Generation of Flag/DYKDDDDK Epitope Tag Knock-In Mice Using i-GONAD Enables Detection of Endogenous CaMKIIα and β Proteins
Abstract
1. Introduction
2. Results
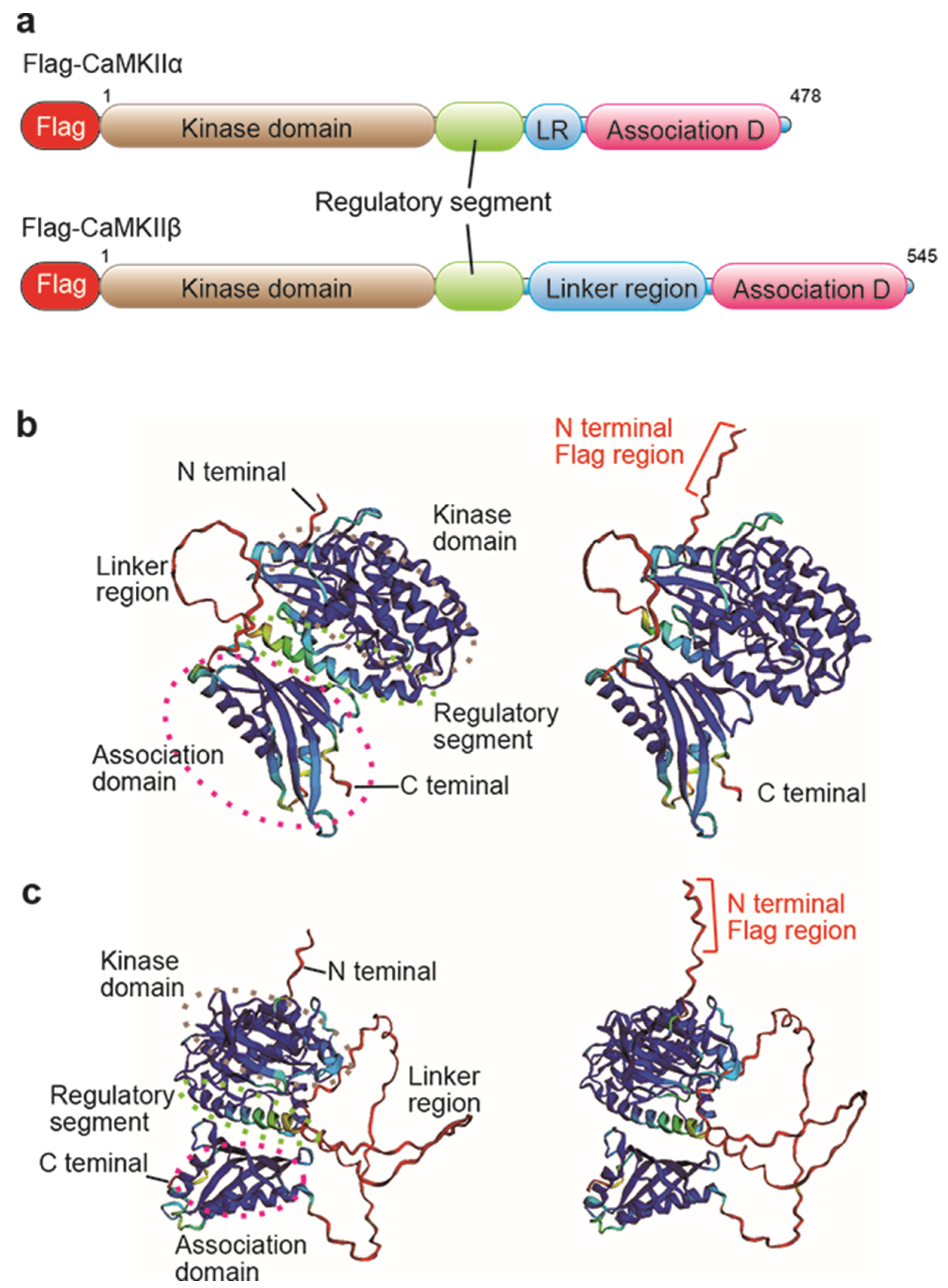
2.1. Investigation of Flag/DYKDDDDK Epitope Tag Position by AlphaFold2 Protein Structure Database
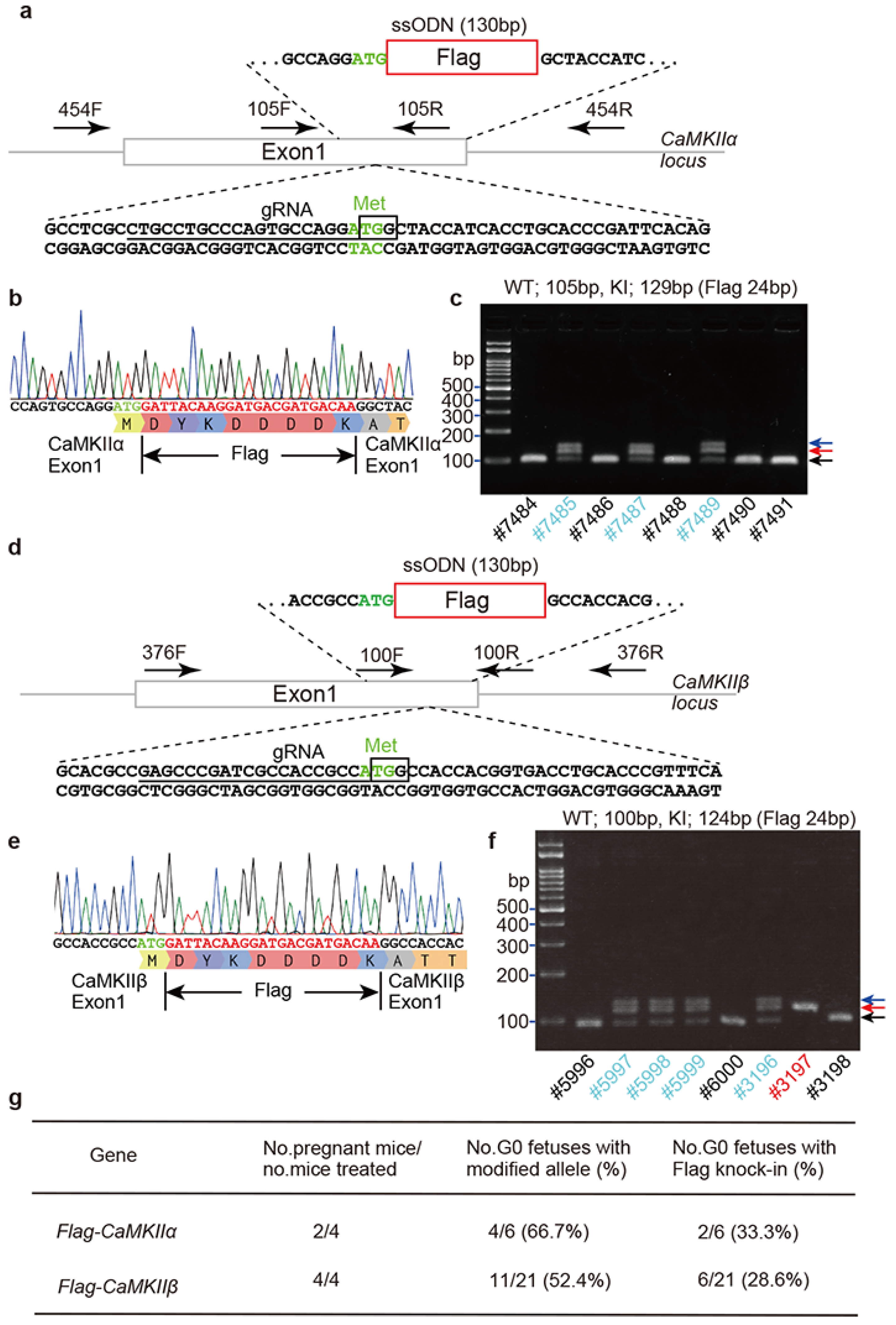
2.2. Flag Knock-In Mouse Generation Using i-GONAD Method
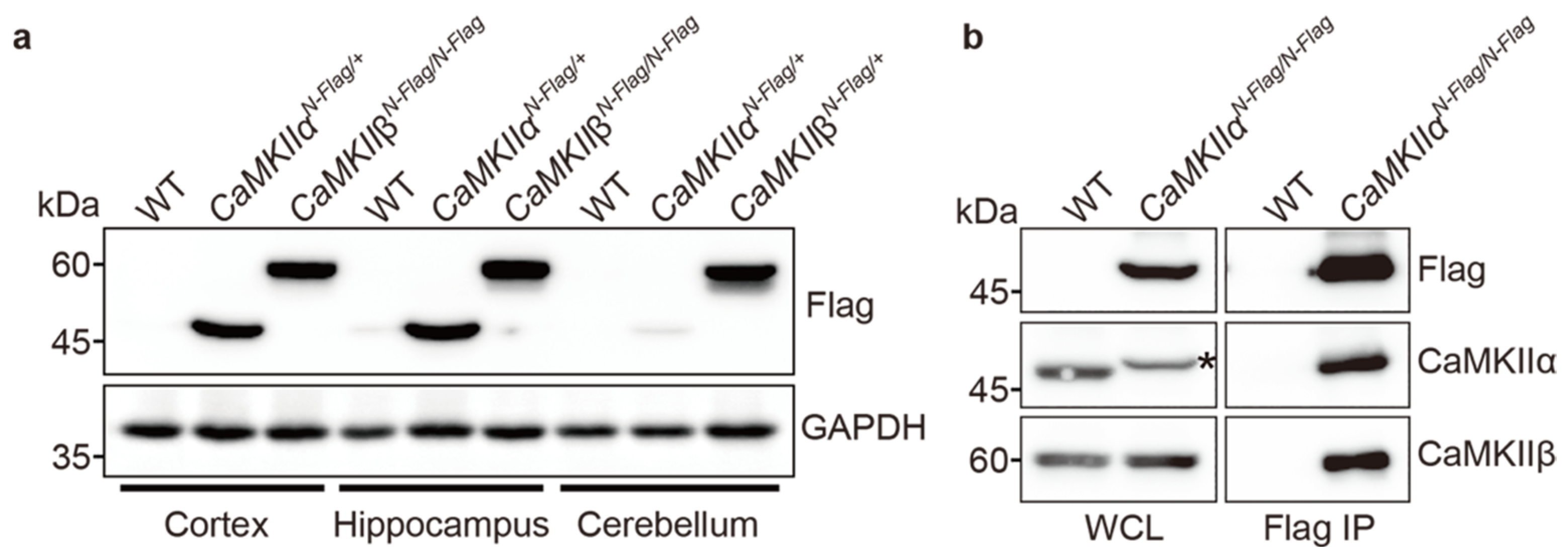
2.3. Biochemical Assay Using CaMKIIαFla and CaMKIIβFlag Mice
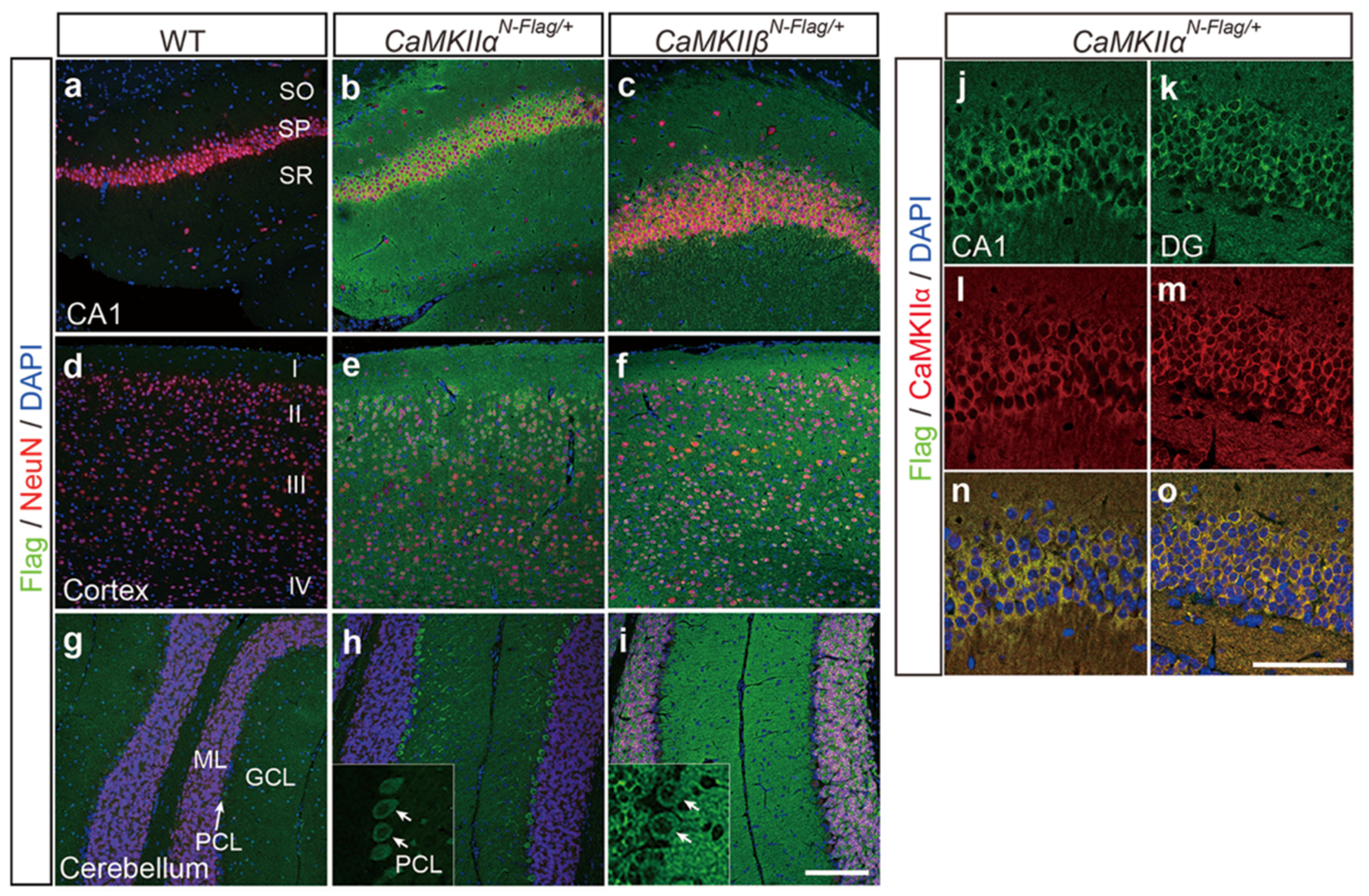
2.4. Immunostaining Using CaMKIIαN-Flag and CaMKIIβN-Flag Mice
3. Discussion
4. Materials and Methods
4.1. AlphaFold2 Analysis for Prediction of Flag Epitope Tag Position
4.2. Flag Knock-In Mouse Generation Using i-GONAD Method
4.3. Genotyping of Flag-CaMKIIα and Flag-CaMKIIβ
4.4. Immunoblotting and Immunoprecipitaion
4.5. Immunohistochemistry
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Li, H.; Haurigot, V.; Doyon, Y.; Li, T.; Wong, S.Y.; Bhagwat, A.S.; Malani, N.; Anguela, X.M.; Sharma, R.; Ivanciu, L.; et al. In vivo genome editing restores haemostasis in a mouse model of haemophilia. Nature 2011, 475, 217–221. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Hu, Y.C.; Markoulaki, S.; Welstead, G.G.; Cheng, A.W.; Shivalila, C.S.; Pyntikova, T.; Dadon, D.B.; Voytas, D.F.; Bogdanove, A.J.; et al. TALEN-mediated editing of the mouse Y chromosome. Nat. Biotechnol. 2013, 31, 530–532. [Google Scholar] [CrossRef]
