Allosteric Inhibition of Neutral Sphingomyelinase 2 (nSMase2) by DPTIP: From Antiflaviviral Activity to Deciphering Its Binding Site through In Silico Studies and Experimental Validation
Abstract
1. Introduction
2. Results and Discussion
2.1. Antiviral Activity of DPTIP
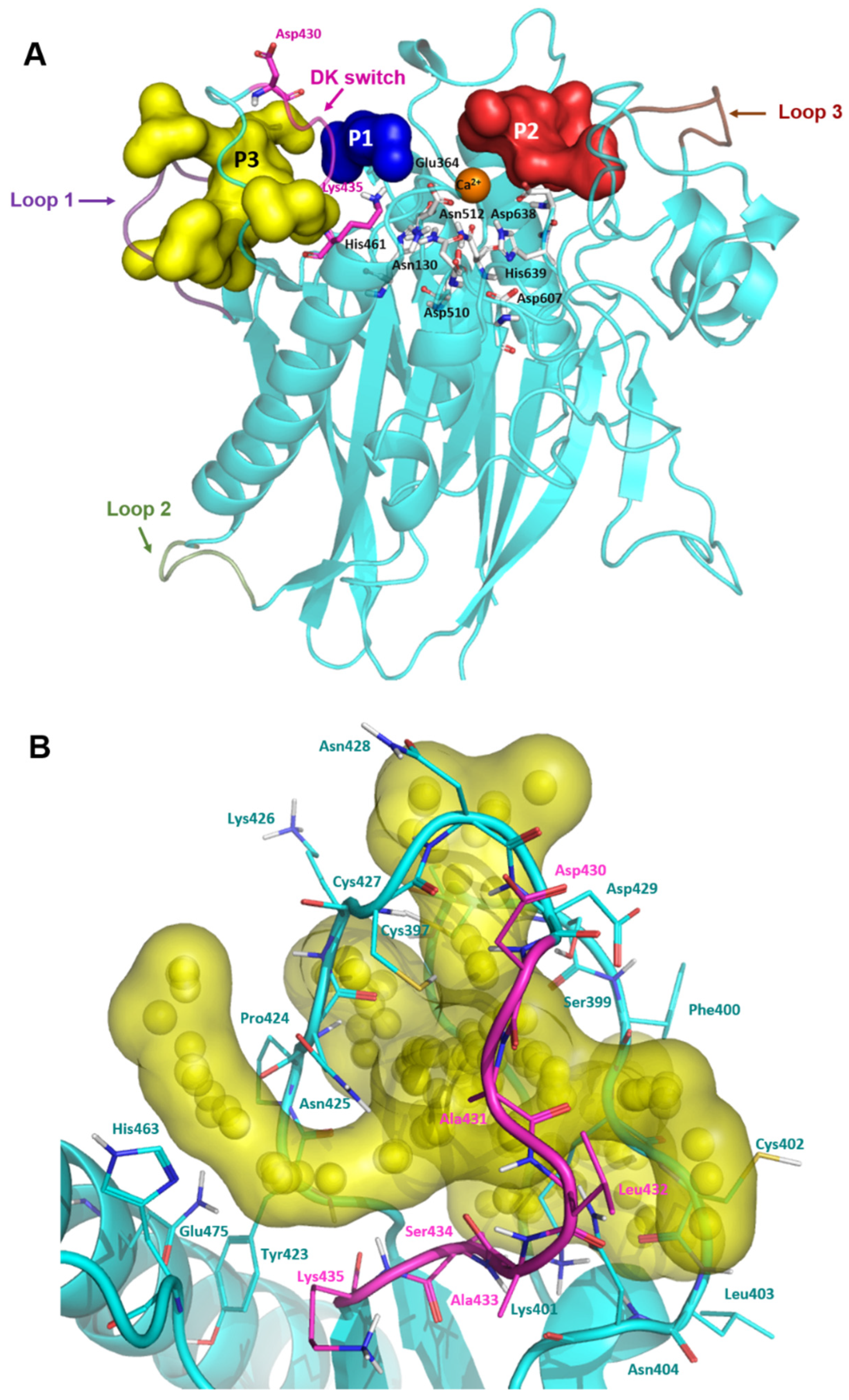
2.2. Modelling of the Missing Loops of nSMase2 Structure and Cavity Probing
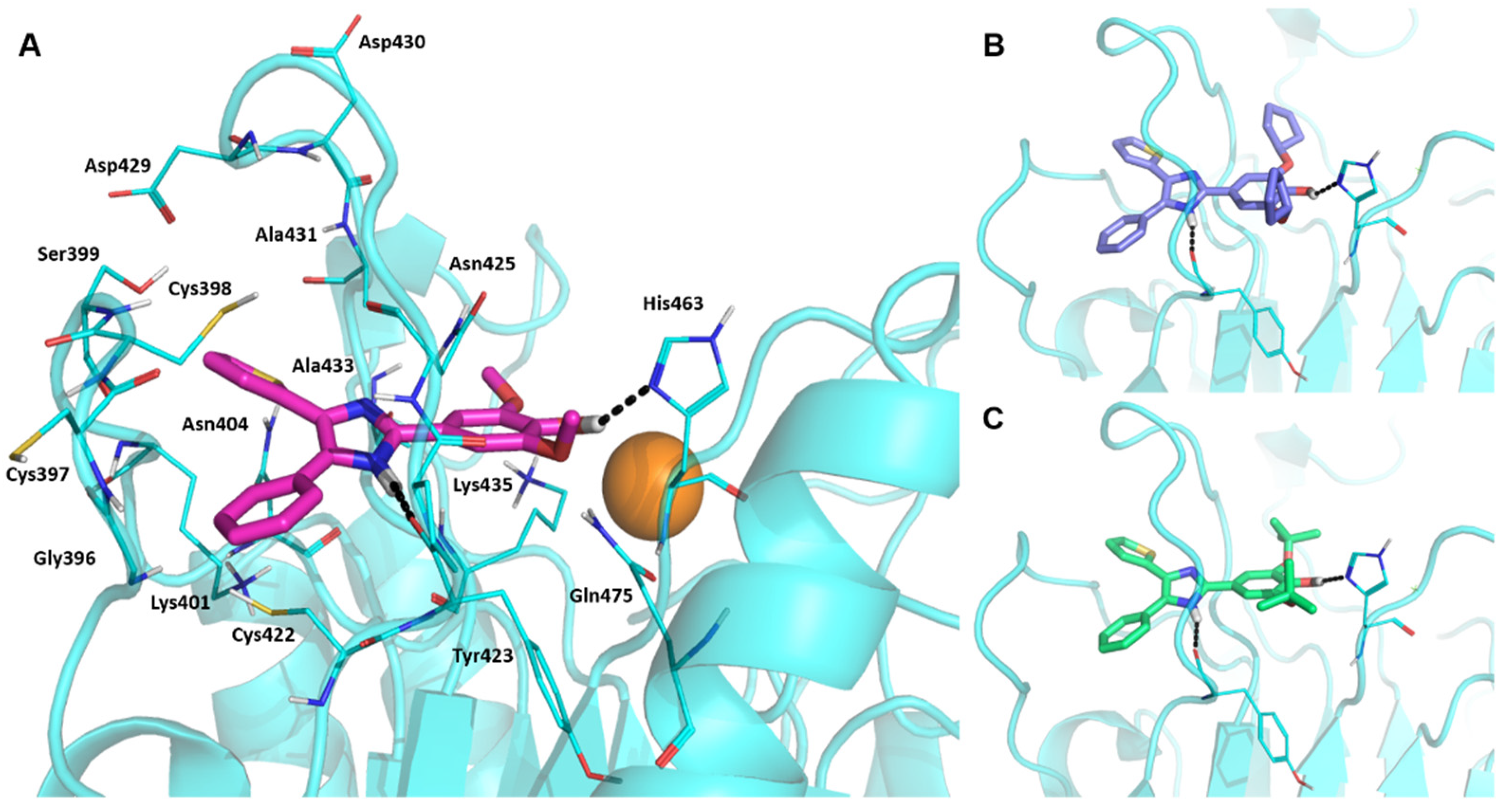
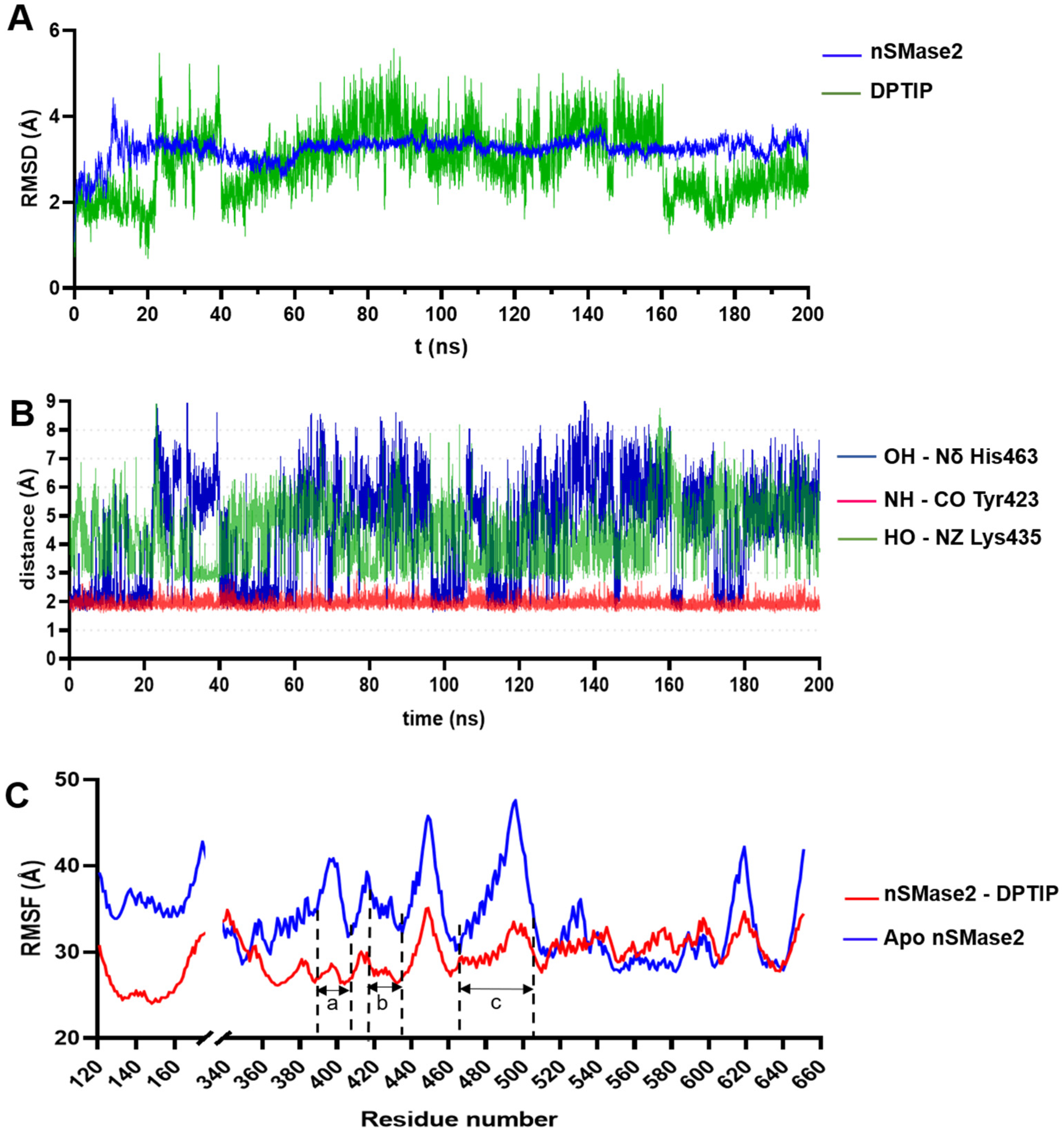
2.3. Binding Mode of DPTIP: Docking Studies and Molecular Dynamics Simulations
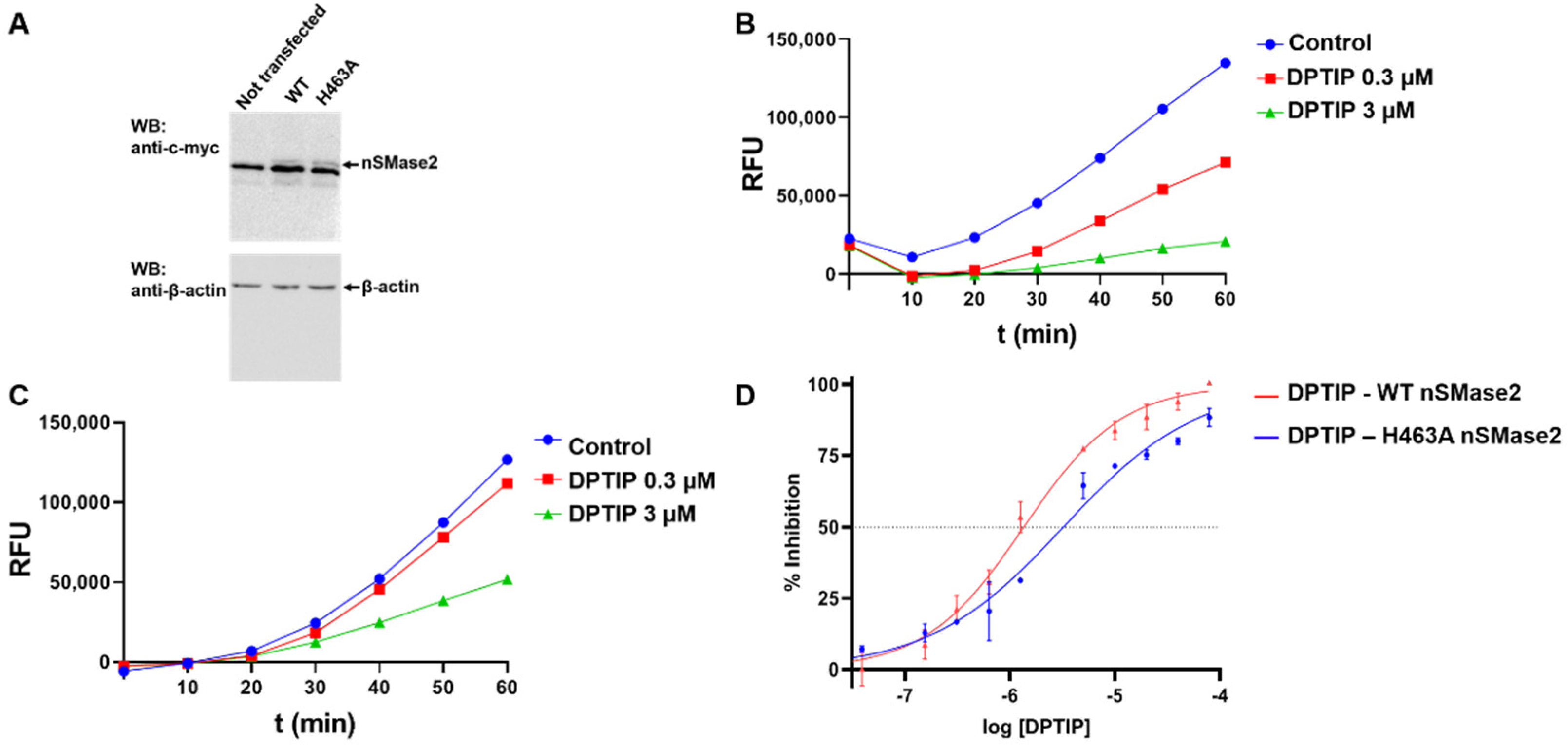
2.4. nSMase2 Inhibition Assay
3. Materials and Methods
3.1. DPTIP
3.2. Virus Infections and Drug Treatments
3.3. Production of Recombinant nSMase2
3.4. nSMase2 3D Model: Modelling, Docking and Molecular Dynamics Simulations
3.4.1. Protein Model
3.4.2. Binding Site Prediction
3.4.3. Docking
3.4.4. Molecular Dynamics Simulations
3.5. Fluorescence-Based nSMase2 Activity Assay
3.6. Data Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Pierson, T.C.; Diamond, M.S. The Continued Threat of Emerging Flaviviruses. Nat. Microbiol. 2020, 5, 796–812. [Google Scholar] [CrossRef]
- Gould, E.A.; Solomon, T. Pathogenic Flaviviruses. Lancet 2008, 371, 500–509. [Google Scholar] [CrossRef]
- Cao-Lormeau, V.-M.; Blake, A.; Mons, S.; Lastère, S.; Roche, C.; Vanhomwegen, J.; Dub, T.; Baudouin, L.; Teissier, A.; Larre, P.; et al. Guillain-Barré Syndrome Outbreak Associated with Zika Virus Infection in French Polynesia: A Case-Control Study. Lancet 2016, 387, 1531–1539. [Google Scholar] [CrossRef]
- Boldescu, V.; Behnam, M.A.M.; Vasilakis, N.; Klein, C.D. Broad-Spectrum Agents for Flaviviral Infections: Dengue, Zika and Beyond. Nat. Rev. Drug Discov. 2017, 16, 565–586. [Google Scholar] [CrossRef]
