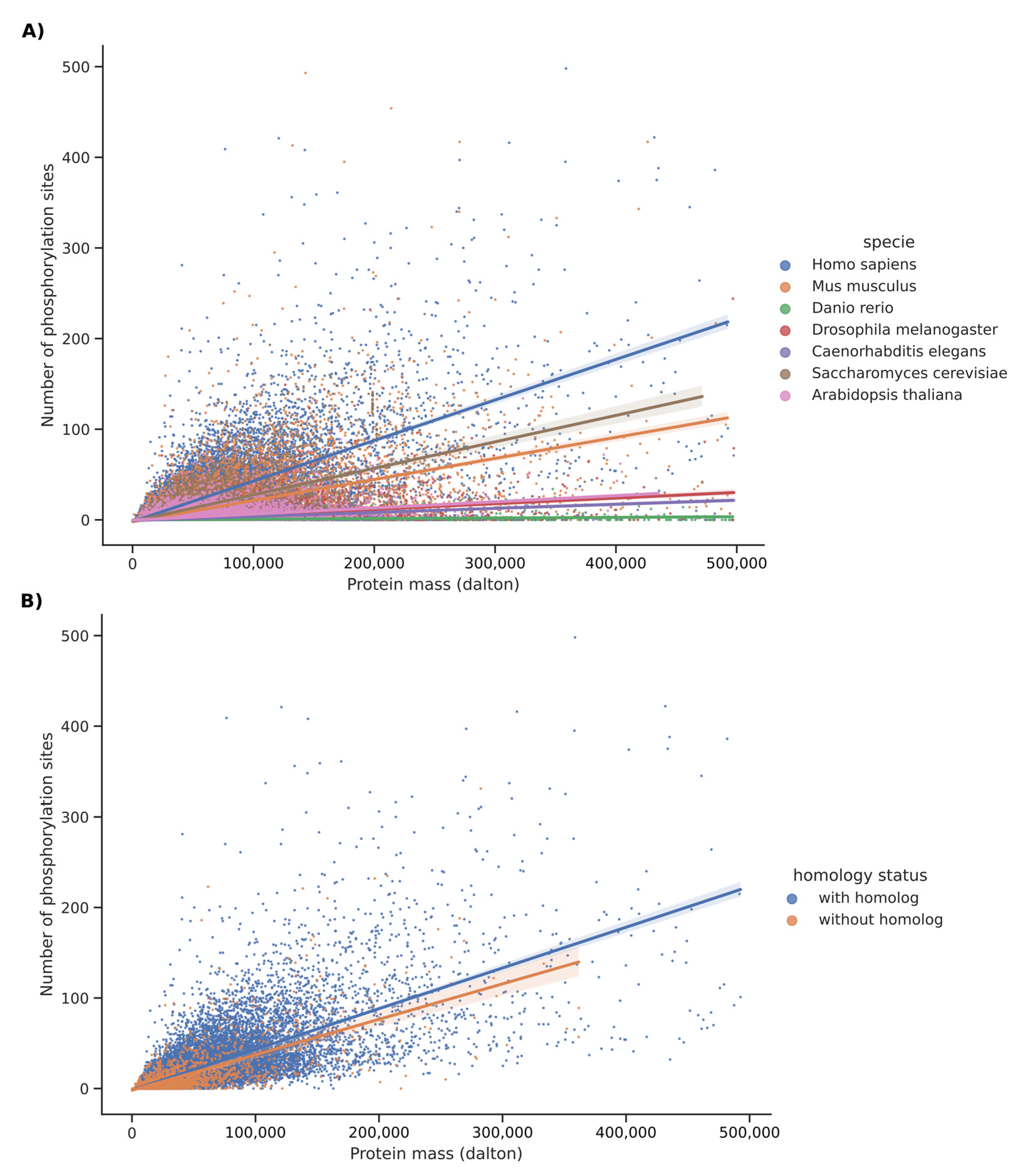
2.2. Comparative Rate of Phospho-Sites in Homolog Versus Unique Proteins for Each of These Species
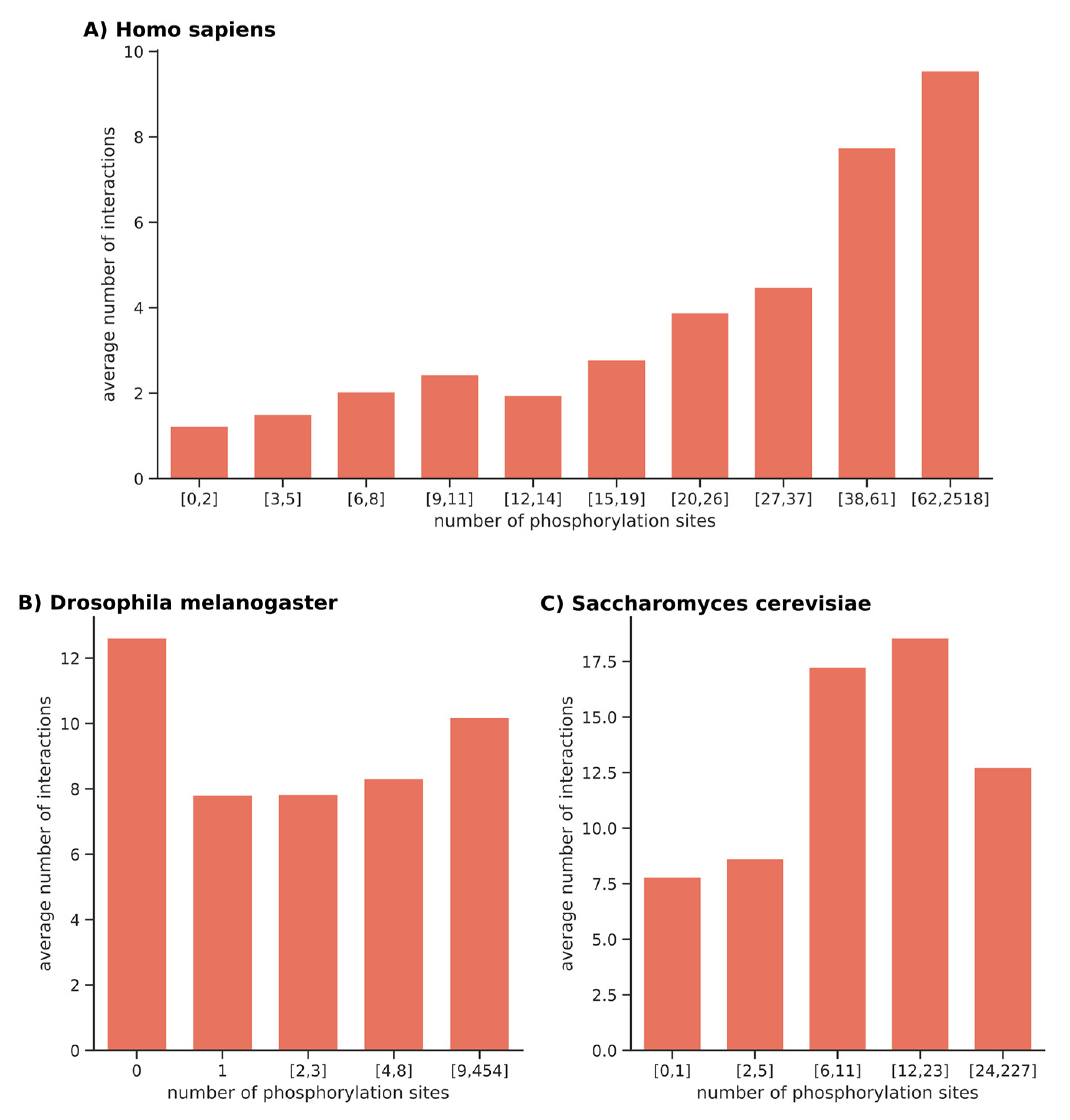
2.3. Correlation between the Degree of Phosphorylation of Proteins and the Size of Interacting Networks
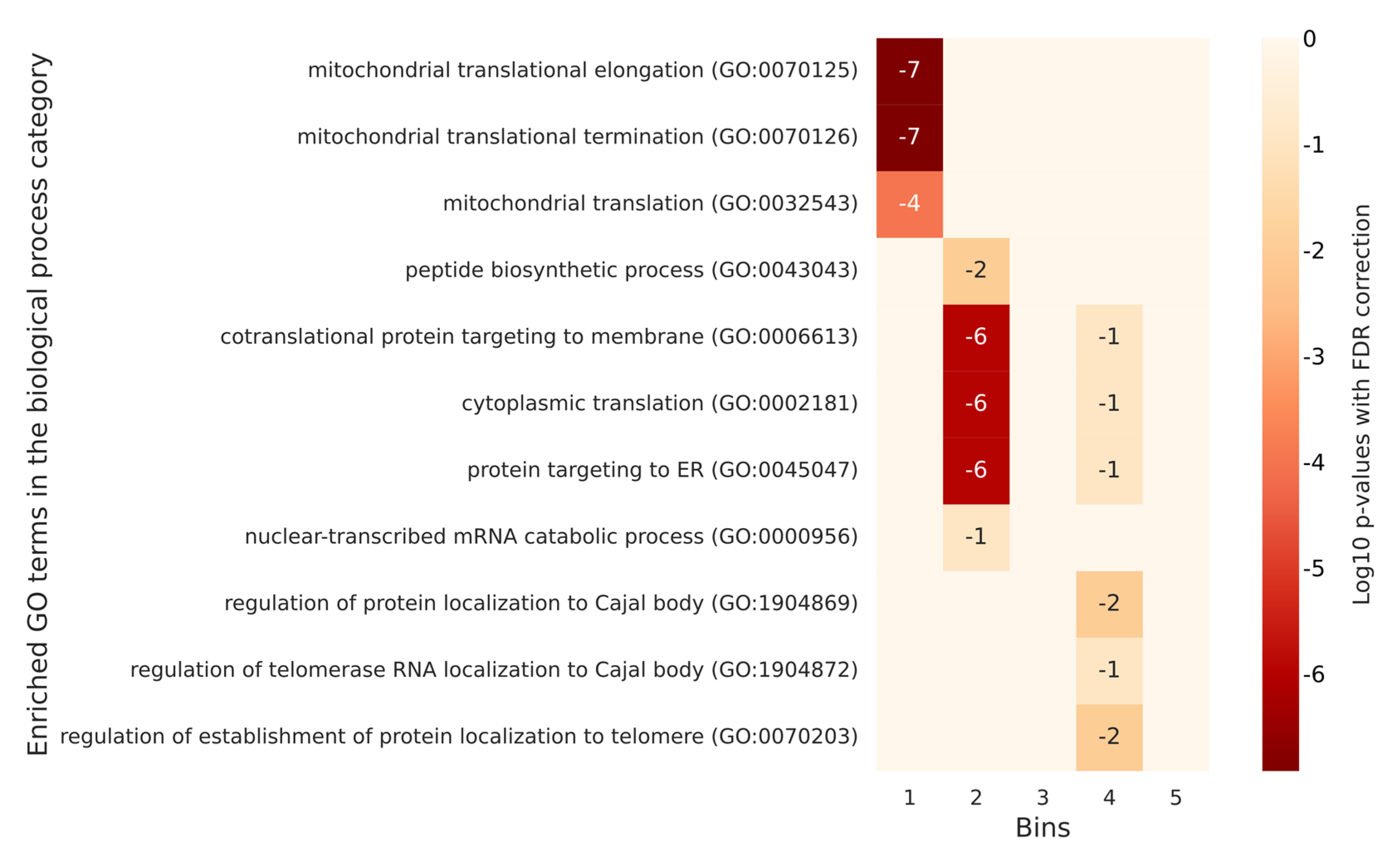
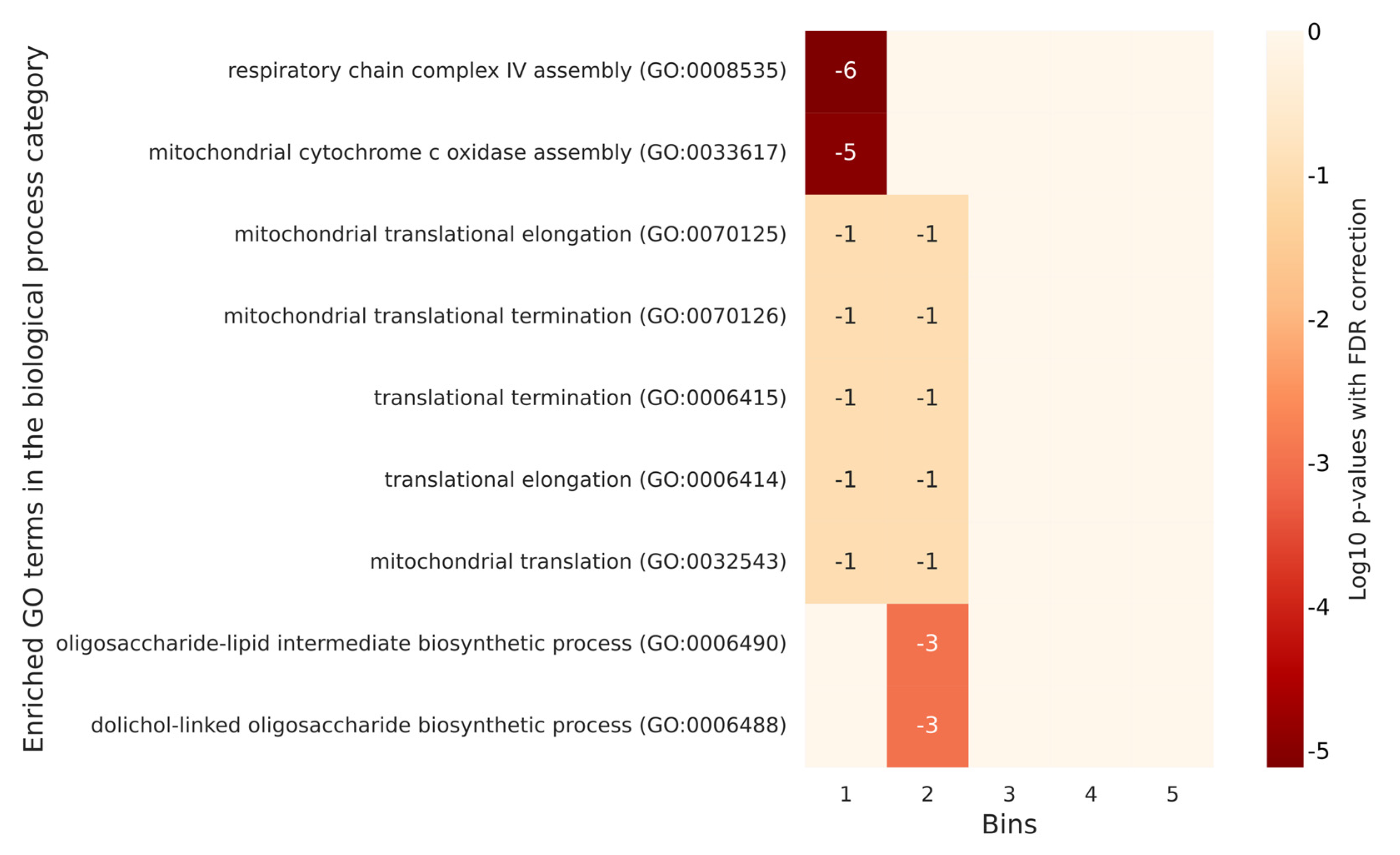
2.4. GO Annotations Analysis Based on the Degree of Phosphorylation of Proteins
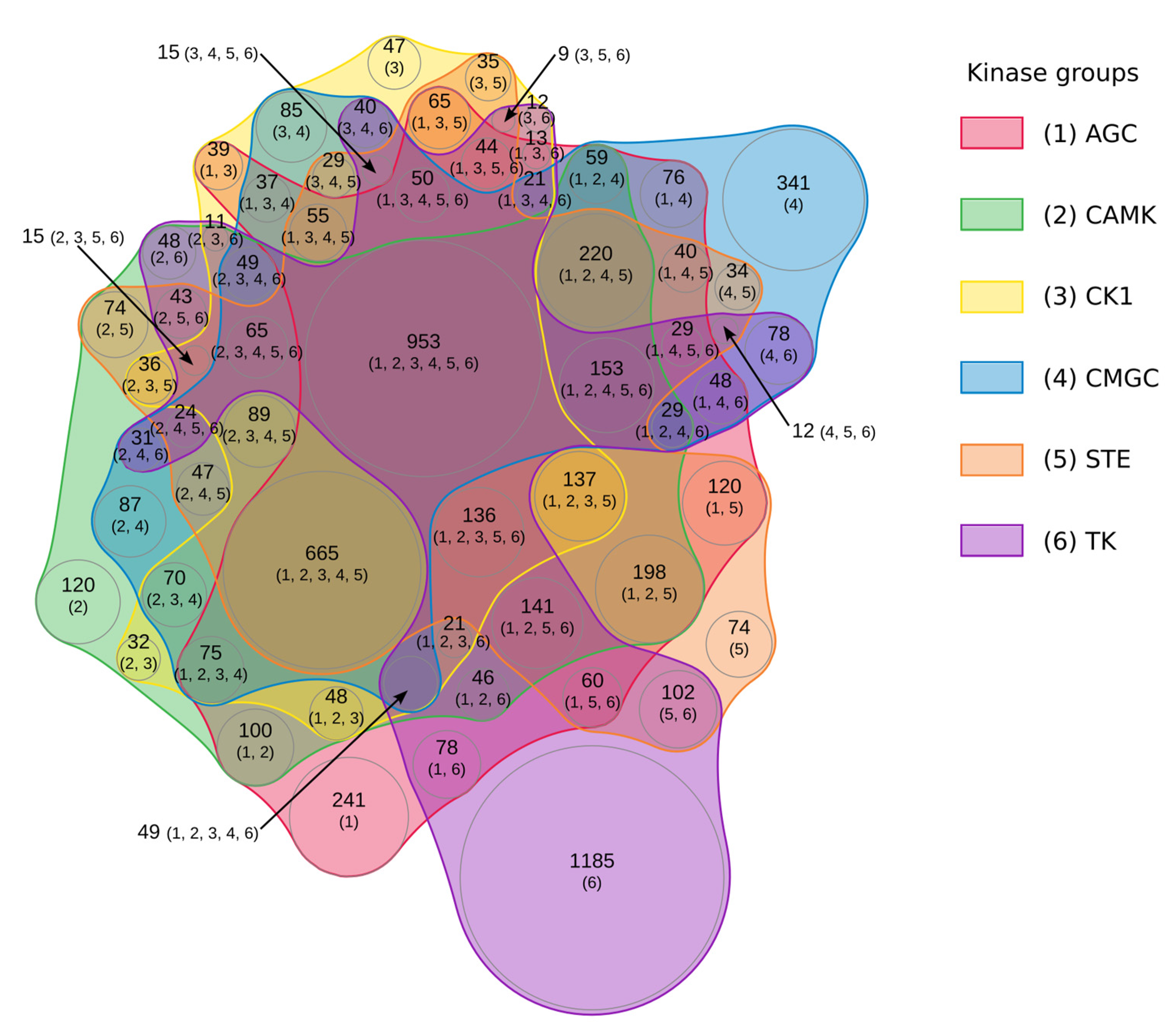
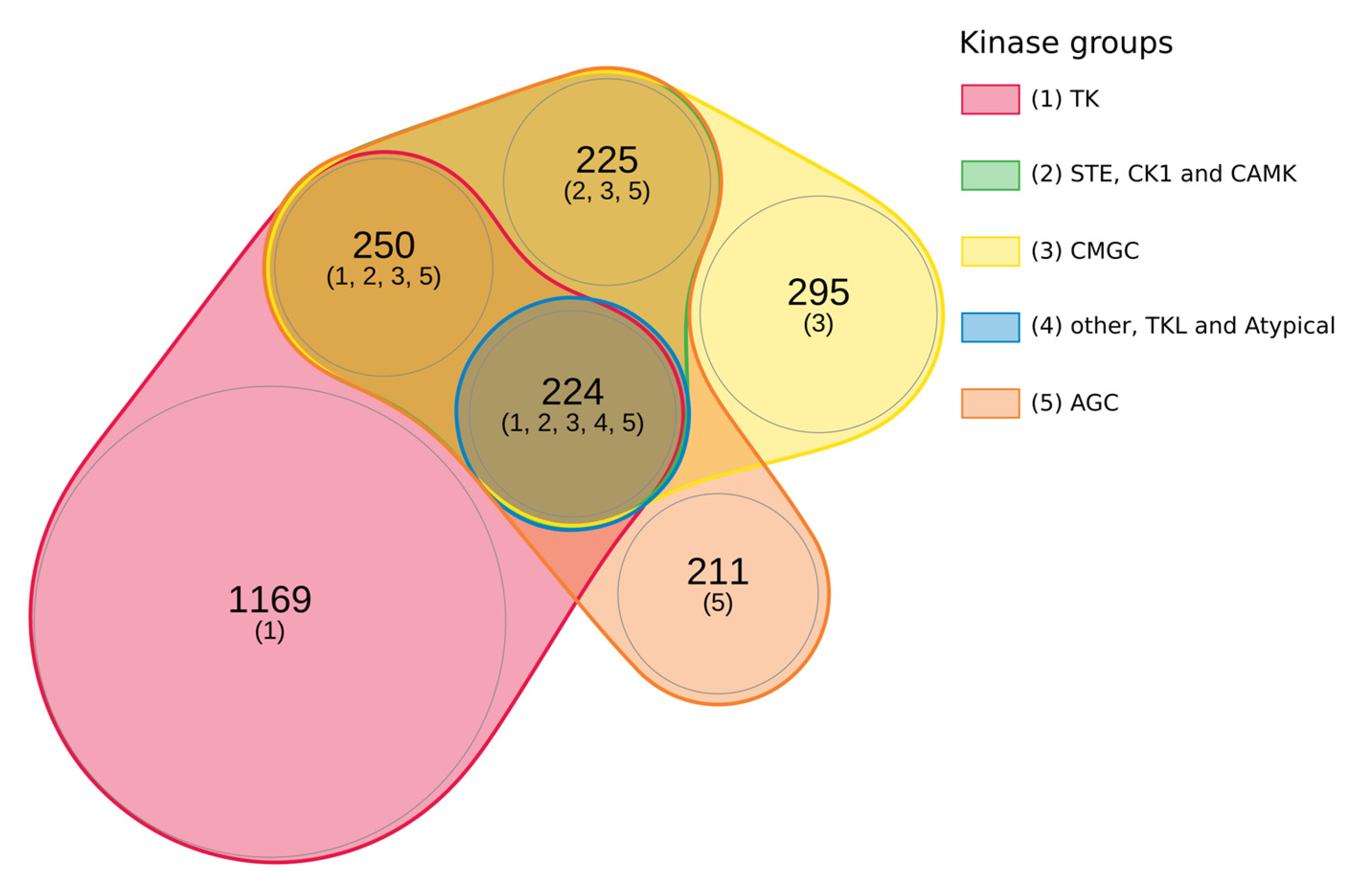
2.5. Integrated Substrate Map for Six Different Kinases Families Highlights a Significant Overlapping
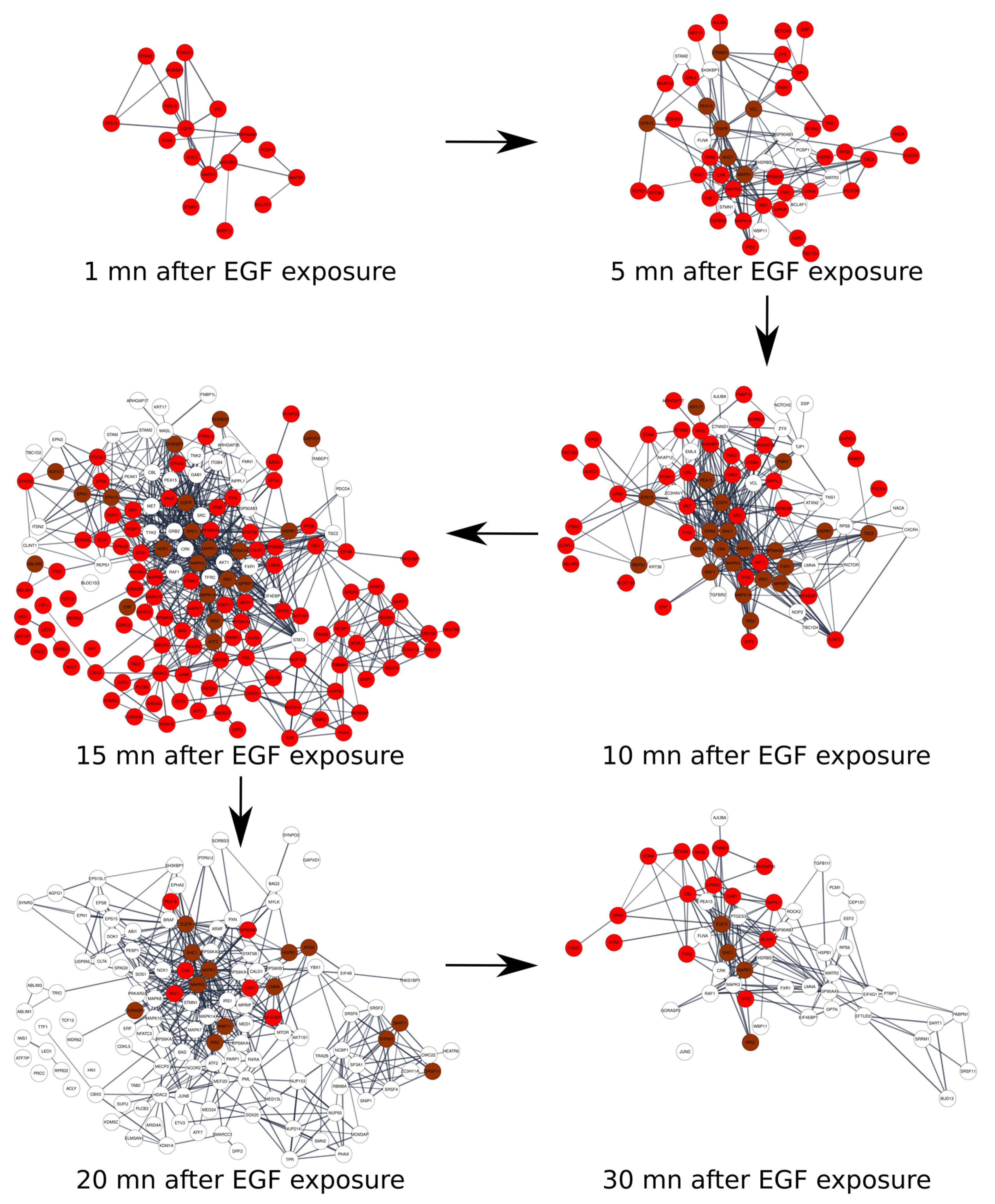
2.6. Time Course of Phospho-Proteome and Time Dependent Specific Changes in Networks Induced by EGF Factor
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Data Gathering
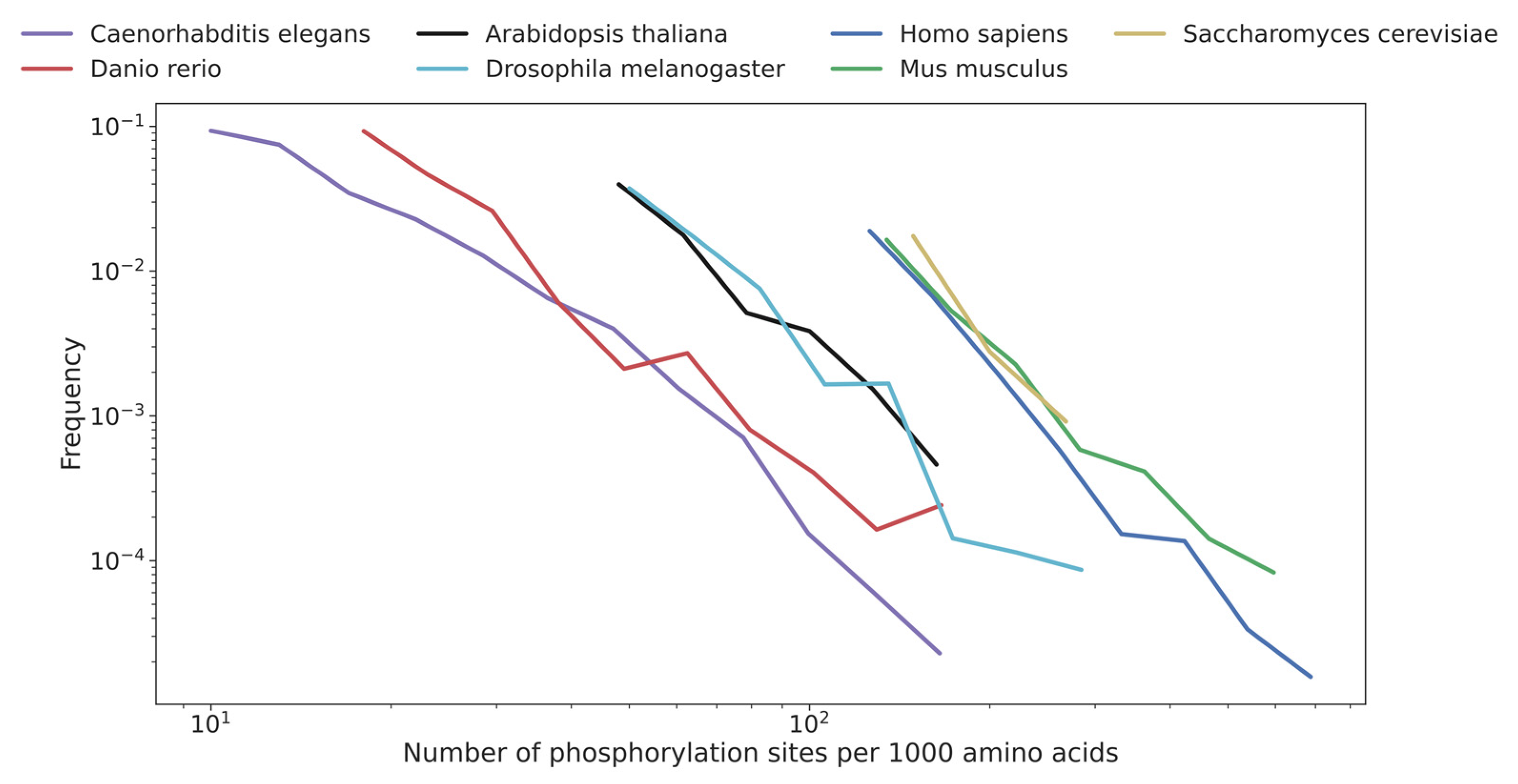
5.2. Computing Correlation between the Distribution of the Number of Phosphorylation Sites Per Protein in Different Species
5.3. Correlation between Phosphorylation and Protein Interactions
5.4. Enrichment Analysis
5.5. Study of Phorphorylated Proteins According to the Kinase group
5.6. Dynamic Analysis
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Johnson, L.N.; Barford, D. The Effects of Phosphorylation on the Structure and Function of Proteins. Annu. Rev. Biophys. Biomol. Struct. 1993, 22, 199–232. [Google Scholar] [CrossRef] [PubMed]
- Johnson, L.N. The Regulation of Protein Phosphorylation. Biochem. Soc. Trans. 2009, 37, 627–641. [Google Scholar] [CrossRef] [PubMed]
- Vlastaridis, P.; Kyriakidou, P.; Chaliotis, A.; Van de Peer, Y.; Oliver, S.G.; Amoutzias, G.D. Estimating the Total Number of Phosphoproteins and Phosphorylation Sites in Eukaryotic Proteomes. GigaScience 2017, 6, giw015. [Google Scholar] [CrossRef] [PubMed]
- Swaney, D.L.; Beltrao, P.; Starita, L.; Guo, A.; Rush, J.; Fields, S.; Krogan, N.J.; Villén, J. Global Analysis of Phosphorylation and Ubiquitylation Cross-Talk in Protein Degradation. Nat. Methods 2013, 10, 676–682. [Google Scholar] [CrossRef]
- Nishi, H.; Shaytan, A.; Panchenko, A.R. Physicochemical Mechanisms of Protein Regulation by Phosphorylation. Front. Genet. 2014, 5, 270. [Google Scholar] [CrossRef]
- Adam, K.; Hunter, T. Histidine Kinases and the Missing Phosphoproteome from Prokaryotes to Eukaryotes. Lab. Investig. 2018, 98, 233–247. [Google Scholar] [CrossRef]
- Cohen, P. The Regulation of Protein Function by Multisite Phosphorylation—A 25 Year Update. Trends Biochem. Sci. 2000, 25, 596–601. [Google Scholar] [CrossRef]
- Pearlman, S.M.; Serber, Z.; Ferrell, J.E. A Mechanism for the Evolution of Phosphorylation Sites. Cell 2011, 147, 934–946. [Google Scholar] [CrossRef]
- Levchenko, A.; Bruck, J.; Sternberg, P.W. Scaffold Proteins May Biphasically Affect the Levels of Mitogen-Activated Protein Kinase Signaling and Reduce Its Threshold Properties. Proc. Natl. Acad. Sci. USA 2000, 97, 5818–5823. [Google Scholar] [CrossRef]
