The Cross-Resistance Pattern and the Metabolic Resistance Mechanism of Acetamiprid in the Brown Planthopper, Nilaparvata lugens (Stål)
Abstract
1. Introduction
2. Results
2.1. Acetamiprid Resistance Selection
2.2. The Cross-Resistance Pattern
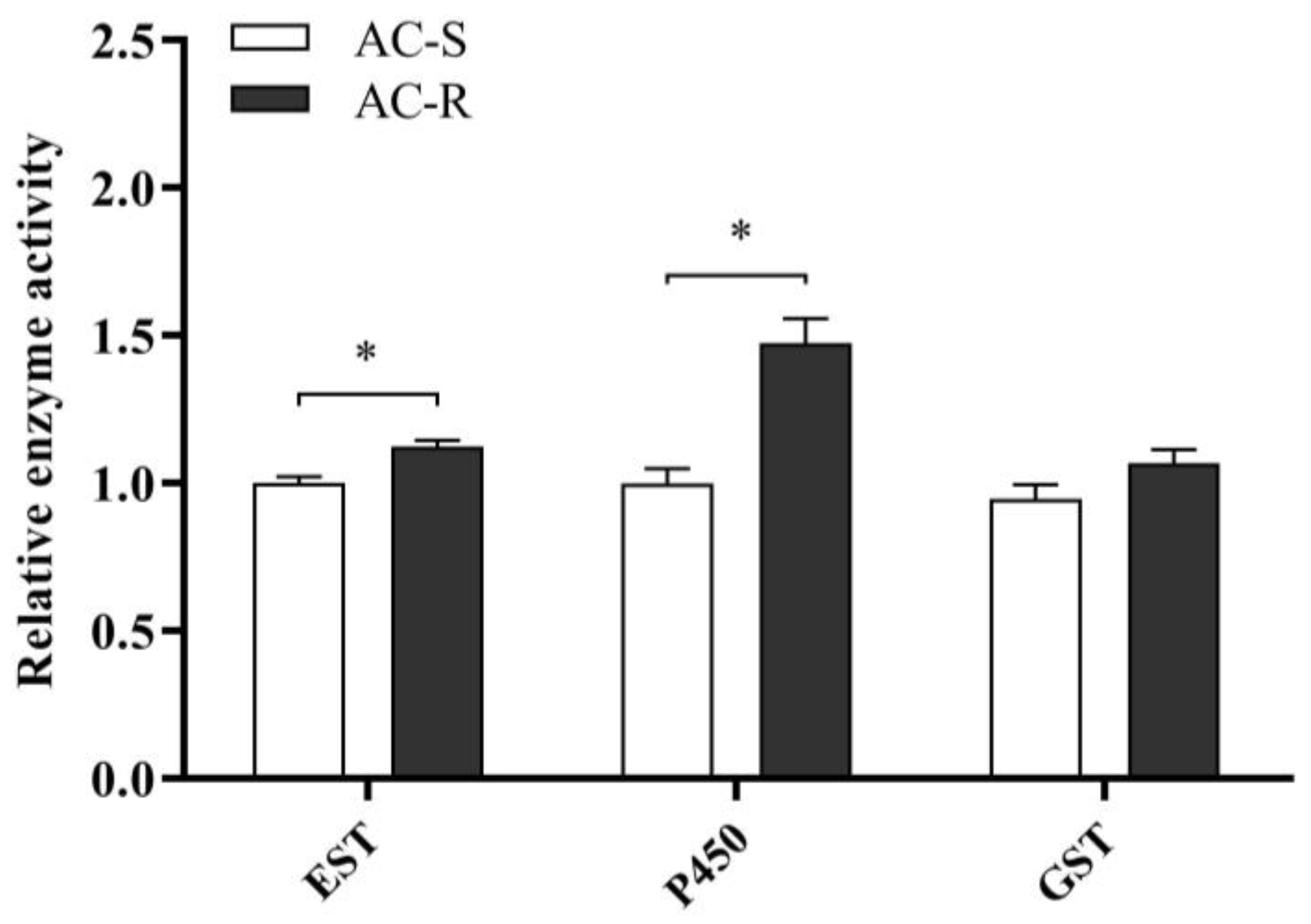
2.3. Synergistic Effects and Enzyme Activity Evaluation
2.4. Assessment of Expression Levels of P450 Genes
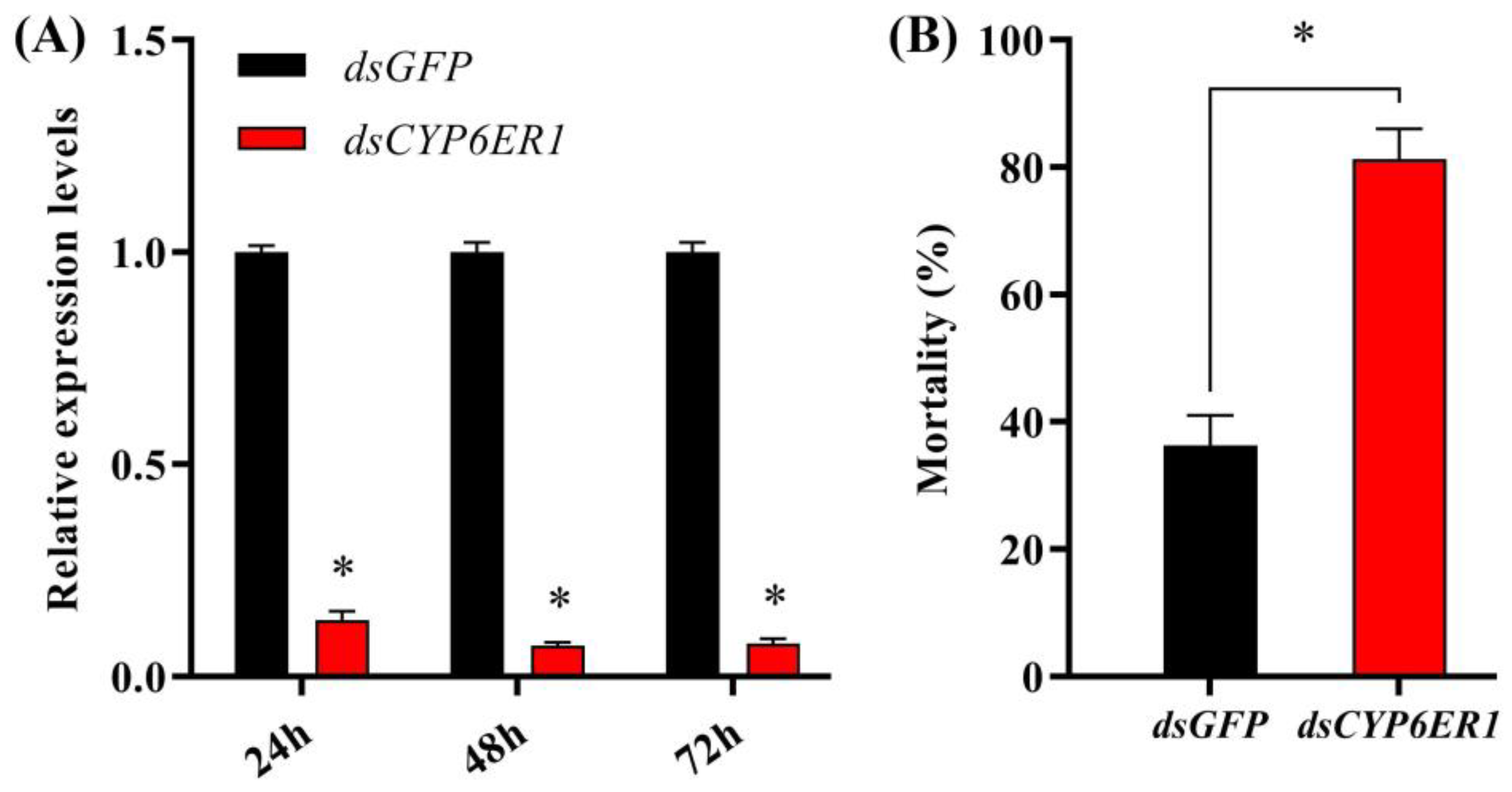
2.5. Silencing of CYP6ER1 Increases Susceptibility to Acetamiprid in the Resistant Strain
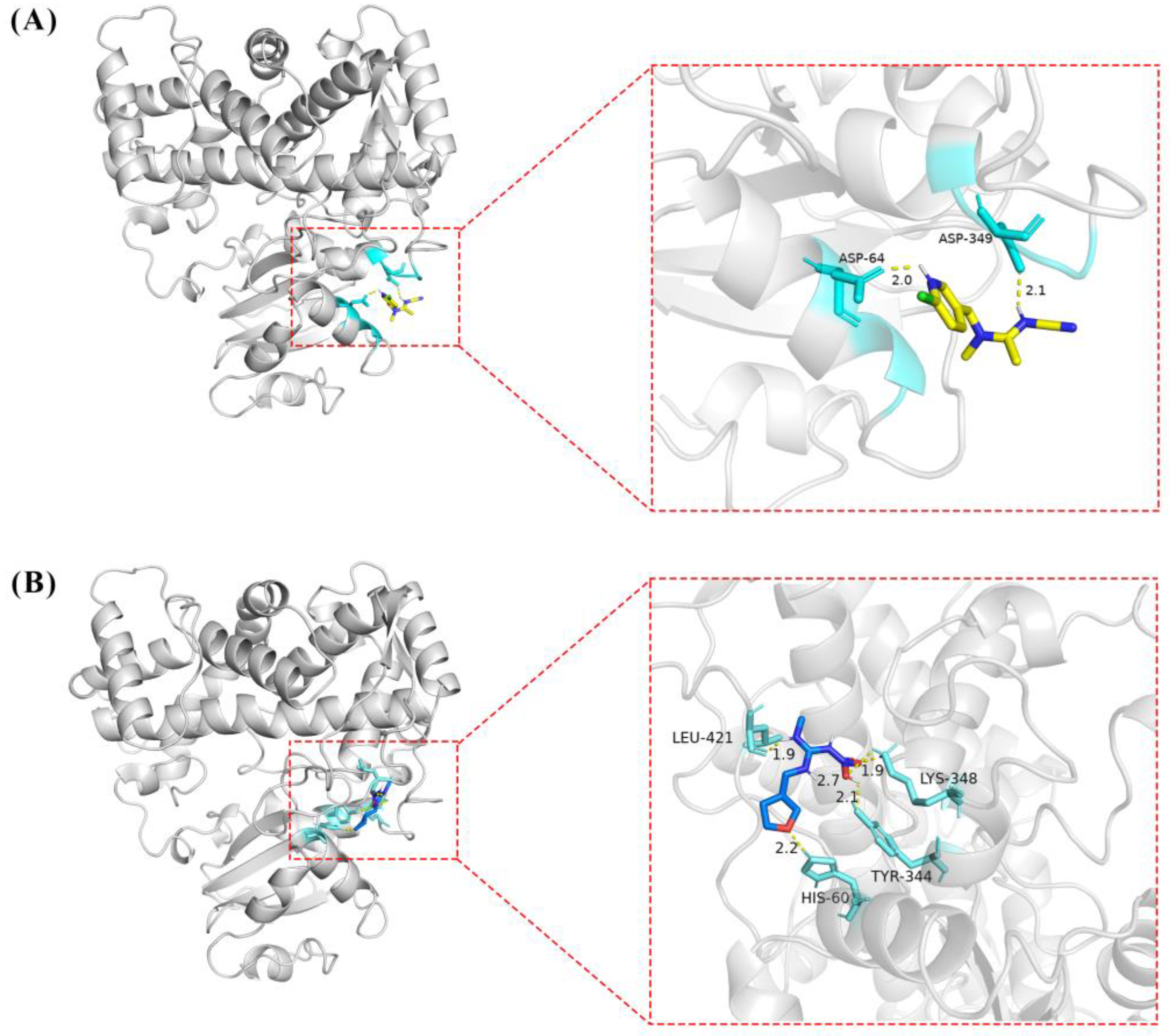
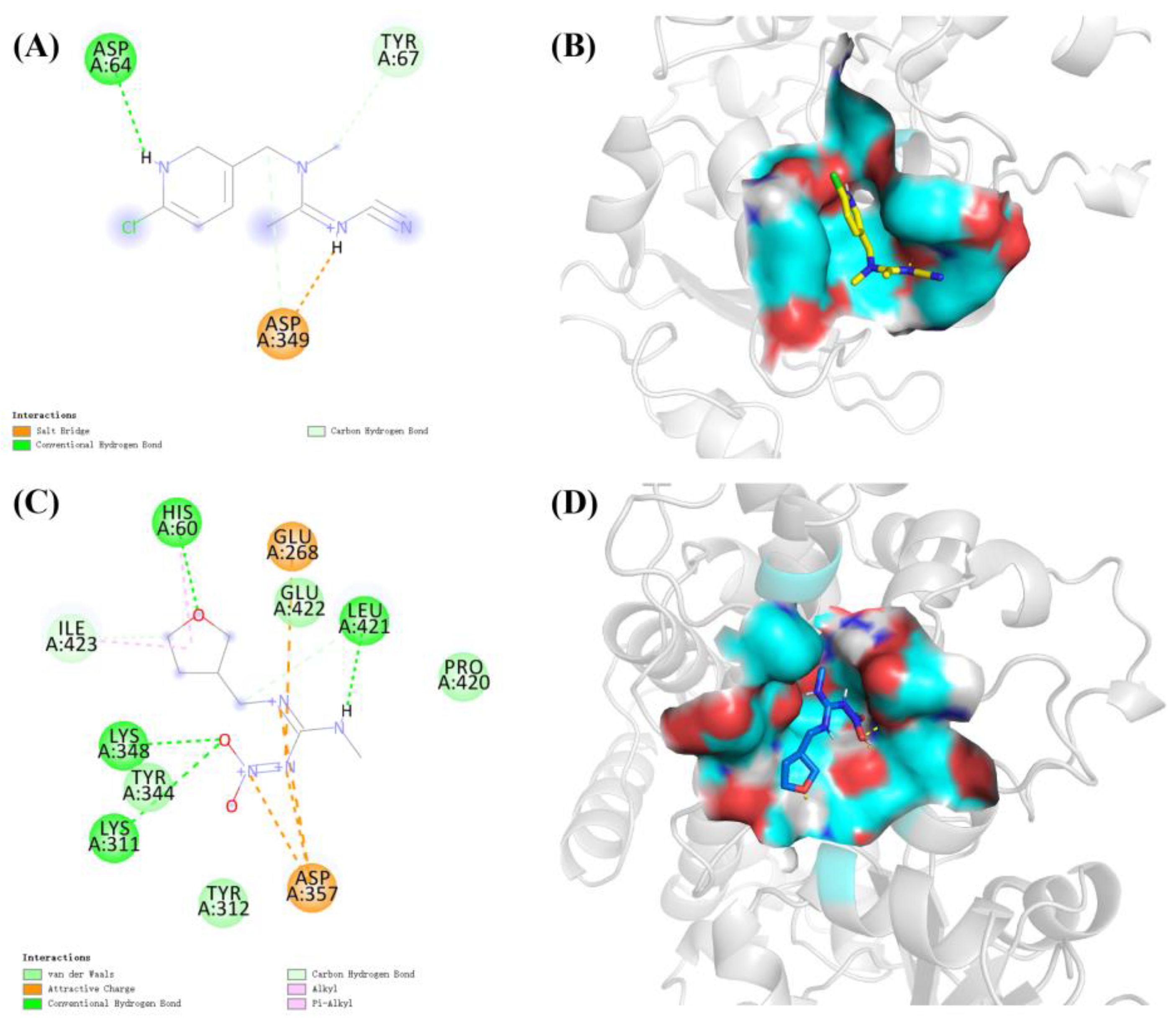
2.6. In Silico Binding of Acetamiprid to CYP6ER1
3. Discussion
4. Materials and Methods
4.1. Insects
4.2. Insecticides and Chemicals
4.3. Bioassays
4.4. Enzyme Activity Measurements
4.5. RNA Isolation, cDNA Preparation, and RT-qPCR
4.6. The RNA Interference of CYP6ER1
4.7. Homology Modelling and Molecular Docking
4.8. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cheng, J. Rice planthopper problems and relevant causes in China. In Planthoppers: New Threats to the Sustainability of Intensive Rice Production Systems in Asia; Heong, K.L., Hardy, B., Eds.; International Rice Research Institute: Los Baños, Philippines, 2009; pp. 157–178. [Google Scholar]
- Zha, W.; Zhou, L.; Li, S.; Liu, K.; Yang, G.; Chen, Z.; Liu, K.; Xu, H.; Li, P.; Hussain, S.; et al. Characterization and comparative profiling of the small RNA transcriptomes in the Hemipteran insect Nilaparvata Lugens. Gene 2016, 595, 83–91. [Google Scholar] [CrossRef]
- Hogenhout, S.; Ammar, E.-D.; Whitfield, A.; Redinbaugh, M.G. Insect vector interactions with persistently transmitted viruses. Annu. Rev. Phytopathol. 2008, 46, 327–359. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.; Guo, J.; Fu, X.; Huang, Y.; Gao, X.; Wu, K. Seasonal migration pattern of Nilaparvata lugens (Hemiptera: Delphacidae) over the bohai sea in northern China. J. Econ. Entomol. 2018, 111, 2129–2135. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Wang, B.; Chen, L.; Li, P.; Cao, C. The different influences of drought stress at the flowering stage on rice physiological traits, grain yield, and quality. Sci. Rep. 2019, 9, 3742. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Zeng, B.; Zheng, C.; Mu, X.C.; Zhang, Y.; Hu, J.; Zhang, S.; Gao, C.F.; Shen, J.L. The evolution of insecticide resistance in the brown planthopper (Nilaparvata lugens Stål) of China in the period 2012–2016. Sci. Rep. 2018, 8, 4586. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Liao, X.; Mao, K.; Zhang, K.; Wan, H.; Li, J. Insecticide resistance monitoring and correlation analysis of insecticides in field populations of the brown planthopper Nilaparvata lugens (stål) in China 2012–2014. Pestic. Biochem. Physiol. 2016, 132, 13–20. [Google Scholar] [CrossRef]
