Implication of microRNAs in Carcinogenesis with Emphasis on Hematological Malignancies and Clinical Translation
Abstract
1. Introduction
2. Implication of miRNAs in Carcinogenesis and Leukemogenesis
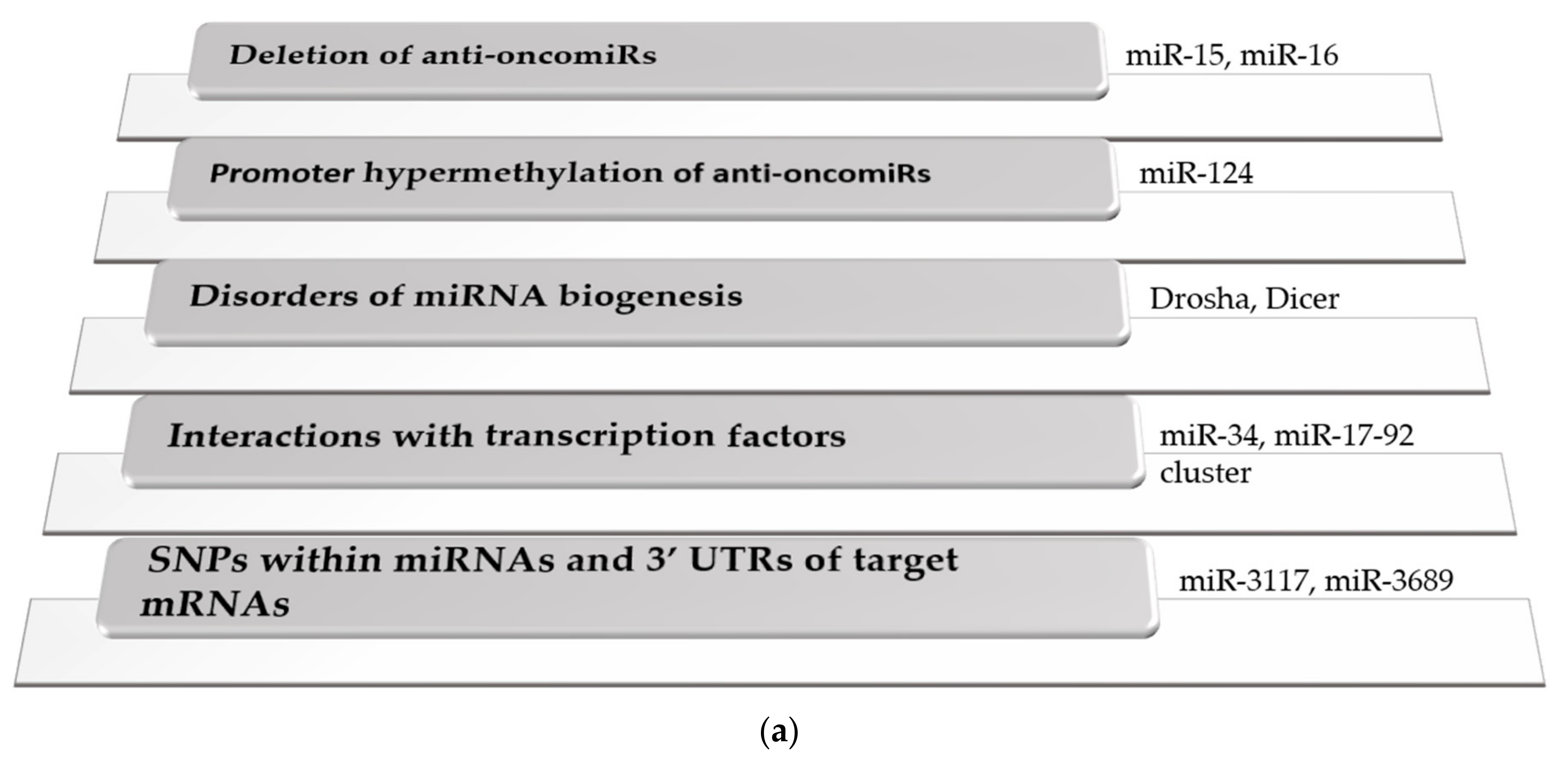
2.1. Dysregulation of miRNA Expression Levels in Cancer
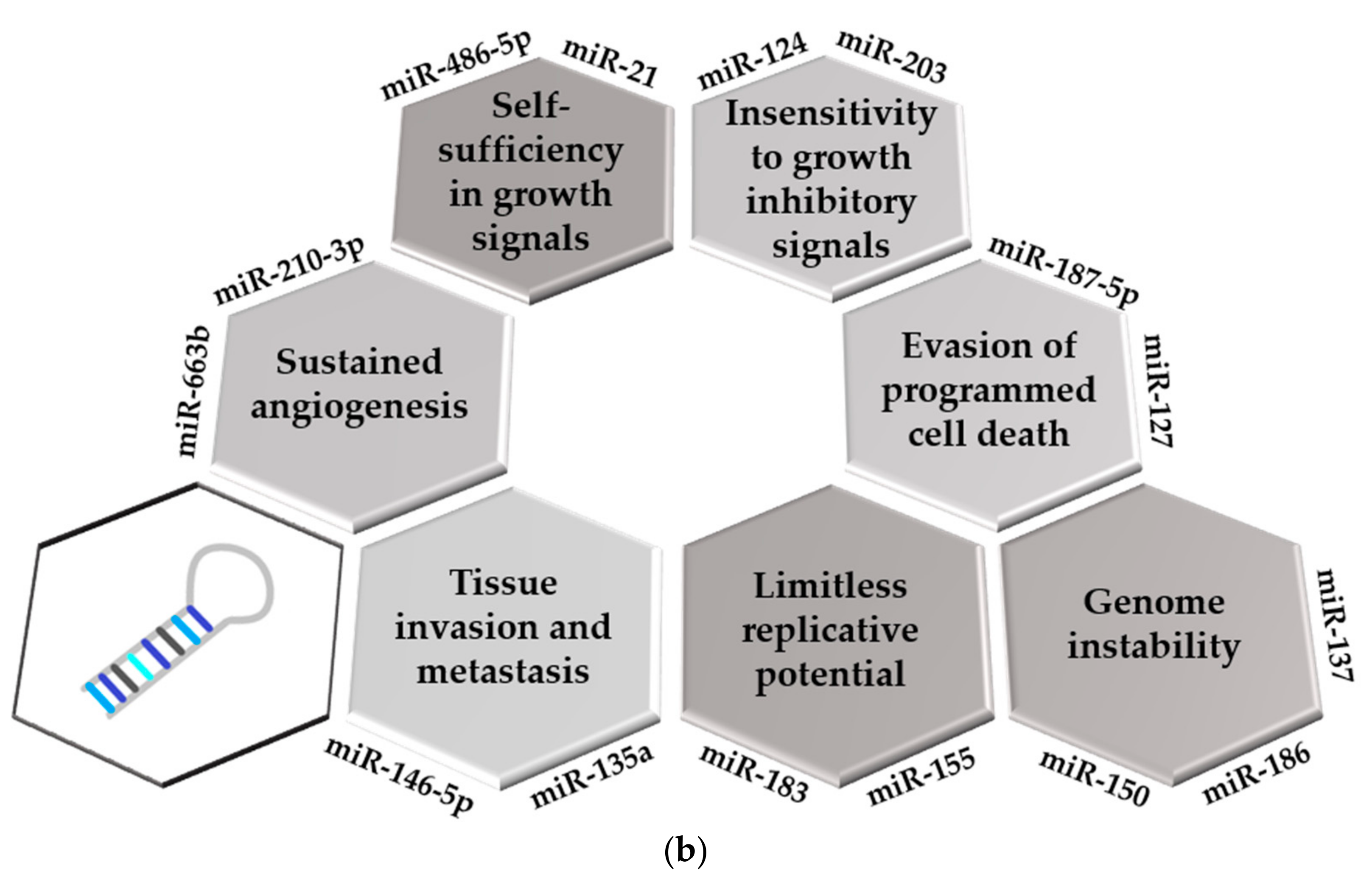
2.2. Implication of miRNAs in the Development of the Hallmarks of Cancer
2.3. miRNAs and Metabolic Reprogramming in Leukemia
2.4. Interactions between miRNAs and Epigenetic Regulatory Circuits
3. Regulation of Hematopoiesis by miRNAs
4. Clinical Applications of miRNAs Focusing on Hematological Malignancies
4.1. Differential Diagnosis and Early Detection of Hematological Malignancies
4.2. Advances in Prognostic Stratification of Hematological Malignancies
4.3. Implication of miRNAs in Chemosensitivity, Prediction of Chemoresistance in Hematological Malignancies
4.4. Changes of miRNA Expression Levels during Chemotherapy
4.5. miRNA-Based Anticancer Therapeutic Approaches
4.6. Clinical Applications of Exosomal miRNAs
5. Concluding Remarks and Future Perspectives
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Tafrihi, M.; Hasheminasab, E. miRNAs: Biology, Biogenesis, their Web-based Tools, and Databases. Microrna 2019, 8, 4–27. [Google Scholar] [CrossRef] [PubMed]
- Lai, P.; Wang, Y. Epigenetics of cutaneous T-cell lymphoma: Biomarkers and therapeutic potentials. Cancer Biol. Med. 2021, 18, 34–51. [Google Scholar] [CrossRef] [PubMed]
- Rupaimoole, R.; Slack, F. MicroRNA therapeutics: Towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 2017, 16, 203–222. [Google Scholar] [CrossRef] [PubMed]
- Matsuyama, H.; Suzuki, H.I. Systems and Synthetic microRNA Biology: From Biogenesis to Disease Pathogenesis. Int. J. Mol. Sci. 2019, 21, 132. [Google Scholar] [CrossRef]
- Calin, G.A.; Ferracin, M.; Cimmino, A.; Di Leva, G.; Shimizu, M.; Wojcik, S.E.; Iorio, M.V.; Visone, R.; Sever, N.I.; Fabbri, M.; et al. A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N. Engl. J. Med. 2005, 353, 1793–1801. [Google Scholar] [CrossRef]
- Huang, Z.; Xu, Y.; Wan, M.; Zeng, X.; Wu, J. miR-340: A multifunctional role in human malignant diseases. Int. J. Biol. Sci. 2021, 17, 236–246. [Google Scholar] [CrossRef]
- Artemaki, P.I.; Letsos, P.A.; Zoupa, I.C.; Katsaraki, K.; Karousi, P.; Papageorgiou, S.G.; Pappa, V.; Scorilas, A.; Kontos, C.K. The Multifaceted Role and Utility of MicroRNAs in Indolent B-Cell Non-Hodgkin Lymphomas. Biomedicines 2021, 9, 333. [Google Scholar] [CrossRef]
- Yeh, C.H.; Moles, R.; Nicot, C. Clinical significance of microRNAs in chronic and acute human leukemia. Mol. Cancer 2016, 15, 37. [Google Scholar] [CrossRef]
- Calin, G.A.; Sevignani, C.; Dumitru, C.D.; Hyslop, T.; Noch, E.; Yendamuri, S.; Shimizu, M.; Rattan, S.; Bullrich, F.; Negrini, M.; et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl. Acad. Sci. USA 2004, 101, 2999–3004. [Google Scholar] [CrossRef]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 15524–15529. [Google Scholar] [CrossRef]
- Peng, Y.; Croce, C. The role of MicroRNAs in human cancer. Signal Transduct. Target. Ther. 2016, 1, 15004. [Google Scholar] [CrossRef]
- Szczepanek, J. Role of microRNA dysregulation in childhood acute leukemias: Diagnostics, monitoring and therapeutics: A comprehensive review. World J. Clin. Oncol. 2020, 11, 348–369. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef]
- Schotte, D.; De Menezes, R.X.; Akbari Moqadam, F.; Khankahdani, L.M.; Lange-Turenhout, E.; Chen, C.; Pieters, R.; Den Boer, M.L. MicroRNA characterize genetic diversity and drug resistance in pediatric acute lymphoblastic leukemia. Haematologica 2011, 96, 703–711. [Google Scholar] [CrossRef]
- Zhang, L.; Nguyen, L.X.T.; Chen, Y.C.; Wu, D.; Cook, G.J.; Hoang, D.H.; Brewer, C.J.; He, X.; Dong, H.; Li, S.; et al. Targeting miR-126 in inv(16) acute myeloid leukemia inhibits leukemia development and leukemia stem cell maintenance. Nat. Commun. 2021, 12, 6154. [Google Scholar] [CrossRef]
- Havelange, V.; Ranganathan, P.; Geyer, S.; Nicolet, D.; Huang, X.; Yu, X.; Volinia, S.; Kornblau, S.M.; Andreeff, M.; Croce, C.M.; et al. Implications of the miR-10 family in chemotherapy response of NPM1-mutated AML. Blood 2014, 123, 2412–2415. [Google Scholar] [CrossRef]
- Malouf, C.; Antunes, E.T.B.; O’Dwyer, M.; Jakobczyk, H.; Sahm, F.; Landua, S.L.; Anderson, R.A.; Soufi, A.; Halsey, C.; Ottersbach, K. miR-130b and miR-128a are essential lineage-specific codrivers of t(4;11) MLL-AF4 acute leukemia. Blood 2021, 138, 2066–2092. [Google Scholar] [CrossRef]
- Schwaller, J. MLL-AF4+ infant leukemia: A microRNA affair. Blood 2021, 138, 2014–2015. [Google Scholar] [CrossRef]
- Mardani, R.; Jafari Najaf Abadi, M.H.; Motieian, M.; Taghizadeh-Boroujeni, S.; Bayat, A.; Farsinezhad, A.; Hayat, S.M.G.; Motieian, M.; Pourghadamyari, H. MicroRNA in leukemia: Tumor suppressors and oncogenes with prognostic potential. J. Cell. Physiol. 2019, 234, 8465–8486. [Google Scholar] [CrossRef]
- Ryu, J.K.; Hong, S.M.; Karikari, C.A.; Hruban, R.H.; Goggins, M.G.; Maitra, A. Aberrant MicroRNA-155 expression is an early event in the multistep progression of pancreatic adenocarcinoma. Pancreatology 2010, 10, 66–73. [Google Scholar] [CrossRef]
- Manfe, V.; Biskup, E.; Willumsgaard, A.; Skov, A.G.; Palmieri, D.; Gasparini, P.; Lagana, A.; Woetmann, A.; Odum, N.; Croce, C.M.; et al. cMyc/miR-125b-5p signalling determines sensitivity to bortezomib in preclinical model of cutaneous T-cell lymphomas. PLoS ONE 2013, 8, e59390. [Google Scholar] [CrossRef] [PubMed]
- Hermeking, H. MicroRNAs in the p53 network: Micromanagement of tumour suppression. Nat. Rev. Cancer 2012, 12, 613–626. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, K.A.; Wentzel, E.A.; Zeller, K.I.; Dang, C.V.; Mendell, J.T. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005, 435, 839–843. [Google Scholar] [CrossRef] [PubMed]
- Szczyrek, M.; Grenda, A.; Kuźnar-Kamińska, B.; Krawczyk, P.; Sawicki, M.; Batura-Gabryel, H.; Mlak, R.; Szudy-Szczyrek, A.; Krajka, T.; Krajka, A.; et al. Methylation of DROSHA and DICER as a Biomarker for the Detection of Lung Cancer. Cancers 2021, 13, 6139. [Google Scholar] [CrossRef]