- Wang, H.; Yang, H.; Shivalila, C.S.; Dawlaty, M.M.; Cheng, A.W.; Zhang, F.; Jaenisch, R. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 2013, 153, 910–918. [Google Scholar] [CrossRef]
- Yang, H.; Wang, H.; Shivalila, C.S.; Cheng, A.W.; Shi, L.; Jaenisch, R. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 2013, 154, 1370–1379. [Google Scholar] [CrossRef]
- Hashimoto, M.; Takemoto, T. Electroporation enables the efficient mRNA delivery into the mouse zygotes and facilitates CRISPR/Cas9-based genome editing. Sci. Rep. 2015, 5, 11315. [Google Scholar] [CrossRef] [PubMed]
- Aoto, K.; Kato, M.; Akita, T.; Nakashima, M.; Mutoh, H.; Akasaka, N.; Tohyama, J.; Nomura, Y.; Hoshino, K.; Ago, Y.; et al. ATP6V0A1 encoding the a1-subunit of the V0 domain of vacuolar H(+)-ATPases is essential for brain development in humans and mice. Nat. Commun. 2021, 12, 2107. [Google Scholar] [CrossRef]
- Gurumurthy, C.B.; Sato, M.; Nakamura, A.; Inui, M.; Kawano, N.; Islam, M.A.; Ogiwara, S.; Takabayashi, S.; Matsuyama, M.; Nakagawa, S.; et al. Creation of CRISPR-based germline-genome-engineered mice without ex vivo handling of zygotes by i-GONAD. Nat. Protoc. 2019, 14, 2452–2482. [Google Scholar] [CrossRef]
- Ohtsuka, M.; Sato, M.; Miura, H.; Takabayashi, S.; Matsuyama, M.; Koyano, T.; Arifin, N.; Nakamura, S.; Wada, K.; Gurumurthy, C.B. i-GONAD: A robust method for in situ germline genome engineering using CRISPR nucleases. Genome Biol. 2018, 19, 25. [Google Scholar] [CrossRef]
- Takabayashi, S.; Aoshima, T.; Kabashima, K.; Aoto, K.; Ohtsuka, M.; Sato, M. i-GONAD (improved genome-editing via oviductal nucleic acids delivery), a convenient in vivo tool to produce genome-edited rats. Sci. Rep. 2018, 8, 12059. [Google Scholar] [CrossRef]
- Hirose, M.; Honda, A.; Fulka, H.; Tamura-Nakano, M.; Matoba, S.; Tomishima, T.; Mochida, K.; Hasegawa, A.; Nagashima, K.; Inoue, K.; et al. Acrosin is essential for sperm penetration through the zona pellucida in hamsters. Proc. Natl. Acad. Sci. USA 2020, 117, 2513–2518. [Google Scholar] [CrossRef]
- Zhang, J.P.; Li, X.L.; Li, G.H.; Chen, W.; Arakaki, C.; Botimer, G.D.; Baylink, D.; Zhang, L.; Wen, W.; Fu, Y.W.; et al. Efficient precise knockin with a double cut HDR donor after CRISPR/Cas9-mediated double-stranded DNA cleavage. Genome Biol. 2017, 18, 35. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, S.; Amaishi, Y.; Maki, I.; Enoki, T.; Mineno, J. Highly efficient genome editing for single-base substitutions using optimized ssODNs with Cas9-RNPs. Sci. Rep. 2019, 9, 4811. [Google Scholar] [CrossRef] [PubMed]
- Ferrando, R.E.; Newton, K.; Chu, F.; Webster, J.D.; French, D.M. Immunohistochemical detection of FLAG-tagged endogenous proteins in knock-in mice. J. Histochem. Cytochem. 2015, 63, 244–255. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Liang, P.; Ding, C.; Zhang, Z.; Zhou, J.; Xie, X.; Huang, R.; Sun, Y.; Sun, H.; Zhang, J.; et al. Efficient Production of Gene-Modified Mice using Staphylococcus aureus Cas9. Sci. Rep. 2016, 6, 32565. [Google Scholar] [CrossRef] [PubMed]
- Imainatsu, K.; Fujii, W.; Hiramatsu, R.; Miura, K.; Kurohmaru, M.; Kanai, Y. CRISPR/Cas9-mediated knock-in of the murine Y chromosomal Sry gene. J. Reprod. Dev. 2018, 64, 283–287. [Google Scholar] [CrossRef] [PubMed]
- Vyas, P.; Wood, M.B.; Zhang, Y.; Goldring, A.C.; Chakir, F.Z.; Fuchs, P.A.; Hiel, H. Characterization of HA-tagged alpha9 and alpha10 nAChRs in the mouse cochlea. Sci. Rep. 2020, 10, 21814. [Google Scholar] [CrossRef]
- Nieto-Rostro, M.; Ramgoolam, K.; Pratt, W.S.; Kulik, A.; Dolphin, A.C. Ablation of alpha2delta-1 inhibits cell-surface trafficking of endogenous N-type calcium channels in the pain pathway in vivo. Proc. Natl. Acad. Sci. USA 2018, 115, E12043–E12052. [Google Scholar] [CrossRef] [PubMed]
- Nakano, H.; Kawai, S.; Ooki, Y.; Chiba, T.; Ishii, C.; Nozawa, T.; Utsuki, H.; Umemura, M.; Takahashi, S.; Takahashi, Y. Functional validation of epitope-tagged ATF5 knock-in mice generated by improved genome editing of oviductal nucleic acid delivery (i-GONAD). Cell Tissue Res. 2021, 385, 239–249. [Google Scholar] [CrossRef] [PubMed]
- Paul, W.; Frankland, C.O.B.; Ohno, M.; Kirkwood, A.; Silva, A.J. α-CaMKII-dependent plasticity in the cortex is required for permanent memory. Nature 2001, 411, 310–313. [Google Scholar]
- Akita, T.; Aoto, K.; Kato, M.; Shiina, M.; Mutoh, H.; Nakashima, M.; Kuki, I.; Okazaki, S.; Magara, S.; Shiihara, T.; et al. De novo variants in CAMK2A and CAMK2B cause neurodevelopmental disorders. Ann. Clin. Transl. Neurol. 2018, 5, 280–296. [Google Scholar] [CrossRef]
- Mutoh, H.; Aoto, K.; Miyazaki, T.; Fukuda, A.; Saitsu, H. Elucidation of pathological mechanism caused by human disease mutation in CaMKIIβ. J. NeuroSci. Res. 2022, 100, 880–896. [Google Scholar] [CrossRef]
- Chao, L.H.; Stratton, M.M.; Lee, I.H.; Rosenberg, O.S.; Levitz, J.; Mandell, D.J.; Kortemme, T.; Groves, J.T.; Schulman, H.; Kuriyan, J. A mechanism for tunable autoinhibition in the structure of a human Ca2+/calmodulin- dependent kinase II holoenzyme. Cell 2011, 146, 732–745. [Google Scholar] [CrossRef]
- Mikuni, T.; Nishiyama, J.; Sun, Y.; Kamasawa, N.; Yasuda, R. High-Throughput, High-Resolution Mapping of Protein Localization in Mammalian Brain by In Vivo Genome Editing. Cell 2016, 165, 1803–1817. [Google Scholar] [CrossRef]
- Tunyasuvunakool, K.; Adler, J.; Wu, Z.; Green, T.; Zielinski, M.; Zidek, A.; Bridgland, A.; Cowie, A.; Meyer, C.; Laydon, A.; et al. Highly accurate protein structure prediction for the human proteome. Nature 2021, 596, 590–596. [Google Scholar] [CrossRef]
- Mirdita, M.; Schütze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: Making protein folding accessible to all. Nat. Methods 2022, 19, 679–682. [Google Scholar] [CrossRef]
- Fritzwanker, S.; Mouledous, L.; Mollereau, C.; Froment, C.; Burlet-Schiltz, O.; Effah, F.; Bailey, A.; Spetea, M.; Reinscheid, R.K.; Schulz, S.; et al. HA-MOP knockin mice express the canonical micro-opioid receptor but lack detectable splice variants. Commun. Biol. 2021, 4, 1070. [Google Scholar] [CrossRef]
- Chamberlain, C.E.; Jeong, J.; Guo, C.; Allen, B.L.; McMahon, A.P. Notochord-derived Shh concentrates in close association with the apically positioned basal body in neural target cells and forms a dynamic gradient during neural patterning. Development 2008, 135, 1097–1106. [Google Scholar] [CrossRef]
- Sasaki, F.; Okuno, T.; Saeki, K.; Min, L.; Onohara, N.; Kato, H.; Shimizu, T.; Yokomizo, T. A high-affinity monoclonal antibody against the FLAG tag useful for G-protein-coupled receptor study. Anal. Biochem. 2012, 425, 157–165. [Google Scholar] [CrossRef]
- Yamagata, Y.; Kobayashi, S.; Umeda, T.; Inoue, A.; Sakagami, H.; Fukaya, M.; Watanabe, M.; Hatanaka, N.; Totsuka, M.; Yagi, T.; et al. Kinase-dead knock-in mouse reveals an essential role of kinase activity of Ca2+/calmodulin-dependent protein kinase IIalpha in dendritic spine enlargement, long-term potentiation, and learning. J. NeuroSci. 2009, 29, 7607–7618. [Google Scholar] [CrossRef]
- Carme Solà, J.M.T.; Serratosa, J. Decreased Expression of Calmodulin Kinase II and Calcineurin Messenger RNAs in the Mouse Hippocampus After Kainic Acid-Induced Seizures. J. Neurochem. 1998, 70, 1600–1608. [Google Scholar] [CrossRef] [PubMed]
- van Breugel, M.; Rosa, E.S.I.; Andreeva, A. Structural validation and assessment of AlphaFold2 predictions for centrosomal and centriolar proteins and their complexes. Commun. Biol. 2022, 5, 312. [Google Scholar] [CrossRef]
- Alerasool, N.; Leng, H.; Lin, Z.Y.; Gingras, A.C.; Taipale, M. Identification and functional characterization of transcriptional activators in human cells. Mol. Cell 2022, 82, 677–695. [Google Scholar] [CrossRef]
- Schmidt, A.; Röner, S.; Mai, K.; Klinkhammer, H.; Kircher, M.; Ludwig, K.U. Predicting the pathogenicity of missense variants using features derived from AlphaFold2. bioRxiv 2022. [Google Scholar] [CrossRef]
- Diwan, G.D.; Gonzalez-Sanchez, J.C.; Apic, G.; Russell, R.B. Next Generation Protein Structure Predictions and Genetic Variant Interpretation. J. Mol. Biol. 2021, 433, 167180. [Google Scholar] [CrossRef]
- McBride, J.M.; Polev, K.; Reinharz, V.; Grzybowski, B.A.; Tlusty, T. AlphaFold2 can predict structural and phenotypic effects of single mutations. arXiv 2022, arXiv:2204.06860. [Google Scholar]
- Li, K.; Wang, G.; Andersen, T.; Zhou, P.; Pu, W.T. Optimization of genome engineering approaches with the CRISPR/Cas9 system. PLoS ONE 2014, 9, e105779. [Google Scholar] [CrossRef] [PubMed]
- Han, J.P.; Chang, Y.J.; Song, D.W.; Choi, B.S.; Koo, O.J.; Yi, S.Y.; Park, T.S.; Yeom, S.C. High Homology-Directed Repair Using Mitosis Phase and Nucleus Localizing Signal. Int. J. Mol. Sci. 2020, 21, 3747. [Google Scholar] [CrossRef]
- Kurihara, T.; Kouyama-Suzuki, E.; Satoga, M.; Li, X.; Badawi, M.; Baig, D.N.; Yanagawa, T.; Uemura, T.; Mori, T.; Tabuchi, K. DNA repair protein RAD51 enhances the CRISPR/Cas9-mediated knock-in efficiency in brain neurons. Biochem. Biophys. Res. Commun. 2020, 524, 621–628. [Google Scholar] [CrossRef]
- Sato, M.; Miyagasako, R.; Takabayashi, S.; Ohtsuka, M.; Hatada, I.; Horii, T. Sequential i-GONAD: An Improved In Vivo Technique for CRISPR/Cas9-Based Genetic Manipulations in Mice. Cells 2020, 9, 546. [Google Scholar] [CrossRef] [PubMed]
- Mizuno, N.; Mizutani, E.; Sato, H.; Kasai, M.; Ogawa, A.; Suchy, F.; Yamaguchi, T.; Nakauchi, H. Intra-embryo Gene Cassette Knockin by CRISPR/Cas9-Mediated Genome Editing with Adeno-Associated Viral Vector. iScience 2018, 9, 286–297. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Sun, S.; Moonen, D.; Lee, C.; Lee, A.Y.; Schaffer, D.V.; He, L. CRISPR-READI: Efficient Generation of Knockin Mice by CRISPR RNP Electroporation and AAV Donor Infection. Cell Rep. 2019, 27, 3780–3789.e4. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aoto, K.; Takabayashi, S.; Mutoh, H.; Saitsu, H. Generation of Flag/DYKDDDDK Epitope Tag Knock-In Mice Using i-GONAD Enables Detection of Endogenous CaMKIIα and β Proteins. Int. J. Mol. Sci. 2022, 23, 11915. https://doi.org/10.3390/ijms231911915
Aoto K, Takabayashi S, Mutoh H, Saitsu H. Generation of Flag/DYKDDDDK Epitope Tag Knock-In Mice Using i-GONAD Enables Detection of Endogenous CaMKIIα and β Proteins. International Journal of Molecular Sciences. 2022; 23(19):11915. https://doi.org/10.3390/ijms231911915
Chicago/Turabian StyleAoto, Kazushi, Shuji Takabayashi, Hiroki Mutoh, and Hirotomo Saitsu. 2022. "Generation of Flag/DYKDDDDK Epitope Tag Knock-In Mice Using i-GONAD Enables Detection of Endogenous CaMKIIα and β Proteins" International Journal of Molecular Sciences 23, no. 19: 11915. https://doi.org/10.3390/ijms231911915
APA StyleAoto, K., Takabayashi, S., Mutoh, H., & Saitsu, H. (2022). Generation of Flag/DYKDDDDK Epitope Tag Knock-In Mice Using i-GONAD Enables Detection of Endogenous CaMKIIα and β Proteins. International Journal of Molecular Sciences, 23(19), 11915. https://doi.org/10.3390/ijms231911915





