Nanomechanical Atomic Force Microscopy to Probe Cellular Microplastics Uptake and Distribution
Abstract
:1. Introduction
2. Results
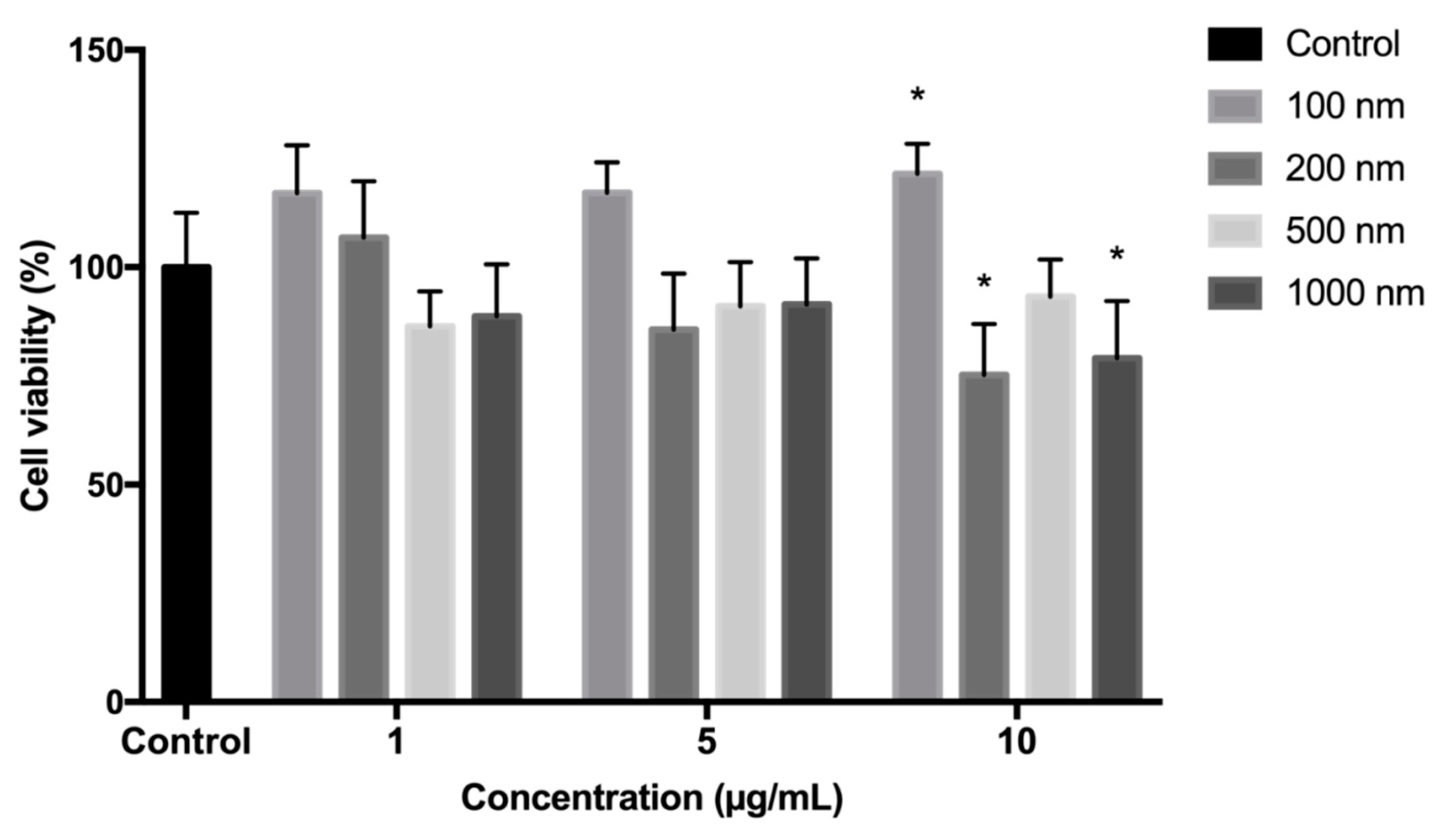
2.1. Polystyrene Microplastics Cytotoxicity Evaluation
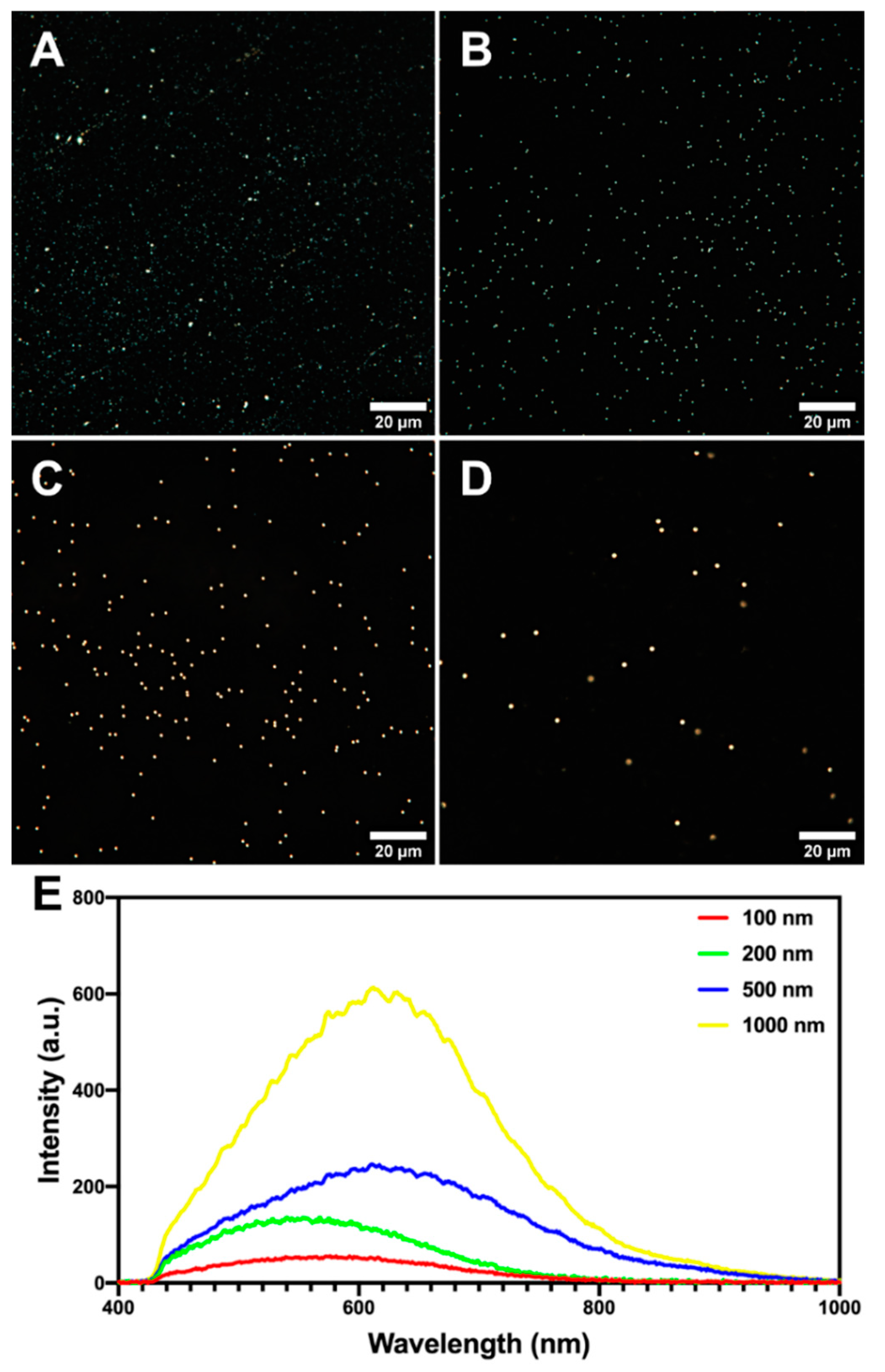
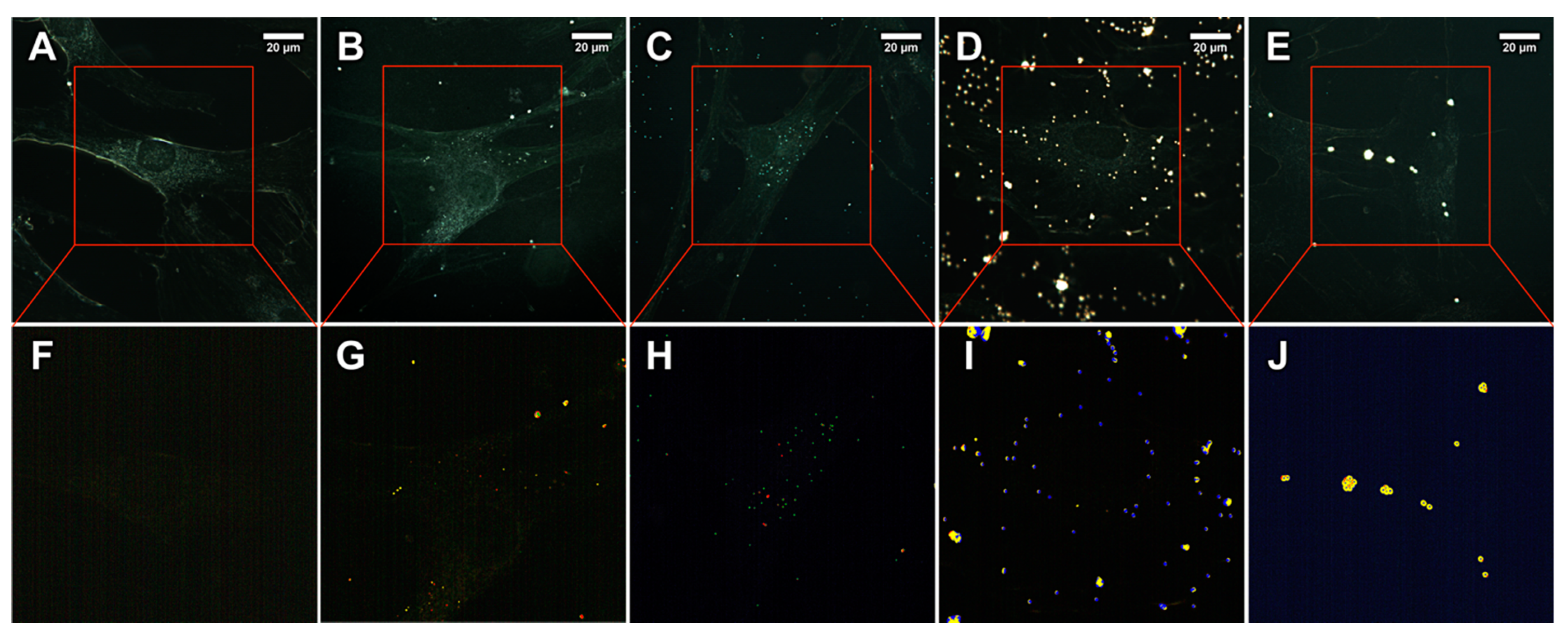
2.2. Microplastics Characterisation and Uptake Imaging with Dark-Field Hyperspectral Microscopy
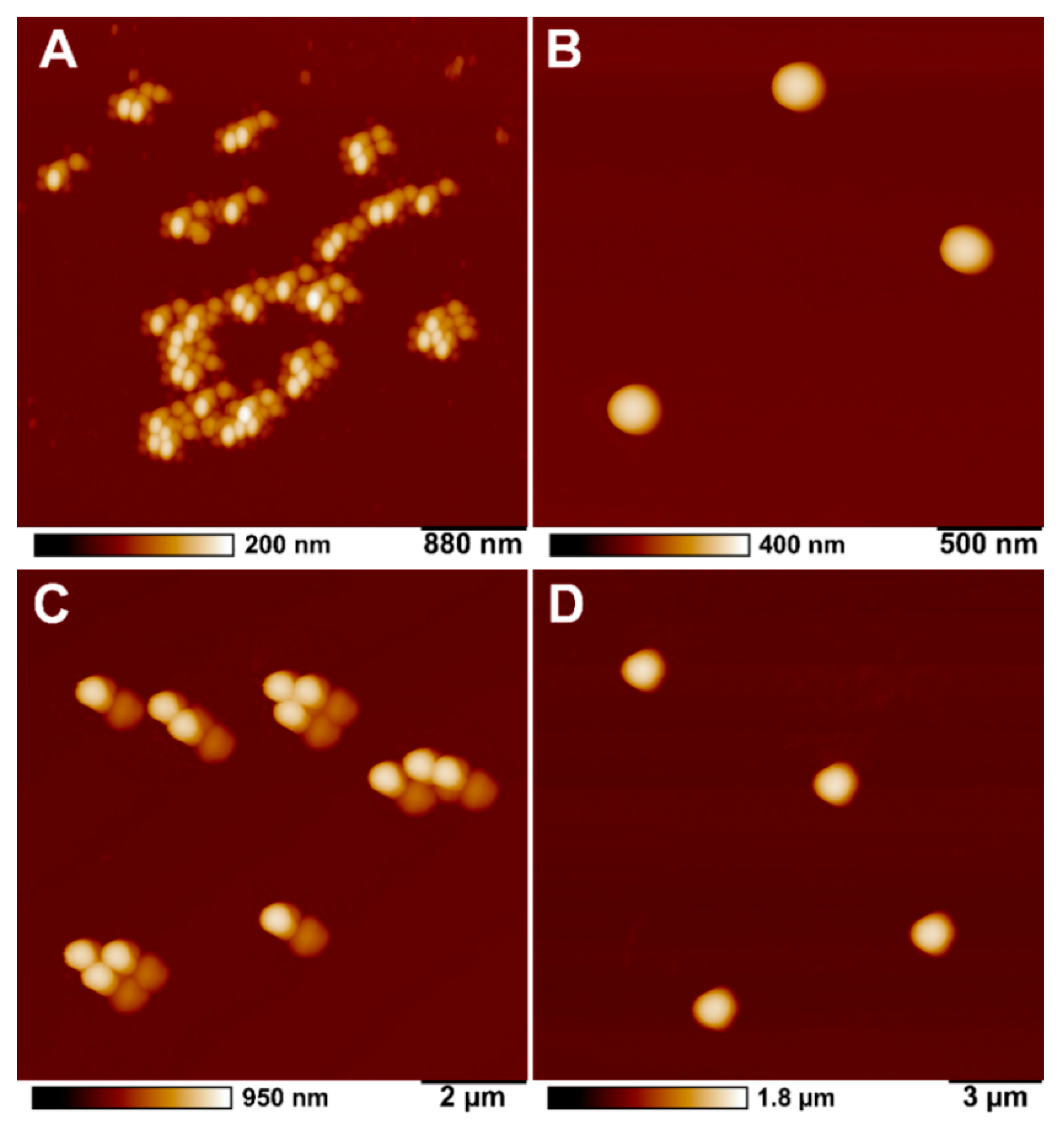
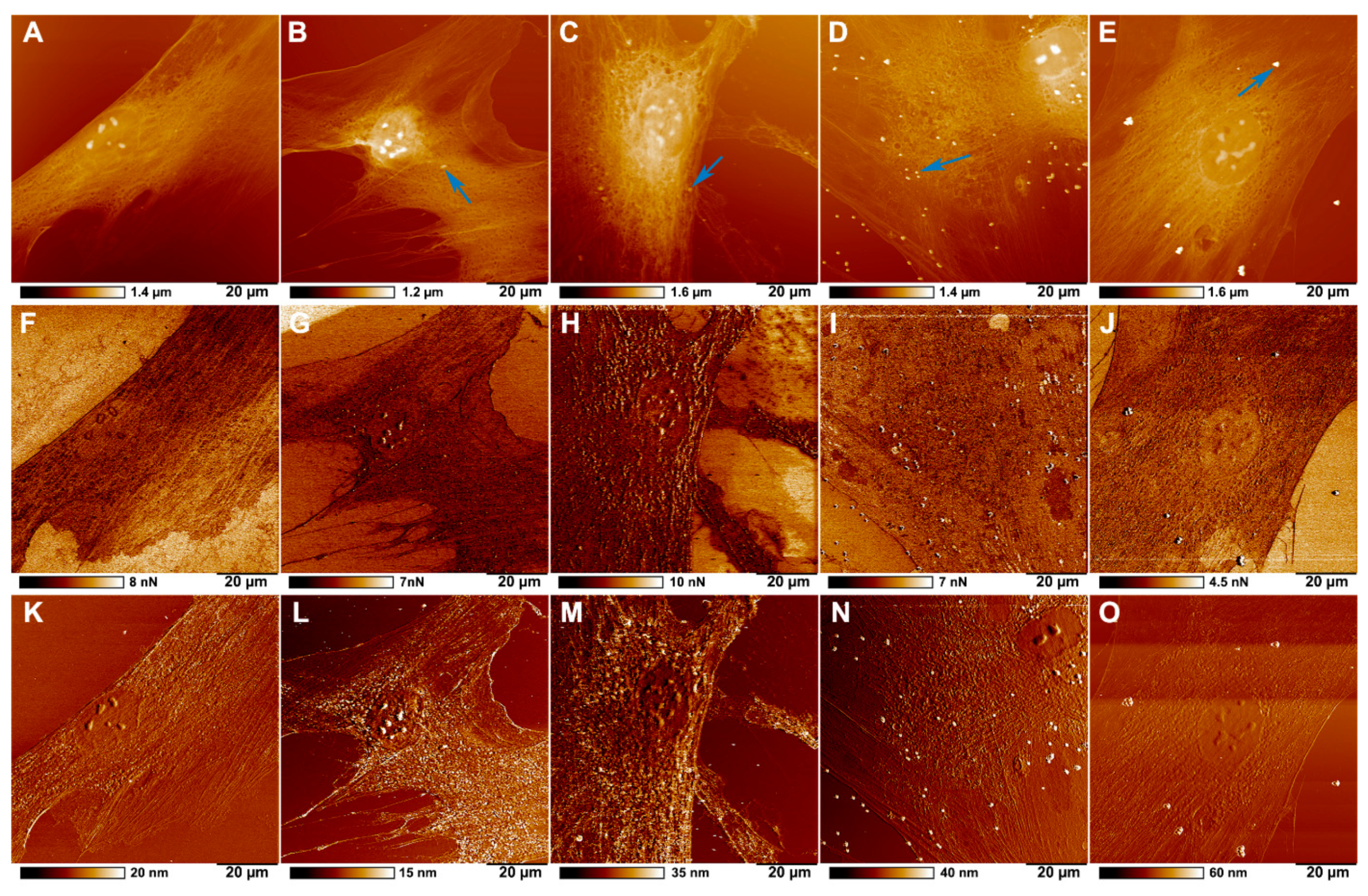
2.3. Microplastics Characterisation and Uptake Imaging with Atomic Force Microscopy
3. Discussion
4. Materials and Methods
4.1. Fibroblasts Cell Culture
4.2. Fibroblasts Viability Evaluation
4.3. Hyperspectral Enhanced Dark-Field Microscopy
4.4. Fluorescence Microscopy
4.5. Atomic Force Microscopy
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Patil, S.M.; Rane, N.R.; Bankole, P.O.; Krishnaiah, P.; Ahn, Y.; Park, Y.-K.; Yadav, K.K.; Amin, M.A.; Jeon, B.-H. An Assessment of Micro- and Nanoplastics in the Biosphere: A Review of Detection, Monitoring, and Remediation Technology. Chem. Eng. J. 2022, 430, 132913. [Google Scholar] [CrossRef]
- Koelmans, A.A.; Mohamed Nor, N.H.; Hermsen, E.; Kooi, M.; Mintenig, S.M.; De France, J. Microplastics in Freshwaters and Drinking Water: Critical Review and Assessment of Data Quality. Water Res. 2019, 155, 410–422. [Google Scholar] [CrossRef]
- Hwang, J.; Choi, D.; Han, S.; Jung, S.Y.; Choi, J.; Hong, J. Potential Toxicity of Polystyrene Microplastic Particles. Sci. Rep. 2020, 10, 7391. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Zhang, Y.; Deng, Y.; Jiang, W.; Zhao, Y.; Geng, J.; Ding, L.; Ren, H. Uptake and Accumulation of Polystyrene Microplastics in Zebrafish (Danio Rerio) and Toxic Effects in Liver. Environ. Sci. Technol. 2016, 50, 4054–4060. [Google Scholar] [CrossRef] [PubMed]