- Lee, S.S.; Min, H.; Ha, J.Y.; Kim, B.H.; Choi, M.S.; Kim, S. Dysregulation of the miRNA biogenesis components DICER1, DROSHA, DGCR8 and AGO2 in clear cell renal cell carcinoma in both a Korean cohort and the cancer genome atlas kidney clear cell carcinoma cohort. Oncol. Lett. 2019, 18, 4337–4345. [Google Scholar] [CrossRef]
- Vardapour, R.; Kehl, T.; Kneitz, S.; Ludwig, N.; Meese, E.; Lenhof, H.P.; Gessler, M. The DGCR8 E518K mutation found in Wilms tumors leads to a partial miRNA processing defect that alters gene expression patterns and biological processes. Carcinogenesis 2021, 43, 82–93. [Google Scholar] [CrossRef]
- Ramirez-Moya, J.; Wert-Lamas, L.; Riesco-Eizaguirre, G.; Santisteban, P. Impaired microRNA processing by DICER1 downregulation endows thyroid cancer with increased aggressiveness. Oncogene 2019, 38, 5486–5499. [Google Scholar] [CrossRef]
- Ali Syeda, Z.; Langden, S.S.S.; Munkhzul, C.; Lee, M.; Song, S.J. Regulatory Mechanism of MicroRNA Expression in Cancer. Int. J. Mol. Sci. 2020, 21, 1723. [Google Scholar] [CrossRef]
- Wen, J.; Lv, Z.; Ding, H.; Fang, X.; Sun, M. Association of miRNA biosynthesis genes DROSHA and DGCR8 polymorphisms with cancer susceptibility: A systematic review and meta-analysis. Biosci. Rep. 2018, 38, BSR20180072. [Google Scholar] [CrossRef]
- Kishan, A.U.; Marco, N.; Schulz-Jaavall, M.B.; Steinberg, M.L.; Tran, P.T.; Juarez, J.E.; Dang, A.; Telesca, D.; Lilleby, W.A.; Weidhaas, J.B. Germline variants disrupting microRNAs predict long-term genitourinary toxicity after prostate cancer radiation. Radiother. Oncol. 2022, 167, 226–232. [Google Scholar] [CrossRef]
- Mosallaei, M.; Simonian, M.; Esmaeilzadeh, E.; Bagheri, H.; Miraghajani, M.; Salehi, A.R.; Mehrzad, V.; Salehi, R. Single nucleotide polymorphism rs10889677 in miRNAs Let-7e and Let-7f binding site of IL23R gene is a strong colorectal cancer determinant: Report and meta-analysis. Cancer Genet. 2019, 239, 46–53. [Google Scholar] [CrossRef]
- Mohammadi, M.; Salehzadeh, A.; Talesh Sasani, S.; Tarang, A. rs6426881 in the 3′-UTR of PBX1 is involved in breast and gastric cancers via altering the binding potential of miR-522-3p. Mol. Biol. Rep. 2021, 48, 7405–7414. [Google Scholar] [CrossRef]
- Gutierrez-Camino, A.; Richer, C.; St-Onge, P.; Lopez-Lopez, E.; Bañeres, A.C.; de Andoin, N.G.; Sastre, A.; Astigarraaga, I.; Martin-Guerrero, I.; Sinnett, D.; et al. Role of rs10406069 in miR-5196 in hyperdiploid childhood acute lymphoblastic leukemia. Epigenomics 2020, 12, 1949–1955. [Google Scholar] [CrossRef]
- Gutierrez-Camino, A.; Martin-Guerrero, I.; Dolzan, V.; Jazbec, J.; Carbone-Bañeres, A.; Garcia de Andoin, N.; Sastre, A.; Astigarraga, I.; Navajas, A.; Garcia-Orad, A. Involvement of SNPs in miR-3117 and miR-3689d2 in childhood acute lymphoblastic leukemia risk. Oncotarget 2018, 9, 22907–22914. [Google Scholar] [CrossRef]
- Hassan, F.M. The Association of rs2114358 in the miR-1206 Polymorphism to Chronic Myeloid Leukemia. Microrna 2019, 8, 248–252. [Google Scholar] [CrossRef]
- Gutierrez-Camino, A.; Umerez, M.; Santos, B.; Martin-Guerrero, I.; García de Andoin, N.; Sastre, A.; Navajas, A.; Astigarraga, I.; Garcia-Orad, A. Pharmacoepigenetics in childhood acute lymphoblastic leukemia: Involvement of miRNA polymorphisms in hepatotoxicity. Epigenomics 2018, 10, 409–417. [Google Scholar] [CrossRef]
- Gutierrez-Camino, A.; Oosterom, N.; den Hoed, M.A.H.; Lopez-Lopez, E.; Martin-Guerrero, I.; Plujim, S.M.F.; Pieters, R.; de Jonge, R.; Tissing, W.J.E.; Heil, S.G.; et al. The miR1206 microRNA variant is associated with methotrexate-induced oral mucositis in pediatric acute lymphoblastic leukemia. Pharmacogenet. Genom. 2017, 27, 303–306. [Google Scholar] [CrossRef]
- Katsaraki, K.; Karousi, P.; Artemaki, P.I.; Scorilas, A.; Pappa, V.; Kontos, C.K.; Papageorgiou, S.G. MicroRNAs: Tiny Regulators of Gene Expression with Pivotal Roles in Normal B-Cell Development and B-Cell Chronic Lymphocytic Leukemia. Cancers 2021, 13, 593. [Google Scholar] [CrossRef]
- Chim, C.S.; Wan, T.S.; Wong, K.Y.; Fung, T.K.; Drexler, H.G.; Wong, K.F. Methylation of miR-34a, miR-34b/c, miR-124-1 and miR-203 in Ph-negative myeloproliferative neoplasms. J. Transl. Med. 2011, 9, 197. [Google Scholar] [CrossRef]
- Wong, K.Y.; So, C.C.; Loong, F.; Chung, L.P.; Lam, W.W.; Liang, R. Epigenetic inactivation of the miR-124-1 in haematological malignancies. PLoS ONE 2011, 6, e19027. [Google Scholar] [CrossRef]
- Paczkowska, J.; Janiszewska, J.; Bein, J.; Schneider, M.; Bednarek, K.; Ustaszewski, A.; Hartmann, S.; Hansmann, M.L.; Giefing, M. The Tumor Suppressive mir-148a Is Epigenetically Inactivated in Classical Hodgkin Lymphoma. Cells 2020, 9, 2292. [Google Scholar] [CrossRef] [PubMed]
- Wong, K.Y.; Liang, R.; So, C.C.; Jin, D.Y.; Costello, J.F.; Chim, C.S. Epigenetic silencing of MIR203 in multiple myeloma. Br. J. Haematol. 2011, 154, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- Shahid, S.; Shahid, W.; Shaheen, J.; Akhtar, M.W.; Sadaf, S. Circulating miR-146a expression as a non-invasive predictive biomarker for acute lymphoblastic leukemia. Sci. Rep. 2021, 11, 22783. [Google Scholar] [CrossRef]
- Cimmino, A.; Calin, G.A.; Fabbri, M.; Iorio, M.V.; Ferracin, M.; Shimizu, M.; Wojcik, S.E.; Aqeilan, R.I.; Zupo, S.; Dono, M.; et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc. Natl. Acad. Sci. USA 2005, 102, 13944–13949. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. MicroRNA-cancer connection: The beginning of a new tale. Cancer Res. 2006, 66, 7390–7394. [Google Scholar] [CrossRef]
- Zhu, S.; Cheng, X.; Wang, R.; Tan, Y.; Ge, M.; Li, D.; Xu, Q.; Sun, Y.; Zhao, C.; Chen, S.; et al. Restoration of microRNA function impairs MYC-dependent maintenance of MLL leukemia. Leukemia 2020, 34, 2484–2488. [Google Scholar] [CrossRef]
- Bao, F.; Zhang, L.; Pei, X.; Lian, C.; Liu, Y.; Tan, H.; Lei, P. MiR-20a-5p functions as a potent tumor suppressor by targeting PPP6C in acute myeloid leukemia. PLoS ONE 2021, 16, e0256995. [Google Scholar] [CrossRef]
- Garbin, A.; Lovisa, F.; Holmes, A.B.; Damanti, C.C.; Gallingani, I.; Carraro, E.; Accordi, B.; Veltri, G.; Pizzi, M.; d’Amore, E.S.G.; et al. miR-939 acts as tumor suppressor by modulating JUNB transcriptional activity in pediatric anaplastic large cell lymphoma. Haematologica 2021, 106, 610–613. [Google Scholar] [CrossRef]