- Markevich, N.I.; Hoek, J.B.; Kholodenko, B.N. Signaling switches and bistability arising from multisite phosphorylation in protein kinase cascades. J. Cell Biol. 2004, 164, 353–359. [Google Scholar] [CrossRef]
- Good, M.C.; Zalatan, J.G.; Lim, W.A. Scaffold Proteins: Hubs for Controlling the Flow of Cellular Information. Science 2011, 332, 680–686. [Google Scholar] [CrossRef] [PubMed]
- Miller, C.J.; Turk, B.E. Homing in: Mechanisms of Substrate Targeting by Protein Kinases. Trends Biochem. Sci. 2018, 43, 380–394. [Google Scholar] [CrossRef] [PubMed]
- Rust, H.L.; Thompson, P.R. Kinase Consensus Sequences: A Breeding Ground for Crosstalk. ACS Chem. Biol. 2011, 6, 881–892. [Google Scholar] [CrossRef] [PubMed]
- Sims, R.J.; Rojas, L.A.; Beck, D.; Bonasio, R.; Schüller, R.; Drury, W.J.; Eick, D.; Reinberg, D. The C-Terminal Domain of RNA Polymerase II Is Modified by Site-Specific Methylation. Science 2011, 332, 99–103. [Google Scholar] [CrossRef]
- Kowenz-Leutz, E.; Pless, O.; Dittmar, G.; Knoblich, M.; Leutz, A. Crosstalk between C/EBPβ Phosphorylation, Arginine Methylation, and SWI/SNF/Mediator Implies an Indexing Transcription Factor Code. EMBO J. 2010, 29, 1105–1115. [Google Scholar] [CrossRef] [PubMed]
- Mowen, K.A.; Tang, J.; Zhu, W.; Schurter, B.T.; Shuai, K.; Herschman, H.R.; David, M. Arginine Methylation of STAT1 Modulates IFNα/β-Induced Transcription. Cell 2001, 104, 731–741. [Google Scholar] [CrossRef]
- Bodenmiller, B.; Malmstrom, J.; Gerrits, B.; Campbell, D.; Lam, H.; Schmidt, A.; Rinner, O.; Mueller, L.N.; Shannon, P.T.; Pedrioli, P.G. PhosphoPep—A Phosphoproteome Resource for Systems Biology Research in Drosophila Kc167 Cells. Mol. Syst. Biol. 2007, 3, 139. [Google Scholar] [CrossRef]
- Ullah, S.; Lin, S.; Xu, Y.; Deng, W.; Ma, L.; Zhang, Y.; Liu, Z.; Xue, Y. DbPAF: An Integrative Database of Protein Phosphorylation in Animals and Fungi. Sci. Rep. 2016, 6, 23534. [Google Scholar] [CrossRef]
- Zolg, D.P.; Wilhelm, M.; Schnatbaum, K.; Zerweck, J.; Knaute, T.; Delanghe, B.; Bailey, D.J.; Gessulat, S.; Ehrlich, H.-C.; Weininger, M.; et al. Building Proteome Tools Based on a Complete Synthetic Human Proteome. Nat. Methods 2017, 14, 259–262. [Google Scholar] [CrossRef]
- Amano, M.; Hamaguchi, T.; Shohag, M.H.; Kozawa, K.; Kato, K.; Zhang, X.; Yura, Y.; Matsuura, Y.; Kataoka, C.; Nishioka, T.; et al. Kinase-Interacting Substrate Screening Is a Novel Method to Identify Kinase Substrates. J. Cell Biol. 2015, 209, 895–912. [Google Scholar] [CrossRef]
- Hardman, G.; Perkins, S.; Ruan, Z.; Kannan, N.; Brownridge, P.; Byrne, D.P.; Eyers, P.A.; Jones, A.R.; Eyers, C.E. Extensive Non-Canonical Phosphorylation in Human Cells Revealed Using Strong-Anion Exchange-Mediated Phosphoproteomics. BioRxiv 2017, 202820. [Google Scholar] [CrossRef]
