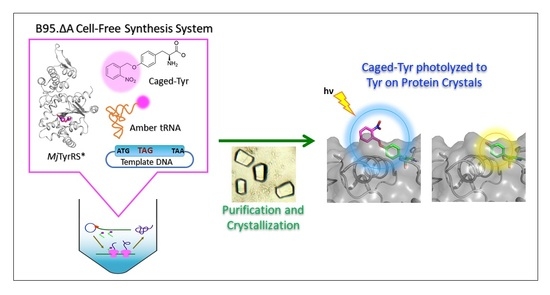
Crystal Structure of an Archaeal Tyrosyl-tRNA Synthetase Bound to Photocaged L-Tyrosine and Its Potential Application to Time-Resolved X-ray Crystallography
Abstract
:1. Introduction
2. Results and Discussion
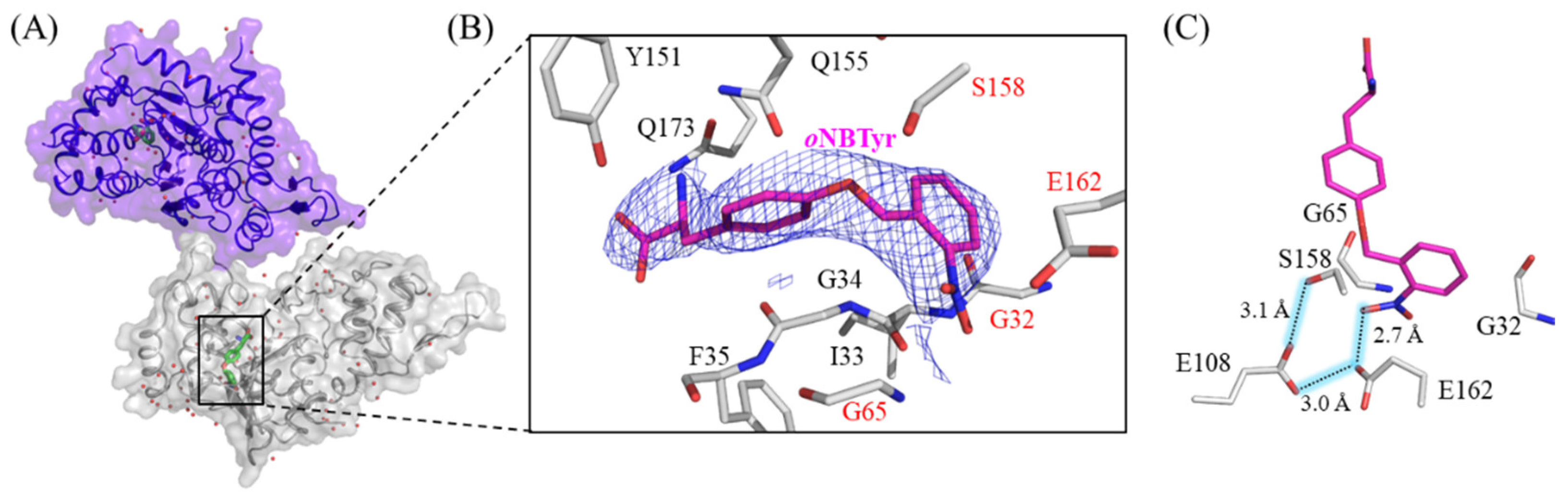
2.1. X-ray Crystallographic Study on oNBTyrRS Complexed with oNBTyr
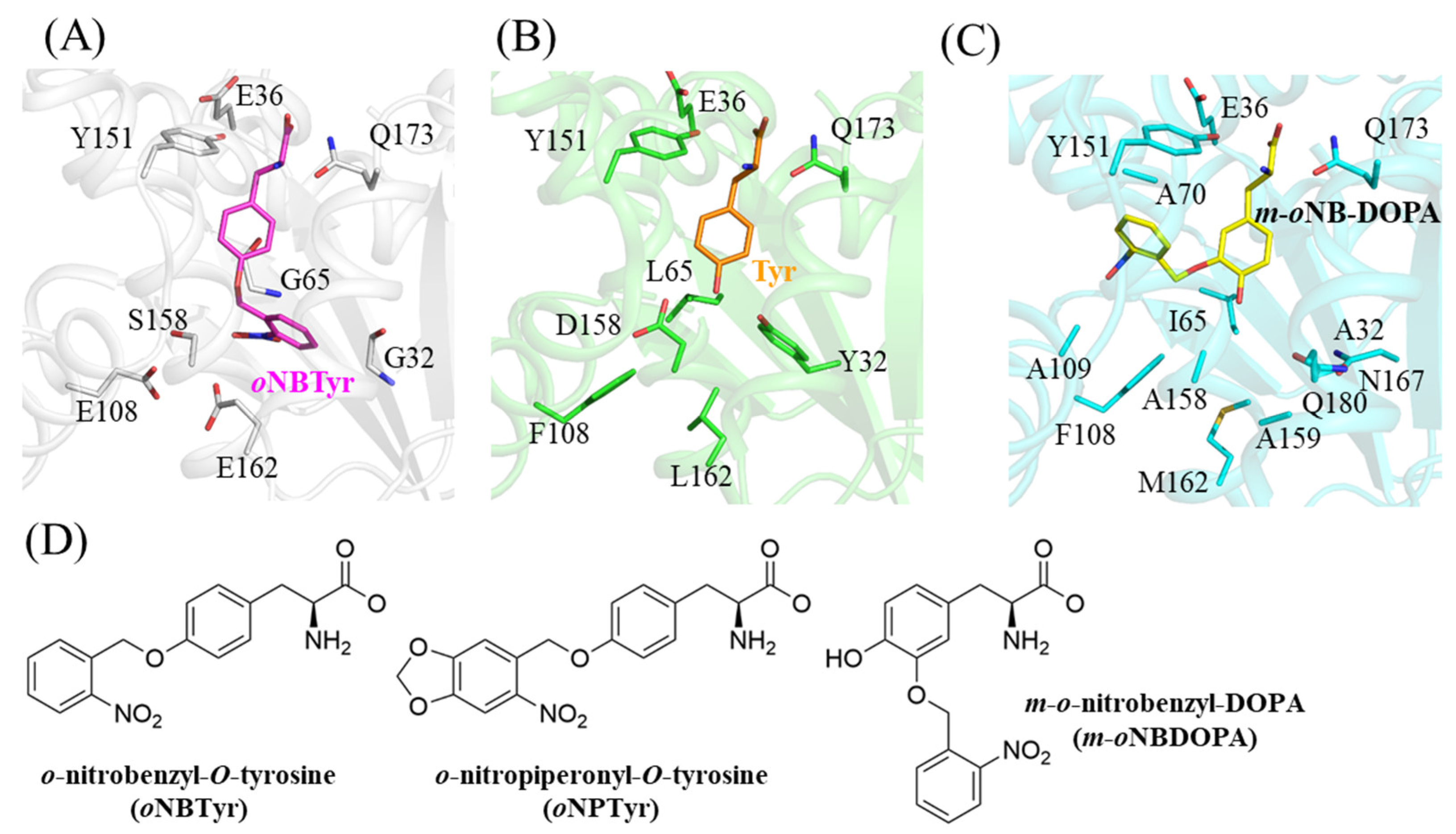
2.2. Structural Comparison with the Wild Type and the Other Mutant of MjTyrRS
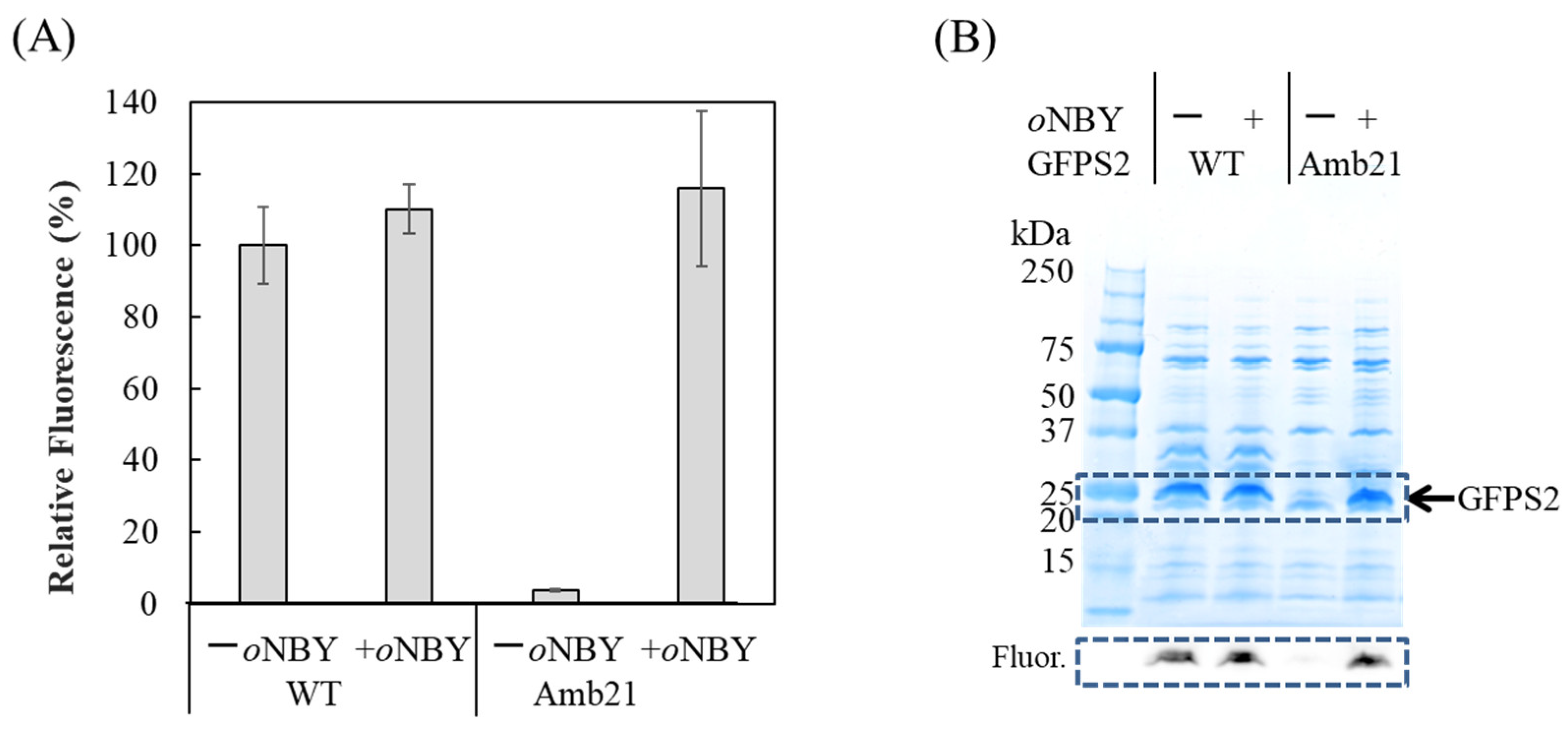
2.3. Synthesis of Caged Proteins in a Cell-Free Protein Synthesis System
2.4. Crystallization of Caged Lysozyme and Decaging in the Crystal
3. Materials and Methods
3.1. Strain, Chemicals, and Enzymes
3.2. Construction of Plasmids Vectors for Protein Production
3.3. Expression, Purification, and Crystallization of the oNBTyr-tRNA Synthetase
3.4. Data Collection, Structure Determination, and Refinement of the oNBTyr-tRNA Synthetase
3.5. Preparation of E. coli S30 Extract and Construction of Cell-Free Protein Synthesis System for Site-Specific Incorporation of Nonnatural Amino Acids
3.6. A Site-Specific Introduction of oNBTyr to HEWL by a Cell-Free System and Its Purification
3.7. Crystallization, Data Collection, Structure Determination, and Refinement of HEWL_Y20oNBTyr
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Engels, J.; Schlaeger, E.J. Synthesis, structure, and reactivity of adenosine cyclic 3′,5′-phosphate benzyl triesters. J. Med. Chem. 1977, 20, 907–911. [Google Scholar] [CrossRef]
- Kaplan, J.H.; Forbush, B., 3rd; Hoffman, J.F. Rapid photolytic release of adenosine 5′-triphosphate from a protected analogue: Utilization by the Na: K pump of human red blood cell ghosts. Biochemistry 1978, 17, 1929–1935. [Google Scholar] [CrossRef]
- Mayer, G.; Heckel, A. Biologically active molecules with a “light switch”. Angew. Chem. Int. Ed. Engl. 2006, 45, 4900–4921. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.M.; Larson, D.R.; Lawrence, D.S. Illuminating the chemistry of life: Design, synthesis, and applications of “caged” and related photoresponsive compounds. ACS Chem. Biol. 2009, 4, 409–427. [Google Scholar] [CrossRef] [PubMed]
