Isolation and Characterization of Urinary Extracellular Vesicles from Healthy Donors and Patients with Castration-Resistant Prostate Cancer
Abstract
:1. Introduction
2. Materials and Methods
2.1. Urine Collection and Pretreatment
2.2. Urinary EV Isolation Using Ultrafiltration and SEC
2.3. Urinary EV Isolation Using Polyethylene Glycol-Based Precipitation
2.4. Quantification of EV Particles and Proteins
2.5. Transmission Electron Microscopy (TEM)
2.6. Western Blot Analysis
2.7. RNA Extraction and Real-Time Polymerase Chain Reaction (RT-PCR)
2.8. Statistical Analysis
3. Results and Discussion
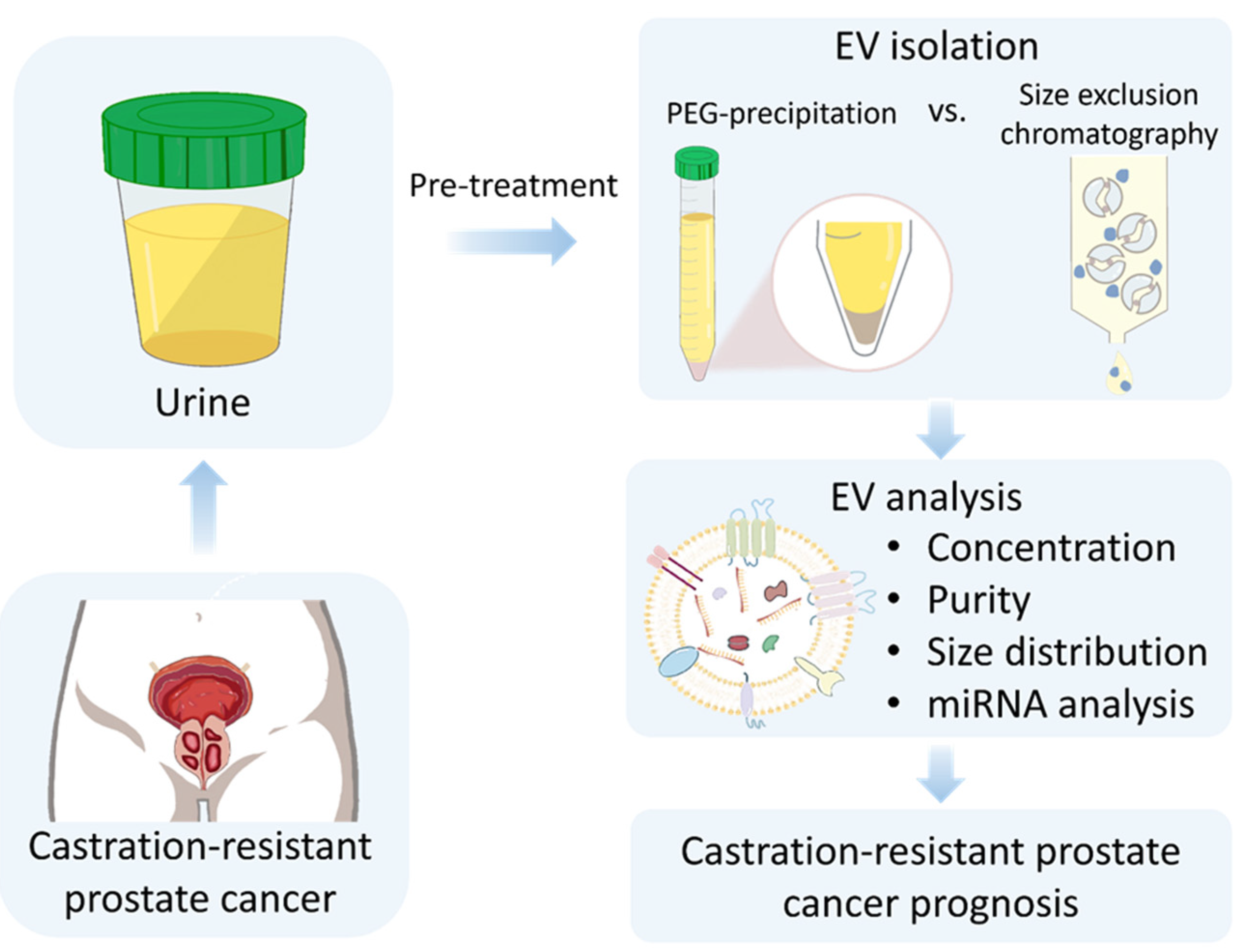
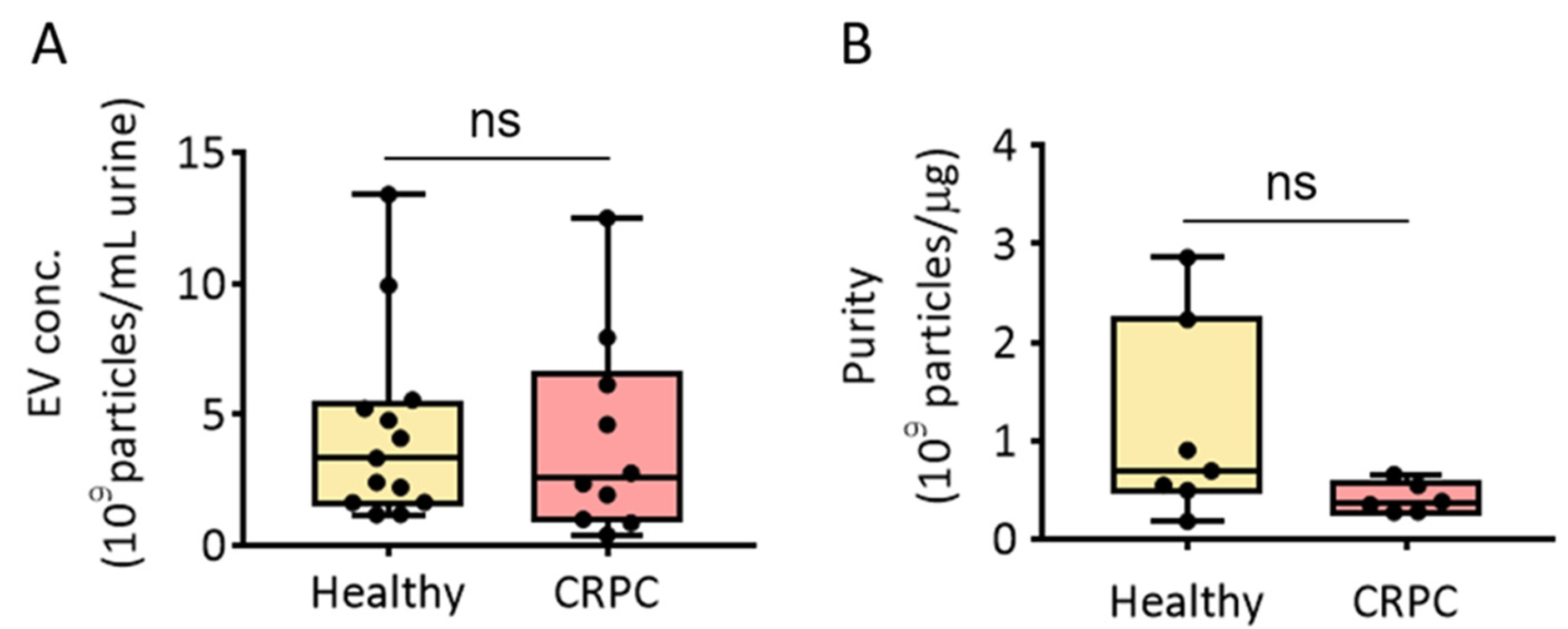
3.1. PEG Precipitation-Based Isolation of EVs from Human Urine
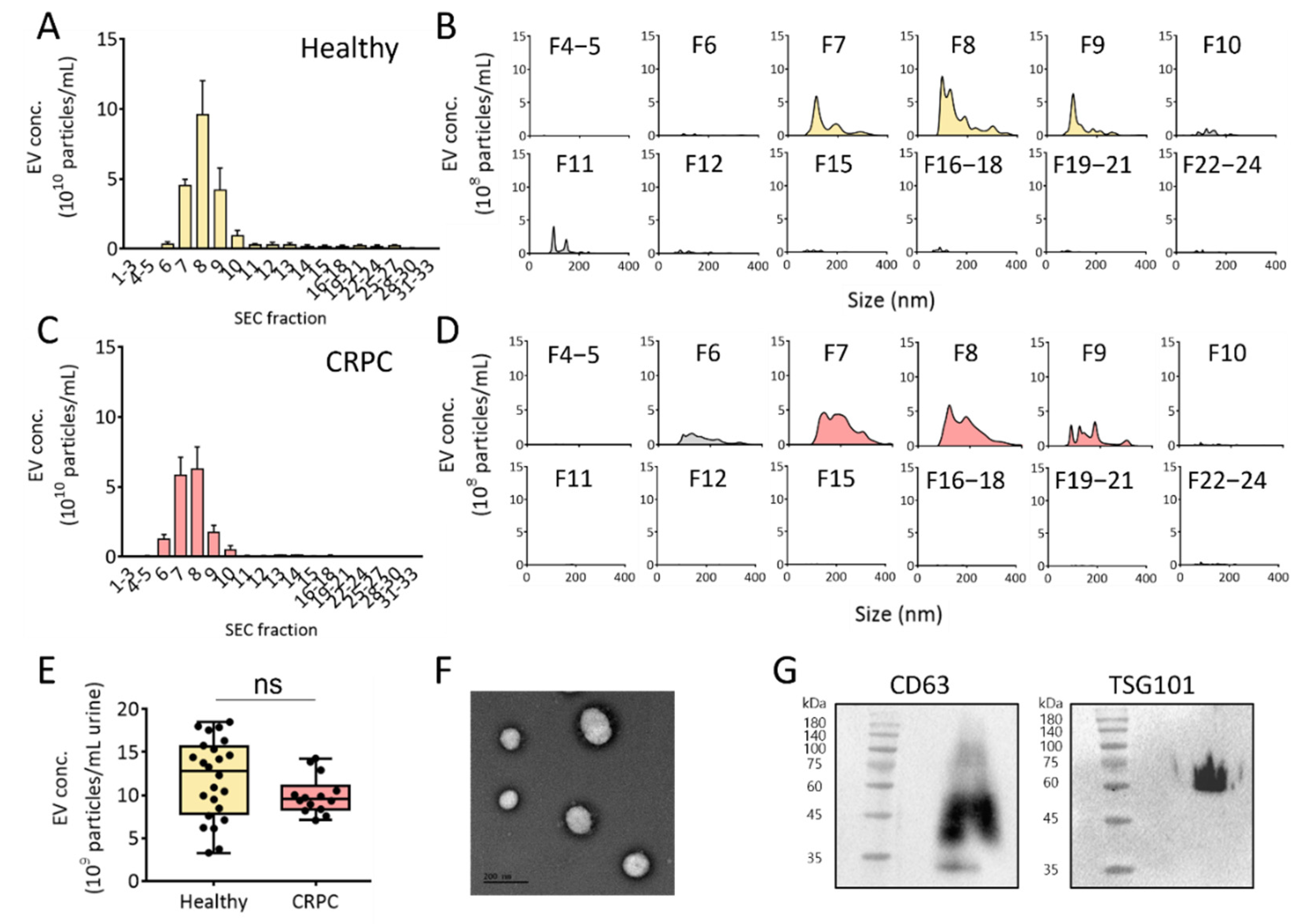
3.2. SEC-Based Isolation of EVs from Human Urine
3.3. Comparison of PEG Precipitation and SEC for Urinary EV Isolation
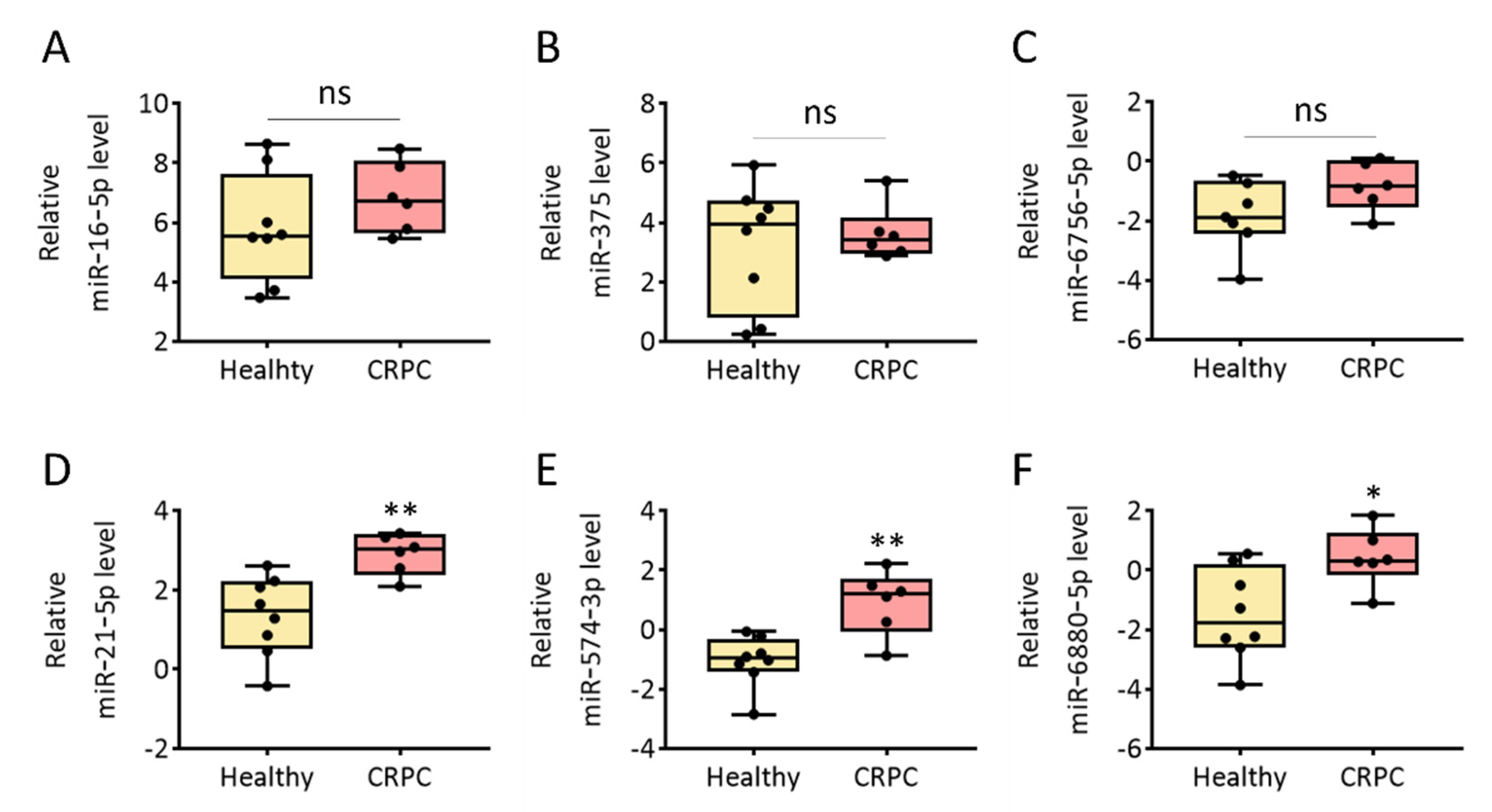
3.4. Evaluation of Cancer-Related miRNA Levels in Urinary EVs from Healthy Donors and Patients with CRPC
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer Statistics, 2017. CA Cancer J. Clin. 2017, 67, 7–30. [Google Scholar] [CrossRef] [Green Version]
- Conteduca, V.; Mosca, A.; Brighi, N.; de Giorgi, U.; Rescigno, P. New Prognostic Biomarkers in Metastatic Castration-Resistant Prostate Cancer. Cells 2021, 10, 193. [Google Scholar] [CrossRef] [PubMed]
- Kirby, M.; Hirst, C.; Crawford, E. Characterising the castration-resistant prostate cancer population: A systematic review. Int. J. Clin. Pract. 2011, 65, 1180–1192. [Google Scholar] [CrossRef] [PubMed]