- Stock, V.; Laurisch, C.; Franke, J.; Dönmez, M.H.; Voss, L.; Böhmert, L.; Braeuning, A.; Sieg, H. Uptake and Cellular Effects of PE, PP, PET and PVC Microplastic Particles. Toxicol. Vitr. 2021, 70, 105021. [Google Scholar] [CrossRef] [PubMed]
- Busch, M.; Bredeck, G.; Kämpfer, A.A.M.; Schins, R.P.F. Investigations of Acute Effects of Polystyrene and Polyvinyl Chloride Micro- and Nanoplastics in an Advanced in Vitro Triple Culture Model of the Healthy and Inflamed Intestine. Environ. Res. 2021, 193, 110536. [Google Scholar] [CrossRef] [PubMed]
- Domenech, J.; Hernández, A.; Rubio, L.; Marcos, R.; Cortés, C. Interactions of Polystyrene Nanoplastics with in Vitro Models of the Human Intestinal Barrier. Arch. Toxicol. 2020, 94, 2997–3012. [Google Scholar] [CrossRef]
- Wu, S.; Wu, M.; Tian, D.; Qiu, L.; Li, T. Effects of Polystyrene Microbeads on Cytotoxicity and Transcriptomic Profiles in Human Caco-2 Cells. Environ. Toxicol. 2020, 35, 495–506. [Google Scholar] [CrossRef]
- Dong, C.-D.; Chen, C.-W.; Chen, Y.-C.; Chen, H.-H.; Lee, J.-S.; Lin, C.-H. Polystyrene Microplastic Particles: In Vitro Pulmonary Toxicity Assessment. J. Hazard. Mater. 2020, 385, 121575. [Google Scholar] [CrossRef]
- Cheng, W.; Li, X.; Zhou, Y.; Yu, H.; Xie, Y.; Guo, H.; Wang, H.; Li, Y.; Feng, Y.; Wang, Y. Polystyrene Microplastics Induce Hepatotoxicity and Disrupt Lipid Metabolism in the Liver Organoids. Sci. Total Environ. 2022, 806, 150328. [Google Scholar] [CrossRef]