- Liang, C.; Li, Y.; Wang, L.N.; Zhang, L.; Luo, J.S.; Peng, C.J.; Tang, W.Y.; Huang, L.B.; Tang, Y.L.; Luo, X.Q. Up-regulated miR-155 is associated with poor prognosis in childhood acute lymphoblastic leukemia and promotes cell proliferation targeting ZNF238. Hematology 2021, 26, 16–25. [Google Scholar] [CrossRef]
- Wang, X.; Zuo, D.; Yuan, Y.; Yang, X.; Hong, Z.; Zhang, R. MicroRNA-183 promotes cell proliferation via regulating programmed cell death 6 in pediatric acute myeloid leukemia. J. Cancer Res. Clin. Oncol. 2017, 143, 169–180. [Google Scholar] [CrossRef]
- Shaham, L.; Vendramini, E.; Ge, Y.; Goren, Y.; Birger, Y.; Tijssen, M.R.; McNulty, M.; Geron, I.; Schwartzman, O.; Goldberg, L.; et al. MicroRNA-486-5p is an erythroid oncomiR of the myeloid leukemias of Down syndrome. Blood 2015, 125, 1292–1301. [Google Scholar] [CrossRef]
- Tapeh, B.E.G.; Alivand, M.R.; Solali, S. The role of microRNAs in acute lymphoblastic leukaemia: From biology to applications. Cell Biochem. Funct. 2020, 38, 334–346. [Google Scholar] [CrossRef]
- Saito, Y.; Liang, G.; Egger, G.; Friedman, J.M.; Chuang, J.C.; Coetzee, G.A.; Jones, P.A. Specific activation of microRNA-127 with downregulation of the proto-Oncogene BCL6 by chromatin-Modifying drugs in human cancer cells. Cancer Cell 2006, 9, 435–443. [Google Scholar] [CrossRef]
- Kong, W.; He, L.; Richards, E.J.; Challa, S.; Xu, C.X.; Permuth-Wey, J.; Lancaster, J.M.; Coppola, D.; Sellers, T.A.; Djeu, J.Y.; et al. Upregulation of miRNA-155 promotes tumour angiogenesis by targeting VHL and is associated with poor prognosis and triple-negative breast cancer. Oncogene 2014, 33, 679–689. [Google Scholar] [CrossRef]
- You, X.; Sun, W.; Wang, Y.; Liu, X.; Wang, A.; Liu, L.; Han, S.; Sun, Y.; Zhang, J.; Guo, L.; et al. Cervical cancer-derived exosomal miR-663b promotes angiogenesis by inhibiting vinculin expression in vascular endothelial cells. Cancer Cell Int. 2021, 21, 684. [Google Scholar] [CrossRef]
- Wang, H.; Wang, L.; Zhou, X.; Luo, X.; Liu, K.; Jiang, E.; Chen, Y.; Shao, Z.; Shang, Z. OSCC Exosomes Regulate miR-210-3p Targeting EFNA3 to Promote Oral Cancer Angiogenesis through the PI3K/AKT Pathway. BioMed Res. Int. 2020, 2020, 2125656. [Google Scholar] [CrossRef]
- Lu, H.Q.; Wang, R.K.; Wang, H.R.; Zhou, G.Q.; Zhang, Y.S. miR-137 Inhibition of the Invasion, Metastasis, and Epithelial-Mesenchymal Transition of Nasopharyngeal Cancer by Regulating KDM1A. J. Oncol. 2021, 2021, 6060762. [Google Scholar] [CrossRef]
- Yuan, H.; Liu, F.; Ma, T.; Zeng, Z.; Zhang, N. miR-338-3p inhibits cell growth, invasion, and EMT process in neuroblastoma through targeting MMP-2. Open Life Sci. 2021, 16, 198–209. [Google Scholar] [CrossRef]
- Tu, Z.; Xiong, J.; Xiao, R.; Shao, L.; Yang, X.; Zhou, L.; Yuan, W.; Wang, M.; Yin, Q.; Wu, Y.; et al. Loss of miR-146b-5p promotes T cell acute lymphoblastic leukemia migration and invasion via the IL-17A pathway. J. Cell. Biochem. 2019, 120, 5936–5948. [Google Scholar] [CrossRef]
- Deng, L.; Petrek, H.; Tu, M.J.; Batra, N.; Yu, A.X.; Yu, A.M. Bioengineered miR-124-3p prodrug selectively alters the proteome of human carcinoma cells to control multiple cellular components and lung metastasis in vivo. Acta Pharm. Sin. B 2021, 11, 3950–3965. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; Brane, A.C.; Tollefsbol, T.O. MicroRNAs and Epigenetics Strategies to Reverse Breast Cancer. Cells 2019, 8, 1214. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Yang, X.; Liu, M.; Zhang, Z.; Xing, E. Roles of miRNA dysregulation in the pathogenesis of multiple myeloma. Cancer Gene Ther. 2021, 28, 1256–1268. [Google Scholar] [CrossRef] [PubMed]
- Muvarak, N.; Kelley, S.; Robert, C.; Baer, M.R.; Perrotti, D.; Gambacorti-Passerini, C.; Civin, C.; Scheibner, K.; Rassool, F.V. c-MYC Generates Repair Errors via Increased Transcription of Alternative-NHEJ Factors, LIG3 and PARP1, in Tyrosine Kinase-Activated Leukemias. Mol. Cancer Res. 2015, 13, 699–712. [Google Scholar] [CrossRef]
- Wu, J.; Ferragut Cardoso, A.P.; States, V.A.R.; Al-Eryani, L.; Doll, M.; Wise, S.S.; Rai, S.N.; States, C. Overexpression of hsa-miR-186 induces chromosomal instability in arsenic-exposed human keratinocytes. Toxicol. Appl. Pharmacol. 2019, 378, 114614. [Google Scholar] [CrossRef]
- Cosentino, G.; Plantamura, I.; Tagliabue, E.; Iorio, M.V.; Cataldo, A. Breast Cancer Drug Resistance: Overcoming the Challenge by Capitalizing on MicroRNA and Tumor Microenvironment Interplay. Cancers 2021, 13, 3691. [Google Scholar] [CrossRef]
- Mitra, A.K.; Zillhardt, M.; Hua, Y.; Tiwari, P.; Murmann, A.E.; Peter, M.E.; Lengyel, E. MicroRNAs reprogram normal fibroblasts into cancer-associated fibroblasts in ovarian cancer. Cancer Discov. 2012, 2, 1100–1108. [Google Scholar] [CrossRef]
- Frassanito, M.A.; Desantis, V.; Di Marzo, L.; Craparotta, I.; Beltrame, L.; Marchini, S.; Annese, T.; Visino, F.; Arciuli, M.; Saltarella, I.; et al. Bone marrow fibroblasts overexpress miR-27b and miR-214 in step with multiple myeloma progression, dependent on tumour cell-derived exosomes. J. Pathol. 2019, 247, 241–253. [Google Scholar] [CrossRef]
- Xi, Q.; Zhang, J.; Yang, G.; Zhang, L.; Chen, Y.; Wang, C.; Zhang, Z.; Guo, X.; Zhao, J.; Xue, Z.; et al. Restoration of miR-340 controls pancreatic cancer cell CD47 expression to promote macrophage phagocytosis and enhance antitumor immunity. J. Immunother. Cancer 2020, 8, e000253. [Google Scholar] [CrossRef]
- Del Gaizo, M.; Sergio, I.; Lazzari, S.; Cialfi, S.; Pelullo, M.; Screpanti, I.; Felli, M.P. MicroRNAs as Modulators of the Immune Response in T-Cell Acute Lymphoblastic Leukemia. Int. J. Mol. Sci. 2022, 23, 829. [Google Scholar] [CrossRef]
- Rashkovan, M.; Ferrando, A. Metabolic dependencies and vulnerabilities in leukemia. Genes Dev. 2019, 33, 1460–1474. [Google Scholar] [CrossRef]
- Bao, L.; Xu, T.; Lu, X.; Huang, P.; Pan, Z.; Ge, M. Metabolic Reprogramming of Thyroid Cancer Cells and Crosstalk in Their Microenvironment. Front. Oncol. 2021, 11, 773028. [Google Scholar] [CrossRef]
- Lo Presti, C.; Fauvelle, F.; Jacob, M.C.; Mondet, J.; Mossuz, P. The metabolic reprogramming in acute myeloid leukemia patients depends on their genotype and is a prognostic marker. Blood Adv. 2021, 5, 156–166. [Google Scholar] [CrossRef]
- Wojcicki, A.V.; Kasowski, M.M.; Sakamoto, K.M.; Lacayo, N. Metabolomics in acute myeloid leukemia. Mol. Genet. Metab. 2020, 130, 230–238. [Google Scholar] [CrossRef]
