Prognostic and Immunological Role of STK38 across Cancers: Friend or Foe?
Abstract
1. Introduction
2. Results
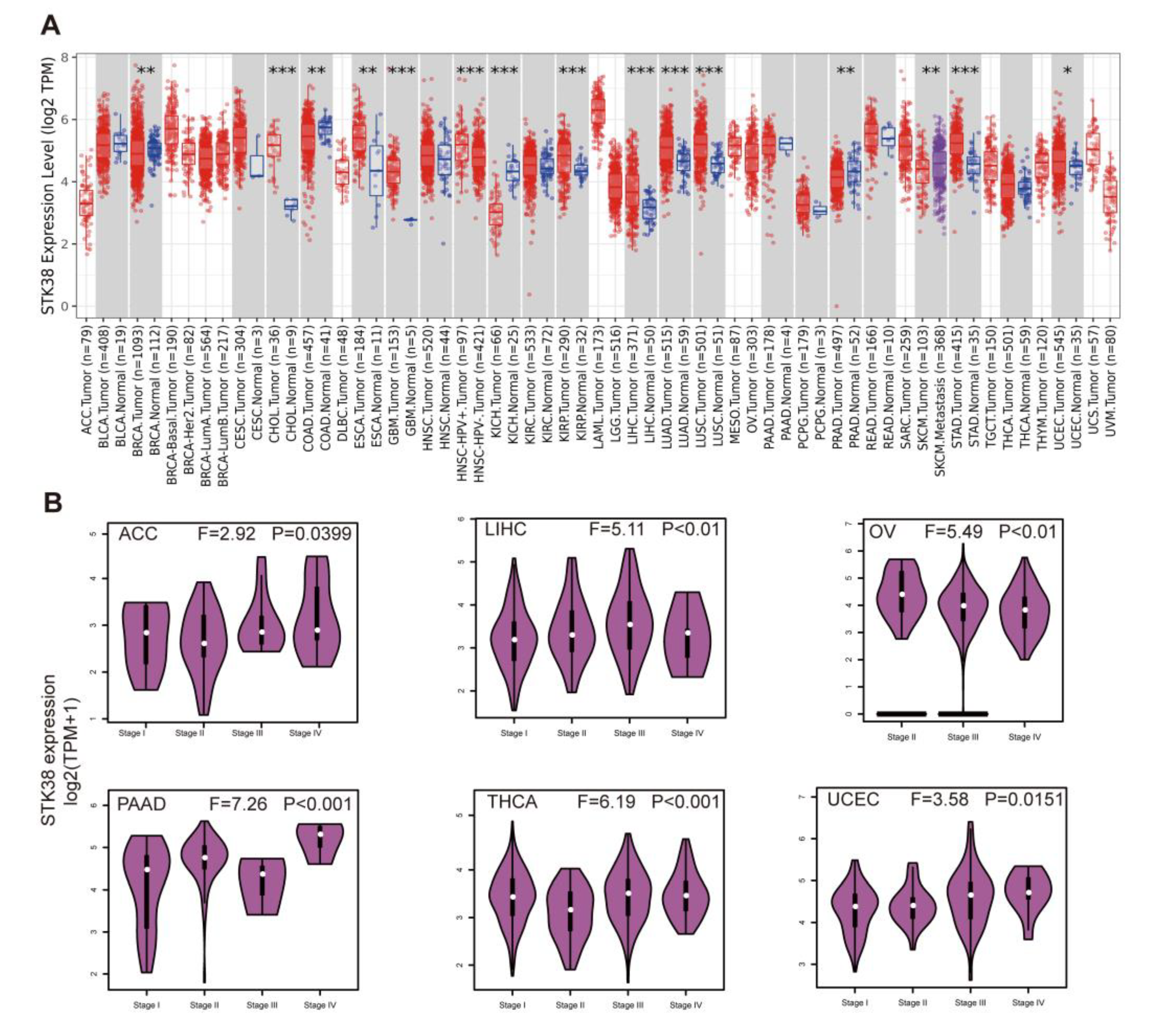
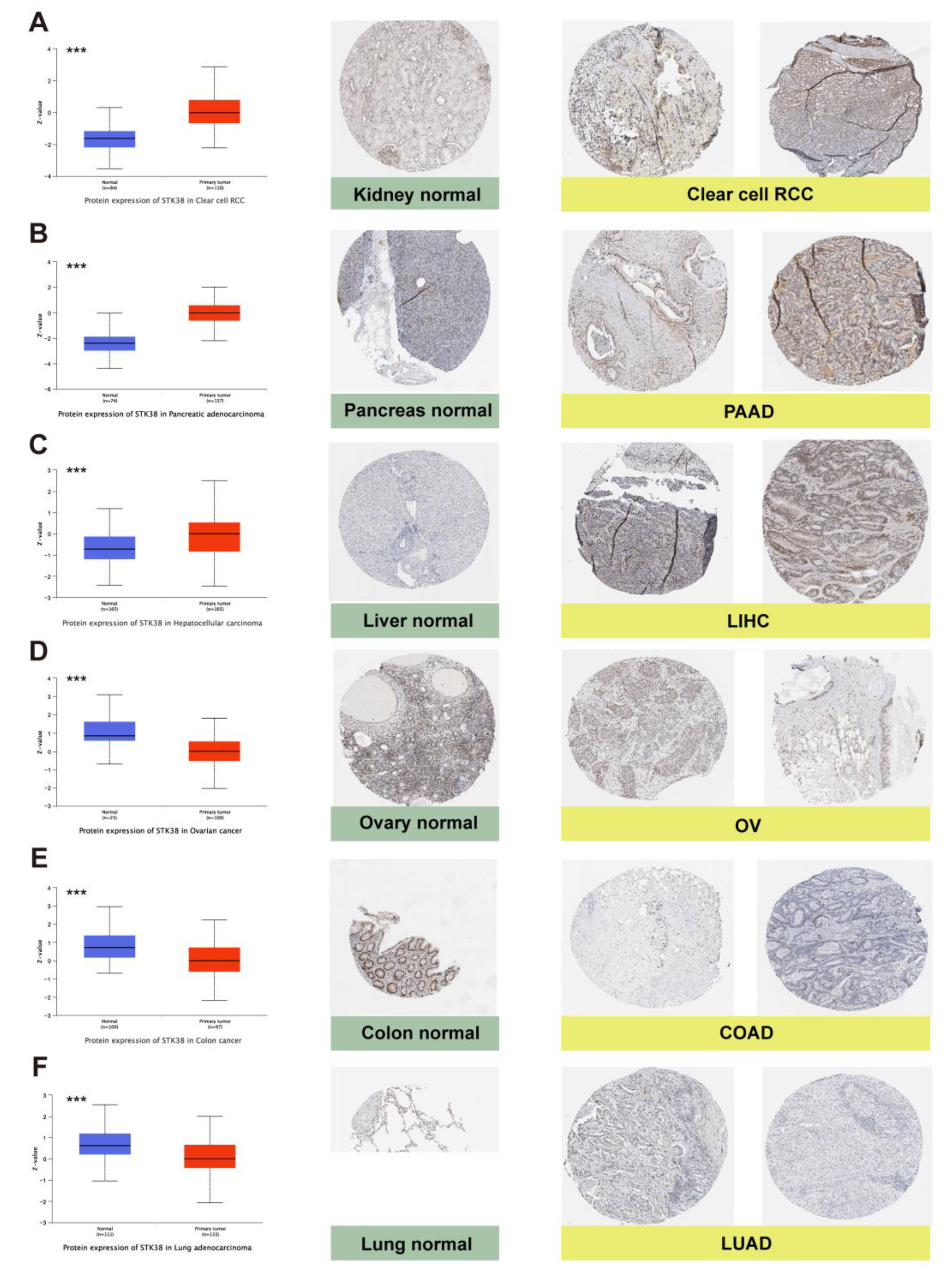
2.1. STK38 Expression Analysis in Human Normal Tissues and Cancers
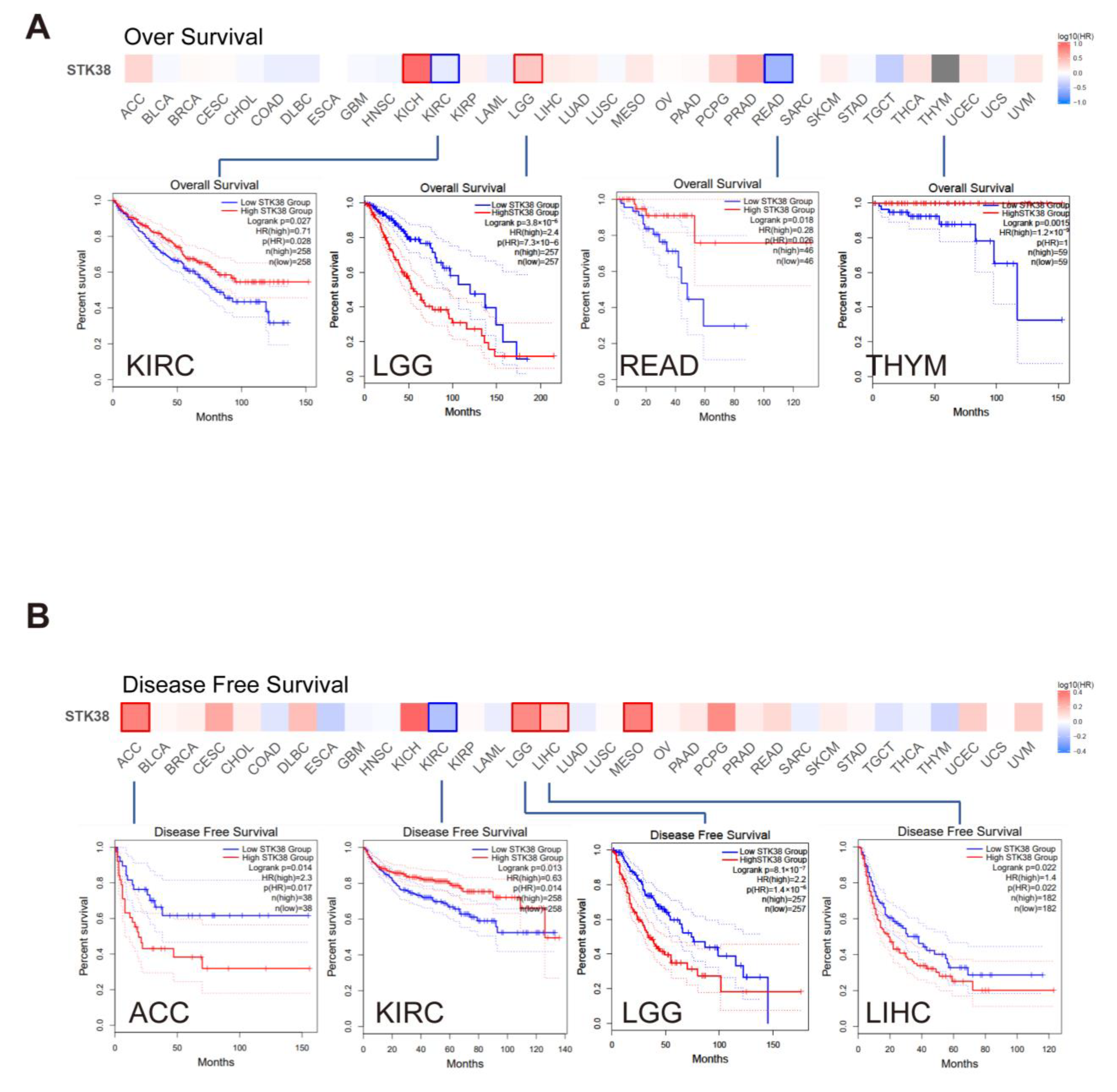
2.2. Survival Prognosis Analysis of STK38 in Cancers
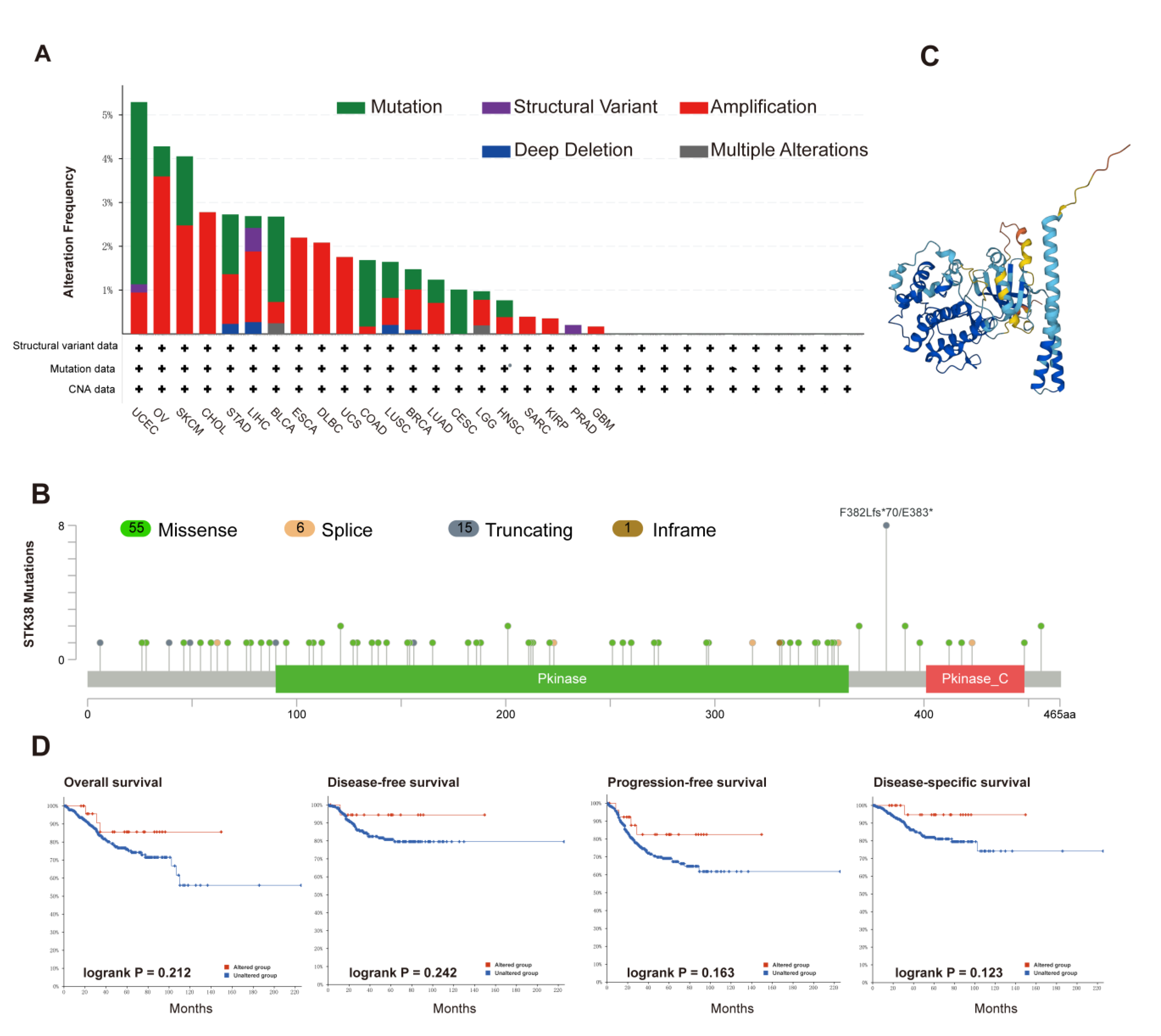
2.3. Genetic Alteration and DNA Methylation Analysis of STK38
2.4. STK38 Is Correlated with the TMB and MSI
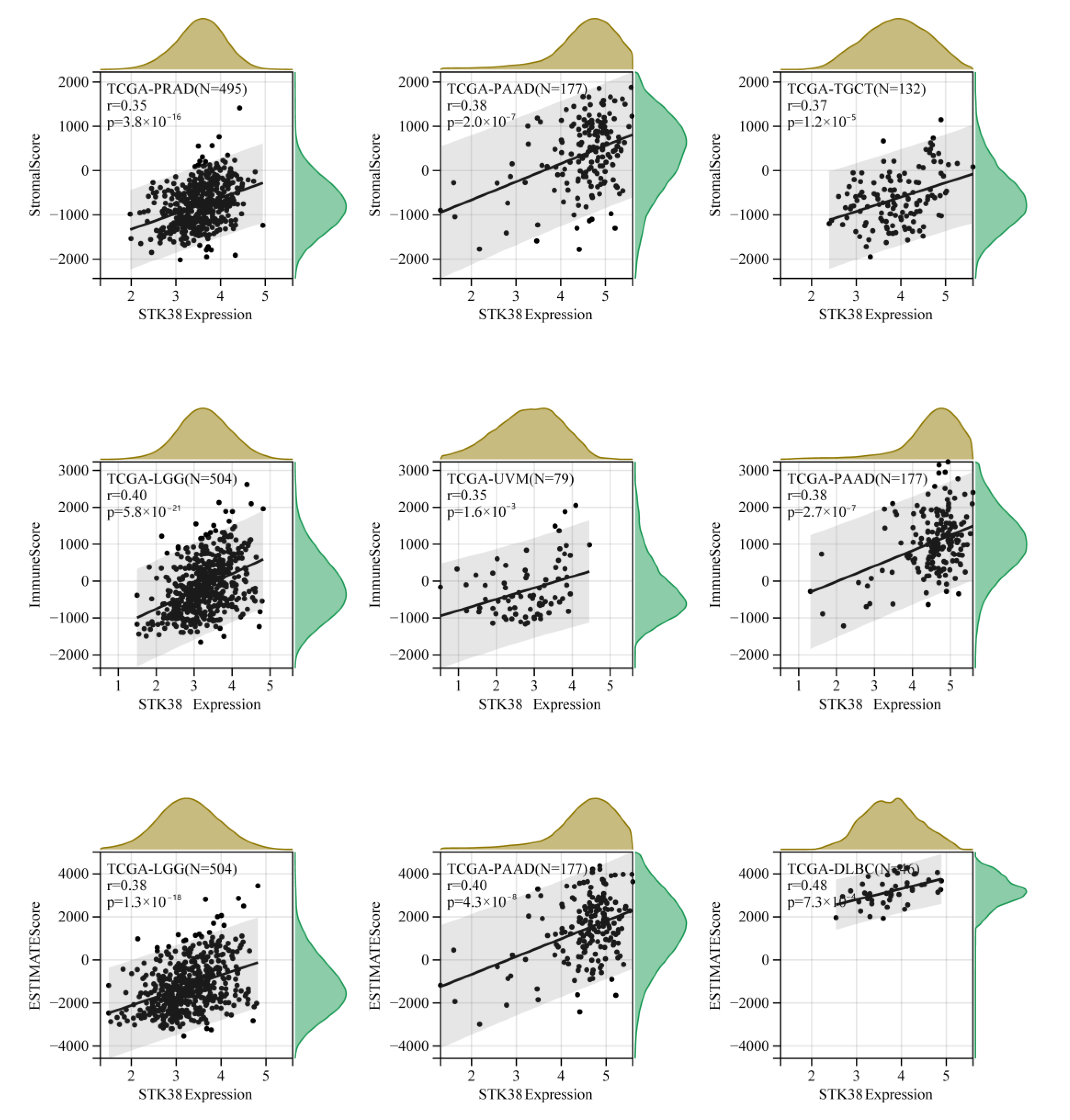
2.5. STK38 Expression Correlates with Tumor Immune Infiltration
2.6. Enrichment Analysis of STK38-Related Partners
3. Discussion
4. Materials and Methods
4.1. Gene Expression Analysis
4.2. Immunohistochemistry (IHC) Staining
4.3. Survival Prognosis Analysis
4.4. Genetic Alteration Analysis
4.5. Immune Infiltration Analysis
4.6. STK38-Related Gene Enrichment Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Ferlay, J.; Soerjomataram, I.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.M.; Forman, D.; Bray, F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 2015, 136, E359–E386. [Google Scholar] [CrossRef] [PubMed]
- Yuan, S.; Norgard, R.J.; Stanger, B.Z. Cellular Plasticity in Cancer. Cancer Discov. 2019, 9, 837–851. [Google Scholar] [CrossRef] [PubMed]
- Pitt, J.M.; Marabelle, A.; Eggermont, A.; Soria, J.C.; Kroemer, G.; Zitvogel, L. Targeting the tumor microenvironment: Removing obstruction to anticancer immune responses and immunotherapy. Ann. Oncol. 2016, 27, 1482–1492. [Google Scholar] [CrossRef]