- Kwon, W.; Kim, D.; Kim, H.-Y.; Jeong, S.W.; Lee, S.-G.; Kim, H.-C.; Lee, Y.-J.; Kwon, M.K.; Hwang, J.-S.; Han, J.E.; et al. Microglial Phagocytosis of Polystyrene Microplastics Results in Immune Alteration and Apoptosis In Vitro and In Vivo. Sci. Total Environ. 2022, 807, 150817. [Google Scholar] [CrossRef] [PubMed]
- Primpke, S.; Dias, P.A.; Gerdts, G. Automated Identification and Quantification of Microfibres and Microplastics. Anal. Methods 2019, 11, 2138–2147. [Google Scholar] [CrossRef]
- Abel, S.M.; Primpke, S.; Int-Veen, I.; Brandt, A.; Gerdts, G. Systematic Identification of Microplastics in Abyssal and Hadal Sediments of the Kuril Kamchatka Trench. Environ. Pollut. 2021, 269, 116095. [Google Scholar] [CrossRef] [PubMed]
- Jeon, S.; Lee, D.-K.; Jeong, J.; Yang, S.I.; Kim, J.-S.; Kim, J.; Cho, W.-S. The Reactive Oxygen Species as Pathogenic Factors of Fragmented Microplastics to Macrophages. Environ. Pollut. 2021, 281, 117006. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Xu, K.; Zhang, B.; Ye, Y.; Zhang, Q.; Jiang, W. Cellular Internalization and Release of Polystyrene Microplastics and Nanoplastics. Sci. Total Environ. 2021, 779, 146523. [Google Scholar] [CrossRef] [PubMed]
- DeLoid, G.M.; Cao, X.; Bitounis, D.; Singh, D.; Llopis, P.M.; Buckley, B.; Demokritou, P. Toxicity, Uptake, and Nuclear Translocation of Ingested Micro-Nanoplastics in an in Vitro Model of the Small Intestinal Epithelium. Food Chem. Toxicol. 2021, 158, 112609. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Ling, Y.; Li, X.; Hu, J.; Cao, C.; He, D. Size-Dependent Cellular Internalization and Effects of Polystyrene Microplastics in Microalgae P. Helgolandica Var. Tsingtaoensis and S. Quadricauda. J. Hazard. Mater. 2020, 399, 123092. [Google Scholar] [CrossRef]