- Stockard, B.; Garrett, T.; Guingab-Cagmat, J.; Meshinchi, S.; Lamba, J. Distinct Metabolic features differentiating FLT3-ITD AML from FLT3-WT childhood Acute Myeloid Leukemia. Sci. Rep. 2018, 8, 5534. [Google Scholar] [CrossRef]
- Chen, W.L.; Wang, J.H.; Zhao, A.H.; Xu, X.; Wang, Y.H.; Chen, T.L.; Li, J.M.; Mi, J.Q.; Zhu, Y.M.; Liu, Y.F.; et al. A distinct glucose metabolism signature of acute myeloid leukemia with prognostic value. Blood 2014, 124, 1645–1654. [Google Scholar] [CrossRef]
- Gao, P.; Sun, L.; He, X.; Cao, Y.; Zhang, H. MicroRNAs and the Warburg Effect: New players in an old arena. Curr. Gene Ther. 2012, 12, 285–291. [Google Scholar] [CrossRef]
- Ozfiliz-Kilbas, P.; Sonmez, O.; Obakan-Yerlikaya, P.; Coker-Gurkan, A.; Palavan-Ünsal, N.; Uysal-Onganer, P.; Arisan, E.D. In Vitro Investigations of miR-33a Expression in Estrogen Receptor-Targeting Therapies in Breast Cancer Cells. Cancers 2021, 13, 5322. [Google Scholar] [CrossRef]
- Liu, S.; Wang, H.; Guo, W.; Zhou, X.; Shu, Y.; Liu, H.; Yang, L.; Tang, S.; Su, H.; Liu, Z.; et al. MiR-652-5p elevated glycolysis level by targeting TIGAR in T-cell acute lymphoblastic leukemia. Cell Death Dis. 2022, 13, 148. [Google Scholar] [CrossRef]
- Rencelj, A.; Gvozdenovic, N.; Cemazar, M. MitomiRs: Their roles in mitochondria and importance in cancer cell metabolism. Radiol. Oncol. 2021, 55, 379–392. [Google Scholar] [CrossRef]
- Prada-Arismendy, J.; Arroyave, J.C.; Röthlisberger, S. Molecular biomarkers in acute myeloid leukemia. Blood Rev. 2017, 31, 63–76. [Google Scholar] [CrossRef] [PubMed]
- Mortazavi, D.; Sohrabi, B.; Mosallaei, M.; Nariman-Saleh-Fam, Z.; Bastami, M.; Mansoori, Y.; Daraei, A.; Vahed, S.Z.; Navid, S.; Saadatian, Z.; et al. Epi-miRNAs: Regulators of the Histone Modification Machinery in Human Cancer. J. Oncol. 2022, 2022, 4889807. [Google Scholar] [CrossRef] [PubMed]
- Karimzadeh, M.R.; Pourdavoud, P.; Ehtesham, N.; Qadbeigi, M.; Asl, M.M.; Alani, B.; Mosallaei, M.; Pakzad, B. Regulation of DNA methylation machinery by epi-miRNAs in human cancer: Emerging new targets in cancer therapy. Cancer Gene Ther. 2021, 28, 157–174. [Google Scholar] [CrossRef] [PubMed]
- Garzon, R.; Liu, S.; Fabbri, M.; Liu, Z.; Heaphy, C.E.; Callegari, E.; Schwind, S.; Pang, J.; Yu, J.; Muthusamy, N.; et al. MicroRNA-29b induces global DNA hypomethylation and tumor suppressor gene reexpression in acute myeloid leukemia by targeting directly DNMT3A and 3B and indirectly DNMT1. Blood 2009, 113, 6411–6418. [Google Scholar] [CrossRef]
- Aure, M.R.; Fleischer, T.; Bjørklund, S.; Ankill, J.; Castro-Mondragon, J.A.; Børresen-Dale, A.L.; Tost, J.; Sahlberg, K.K.; Mathelier, A.; Tekpli, X.; et al. Crosstalk between microRNA expression and DNA methylation drives the hormone-dependent phenotype of breast cancer. Genome Med. 2021, 13, 72. [Google Scholar] [CrossRef]
- Stumpel, D.; Schotte, D.; Lange-Turenhout, E.; Schneider, P.; Seslija, L.; de Menezes, R.X.; Marquez, V.E.; Pieters, R.; den Boer, M.L.; Stam, R.W. Hypermethylation of specific microRNA genes in MLL-rearranged infant acute lymphoblastic leukemia: Major matters at a micro scale. Leukemia 2011, 25, 429–439. [Google Scholar] [CrossRef]
- Humphries, B.; Wang, Z.; Yang, C. MicroRNA Regulation of Epigenetic Modifiers in Breast Cancer. Cancers 2019, 11, 897. [Google Scholar] [CrossRef]
- Xia, P.; Gu, R.; Zhang, W.; Shao, L.; Li, F.; Wu, C.; Sun, Y. MicroRNA-377 exerts a potent suppressive role in osteosarcoma through the involvement of the histone acetyltransferase 1-mediated Wnt axis. J. Cell. Physiol. 2019, 234, 22787–22798. [Google Scholar] [CrossRef]
- Roccaro, A.M.; Sacco, A.; Jia, X.; Azab, A.K.; Maiso, P.; Ngo, H.T.; Azab, F.; Runnels, J.; Quang, P.; Ghobrial, I.M. microRNA dependent modulation of histone acetylation in Waldenstrom macroglobulinemia. Blood 2010, 116, 1506–1514. [Google Scholar] [CrossRef]
- Zhang, Q.; Deng, X.; Tang, X.; You, Y.; Mei, M.; Liu, D.; Gui, L.; Cai, Y.; Xin, X.; He, X.; et al. MicroRNA-20a Suppresses Tumor Proliferation and Metastasis in Hepatocellular Carcinoma by Directly Targeting EZH1. Front. Oncol. 2021, 11, 737986. [Google Scholar] [CrossRef]
- Chen, C.Z.; Li, L.; Lodish, H.F.; Bartel, D.P. MicroRNAs modulate hematopoietic lineage differentiation. Science 2004, 303, 83–86. [Google Scholar] [CrossRef]
- Kim, M.; Civin, C.I.; Kingsbury, T.J. MicroRNAs as regulators and effectors of hematopoietic transcription factors. Wiley Interdiscip. Rev. RNA 2019, 10, e1537. [Google Scholar] [CrossRef]
- Surdziel, E.; Cabanski, M.; Dallmann, I.; Lyszkiewicz, M.; Krueger, A.; Ganser, A.; Scherr, M.; Eder, M. Enforced expression of miR-125b affects myelopoiesis by targeting multiple signaling pathways. Blood 2011, 117, 4338–4348. [Google Scholar] [CrossRef]
- Raghuwanshi, S.; Dahariya, S.; Musvi, S.S.; Gutti, U.; Kandi, R.; Undi, R.B.; Sahu, I.; Gautam, D.K.; Paddibhatla, I.; Gutti, R.K. MicroRNA function in megakaryocytes. Platelets 2019, 30, 809–816. [Google Scholar] [CrossRef]
- Fazi, F.; Rosa, A.; Fatica, A.; Gelmetti, V.; De Marchis, M.L.; Nervi, C.; Bozzoni, I. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPalpha regulates human granulopoiesis. Cell 2005, 123, 819–831. [Google Scholar] [CrossRef]
- Rao, D.S.; O’Connell, R.M.; Chaudhuri, A.A.; Garcia-Flores, Y.; Geiger, T.L.; Baltimore, D. MicroRNA-34a perturbs B lymphocyte development by repressing the forkhead box transcription factor Foxp1. Immunity 2010, 33, 48–59. [Google Scholar] [CrossRef]
- Koenecke, C.; Krueger, A. MicroRNA in T-Cell Development and T-Cell Mediated Acute Graft-Versus-Host Disease. Front. Immunol. 2018, 9, 992. [Google Scholar] [CrossRef]
- Ghisi, M.; Corradin, A.; Basso, K.; Frasson, C.; Serafin, V.; Mukherjee, S.; Mussolin, L.; Ruggero, K.; Bonanno, L.; Guffanti, A.; et al. Modulation of microRNA expression in human T-cell development: Targeting of NOTCH3 by miR-150. Blood 2011, 117, 7053–7062. [Google Scholar] [CrossRef]
- Saultz, J.N.; Freud, A.G.; Mundy-Bosse, B.L. MicroRNA regulation of natural killer cell development and function in leukemia. Mol. Immunol. 2019, 115, 12–20. [Google Scholar] [CrossRef]
- Mi, S.; Lu, J.; Sun, M.; Li, Z.; Zhang, H.; Neilly, M.B.; Wang, Y.; Qian, Z.; Jin, J.; Zhang, Y.; et al. MicroRNA expression signatures accurately discriminate acute lymphoblastic leukemia from acute myeloid leukemia. Proc. Natl. Acad. Sci. USA 2007, 104, 19971–19976. [Google Scholar] [CrossRef]