- Corwin, T.; Woodsmith, J.; Apelt, F.; Fontaine, J.-F.; Meierhofer, D.; Helmuth, J.; Grossmann, A.; Andrade-Navarro, M.A.; Ballif, B.A.; Stelzl, U. Defining Human Tyrosine Kinase Phosphorylation Networks Using Yeast as an in Vivo Model Substrate. Cell Syst. 2017, 5, 128–139. [Google Scholar] [CrossRef]
- Lee, T.-Y.; Huang, H.-D.; Hung, J.-H.; Huang, H.-Y.; Yang, Y.-S.; Wang, T.-H. DbPTM: An Information Repository of Protein Post-Translational Modification. Nucleic Acids Res. 2006, 34, D622–D627. [Google Scholar] [CrossRef] [PubMed]
- Roux, P.P.; Blenis, J. ERK and P38 MAPK-Activated Protein Kinases: A Family of Protein Kinases with Diverse Biological Functions. Microbiol. Mol. Biol. Rev. 2004, 68, 320–344. [Google Scholar] [CrossRef] [PubMed]
- Peti, W.; Page, R. Molecular Basis of MAP Kinase Regulation. Protein Sci. 2013, 22, 1698–1710. [Google Scholar] [CrossRef] [PubMed]
- Bandyopadhyay, S.; Chiang, C.; Srivastava, J.; Gersten, M.; White, S.; Bell, R.; Kurschner, C.; Martin, C.H.; Smoot, M.; Sahasrabudhe, S.; et al. A Human MAP Kinase Interactome. Nat. Methods 2010, 7, 801–805. [Google Scholar] [CrossRef]
- Diella, F.; Cameron, S.; Gemünd, C.; Linding, R.; Via, A.; Kuster, B.; Sicheritz-Pontén, T.; Blom, N.; Gibson, T.J. Phospho. ELM: A Database of Experimentally Verified Phosphorylation Sites in Eukaryotic Proteins. BMC Bioinform. 2004, 5, 79. [Google Scholar] [CrossRef] [PubMed]
- Dinkel, H.; Chica, C.; Via, A.; Gould, C.M.; Jensen, L.J.; Gibson, T.J.; Diella, F. Phospho. ELM: A Database of Phosphorylation Sites—Update 2011. Nucleic Acids Res. 2010, 39, D261–D267. [Google Scholar] [CrossRef]
- Huang, K.-Y.; Su, M.-G.; Kao, H.-J.; Hsieh, Y.-C.; Jhong, J.-H.; Cheng, K.-H.; Huang, H.-D.; Lee, T.-Y. DbPTM 2016: 10-Year Anniversary of a Resource for Post-Translational Modification of Proteins. Nucleic Acids Res. 2016, 44, D435–D446. [Google Scholar] [CrossRef]
- Gnad, F.; Gunawardena, J.; Mann, M. PHOSIDA 2011: The Posttranslational Modification Database. Nucleic Acids Res. 2010, 39, D253–D260. [Google Scholar] [CrossRef]
- Hornbeck, P.V.; Kornhauser, J.M.; Tkachev, S.; Zhang, B.; Skrzypek, E.; Murray, B.; Latham, V.; Sullivan, M. PhosphoSitePlus: A Comprehensive Resource for Investigating the Structure and Function of Experimentally Determined Post-Translational Modifications in Man and Mouse. Nucleic Acids Res. 2012, 40, D261–D270. [Google Scholar] [CrossRef] [PubMed]
- Hornbeck, P.V.; Zhang, B.; Murray, B.; Kornhauser, J.M.; Latham, V.; Skrzypek, E. PhosphoSitePlus, 2014: Mutations, PTMs and Recalibrations. Nucleic Acids Res. 2015, 43, D512–D520. [Google Scholar] [CrossRef] [PubMed]
- Hornbeck, P.V.; Kornhauser, J.M.; Latham, V.; Murray, B.; Nandhikonda, V.; Nord, A.; Skrzypek, E.; Wheeler, T.; Zhang, B.; Gnad, F. 15 Years of PhosphoSitePlus®: Integrating Post-Translationally Modified Sites, Disease Variants and Isoforms. Nucleic Acids Res. 2019, 47, D433–D441. [Google Scholar] [CrossRef] [PubMed]