- Obinata, D.; Takayama, K.; Takahashi, S.; Inoue, S. Crosstalk of the Androgen Receptor with Transcriptional Collaborators: Potential Therapeutic Targets for Castration-Resistant Prostate Cancer. Cancers 2017, 9, 22. [Google Scholar] [CrossRef] [Green Version]
- Prensner, J.R.; Rubin, M.A.; Wei, J.T.; Chinnaiyan, A.M. Beyond PSA: The next generation of prostate cancer biomarkers. Sci. Transl. Med. 2012, 4, 127rv3. [Google Scholar] [CrossRef] [Green Version]
- Hoffman, R.M. Screening for prostate cancer. New Engl. J. Med. 2011, 365, 2013–2019. [Google Scholar] [CrossRef] [PubMed]
- Kuner, R.; Brase, J.C.; Sültmann, H.; Wuttig, D. microRNA biomarkers in body fluids of prostate cancer patients. Methods 2013, 59, 132–137. [Google Scholar] [CrossRef] [PubMed]
- Ochodo, E.A.; Gopalakrishna, G.; Spek, B.; Reitsma, J.B.; Van Lieshout, L.; Polman, K.; Lamberton, P.; Bossuyt, P.M.; Leeflang, M.M. Circulating antigen tests and urine reagent strips for diagnosis of active schistosomiasis in endemic areas. Cochrane Database Syst. Rev. 2015, 2015, CD009579. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, A.; Péoc’h, M.; Cottier, M.; Genin, C.; Mottet, N.; Li, G. Cell-free RNA content in urine as a possible molecular diagnostic tool for clear cell renal cell carcinoma. Int. J. Cancer 2015, 136, 2610–2615. [Google Scholar] [CrossRef]
- Yu, W.; Hurley, J.; Roberts, D.; Chakrabortty, S.K.; Enderle, D.; Noerholm, M.; Breakefield, X.O.; Skog, J.K. Exosome-based liquid biopsies in cancer: Opportunities and challenges. Ann. Oncol. 2021, 32, 466–477. [Google Scholar] [CrossRef]