- Gautier, A.; Gauron, C.; Volovitch, M.; Bensimon, D.; Jullien, L.; Vriz, S. How to control proteins with light in living systems. Nat. Chem. Biol. 2014, 10, 533–541. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Liu, Y.; Liu, Y.; Zheng, S.; Wang, X.; Zhao, J.; Yang, F.; Zhang, G.; Wang, C.; Chen, P.R. Time-resolved protein activation by proximal decaging in living systems. Nature 2019, 569, 509–513. [Google Scholar] [CrossRef] [PubMed]
- Ellis-Davies, G.C. Caged compounds: Photorelease technology for control of cellular chemistry and physiology. Nat. Met. 2007, 4, 619–628. [Google Scholar] [CrossRef] [PubMed]
- Hauf, M.; Richter, F.; Schneider, T.; Faidt, T.; Martins, B.M.; Baumann, T.; Durkin, P.; Dobbek, H.; Jacobs, K.; Möglich, A.; et al. Photoactivatable Mussel-Based Underwater Adhesive Proteins by an Expanded Genetic Code. ChemBioChem 2017, 18, 1819–1823. [Google Scholar] [CrossRef] [PubMed]
- Kneuttinger, A.C.; Straub, K.; Bittner, P.; Simeth, N.A.; Bruckmann, A.; Busch, F.; Rajendran, C.; Hupfeld, E.; Wysocki, V.H.; Horinek, D.; et al. Light Regulation of Enzyme Allostery through Photo-responsive Unnatural Amino Acids. Cell Chem. Biol. 2019, 26, 1501–1514.e9. [Google Scholar] [CrossRef]
- Liu, C.C.; Schultz, P.G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 2010, 79, 413–444. [Google Scholar] [CrossRef] [Green Version]
- Deiters, A.; Groff, D.; Ryu, Y.; Xie, J.; Schultz, P.G. A genetically encoded photocaged tyrosine. Angew. Chem. Int. Ed. Engl. 2006, 45, 2728–2731. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Nureki, O.; Ishitani, R.; Yaremchuk, A.; Tukalo, M.; Cusack, S.; Sakamoto, K.; Yokoyama, S. Structural basis for orthogonal tRNA specificities of tyrosyl-tRNA synthetases for genetic code expansion. Nat. Struct. Biol. 2003, 10, 425–432. [Google Scholar] [CrossRef] [PubMed]