- Goel, R.; Harsha, H.C.; Pandey, A.; Prasad, T.K. Human Protein Reference Database and Human Proteinpedia as Resources for Phosphoproteome Analysis. Mol. BioSyst. 2012, 8, 453–463. [Google Scholar] [CrossRef] [PubMed]
- The UniProt Consortium UniProt: A Worldwide Hub of Protein Knowledge. Nucleic Acids Res. 2019, 47, D506–D515. [CrossRef] [PubMed]
- Safaei, J.; Maňuch, J.; Gupta, A.; Stacho, L.; Pelech, S. Prediction of 492 Human Protein Kinase Substrate Specificities. In Proceedings of the Proteome Science; Springer: Berlin/Heidelberg, Germany, 2011; Volume 9, pp. 1–13. [Google Scholar]
- Sadowski, I.; Breitkreutz, B.-J.; Stark, C.; Su, T.-C.; Dahabieh, M.; Raithatha, S.; Bernhard, W.; Oughtred, R.; Dolinski, K.; Barreto, K.; et al. The PhosphoGRID Saccharomyces Cerevisiae Protein Phosphorylation Site Database: Version 2.0 Update. Database 2013, 2013, bat026. [Google Scholar] [CrossRef]
- Cheng, H.; Deng, W.; Wang, Y.; Ren, J.; Liu, Z.; Xue, Y. DbPPT: A Comprehensive Database of Protein Phosphorylation in Plants. Database 2014, 2014, bau121. [Google Scholar] [CrossRef]
- Lin, S.; Wang, C.; Zhou, J.; Shi, Y.; Ruan, C.; Tu, Y.; Yao, L.; Peng, D.; Xue, Y. EPSD: A Well-Annotated Data Resource of Protein Phosphorylation Sites in Eukaryotes. Brief. Bioinform. 2021, 22, 298–307. [Google Scholar] [CrossRef]
- Bai, Y.; Chen, B.; Li, M.; Zhou, Y.; Ren, S.; Xu, Q.; Chen, M.; Wang, S. FPD: A Comprehensive Phosphorylation Database in Fungi. Fungal Biol. 2017, 121, 869–875. [Google Scholar] [CrossRef]
- Rose, C.; Venkateshwaran, M.; Grimsrud, P.; Westphall, M.; Sussman, M.; Coon, J.; Ané, J.-M. Medicago PhosphoProtein Database: A Repository for Medicago Truncatula Phosphoprotein Data. Front. Plant Sci. 2012, 3, 122. [Google Scholar] [CrossRef]
- Gao, J.; Agrawal, G.K.; Thelen, J.J.; Xu, D. P3DB: A Plant Protein Phosphorylation Database. Nucleic Acids Res. 2009, 37, D960–D962. [Google Scholar] [CrossRef]
- Heazlewood, J.L.; Durek, P.; Hummel, J.; Selbig, J.; Weckwerth, W.; Walther, D.; Schulze, W.X. PhosPhAt: A Database of Phosphorylation Sites in Arabidopsis Thaliana and a Plant-Specific Phosphorylation Site Predictor. Nucleic Acids Res. 2007, 36, D1015–D1021. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Xing, X.; Ding, G.; Li, Q.; Wang, C.; Xie, L.; Zeng, R.; Li, Y. SysPTM: A Systematic Resource for Proteomic Research on Post-Translational Modifications. Mol. Cell. Proteom. 2009, 8, 1839–1849. [Google Scholar] [CrossRef] [PubMed]
- Rapaport, F.; Zinovyev, A.; Dutreix, M.; Barillot, E.; Vert, J.-P. Classification of Microarray Data Using Gene Networks. BMC Bioinform. 2007, 8, 35. [Google Scholar] [CrossRef] [PubMed]
- Pasquier, C.; Guerlais, V.; Pallez, D.; Rapetti-Mauss, R.; Soriani, O. Identification of Active Modules in Interaction Networks Using Node2vec Network Embedding. BioRxiv 2021. [Google Scholar] [CrossRef]