- Liao, X.; Xu, P.; Gong, P.; Wan, H.; Li, J. Current susceptibilities of brown planthopper Nilaparvata lugens to triflumezopyrim and other frequently used insecticides in China. Insect Sci. 2021, 28, 115–126. [Google Scholar] [CrossRef]
- Takahashi, H.; Takakusa, N.; Suzuki, J.; Kishimoto, T. Development of a new insecticide, acetamiprid. J. Pest. Sci. 1998, 23, 193–200. [Google Scholar] [CrossRef][Green Version]
- Tomizawa, M.; Casida, J.E. Neonicotinoid insecticide toxicology: Mechanisms of selective action. Annu. Rev. Pharmacol. Toxicol. 2005, 45, 247–268. [Google Scholar] [CrossRef]
- Otoidobiga, L.; Vincent, L.; Stewart, R.K. Field efficacy and baseline toxicities of pyriproxifen, acetamiprid, and diafenthiuron against Bemisia tabaci Gennadius (Homoptera: Aleyrodidae) in Burkina Faso (West Africa). J. Environ. Sci. Health B 2003, 38, 757–769. [Google Scholar] [CrossRef]
- Mokbel, E.-S. Resistance risk assessment: Realized heritability, cross resistance and resistance stability of acetamiprid in the cotton aphid, Aphis gossypii Glover (Homoptera: Aphididae). J. Plant Protect. Res. 2019, 58, 328–334. [Google Scholar]
- Ijaz, M.; Afzal, M.B.S.; Shad, S.A. Resistance risk analysis to acetamiprid and other insecticides in Acetamiprid-Selected population of Phenacoccus solenopsis. Phytoparasitica 2016, 44, 177–186. [Google Scholar] [CrossRef]
- Ninsin, K.; Tanaka, T. Synergism and stability of acetamiprid resistance in a laboratory colony of Plutella xylostella. Pest Manag. Sci. 2005, 61, 723–727. [Google Scholar] [CrossRef] [PubMed]
- Ullah, F.; Gul, H.; Tariq, K.; Desneux, N.; Gao, X.; Song, D. Functional analysis of cytochrome P450 genes linked with acetamiprid resistance in melon aphid, Aphis gossypii. Pestic. Biochem. Physiol. 2020, 170, 104687. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Cao, Q.; Li, G.; Ma, D. Role of several cytochrome P450s in the resistance and cross-resistance; against imidacloprid and acetamiprid of Bemisia tabaci (Hemiptera: Aleyrodidae) MEAM1 cryptic species in Xinjiang, China. Pestic. Biochem. Physiol. 2020, 163, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Heckel, D.G. Insecticide Resistance After Silent Spring. Science 2012, 337, 1613–1614. [Google Scholar] [CrossRef]
- Li, X.; Schuler, M.A.; Berenbaum, M.R. Molecular mechanisms of metabolic resistance to synthetic and natural xenobiotics. Annu. Rev. Entomol. 2007, 52, 231–253. [Google Scholar] [CrossRef]
- Zhang, Y.X.; Yang, B.; Li, J.; Liu, M.; Liu, Z.W. Point mutations in acetylcholinesterase 1 associated with chlorpyrifos resistance in the brown planthopper, Nilaparvata lugens Stål. Insect Mol. Biol. 2017, 26, 453–460. [Google Scholar] [CrossRef]