- Gun, S.Y.; Lee, S.W.L.; Sieow, J.L.; Wong, S.C. Targeting immune cells for cancer therapy. Redox Biol. 2019, 25, 101174. [Google Scholar] [CrossRef] [PubMed]
- Binnewies, M.; Roberts, E.W.; Kersten, K.; Chan, V.; Fearon, D.F.; Merad, M.; Coussens, L.M.; Gabrilovich, D.I.; Ostrand-Rosenberg, S.; Hedrick, C.C.; et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 2018, 24, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Hergovich, A. Hippo Signaling in Mitosis: An Updated View in Light of the MEN Pathway. Methods Mol. Biol. 2017, 1505, 265–277. [Google Scholar] [PubMed]
- Hergovich, A.; Hemmings, B.A. Hippo signalling in the G2/M cell cycle phase: Lessons learned from the yeast MEN and SIN pathways. Semin. Cell. Dev. Biol. 2012, 23, 794–802. [Google Scholar] [CrossRef]
- Sharif, A.A.D.; Hergovich, A. The NDR/LATS protein kinases in immunology and cancer biology. Semin. Cancer Biol. 2018, 48, 104–114. [Google Scholar] [CrossRef] [PubMed]
- Hergovich, A.; Bichsel, S.J.; Hemmings, B.A. Human NDR kinases are rapidly activated by MOB proteins through recruitment to the plasma membrane and phosphorylation. Mol. Cell. Biol. 2005, 25, 8259–8272. [Google Scholar] [CrossRef]
- Cornils, H.; Kohler, R.S.; Hergovich, A.; Hemmings, B.A. Human NDR kinases control G(1)/S cell cycle transition by directly regulating p21 stability. Mol. Cell. Biol. 2011, 31, 1382–1395. [Google Scholar] [CrossRef]
- Du, Y.; Ge, M.-M.; Xue, W.; Yang, Q.-Q.; Wang, S.; Xu, Y.; Wang, H.-L. Chronic Lead Exposure and Mixed Factors of Gender×Age×Brain Regions Interactions on Dendrite Growth, Spine Maturity and NDR Kinase. PLoS ONE 2015, 10, e0138112. [Google Scholar]