- Lahiry, P.; Torkamani, A.; Schork, N.J.; Hegele, R.A. Kinase Mutations in Human Disease: Interpreting Genotype–Phenotype Relationships. Nat. Rev. Genet. 2010, 11, 60–74. [Google Scholar] [CrossRef] [PubMed]
- Dammer, E.B.; Lee, A.K.; Duong, D.M.; Gearing, M.; Lah, J.J.; Levey, A.I.; Seyfried, N.T. Quantitative Phosphoproteomics of Alzheimer’s Disease Reveals Cross-Talk between Kinases and Small Heat Shock Proteins. Proteomics 2015, 15, 508–519. [Google Scholar] [CrossRef] [PubMed]
- Sathe, G.; Mangalaparthi, K.K.; Jain, A.; Darrow, J.; Troncoso, J.; Albert, M.; Moghekar, A.; Pandey, A. Multiplexed Phosphoproteomic Study of Brain in Patients with Alzheimer’s Disease and Age-Matched Cognitively Healthy Controls. OMICS A J. Integr. Biol. 2020, 24, 216–227. [Google Scholar] [CrossRef]
- Tagawa, K.; Homma, H.; Saito, A.; Fujita, K.; Chen, X.; Imoto, S.; Oka, T.; Ito, H.; Motoki, K.; Yoshida, C.; et al. Comprehensive Phosphoproteome Analysis Unravels the Core Signaling Network That Initiates the Earliest Synapse Pathology in Preclinical Alzheimer’s Disease Brain. Hum. Mol. Genet. 2015, 24, 540–558. [Google Scholar] [CrossRef]
- Galleano, I.; Harms, H.; Choudhury, K.; Khoo, K.; Delemotte, L.; Pless, S.A. Functional Cross-Talk between Phosphorylation and Disease-Causing Mutations in the Cardiac Sodium Channel Nav1. 5. Proc. Natl. Acad. Sci. USA 2021, 118, e2025320118. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING V11: Protein–Protein Association Networks with Increased Coverage, Supporting Functional Discovery in Genome-Wide Experimental Datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Li, H.; He, Y.; Sun, H.; Xia, L.; Wang, L.; Sun, B.; Ma, L.; Zhang, G.; Li, J.; et al. Construction and Deciphering of Human Phosphorylation-Mediated Signaling Transduction Networks. J. Proteome Res. 2015, 14, 2745–2757. [Google Scholar] [CrossRef] [PubMed]
- Trost, B.; Kusalik, A.; Napper, S. Computational Analysis of the Predicted Evolutionary Conservation of Human Phosphorylation Sites. PLoS ONE 2016, 11, e0152809. [Google Scholar] [CrossRef] [PubMed]
- Taylor, S.S.; Kornev, A.P. Protein Kinases: Evolution of Dynamic Regulatory Proteins. Trends Biochem. Sci. 2011, 36, 65–77. [Google Scholar] [CrossRef]
- Bradley, D.; Beltrao, P. Evolution of Protein Kinase Substrate Recognition at the Active Site. PLoS Biol. 2019, 17, e3000341. [Google Scholar] [CrossRef]
- Howard, C.J.; Hanson-Smith, V.; Kennedy, K.J.; Miller, C.J.; Lou, H.J.; Johnson, A.D.; Turk, B.E.; Holt, L.J. Ancestral Resurrection Reveals Evolutionary Mechanisms of Kinase Plasticity. eLife 2014, 3, e04126. [Google Scholar] [CrossRef]
- Pereira, S.F.F.; Goss, L.; Dworkin, J. Eukaryote-Like Serine/Threonine Kinases and Phosphatases in Bacteria. Microbiol. Mol. Biol. Rev. 2011, 75, 192–212. [Google Scholar] [CrossRef]
- Kannan, N.; Taylor, S.S.; Zhai, Y.; Venter, J.C.; Manning, G. Structural and Functional Diversity of the Microbial Kinome. PLoS Biol. 2007, 5, e17. [Google Scholar] [CrossRef]