- Mao, K.; Ren, Z.; Li, W.; Cai, T.; Qin, X.; Wan, H.; Jin, B.; He, S.; Li, J. Carboxylesterase genes in nitenpyramresistant brown planthoppers, Nilaparvata lugens. Insect Sci. 2020, 28, 1049–1060. [Google Scholar] [CrossRef]
- Vontas, J.G.; Small, G.J.; Hemingway, J. Glutathione S-transferases as antioxidant defence agents confer pyrethroid resistance in Nilaparvata lugens. Biochem. J. 2001, 357, 65–72. [Google Scholar] [CrossRef]
- Feyereisen, R. 4.1-Insect Cytochrome P450. In Comprehensive Molecular Insect Science; Gilbert, L.I., Ed.; Elsevier: Amsterdam, The Netherlands, 2005; pp. 1–77. [Google Scholar]
- Chen, A.; Zhang, H.; Shan, T.; Shi, X.; Gao, X. The overexpression of three cytochrome P450 genes CYP6CY14, CYP6CY22 and CYP6UN1 contributed to metabolic resistance to dinotefuran in melon/cotton aphid, Aphis gossypii Glover. Pestic. Biochem. Physiol. 2020, 167, 104601. [Google Scholar] [CrossRef] [PubMed]
- Zhu, F.; Parthasarathy, R.; Bai, H.; Woithe, K.; Kaussmann, M.; Nauen, R.; Harrison, D.A.; Palli, S.R. A brain-specific cytochrome P450 responsible for the majority of deltamethrin resistance in the QTC279 strain of Tribolium castaneum. Proc. Natl. Acad. Sci. USA 2010, 107, 8557–8562. [Google Scholar] [CrossRef] [PubMed]
- Jin, R.; Mao, K.; Liao, X.; Xu, P.; Li, Z.; Ali, E.; Wan, H.; Li, J. Overexpression of CYP6ER1 associated with clothianidin resistance in Nilaparvata lugens (Stal). Pestic. Biochem. Physiol. 2019, 154, 39–45. [Google Scholar] [CrossRef]
- Mao, K.; Zhang, X.; Ali, E.; Liao, X.; Jin, R.; Ren, Z.; Wan, H.; Li, J. Characterization of nitenpyram resistance in Nilaparvata lugens (Stål). Pestic. Biochem. Physiol. 2019, 157, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.; Wen, Y.; Yang, B.; Zhang, Y.; Liu, S.; Liu, Z.; Han, Z. Biochemical mechanisms of imidacloprid resistance in Nilaparvata lugens: Over-expression of cytochrome P450 CYP6AY1. Insect Biochem. Mol. 2013, 43, 1021–1027. [Google Scholar] [CrossRef] [PubMed]
- Liao, X.; Jin, R.; Zhang, X.; Ali, E.; Mao, K.; Xu, P.; Li, J.; Wan, H. Characterization of sulfoxaflor resistance in the brown planthopper, Nilaparvata lugens (Stål). Pest Manag. Sci. 2019, 75, 1646–1654. [Google Scholar] [CrossRef]
- Sun, X.; Gong, Y.; Ali, S.; Hou, M. Mechanisms of resistance to thiamethoxam and dinotefuran compared to imidacloprid in the brown planthopper: Roles of cytochrome P450 monooxygenase and a P450 gene CYP6ER1. Pestic. Biochem. Physiol. 2018, 150, 17–26. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, B.; Zhang, Z.; Wang, L.; Guo, H.; Li, Z.; He, P.; Liu, Z.; Fang, J. Differential expression of P450 genes and nAChR subunits associated with imidacloprid resistance in Laodelphax striatellus (Hemiptera: Delphacidae). J. Econ. Entomol. 2018, 111, 1382–1387. [Google Scholar] [CrossRef]