- Calendar, R.; Berg, P. Purification and physical characterization of tyrosyl ribonucleic acid synthetases from Escherichia coli and Bacillus subtilis. Biochemistry 1966, 5, 1681–1690. [Google Scholar] [CrossRef]
- Koch, G.L. Tyrosyl transfer ribonucleic acid synthetase from Bacillus stearothermophilus. Preparation and properties of the crystallizable enzyme. Biochemistry 1974, 13, 2307–2312. [Google Scholar] [CrossRef] [PubMed]
- Brick, P.; Bhat, T.N.; Blow, D.M. Structure of tyrosyl-tRNA synthetase refined at 2.3 A resolution. Interaction of the enzyme with the tyrosyl adenylate intermediate. J. Mol. Biol. 1989, 208, 83–98. [Google Scholar] [CrossRef]
- D’Ascenzo, L.; Auffinger, P. A comprehensive classification and nomenclature of carboxyl-carboxyl(ate) supramolecular motifs and related catemers: Implications for biomolecular systems. Acta Crystallogr. B Struct. Sci. Cryst. Eng. Mater. 2015, 71, 164–175. [Google Scholar] [CrossRef] [PubMed]
- Søndergaard, C.R.; Olsson, M.H.; Rostkowski, M.; Jensen, J.H. Improved Treatment of Ligands and Coupling Effects in Empirical Calculation and Rationalization of pKa Values. J. Chem. Theory Comput. 2011, 7, 2284–2295. [Google Scholar] [CrossRef] [PubMed]
- Olsson, M.H.; Søndergaard, C.R.; Rostkowski, M.; Jensen, J.H. PROPKA3: Consistent Treatment of Internal and Surface Residues in Empirical pKa Predictions. J. Chem. Theory Comput. 2011, 7, 525–537. [Google Scholar] [CrossRef]
- Luo, J.; Torres-Kolbus, J.; Liu, J.; Deiters, A. Genetic Encoding of Photocaged Tyrosines with Improved Light-Activation Properties for the Optical Control of Protease Function. ChemBioChem 2017, 18, 1442–1447. [Google Scholar] [CrossRef]
- Mukai, T.; Yanagisawa, T.; Ohtake, K.; Wakamori, M.; Adachi, J.; Hino, N.; Sato, A.; Kobayashi, T.; Hayashi, A.; Shirouzu, M.; et al. Genetic-code evolution for protein synthesis with non-natural amino acids. Biochem. Biophys. Res. Commun. 2011, 411, 757–761. [Google Scholar] [CrossRef] [Green Version]