- Martín-Acebes, M.A.; Vázquez-Calvo, Á.; Saiz, J.C. Lipids and Flaviviruses, Present and Future Perspectives for the Control of Dengue, Zika, and West Nile Viruses. Prog. Lipid Res. 2016, 64, 123–137. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Pérez, M.J.; Saiz, J.C.; Priego, E.M.; Martín-Acebes, M.A. Antivirals against (Re)Emerging Flaviviruses: Should We Target the Virus or the Host? ACS Med. Chem. Lett. 2022, 13, 5–10. [Google Scholar] [CrossRef]
- Collenburg, L.; Beyersdorf, N.; Wiese, T.; Arenz, C.; Saied, E.M.; Becker-Flegler, K.A.; Schneider-Schaulies, S.; Avota, E. The Activity of the Neutral Sphingomyelinase Is Important in T Cell Recruitment and Directional Migration. Front. Immunol. 2017, 8, 1007. [Google Scholar] [CrossRef] [PubMed]
- Xiang, H.; Jin, S.; Tan, F.; Xu, Y.; Lu, Y.; Wu, T. Physiological Functions and Therapeutic Applications of Neutral Sphingomyelinase and Acid Sphingomyelinase. Biomed. Pharmacother. 2021, 139, 111610. [Google Scholar] [CrossRef]
- Shamseddine, A.A.; Airola, M.V.; Hannun, Y.A. Roles and Regulation of Neutral Sphingomyelinase-2 in Cellular and Pathological Processes. Adv. Biol. Regul. 2015, 57, 24–41. [Google Scholar] [CrossRef]
- Ong, W.-Y.; Herr, D.R.; Farooqui, T.; Ling, E.-A.; Farooqui, A.A. Role of Sphingomyelinases in Neurological Disorders. Expert Opin. Ther. Targets 2015, 19, 1725–1742. [Google Scholar] [CrossRef]
- Kalinichenko, L.S.; Mühle, C.; Jia, T.; Anderheiden, F.; Datz, M.; Eberle, A.-L.; Eulenburg, V.; Granzow, J.; Hofer, M.; Hohenschild, J.; et al. Adult Alcohol Drinking and Emotional Tone Are Mediated by Neutral Sphingomyelinase during Development in Males. Cereb. Cortex 2022, bhac106. [Google Scholar] [CrossRef]