- Arraud, N.; Linares, R.; Tan, S.; Gounou, C.; Pasquet, J.M.; Mornet, S.; Brisson, A.R. Extracellular vesicles from blood plasma: Determination of their morphology, size, phenotype and concentration. J. Thromb. Haemost. 2014, 12, 614–627. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.; Yang, H.C.; Rhee, W.J. Development and comparative analysis of human urine exosome isolation strategies. Process Biochem. 2020, 88, 197–203. [Google Scholar] [CrossRef]
- Yang, H.C.; Ham, Y.M.; Kim, J.A.; Rhee, W.J. Single-step equipment-free extracellular vesicle concentration using super absorbent polymer beads. J. Extracell. Vesicles 2021, 10, e12074. [Google Scholar] [CrossRef] [PubMed]
- El Andaloussi, S.; Mäger, I.; Breakefield, X.O.; Wood, M.J. Extracellular vesicles: Biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Macias, M.; Alegre, E.; Diaz-Lagares, A.; Patino, A.; Perez-Gracia, J.L.; Sanmamed, M.; Lopez-Lopez, R.; Varo, N.; Gonzalez, A. Liquid Biopsy: From Basic Research to Clinical Practice. Adv. Clin. Chem. 2018, 83, 73–119. [Google Scholar] [PubMed]
- Erdbrugger, U.; Blijdorp, C.J.; Bijnsdorp, I.V.; Borras, F.E.; Burger, D.; Bussolati, B.; Byrd, J.B.; Clayton, A.; Dear, J.W.; Falcon-Perez, J.M.; et al. Urinary extracellular vesicles: A position paper by the Urine Task Force of the International Society for Extracellular Vesicles. J. Extracell. Vesicles 2021, 10, e12093. [Google Scholar] [CrossRef]
- Jeong, K.; Yu, Y.J.; You, J.Y.; Rhee, W.J.; Kim, J.A. Exosome-mediated microRNA-497 delivery for anti-cancer therapy in a microfluidic 3D lung cancer model. Lab Chip 2020, 20, 548–557. [Google Scholar] [CrossRef]
- Kim, H.; Kang, S.J.; Rhee, W.J. Phenylboronic Acid-conjugated Exosomes for Enhanced Anticancer Therapeutic Effect by Increasing Doxorubicin Loading Efficiency. Biotechnol. Bioprocess Eng. 2021, 26, 78–85. [Google Scholar] [CrossRef]