- Wang, X.; Ruan, Y.; Gong, C.; Xiang, X.; Xu, X.; Zhang, Y.; Shen, L. Transcriptome analysis of Sogatella furcifera (Homoptera: Delphacidae) in response to sulfoxaflor and functional verification of resistance-related P450 genes. Int. J. Mol. Sci. 2019, 20, 4573. [Google Scholar] [CrossRef]
- Liao, X.; Mao, K.; Ali, E.; Zhang, X.; Wan, H.; Li, J. Temporal variability and resistance correlation of sulfoxaflor susceptibility among. Chinese populations of the brown planthopper Nilaparvata lugens (Stål). Crop Prot. 2017, 102, 141–146. [Google Scholar] [CrossRef]
- Bass, C.; Denholm, I.; Williamson, M.S.; Nauen, R. The global status of insect resistance to neonicotinoid insecticides. Pestic. Biochem. Physiol. 2015, 121, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Wu, Q.; Wang, S.; Chang, X.; Xie, W.; Xu, B.; Zhang, Y. Cross-resistance study and biochemical mechanisms of thiamethoxam resistance in B-biotype Bemisia tabaci (Hemiptera: Aleyrodidae). Pest Manag. Sci. 2010, 66, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Horowitz, A.R.; Ghanim, M.; Roditakis, E.; Nauen, R.; Ishaaya, I. Insecticide resistance and its management in Bemisia tabaci species. J. Pest Sci. 2020, 93, 893–910. [Google Scholar] [CrossRef]
- Liu, N. Insecticide Resistance in Mosquitoes: Impact, Mechanisms, and Research Directions. Annu. Rev. Entomol. 2015, 60, 537–559. [Google Scholar] [CrossRef]
- Scott, E.E.; Wolf, C.R.; Otyepka, M.; Humphreys, S.C.; Reed, J.R.; Henderson, C.J.; McLaughlin, L.A.; Paloncyova, M.; Navratilova, V.; Berka, K.; et al. The role of protein-protein and protein-membrane interactions on P450 function. Drug Metab. Dispos. 2016, 44, 576–590. [Google Scholar] [CrossRef]
- Karunker, I.; Benting, J.; Lueke, B.; Ponge, T.; Nauen, R.; Roditakis, E.; Vontas, J.; Gorman, K.; Denholm, I.; Morin, S. Over-expression of cytochrome P450 CYP6CM1 is associated with high resistance to imidacloprid in the B and Q biotypes of Bemisia tabaci (Hemiptera: Aleyrodidae). Insect Biochem. Mol. Biol. 2008, 38, 634–644. [Google Scholar] [CrossRef]
- Li, X.; Li, R.; Zhu, B.; Gao, X.; Liang, P. Overexpression of cytochrome P450 CYP6BG1 may contribute to chlorantraniliprole resistance in Plutella xylostella (L.). Pest Manag. Sci. 2018, 74, 1386–1393. [Google Scholar] [CrossRef]
- Ma, K.; Tang, Q.; Zhang, B.; Liang, P.; Wang, B.; Gao, X. Overexpression of multiple cytochrome P450 genes associated with sulfoxaflor resistance in Aphis gossypii Glover. Pestic. Biochem. Physiol. 2019, 157, 204–210. [Google Scholar] [CrossRef]