- Ghodousi, E.S.; Rahgozar, S. MicroRNA-326 and microRNA-200c: Two novel biomarkers for diagnosis and prognosis of pediatric acute lymphoblastic leukemia. J. Cell. Biochem. 2018, 119, 6024–6032. [Google Scholar] [CrossRef] [PubMed]
- Almeida, R.S.; Costa e Silva, M.; Coutinho, L.L.; Garcia Gomes, R.; Pedrosa, F.; Massaro, J.D.; Donadi, E.A.; Lucena-Silva, N. MicroRNA expression profiles discriminate childhood T- from B-acute lymphoblastic leukemia. Hematol. Oncol. 2019, 37, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, M.; Oikawa, K.; Takanashi, M.; Kudo, M.; Ohyashiki, J.; Ohyashiki, K.; Kuroda, M. Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PLoS ONE 2009, 4, e5532. [Google Scholar] [CrossRef] [PubMed]
- Yan, W.; Xu, L.; Sun, Z.; Lin, Y.; Zhang, W.; Chen, J.; Hu, S.; Shen, B. MicroRNA biomarker identification for pediatric acute myeloid leukemia based on a novel bioinformatics model. Oncotarget 2015, 6, 26424–26436. [Google Scholar] [CrossRef]
- Lin, X.; Wang, Z.; Wang, Y.; Feng, W. Serum MicroRNA-370 as a potential diagnostic and prognostic biomarker for pediatric acute myeloid leukemia. Int. J. Clin. Exp. Pathol. 2015, 8, 14658–14666. [Google Scholar]
- Rahimi, Z.; Ghorbani, Z.; Motamed, H.; Jalilian, N. Aberrant expression profile of miR-32, miR-98 and miR-374 in chronic lymphocytic leukemia. Leuk. Res. 2021, 111, 106691. [Google Scholar] [CrossRef]
- Bagheri, M.; Khansarinejad, B.; Mosayebi, G.; Moradabadi, A.; Mondanizadeh, M. Diagnostic Value of Plasma miR-145 and miR-185 as Targeting of the APRIL Oncogene in the B-cell Chronic Lymphocytic Leukemia. Asian Pac. J. Cancer Prev. 2021, 22, 111–117. [Google Scholar] [CrossRef]
- Keramati, F.; Jafarian, A.; Soltani, A.; Javandoost, E.; Mollaei, M.; Fallah, P. Circulating miRNAs can serve as potential diagnostic biomarkers in chronic myelogenous leukemia patients. Leuk. Res. Rep. 2021, 16, 100257. [Google Scholar] [CrossRef]
- Poláková, K.M.; Lopotová, T.; Klamová, H.; Burda, P.; Trněný, M.; Stopka, T.; Moravcová, J. Expression patterns of microRNAs associated with CML phases and their disease related targets. Mol. Cancer 2011, 10, 41. [Google Scholar] [CrossRef]
- Paczkowska, J.; Giefing, M. MicroRNA signature in classical Hodgkin lymphoma. J. Appl. Genet. 2021, 62, 281–288. [Google Scholar] [CrossRef]
- Culpin, R.E.; Proctor, S.J.; Angus, B.; Crosier, S.; Anderson, J.J.; Mainou-Fowler, T. A 9 series microRNA signature differentiates between germinal centre and activated B-cell-like diffuse large B-cell lymphoma cell lines. Int. J. Oncol. 2010, 37, 367–376. [Google Scholar] [CrossRef]
- Denaro, M.; Navari, E.; Ugolini, C.; Seccia, V.; Donati, V.; Casani, A.P.; Basolo, F. A microRNA signature for the differential diagnosis of salivary gland tumors. PLoS ONE 2019, 14, e0210968. [Google Scholar] [CrossRef]
- Fernández, C.; Bellosillo, B.; Ferraro, M.; Seoane, A.; Sánchez-González, B.; Pairet, S.; Pons, A.; Barranco, L.; Vela, M.C.; Gimeno, E.; et al. MicroRNAs 142-3p, miR-155 and miR-203 Are Deregulated in Gastric MALT Lymphomas Compared to Chronic Gastritis. Cancer Genom. Proteom. 2017, 14, 75–82. [Google Scholar] [CrossRef][Green Version]
- Baraniskin, A.; Chomiak, M.; Ahle, G.; Gress, T.; Buchholz, M.; Turewicz, M.; Eisenacher, M.; Margold, M.; Schlegel, U.; Schmiegel, W.; et al. MicroRNA-30c as a novel diagnostic biomarker for primary and secondary B-cell lymphoma of the CNS. J. Neurooncol. 2018, 137, 463–468. [Google Scholar] [CrossRef]
- Xiang, Y.; Zhang, L.; Xiang, P.; Zhang, J. Circulating miRNAs as Auxiliary Diagnostic Biomarkers for Multiple Myeloma: A Systematic Review, Meta-Analysis, and Recommendations. Front. Oncol. 2021, 11, 698197. [Google Scholar] [CrossRef]
- Kubiczkova, L.; Kryukov, F.; Slaby, O.; Dementyeva, E.; Jarkovsky, J.; Nekvindova, J.; Radova, L.; Greslikova, H.; Kuglik, P.; Vetesnikova, E.; et al. Circulating serum microRNAs as novel diagnostic and prognostic biomarkers for multiple myeloma and monoclonal gammopathy of undetermined significance. Haematologica 2014, 99, 511–518. [Google Scholar] [CrossRef]
- Zhang, L.C.; Zhong, C.; Ma, Y.J.; Liu, M.; Nong, W.X. Expression of MiR-429 in Patients with Acute Lymphoblastic Leukemia and Its Prognostic Value. Zhongguo Shi Yan Xue Ye Xue Za Zhi 2020, 28, 119–124. [Google Scholar] [CrossRef]
- Gębarowska, K.; Mroczek, A.; Kowalczyk, J.R.; Lejman, M. MicroRNA as a Prognostic and Diagnostic Marker in T-Cell Acute Lymphoblastic Leukemia. Int. J. Mol. Sci. 2021, 22, 5317. [Google Scholar] [CrossRef]
- Agirre, X.; Vilas-Zornoza, A.; Jiménez-Velasco, A.; Martin-Subero, J.I.; Cordeu, L.; Gárate, L.; San José-Eneriz, E.; Abizanda, G.; Rogríguez-Otero, P.; Fortes, P.; et al. Epigenetic silencing of the tumor suppressor microRNA Hsa-miR-124a regulates CDK6 expression and confers a poor prognosis in acute lymphoblastic leukemia. Cancer Res. 2009, 69, 4443–4453. [Google Scholar] [CrossRef]
- Gutierrez-Camino, A.; Garcia-Obregon, S.; Lopez-Lopez, E.; Astigarraga, I.; Garcia-Orad, A. miRNA deregulation in childhood acute lymphoblastic leukemia: A systematic review. Epigenomics 2020, 12, 69–80. [Google Scholar] [CrossRef]
- El-Khazragy, N.; Noshi, M.A.; Abdel-Malak, C.; Zahran, R.F.; Swellam, M. miRNA-155 and miRNA-181a as prognostic biomarkers for pediatric acute lymphoblastic leukemia. J. Cell. Biochem. 2019, 120, 6315–6321. [Google Scholar] [CrossRef] [PubMed]
- Avigad, S.; Verly, I.R.; Lebel, A.; Kordi, O.; Shichrur, K.; Ohali, A.; Hameiri-Grossman, M.; Kaspers, G.J.L.; Cloos, J.; Fronkova, E.; et al. miR expression profiling at diagnosis predicts relapse in pediatric precursor B-cell acute lymphoblastic leukemia. Genes Chromosomes Cancer 2016, 55, 328–339. [Google Scholar] [CrossRef] [PubMed]
- Piatopoulou, D.; Avgeris, M.; Drakaki, I.; Marmarinos, A.; Xagorari, M.; Baka, M.; Pourtsidis, A.; Kossiva, L.; Gourgiotis, D.; Scorilas, A. Clinical utility of miR-143/miR-182 levels in prognosis and risk stratification specificity of BFM-treated childhood acute lymphoblastic leukemia. Ann. Hematol. 2018, 97, 1169–1182. [Google Scholar] [CrossRef] [PubMed]