- Cao, T.G.N.; Kang, J.H.; Kim, W.; Lim, J.; Kang, S.J.; You, J.Y.; Hoang, Q.T.; Kim, W.J.; Rhee, W.J.; Kim, C. Engineered extracellular vesicle-based sonotheranostics for dual stimuli-sensitive drug release and photoacoustic imaging-guided chemo-sonodynamic cancer therapy. Theranostics 2022, 12, 1247. [Google Scholar]
- You, J.Y.; Kang, S.J.; Rhee, W.J. Isolation of cabbage exosome-like nanovesicles and investigation of their biological activities in human cells. Bioact. Mater. 2021, 6, 4321–4332. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Rhee, W.J. Exosome-mediated let7c-5p delivery for breast cancer therapeutic development. Biotechnol. Bioprocess Eng. 2020, 25, 513–520. [Google Scholar] [CrossRef]
- Dhondt, B.; Geeurickx, E.; Tulkens, J.; Van Deun, J.; Vergauwen, G.; Lippens, L.; Miinalainen, I.; Rappu, P.; Heino, J.; Ost, P.; et al. Unravelling the proteomic landscape of extracellular vesicles in prostate cancer by density-based fractionation of urine. J. Extracell. Vesicles 2020, 9, 1736935. [Google Scholar] [CrossRef] [PubMed]
- Merchant, M.L.; Rood, I.M.; Deegens, J.K.; Klein, J.B. Isolation and characterization of urinary extracellular vesicles: Implications for biomarker discovery. Nat. Rev. Nephrol. 2017, 13, 731–749. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rider, M.A.; Hurwitz, S.N.; Meckes, D.G., Jr. ExtraPEG: A Polyethylene Glycol-Based Method for Enrichment of Extracellular Vesicles. Sci. Rep. 2016, 6, 23978. [Google Scholar] [CrossRef]
- Lourenco, C.; Constancio, V.; Henrique, R.; Carvalho, A.; Jeronimo, C. Urinary Extracellular Vesicles as Potential Biomarkers for Urologic Cancers: An Overview of Current Methods and Advances. Cancers 2021, 13, 1529. [Google Scholar] [CrossRef]
- Fernandez-Llama, P.; Khositseth, S.; Gonzales, P.A.; Star, R.A.; Pisitkun, T.; Knepper, M.A. Tamm-Horsfall protein and urinary exosome isolation. Kidney Int. 2010, 77, 736–742. [Google Scholar] [CrossRef] [Green Version]
- Gheinani, A.H.; Vögeli, M.; Baumgartner, U.; Vassella, E.; Draeger, A.; Burkhard, F.C.; Monastyrskaya, K. Improved isolation strategies to increase the yield and purity of human urinary exosomes for biomarker discovery. Sci. Rep. 2018, 8, 3945. [Google Scholar] [CrossRef]
- Cho, S.; Yang, H.C.; Rhee, W.J. Simultaneous multiplexed detection of exosomal microRNAs and surface proteins for prostate cancer diagnosis. Biosens. Bioelectron. 2019, 146, 111749. [Google Scholar] [CrossRef]
- Kojima, M.; Sudo, H.; Kawauchi, J.; Takizawa, S.; Kondou, S.; Nobumasa, H.; Ochiai, A. MicroRNA markers for the diagnosis of pancreatic and biliary-tract cancers. PLoS ONE 2015, 10, e0118220. [Google Scholar] [CrossRef] [Green Version]
- Cui, X.; Li, Z.; Zhao, Y.; Song, A.; Shi, Y.; Hai, X.; Zhu, W. Breast cancer identification via modeling of peripherally circulating miRNAs. PeerJ 2018, 6, e4551. [Google Scholar] [CrossRef]
- Hessvik, N.P.; Sandvig, K.; Llorente, A. Exosomal miRNAs as Biomarkers for Prostate Cancer. Front. Genet. 2013, 4, 36. [Google Scholar] [CrossRef] [Green Version]
- Salehi, M.; Sharifi, M. Exosomal miRNAs as novel cancer biomarkers: Challenges and opportunities. J. Cell. Physiol. 2018, 233, 6370–6380. [Google Scholar] [CrossRef] [PubMed]
- Foj, L.; Ferrer, F.; Serra, M.; Arevalo, A.; Gavagnach, M.; Gimenez, N.; Filella, X. Exosomal and Non-Exosomal Urinary miRNAs in Prostate Cancer Detection and Prognosis. Prostate 2017, 77, 573–583. [Google Scholar] [CrossRef] [PubMed]
- Bryzgunova, O.E.; Zaripov, M.M.; Skvortsova, T.E.; Lekchnov, E.A.; Grigor’eva, A.E.; Zaporozhchenko, I.A.; Morozkin, E.S.; Ryabchikova, E.I.; Yurchenko, Y.B.; Voitsitskiy, V.E.; et al. Comparative Study of Extracellular Vesicles from the Urine of Healthy Individuals and Prostate Cancer Patients. PLoS ONE 2016, 11, e0157566. [Google Scholar] [CrossRef]
- Kinoshita, T.; Yip, K.W.; Spence, T.; Liu, F.F. MicroRNAs in extracellular vesicles: Potential cancer biomarkers. J. Hum. Genet. 2017, 62, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Lange, T.; Stracke, S.; Rettig, R.; Lendeckel, U.; Kuhn, J.; Schluter, R.; Rippe, V.; Endlich, K.; Endlich, N. Identification of miR-16 as an endogenous reference gene for the normalization of urinary exosomal miRNA expression data from CKD patients. PLoS ONE 2017, 12, e0183435. [Google Scholar] [CrossRef]
- Lin, H.M.; Mahon, K.L.; Spielman, C.; Gurney, H.; Mallesara, G.; Stockler, M.R.; Bastick, P.; Briscoe, K.; Marx, G.; Swarbrick, A.; et al. Phase 2 study of circulating microRNA biomarkers in castration-resistant prostate cancer. Br. J. Cancer 2017, 116, 1002–1011. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Lieberman, R.; Pan, J.; Zhang, Q.; Du, M.; Zhang, P.; Nevalainen, M.; Kohli, M.; Shenoy, N.K.; Meng, H.; et al. miR-375 induces docetaxel resistance in prostate cancer by targeting SEC23A and YAP1. Mol. Cancer 2016, 15, 70. [Google Scholar] [CrossRef] [Green Version]