- Yang, X.; Deng, S.; Wei, X.; Yang, J.; Zhao, Q.; Yin, C.; Du, T.; Guo, Z.; Xia, J.; Yang, Z.; et al. MAPK-directed activation of the whitefly transcription factor CREB leads to P450-mediated imidacloprid resistance. Proc. Natl. Acad. Sci. USA 2020, 117, 10246–10253. [Google Scholar] [CrossRef]
- Zimmer, C.T.; Garrood, W.T.; Singh, K.S.; Randall, E.; Lueke, B.; Gutbrod, O.; Matthiesen, S.; Kohler, M.; Nauen, R.; Davies, T.G.E.; et al. Neofunctionalization of duplicated P450 genes drives the evolution of insecticide resistance in the brown planthopper. Curr. Biol. 2018, 28, 268–274. [Google Scholar] [CrossRef]
- Bass, C.; Field, L.M. Gene amplification and insecticide resistance. Pest Manag. Sci. 2011, 67, 886–890. [Google Scholar] [CrossRef] [PubMed]
- Bass, C.; Carvalho, R.A.; Oliphant, L.; Puinean, A.M.; Field, L.M.; Nauen, R.; Williamson, M.S.; Moores, G.; Gorman, K. Overexpression of a cytochrome P450 monooxygenase, CYP6ER1, is associated with resistance to imidacloprid in the brown planthopper, Nilaparvata lugens. Insect Mol. Biol. 2011, 20, 763–773. [Google Scholar] [CrossRef] [PubMed]
- Pang, R.; Chen, M.; Liang, Z.; Yue, X.; Ge, H.; Zhang, W. Functional analysis of CYP6ER1, a P450 gene associated with imidacloprid resistance in Nilaparvata lugens. Sci. Rep. 2016, 6, 34992. [Google Scholar] [CrossRef]
- Habig, W.H.; Pabst, M.J.; Jakoby, W.B. Glutathione S-transferases. The first enzymatic step in mercapturic acid formation. J. Biol. Chem. 1974, 249, 7130–7139. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Schmittgen, T.D. Real-Time Quantitative PCR. Methods 2001, 25, 383–385. [Google Scholar] [CrossRef] [PubMed]
- Polyakov, K.M.; Gavryushov, S.; Fedorova, T.V.; Glazunova, O.A.; Popov, A.N. The subatomic resolution study of laccase inhibition by chloride and fluoride anions using single-crystal serial crystallography: Insights into the enzymatic reaction mechanism. Acta Crystallogr. Sect. D Struct. Biol. 2019, 75, 804–816. [Google Scholar] [CrossRef]
- Glazunova, O.A.; Polyakov, K.M.; Moiseenko, K.V.; Kurzeev, S.A.; Fedorova, T.V. Structure-function study of two new middle-redox potential laccases from basidiomycetes Antrodiella faginea and Steccherinum murashkinskyi. Int. J. Biol. Macromol. 2018, 118, 406–418. [Google Scholar] [CrossRef]
- Schwede, T.; Kopp, J.; Guex, N.; Peitsch, M.C. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res. 2003, 31, 3381–3385. [Google Scholar] [CrossRef]
- Pontius, J.; Richelle, J.; Wodak, S.J. Deviations from standard atomic volumes as a quality measure for protein crystal structures. J. Mol. Biol. 1996, 264, 121–136. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed]
