Successful i-GONAD in Mice at Early Zygote Stage through In Vivo Electroporation Three Min after Intraoviductal Instillation of CRISPR-Ribonucleoprotein
Abstract
:1. Introduction
2. Results
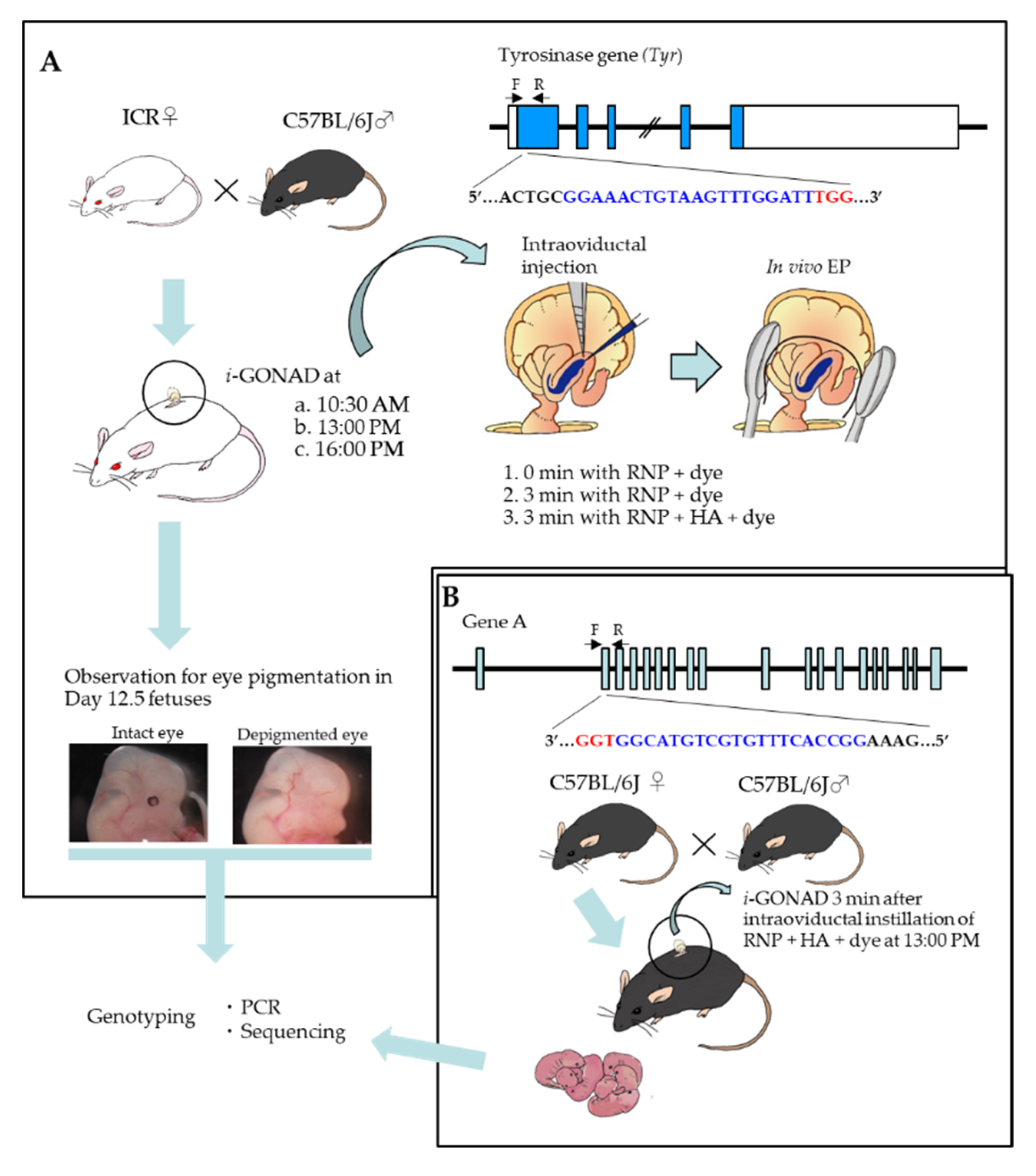
2.1. Three Min Interval before In Vivo EP Is Effective to Induce Indels in Early Zygotes
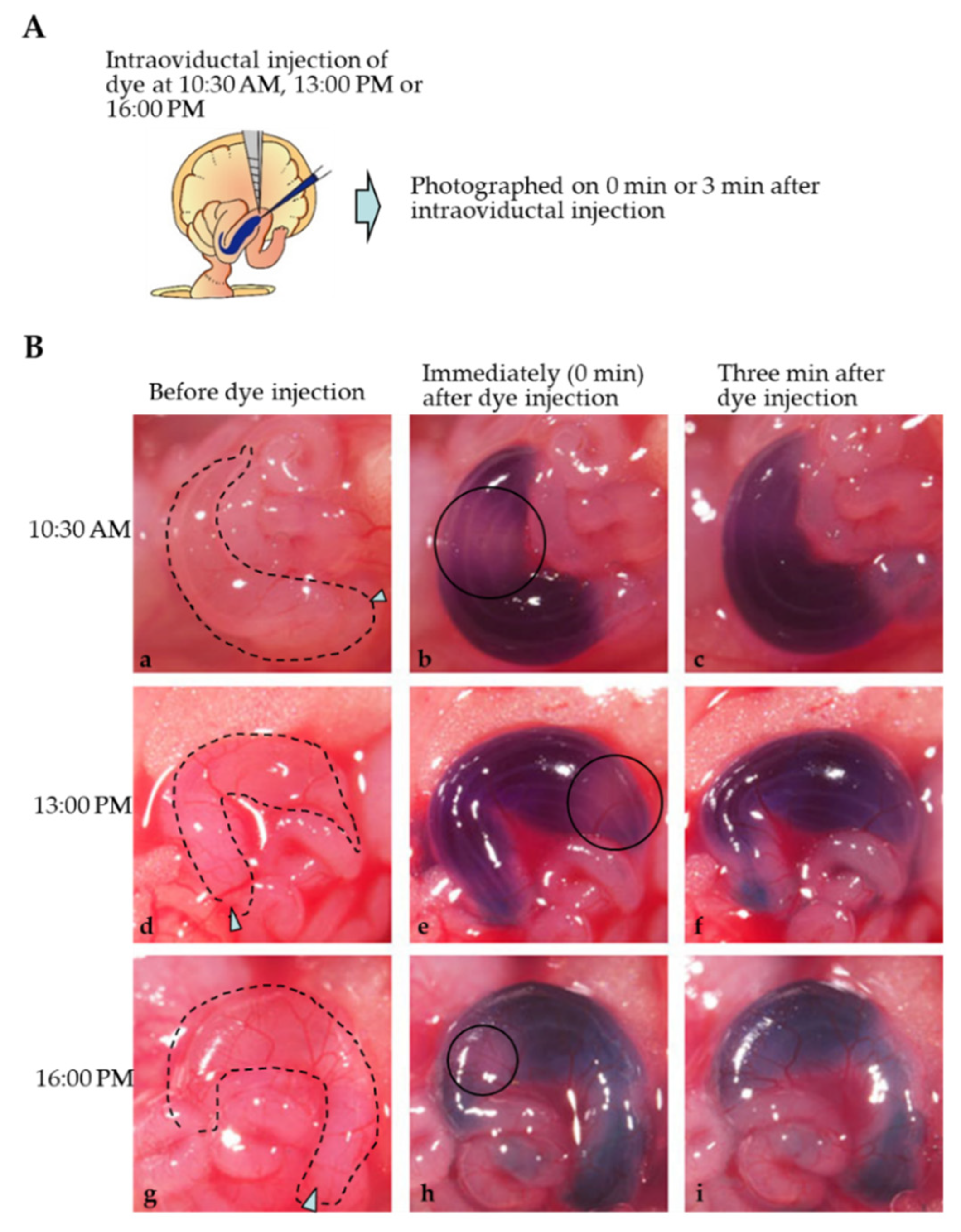
2.2. Mode of Distribution of a Solution Injected into the Oviductal Lumen
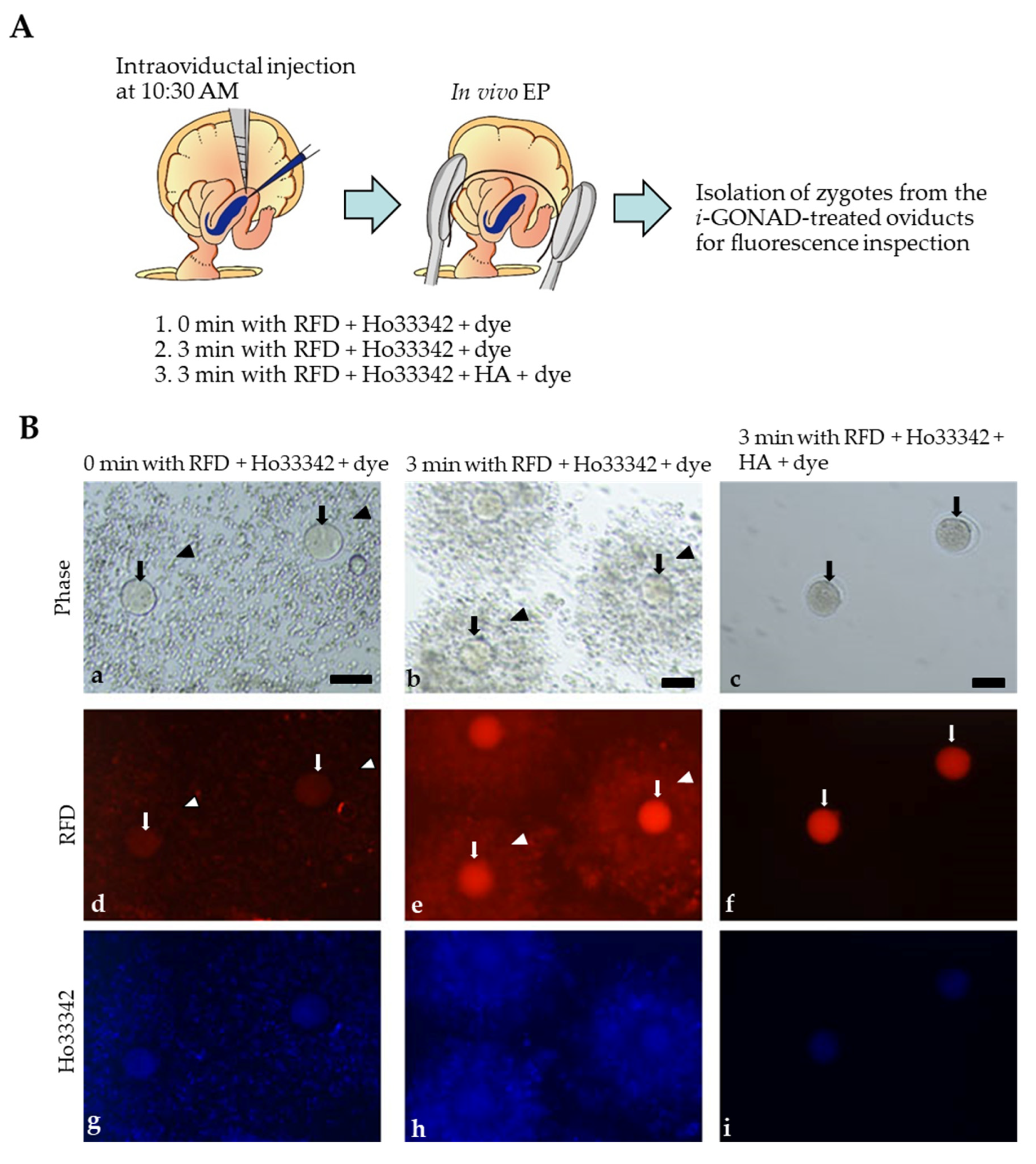
2.3. Visualization of i-GONAD-Mediated Delivery of Fluorescent Reagents to Early Zygotes
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. Mating Protocol
4.3. Preparation of Solutions Used for i-GONAD
4.4. i-GONAD Procedure and Post-i-GONAD Treatment
4.5. Analysis of CRISPR/Cas9-Induced Indels
4.6. Fluorescence Analysis
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CRISPR/Cas9 | clustered regularly interspaced short palindromic repeat-associated protein 9 |
| ET | egg transfer |
| Fgf10 | fibroblast growth factor 10 |
| GM | genetically modified |
| i-GONAD | improved genome editing via oviductal nucleic acid delivery |
| Indel | insertion and deletion mutation |
| HA | hyaluronidase |
| Ho33342 | Hoechst33342 |
| KI | knock-in |
| KO | knock-out |
| Pa.m. | protospacer adjacent motif |
| PCR | Polymerase chain reaction |
| RFD | tetramethylrhodamine-labeled dextran 3 kDa |
| tracrRNA | trans-activating cr RNA |
| Tyr | tyrosinase |
References
