Tolerance, Adaptation, and Cell Response Elicited by Micromonospora sp. Facing Tellurite Toxicity: A Biological and Physical-Chemical Characterization
Abstract
1. Introduction
2. Results
2.1. Micromonospora Exponentially Grown Cells Facing Tellurite
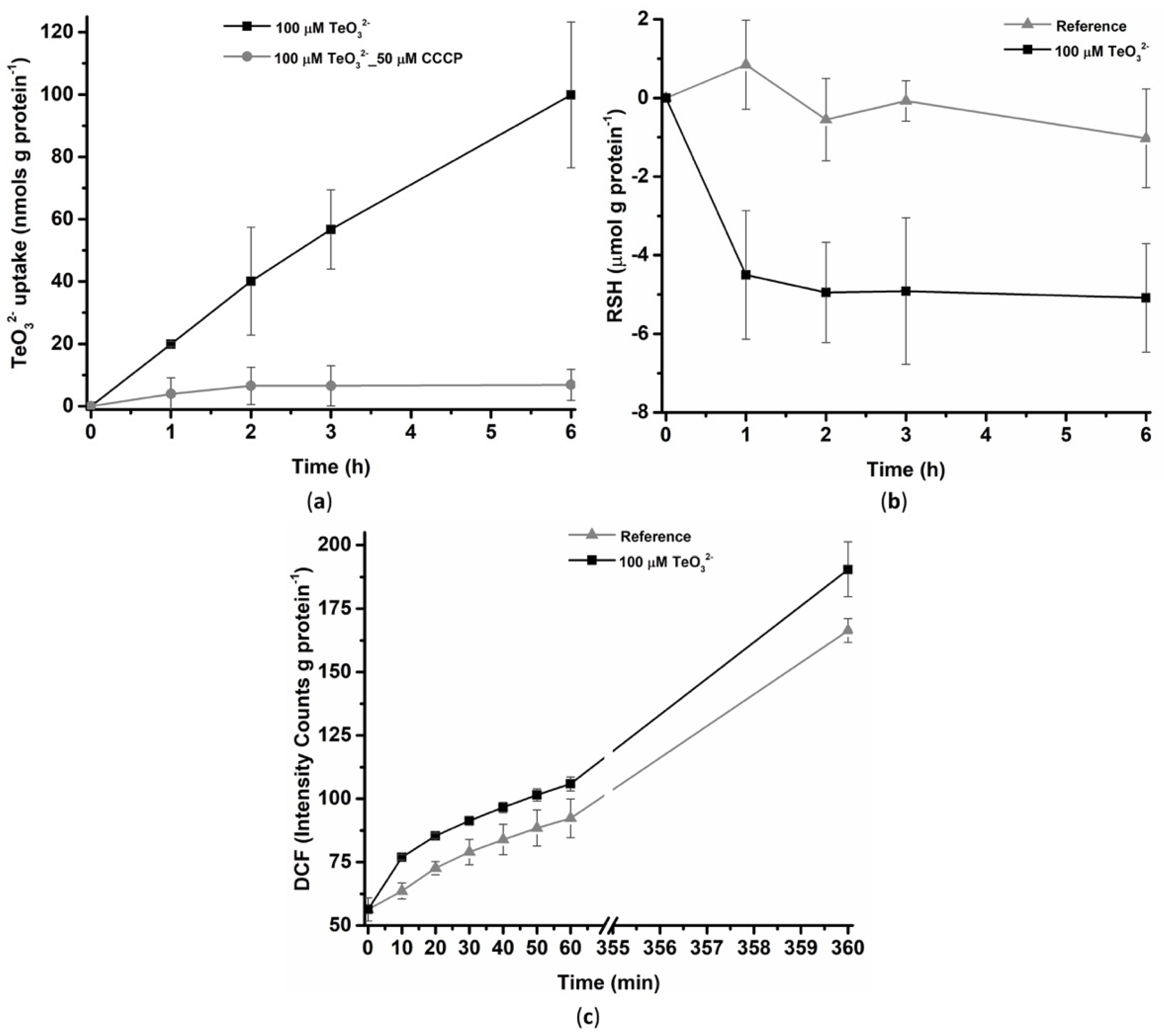
2.1.1. Tellurite Uptake, Thiol Oxidation, and Reactive Oxygen Species Production
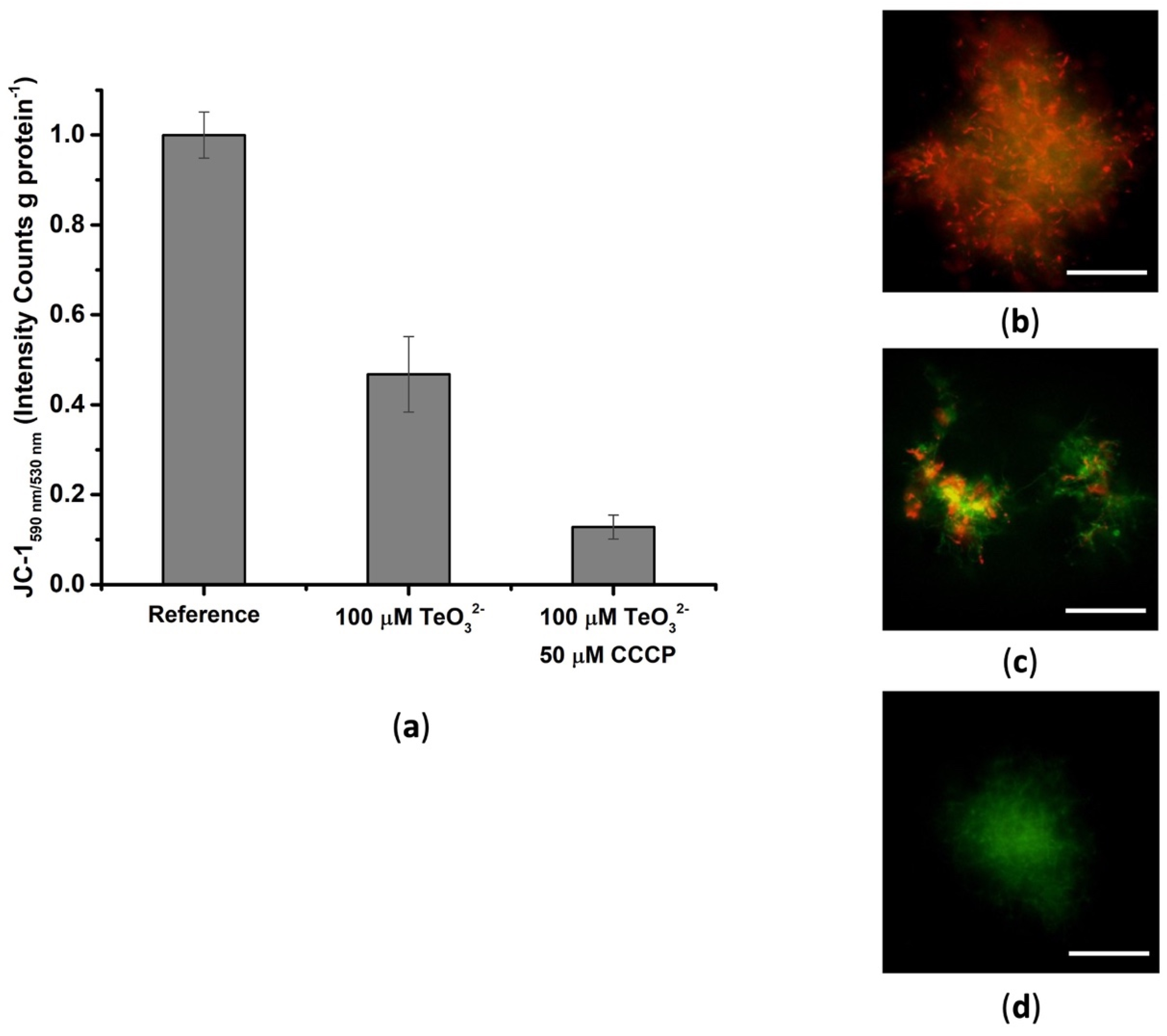
2.1.2. Effect of Tellurite on the Membrane Potential
2.1.3. Tellurite Effect on the Fatty Acid Profile
2.2. Micromonospora Cells Growing in the Presence of Different Tellurite Concentrations
2.2.1. Bacterial Tolerance towards TeO32−
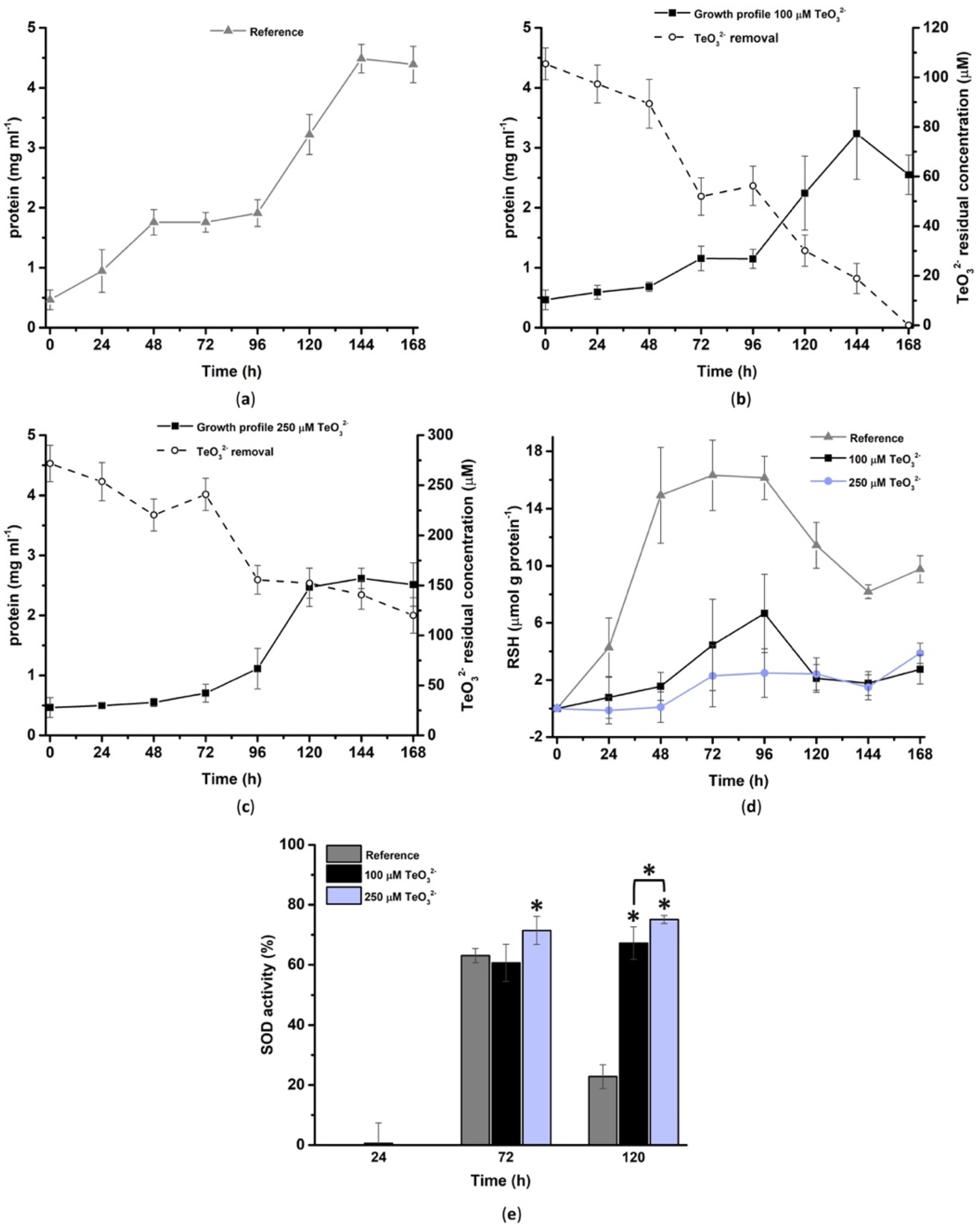

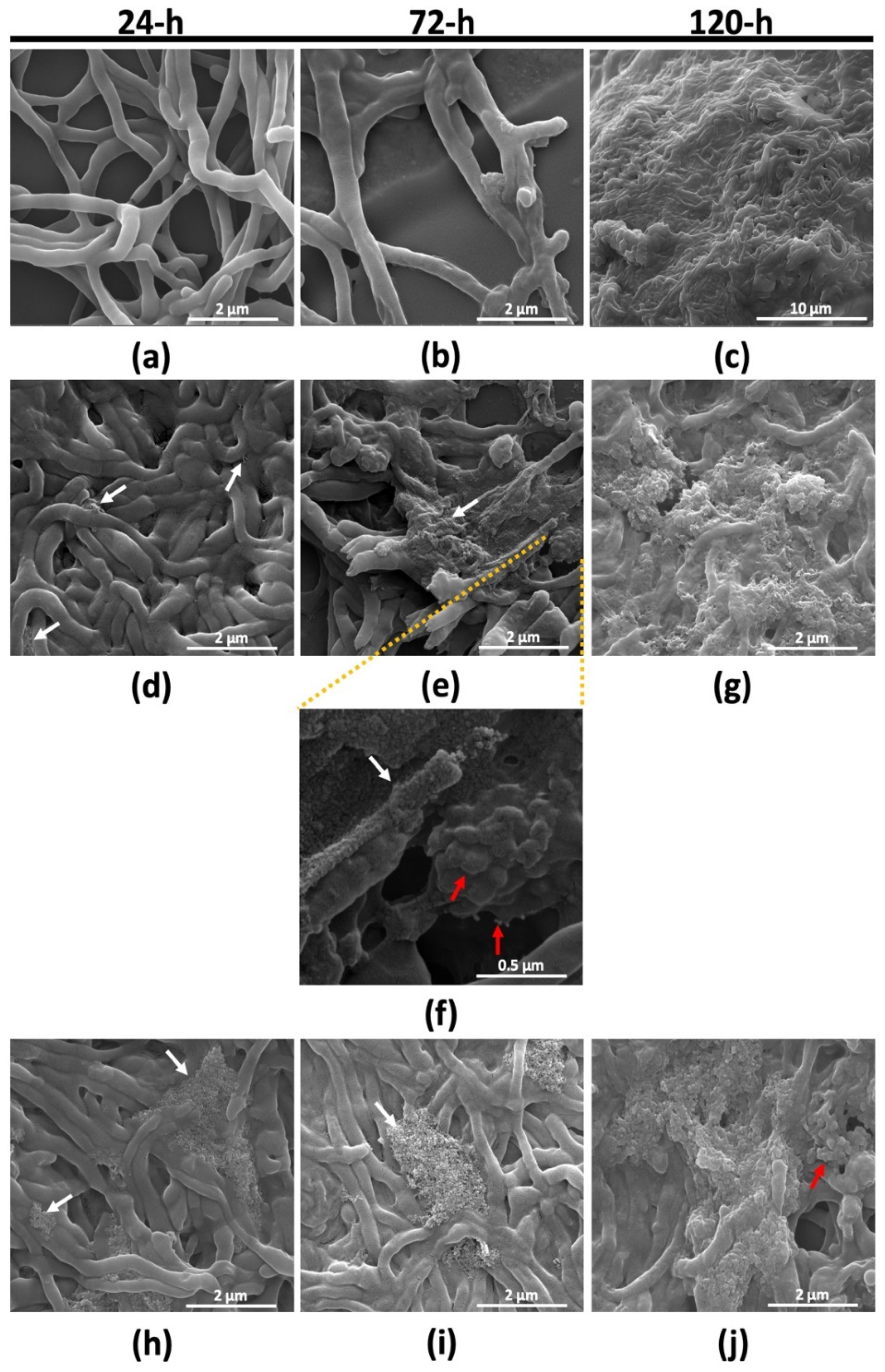
2.2.2. Morphological Characterization of Micromonospora Cells under Tellurite Stress
2.2.3. Fourier Transform Infrared Spectroscopy in Attenuated Total Reflectance (ATR-FTIR) Mode

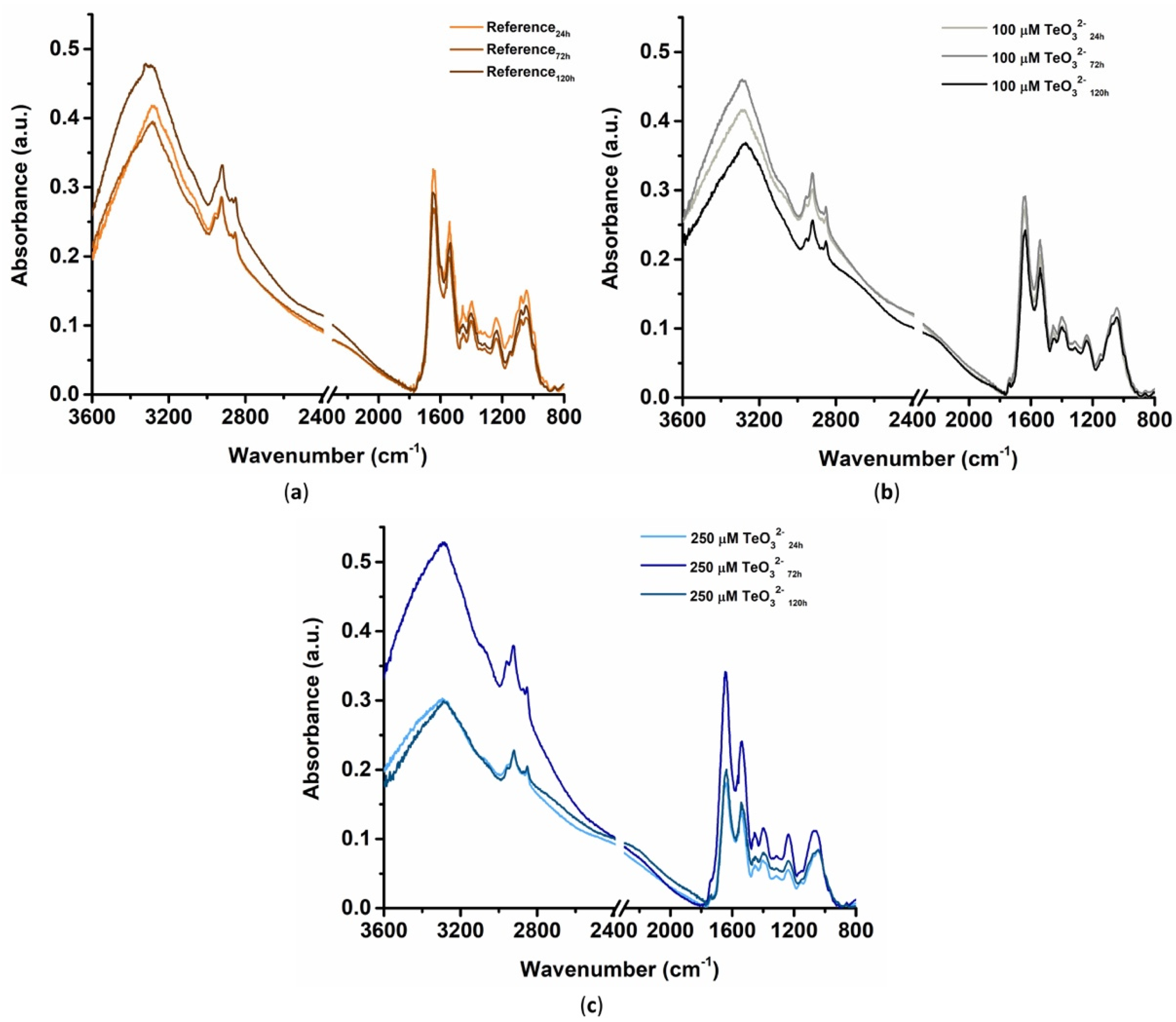
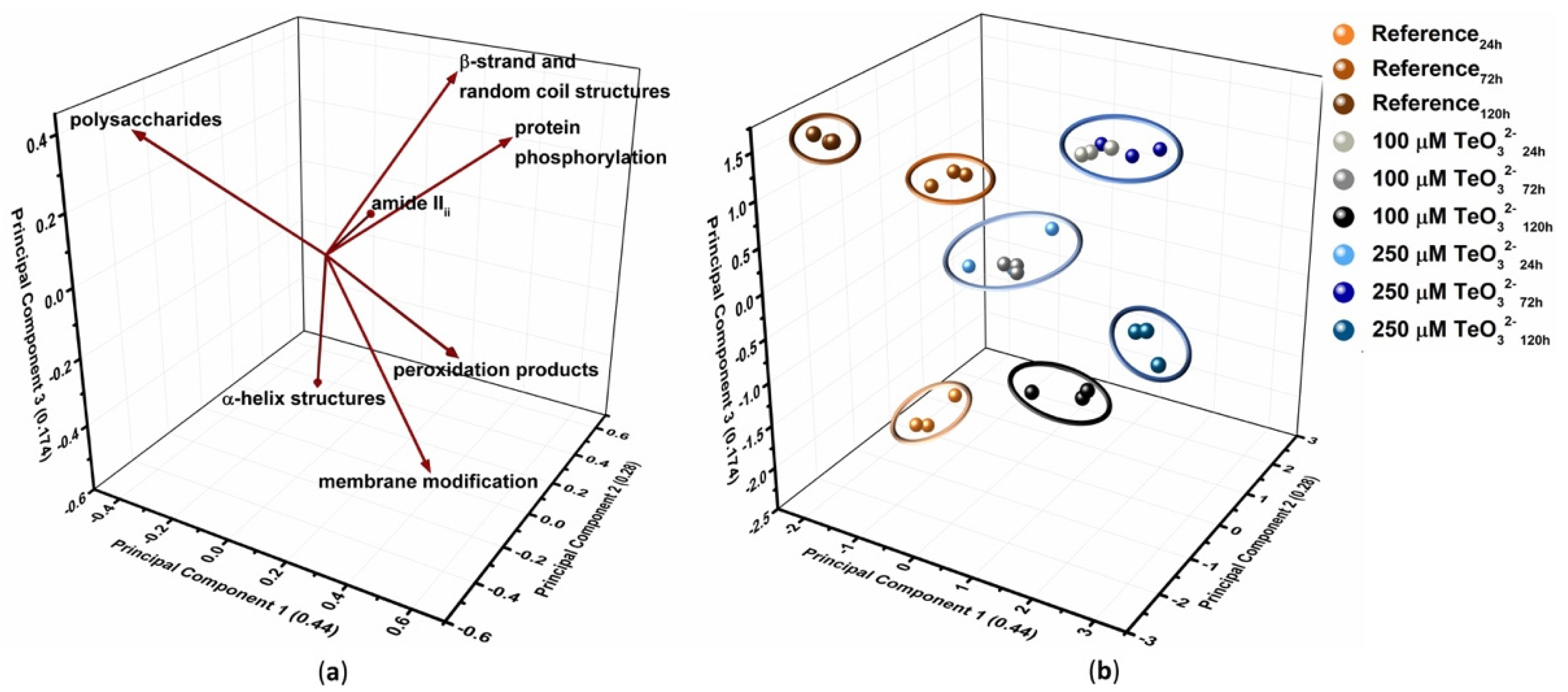
2.2.4. Multivariate Statistical Analysis
3. Discussion
3.1. TeO32− Effects on Exponentially Grown Cells
3.2. Toxicity, Adaptation, and Recovery of Micromonospora Cells Growing in the Presence of Tellurite
3.2.1. TeO32− Targets Micromonospora Cell Membrane
3.2.2. Involvement of Thiol-Containing Molecules
3.2.3. Peroxidation Products Deriving from Oxidative Stress
3.2.4. Protein Aggregation and Phosphorylation
4. Materials and Methods
4.1. Bacterial Strain, Growth Medium, and Culture Conditions
4.2. Tellurite Consumption and Uptake Assays
4.3. Thiol Oxidation Assay
4.4. Cell Viability Assay
4.5. Assessment of the Membrane Potential
4.6. Fatty Acid Methyl Esters Analysis
4.7. ROS Determination
4.8. SOD Activity Evaluation
4.9. Scanning Electron Microscopy (SEM) Imaging
4.10. ATR-FTIR Spectroscopy
4.11. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Piacenza, E.; Presentato, A.; Zonaro, E.; Lampis, S.; Vallini, G.; Turner, R.J. Selenium and Tellurium Nanomaterials. Phys. Sci. Rev. 2018, 3, 20170100. [Google Scholar] [CrossRef]
- Li, Z.; Qiu, F.; Tian, Q.; Yue, X.; Zhang, T. Production and Recovery of Tellurium from Metallurgical Intermediates and Electronic Waste-A Comprehensive Review. J. Clean. Prod. 2022, 366, 132796. [Google Scholar] [CrossRef]
- Ilyas, S.; Kim, M.S.; Lee, J.C.; Jabeen, A.; Bhatti, H.N. Bio-Reclamation of Strategic and Energy Critical Metals from Secondary Resources. Metals 2017, 7, 207. [Google Scholar] [CrossRef]
- Grygoyć, K.; Jabłońska-Czapla, M. Development of a Tellurium Speciation Study Using IC-ICP-MS on Soil Samples Taken from an Area Associated with the Storage, Processing, and Recovery of Electrowaste. Molecules 2021, 26, 2651. [Google Scholar] [CrossRef]
- Farias, P.; Francisco, R.; Morais, P.V. Potential of Tellurite Resistance in Heterotrophic Bacteria from Mining Environments. iScience 2022, 25, 104566. [Google Scholar] [CrossRef]
- Presentato, A.; Turner, R.J.; Vásquez, C.C.; Yurkov, V.; Zannoni, D. Tellurite-Dependent Blackening of Bacteria Emerges from the Dark Ages. Environ. Chem. 2019, 16, 266–288. [Google Scholar] [CrossRef]
- Curtin, A.M.; Vail, C.A.; Buckley, H.L. CdTe in Thin Film Photovoltaic Cells: Interventions to Protect Drinking Water in Production and End-of-Life. Water-Energy Nexus 2020, 3, 15–28. [Google Scholar] [CrossRef]
- Kessi, J.; Turner, R.J.; Zannoni, D. Tellurite and Selenite: How Can These Two Oxyanions Be Chemically Different yet so Similar in the Way They Are Transformed to Their Metal Forms by Bacteria? Biol. Res. 2022, 55, 17. [Google Scholar] [CrossRef]
- Presentato, A.; Piacenza, E.; Turner, R.J.; Zannoni, D.; Cappelletti, M. Processing of Metals and Metalloids by Actinobacteria: Cell Resistance Mechanisms and Synthesis of Metal(Loid)-Based Nanostructures. Microorganisms 2020, 8, 2027. [Google Scholar] [CrossRef]
- El-Tarabily, K.A.; Nassar, A.H.; Sivasithamparam, K. Promotion of Growth of Bean (Phaseolus vulgaris L.) in a Calcareous Soil by a Phosphate-Solubilizing, Rhizosphere-Competent Isolate of Micromonospora endolithica. Appl. Soil Ecol. 2008, 39, 161–171. [Google Scholar] [CrossRef]
- Ortúzar, M.; Trujillo, M.E.; Román-Ponce, B.; Carro, L. Micromonospora metallophores: A Plant Growth Promotion Trait Useful for Bacterial-Assisted Phytoremediation? Sci. Total Environ. 2020, 739, 139850. [Google Scholar] [CrossRef]
- Corsini, A.; Cavalca, L.; Crippa, L.; Zaccheo, P.; Andreoni, V. Impact of Glucose on Microbial Community of a Soil Containing Pyrite Cinders: Role of Bacteria in Arsenic Mobilization under Submerged Condition. Soil Biol. Biochem. 2010, 42, 699–707. [Google Scholar] [CrossRef]
- Raja, M.M.M.; John, S.A. Biosynthesis of Silver Nanoparticles by Novel Isolate of Marine Micromonospora Species (KU 867645) and Its Antibacterial Activity against Multidrug Resistant Hospital-Acquired Uropathogens in Reference with Standard Antibiotics. Indian J. Pharm. Sci. 2017, 79, 369–376. [Google Scholar] [CrossRef]
- Piacenza, E.; Presentato, A.; di Salvo, F.; Alduina, R.; Ferrara, V.; Minore, V.; Giannusa, A.; Sancataldo, G.; Chillura Martino, D.F. A Combined Physical–Chemical and Microbiological Approach to Unveil the Fabrication, Provenance, and State of Conservation of the Kinkarakawa-Gami Art. Sci. Rep. 2020, 10, 16072. [Google Scholar] [CrossRef]
- Borsetti, F.; Toninello, A.; Zannoni, D. Tellurite Uptake by Cells of the Facultative Phototroph Rhodobacter capsulatus Is a ΔpH-Dependent Process. FEBS Lett. 2003, 554, 315–318. [Google Scholar] [CrossRef]
- Becker, L.A.; Bang, I.S.; Crouch, M.L.; Fang, F.C. Compensatory Role of PspA, a Member of the Phage Shock Protein Operon, in RpoE Mutant Salmonella enterica serovar Typhimurium. Mol. Microbiol. 2005, 56, 1004–1016. [Google Scholar] [CrossRef]
- Singh, R.; Cheng, S.; Singh, S. Oxidative Stress-Mediated Genotoxic Effect of Zinc Oxide Nanoparticles on Deinococcus radiodurans. 3 Biotech 2020, 10, 66. [Google Scholar] [CrossRef]
- Tugarova, A.V.; Mamchenkova, P.V.; Dyatlova, Y.A.; Kamnev, A.A. FTIR and Raman Spectroscopic Studies of Selenium Nanoparticles Synthesised by the Bacterium Azospirillum thiophilum. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2018, 192, 458–463. [Google Scholar] [CrossRef]
- Kamnev, A.A.; Mamchenkova, P.V.; Dyatlova, Y.A.; Tugarova, A.V.; Kamnev, A.A.; Mamchenkova, P.V.; Dyatlova, Y.A.; Tugarova, A.V. FTIR Spectroscopic Studies of Selenite Reduction by Cells of the Rhizobacterium Azospirillum brasilense Sp7 and the Formation of Selenium Nanoparticles. J. Mol. Struct. 2017, 1140, 106–112. [Google Scholar] [CrossRef]
- Benseny-Cases, N.; Klementieva, O.; Cotte, M.; Ferrer, I.; Cladera, J. Microspectroscopy (ΜFTIR) Reveals Co-Localization of Lipid Oxidation and Amyloid Plaques in Human Alzheimer Disease Brains. Anal. Chem. 2014, 86, 12047–12054. [Google Scholar] [CrossRef]
- Barraza-Garza, G.; Castillo-Michel, H.; de La Rosa, L.A.; Martinez-Martinez, A.; Pérez-León, J.A.; Cotte, M.; Alvarez-Parrilla, E. Infrared Spectroscopy as a Tool. to Study the Antioxidant Activity of Polyphenolic Compounds in Isolated Rat Enterocytes. Oxid. Med. Cell. Longev. 2016, 2016, 9245150. [Google Scholar] [CrossRef]
- Gupta, A.D.; Karthikeyan, S. Individual and Combined Toxic Effect of Nickel and Chromium on Biochemical Constituents in E. coli Using FTIR Spectroscopy and Principle Component Analysis. Ecotoxicol. Environ. Saf. 2016, 130, 289–294. [Google Scholar] [CrossRef]
- Kiwi, J.; Nadtochenko, V. Evidence for the Mechanism of Photocatalytic Degradation of the Bacterial Wall Membrane at the TiO2 Interface by ATR-FTIR and Laser Kinetic Spectroscopy. Langmuir 2005, 21, 4631–4641. [Google Scholar] [CrossRef]
- Nadtochenko, V.A.; Rincon, A.G.; Stanca, S.E.; Kiwi, J. Dynamics of E. coli Membrane Cell Peroxidation during TiO2 Photocatalysis Studied by ATR-FTIR Spectroscopy and AFM Microscopy. J. Photochem. Photobiol. A 2005, 169, 131–137. [Google Scholar] [CrossRef]
- Vileno, B.; Jeney, S.; Sienkiewicz, A.; Marcoux, P.R.; Miller, L.M.; Forró, L. Evidence of Lipid Peroxidation and Protein Phosphorylation in Cells upon Oxidative Stress Photo-Generated by Fullerols. Biophys. Chem. 2010, 152, 164–169. [Google Scholar] [CrossRef]
- Nikonenko, N.A.; Buslov, D.K.; Sushko, N.I.; Zhbankov, R.G.; Stepanov, B.I. Investigation of Stretching Vibrations of Glycosidic Linkages in Disaccharides and Polysaccarides with Use of IR Spectra Deconvolution. Biopolymers 2000, 57, 257–262. [Google Scholar] [CrossRef]
- Piacenza, E.; Presentato, A.; Ferrante, F.; Cavallaro, G.; Alduina, R.; Chillura Martino, D.F. Biogenic Selenium Nanoparticles: A Fine Characterization to Unveil Their Thermodynamic Stability. Nanomaterials 2021, 11, 1195. [Google Scholar] [CrossRef]
- El-Meleigy, M.; Mohamed, H.; Mokhtar, M.; Salem, M. Streptomyces anulatus, a Tellurium Tolerant Actinomycete, Some Modes of Tolerance. Nat. Sci. 2011, 9, 141–153. [Google Scholar]
- Amoozegar, M.A.; Khoshnoodi, M.; Didari, M.; Hamedi, J.; Ventosa, A.; Baldwin, S.A. Tellurite Removal by a Tellurium-Tolerant Halophilic Bacterial Strain, Thermoactinomyces sp. QS-2006. Ann. Microbiol. 2012, 62, 1031–1037. [Google Scholar] [CrossRef]
- Furnholm, T.R.; Tisa, L.S. The Ins and Outs of Metal Homeostasis by the Root Nodule Actinobacterium Frankia. BMC Genom. 2014, 15, 1092. [Google Scholar] [CrossRef]
- Undabarrena, A.; Ugalde, J.A.; Seeger, M.; Cámara, B. Genomic Data Mining of the Marine Actinobacteria Streptomyces sp. H-KF8 Unveils Insights into Multi-Stress Related Genes and Metabolic Pathways Involved in Antimicrobial Synthesis. PeerJ 2017, 5, e2912. [Google Scholar] [CrossRef] [PubMed]
- Presentato, A.; Piacenza, E.; Anikovskiy, M.; Cappelletti, M.; Zannoni, D.; Turner, R.J. Rhodococcus aetherivorans BCP1 as Cell Factory for the Production of Intracellular Tellurium Nanorods under Aerobic Conditions. Microb. Cell Fact. 2016, 15, 204. [Google Scholar] [CrossRef] [PubMed]
- Borghese, R.; Marchetti, D.; Zannoni, D. The Highly Toxic Oxyanion Tellurite (TeO32−) Enters the Phototrophic Bacterium Rhodobacter capsulatus via an as yet Uncharacterized Monocarboxylate Transport System. Arch. Microbiol. 2008, 189, 93–100. [Google Scholar] [CrossRef] [PubMed]
- Borghese, R.; Zannoni, D. Acetate Permease (ActP) Is Responsible for Tellurite (TeO32−) Uptake and Resistance in Cells of the Facultative Phototroph Rhodobacter capsulatus. Appl. Environ. Microbiol. 2010, 76, 942–944. [Google Scholar] [CrossRef]
- Elías, A.; Díaz-Vásquez, W.; Abarca-Lagunas, M.J.; Chasteen, T.G.; Arenas, F.; Vásquez, C.C. The ActP Acetate Transporter Acts Prior to the PitA Phosphate Carrier in Tellurite Uptake by Escherichia coli. Microbiol. Res. 2015, 177, 15–21. [Google Scholar] [CrossRef]
- Borghese, R.; Brucale, M.; Fortunato, G.; Lanzi, M.; Mezzi, A.; Valle, F.; Cavallini, M.; Zannoni, D. Extracellular Production of Tellurium Nanoparticles by the Photosynthetic Bacterium Rhodobacter capsulatus. J. Hazard. Mater. 2016, 309, 202–209. [Google Scholar] [CrossRef]
- Martín, J.F.; Liras, P. Molecular Mechanisms of Phosphate Sensing, Transport and Signalling in Streptomyces and Related Actinobacteria. Int. J. Mol. Sci. 2021, 22, 1129. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, J.; Zhang, J.; Xu, W.; Mou, Z. Characteristics of Inorganic Phosphate-Solubilizing Bacteria from the Sediments of a Eutrophic Lake. Int. J. Environ. Res. Public Health 2019, 16, 2141. [Google Scholar] [CrossRef]
- Tian, J.; Ge, F.; Zhang, D.; Deng, S.; Liu, X. Roles of Phosphate Solubilizing Microorganisms from Managing Soil Phosphorus Deficiency to Mediating Biogeochemical P Cycle. Biology 2021, 10, 158. [Google Scholar] [CrossRef]
- Turner, R.J.; Weiner, J.H.; Taylor, D.E. Tellurite-Mediated Thiol. Oxidation in Escherichia coli. Microbiology 1999, 145, 2549–2557. [Google Scholar] [CrossRef]
- Vrionis, H.A.; Wang, S.; Haslam, B.; Turner, R.J. Selenite Protection of Tellurite Toxicity toward Escherichia coli. Front. Mol. Biosci. 2015, 2, 69. [Google Scholar] [CrossRef]
- Contreras, F.; Vargas, E.; Jiménez, K.; Muñoz-Villagrán, C.; Figueroa, M.; Vásquez, C.; Arenas, F. Reduction of Gold (III) and Tellurium (IV) by Enterobacter cloacae MF01 Results in Nanostructure Formation Both in Aerobic and Anaerobic Conditions. Front. Microbiol. 2018, 9, 3118. [Google Scholar] [CrossRef]
- Newton, G.L.; Buchmeier, N.; Fahey, R.C. Biosynthesis and Functions of Mycothiol, the Unique Protective Thiol. of Actinobacteria. Microbiol. Mol. Biol. Rev. 2008, 72, 471–494. [Google Scholar] [CrossRef]
- Morales, E.H.; Pinto, C.A.; Luraschi, R.; Muñoz-Villagrán, C.M.; Cornejo, F.A.; Simpkins, S.W.; Nelson, J.; Arenas, F.A.; Piotrowski, J.S.; Myers, C.L.; et al. Accumulation of Heme Biosynthetic Intermediates Contributes to the Antibacterial Action of the Metalloid Tellurite. Nat. Commun. 2017, 8, 15320. [Google Scholar] [CrossRef]
- Borsetti, F.; Tremaroli, V.; Michelacci, F.; Borghese, R.; Winterstein, C.; Daldal, F.; Zannoni, D. Tellurite Effects on Rhodobacter capsulatus Cell Viability and Superoxide Dismutase Activity under Oxidative Stress Conditions. Res. Microbiol. 2005, 156, 807–813. [Google Scholar] [CrossRef]
- Pérez, J.M.; Calderón, I.L.; Arenas, F.A.; Fuentes, D.E.; Pradenas, G.A.; Fuentes, E.L.; Sandoval, J.M.; Castro, M.E.; Elías, A.O.; Vásquez, C.C. Bacterial Toxicity of Potassium Tellurite: Unveiling an Ancient Enigma. PLoS ONE 2007, 2, e211. [Google Scholar] [CrossRef]
- Tremaroli, V.; Fedi, S.; Zannoni, D. Evidence for a Tellurite-Dependent Generation of Reactive Oxygen Species and Absence of a Tellurite-Mediated Adaptive Response to Oxidative Stress in Cells of Pseudomonas pseudoalcaligenes KF707. Arch. Microbiol. 2007, 187, 127–135. [Google Scholar] [CrossRef]
- Tremaroli, V.; Workentine, M.L.; Weljie, A.M.; Vogel, H.J.; Ceri, H.; Viti, C.; Tatti, E.; Zhang, P.; Hynes, A.P.; Turner, R.J.; et al. Metabolomic Investigation of the Bacterial Response to a Metal Challenge. Appl. Environ. Microbiol. 2009, 75, 719–728. [Google Scholar] [CrossRef]
- Arenas, F.A.; Pugin, B.; Henríquez, N.A.; Arenas-Salinas, M.A.; Díaz-Vásquez, W.A.; Pozo, M.F.; Muñoz, C.M.; Chasteen, T.G.; Pérez-Donoso, J.M.; Vásquez, C.C. Isolation, Identification and Characterization of Highly Tellurite-Resistant, Tellurite-Reducing Bacteria from Antarctica. Polar Sci. 2014, 8, 40–52. [Google Scholar] [CrossRef]
- Konings, W.N.; Albers, S.-V.; Koning, S.; Driessen, A.J.M. The Cell Membrane Plays a Crucial Role in Survival of Bacteria and Archaea in Extreme Environments. Antonie Van Leeuwenhoek 2002, 81, 61–72. [Google Scholar] [CrossRef]
- de Carvalho, C.C.C.R.; Caramujo, M.J. The Various Roles of Fatty Acids. Molecules 2018, 23, 2583. [Google Scholar] [CrossRef]
- Männistö, M.K.; Puhakka, J.A. Temperature- and Growth-Phase-Regulated Changes in Lipid Fatty Acid Structures of Psychrotolerant Groundwater Proteobacteria. Arch. Microbiol. 2001, 177, 41–46. [Google Scholar] [CrossRef]
- Nichols, D.S.; Russell, N.J. Fatty Acid Adaptation in an Antarctic Bacterium-Changes in Primer Utilization. Microbiology 1996, 142, 747–754. [Google Scholar] [CrossRef]
- Bowman, J.P.; McCammon, S.A.; Nichols, D.S.; Skerrat, J.H.; Rea, S.M.; Nichols, P.D.; McMeekin, T.A. Shewanella gelidimarina sp. nov. and Shewanella frigidimarina sp. nov., Novel Antarctic Species with the Ability to Produce Eicosapentaenoic Acid (20:5 Omega 3) and Grow Anaerobically by Dissimilatory Fe(III) Reduction. Int. J. Syst. Evol. Microbiol. 1997, 47, 1040–1047. [Google Scholar] [CrossRef]
- Suutari, M.; Laakso, S. Microbial Fatty Acids and Thermal Adaptation. Crit. Rev. Biotechnol. 1994, 20, 285–328. [Google Scholar] [CrossRef]
- Markowicz, A.; Plociniczak, T.; Piotrovska-Seget, Z. Response of Bacteria to Heavy Metals Measured as Changes in FAME Profiles. Pol. J. Environ. Stud. 2010, 19, 957–965. [Google Scholar]
- Pradenas, G.A.; Paillavil, B.A.; Reyes-Cerpa, S.; Pérez-Donoso, J.M.; Vásquez, C.C. Reduction of the Monounsaturated Fatty Acid Content of Escherichia coli Results in Increased Resistance to Oxidative Damage. Microbiology 2012, 158, 1279–1283. [Google Scholar] [CrossRef]
- Borghese, R.; Borsetti, F.; Foladori, P.; Ziglio, G.; Zannoni, D. Effects of the Metalloid Oxyanion Tellurite (TeO32-) on Growth Characteristics of the Phototrophic Bacterium Rhodobacter capsulatus. Appl. Environ. Microbiol. 2004, 70, 6595–6602. [Google Scholar] [CrossRef]
- Murínová, S.; Dercová, K. Response Mechanisms of Bacterial Degraders to Environmental Contaminants on the Level of Cell Walls and Cytoplasmic Membrane. Int. J. Microbiol. 2014, 2014, 873081. [Google Scholar] [CrossRef]
- Mrozik, A.; Łabuzek, S.; Piotrowska-Seget, Z. Changes in Fatty Acid Composition in Pseudomonas putida and Pseudomonas stutzeri during Naphthalene Degradation. Microbiol. Res. 2005, 160, 149–157. [Google Scholar] [CrossRef]
- Mrozik, A.; Cycoń, M.; Piotrowska-Seget, Z. Changes of FAME Profiles as a Marker of Phenol. Degradation in Different Soils Inoculated with Pseudomonas sp. CF600. Int. Biodeterior. Biodegrad. 2010, 64, 86–96. [Google Scholar] [CrossRef]
- Unell, M.; Kabelitz, N.; Jansson, J.K.; Heipieper, H.J. Adaptation of the Psychrotroph Arthrobacter chlorophenolicus A6 to Growth Temperature and the Presence of Phenols by Changes in the Anteiso/Iso Ratio of Branched Fatty Acids. FEMS Microbiol. Lett. 2007, 266, 138–143. [Google Scholar] [CrossRef] [PubMed]
- Antony, R.; Sujith, P.P.; Fernandes, S.O.; Verma, P.; Khedekar, V.D.; Loka Bharathi, P.A. Cobalt Immobilization by Manganese Oxidizing Bacteria from the Indian Ridge System. Curr. Microbiol. 2011, 62, 840–849. [Google Scholar] [CrossRef] [PubMed]
- Nepple, B.B.; Bachofen, R. Morphological Changes in Phototrophic Bacteria Induced by Metalloid Oxyanions. Microbiol. Res. 1999, 154, 191–198. [Google Scholar] [CrossRef]
- Chakravarty, R.; Manna, S.; Ghosh, A.K.; Banerjee, P.C. Morphological Changes in an Acidocella Strain in Response to Heavy Metal Stress. Res. J. Microbiol. 2007, 2, 742–748. [Google Scholar]
- Colin, V.L.; Villegas, L.B.; Pereira, C.E.; Amoroso, M.J.; Abate, C.M. Morphological Changes and Oxidative Stress in Actinobacteria during Removal of Heavy Metals. In Actinobacteria, 1st ed.; Maroso, M.J., Benimeli, C.S., Cuozzo, S.A., Eds.; CRC Press—Taylor & Francis Group: Boca Raton, FL, USA, 2013; pp. 52–63. ISBN 978-0-42907-137-9. [Google Scholar]
- de Carvalho, C.C.C.R. Adaptation of Rhodococcus erythropolis Cells for Growth and Bioremediation under Extreme Conditions. Res. Microbiol. 2012, 163, 125–136. [Google Scholar] [CrossRef]
- Neumann, G.; Veeranagouda, Y.; Karegoudar, T.B.; Sahin, Ö.; Mäusezahl, I.; Kabelitz, N.; Kappelmeyer, U.; Heipieper, H.J. Cells of Pseudomonas putida and Enterobacter sp. Adapt to Toxic Organic Compounds by Increasing Their Size. Extremophiles 2005, 9, 163–168. [Google Scholar] [CrossRef]
- Kepenek, E.S.; Gozen, A.G.; Severcan, F. Molecular Characterization of Acutely and Gradually Heavy Metal Acclimated Aquatic Bacteria by FTIR Spectraoscopy. J. Biophotonics 2019, 12, e201800301. [Google Scholar] [CrossRef]
- Shuhong, Y.; Meiping, Z.; Hong, Y.; Han, W.; Shan, X.; Yan, L.; Jihui, W. Biosorption of Cu2+, Pb2+ and Cr6+ by a Novel Exopolysaccharide from Arthrobacter Ps-5. Carbohydr. Polym. 2014, 101, 50–56. [Google Scholar] [CrossRef]
- Bankar, A.; Nagaraja, G. Recent Trends in Biosorption of Heavy Metals by Actinobacteria. In New and Future Developments in Microbial Biotechnology and Bioengineering: Actinobacteria: Diversity and Biotechnological Applications; Singh, B.P., Gupta, V.K., Passari, A.K., Eds.; Elsevier: Amsterdam, The Netherlands, 2018; Chapter 18; pp. 257–275. ISBN 978-0-44463-995-0. [Google Scholar]
- Xiong, Y.W.; Ju, X.Y.; Li, X.W.; Gong, Y.; Xu, M.J.; Zhang, C.M.; Yuan, B.; Lv, Z.P.; Qin, S. Fermentation Conditions Optimization, Purification, and Antioxidant Activity of Exopolysaccharides Obtained from the Plant Growth-Promoting Endophytic Actinobacterium Glutamicibacter halophytocola KLBMP 5180. Int. J. Biol. Macromol. 2020, 153, 1176–1185. [Google Scholar] [CrossRef]
- Kardas, M.; Gozen, A.G.; Severcan, F. FTIR Spectroscopy Offers Hints towards Widespread Molecular Changes in Cobalt-Acclimated Freshwater Bacteria. Aquat. Toxicol. 2014, 155, 15–23. [Google Scholar] [CrossRef]
- Vishnoi, N.; Dixit, S.; Singh, D.P. Surface Binding and Intracellular Uptake of Arsenic in Bacteria Isolated from Arsenic Contaminated Site. Ecol. Eng. 2014, 73, 569–578. [Google Scholar] [CrossRef]
- Gupta, A.D.; Kavitha, E.; Singh, S.; Karthikeyan, S. Toxicity Mechanism of Cu2+ Ion Individually and in Combination with Zn2+ Ion in Characterizing the Molecular Changes of Staphylococcus aureus Studied Using FTIR Coupled with Chemometric Analysis. J. Biol. Phys. 2020, 46, 395–414. [Google Scholar] [CrossRef]
- Goff, J.L.; Wang, Y.; Boyanov, M.I.; Yu, Q.; Kemner, K.M.; Fein, J.B.; Yee, N. Tellurite Adsorption onto Bacterial Surfaces. Environ. Sci. Technol. 2021, 55, 10378–10386. [Google Scholar] [CrossRef]
- Gurbanov, R.; Simsek Ozek, N.; Gozen, A.G.; Severcan, F. Quick Discrimination of Heavy Metal Resistant Bacterial Populations Using Infrared Spectroscopy Coupled with Chemometrics. Anal. Chem. 2015, 87, 9653–9661. [Google Scholar] [CrossRef]
- Castro, M.E.; Molina, R.; Díaz, W.; Pichuantes, S.E.; Vásquez, C.C. The Dihydrolipoamide Dehydrogenase of Aeromonas caviae ST Exhibits NADH-Dependent Tellurite Reductase Activity. Biochem. Biophys. Res. Commun. 2008, 375, 91–94. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, L.; Peng, J.; Yuan, F.; Li, J.; Wang, J.; Chen, J.; Wang, H.; Tyagi, R.D. Heavy Metal Impact on Lipid Production from Oleaginous Microorganism Cultivated with Wastewater Sludge. Bioresour. Technol. 2022, 344, 126356. [Google Scholar] [CrossRef]
- Ami, D.; Natalello, A.; Schultz, T.; Gatti-Lafranconi, P.; Lotti, M.; Doglia, S.M.; de Marco, A. Effects of Recombinant Protein Misfolding and Aggregation on Bacterial Membranes. Biochim. Biophys. Acta-Proteins Proteom. 2009, 1794, 263–269. [Google Scholar] [CrossRef]
- Schwechheimer, C.; Kuehn, M.J. Outer-Membrane Vesicles from Gram-Negative Bacteria: Biogenesis and Functions. Nat. Rev. Microbiol. 2015, 13, 605–619. [Google Scholar] [CrossRef]
- Faddetta, T.; Renzone, G.; Vassallo, A.; Rimini, E.; Nasillo, G.; Buscarino, G.; Agnello, S.; Licciardi, M.; Botta, L.; Scaloni, A.; et al. Streptomyces coelicolor Vesicles: Many Molecules To Be Delivered. Appl. Environ. Microbiol. 2022, 88, e01881-21. [Google Scholar] [CrossRef]
- Schrempf, H.; Koebsch, I.; Walter, S.; Engelhardt, H.; Meschke, H. Extracellular Streptomyces Vesicles: Amphorae for Survival and Defence. Microb. Biotechnol. 2011, 4, 286–299. [Google Scholar] [CrossRef]
- Chasteen, T.G.; Fuentes, D.E.; Tantaleán, J.C.; Vásquez, C.C. Tellurite: History, Oxidative Stress, and Molecular Mechanisms of Resistance. FEMS Microbiol. Rev. 2009, 33, 820–832. [Google Scholar] [CrossRef]
- Kessi, J.; Hanselmann, K.W. Similarities between the Abiotic Reduction of Selenite with Glutathione and the Dissimilatory Reaction Mediated by Rhodospirillum rubrum and Escherichia coli. J. Biol. Chem. 2004, 279, 50662–50669. [Google Scholar] [CrossRef]
- Pérez, J.M.; Arenas, F.A.; Pradenas, G.A.; Sandoval, J.M.; Vásquez, C.C. Escherichia coli YqhD Exhibits Aldehyde Reductase Activity and Protects from the Harmful Effect of Lipid Peroxidation-Derived Aldehydes. J. Biol. Chem. 2008, 283, 7346–7353. [Google Scholar] [CrossRef]
- Pradenas, G.A.; Díaz-Vásquez, W.A.; Pérez-Donoso, J.M.; Vásquez, C.C. Monounsaturated Fatty Acids Are Substrates for Aldehyde Generation in Tellurite-Exposed Escherichia coli. BioMed Res. Int. 2013, 2013, 563756. [Google Scholar] [CrossRef]
- Contreras, N.D.P.; Vásquez, C.C. Tellurite-Induced Carbonylation of the Escherichia coli Pyruvate Dehydrogenase Multienzyme Complex. Arch. Microbiol. 2010, 192, 969–973. [Google Scholar] [CrossRef]
- Schramm, F.D.; Schroeder, K.; Jonas, K. Protein Aggregation in Bacteria. FEMS Microbiol. Rev. 2020, 44, 54–72. [Google Scholar] [CrossRef]
- Jacobson, T.; Navarrete, C.; Sharma, S.K.; Sideri, T.C.; Ibstedt, S.; Priya, S.; Grant, C.M.; Christen, P.; Goloubinoff, P.; Tamá, M.J. Arsenite Interferes with Protein Folding and Triggers Formation of Protein Aggregates in Yeast. J. Cell Sci. 2012, 125, 5073–5083. [Google Scholar] [CrossRef]
- Gerbino, E.; Mobili, P.; Tymczyszyn, E.; Fausto, R.; Gómez-Zavaglia, A. FTIR Spectroscopy Structural Analysis of the Interaction between Lactobacillus kefir S-Layers and Metal Ions. J. Mol. Struct. 2011, 987, 186–192. [Google Scholar] [CrossRef]
- Parker, J.L.; Jones, A.M.E.; Serazetdinova, L.; Saalbach, G.; Bibb, M.J.; Naldrett, M.J. Analysis of the Phosphoproteome of the Multicellular Bacterium Streptomyces coelicolor A3(2) by Protein/Peptide Fractionation, Phosphopeptide Enrichment and High-Accuracy Mass Spectrometry. Proteomics 2010, 10, 2486–2497. [Google Scholar] [CrossRef]
- Ortiz De Orué Lucana, D.; Groves, M.R. The Three-Component Signalling System HbpS-SenS-SenR as an Example of a Redox Sensing Pathway in Bacteria. Amino Acids 2009, 37, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Neu, J.M.; MacMillan, S.V.; Nodwell, J.R.; Wright, G.D. StoPK-1, a Serine/Threonine Protein Kinase from the Glycopeptide Antibiotic Producer Streptomyces toyocaensis NRRL 15009, Affects Oxidative Stress Response. Mol. Microbiol. 2002, 44, 417–430. [Google Scholar] [CrossRef] [PubMed]
- Piacenza, E.; Presentato, A.; Bardelli, M.; Lampis, S.; Vallini, G.; Turner, R.J. Influence of Bacterial Physiology on Processing of Selenite, Biogenesis of Nanomaterials and Their Thermodynamic Stability. Molecules 2019, 24, 2532. [Google Scholar] [CrossRef] [PubMed]
- Turner, R.J.; Weiner, J.H.; Taylor, D.E. Use of Diethyldithiocarbamate for Quantitative Determination of Tellurite Uptake by Bacteria. Anal. Biochem. 1992, 204, 292–295. [Google Scholar] [CrossRef]
- Piacenza, E.; Presentato, A.; Zonaro, E.; Lemire, J.A.; Demeter, M.; Vallini, G.; Turner, R.J.; Lampis, S. Antimicrobial Activity of Biogenically Produced Spherical Se-Nanomaterials Embedded in Organic Material against Pseudomonas aeruginosa and Staphylococcus aureus Strains on Hydroxyapatite-Coated Surfaces. Microb. Biotechnol. 2017, 10, 804–818. [Google Scholar] [CrossRef]
- Hinojosa, M.B.; Laudicina, V.A.; Parra, A.; Albert-Belda, E.; Moreno, J.M. Drought and Its Legacy Modulate the Post-Fire Recovery of Soil Functionality and Microbial Community Structure in a Mediterranean Shrubland. Glob. Chang. Biol. 2019, 25, 1409–1427. [Google Scholar] [CrossRef]
- Presentato, A.; Piacenza, E.; Scurria, A.; Albanese, L.; Zabini, F.; Meneguzzo, F.; Nuzzo, D.; Pagliaro, M.; Chillura Martino, D.F.; Alduina, R.; et al. A New Water-Soluble Bactericidal Agent for the Treatment of Infections Caused by Gram-Positive and Gram-Negative Bacterial Strains. Antibiotics 2020, 9, 586. [Google Scholar] [CrossRef]
- Zonaro, E.; Piacenza, E.; Presentato, A.; Monti, F.; Dell’Anna, R.; Lampis, S.; Vallini, G. Ochrobactrum Sp. MPV1 from a Dump of Roasted Pyrites Can Be Exploited as Bacterial Catalyst for the Biogenesis of Selenium and Tellurium Nanoparticles. Microb. Cell Fact. 2017, 16, 215. [Google Scholar] [CrossRef]
- Faghihzadeh, F.; Anaya, N.M.; Schifman, L.A.; Oyanedel-Craver, V. Fourier Transform Infrared Spectroscopy to Assess Molecular-Level Changes in Microorganisms Exposed to Nanoparticles. Nanotechnol. Environ. Eng. 2016, 1, 1–16. [Google Scholar] [CrossRef]
- Lasch, P.; Naumann, D. Infrared Spectroscopy in Microbiology. In Encyclopedia of Analytical Chemistry; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2015; pp. 1–32. [Google Scholar]
- Wang, X.; Wang, W.; Liu, P.; Wang, P.; Zhang, L. Photocatalytic Degradation of E.Coli Membrane Cell in the Presence of ZnO Nanowires. J. Wuhan Univ. Technol. Mater. Sci. Ed. 2011, 26, 222–225. [Google Scholar] [CrossRef]
- Jiang, W.; Saxena, A.; Song, B.; Ward, B.B.; Beveridge, T.J.; Myneni, S.C.B. Elucidation of Functional Groups on Gram-Positive and Gram-Negative Bacterial Surfaces Using Infrared Spectroscopy. Langmuir 2004, 20, 11433–11442. [Google Scholar] [CrossRef]
- Buszewski, B.; Dziubakiewicz, E.; Pomastowski, P.; Hrynkiewicz, K.; Ploszaj-Pyrek, J.; Talik, E.; Kramer, M.; Albert, K. Assignment of Functional Groups in Gram-Positive Bacteria. J. Anal. Bioanal. Tech. 2015, 6, 1–8. [Google Scholar] [CrossRef]
- Otari, S.V.; Patil, R.M.; Ghosh, S.J.; Thorat, N.D.; Pawar, S.H. Intracellular Synthesis of Silver Nanoparticle by Actinobacteria and Its Antimicrobial Activity. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2015, 136, 1175–1180. [Google Scholar] [CrossRef]
- Mohamed, M.E.; Mohammed, A.M.A. Experimental and Computational Vibration Study of Amino Acids. Int. Lett. Chem. Phys. Astron. 2013, 15, 1–17. [Google Scholar] [CrossRef]
- Kurihara, T.; Noda, Y.; Takegoshi, K. Capping Structure of Ligand-Cysteine on CdSe Magic-Sized Clusters. ACS Omega 2019, 4, 3476–3483. [Google Scholar] [CrossRef]
- Barth, A. The Infrared Absorption of Amino Acid Side Chains. Prog. Biophys. Mol. Biol. 2000, 74, 141–173. [Google Scholar] [CrossRef]
- Garip, S.; Gozen, A.C.; Severcan, F. Use of Fourier Transform Infrared Spectroscopy for Rapid Comparative Analysis of Bacillus and Micrococcus Isolates. Food Chem. 2009, 113, 1301–1307. [Google Scholar] [CrossRef]
| Relative Abundance (%) | ||
|---|---|---|
| Fatty Acid | Control Cells | TeO32−-Challenged Cells |
| C15:0 | 9.88 ± 0.26 | 3.67 ± 0.80 |
| C16:0 | 4.59 ± 0.07 | 3.64 ± 0.01 |
| C17:0 | 18.0 ± 0.2 | 10.3 ± 0.6 |
| C18:0 | 3.56 ± 0.27 | 8.95 ± 0.97 |
| iso-C15:0 | 6.23 ± 0.03 | 8.27 ± 0.25 |
| anteiso-C15:0 | 3.09 ± 0.08 | 2.78 ± 0.01 |
| iso-C16:0 | 12.4 ±0.9 | 22.6 ± 1.4 |
| iso-C17:0 | 1.93 ± 0.09 | 3.37 ± 0.28 |
| C16:1ω7c | 3.02 ± 0.21 | 4.78 ± 0.11 |
| C17:1 | 28.0 ± 0.9 | 20.2 ± 0.9 |
| C18:1ω9c | 9.09 ± 0.23 | 11.7 ± 1.7 |
| Even saturated fatty acid | 20.5 ± 1.3 | 36.1 ± 0.5 |
| Odd saturated fatty acid | 39.4 ± 1.2 | 28.2 ± 0.5 |
| Saturated fatty acid | 59.9 ± 0.8 | 64.3 ± 0.9 |
| Unsaturated fatty acid | 40.1 ± 0.5 | 35.7 ± 0.9 |
| Saturated/Unsaturated | 1.49 ± 0.04 | 1.80 ± 0.07 |
| PC1 | PC2 | PC3 | |
|---|---|---|---|
| Information (%) | 44.0 | 28.0 | 17.4 |
| Discriminating 1 Vectors | protein phosphorylation | amide IIII | polysaccharides |
| membrane integrity | polysaccharides | β-strand + random coil structure | |
| peroxidation products | α-helix structure | ||
| polysaccharides |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Piacenza, E.; Campora, S.; Carfì Pavia, F.; Chillura Martino, D.F.; Laudicina, V.A.; Alduina, R.; Turner, R.J.; Zannoni, D.; Presentato, A. Tolerance, Adaptation, and Cell Response Elicited by Micromonospora sp. Facing Tellurite Toxicity: A Biological and Physical-Chemical Characterization. Int. J. Mol. Sci. 2022, 23, 12631. https://doi.org/10.3390/ijms232012631
Piacenza E, Campora S, Carfì Pavia F, Chillura Martino DF, Laudicina VA, Alduina R, Turner RJ, Zannoni D, Presentato A. Tolerance, Adaptation, and Cell Response Elicited by Micromonospora sp. Facing Tellurite Toxicity: A Biological and Physical-Chemical Characterization. International Journal of Molecular Sciences. 2022; 23(20):12631. https://doi.org/10.3390/ijms232012631
Chicago/Turabian StylePiacenza, Elena, Simona Campora, Francesco Carfì Pavia, Delia Francesca Chillura Martino, Vito Armando Laudicina, Rosa Alduina, Raymond Joseph Turner, Davide Zannoni, and Alessandro Presentato. 2022. "Tolerance, Adaptation, and Cell Response Elicited by Micromonospora sp. Facing Tellurite Toxicity: A Biological and Physical-Chemical Characterization" International Journal of Molecular Sciences 23, no. 20: 12631. https://doi.org/10.3390/ijms232012631
APA StylePiacenza, E., Campora, S., Carfì Pavia, F., Chillura Martino, D. F., Laudicina, V. A., Alduina, R., Turner, R. J., Zannoni, D., & Presentato, A. (2022). Tolerance, Adaptation, and Cell Response Elicited by Micromonospora sp. Facing Tellurite Toxicity: A Biological and Physical-Chemical Characterization. International Journal of Molecular Sciences, 23(20), 12631. https://doi.org/10.3390/ijms232012631