- Amankwah, E.K.; Devidas, M.; Teachey, D.T.; Rabin, K.R.; Brown, P.A. Six Candidate miRNAs Associated with Early Relapse in Pediatric B-Cell Acute Lymphoblastic Leukemia. Anticancer Res. 2020, 40, 3147–3153. [Google Scholar] [CrossRef]
- Weng, H.; Lal, K.; Yang, F.F.; Chen, J. The pathological role and prognostic impact of miR-181 in acute myeloid leukemia. Cancer Genet. 2015, 208, 225–229. [Google Scholar] [CrossRef]
- Cui, L.; Zeng, T.; Zhang, L.; Liu, Y.; Wu, G.; Fu, L. High expression of miR-195 is related to favorable prognosis in cytogenetically normal acute myeloid leukemia. Int. J. Clin. Oncol. 2021, 26, 1986–1993. [Google Scholar] [CrossRef]
- Li, S.M.; Zhao, Y.Q.; Hao, Y.L.; Liang, Y.Y. Upregulation of miR-504-3p is associated with favorable prognosis of acute myeloid leukemia and may serve as a tumor suppressor by targeting MTHFD2. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 1203–1213. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, N.; Wang, R.; Shao, T.; Feng, Y.; Yao, Y.; Wu, Q.; Zhu, S.; Cao, J.; Zhang, H.; et al. High expression of miR-363 predicts poor prognosis and guides treatment selection in acute myeloid leukemia. J. Transl. Med. 2019, 17, 106. [Google Scholar] [CrossRef]
- Bhayadia, R.; Krowiorz, K.; Haetscher, N.; Jammal, R.; Emmrich, S.; Obulkasim, A.; Fiedler, J.; Schwarzer, A.; Rouhi, A.; Heuser, M.; et al. Endogenous Tumor Suppressor microRNA-193b: Therapeutic and Prognostic Value in Acute Myeloid Leukemia. J. Clin. Oncol. 2018, 36, 1007–1016. [Google Scholar] [CrossRef]
- Zhu, R.; Lin, W.; Zhao, W.; Fan, F.; Tang, L.; Hu, Y. A 4-microRNA signature for survival prognosis in pediatric and adolescent acute myeloid leukemia. J. Cell. Biochem. 2019, 120, 3958–3968. [Google Scholar] [CrossRef]
- Zhu, C.; Wang, Y.; Kuai, W.; Sun, X.; Chen, H.; Hong, Z. Prognostic value of miR-29a expression in pediatric acute myeloid leukemia. Clin. Biochem. 2013, 46, 49–53. [Google Scholar] [CrossRef]
- Qi, X.; Zhang, Y. MicroRNA-199a deficiency relates to higher bone marrow blasts, poor risk stratification and worse prognostication in pediatric acute myeloid leukemia patients. Pediatr. Hematol. Oncol. 2022, 19, 1–8. [Google Scholar] [CrossRef]
- Marcucci, G.; Mrózek, K.; Radmacher, M.D.; Bloomfield, C.D.; Croce, C.M. MicroRNA expression profiling in acute myeloid and chronic lymphocytic leukaemias. Best Pract. Res. Clin. Haematol. 2009, 22, 239–248. [Google Scholar] [CrossRef]
- Khalifa, M.M.; Zaki, N.E.; Nazier, A.A.; Moussa, M.A.; Haleem, R.A.; Rabie, M.A.; Mansour, A.R. Prognostic significance of microRNA 17–92 cluster expression in Egyptian chronic lymphocytic leukemia patients. J. Egypt. Natl. Cancer Inst. 2021, 33, 37. [Google Scholar] [CrossRef]
- Baer, C.; Oakes, C.C.; Ruppert, A.S.; Claus, R.; Kim-Wanner, S.Z.; Mertens, D.; Zenz, T.; Stilgenbauer, S.; Byrd, J.C.; Plass, C. Epigenetic silencing of miR-708 enhances NF-κB signaling in chronic lymphocytic leukemia. Int. J. Cancer 2015, 137, 1352–1361. [Google Scholar] [CrossRef]
- Litwińska, Z.; Machaliński, B. miRNAs in chronic myeloid leukemia: Small molecules, essential function. Leuk. Lymphoma 2017, 58, 1297–1305. [Google Scholar] [CrossRef]
- Habib, E.M.; Nosiar, N.A.; Eid, M.A.; Taha, A.M.; Sherief, D.E.; Hassan, A.E.; Abdel Ghafar, M.T. MiR-150 Expression in Chronic Myeloid Leukemia: Relation to Imatinib Response. Lab Med. 2022, 53, 58–64. [Google Scholar] [CrossRef]
- Ninawe, A.; Guru, S.A.; Yadav, P.; Masroor, M.; Samadhiya, A.; Bhutani, N.; Gupta, N.; Gupta, R.; Saxena, A. miR-486-5p: A Prognostic Biomarker for Chronic Myeloid Leukemia. ACS Omega 2021, 6, 7711–7718. [Google Scholar] [CrossRef]
- Jia, Y.J.; Liu, Z.B.; Wang, W.G.; Sun, C.B.; Wei, P.; Yang, Y.L.; You, M.J.; Yu, B.H.; Li, X.Q.; Zhou, X.Y. HDAC6 regulates microRNA-27b that suppresses proliferation, promotes apoptosis and target MET in diffuse large B-cell lymphoma. Leukemia 2018, 32, 703–711. [Google Scholar] [CrossRef]
- Hedström, G.; Thunberg, U.; Berglund, M.; Simonsson, M.; Amini, R.M.; Enblad, G. Low expression of microRNA-129-5p predicts poor clinical outcome in diffuse large B cell lymphoma (DLBCL). Int. J. Hematol. 2013, 97, 465–471. [Google Scholar] [CrossRef]
- Li, L.; Qiu, L.; Zhang, H.; Qian, Z.; Zhao, H. Overexpression of microRNA-130a predicts adverse prognosis of primary gastrointestinal diffuse large B-cell lymphoma. Oncol. Lett. 2020, 20, 93. [Google Scholar] [CrossRef]
- Marchesi, F.; Regazzo, G.; Palombi, F.; Terrenato, I.; Sacconi, A.; Spagnuolo, M.; Donzelli, S.; Marino, M.; Ercolani, C.; Di Benedetto, A.; et al. Serum miR-22 as potential non-invasive predictor of poor clinical outcome in newly diagnosed, uniformly treated patients with diffuse large B-cell lymphoma: An explorative pilot study. J. Exp. Clin. Cancer Res. 2018, 37, 95. [Google Scholar] [CrossRef] [PubMed]
- Troppan, K.; Wenzl, K.; Pichler, M.; Pursche, B.; Schwarzenbacher, D.; Feichtinger, J.; Thallinger, G.G.; Beham-Schmid, C.; Neumeister, P.; Deutsch, A. miR-199a and miR-497 Are Associated with Better Overall Survival due to Increased Chemosensitivity in Diffuse Large B-Cell Lymphoma Patients. Int. J. Mol. Sci. 2015, 16, 18077–18095. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Gao, L.; Zhang, S.; Tao, L.; Wang, J.; Yang, J.; Zhu, M. Prognostic significance of miR-34a and its target proteins of FOXP1, p53, and BCL2 in gastric MALT lymphoma and DLBCL. Gastric Cancer 2014, 17, 431–441. [Google Scholar] [CrossRef]
- Morales-Martinez, M.; Vega, G.G.; Neri, N.; Nambo, M.J.; Alvarado, I.; Cuadra, I.; Duran-Padilla, M.A.; Huerta-Yepez, S.; Vega, M.I. MicroRNA-7 Regulates Migration and Chemoresistance in Non-Hodgkin Lymphoma Cells Through Regulation of KLF4 and YY1. Front. Oncol. 2020, 10, 588893. [Google Scholar] [CrossRef]
- Zhou, K.; Feng, X.; Wang, Y.; Liu, Y.; Tian, L.; Zuo, W.; Yi, S.; Wei, X.; Song, Y.; Qiu, L. miR-223 is repressed and correlates with inferior clinical features in mantle cell lymphoma through targeting SOX11. Exp. Hematol. 2018, 58, 27–34.e1. [Google Scholar] [CrossRef]
- Papanota, A.M.; Karousi, P.; Kontos, C.K.; Artemaki, P.I.; Liacos, C.I.; Papadimitriou, M.A.; Bagratuni, T.; Eleutherakis-Papaiakovou, E.; Malandrakis, P.; Ntanasis-Stathopoulos, I.; et al. A Cancer-Related microRNA Signature Shows Biomarker Utility in Multiple Myeloma. Int. J. Mol. Sci. 2021, 22, 13144. [Google Scholar] [CrossRef]