- Lin, M.-H.; Sugiyama, N.; Ishihama, Y. Systematic Profiling of the Bacterial Phosphoproteome Reveals Bacterium-Specific Features of Phosphorylation. Sci. Signal. 2015, 8, rs10. [Google Scholar] [CrossRef]
- Gnad, F.; Forner, F.; Zielinska, D.F.; Birney, E.; Gunawardena, J.; Mann, M. Evolutionary Constraints of Phosphorylation in Eukaryotes, Prokaryotes, and Mitochondria. Mol. Cell. Proteom. 2010, 9, 2642–2653. [Google Scholar] [CrossRef]
- Gabaldón, T.; Koonin, E.V. Functional and Evolutionary Implications of Gene Orthology. Nat. Rev. Genet. 2013, 14, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Beltrao, P.; Bork, P.; Krogan, N.J.; van Noort, V. Evolution and Functional Cross-Talk of Protein Post-Translational Modifications. Mol. Syst. Biol. 2013, 9, 714. [Google Scholar] [CrossRef] [PubMed]
- Wagih, O.; Sugiyama, N.; Ishihama, Y.; Beltrao, P. Uncovering Phosphorylation-Based Specificities through Functional Interaction Networks. Mol. Cell. Proteom. 2016, 15, 236–245. [Google Scholar] [CrossRef] [PubMed]
- Tan, D.J.; Dvinge, H.; Christoforou, A.; Bertone, P.; Martinez Arias, A.; Lilley, K.S. Mapping organelle proteins and protein complexes in Drosophila melanogaster. J. Proteome Res. 2009, 8, 2667–2678. [Google Scholar] [CrossRef]
- Telford, M.J.; Budd, E.; Philippe, H. Phylogenomic Insights into Animal Evolution. Curr. Biol. 2015, 25, R876–R887. [Google Scholar] [CrossRef] [PubMed]
- Yoshizaki, H.; Okuda, S. Large-Scale Analysis of the Evolutionary Histories of Phosphorylation Motifs in the Human Genome. GigaScience 2015, 4, s13742-015-0057–6. [Google Scholar] [CrossRef]
- Yoshizaki, H.; Okuda, S. Elucidation of the Evolutionary Expansion of Phosphorylation Signaling Networks Using Comparative Phosphomotif Analysis. BMC Genom. 2014, 15, 546. [Google Scholar] [CrossRef]
- Strumillo, M.J.; Oplová, M.; Viéitez, C.; Ochoa, D.; Shahraz, M.; Busby, B.P.; Sopko, R.; Studer, R.A.; Perrimon, N.; Panse, V.G.; et al. Conserved Phosphorylation Hotspots in Eukaryotic Protein Domain Families. Nat. Commun. 2019, 10, 1977. [Google Scholar] [CrossRef]
- Manning, G.; Whyte, D.B.; Martinez, R.; Hunter, T.; Sudarsanam, S. The Protein Kinase Complement of the Human Genome. Science 2002, 298, 1912–1934. [Google Scholar] [CrossRef]
- Yu, K.; Zhang, Q.; Liu, Z.; Zhao, Q.; Zhang, X.; Wang, Y.; Wang, Z.-X.; Jin, Y.; Li, X.; Liu, Z.-X.; et al. QPhos: A Database of Protein Phosphorylation Dynamics in Humans. Nucleic Acids Res. 2019, 47, D451–D458. [Google Scholar] [CrossRef]
- Hedges, S.B. The Origin and Evolution of Model Organisms. Nat. Rev. Genet. 2002, 3, 838–849. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Doncheva, N.T.; Morris, J.H.; Gorodkin, J.; Jensen, L.J. Cytoscape StringApp: Network Analysis and Visualization of Proteomics Data. J. Proteome Res. 2018, 18, 623–632. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Silva, J.G.; Araujo-Voces, M.; Quesada, V. NVenn: Generalized, Quasi-Proportional Venn and Euler Diagrams. Bioinformatics 2018, 34, 2322–2324. [Google Scholar] [CrossRef]