- Adachi, J.; Katsura, K.; Seki, E.; Takemoto, C.; Shirouzu, M.; Terada, T.; Mukai, T.; Sakamoto, K.; Yokoyama, S. Cell-Free Protein Synthesis Using S30 Extracts from Escherichia coli RFzero Strains for Efficient Incorporation of Non-Natural Amino Acids into Proteins. Int. J. Mol. Sci. 2019, 20, 492. [Google Scholar] [CrossRef] [PubMed]
- Mukai, T.; Hoshi, H.; Ohtake, K.; Takahashi, M.; Yamaguchi, A.; Hayashi, A.; Yokoyama, S.; Sakamoto, K. Highly reproductive Escherichia coli cells with no specific assignment to the UAG codon. Sci. Rep. 2015, 5, 9699. [Google Scholar] [CrossRef] [PubMed]
- Katsura, K.; Matsuda, T.; Tomabechi, Y.; Yonemochi, M.; Hanada, K.; Ohsawa, N.; Sakamoto, K.; Takemoto, C.; Shirouzu, M. A reproducible and scalable procedure for preparing bacterial extracts for cell-free protein synthesis. J. Biochem. 2017, 162, 357–369. [Google Scholar] [CrossRef] [PubMed]
- Kigawa, T.; Yabuki, T.; Matsuda, N.; Matsuda, T.; Nakajima, R.; Tanaka, A.; Yokoyama, S. Preparation of Escherichia coli cell extract for highly productive cell-free protein expression. J. Struct. Funct. Genom. 2004, 5, 63–68. [Google Scholar] [CrossRef]
- Matsuda, T.; Ito, T.; Takemoto, C.; Katsura, K.; Ikeda, M.; Wakiyama, M.; Kukimoto-Niino, M.; Yokoyama, S.; Kurosawa, Y.; Shirouzu, M. Cell-free synthesis of functional antibody fragments to provide a structural basis for antibody-antigen interaction. PLoS ONE 2018, 13, e0193158. [Google Scholar] [CrossRef]
- Liebschner, D.; Afonine, P.V.; Baker, M.L.; Bunkoczi, G.; Chen, V.B.; Croll, T.I.; Hintze, B.; Hung, L.-W.; Jain, S.; McCoy, A.J.; et al. Macromolecular structure determination using X-rays, neutrons and electrons: Recent developments in Phenix. Acta Crystallogr. D Biol. Crystallogr. 2019, 75, 861–877. [Google Scholar] [CrossRef]
- Sugahara, M.; Mizohata, E.; Nango, E.; Suzuki, M.; Tanaka, T.; Masuda, T.; Tanaka, R.; Shimamura, T.; Tanaka, Y.; Suno, C.; et al. Grease matrix as a versatile carrier of proteins for serial crystallography. Nat. Methods 2015, 12, 61–63. [Google Scholar] [CrossRef]
- Kim, D.-M.; Swartz, J.R. Efficient production of a bioactive, multiple disulfide-bonded protein using modified extracts of Escherichia coli. Biotechnol. Bioeng. 2004, 85, 122–129. [Google Scholar] [CrossRef]