- Eby, J.M.; Smith, A.R.; Riley, T.P.; Cosgrove, C.; Ankney, C.M.; Henning, S.W.; Paulos, C.M.; Garrett-Mayer, E.; Luiten, R.M.; Nishimura, M.I.; et al. Molecular properties of gp100-reactive T-cell receptors drive the cytokine profile and antitumor efficacy of transgenic host T cells. Pigm. Cell Melanoma Res. 2019, 32, 68–78. [Google Scholar] [CrossRef] [PubMed]


| Generation | No. | Slope (±SE) | χ2 (df) | p Value | LC50 (95%CI) (mg/L) | RR | RR’ |
|---|---|---|---|---|---|---|---|
| AC-S | 315 | 2.73 (±0.26) | 2.74 (4) | 0.603 | 16.20 (13.65–18.98) | 1 | - |
| G0 | 315 | 2.02 (±0.22) | 1.41 (4) | 0.842 | 39.13 (31.56–47.55) | 2.4 | 1 |
| G1 | 315 | 2.28 (±0.24) | 1.28 (4) | 0.864 | 44.24 (36.75–52.87) | 2.7 | 1.1 |
| G2 | 315 | 1.68 (±0.21) | 2.39 (4) | 0.664 | 40.11 (31.44–53.88) | 2.5 | 1.0 |
| G3 | 270 | 2.30 (±0.35) | 1.40 (3) | 0.706 | 64.47 (52.35–79.10) | 4.0 | 1.7 |
| G4 | 315 | 2.53 (±0.27) | 3.15 (4) | 0.533 | 66.91 (56.03–78.42) | 4.1 | 1.7 |
| G5 | 315 | 1.88 (±0.22) | 5.48 (4) | 0.241 | 69.90 (54.38–86.63) | 4.3 | 1.8 |
| G6 | 270 | 2.45 (±0.32) | 2.71 (3) | 0.438 | 102.22 (80.48–123.93) | 6.3 | 2.6 |
| G7 | 315 | 3.78 (±0.46) | 3.71 (4) | 0.446 | 143.35 (127.85–162.68) | 8.9 | 3.7 |
| G8 | 315 | 2.74 (±0.32) | 0.67 (4) | 0.956 | 165.83 (141.86–194.22) | 10.2 | 4.2 |
| G9 | 315 | 1.71 (±0.20) | 4.58 (4) | 0.644 | 110.35 (86.87–140.56) | 6.8 | 2.8 |
| G10 | 315 | 2.97 (±0.34) | 1.94 (4) | 0.746 | 220.39 (191.08–262.42) | 13.6 | 5.6 |
| G11 | 315 | 3.02 (±0.40) | 1.24 (4) | 0.872 | 125.32 (107.87–144.89) | 7.7 | 3.2 |
| G12 | 315 | 1.64 (±0.24) | 0.96 (4) | 0.915 | 134.95 (107.67–172.07) | 8.3 | 3.5 |
| G13 | 315 | 3.32 (±0.27) | 0.72 (4) | 0.949 | 231.45 (191.95–293.64) | 14.3 | 5.9 |
| G14 | 315 | 1.41 (±0.25) | 1.96 (4) | 0.743 | 271.45 (205.26–414.05) | 16.8 | 6.9 |
| G15 | 315 | 2.06 (±0.30) | 1.78 (4) | 0.775 | 315.42 (263.68–390.67) | 19.5 | 8.1 |
| G16 | 315 | 3.55 (±0.44) | 2.14 (4) | 0.711 | 363.72(325.08–417.12) | 22.5 | 9.3 |
| G17 | 315 | 3.39 (±0.40) | 3.00 (4) | 0.559 | 343.50 (303.20–401.48) | 21.2 | 8.8 |
| G18 | 315 | 2.72 (±0.40) | 2.38 (4) | 0.667 | 421.36 (362.82–519.72) | 26.0 | 10.8 |
| G19 | 315 | 3.28 (±0.47) | 5.86 (4) | 0.210 | 382.19 (336.98–427.66) | 23.6 | 9.8 |
| G20 | 315 | 2.10 (±0.33) | 5.41 (4) | 0.248 | 559.62 (463.01–667.15) | 34.5 | 14.3 |
| G21 | 270 | 4.07 (±0.65) | 6.01 (3) | 0.111 | 858.75 (773.01–983.36) | 53.0 | 22.0 |
| G22 | 315 | 3.44 (±0.49) | 4.00 (4) | 0.406 | 763.95 (674.62–853.34) | 47.2 | 19.5 |
| G23 | 315 | 5.70 (±0.69) | 2.80 (4) | 0.592 | 867.57 (805.36–939.79) | 53.6 | 22.2 |
| G24 | 315 | 2.70 (±0.45) | 1.96 (4) | 0.744 | 769.81 (661.09–880.96) | 47.5 | 19.7 |
| G25 | 270 | 3.87 (±0.48) | 0.87 (3) | 0.833 | 805.16 (718.30–900.54) | 49.7 | 20.6 |
| G26 | 315 | 2.80 (±0.28) | 5.40 (4) | 0.248 | 755.55 (638.24–887.94) | 46.6 | 19.3 |