- Martín-Acebes, M.A.; Merino-Ramos, T.; Blázquez, A.-B.; Casas, J.; Escribano-Romero, E.; Sobrino, F.; Saiz, J.-C. The Composition of West Nile Virus Lipid Envelope Unveils a Role of Sphingolipid Metabolism in Flavivirus Biogenesis. J. Virol. 2014, 88, 12041–12054. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Woodson, M.; Sherman, M.B.; Neelakanta, G.; Sultana, H. Exosomes Mediate Zika Virus Transmission through SMPD3 Neutral Sphingomyelinase in Cortical Neurons. Emerg. Microbes Infect. 2019, 8, 307–326. [Google Scholar] [CrossRef]
- Huang, Y.; Li, Y.; Zhang, H.; Zhao, R.; Jing, R.; Xu, Y.; He, M.; Peer, J.; Kim, Y.C.; Luo, J.; et al. Zika Virus Propagation and Release in Human Fetal Astrocytes Can Be Suppressed by Neutral Sphingomyelinase-2 Inhibitor GW4869. Cell Discov. 2018, 4, 1–16. [Google Scholar] [CrossRef]
- Shanbhogue, P.; Hannun, Y.A. Exploring the Therapeutic Landscape of Sphingomyelinases BT—Lipid Signaling in Human Diseases; Gomez-Cambronero, J., Frohman, M.A., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 19–47. ISBN 978-3-030-33668-4. [Google Scholar]
- Airola, M.V.; Shanbhogue, P.; Shamseddine, A.A.; Guja, K.E.; Senkal, C.E.; Maini, R.; Bartke, N.; Wu, B.X.; Obeid, L.M.; Garcia-Diaz, M.; et al. Structure of Human NSMase2 Reveals an Interdomain Allosteric Activation Mechanism for Ceramide Generation. Proc. Natl. Acad. Sci. USA 2017, 114, E5549–E5558. [Google Scholar] [CrossRef]
- Shanbhogue, P.; Hoffmann, R.M.; Airola, M.V.; Maini, R.; Hamelin, D.J.; Garcia-Diaz, M.; Burke, J.E.; Hannun, Y.A. The Juxtamembrane Linker in Neutral Sphingomyelinase-2 Functions as an Intramolecular Allosteric Switch That Activates the Enzyme. J. Biol. Chem. 2019, 294, 7488–7502. [Google Scholar] [CrossRef]
- Skácel, J.; Slusher, B.S.; Tsukamoto, T. Small Molecule Inhibitors Targeting Biosynthesis of Ceramide, the Central Hub of the Sphingolipid Network. J. Med. Chem. 2021, 64, 279–297. [Google Scholar] [CrossRef]