- Desantis, V.; Solimando, A.G.; Saltarella, I.; Sacco, A.; Giustini, V.; Bento, M.; Lamanuzzi, A.; Melaccio, A.; Frassanito, M.A.; Paradiso, A.; et al. MicroRNAs as a Potential New Preventive Approach in the Transition from Asymptomatic to Symptomatic Multiple Myeloma Disease. Cancers 2021, 13, 3650. [Google Scholar] [CrossRef]
- Ren, Y.; Li, X.; Wang, W.; He, W.; Wang, J.; Wang, Y. Expression of Peripheral Blood miRNA-720 and miRNA-1246 Can Be Used as a Predictor for Outcome in Multiple Myeloma Patients. Clin. Lymphoma Myeloma Leuk. 2017, 17, 415–423. [Google Scholar] [CrossRef]
- Mondal, P.; Meeran, S.M. microRNAs in cancer chemoresistance: The sword and the shield. Noncoding RNA Res. 2021, 6, 200–210. [Google Scholar] [CrossRef]
- Sakurai, N.; Komada, Y.; Hanaki, R.; Morimoto, M.; Ito, T.; Nakato, D.; Hirayama, M. Role of microRNAs in glucocorticoid-resistant B-cell precursor acute lymphoblastic leukemia. Oncol. Rep. 2019, 42, 708–716. [Google Scholar] [CrossRef]
- Najjary, S.; Mohammadzadeh, R.; Mansoori, B.; Vahidian, F.; Mohammadi, A.; Doustvandi, M.A.; Khaze, V.; Hajiasgharzadeh, K.; Baradaran, B. Combination therapy with miR-34a and doxorubicin synergistically induced apoptosis in T-cell acute lymphoblastic leukemia cell line. Med. Oncol. 2021, 38, 142. [Google Scholar] [CrossRef]
- Sun, H.; Zhang, Z.; Luo, W.; Liu, J.; Lou, Y.; Xia, S. NET1 Enhances Proliferation and Chemoresistance in Acute Lymphoblastic Leukemia Cells. Oncol. Res. 2019, 27, 935–944. [Google Scholar] [CrossRef]
- Zhang, H.; Luo, X.Q.; Zhang, P.; Huang, L.B.; Zheng, Y.S.; Wu, J.; Zhou, H.; Qu, L.H.; Xu, L.; Chen, Y.Q. MicroRNA patterns associated with clinical prognostic parameters and CNS relapse prediction in pediatric acute leukemia. PLoS ONE 2009, 4, e7826. [Google Scholar] [CrossRef]
- Li, S.W.; Li, H.; Zhang, Z.P.; Zhuo, F.; Li, Z.X. Expression and Clinical Significance of MiR-146a and MiR-221 in Childhood Acute T Lymphoblastic Leukemia. Zhongguo Shi Yan Xue Ye Xue Za Zhi 2020, 28, 436–441. [Google Scholar] [CrossRef]
- Labib, H.A.; Elantouny, N.G.; Ibrahim, N.F.; Alnagar, A.A. Upregulation of microRNA-21 is a poor prognostic marker in patients with childhood B cell acute lymphoblastic leukemia. Hematology 2017, 22, 392–397. [Google Scholar] [CrossRef]
- Zamani, A.; Fattahi Dolatabadi, N.; Houshmand, M.; Nabavizadeh, N. miR-324-3p and miR-508-5p expression levels could serve as potential diagnostic and multidrug-resistant biomarkers in childhood acute lymphoblastic leukemia. Leuk. Res. 2021, 109, 106643. [Google Scholar] [CrossRef]
- Zhang, H.; Liu, L.; Chen, L.; Liu, H.; Ren, S.; Tao, Y. Long noncoding RNA DANCR confers cytarabine resistance in acute myeloid leukemia by activating autophagy via the miR-874-3P/ATG16L1 axis. Mol. Oncol. 2021, 15, 1203–1216. [Google Scholar] [CrossRef]
- Vandewalle, V.; Essaghir, A.; Bollaert, E.; Lenglez, S.; Graux, C.; Schoemans, H.; Saussoy, P.; Michaux, L.; Valk, P.J.M.; Demoulin, J.B.; et al. miR-15a-5p and miR-21-5p contribute to chemoresistance in cytogenetically normal acute myeloid leukaemia by targeting PDCD4, ARL2 and BTG2. J. Cell. Mol. Med. 2021, 25, 575–585. [Google Scholar] [CrossRef]
- Zhang, H.; Kang, J.; Liu, L.; Chen, L.; Ren, S.; Tao, Y. MicroRNA-143 sensitizes acute myeloid leukemia cells to cytarabine via targeting ATG7- and ATG2B-dependent autophagy. Aging 2020, 12, 20111–20126. [Google Scholar] [CrossRef]
- Bollaert, E.; Claus, M.; Vandewalle, V.; Lenglez, S.; Essaghir, A.; Demoulin, J.B.; Havelange, V. MiR-15a-5p Confers Chemoresistance in Acute Myeloid Leukemia by Inhibiting Autophagy Induced by Daunorubicin. Int. J. Mol. Sci. 2021, 22, 5153. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Lei, P.; Qiao, H.; Sun, K.; Lu, X.; Bao, F.; Yu, R.; Lian, C.; Li, Y.; Chen, W.; et al. miR-9 Enhances the Chemosensitivity of AML Cells to Daunorubicin by Targeting the EIF5A2/MCL-1 Axis. Int. J. Biol. Sci. 2019, 15, 579–586. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Deng, T.; Su, C.; Shang, Z. MicroRNA 217 inhibits cell proliferation and enhances chemosensitivity to doxorubicin in acute myeloid leukemia by targeting KRAS. Oncol. Lett. 2017, 13, 4986–4994. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Fang, Z.; Lu, R.; Zhao, H.; Gong, T.; Liu, D.; Hong, L.; Ma, J.; Zhang, M. MicroRNA-204 Potentiates the Sensitivity of Acute Myeloid Leukemia Cells to Arsenic Trioxide. Oncol. Res. 2019, 27, 1035–1042. [Google Scholar] [CrossRef]
- Said, F.; Tantawy, M.; Sayed, A.; Ahmed, S. Clinical Significance of MicroRNA-29a and MicroRNA-100 Gene Expression in Pediatric Acute Myeloid Leukemia. J. Pediatr. Hematol. Oncol. 2022, 44, e391–e395. [Google Scholar] [CrossRef]
- Zhang, H.; Luo, X.Q.; Feng, D.D.; Zhang, X.J.; Wu, J.; Zheng, Y.S.; Chen, X.; Xu, L.; Chen, Y.Q. Upregulation of microRNA-125b contributes to leukemogenesis and increases drug resistance in pediatric acute promyelocytic leukemia. Mol. Cancer 2011, 10, 108. [Google Scholar] [CrossRef]
- Fonte, E.; Apollonio, B.; Scarfò, L.; Ranghetti, P.; Fazi, C.; Ghia, P.; Caligaris-Cappio, F.; Muzio, M. In vitro sensitivity of CLL cells to fludarabine may be modulated by the stimulation of Toll-like receptors. Clin. Cancer Res. 2013, 19, 367–379. [Google Scholar] [CrossRef]
- Zhu, D.X.; Zhu, W.; Fang, C.; Fan, L.; Zou, Z.J.; Wang, Y.H.; Liu, P.; Hong, M.; Miao, K.R.; Liu, P.; et al. miR-181a/b significantly enhances drug sensitivity in chronic lymphocytic leukemia cells via targeting multiple anti-apoptosis genes. Carcinogenesis 2012, 33, 1294–1301. [Google Scholar] [CrossRef]
- Li, Y.; Zhao, L.; Li, N.; Miao, Y.; Zhou, H.; Jia, L. miR-9 regulates the multidrug resistance of chronic myelogenous leukemia by targeting ABCB1. Oncol. Rep. 2017, 37, 2193–2200. [Google Scholar] [CrossRef]
- Jiang, X.; Cheng, Y.; Hu, C.; Zhang, A.; Ren, Y.; Xu, X. MicroRNA-221 sensitizes chronic myeloid leukemia cells to imatinib by targeting STAT5. Leuk. Lymphoma 2019, 60, 1709–1720. [Google Scholar] [CrossRef]
- Klümper, T.; Bruckmueller, H.; Diewock, T.; Kaehler, M.; Haenisch, S.; Pott, C.; Bruhn, O.; Cascorbi, I. Expression differences of miR-142-5p between treatment-naïve chronic myeloid leukemia patients responding and non-responding to imatinib therapy suggest a link to oncogenic ABL2, SRI, cKIT and MCL1 signaling pathways critical for development of therapy resistance. Exp. Hematol. Oncol. 2020, 9, 26. [Google Scholar] [CrossRef]