- Das, D.K.; Persaud, L.; Sauane, M. MicroRNA-4719 and microRNA-6756-5p Correlate with Castration-Resistant Prostate Cancer Progression through Interleukin-24 Regulation. Noncoding RNA 2019, 5, 10. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, D.P.; Li, J.; Tewari, A.K. Inflammation and prostate cancer: The role of interleukin 6 (IL-6). BJU Int. 2014, 113, 986–992. [Google Scholar] [CrossRef]
- Guan, Y.; Wu, Y.; Liu, Y.; Ni, J.; Nong, S. Association of microRNA-21 expression with clinicopathological characteristics and the risk of progression in advanced prostate cancer patients receiving androgen deprivation therapy. Prostate 2016, 76, 986–993. [Google Scholar] [CrossRef]
- Zennami, K.; Choi, S.M.; Liao, R.; Li, Y.; Dinalankara, W.; Marchionni, L.; Rafiqi, F.H.; Kurozumi, A.; Hatano, K.; Lupold, S.E. PDCD4 is an androgen-repressed tumor suppressor that regulates prostate cancer growth and castration resistance. Mol. Cancer Res. 2019, 17, 618–627. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, L.-Q.; Yang, X.-W.; Chen, Y.-B.; Zhang, D.-W.; Jiang, X.-F.; Xue, P. Exosomal miR-21 regulates the TETs/PTENp1/PTEN pathway to promote hepatocellular carcinoma growth. Mol. Cancer 2019, 18, 148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Samsonov, R.; Shtam, T.; Burdakov, V.; Glotov, A.; Tsyrlina, E.; Berstein, L.; Nosov, A.; Evtushenko, V.; Filatov, M.; Malek, A. Lectin-induced agglutination method of urinary exosomes isolation followed by mi-RNA analysis: Application for prostate cancer diagnostic. Prostate 2016, 76, 68–79. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, N.M.; Imran, K.; Kastner, D.W.; Ikram, K.; Mushtaq, A.; Hussain, A.; Zeeshan, N. Potential involvement of mi-RNA 574-3p in progression of prostate cancer: A bioinformatic study. Mol. Cell. Probes 2017, 36, 21–28. [Google Scholar] [CrossRef]