- Wang, H.; Yang, H.; Shivalila, C.S.; Dawlaty, M.M.; Cheng, A.W.; Zhang, F.; Jaenisch, R. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 2013, 153, 910–918. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Wang, H.; Shivalila, C.S.; Cheng, A.W.; Shi, L.; Jaenisch, R. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 2013, 154, 1370–1379. [Google Scholar] [CrossRef] [PubMed]
- Mashiko, D.; Fujihara, Y.; Satouh, Y.; Miyata, H.; Isotani, A.; Ikawa, M. Generation of mutant mice by pronuclear injection of circular plasmid expressing Cas9 and single guided RNA. Sci. Rep. 2013, 3, 3355. [Google Scholar] [CrossRef] [PubMed]
- Horii, T.; Arai, Y.; Yamazaki, M.; Morita, S.; Kimura, M.; Itoh, M.; Abe, Y.; Hatada, I.I. Validation of microinjection methods for generating knockout mice by CRISPR/Cas-mediated genome engineering. Sci. Rep. 2014, 4, 4513. [Google Scholar] [CrossRef] [PubMed]
- Yen, S.T.; Zhang, M.; Deng, J.M.; Usman, S.J.; Smith, C.N.; Parker-Thornburg, J.; Swinton, P.G.; Martin, J.F.; Behringer, R.R. Somatic mosaicism and allele complexity induced by CRISPR/Cas9 RNA injections in mouse zygotes. Dev. Biol. 2014, 393, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Ohtsuka, M.; Sato, M.; Miura, H.; Takabayashi, S.; Matsuyama, M.; Koyano, T.; Arifin, N.; Nakamura, S.; Wada, K.; Gurumurthy, C.B. i-GONAD: A robust method for in situ germline genome engineering using CRISPR nucleases. Genome Biol. 2018, 19, 25. [Google Scholar] [CrossRef] [PubMed]
- Hogan, B.; Beddington, R.; Constantini, F.; Lacy, E. (Eds.) Manipulating the Mouse Embryo, 2nd ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1994. [Google Scholar]
- Sato, M.; Akasaka, E.; Saitoh, I.; Ohtsuka, M.; Watanabe, S. In vivo gene transfer in mouse preimplantation embryos after intraoviductal injection of plasmid DNA and subsequent in vivo electroporation. Syst. Biol. Reprod. Med. 2012, 58, 278–287. [Google Scholar] [CrossRef] [PubMed]
- Sato, M. Intraoviductal introduction of plasmid DNA and subsequent electroporation for efficient in vivo gene transfer to murine oviductal epithelium. Mol. Reprod. Dev. 2005, 71, 321–330. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, G.; Gurumurthy, C.B.; Wada, K.; Miura, H.; Sato, M.; Ohtsuka, M. GONAD: Genome-editing via Oviductal Nucleic Acids Delivery system: A novel microinjection independent genome engineering method in mice. Sci. Rep. 2015, 5, 11406. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, T.; Tanaka, S. Improvement of genome editing by electroporation using embryos artificially removed cumulus cells in the oviducts. Biochem. Biophys. Res. Commun. 2020, 527, 1039–1042. [Google Scholar] [CrossRef] [PubMed]
- Gurumurthy, C.B.; Sato, M.; Nakamura, A.; Inui, M.; Kawano, N.; Islam, M.; Ogiwara, S.; Takabayashi, S.; Matsuyama, M.; Nakagawa, S.; et al. Creation of CRISPR-based germline-genome-engineered mice without ex vivo handling of zygotes by i-GONAD. Nat. Protoc. 2019, 14, 2452–2482. [Google Scholar] [CrossRef] [PubMed]

| Target Locus | Treatment | No. Mice Treated | No. Pregnant Mice | No. Pups (A) | No. Indels (B) | % (B/A) |
|---|---|---|---|---|---|---|
| Gene A | 0 min with RNP + dye | 7 | 3 | 13 | 3 | 23 |
| 3 min with RNP + HA + dye | 6 | 3 | 13 | 6 | 46 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Takabayashi, S.; Iijima, K.; Tsujimura, M.; Aoshima, T.; Takagi, H.; Aoto, K.; Sato, M. Successful i-GONAD in Mice at Early Zygote Stage through In Vivo Electroporation Three Min after Intraoviductal Instillation of CRISPR-Ribonucleoprotein. Int. J. Mol. Sci. 2022, 23, 10678. https://doi.org/10.3390/ijms231810678
Takabayashi S, Iijima K, Tsujimura M, Aoshima T, Takagi H, Aoto K, Sato M. Successful i-GONAD in Mice at Early Zygote Stage through In Vivo Electroporation Three Min after Intraoviductal Instillation of CRISPR-Ribonucleoprotein. International Journal of Molecular Sciences. 2022; 23(18):10678. https://doi.org/10.3390/ijms231810678
Chicago/Turabian StyleTakabayashi, Shuji, Kenta Iijima, Masumi Tsujimura, Takuya Aoshima, Hisayoshi Takagi, Kazushi Aoto, and Masahiro Sato. 2022. "Successful i-GONAD in Mice at Early Zygote Stage through In Vivo Electroporation Three Min after Intraoviductal Instillation of CRISPR-Ribonucleoprotein" International Journal of Molecular Sciences 23, no. 18: 10678. https://doi.org/10.3390/ijms231810678
APA StyleTakabayashi, S., Iijima, K., Tsujimura, M., Aoshima, T., Takagi, H., Aoto, K., & Sato, M. (2022). Successful i-GONAD in Mice at Early Zygote Stage through In Vivo Electroporation Three Min after Intraoviductal Instillation of CRISPR-Ribonucleoprotein. International Journal of Molecular Sciences, 23(18), 10678. https://doi.org/10.3390/ijms231810678