| H. sapiens | M. musculus | D. rerio | D. melanogaster | C. elegans | S. cerevisiae | A. thaliana | |
|---|---|---|---|---|---|---|---|
| All proteins | |||||||
| count (%) | 157,160 (100%) | 169,976 (100.00%) | 175,752 (100.00%) | 95,752 (100.00%) | 81,312 (100.00%) | 40,952 (100.00%) | 122,984 (100.00%) |
| avg. length | 567.70 | 545.88 | 569.25 | 553.17 | 494.47 | 519.80 | 453.27 |
| avg. weight (dalton) | 63,239.35 | 60,899.63 | 63,738.36 | 61,708.17 | 55,752.23 | 58,769.06 | 50,616.29 |
| avg. nb of phos. sites | 26.33 | 12.16 | 0.40 | 3.00 | 1.78 | 15.60 | 3.10 |
| avg. nb of phos. sites/1000aa | 44.08 | 20.14 | 0.68 | 4.44 | 2.87 | 28.76 | 6.79 |
| Non-phosphorylated | |||||||
| count (%) | 8760 (5.57%) | 34,880 (20.52%) | 158,816 (90.36%) | 58,704 (61.31%) | 58,504 (71.95%) | 5312 (12.97%) | 56,992 (46.34%) |
| avg. length | 189.81 | 286.04 | 535.28 | 408.51 | 403.88 | 275.86 | 349.89 |
| avg. weight (dalton) | 21,073.58 | 32,059.07 | 60,001.19 | 45,756.87 | 45,696.83 | 31,230.69 | 39,157.83 |
| Non-phos. & homologous | |||||||
| count (%) | 4752 (3.02%) | 24,856 (14.62%) | 84,888 (48.30%) | 14,392 (15.03%) | 22,768 (28.00%) | 3672 (8.97%) | 17,920 (14.57%) |
| avg. length | 217.63 | 308.83 | 537.63 | 467.38 | 462.18 | 306.98 | 394.12 |
| avg. weight (dalton) | 24,179.38 | 34,604.76 | 60,277.16 | 52,470.09 | 52,047.64 | 34,790.76 | 44,152.47 |
| Non-phos. & non-homologous | |||||||
| count (%) | 4008 (2.55%) | 10,024 (5.90%) | 73,928 (42.06%) | 44,312 (46.28%) | 35,736 (43.95%) | 1640 (4.00%) | 39,072 (31.77%) |
| avg. length | 156.82 | 229.55 | 532.57 | 389.38 | 366.73 | 206.17 | 329.60 |
| avg. weight (alton) | 17,391.26 | 25,746.64 | 59,684.32 | 43,576.50 | 41,650.63 | 23,259.61 | 36,867.08 |
| Phosphorylated | |||||||
| count (%) | 148,400 (94.43%) | 135,096 (79.48%) | 16,936 (9.64%) | 37,048 (38.69%) | 22,808 (28.05%) | 35,640 (87.03%) | 65,992 (53.66%) |
| avg. length | 590.01 | 612.97 | 887.84 | 782.39 | 726.85 | 556.15 | 542.56 |
| avg. weight (alton) | 65,728.38 | 68,345.88 | 98,783.34 | 86,983.62 | 81,544.98 | 62,873.55 | 60,512.04 |
| avg. nb of phos. Sites | 27.88 | 15.31 | 4.14 | 7.75 | 6.34 | 17.93 | 5.78 |
| avg. nb of phos. Sites/1000aa | 46.69 | 25.34 | 7.02 | 11.49 | 10.25 | 33.05 | 12.65 |
| Phos. & homologous | |||||||
| count (%) | 135,000 (85.90%) | 118,872 (69.93%) | 10,576 (6.02%) | 16,248 (16.97%) | 11,712 (14.40%) | 28,584 (69.80%) | 24,152 (19.64%) |
| avg. length | 603.56 | 623.26 | 824.95 | 814.92 | 702.19 | 561.02 | 554.76 |
| avg. weight (alton) | 67,246.82 | 69,484.93 | 91,920.53 | 90,881.20 | 78,902.92 | 63,482.53 | 61,901.72 |
| avg. nb of phos. Sites | 28.73 | 15.74 | 4.05 | 8.11 | 5.57 | 15.97 | 5.65 |
| avg. nb of phos. Sites/1000aa | 46.93 | 25.43 | 7.18 | 11.92 | 9.68 | 30.14 | 11.75 |
| Phos. & non-homologous | |||||||
| count (%) | 13,400 (8.53%) | 16,224 (9.54%) | 6360 (3.62%) | 20,800 (21.72%) | 11,096 (13.65%) | 7056 (17.23%) | 41,840 (34.02%) |
| avg. length | 453.47 | 537.58 | 992.41 | 756.98 | 752.89 | 536.44 | 535.52 |
| avg. weight (alton) | 50,430.62 | 60,000.12 | 110,195.46 | 83,939.00 | 84,333.71 | 60,406.56 | 59,709.85 |
| avg. nb of phos. Sites | 19.35 | 12.13 | 4.30 | 7.47 | 7.14 | 25.86 | 5.85 |
| avg. nb of phos. Sites/1000aa | 44.19 | 24.63 | 6.76 | 11.15 | 10.84 | 44.87 | 13.18 |
| Decile | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|---|
| ranges | [0, 2] | [3, 5] | [6, 8] | [9, 11] | [12, 14] | [15, 19] | [20, 26] | [27, 37] | [38, 61] | [62, 2518] |
| nb nodes | 2368 | 2130 | 1985 | 1818 | 1538 | 2042 | 1964 | 1892 | 1984 | 1924 |
| nb edges | 1455 | 1607 | 2021 | 2217 | 1499 | 2840 | 3819 | 4240 | 7688 | 9188 |
| avg. neighbors | 1.229 | 1.509 | 2.036 | 2.439 | 1.950 | 2.781 | 3.889 | 4.482 | 7.750 | 9.551 |
| nb connected | 120 | 135 | 89 | 83 | 80 | 76 | 52 | 39 | 33 | 22 |
| % singletons | 67.95 | 55.77 | 48.51 | 46.97 | 48.57 | 42.36 | 37.58 | 33.51 | 23.14 | 15.96 |
| Ranges | 0 | 1 | [2, 3] | [4, 8] | [9, 454] |
|---|---|---|---|---|---|
| nb nodes | 7338 | 1111 | 1273 | 1110 | 1137 |
| nb edges | 28,323 | 2232 | 2965 | 3156 | 4514 |
| avg. neighbors | 7.72 | 4.02 | 4.66 | 5.69 | 7.94 |
| nb connected | 123 | 38 | 36 | 15 | 17 |
| % singletons | 35.12 | 41.49 | 34.72 | 29.46 | 19.88 |
| Ranges | [0, 1] | [2, 5] | [6, 11] | [12, 23] | [24, 227] |
|---|---|---|---|---|---|
| nb nodes | 1020 | 1029 | 1072 | 987 | 1011 |
| nb edges | 3190 | 3628 | 7945 | 8366 | 5508 |
| avg. neighbors | 6.25 | 7.05 | 14.82 | 16.95 | 10.90 |
| nb connected | 29 | 29 | 17 | 16 | 8 |
| % singletons | 14.12 | 11.47 | 20.15 | 6.18 | 12.56 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pasquier, C.; Robichon, A. Evolutionary Divergence of Phosphorylation to Regulate Interactive Protein Networks in Lower and Higher Species. Int. J. Mol. Sci. 2022, 23, 14429. https://doi.org/10.3390/ijms232214429
Pasquier C, Robichon A. Evolutionary Divergence of Phosphorylation to Regulate Interactive Protein Networks in Lower and Higher Species. International Journal of Molecular Sciences. 2022; 23(22):14429. https://doi.org/10.3390/ijms232214429
Chicago/Turabian StylePasquier, Claude, and Alain Robichon. 2022. "Evolutionary Divergence of Phosphorylation to Regulate Interactive Protein Networks in Lower and Higher Species" International Journal of Molecular Sciences 23, no. 22: 14429. https://doi.org/10.3390/ijms232214429
APA StylePasquier, C., & Robichon, A. (2022). Evolutionary Divergence of Phosphorylation to Regulate Interactive Protein Networks in Lower and Higher Species. International Journal of Molecular Sciences, 23(22), 14429. https://doi.org/10.3390/ijms232214429