- Malakhov, M.P.; Mattern, M.R.; Malakhova, O.A.; Drinker, M.; Weeks, S.D.; Butt, T.R. SUMO fusions and SUMO-specific protease for efficient expression and purification of proteins. J. Struct. Funct. Genom. 2004, 5, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Katsura, K.; Tomabechi, Y.; Matsuda, T.; Yonemochi, M.; Mikuni, J.; Ohsawa, N.; Terada, T.; Yokoyama, S.; Kukimoto-Niino, M.; Takemoto, C.; et al. Phosphorylated and non-phosphorylated HCK kinase domains produced by cell-free protein expression. Protein Expr. Purif. 2018, 150, 92–99. [Google Scholar] [CrossRef]
- Shinoda, T.; Shinya, N.; Ito, K.; Ishizuka-Katsura, Y.; Ohsawa, N.; Terada, T.; Hirata, K.; Kawano, Y.; Yamamoto, M.; Tomita, T.; et al. Cell-free methods to produce structurally intact mammalian membrane proteins. Sci. Rep. 2016, 6, 30442. [Google Scholar] [CrossRef] [PubMed]
- Hirata, K.; Kawano, Y.; Ueno, G.; Hashimoto, K.; Murakami, H.; Hasegawa, K.; Hikima, T.; Kumasaka, T.; Yamamoto, M. Achievement of protein micro-crystallography at SPring-8 beamline BL32XU. J. Phys. Conf. Ser. 2013, 425, 012002. [Google Scholar] [CrossRef]
- Hirata, K.; Yamashita, K.; Ueno, G.; Kawano, Y.; Hasegawa, K.; Kumasaka, T.; Yamamoto, M. ZOO: An automatic data-collection system for high-throughput structure analysis in protein microcrystallography. Acta Crystallogr. D Biol. Crystallogr. 2019, 75, 138–150. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, K.; Hirata, K.; Yamamoto, M. KAMO: Towards automated data processing for microcrystals. Acta Crystallogr. D Biol. Crystallogr. 2018, 74, 441–449. [Google Scholar] [CrossRef]
- Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 125–132. [Google Scholar] [CrossRef]
- McCoy, A.J.; Grosse-Kunstleve, R.W.; Adams, P.D.; Winn, M.D.; Storoni, L.C.; Read, R.J. Phaser crystallographic software. J. Appl. Crystallogr. 2007, 40, 658–674. [Google Scholar] [CrossRef] [PubMed]
- Emsley, P.; Cowtan, K. Coot: Model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 2004, 60, 2126–2132. [Google Scholar] [CrossRef]
- Seki, E.; Yanagisawa, T.; Yokoyama, S. Cell-Free Protein Synthesis for Multiple Site-Specific Incorporation of Noncanonical Amino Acids Using Cell Extracts from RF-1 Deletion E. coli Strains. Methods Mol. Biol. 2018, 1728, 49–65. [Google Scholar]
- San Miguel, V.; Bochet, C.G.; del Campo, A. Wavelength-selective caged surfaces: How many functional levels are possible? J. Am. Chem. Soc. 2011, 133, 5380–5388. [Google Scholar] [CrossRef]
- Liu, Q.; He, Q.T.; Lyu, X.; Yang, F.; Zhu, Z.L.; Xiao, P.; Yang, Z.; Zhang, F.; Yang, Z.Y.; Wang, X.Y.; et al. DeSiphering receptor core-induced and ligand-dependent conformational changes in arrestin via genetic encoded trimethylsilyl (1)H-NMR probe. Nat. Commun. 2020, 11, 4857. [Google Scholar] [CrossRef] [PubMed]
- Beyer, J.N.; Hosseinzadeh, P.; Gottfried-Lee, I.; Van Fossen, E.M.; Zhu, P.; Bednar, R.M.; Karplus, P.A.; Mehl, R.A.; Cooley, R.B. Overcoming Near-Cognate Suppression in a Release Factor 1-Deficient Host with an Improved Nitro-Tyrosine tRNA Synthetase. J. Mol. Biol. 2020, 432, 4690–4704. [Google Scholar] [CrossRef]
- Schiefner, A.; Nastle, L.; Landgraf, M.; Reichert, A.J.; Skerra, A. Structural Basis for the Specific Cotranslational Incorporation of p-Boronophenylalanine into Biosynthetic Proteins. Biochemistry 2018, 57, 2597–2600. [Google Scholar] [CrossRef]
- Baumann, T.; Hauf, M.; Schildhauer, F.; Eberl, K.B.; Durkin, P.M.; Deniz, E.; Löffler, J.G.; Acevedo-Rocha, C.G.; Jaric, J.; Martins, B.M.; et al. Site-Resolved Observation of Vibrational Energy Transfer Using a Genetically Encoded Ultrafast Heater. Angew. Chem. Int. Ed. Engl. 2019, 58, 2899–2903. [Google Scholar] [CrossRef]
- Luo, X.; Fu, G.; Wang, R.E.; Zhu, X.; Zambaldo, C.; Liu, R.; Liu, T.; Lyu, X.; Du, J.; Xuan, W.; et al. Genetically encoding phosphotyrosine and its nonhydrolyzable analog in bacteria. Nat. Chem. Biol. 2017, 13, 845–849. [Google Scholar] [CrossRef]
- Liu, X.; Jiang, L.; Li, J.; Wang, L.; Yu, Y.; Zhou, Q.; Lv, X.; Gong, W.; Lu, Y.; Wang, J. Significant expansion of fluorescent protein sensing ability through the genetic incorporation of superior photo-induced electron-transfer quenchers. J. Am. Chem. Soc. 2014, 136, 13094–13097. [Google Scholar] [CrossRef]
- Cooley, R.B.; Karplus, P.A.; Mehl, R.A. Gleaning unexpected fruits from hard-won synthetases: Probing principles of permissivity in non-canonical amino acid-tRNA synthetases. ChemBioChem 2014, 15, 1810–1819. [Google Scholar] [CrossRef] [PubMed]
- Cooley, R.B.; Feldman, J.L.; Driggers, C.M.; Bundy, T.A.; Stokes, A.L.; Karplus, P.A.; Mehl, R.A. Structural basis of improved second-generation 3-nitro-tyrosine tRNA synthetases. Biochemistry 2014, 53, 1916–1924. [Google Scholar] [CrossRef]
- Li, F.; Shi, P.; Li, J.; Yang, F.; Wang, T.; Zhang, W.; Gao, F.; Ding, W.; Li, D.; Li, J.; et al. A genetically encoded 19F NMR probe for tyrosine phosphorylation. Angew. Chem. Int. Ed. Engl. 2013, 52, 3958–3962. [Google Scholar] [CrossRef]
- Young, D.D.; Young, T.S.; Jahnz, M.; Ahmad, I.; Spraggon, G.; Schultz, P.G. An evolved aminoacyl-tRNA synthetase with atypical polysubstrate specificity. Biochemistry 2011, 50, 1894–1900. [Google Scholar] [CrossRef]
- Wang, J.; Zhang, W.; Song, W.; Wang, Y.; Yu, Z.; Li, J.; Wu, M.; Wang, L.; Zang, J.; Lin, Q. A biosynthetic route to photoclick chemistry on proteins. J. Am. Chem. Soc. 2010, 132, 14812–14818. [Google Scholar] [CrossRef]
- Tippmann, E.M.; Liu, W.; Summerer, D.; Mack, A.V.; Schultz, P.G. A genetically encoded diazirine photocrosslinker in Escherichia coli. ChemBioChem 2007, 8, 2210–2214. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Liu, W.; Schultz, P.G. A genetically encoded bidentate, metal-binding amino acid. Angew. Chem. Int. Ed. Engl. 2007, 46, 9239–9242. [Google Scholar] [CrossRef]
- Liu, W.; Alfonta, L.; Mack, A.V.; Schultz, P.G. Structural basis for the recognition of para-benzoyl-L-phenylalanine by evolved aminoacyl-tRNA synthetases. Angew. Chem. Int. Ed. Engl. 2007, 46, 6073–6075. [Google Scholar] [CrossRef]