- Deville, S.; Penjweini, R.; Smisdom, N.; Notelaers, K.; Nelissen, I.; Hooyberghs, J.; Ameloot, M. Intracellular Dynamics and Fate of Polystyrene Nanoparticles in A549 Lung Epithelial Cells Monitored by Image (Cross-) Correlation Spectroscopy and Single Particle Tracking. Biochim. Biophys. Acta-Mol. Cell Res. 2015, 1853, 2411–2419. [Google Scholar] [CrossRef] [Green Version]
- Piarulli, S.; Sciutto, G.; Oliveri, P.; Malegori, C.; Prati, S.; Mazzeo, R.; Airoldi, L. Rapid and Direct Detection of Small Microplastics in Aquatic Samples by a New near Infrared Hyperspectral Imaging (NIR-HSI) Method. Chemosphere 2020, 260, 127655. [Google Scholar] [CrossRef]
- Zhu, C.; Kanaya, Y.; Nakajima, R.; Tsuchiya, M.; Nomaki, H.; Kitahashi, T.; Fujikura, K. Characterization of Microplastics on Filter Substrates Based on Hyperspectral Imaging: Laboratory Assessments. Environ. Pollut. 2020, 263, 114296. [Google Scholar] [CrossRef]
- Lv, L.; He, L.; Jiang, S.; Chen, J.; Zhou, C.; Qu, J.; Lu, Y.; Hong, P.; Sun, S.; Li, C. In Situ Surface-Enhanced Raman Spectroscopy for Detecting Microplastics and Nanoplastics in Aquatic Environments. Sci. Total Environ. 2020, 728, 138449. [Google Scholar] [CrossRef]
- Nigamatzyanova, L.; Fakhrullin, R. Dark-Field Hyperspectral Microscopy for Label-Free Microplastics and Nanoplastics Detection and Identification in Vivo: A Caenorhabditis Elegans Study. Environ. Pollut. 2021, 271, 116337. [Google Scholar] [CrossRef]
- Ishmukhametov, I.; Nigamatzyanova, L.; Fakhrullina, G.; Fakhrullin, R. Label-Free Identification of Microplastics in Human Cells: Dark-Field Microscopy and Deep Learning Study. Anal. Bioanal. Chem. 2021, 414, 1297–1312. [Google Scholar] [CrossRef] [PubMed]
- Akhatova, F.; Fakhrullina, G.; Khakimova, E.; Fakhrullin, R. Atomic Force Microscopy for Imaging and Nanomechanical Characterisation of Live Nematode Epicuticle: A Comparative Caenorhabditis Elegans and Turbatrix Aceti Study. Ultramicroscopy 2018, 194, 40–47. [Google Scholar] [CrossRef]
- Bhagwat, G.; O’Connor, W.; Grainge, I.; Palanisami, T. Understanding the Fundamental Basis for Biofilm Formation on Plastic Surfaces: Role of Conditioning Films. Front. Microbiol. 2021, 12, 687118. [Google Scholar] [CrossRef] [PubMed]
- Tarafdar, A.; Lee, J.-U.; Jeong, J.-E.; Lee, H.; Jung, Y.; Bin Oh, H.; Woo, H.Y.; Kwon, J.-H. Biofilm Development of Bacillus Siamensis ATKU1 on Pristine Short Chain Low-Density Polyethylene: A Case Study on Microbe-Microplastics Interaction. J. Hazard. Mater. 2021, 409, 124516. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wang, Z.; Guan, B. Separation and Identification of Nanoplastics in Tap Water. Environ. Res. 2022, 204, 112134. [Google Scholar] [CrossRef]
- Barlow, D.E.; Biffinger, J.C.; Estrella, L.; Lu, Q.; Hung, C.-S.; Nadeau, L.J.; Crouch, A.L.; Russell, J.N.; Crookes-Goodson, W.J. Edge-Localized Biodeterioration and Secondary Microplastic Formation by Papiliotrema Laurentii Unsaturated Biofilm Cells on Polyurethane Films. Langmuir 2020, 36, 1596–1607. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Fu, J.; Wang, S.; Xiang, Y.; Pan, X. Microplastics Generated under Simulated Fire Scenarios: Characteristics, Antimony Leaching, and Toxicity. Environ. Pollut. 2021, 269, 115905. [Google Scholar] [CrossRef]
- Luo, H.; Xiang, Y.; Li, Y.; Zhao, Y.; Pan, X. Photocatalytic Aging Process of Nano-TiO2 Coated Polypropylene Microplastics: Combining Atomic Force Microscopy and Infrared Spectroscopy (AFM-IR) for Nanoscale Chemical Characterization. J. Hazard. Mater. 2021, 404, 124159. [Google Scholar] [CrossRef]
- Choi, D.; Hwang, J.; Bang, J.; Han, S.; Kim, T.; Oh, Y.; Hwang, Y.; Choi, J.; Hong, J. In Vitro Toxicity from a Physical Perspective of Polyethylene Microplastics Based on Statistical Curvature Change Analysis. Sci. Total Environ. 2021, 752, 142242. [Google Scholar] [CrossRef]
- Han, S.-W.; Choi, J.; Ryu, K.-Y. Stress Response of Mouse Embryonic Fibroblasts Exposed to Polystyrene Nanoplastics. Int. J. Mol. Sci. 2021, 22, 2094. [Google Scholar] [CrossRef] [PubMed]
- Lotfi, C.F.P.; Machado-Santelli, G.M. Comparative analysis of colchicine induced micronuclei in different cell types in vitro. Mutat. Res. Mol. Mech. Mutagen. 1996, 349, 77–83. [Google Scholar] [CrossRef]
- Rozhina, E.; Panchal, A.; Akhatova, F.; Lvov, Y.; Fakhrullin, R. Cytocompatibility and Cellular Uptake of Alkylsilane-Modified Hydrophobic Halloysite Nanotubes. Appl. Clay Sci. 2020, 185, 105371. [Google Scholar] [CrossRef]
- Kuku, G.; Saricam, M.; Akhatova, F.; Danilushkina, A.; Fakhrullin, R.; Culha, M. Surface-Enhanced Raman Scattering to Evaluate Nanomaterial Cytotoxicity on Living Cells. Anal. Chem. 2016, 88, 9813–9820. [Google Scholar] [CrossRef]
- Dzamukova, M.R.; Naumenko, E.A.; Rozhina, E.V.; Trifonov, A.A.; Fakhrullin, R.F. Cell Surface Engineering with Polyelectrolyte-Stabilized Magnetic Nanoparticles: A Facile Approach for Fabrication of Artificial Multicellular Tissue-Mimicking Clusters. Nano Res. 2015, 8, 2515–2532. [Google Scholar] [CrossRef] [Green Version]
- Rozhina, E.; Batasheva, S.; Miftakhova, R.; Yan, X.; Vikulina, A.; Volodkin, D.; Fakhrullin, R. Comparative Cytotoxicity of Kaolinite, Halloysite, Multiwalled Carbon Nanotubes and Graphene Oxide. Appl. Clay Sci. 2021, 205, 106041. [Google Scholar] [CrossRef]
- Fakhrullin, R.; Nigamatzyanova, L.; Fakhrullina, G. Dark-Field/Hyperspectral Microscopy for Detecting Nanoscale Particles in Environmental Nanotoxicology Research. Sci. Total Environ. 2021, 772, 145478. [Google Scholar] [CrossRef]
- Akhatova, F.; Danilushkina, A.; Kuku, G.; Saricam, M.; Culha, M.; Fakhrullin, R. Simultaneous Intracellular Detection of Plasmonic and Non-Plasmonic Nanoparticles Using Dark-Field Hyperspectral Microscopy. Bull. Chem. Soc. Jpn. 2018, 91, 1640–1645. [Google Scholar] [CrossRef]
- Liu, Y.; Naumenko, E.; Akhatova, F.; Zou, Q.; Fakhrullin, R.; Yan, X. Self-Assembled Peptide Nanoparticles for Enhanced Dark-Field Hyperspectral Imaging at the Cellular and Invertebrate Level. Chem. Eng. J. 2021, 424, 130348. [Google Scholar] [CrossRef]
- Guryanov, I.; Naumenko, E.; Akhatova, F.; Lazzara, G.; Cavallaro, G.; Nigamatzyanova, L.; Fakhrullin, R. Selective Cytotoxic Activity of Prodigiosin@halloysite Nanoformulation. Front. Bioeng. Biotechnol. 2020, 8, 424. [Google Scholar] [CrossRef] [PubMed]
- Cavallaro, G.; Milioto, S.; Konnova, S.; Fakhrullina, G.; Akhatova, F.; Lazzara, G.; Fakhrullin, R.; Lvov, Y. Halloysite/Keratin Nanocomposite for Human Hair Photoprotection Coating. ACS Appl. Mater. Interfaces 2020, 12, 24348–24362. [Google Scholar] [CrossRef]
- Xu, K.; Sun, W.; Shao, Y.; Wei, F.; Zhang, X.; Wang, W.; Li, P. Recent Development of PeakForce Tapping Mode Atomic Force Microscopy and its Applications on Nanoscience. Nanotechnol. Rev. 2018, 7, 605–621. [Google Scholar] [CrossRef]
- Rahman, N.; Scott, F.H.; Lvov, Y.; Stavitskaya, A.; Akhatova, F.; Konnova, S.; Fakhrullina, G.; Fakhrullin, R. Clay Nanotube Immobilization on Animal Hair for Sustained Anti-Lice Protection. Pharmaceutics 2021, 13, 1477. [Google Scholar] [CrossRef] [PubMed]

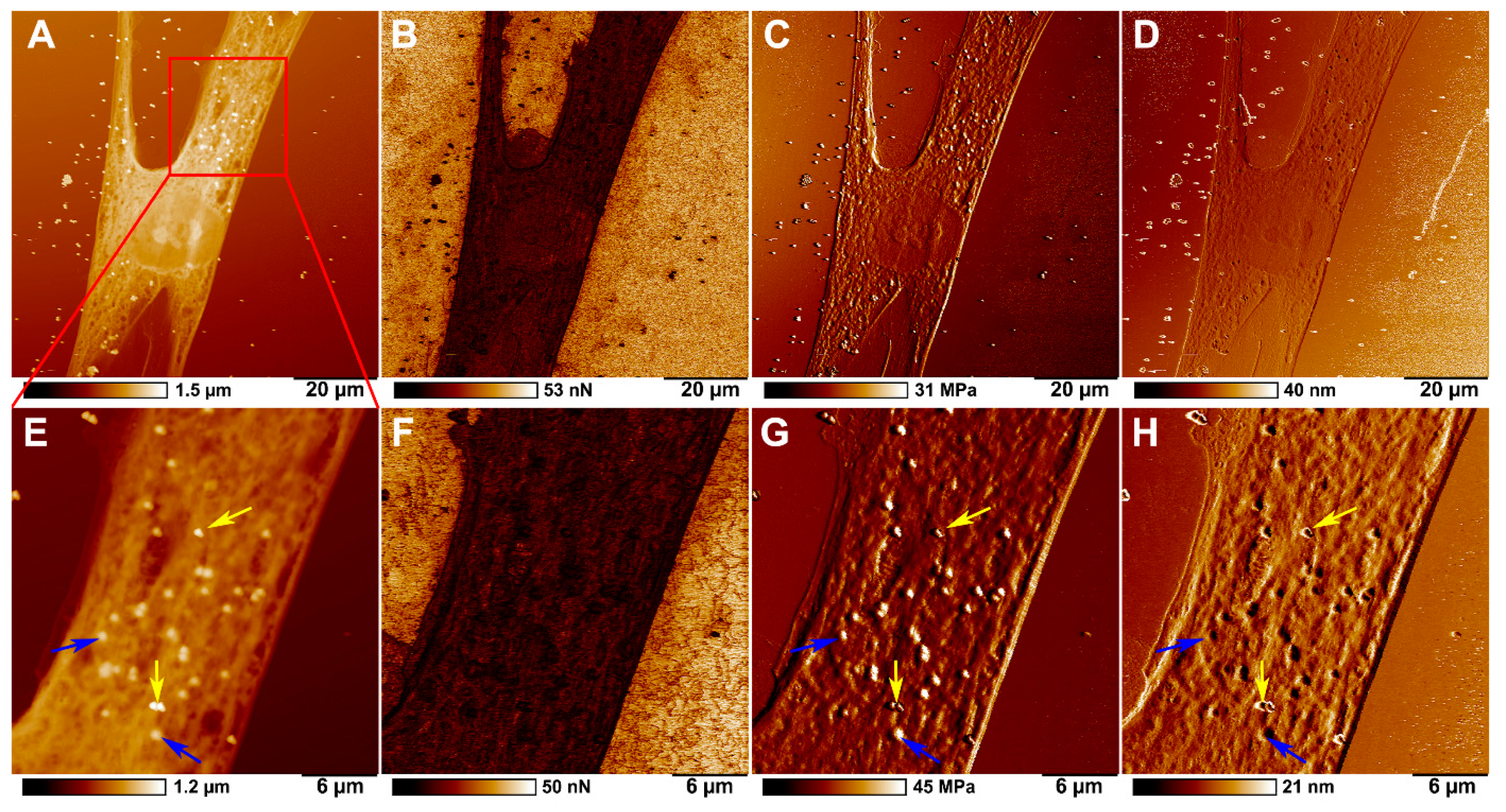
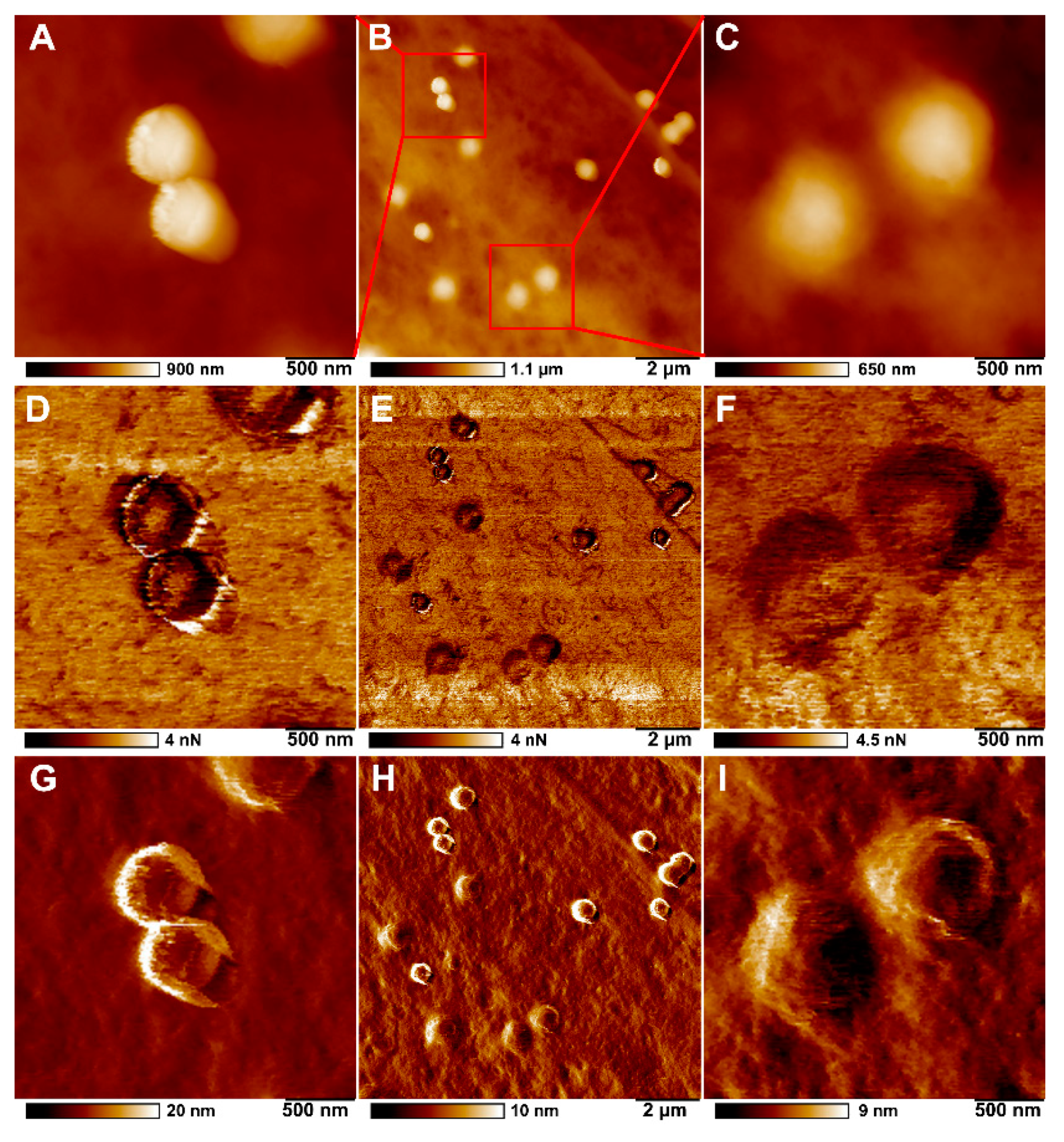
| Adhesion (nN) | DMT Modulus (MPa) | Deformation (nm) | |
|---|---|---|---|
| Cells | 3.2 ± 0.2 | 120.3 ± 19.5 | 7 ± 0.2 *** |
| 500 nm PS on cells | 2.3 ± 0.1 *** | 135.2 ± 19.7 | 10 ± 0.5 *** |
| 500 nm PS in cells | 2.6 ± 0.3 *** | 135.7 ± 30.9 | 8 ± 0.4 *** |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Akhatova, F.; Ishmukhametov, I.; Fakhrullina, G.; Fakhrullin, R. Nanomechanical Atomic Force Microscopy to Probe Cellular Microplastics Uptake and Distribution. Int. J. Mol. Sci. 2022, 23, 806. https://doi.org/10.3390/ijms23020806
Akhatova F, Ishmukhametov I, Fakhrullina G, Fakhrullin R. Nanomechanical Atomic Force Microscopy to Probe Cellular Microplastics Uptake and Distribution. International Journal of Molecular Sciences. 2022; 23(2):806. https://doi.org/10.3390/ijms23020806
Chicago/Turabian StyleAkhatova, Farida, Ilnur Ishmukhametov, Gölnur Fakhrullina, and Rawil Fakhrullin. 2022. "Nanomechanical Atomic Force Microscopy to Probe Cellular Microplastics Uptake and Distribution" International Journal of Molecular Sciences 23, no. 2: 806. https://doi.org/10.3390/ijms23020806
APA StyleAkhatova, F., Ishmukhametov, I., Fakhrullina, G., & Fakhrullin, R. (2022). Nanomechanical Atomic Force Microscopy to Probe Cellular Microplastics Uptake and Distribution. International Journal of Molecular Sciences, 23(2), 806. https://doi.org/10.3390/ijms23020806