- Schmitz-Rohmer, D.; Probst, S.; Yang, Z.-Z.; Laurent, F.; Stadler, M.B.; Zuniga, A.; Zeller, R.; Hynx, D.; Hemmings, B.A.; Hergovich, A. NDR Kinases Are Essential for Somitogenesis and Cardiac Looping during Mouse Embryonic Development. PLoS ONE 2015, 10, e0136566. [Google Scholar] [CrossRef] [PubMed]
- Martin, A.P.; Aushev, V.N.; Zalcman, G.; Camonis, J.H. The STK38-XPO1 axis, a new actor in physiology and cancer. Cell. Mol. Life Sci. 2021, 78, 1943–1955. [Google Scholar] [CrossRef] [PubMed]
- Adeyinka, A.; Emberley, E.; Niu, Y.; Snell, L.; Murphy, L.C.; Sowter, H.; Wykoff, C.C.; Harris, A.L.; Watson, P.H. Analysis of gene expression in ductal carcinoma in situ of the breast. Clin. Cancer Res. 2002, 8, 3788–3795. [Google Scholar]
- Bhattacharjee, A.; Richards, W.G.; Staunton, J.; Li, C.; Monti, S.; Vasa, P.; Ladd, C.; Beheshti, J.; Bueno, R.; Gillette, M.; et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc. Natl. Acad. Sci. USA 2001, 98, 13790–13795. [Google Scholar] [CrossRef]
- Garber, M.E.; Troyanskaya, O.G.; Schluens, K.; Petersen, S.; Thaesler, Z.; Pacyna-Gengelbach, M.; van de Rijn, M.; Rosen, G.D.; Perou, C.M.; Whyte, R.I.; et al. Diversity of gene expression in adenocarcinoma of the lung. Proc. Natl. Acad. Sci. USA 2001, 98, 13784–13789. [Google Scholar] [CrossRef] [PubMed]
- Welsh, J.B.; Zarrinkar, P.P.; Sapinoso, L.M.; Kern, S.G.; Behling, C.A.; Monk, B.J.; Lockhart, D.J.; Burger, R.A.; Hampton, G.M. Analysis of gene expression profiles in normal and neoplastic ovarian tissue samples identifies candidate molecular markers of epithelial ovarian cancer. Proc. Natl. Acad. Sci. USA 2001, 98, 1176–1181. [Google Scholar] [CrossRef]