- Alves, R.; Gonçalves, A.C.; Jorge, J.; Marques, G.; Luís, D.; Ribeiro, A.B.; Freitas-Tavares, P.; Oliveiros, B.; Almeida, A.M.; Sarmento-Ribeiro, A.B. MicroRNA signature refine response prediction in CML. Sci. Rep. 2019, 9, 9666. [Google Scholar] [CrossRef]
- Habib, E.M.; Nosiar, N.A.; Eid, M.A.; Taha, A.M.; Sherief, D.E.; Hassan, A.E.; Ghafar, M.T.A. Circulating miR-146a expression predicts early treatment response to imatinib in adult chronic myeloid leukemia. J. Investig. Med. 2021, 69, 333–337. [Google Scholar] [CrossRef]
- Zhang, X.T.; Dong, S.H.; Zhang, J.Y.; Shan, B. MicroRNA-577 promotes the sensitivity of chronic myeloid leukemia cells to imatinib by targeting NUP160. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 7008–7015. [Google Scholar] [CrossRef]
- Iqbal, J.; Shen, Y.; Huang, X.; Liu, Y.; Wake, L.; Liu, C.; Deffenbacher, K.; Lachel, C.M.; Wang, C.; Rohr, J.; et al. Global microRNA expression profiling uncovers molecular markers for classification and prognosis in aggressive B-cell lymphoma. Blood 2015, 125, 1137–1145. [Google Scholar] [CrossRef]
- Bento, L.; Vögler, O.; Sas-Barbeito, A.; Muncunill, J.; Ros, T.; Martínez, J.; Quintero-Duarte, A.; Ramos, R.; Asensio, V.J.; Fernández-Rodríguez, C.; et al. Screening for Prognostic microRNAs Associated with Treatment Failure in Diffuse Large B Cell Lymphoma. Cancers 2022, 14, 1065. [Google Scholar] [CrossRef]
- Yuan, J.; Zhang, Q.; Wu, S.; Yan, S.; Zhao, R.; Sun, Y.; Tian, X.; Zhou, K. miRNA-223-3p modulates ibrutinib resistance through regulation of the CHUK/Nf-κb signaling pathway in mantle cell lymphoma. Exp. Hematol. 2021, 103, 52–59.e2. [Google Scholar] [CrossRef]
- Wu, H.; Liu, C.; Yang, Q.; Xin, C.; Du, J.; Sun, F.; Zhou, L. MIR145-3p promotes autophagy and enhances bortezomib sensitivity in multiple myeloma by targeting HDAC4. Autophagy 2020, 16, 683–697. [Google Scholar] [CrossRef]
- Amodio, N.; Gallo Cantafio, M.E.; Botta, C.; Agosti, V.; Federico, C.; Caracciolo, D.; Ronchetti, D.; Rossi, M.; Driessen, C.; Neri, A.; et al. Replacement of miR-155 elicits tumor suppressive activity and antagonizes bortezomib resistance in multiple myeloma. Cancers 2019, 11, 236. [Google Scholar] [CrossRef]
- Xu, J.; Su, Y.; Xu, A.; Fan, F.; Mu, S.; Chen, L.; Chu, Z.; Zhang, B.; Huang, H.; Zhang, J.; et al. miR-221/222-mediated inhibition of autophagy promotes dexamethasone resistance in multiple myeloma. Molecules 2019, 27, 559–570. [Google Scholar] [CrossRef]
- Meng, F.; Henson, R.; Lang, M.; Wehbe, H.; Maheshwari, S.; Mendell, J.T.; Jiang, J.; Schmittgen, T.D.; Patel, T. Involvement of human microRNAs in growth and response to chemotherapy in human cholangiocarcinoma cell lines. Gastroenterology 2006, 130, 2113–2129. [Google Scholar] [CrossRef] [PubMed]
- Azimi, A.; Hagh, M.F.; Yousefi, B.; Rahnama, M.A.; Khorrami, A.; Heydarabad, M.Z.; Najafpour, M.; Hallajzadeh, J.; Ghahremani, A. The Effect of Prednisolone on miR 15a and miR16-1 Expression Levels and Apoptosis in Acute Lymphoblastic Leukemia Cell Line: CCRF-CEM. Drug Res. 2016, 66, 432–435. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, J.C.; Scrideli, C.A.; Brassesco, M.S.; Morales, A.G.; Pezuk, J.A.; de Queiroz, P.; Yunes, J.A.; Brandalise, S.R.; Tone, L.G. Differential miRNA expression in childhood acute lymphoblastic leukemia and association with clinical and biological features. Leuk. Res. 2012, 36, 293–298. [Google Scholar] [CrossRef] [PubMed]
- Capuano, C.; Pighi, C.; Maggio, R.; Battella, S.; Morrone, S.; Palmieri, G.; Santoni, A.; Klein, C.; Galandrini, R. CD16 pre-ligation by defucosylated tumor-targeting mAb sensitizes human NK cells to gammac cytokine stimulation via PI3K/mTOR axis. Cancer Immunol. Immunother. 2020, 69, 501–512. [Google Scholar] [CrossRef] [PubMed]
- Kohnken, R.; Wen, J.; Mundy-Bosse, B.; McConnell, K.; Keiter, A.; Grinshpun, L.; Hartlage, A.; Yano, M.; McNeil, B.; Chakravarti, N.; et al. Diminished microRNA-29b level is associated with BRD4-mediated activation of oncogenes in cutaneous T-cell lymphoma. Blood 2018, 131, 771–781. [Google Scholar] [CrossRef]
- Ferreira, A.F.; Moura, L.G.; Tojal, I.; Ambrósio, L.; Pinto-Simões, B.; Hamerschlak, N.; Calin, G.A.; Ivan, C.; Covas, D.T.; Kashima, S.; et al. ApoptomiRs expression modulated by BCR-ABL is linked to CML progression and imatinib resistance. Blood Cells Mol. Dis. 2014, 53, 47–55. [Google Scholar] [CrossRef]
- Ali Beg, M.M.; Guru, S.A.; Abdullah, S.M.; Ahmad, I.; Rizvi, A.; Akhter, J.; Goyal, J.; Verma, A.K. Regulation of miR-126 and miR-122 Expression and Response of Imatinib Treatment on Its Expression in Chronic Myeloid Leukemia Patients. Oncol. Res. Treat. 2021, 44, 530–537. [Google Scholar] [CrossRef]
- Mirza, M.A.B.; Guru, S.A.; Abdullah, S.M.; Rizvi, A.; Saxena, A. microRNA-21 Expression as Prognostic and Therapeutic Response Marker in Chronic Myeloid Leukaemia Patients. Asian Pac. J. Cancer Prev. 2019, 20, 2379–2383. [Google Scholar] [CrossRef]
- Brüggenwirth, I.M.A.; Martins, P.N. RNA interference therapeutics in organ transplantation: The dawn of a new era. Am. J. Transplant. 2020, 20, 931–941. [Google Scholar] [CrossRef]
- Chakraborty, C.; Sharma, A.R.; Sharma, G.; Lee, S.S. Therapeutic advances of miRNAs: A preclinical and clinical update. J. Adv. Res. 2020, 28, 127–138. [Google Scholar] [CrossRef]
- Van der Ree, M.H.; van der Meer, A.J.; van Nuenen, A.C.; de Bruijne, J.; Ottosen, S.; Janssen, H.L.; Kootstra, N.A.; Reesink, H.W. Miravirsen dosing in chronic hepatitis C patients results in decreased microRNA-122 levels without affecting other microRNAs in plasma. Aliment. Pharmacol. Ther. 2016, 43, 102–113. [Google Scholar] [CrossRef]
- Van Roosbroeck, K.; Calin, G.A. MicroRNAs in chronic lymphocytic leukemia: miRacle or miRage for prognosis and targeted therapies? Semin. Oncol. 2016, 43, 209–214. [Google Scholar] [CrossRef]
- Wu, S.J.; Chen, J.; Wu, B.; Wang, Y.J.; Guo, K.Y. MicroRNA-150 enhances radiosensitivity by inhibiting the AKT pathway in NK/T cell lymphoma. J. Exp. Clin. Cancer Res. 2018, 37, 18. [Google Scholar] [CrossRef]
- Yan, S.; Chen, Y.; Yang, M.; Zhang, Q.; Ma, J.; Liu, B.; Yang, L.; Li, X. hsa-MicroRNA-28-5p Inhibits Diffuse Large B-Cell Lymphoma Cell Proliferation by Downregulating 14-3-3ζ Expression. Evid.-Based Complement. Altern. