Identification of the Major Effector StSROs in Potato: A Potential StWRKY-SRO6 Regulatory Pathway Enhances Plant Tolerance to Cadmium Stress
Abstract
1. Introduction
2. Results
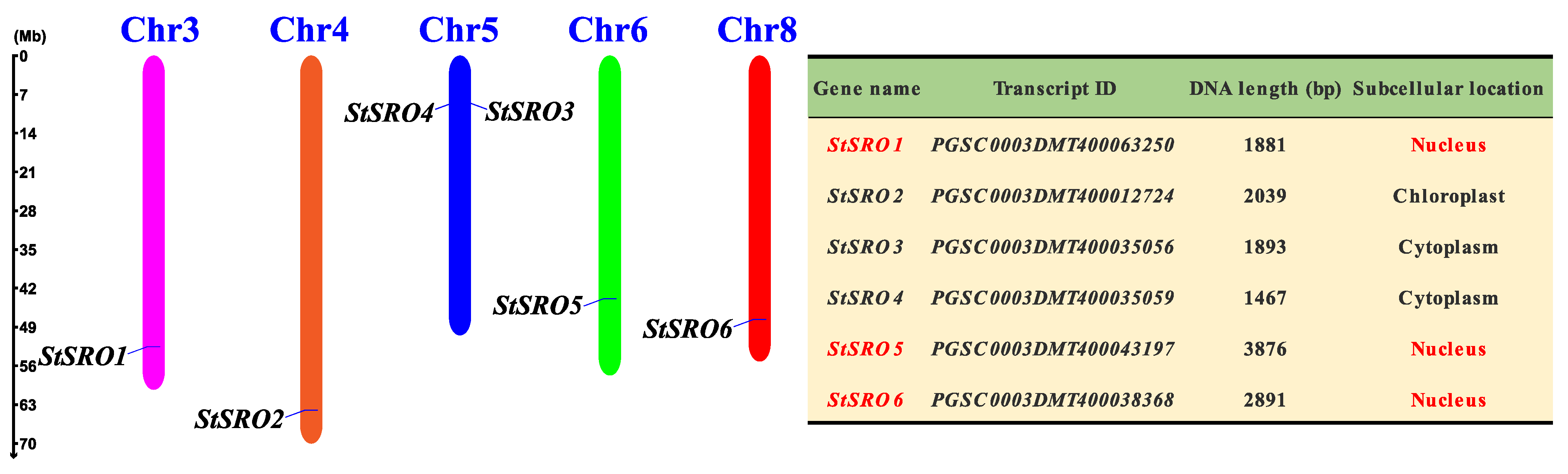
2.1. Genome-Wide Identification and Physicochemical Properties of SROs in Potato
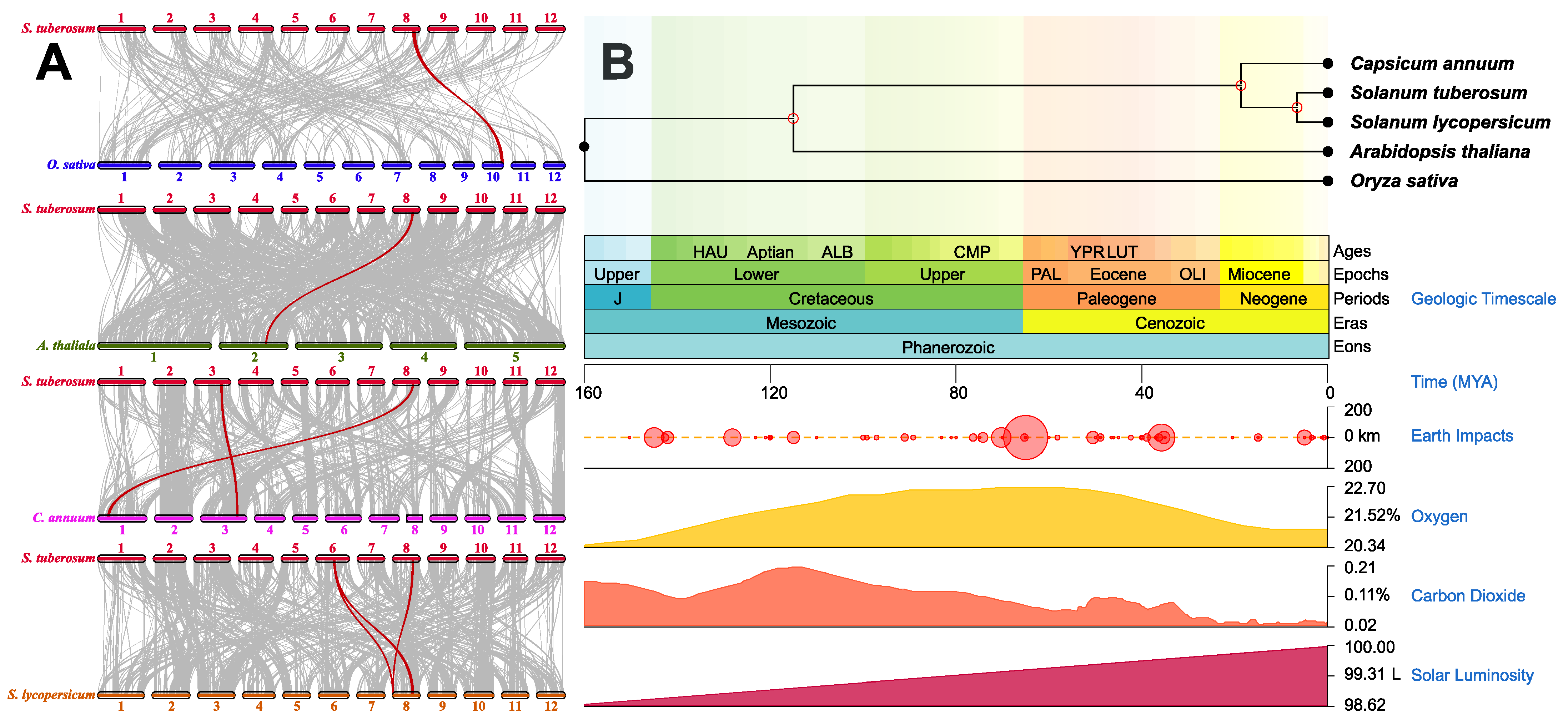
2.2. Evolutionary Exploration of SROs in Potato and Different Species
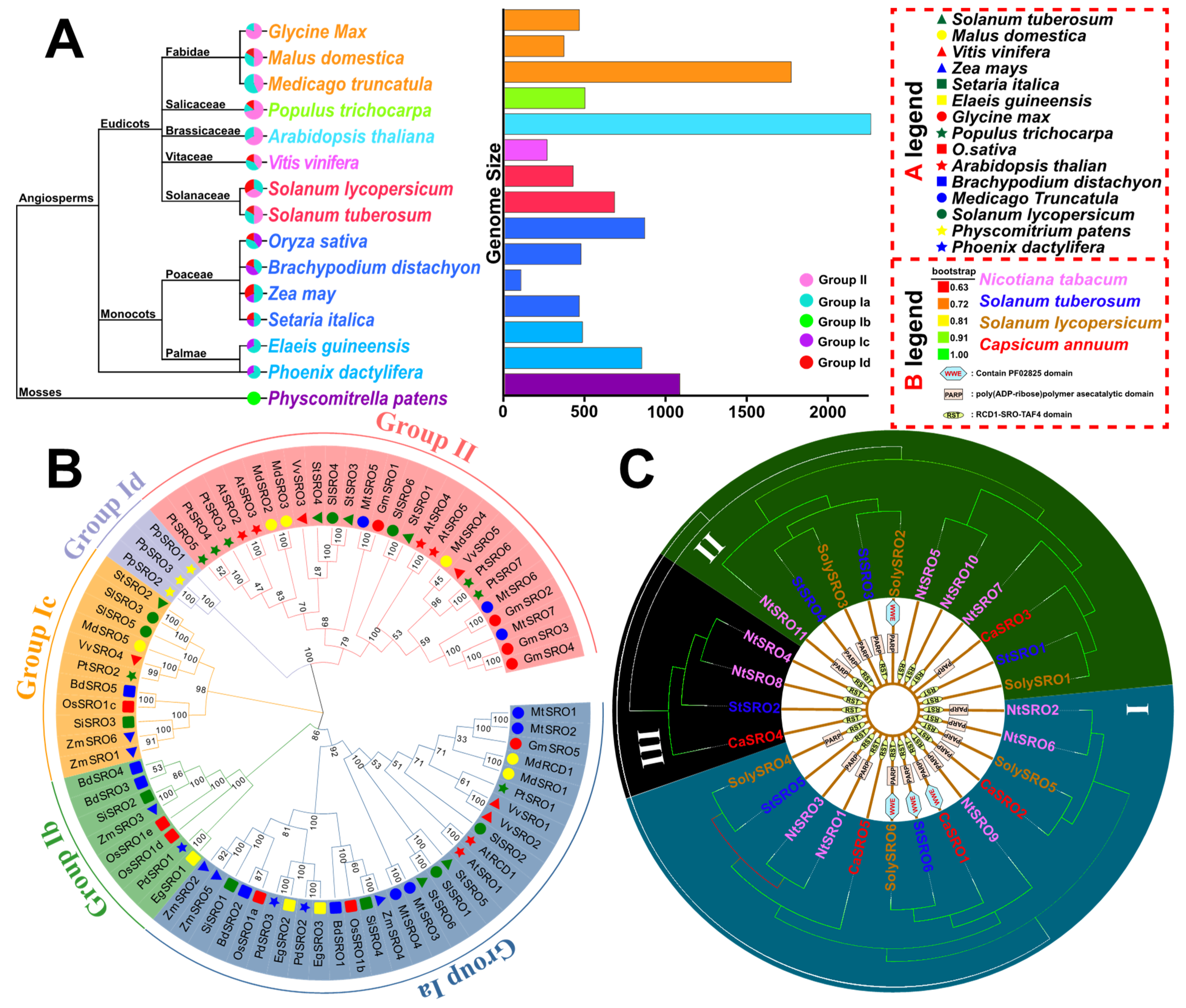
2.3. Phylogenetic Tree Analysis and Classification of SROs in Many Types of Plants
2.4. Gene Structures and Conserved Motif Analysis
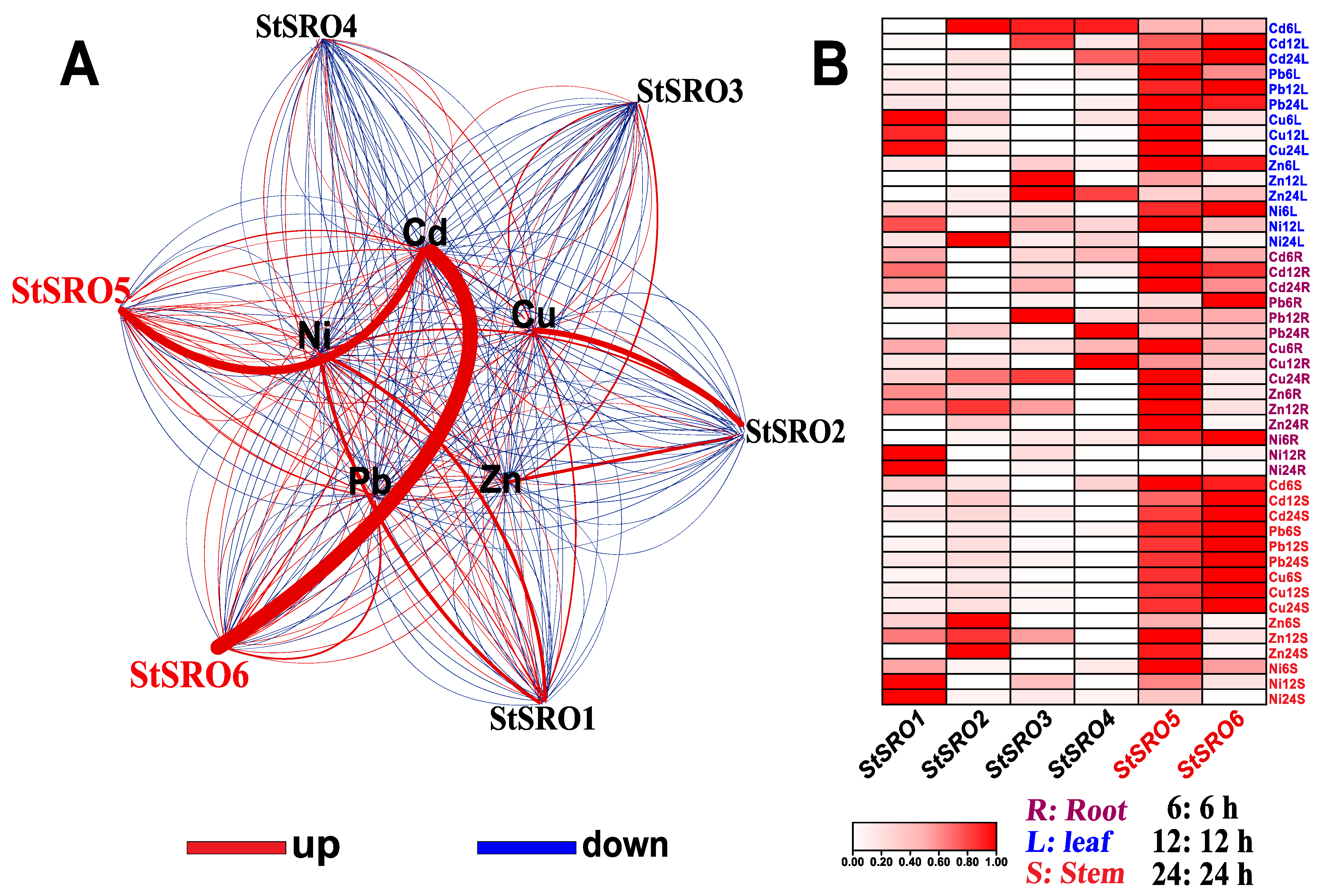
2.5. Expression Profile of SROs in Different Potato Tissues and Expression Response Analysis of Five Heavy Metals under Stress
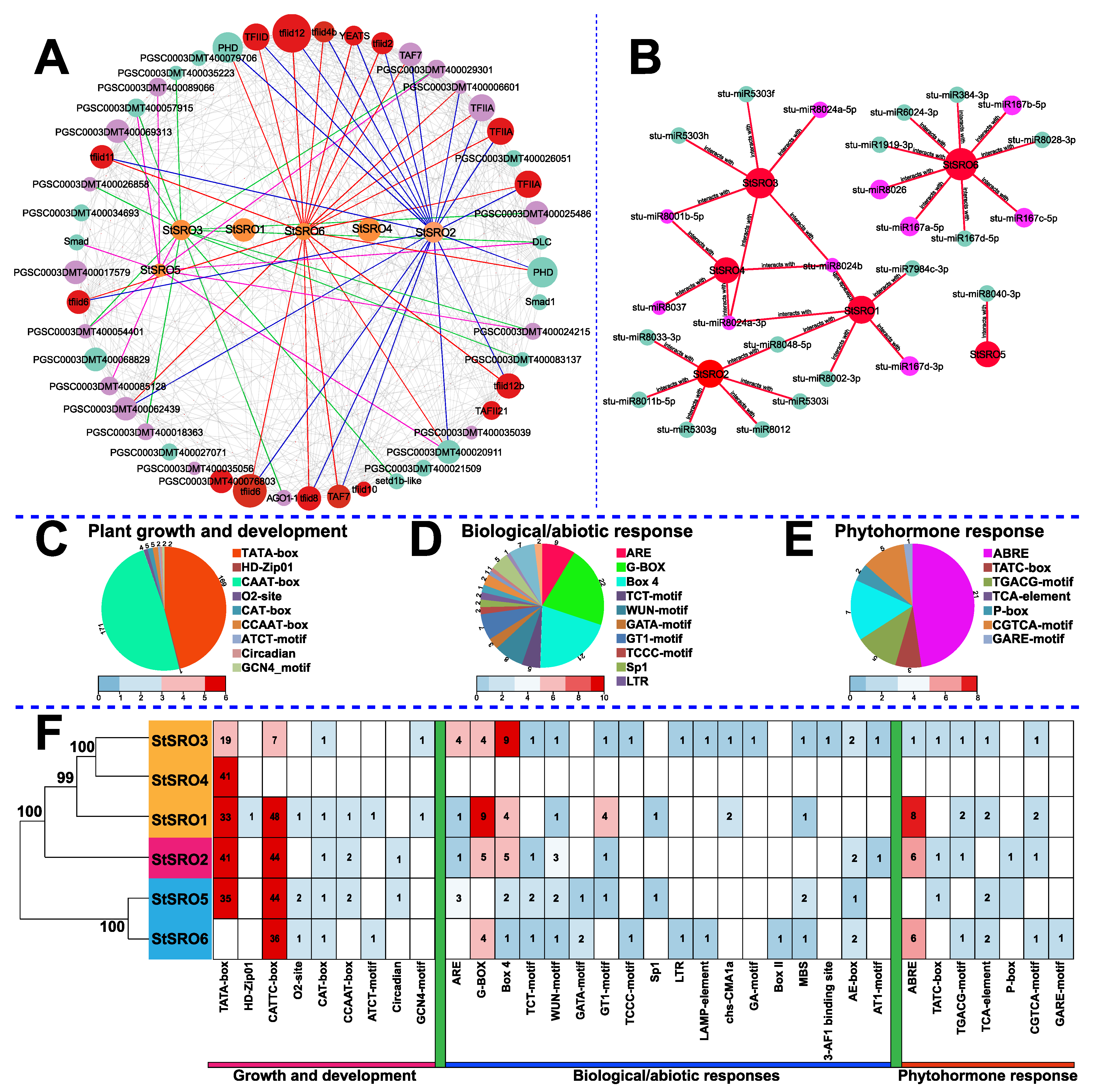
2.6. StSRO Co-Expression Framework
2.7. Prediction of Protein Folding Structure
2.8. StSRO Regulatory Network Analysis
2.9. Effect of Agrobacterium-Mediated Transient Expression of StWRKY6 on Transcription of StSROs 5/6
2.10. In Vitro Culture and Determination of Cd Tolerance of Potato Leaves
2.11. Verification of Cd Tolerance Function of the Heteroexpressed StSRO6 Gene in Yeast Cells
3. Discussion
3.1. Special StSROs 5/6 in the StSRO Family
3.2. OE-StWRKY6 Leads to Co-Expression of StSROs 5/6 to Enhance Potato Cd Tolerance
3.3. Potato MEG (StSRO6) Can Restore Cd Tolerance of Δycf1
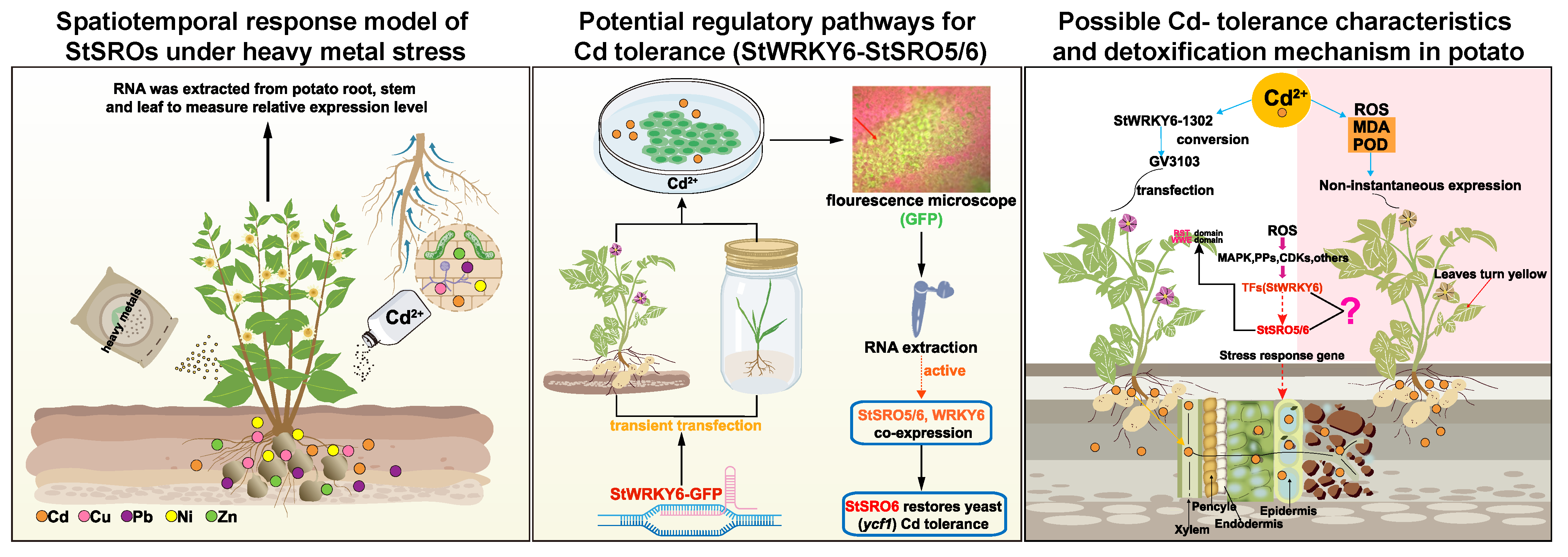
3.4. A possible New Pathway for Potato to Cope with Cd Stress
3.5. Conclusions
4. Materials and Methods
4.1. Toxicological Test and Treatment Method
4.2. Identification, Basic Characteristics and Interspecies Collinearity of SRO Family Members
4.3. Interspecies Phylogenetic and Evolutionary Analysis of SROs
4.4. The Functional Analysis of SRO Family Members was Carried out Based on Structures, Motifs, and Cis-Acting Elements
4.5. StSROs under/without Heavy Metal Stress and Co-Expression Network Analysis of Five Heavy Metals
4.6. StSRO Family Interaction Network Analysis and Proteins Tertiary Structure Prediction
4.7. Agrobacterium-Mediated Transient Expression in Potato
4.8. In Vitro Culture Test of Potato Leaf and Determination of Related Enzyme Activity
4.9. PI staining and Cd Localization Staining
4.10. StSRO6 Yeast Functional Complementarity Validation Test
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Shapiguzov, A.; Vainonen, J.P.; Hunter, K.; Tossavainen, H.; Tiwari, A.; Järvi, S.; Hellman, M.; Aarabi, F.; Alseekh, S.; Wybouw, B.; et al. Arabidopsis RCD1 coordinates chloroplast and mitochondrial functions through interaction with ANAC transcription factors. eLife 2019, 8, e43284. [Google Scholar] [CrossRef]
- Belles-Boix, E.; Babiychuk, E.; Van Montagu, M.; Inzé, D.; Kushnir, S. CEO1, a new protein from Arabidopsis thaliana, protects yeast against oxidative damage. FEBS Lett. 2000, 482, 19–24. [Google Scholar] [CrossRef]
- Overmyer, K.; Tuominen, H.; Kettunen, R.; Betz, C.; Langebartels, C.; Sandermann, H.J.; Kangasjärvi, J. Ozone-sensitive arabidopsis rcd1 mutant reveals opposite roles for ethylene and jasmonate signaling pathways in regulating superoxide-dependent cell death. Plant Cell 2000, 12, 1849–1862. [Google Scholar] [CrossRef] [PubMed]
- Jaspers, P.; Blomster, T.; Brosché, M.; Salojärvi, J.; Ahlfors, R.; Vainonen, J.P.; Reddy, R.A.; Immink, R.; Angenent, G.; Turck, F.; et al. Unequally redundant RCD1 and SRO1 mediate stress and developmental responses and interact with transcription factors. Plant J. 2009, 60, 268–279. [Google Scholar] [CrossRef] [PubMed]
- Teotia, S.; Lamb, R.S. The paralogous genes RADICAL-INDUCED CELL DEATH1 and SIMILAR TO RCD ONE1 have partially redundant functions during Arabidopsis development. Plant Physiol. 2009, 151, 180–198. [Google Scholar] [CrossRef]
- Briggs, G.C.; Osmont, K.S.; Shindo, C.; Sibout, R.; Hardtke, C.S. Unequal genetic redundancies in Arabidopsis--a neglected phenomenon? Trends Plant Sci. 2006, 11, 492–498. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhou, D.; Hu, H.; Li, W.; Hu, Y.; Xie, J.; Huang, S.; Wang, W. Genome-wide characterization of a SRO gene family involved in response to biotic and abiotic stresses in banana (Musa spp.). BMC Plant Biol. 2019, 19, 211. [Google Scholar] [CrossRef]
- Citarelli, M.; Teotia, S.; Lamb, R.S. Evolutionary history of the poly(ADP-ribose) polymerase gene family in eukaryotes. BMC Evol. Biol. 2010, 10, 308. [Google Scholar] [CrossRef]
- Hassa, P.O.; Haenni, S.S.; Elser, M.; Hottiger, M.O. Nuclear ADP-ribosylation reactions in mammalian cells: Where are we today and where are we going? Microbiol. Mol. Biol. Rev. 2006, 70, 789–829. [Google Scholar] [CrossRef]
- Otto, H.; Reche, P.A.; Bazan, F.; Dittmar, K.; Haag, F.; Koch-Nolte, F. In silico characterization of the family of PARP-like poly(ADP-ribosyl)transferases (pARTs). BMC Genom. 2005, 6, 139. [Google Scholar] [CrossRef]
- Jaspers, P.; Brosché, M.; Overmyer, K.; Kangasjärvi, J. The transcription factor interacting protein RCD1 contains a novel conserved domain. Plant Signal Behav. 2010, 5, 78–80. [Google Scholar] [CrossRef] [PubMed]
- He, F.; Tsuda, K.; Takahashi, M.; Kuwasako, K.; Terada, T.; Shirouzu, M.; Watanabe, S.; Kigawa, T.; Kobayashi, N.; Güntert, P.; et al. Structural insight into the interaction of ADP-ribose with the PARP WWE domains. FEBS Lett. 2012, 586, 3858–3864. [Google Scholar] [CrossRef]
- Zhao, X.; Gao, L.; Ren, J.; Pan, F. Arabidopsis SIMILAR TO RCD-ONE genes are ubiquitous and respond to multiple abiotic stresses through diverse signaling pathways. J. Biosci. 2019, 44, 129. [Google Scholar] [CrossRef]
- Liu, W.L.; Coleman, R.A.; Grob, P.; King, D.S.; Florens, L.; Washburn, M.P.; Geles, K.G.; Yang, J.L.; Ramey, V.; Nogales, E.; et al. Structural changes in TAF4b-TFIID correlate with promoter selectivity. Mol. Cell 2008, 29, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Malkowska, M.; Kokoszynska, K.; Rychlewski, L.; Wyrwicz, L. Structural bioinformatics of the general transcription factor TFIID. Biochimie 2013, 95, 680–691. [Google Scholar] [CrossRef] [PubMed]
- You, J.; Zong, W.; Du, H.; Hu, H.; Xiong, L. A special member of the rice SRO family, OsSRO1c, mediates responses to multiple abiotic stresses through interaction with various transcription factors. Plant Mol. Biol. 2014, 84, 693–705. [Google Scholar] [CrossRef]
- Jiang, W.; Geng, Y.; Liu, Y.; Chen, S.; Cao, S.; Li, W.; Chen, H.; Ma, D.; Yin, J. Genome-wide identification and characterization of SRO gene family in wheat: Molecular evolution and expression profiles during different stresses. Plant Physiol. Biochem. 2020, 154, 590–611. [Google Scholar] [CrossRef]
- Qiao, Y.; Gao, X.; Liu, Z.; Wu, Y.; Hu, L.; Yu, J. Genome-Wide Identification and Analysis of SRO Gene Family in Chinese Cabbage (Brassica rapa L.). Plants 2020, 9, 1235. [Google Scholar] [CrossRef]
- Li, D.; He, T.; Saleem, M.; He, G. Metalloprotein-Specific or Critical Amino Acid Residues: Perspectives on Plant-Precise Detoxification and Recognition Mechanisms under Cadmium Stress. Int. J. Mol. Sci. 2022, 23, 1734. [Google Scholar] [CrossRef]
- Gutaker, R.M.; Weiß, C.L.; Ellis, D.; Anglin, N.L.; Knapp, S.; Luis, F.J.; Prat, S.; Burbano, H.A. The origins and adaptation of European potatoes reconstructed from historical genomes. Nat. Ecol. Evol. 2019, 3, 1093–1101. [Google Scholar] [CrossRef]
- Marshall, C.R. Using the Fossil Record to Evaluate Timetree Timescales. Front. Genet. 2019, 10, 1049. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Xu, R.; Wang, B.; Wang, J.; Huang, S.; Yu, Q.; Gao, J. Genome-Wide Identification and Evolutionary Analysis of the SRO Gene Family in Tomato. Front. Genet. 2021, 12, 753638. [Google Scholar] [CrossRef]
- Pydiura, N.; Pirko, Y.; Galinousky, D.; Postovoitova, A.; Yemets, A.; Kilchevsky, A.; Blume, Y. Genome-wide identification, phylogenetic classification, and exon-intron structure characterization of the tubulin and actin genes in flax (Linum usitatissimum). Cell Biol. Int. 2019, 43, 1010–1019. [Google Scholar] [CrossRef]
- Safdari, Y.; Farajnia, S.; Asgharzadeh, M.; Khalili, M.; Jaliani, H.Z. Affinity measurement of single chain antibodies: A mathematical method facilitated by statistical software SigmaPlot. Monoclon. Antibodies Immunodiagn. Immunother. 2014, 33, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Schoch, C.L.; Ciufo, S.; Domrachev, M.; Hotton, C.L.; Kannan, S.; Khovanskaya, R.; Leipe, D.; Mcveigh, R.; O’Neill, K.; Robbertse, B.; et al. NCBI Taxonomy: A comprehensive update on curation, resources and tools. Database 2020, 2020, baaa062. [Google Scholar] [CrossRef] [PubMed]
- Tossavainen, H.; Hellman, M.; Vainonen, J.P.; Kangasjärvi, J.; Permi, P. (1)H, (13)C and (15)N NMR chemical shift assignments of A. thaliana RCD1 RST. Biomol. NMR Assign. 2017, 11, 207–210. [Google Scholar] [CrossRef] [PubMed]
- van Beek, L.; McClay, É.; Patel, S.; Schimpl, M.; Spagnolo, L.; Maia, D.O.T. PARP Power: A Structural Perspective on PARP1, PARP2, and PARP3 in DNA Damage Repair and Nucleosome Remodelling. Int. J. Mol. Sci. 2021, 22, 5112. [Google Scholar] [CrossRef]
- van Dam, S.; Võsa, U.; van der Graaf, A.; Franke, L.; de Magalhães, J.P. Gene co-expression analysis for functional classification and gene-disease predictions. Brief Bioinform. 2018, 19, 575–592. [Google Scholar] [CrossRef]
- Turoverov, K.K.; Kuznetsova, I.M.; Fonin, A.V.; Darling, A.L.; Zaslavsky, B.Y.; Uversky, V.N. Stochasticity of Biological Soft Matter: Emerging Concepts in Intrinsically Disordered Proteins and Biological Phase Separation. Trends Biochem. Sci. 2019, 44, 716–728. [Google Scholar] [CrossRef]
- Chen, L.; Heikkinen, L.; Wang, C.; Yang, Y.; Sun, H.; Wong, G. Trends in the development of miRNA bioinformatics tools. Brief Bioinform. 2019, 20, 1836–1852. [Google Scholar] [CrossRef]
- Perez-Cervantes, C.; Smith, L.A.; Nadadur, R.D.; Hughes, A.; Wang, S.; Corbo, J.C.; Cepko, C.; Lonfat, N.; Moskowitz, I.P. Enhancer transcription identifies cis-regulatory elements for photoreceptor cell types. Development 2020, 147, dev184432. [Google Scholar] [CrossRef] [PubMed]
- Zang, D.; Wang, C.; Ji, X.; Wang, Y. Tamarix hispida zinc finger protein ThZFP1 participates in salt and osmotic stress tolerance by increasing proline content and SOD and POD activities. Plant Sci. 2015, 235, 111–121. [Google Scholar] [CrossRef]
- Cui, X.; Zhu, P.; Li, B. Role of CEO1 gene in Cadmium Chloride-induced oxidative Stress in Arabidopsis thaliana. Plant Physiol. Commun. 2009, 45, 753–756. [Google Scholar]
- Li, X.; Xu, Y.; Liu, F.; Zhao, M.; Sun, Y.; Ma, Q. Maize similar to RCD1 gene induced by salt enhances Arabidopsis thaliana abiotic stress resistance. Biochem. Biophys. Res. Commun. 2018, 503, 2625–2632. [Google Scholar] [CrossRef] [PubMed]
- Teotia, S.; Lata, R.; Khare, S.K.; Gupta, M.N. One-step purification of glucoamylase by affinity precipitation with alginate. J. Mol. Recognit. 2001, 14, 295–299. [Google Scholar] [CrossRef]
- Yuan, B.; Chen, M.; Li, S. Isolation and Identification of Ipomoea cairica (L.) Sweet Gene IcSRO1 Encoding a SIMILAR TO RCD-ONE Protein, Which Improves Salt and Drought Tolerance in Transgenic Arabidopsis. Int. J. Mol. Sci. 2020, 21, 1017. [Google Scholar] [CrossRef]
- He, G.; Tian, W.; Qin, L.; Meng, L.; Wu, D.; Huang, Y.; Li, D.; Zhao, D.; He, T. Identification of novel heavy metal detoxification proteins in Solanum tuberosum: Insights to improve food security protection from metal ion stress. Sci. Total Environ. 2021, 779, 146197. [Google Scholar] [CrossRef]
- He, G.; Qin, L.; Tian, W.; Meng, L.; He, T.; Zhao, D. Heavy Metal Transporters-Associated Proteins in S. tuberosum: Genome-Wide Identification, Comprehensive Gene Feature, Evolution and Expression Analysis. Genes 2020, 11, 1269. [Google Scholar] [CrossRef]
- Wu, D.; He, G.; Tian, W.; Saleem, M.; Li, D.; Huang, Y.; Meng, L.; He, Y.; Liu, Y.; He, T. OPT gene family analysis of potato (Solanum tuberosum) responding to heavy metal stress: Comparative omics and co-expression networks revealed the underlying core templates and specific response patterns. Int. J. Biol. Macromol. 2021, 188, 892–903. [Google Scholar] [CrossRef]
- Tian, W.; He, G.; Qin, L.; Li, D.; Meng, L.; Huang, Y.; He, T. Genome-wide analysis of the NRAMP gene family in potato (Solanum tuberosum): Identification, expression analysis and response to five heavy metals stress. Ecotoxicol. Environ. Saf. 2021, 208, 111661. [Google Scholar] [CrossRef]
- Li, D.; He, G.; Tian, W.; Saleem, M.; Huang, Y.; Meng, L.; Wu, D.; He, T. Comparative and Systematic Omics Revealed Low Cd Accumulation of Potato StMTP9 in Yeast: Suggesting a New Mechanism for Heavy Metal Detoxification. Int. J. Mol. Sci. 2021, 22, 10478. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; He, G.; Tian, W.; Li, D.; Meng, L.; Wu, D.; He, T. Genome-Wide Identification of MATE Gene Family in Potato (Solanum tuberosum L.) and Expression Analysis in Heavy Metal Stress. Front. Genet. 2021, 12, 650500. [Google Scholar] [CrossRef] [PubMed]
- Ahlfors, R.; Lång, S.; Overmyer, K.; Jaspers, P.; Brosché, M.; Tauriainen, A.; Kollist, H.; Tuominen, H.; Belles-Boix, E.; Piippo, M.; et al. Arabidopsis RADICAL-INDUCED CELL DEATH1 belongs to the WWE protein-protein interaction domain protein family and modulates abscisic acid, ethylene, and methyl jasmonate responses. Plant Cell 2004, 16, 1925–1937. [Google Scholar] [CrossRef]
- Cunningham, F.; Allen, J.E.; Allen, J.; Alvarez-Jarreta, J.; Amode, M.R.; Armean, I.M.; Austine-Orimoloye, O.; Azov, A.G.; Barnes, I.; Bennett, R.; et al. Ensembl 2022. Nucleic Acids Res. 2022, 50, D988–D995. [Google Scholar] [CrossRef] [PubMed]
- Mistry, J.; Chuguransky, S.; Williams, L.; Qureshi, M.; Salazar, G.A.; Sonnhammer, E.; Tosatto, S.; Paladin, L.; Raj, S.; Richardson, L.J.; et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 2021, 49, D412–D419. [Google Scholar] [CrossRef]
- Potter, S.C.; Luciani, A.; Eddy, S.R.; Park, Y.; Lopez, R.; Finn, R.D. HMMER web server: 2018 update. Nucleic Acids Res. 2018, 46, W200–W204. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 2018, 46, D493–D496. [Google Scholar] [CrossRef]
- Yang, M.; Derbyshire, M.K.; Yamashita, R.A.; Marchler-Bauer, A. NCBI’s Conserved Domain Database and Tools for Protein Domain Analysis. Curr. Protoc. Bioinform. 2020, 69, e90. [Google Scholar] [CrossRef]
- Mohammad, N.S.; Nazli, R.; Zafar, H.; Fatima, S. Effects of lipid based Multiple Micronutrients Supplement on the birth outcome of underweight pre-eclamptic women: A randomized clinical trial. Pak. J. Med. Sci. 2022, 38, 219–226. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Subramanian, B.; Gao, S.; Lercher, M.J.; Hu, S.; Chen, W.H. Evolview v3: A webserver for visualization, annotation, and management of phylogenetic trees. Nucleic Acids Res. 2019, 47, W270–W275. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Nystrom, S.L.; McKay, D.J. Memes: A motif analysis environment in R using tools from the MEME Suite. PLoS Comput. Biol. 2021, 17, e1008991. [Google Scholar] [CrossRef] [PubMed]
- Sarkans, U.; Füllgrabe, A.; Ali, A.; Athar, A.; Behrangi, E.; Diaz, N.; Fexova, S.; George, N.; Iqbal, H.; Kurri, S.; et al. From ArrayExpress to BioStudies. Nucleic Acids Res. 2021, 49, D1502–D1506. [Google Scholar] [CrossRef]
- Otasek, D.; Morris, J.H.; Bouças, J.; Pico, A.R.; Demchak, B. Cytoscape Automation: Empowering workflow-based network analysis. Genome Biol. 2019, 20, 185. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING database in 2021: Customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef]
- Dai, X.; Zhuang, Z.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server (2017 release). Nucleic Acids Res. 2018, 46, W49–W54. [Google Scholar] [CrossRef]
- Amoozadeh, S.; Johnston, J.; Meisrimler, C.N. Exploiting Structural Modelling Tools to Explore Host-Translocated Effector Proteins. Int. J. Mol. Sci. 2021, 22, 12962. [Google Scholar] [CrossRef]
- Qu, J.; Yin, S.S.; Wang, H. Prediction of Metal Ion Binding Sites of Transmembrane Proteins. Comput. Math. Methods Med. 2021, 2021, 2327832. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Chen, T.; Wang, W.; Liu, H.; Yan, X.; Wu-Zhang, K.; Qin, W.; Xie, L.; Zhang, Y.; Peng, B.; et al. A high-efficiency Agrobacterium-mediated transient expression system in the leaves of Artemisia annua L. Plant Methods 2021, 17, 106. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, C.; Huang, Y.; Wang, C.; Xue, W.; Liu, Z. Effects of cadmium accumulation in vegetative organs of rice on Cadmium content in Rice. J. Agro-Environ. Sci. 2022, 41, 707–715. [Google Scholar]
- Xue, W.; Jiang, Y.; Shang, X.; Zou, J. Characterisation of early responses in lead accumulation and localization of Salix babylonica L. roots. BMC Plant Biol. 2020, 20, 296. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y. Molecular Physiological Basis of Cadmium Uptake and Transport in Steamed Bread; Chinese Academy of Forestry: Beijing, China, 2017; Available online: https://kns.cnki.net/KCMS/detail/detail.aspx?dbname=CDFDLAST2018&filename=1017270635 (accessed on 13 November 2022).


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, Y.; He, G.; Lou, F.; Zhou, Z.; Liu, Y.; Zhang, Y.; He, T. Identification of the Major Effector StSROs in Potato: A Potential StWRKY-SRO6 Regulatory Pathway Enhances Plant Tolerance to Cadmium Stress. Int. J. Mol. Sci. 2022, 23, 14318. https://doi.org/10.3390/ijms232214318
He Y, He G, Lou F, Zhou Z, Liu Y, Zhang Y, He T. Identification of the Major Effector StSROs in Potato: A Potential StWRKY-SRO6 Regulatory Pathway Enhances Plant Tolerance to Cadmium Stress. International Journal of Molecular Sciences. 2022; 23(22):14318. https://doi.org/10.3390/ijms232214318
Chicago/Turabian StyleHe, Yeqing, Guandi He, Fei Lou, Zheng Zhou, Yao Liu, Yule Zhang, and Tengbing He. 2022. "Identification of the Major Effector StSROs in Potato: A Potential StWRKY-SRO6 Regulatory Pathway Enhances Plant Tolerance to Cadmium Stress" International Journal of Molecular Sciences 23, no. 22: 14318. https://doi.org/10.3390/ijms232214318
APA StyleHe, Y., He, G., Lou, F., Zhou, Z., Liu, Y., Zhang, Y., & He, T. (2022). Identification of the Major Effector StSROs in Potato: A Potential StWRKY-SRO6 Regulatory Pathway Enhances Plant Tolerance to Cadmium Stress. International Journal of Molecular Sciences, 23(22), 14318. https://doi.org/10.3390/ijms232214318