- Turner, J.M.; Graziano, J.; Spraggon, G.; Schultz, P.G. Structural plasticity of an aminoacyl-tRNA synthetase active site. Proc. Natl. Acad. Sci. USA 2006, 103, 6483–6488. [Google Scholar] [CrossRef]
- Turner, J.M.; Graziano, J.; Spraggon, G.; Schultz, P.G. Structural characterization of a p-acetylphenylalanyl aminoacyl-tRNA synthetase. J. Am. Chem. Soc. 2005, 127, 14976–14977. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hosaka, T.; Katsura, K.; Ishizuka-Katsura, Y.; Hanada, K.; Ito, K.; Tomabechi, Y.; Inoue, M.; Akasaka, R.; Takemoto, C.; Shirouzu, M. Crystal Structure of an Archaeal Tyrosyl-tRNA Synthetase Bound to Photocaged L-Tyrosine and Its Potential Application to Time-Resolved X-ray Crystallography. Int. J. Mol. Sci. 2022, 23, 10399. https://doi.org/10.3390/ijms231810399
Hosaka T, Katsura K, Ishizuka-Katsura Y, Hanada K, Ito K, Tomabechi Y, Inoue M, Akasaka R, Takemoto C, Shirouzu M. Crystal Structure of an Archaeal Tyrosyl-tRNA Synthetase Bound to Photocaged L-Tyrosine and Its Potential Application to Time-Resolved X-ray Crystallography. International Journal of Molecular Sciences. 2022; 23(18):10399. https://doi.org/10.3390/ijms231810399
Chicago/Turabian StyleHosaka, Toshiaki, Kazushige Katsura, Yoshiko Ishizuka-Katsura, Kazuharu Hanada, Kaori Ito, Yuri Tomabechi, Mio Inoue, Ryogo Akasaka, Chie Takemoto, and Mikako Shirouzu. 2022. "Crystal Structure of an Archaeal Tyrosyl-tRNA Synthetase Bound to Photocaged L-Tyrosine and Its Potential Application to Time-Resolved X-ray Crystallography" International Journal of Molecular Sciences 23, no. 18: 10399. https://doi.org/10.3390/ijms231810399
APA StyleHosaka, T., Katsura, K., Ishizuka-Katsura, Y., Hanada, K., Ito, K., Tomabechi, Y., Inoue, M., Akasaka, R., Takemoto, C., & Shirouzu, M. (2022). Crystal Structure of an Archaeal Tyrosyl-tRNA Synthetase Bound to Photocaged L-Tyrosine and Its Potential Application to Time-Resolved X-ray Crystallography. International Journal of Molecular Sciences, 23(18), 10399. https://doi.org/10.3390/ijms231810399