| G27 | 270 | 2.57 (±0.37) | 6.41 (3) | 0.094 | 730.38 (621.35–888.24) | 45.1 | 18.7 |
| G28 | 315 | 3.06 (±0.33) | 4.28 (4) | 0.370 | 953.99 (833.19–1108.69) | 58.9 | 24.4 |
| G29 | 270 | 2.68 (±0.32) | 3.89 (3) | 0.274 | 793.50 (664.17–953.40) | 49.0 | 20.3 |
| G30 | 315 | 3.37 (±0.38) | 0.66 (4) | 0.956 | 971.23 (861.86–1093.52) | 60.0 | 24.8 |
| Insecticides | Strains | Slope (±SE) | χ2 (df) | p Value | LC50 (95%CI) mg/L | RR a | CR b |
|---|---|---|---|---|---|---|---|
| Acetamiprid | UNSEL | 2.34 (±0.24) | 0.66 (4) | 0.956 | 39.42 (32.61–46.89) | - | |
| G24 | 2.70 (±0.45) | 1.96 (4) | 0.744 | 769.81 (661.09–880.96) | 19.5 | ||
| G30 | 3.37 (±0.38) | 0.66 (4) | 0.956 | 971.23 (861.86–1093.52) | 24.6 | ||
| Thiamethoxam | UNSEL | 1.37 (±0.18) | 6.99 (4) | 0.136 | 32.20 (24.18–45.18) | - | |
| G24 | 3.20 (±0.32) | 2.83 (4) | 0.586 | 722.78 (625.86–826.48) | 22.5 | ||
| G30 | 2.88 (±0.30) | 5.77 (4) | 0.217 | 824.08 (712.78–953.62) | 26.0 | ||
| Nitenpyram | UNSEL | 2.05 (±0.22) | 2.24 (4) | 0.691 | 2.96 (2.40–3.68) | - | |
| G24 | 2.65 (±0.31) | 3.44 (3) | 0.151 | 61.64 (50.49–73.60) | 20.8 | ||
| G30 | 2.30 (±0.26) | 2.29 (4) | 0.665 | 63.20 (51.41–75.36) | 21.4 | ||
| Imidacloprid | UNSEL | 1.91 (±0.26) | 3.64 (4) | 0.457 | 125.54 (110.07–162.34) | ||
| G24 | 2.35 (±0.32) | 2.86 (3) | 0.414 | 1655.61 (1384.60–2051.04) | 13.2 | ||
| G30 | 2.52 (±0.39) | 4.98 (4) | 0.289 | 1837.57 (1571.23–2175.77) | 14.6 | ||
| Cycloxaprid | UNSEL | 1.66 (±0.21) | 2.53 (4) | 0.639 | 14.10 (10.95–18.67) | - | |
| G24 | 2.82 (±0.33) | 4.23 (4) | 0.376 | 159.99 (137.05–186.37) | 11.4 | ||
| G30 | 2.76 (±0.30) | 1.96 (4) | 0.743 | 168.81 (145.55–196.41) | 12.0 | ||
| Dinotefuran | UNSEL | 1.44 (±0.18) | 2.81 (4) | 0.560 | 21.30 (16.04–28.02) | - | |
| G24 | 2.71 (±0.34) | 2.59 (4) | 0.628 | 164.31 (142.28–195.58) | 7.7 | ||
| G30 | 2.63 (±0.29) | 2.02 (4) | 0.732 | 185.93 (159.41–222.21) | 8.7 | ||
| Sulfoxaflor | UNSEL | 2.75 (±0.28) | 1.95 (4) | 0.745 | 7.80 (6.58–9.50) | - | |
| G24 | 3.03 (±0.36) | 1.62 (4) | 0.806 | 40.01 (35.09–46.73) | 5.1 | ||
| G30 | 2.91 (±0.40) | 1.43 (3) | 0.670 | 59.25 (51.32–69.39) | 7.6 | ||
| Clothianidin | UNSEL | 1.55 (±0.19) | 2.52 (4) | 0.640 | 29.81 (23.04–45.18) | - | |
| G24 | 3.28 (±0.42) | 2.01 (3) | 0.570 | 143.64 (124.36–167.85) | 4.8 | ||
| G30 | 2.68 (±0.33) | 4.02 (3) | 0.259 | 151.48 (19.06–181.03) | 5.1 | ||
| Isoprocarb | UNSEL | 2.09 (±0.26) | 1.65 (3) | 0.647 | 77.09 (61.69–95.87) | - | |
| G24 | 2.19 (±0.31) | 5.71 (3) | 0.127 | 602.64 (487.94–733.52) | 7.8 | ||
| G30 | 1.86 (±0.22) | 4.93 (4) | 0.295 | 633.55 (507.88–804.32) | 8.2 | ||
| Chlorpyrifos | UNSEL | 2.77 (±0.37) | 2.23 (3) | 0.526 | 18.99 (16.09–23.64) | - | |
| G24 | 3.82 (±0.45) | 3.50 (3) | 0.321 | 25.45 (22.48–28.61) | 1.3 | ||
| G30 | 2.80 (±0.33) | 6.05 (3) | 0.109 | 32.43 (27.63–38.00) | 1.7 | ||
| Buprofezin | UNSEL | 1.53 (±0.20) | 1.65 (4) | 0.800 | 99.81 (75.78–142.17) | - | |
| G24 | 1.29 (±0.22) | 1.23 (3) | 0.745 | 146.20 (103.14–202.52) | 1.5 | ||
| G30 | 1.70 (±0.23) | 0.20 (3) | 0.978 | 152.44 (118.07–195.39) | 1.5 | ||
| Etofenprox | UNSEL | 1.71 (±0.23) | 3.73 (4) | 0.444 | 121.45 (96.68–157.90) | - | |
| G24 | 1.78 (±0.20) | 0.71 (4) | 0.950 | 110.02 (85.17–138.46) | 0.9 | ||
| G30 | 1.67 (±0.20) | 0.70(4) | 0.951 | 138.67 (108.14–177.42) | 1.1 |
| Strain | Acetamiprid/Synergist | Slope (±SE) | χ2 (df) | p Value | LC50 (95%CI) mg/L | SR a | RSR b |
|---|---|---|---|---|---|---|---|
| AC-S | Acetamiprid | 2.18 (±0.27) | 0.94 (3) | 0.816 | 27.27 (21.74–33.53) | ||
| Acetamiprid + PBO | 1.93 (±0.25) | 0.19 (3) | 0.980 | 21.00 (15.98–26.21) | 1.3 | ||
| Acetamiprid + TPP | 1.83 (±0.20) | 2.06 (4) | 0.726 | 17.23 (13.58–21.38) | 1.6 | ||
| Acetamiprid + DEM | 2.20 (±0.26) | 1.93 (3) | 0.586 | 26.56 (21.50–32.32) | 1.0 | ||
| AC-R(G28) | Acetamiprid | 3.13 (±0.41) | 3.83 (4) | 0.429 | 1898.71 (1645.95–2237.97) | ||
| Acetamiprid + PBO | 2.11 (±0.24) | 2.91 (4) | 0.572 | 676.31 (535.17–825.02) | 2.8 | 2.2 | |
| Acetamiprid + TPP | 1.47 (±0.20) | 0.97 (4) | 0.442 | 871.68 (662.61–1234.75) | 2.2 | 1.4 | |
| Acetamiprid + DEM | 2.48 (±0.28) | 5.80 (4) | 0.215 | 1685.20 (1434.62–1986.06) | 1.1 | 1.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, S.; He, M.; Xia, F.; Zhao, X.; Liao, X.; Li, R.; Li, M. The Cross-Resistance Pattern and the Metabolic Resistance Mechanism of Acetamiprid in the Brown Planthopper, Nilaparvata lugens (Stål). Int. J. Mol. Sci. 2022, 23, 9429. https://doi.org/10.3390/ijms23169429
Wu S, He M, Xia F, Zhao X, Liao X, Li R, Li M. The Cross-Resistance Pattern and the Metabolic Resistance Mechanism of Acetamiprid in the Brown Planthopper, Nilaparvata lugens (Stål). International Journal of Molecular Sciences. 2022; 23(16):9429. https://doi.org/10.3390/ijms23169429
Chicago/Turabian StyleWu, Shuai, Minrong He, Fujin Xia, Xueyi Zhao, Xun Liao, Rongyu Li, and Ming Li. 2022. "The Cross-Resistance Pattern and the Metabolic Resistance Mechanism of Acetamiprid in the Brown Planthopper, Nilaparvata lugens (Stål)" International Journal of Molecular Sciences 23, no. 16: 9429. https://doi.org/10.3390/ijms23169429
APA StyleWu, S., He, M., Xia, F., Zhao, X., Liao, X., Li, R., & Li, M. (2022). The Cross-Resistance Pattern and the Metabolic Resistance Mechanism of Acetamiprid in the Brown Planthopper, Nilaparvata lugens (Stål). International Journal of Molecular Sciences, 23(16), 9429. https://doi.org/10.3390/ijms23169429