| Age (years) | Tumor Stage | Gleason Score | PSA (ng/mL) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Mean ± SD | Range | T4 | 6 | 7 | 8 | 9 | Mean ± SD | Range | |
| Patients with CRPC (n = 6) | 76.8 ± 5.9 | 68–83 | 6 | 1 | 2 | 2 | 1 | 171.1 ± 254.2 | 0.236–600 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, H.; Kang, S.J.; Lee, J.; Park, K.H.; Rhee, W.J. Isolation and Characterization of Urinary Extracellular Vesicles from Healthy Donors and Patients with Castration-Resistant Prostate Cancer. Int. J. Mol. Sci. 2022, 23, 7134. https://doi.org/10.3390/ijms23137134
Lee H, Kang SJ, Lee J, Park KH, Rhee WJ. Isolation and Characterization of Urinary Extracellular Vesicles from Healthy Donors and Patients with Castration-Resistant Prostate Cancer. International Journal of Molecular Sciences. 2022; 23(13):7134. https://doi.org/10.3390/ijms23137134
Chicago/Turabian StyleLee, Haneul, Su Jin Kang, Jimin Lee, Kyong Hwa Park, and Won Jong Rhee. 2022. "Isolation and Characterization of Urinary Extracellular Vesicles from Healthy Donors and Patients with Castration-Resistant Prostate Cancer" International Journal of Molecular Sciences 23, no. 13: 7134. https://doi.org/10.3390/ijms23137134
APA StyleLee, H., Kang, S. J., Lee, J., Park, K. H., & Rhee, W. J. (2022). Isolation and Characterization of Urinary Extracellular Vesicles from Healthy Donors and Patients with Castration-Resistant Prostate Cancer. International Journal of Molecular Sciences, 23(13), 7134. https://doi.org/10.3390/ijms23137134