- Uchida, R.; Tomoda, H.; Dong, Y.; Omura, S. Altenusin, a Specific Neutral Sphingomyelinase Inhibitor, Produced by Penidltium Sp. FO-7436. J. Antibiot. 1999, 52, 572–574. [Google Scholar] [CrossRef]
- Kumar, A.; Henry, R.J.; Stoica, B.A.; Loane, D.J.; Abulwerdi, G.; Bhat, S.A.; Faden, A.I. Neutral Sphingomyelinase Inhibition Alleviates LPS-Induced Microglia Activation and Neuroinflammation after Experimental Traumatic Brain Injury. J. Pharmacol. Exp. Ther. 2019, 368, 338–352. [Google Scholar] [CrossRef]
- Luberto, C.; Hassler, D.F.; Signorelli, P.; Okamoto, Y.; Sawai, H.; Boros, E.; Hazen-Martin, D.J.; Obeid, L.M.; Hannun, Y.A.; Smith, G.K. Inhibition of Tumor Necrosis Factor-Induced Cell Death in MCF7 by a Novel Inhibitor of Neutral Sphingomyelinase. J. Biol. Chem. 2002, 277, 41128–41139. [Google Scholar] [CrossRef]
- Figuera-Losada, M.; Stathis, M.; Dorskind, J.M.; Thomas, A.G.; Ratnam Bandaru, V.V.; Yoo, S.W.; Westwood, N.J.; Rogers, G.W.; McArthur, J.C.; Haughey, N.J.; et al. Cambinol, a Novel Inhibitor of Neutral Sphingomyelinase 2 Shows Neuroprotective Properties. PLoS ONE 2015, 10, e0124481. [Google Scholar] [CrossRef]
- Bilousova, T.; Elias, C.; Miyoshi, E.; Alam, M.P.; Zhu, C.; Campagna, J.; Vadivel, K.; Jagodzinska, B.; Gylys, K.H.; John, V. Suppression of Tau Propagation Using an Inhibitor That Targets the DK-Switch of NSMase2. Biochem. Biophys. Res. Commun. 2018, 499, 751–757. [Google Scholar] [CrossRef] [PubMed]
- Rojas, C.; Sala, M.; Thomas, A.G.; Datta Chaudhuri, A.; Yoo, S.W.; Li, Z.; Dash, R.P.; Rais, R.; Haughey, N.J.; Nencka, R.; et al. A Novel and Potent Brain Penetrant Inhibitor of Extracellular Vesicle Release. Br. J. Pharmacol. 2019, 176, 3857–3870. [Google Scholar] [CrossRef] [PubMed]
- Bilousova, T.; Simmons, B.J.; Knapp, R.R.; Elias, C.J.; Campagna, J.; Melnik, M.; Chandra, S.; Focht, S.; Zhu, C.; Vadivel, K.; et al. Dual Neutral Sphingomyelinase-2/Acetylcholinesterase Inhibitors for the Treatment of Alzheimer’ s Disease. ACS Chem. Biol. 2020, 15, 1671–1684. [Google Scholar] [CrossRef]
- Rojas, C.; Barnaeva, E.; Thomas, A.G.; Hu, X.; Southall, N.; Marugan, J.; Chaudhuri, A.D.; Yoo, S.-W.; Hin, N.; Stepanek, O.; et al. DPTIP, a Newly Identified Potent Brain Penetrant Neutral Sphingomyelinase 2 Inhibitor, Regulates Astrocyte-Peripheral Immune Communication Following Brain Inflammation. Sci. Rep. 2018, 8, 17715. [Google Scholar] [CrossRef] [PubMed]
- Stepanek, O.; Hin, N.; Thomas, A.G.; Dash, R.P.; Alt, J.; Rais, R.; Rojas, C.; Slusher, B.S.; Tsukamoto, T. Neutral Sphingomyelinase 2 Inhibitors Based on the 4-(1H-Imidazol-2-Yl)-2,6-Dialkoxyphenol Scaffold. Eur. J. Med. Chem. 2019, 170, 276–289. [Google Scholar] [CrossRef] [PubMed]
- Fiser, A.; Sali, A. ModLoop: Automated Modeling of Loops in Protein Structures. Bioinformatics 2003, 19, 2500–2501. [Google Scholar] [CrossRef] [PubMed]
- Jacobson, M.P.; Pincus, D.L.; Rapp, C.S.; Day, T.J.F.; Honig, B.; Shaw, D.E.; Friesner, R.A. A Hierarchical Approach to All-Atom Protein Loop Prediction. Proteins Struct. Funct. Genet. 2004, 55, 351–367. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R.A.; MacArthur, M.W.; Moss, D.S.; Thornton, J.M. PROCHECK: A Program to Check the Stereochemical Quality of Protein Structures. J. Appl. Crystallogr. 1993, 26, 283–291. [Google Scholar] [CrossRef]
- Le Guilloux, V.; Schmidtke, P.; Tuffery, P. Fpocket: An Open Source Platform for Ligand Pocket Detection. BMC Bioinform. 2009, 10, 168. [Google Scholar] [CrossRef] [PubMed]
- Schmidtke, P.; Barril, X. Understanding and Predicting Druggability. A High-Throughput Method for Detection of Drug Binding Sites. J. Med. Chem. 2010, 53, 5858–5867. [Google Scholar] [CrossRef] [PubMed]
- Morris, G.M.; Ruth, H.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. Software News and Updates AutoDock4 and AutoDockTools4: Automated Docking with Selective Receptor Flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed]
- Case, D.A.; Betz, R.M.; Cerutti, D.S.; Cheatham, T.E., III; Darden, T.A.; Duke, R.E.; Giese, T.J.; Gohlke, H.; Goetz, A.W.; Homeyer, N.; et al. AMBER 2016; University of California: San Francisco, CA, USA, 2016. [Google Scholar]
- Klett, J.; Núñez-Salgado, A.; Dos Santos, H.G.; Cortés-Cabrera, Á.; Perona, A.; Gil-Redondo, R.; Abia, D.; Gago, F.; Morreale, A. MM-ISMSA: An Ultrafast and Accurate Scoring Function for Protein-Protein Docking. J. Chem. Theory Comput. 2012, 8, 3395–3408. [Google Scholar] [CrossRef]
- Morreale, A.; Gil-Redondo, R.; Ortiz, Á.R. A New Implicit Solvent Model for Protein–Ligand Docking. Proteins Struct. Funct. Bioinform. 2007, 67, 606–616. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Calvo, Á.; Blázquez, A.B.; Escribano-Romero, E.; Merino-Ramos, T.; Saiz, J.C.; Martín-Acebes, M.A.; Jiménez de Oya, N. Zika Virus Infection Confers Protection against West Nile Virus Challenge in Mice: Zika Virus Protects against West Nile Virus. Emerg. Microbes Infect. 2017, 6, e81. [Google Scholar] [CrossRef]
- Martín-Acebes, M.A.; Saiz, J.C. A West Nile Virus Mutant with Increased Resistance to Acid-Induced Inactivation. J. Gen. Virol. 2011, 92, 831–840. [Google Scholar] [CrossRef]
- Sáez-Álvarez, Y.; De Oya, N.J.; Del Águila, C.; Saiz, J.C.; Arias, A.; Agudo, R.; Martín-Acebes, M.A. Novel Nonnucleoside Inhibitors of Zika Virus Polymerase Identified through the Screening of an Open Library of Antikinetoplastid Compounds. Antimicrob. Agents Chemother. 2021, 65, AAC0089421. [Google Scholar] [CrossRef]
- Martín-Acebes, M.A.; Blázquez, A.B.; Jiménez de Oya, N.; Escribano-Romero, E.; Saiz, J.C. West Nile Virus Replication Requires Fatty Acid Synthesis but Is Independent on Phosphatidylinositol-4-Phosphate Lipids. PLoS ONE 2011, 6, e24970. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Jacobson, M.P.; Friesner, R.A.; Xiang, Z.; Honig, B. On the Role of the Crystal Environment in Determining Protein Side-Chain Conformations. J. Mol. Biol. 2002, 320, 597–608. [Google Scholar] [CrossRef]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. Ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from Ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [PubMed]
- Mark, P.; Nilsson, L. Structure and Dynamics of the TIP3P, SPC, and SPC/E Water Models at 298 K. J. Phys. Chem. A 2001, 105, 9954–9960. [Google Scholar] [CrossRef]
- Vassetti, D.; Pagliai, M.; Procacci, P. Assessment of GAFF2 and OPLS-AA General Force Fields in Combination with the Water Models TIP3P, SPCE, and OPC3 for the Solvation Free Energy of Druglike Organic Molecules. J. Chem. Theory Comput. 2019, 15, 1983–1995. [Google Scholar] [CrossRef] [PubMed]
- Aqvist, J. Ion Water Interaction Potential-Derived from Free Energy Perturbation Simulations. J. Phys. Chem. 1990, 94, 8021–8024. [Google Scholar] [CrossRef]
- Darden, T.; York, D.; Pedersen, L. Particle Mesh Ewald—An n.Log(n) Method for Ewald Sums in Large Systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Ryckaert, J.P.; Ciccotti, G.; Berendsen, H.J.C. Numerical-Integration of Cartesian Equations of Motion of a System with Constraints—Molecular Dynamics of N-Alkanes. J. Comput. Phys. 1977, 23, 327–341. [Google Scholar] [CrossRef]
- Schrödinger, L.; DeLano, W. Pymol 2020. Available online: https://www.pymol.org/pymol.html (accessed on 20 October 2022).
- Roe, D.R.; Cheatham, T.E. PTRAJ and CPPTRAJ: Software for Processing and Analysis of Molecular Dynamics Trajectory Data. J. Chem. Theory Comput. 2013, 9, 3084–3095. [Google Scholar] [CrossRef]




| Pocket. | Volume | Druggability Score | Hydrophobicity Score | Location |
|---|---|---|---|---|
| 1 | 321 | 0.104 | 40 | Adjacent to active site and to DK switch |
| 2 | 1000 | 0.168 | 29 | Adjacent to active site |
| 3 | 1618 | 0.820 | 22 | Adjacent to active site. Includes DK switch |
| Compound | Structure | IC50 (µM) |
|---|---|---|
| DPTIP (6) | 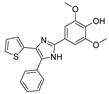 | 0.03 a |
| 7 |  | >100 a |
| 8 |  | >100 a |
| 9 |  | 40 a |
| 10 |  | 0.12 a |
| 11 | 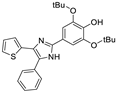 | 0.52 a |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Álvarez-Fernández, H.; Mingo-Casas, P.; Blázquez, A.-B.; Caridi, F.; Saiz, J.C.; Pérez-Pérez, M.-J.; Martín-Acebes, M.A.; Priego, E.-M. Allosteric Inhibition of Neutral Sphingomyelinase 2 (nSMase2) by DPTIP: From Antiflaviviral Activity to Deciphering Its Binding Site through In Silico Studies and Experimental Validation. Int. J. Mol. Sci. 2022, 23, 13935. https://doi.org/10.3390/ijms232213935
Álvarez-Fernández H, Mingo-Casas P, Blázquez A-B, Caridi F, Saiz JC, Pérez-Pérez M-J, Martín-Acebes MA, Priego E-M. Allosteric Inhibition of Neutral Sphingomyelinase 2 (nSMase2) by DPTIP: From Antiflaviviral Activity to Deciphering Its Binding Site through In Silico Studies and Experimental Validation. International Journal of Molecular Sciences. 2022; 23(22):13935. https://doi.org/10.3390/ijms232213935
Chicago/Turabian StyleÁlvarez-Fernández, Hadrián, Patricia Mingo-Casas, Ana-Belén Blázquez, Flavia Caridi, Juan Carlos Saiz, María-Jesús Pérez-Pérez, Miguel A. Martín-Acebes, and Eva-María Priego. 2022. "Allosteric Inhibition of Neutral Sphingomyelinase 2 (nSMase2) by DPTIP: From Antiflaviviral Activity to Deciphering Its Binding Site through In Silico Studies and Experimental Validation" International Journal of Molecular Sciences 23, no. 22: 13935. https://doi.org/10.3390/ijms232213935
APA StyleÁlvarez-Fernández, H., Mingo-Casas, P., Blázquez, A.-B., Caridi, F., Saiz, J. C., Pérez-Pérez, M.-J., Martín-Acebes, M. A., & Priego, E.-M. (2022). Allosteric Inhibition of Neutral Sphingomyelinase 2 (nSMase2) by DPTIP: From Antiflaviviral Activity to Deciphering Its Binding Site through In Silico Studies and Experimental Validation. International Journal of Molecular Sciences, 23(22), 13935. https://doi.org/10.3390/ijms232213935