- Chen, B.; Liu, B.; Yu, T.; Han, Y.-F.; Wu, C.; Wang, Z.-Y. Nuclear Dbf2-related Kinase 1 functions as tumor suppressor in glioblastoma by phosphorylation of Yes-associated protein. Chin. Med. J. (Engl.) 2021, 134, 2054–2065. [Google Scholar] [CrossRef]
- Yue, J.; Sun, H.; Liu, S.; Yu, F.; Wang, S.; Wang, F.; Shen, R.; Zhu, F.; Zhang, L.; Shao, C. Downregulation of NDR1 contributes to metastasis of prostate cancer cells via activating epithelial-mesenchymal transition. Cancer Med. 2018, 7, 3200–3212. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Paredes, M.; Esteller, M. Cancer epigenetics reaches mainstream oncology. Nat. Med. 2011, 17, 330–339. [Google Scholar] [CrossRef] [PubMed]
- Koch, A.; Joosten, S.C.; Feng, Z.; de Ruijter, T.C.; Draht, M.X.; Melotte, V.; Smits, K.M.; Veeck, J.; Herman, J.G.; Van Neste, L.; et al. Analysis of DNA methylation in cancer: Location revisited. Nat. Rev. Clin. Oncol. 2018, 15, 459–466. [Google Scholar] [CrossRef] [PubMed]
- Kwa, M.Q.; Herum, K.M.; Brakebusch, C. Cancer-associated fibroblasts: How do they contribute to metastasis? Clin. Exp. Metastasis 2019, 36, 71–86. [Google Scholar] [CrossRef]
- Chen, X.; Song, E. Turning foes to friends: Targeting cancer-associated fibroblasts. Nat. Rev. Drug Discov. 2019, 18, 99–115. [Google Scholar] [CrossRef]
- Hergovich, A.; Cornils, H.; Hemmings, B.A. Mammalian NDR protein kinases: From regulation to a role in centrosome duplication. Biochim. Biophys. Acta 2008, 1784, 3–15. [Google Scholar] [CrossRef]
- Bisikirska, B.C.; Adam, S.J.; Alvarez, M.J.; Rajbhandari, P.; Cox, R.; Lefebvre, C.; Wang, K.; Rieckhof, G.E.; Felsher, D.W.; Califano, A. STK38 is a critical upstream regulator of MYC’s oncogenic activity in human B-cell lymphoma. Oncogene 2013, 32, 5283–5291. [Google Scholar] [CrossRef]
- Zhang, L.; Tang, F.; Terracciano, L.; Hynx, D.; Kohler, R.; Bichet, S.; Hess, D.; Cron, P.; Hemmings, B.A.; Hergovich, A.; et al. NDR functions as a physiological YAP1 kinase in the intestinal epithelium. Curr. Biol. 2015, 25, 296–305. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Zhang, J.; Guo, W. Tumor purity as a prognosis and immunotherapy relevant feature in gastric cancer. Cancer Med. 2020, 9, 9052–9063. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Ren, R.; Lin, W.; Xiang, L.; Zhao, Z.; Shao, B. Exploring the oncostatin M (OSM) feed-forward signaling of glioblastoma via STAT3 in pan-cancer analysis. Cancer Cell Int. 2021, 21, 565. [Google Scholar] [CrossRef] [PubMed]
- Shiao, S.L.; Chu, G.C.-Y.; Chung, L.W.K. Regulation of prostate cancer progression by the tumor microenvironment. Cancer Lett. 2016, 380, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Tang, F.; Gill, J.; Ficht, X.; Barthlott, T.; Cornils, H.; Schmitz-Rohmer, D.; Hynx, D.; Zhou, D.; Zhang, L.; Xue, G.; et al. The kinases NDR1/2 act downstream of the Hippo homolog MST1 to mediate both egress of thymocytes from the thymus and lymphocyte motility. Sci. Signal. 2015, 8, ra100. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Qin, Q.; Wu, C.; Li, H.; Shou, J.n.; Yang, Y.; Gu, M.; Ma, C.; Lin, W.; Zou, Y.; et al. Downregulated NDR1 protein kinase inhibits innate immune response by initiating an miR146a-STAT1 feedback loop. Nat. Commun. 2018, 9, 2789. [Google Scholar] [CrossRef]
- Galon, J.; Bruni, D. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat. Rev. Drug Discov. 2019, 18, 197–218. [Google Scholar] [CrossRef] [PubMed]
- Tsan, M.-F. Toll-like receptors, inflammation and cancer. Semin. Cancer Biol. 2006, 16, 32–37. [Google Scholar] [CrossRef] [PubMed]
- Rakoff-Nahoum, S.; Medzhitov, R. Toll-like receptors and cancer. Nat. Rev. Cancer 2009, 9, 57–63. [Google Scholar] [CrossRef]
- Wen, M.; Ma, X.; Cheng, H.; Jiang, W.; Xu, X.; Zhang, Y.; Zhang, Y.; Guo, Z.; Yu, Y.; Xu, H.; et al. Stk38 protein kinase preferentially inhibits TLR9-activated inflammatory responses by promoting MEKK2 ubiquitination in macrophages. Nat. Commun. 2015, 6, 7167. [Google Scholar] [CrossRef] [PubMed]
- Nurmik, M.; Ullmann, P.; Rodriguez, F.; Haan, S.; Letellier, E. In search of definitions: Cancer-associated fibroblasts and their markers. Int. J. Cancer 2020, 146, 895–905. [Google Scholar] [CrossRef]
- Snyder, A.; Makarov, V.; Merghoub, T.; Yuan, J.; Zaretsky, J.M.; Desrichard, A.; Walsh, L.A.; Postow, M.A.; Wong, P.; Ho, T.S.; et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 2014, 371, 2189–2199. [Google Scholar] [CrossRef] [PubMed]
- Balar, A.V.; Galsky, M.D.; Rosenberg, J.E.; Powles, T.; Petrylak, D.P.; Bellmunt, J.; Loriot, Y.; Necchi, A.; Hoffman-Censits, J.; Perez-Gracia, J.L.; et al. Atezolizumab as first-line treatment in cisplatin-ineligible patients with locally advanced and metastatic urothelial carcinoma: A single-arm, multicentre, phase 2 trial. Lancet 2017, 389, 67–76. [Google Scholar] [CrossRef]
- Gavrielatou, N.; Doumas, S.; Economopoulou, P.; Foukas, P.G.; Psyrri, A. Biomarkers for immunotherapy response in head and neck cancer. Cancer Treat Rev. 2020, 84, 101977. [Google Scholar] [CrossRef]
- Chen, F.; Chandrashekar, D.S.; Varambally, S.; Creighton, C.J. Pan-cancer molecular subtypes revealed by mass-spectrometry-based proteomic characterization of more than 500 human cancers. Nat. Commun. 2019, 10, 5679. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Kang, B.; Li, C.; Chen, T.; Zhang, Z. GEPIA2: An enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019, 47, W556–W560. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef] [PubMed]
- Knijnenburg, T.A.; Wang, L.; Zimmermann, M.T.; Chambwe, N.; Gao, G.F.; Cherniack, A.D.; Fan, H.; Shen, H.; Way, G.P.; Greene, C.S.; et al. Genomic and Molecular Landscape of DNA Damage Repair Deficiency across The Cancer Genome Atlas. Cell Rep. 2018, 23, 239–254. [Google Scholar] [CrossRef] [PubMed]
- Siemers, N.O.; Holloway, J.L.; Chang, H.; Chasalow, S.D.; Ross-MacDonald, P.B.; Voliva, C.F.; Szustakowski, J.D. Genome-wide association analysis identifies genetic correlates of immune infiltrates in solid tumors. PLoS ONE 2017, 12, e0179726. [Google Scholar] [CrossRef] [PubMed]


| Cancer | Methylation | |
|---|---|---|
| p-Value | Correlation | |
| ACC | 0.082 | 0.197 |
| BLCA | 0.054 | −0.095 |
| BRCA | <0.001 | −0.25 |
| CESC | <0.001 | −0.317 |
| CHOL | 0.438 | −0.133 |
| COAD | <0.001 | −0.479 |
| READ | <0.001 | −0.378 |
| DLBC | 0.627 | 0.072 |
| ESCA | 0.02 | −0.291 |
| GBM | 0.1 | 0.231 |
| HNSC | <0.001 | −0.176 |
| KICH | 0.401 | −0.106 |
| KIRC | 0.011 | −0.142 |
| KIRP | 0.019 | 0.142 |
| LAML | 0.047 | 0.188 |
| LGG | 0.008 | −0.117 |
| LIHC | <0.001 | −0.272 |
| LUAD | 0.042 | −0.096 |
| LUSC | <0.001 | −0.222 |
| MESO | 0.437 | −0.085 |
| OV | 0.396 | −0.393 |
| PAAD | 0.001 | 0.294 |
| PCPG | <0.001 | 0.363 |
| PRAD | <0.001 | 0.216 |
| SARC | 0.005 | 0.175 |
| SKCM | <0.001 | −0.218 |
| STAD | 0.033 | −0.116 |
| TGCT | <0.001 | 0.349 |
| THCA | <0.001 | −0.419 |
| THYM | <0.001 | 0.548 |
| UCEC | 0.571 | −0.027 |
| UCS | 0.001 | −0.423 |
| UVM | 0.032 | −0.241 |
| KIRC | 0.006 | −0.156 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Shi, Z.; Zheng, Z.; Li, J.; Yang, K.; Xu, C.; Liu, Q.; Gong, Z.; Yang, Y.; Zhao, Y.; et al. Prognostic and Immunological Role of STK38 across Cancers: Friend or Foe? Int. J. Mol. Sci. 2022, 23, 11590. https://doi.org/10.3390/ijms231911590
Liu Y, Shi Z, Zheng Z, Li J, Yang K, Xu C, Liu Q, Gong Z, Yang Y, Zhao Y, et al. Prognostic and Immunological Role of STK38 across Cancers: Friend or Foe? International Journal of Molecular Sciences. 2022; 23(19):11590. https://doi.org/10.3390/ijms231911590
Chicago/Turabian StyleLiu, Yankuo, Zhiyuan Shi, Zeyuan Zheng, Jinxin Li, Kunao Yang, Chunlan Xu, Qing Liu, Zhicheng Gong, Yi Yang, Yue Zhao, and et al. 2022. "Prognostic and Immunological Role of STK38 across Cancers: Friend or Foe?" International Journal of Molecular Sciences 23, no. 19: 11590. https://doi.org/10.3390/ijms231911590
APA StyleLiu, Y., Shi, Z., Zheng, Z., Li, J., Yang, K., Xu, C., Liu, Q., Gong, Z., Yang, Y., Zhao, Y., Xuan, Z., Sun, H., & Shao, C. (2022). Prognostic and Immunological Role of STK38 across Cancers: Friend or Foe? International Journal of Molecular Sciences, 23(19), 11590. https://doi.org/10.3390/ijms231911590






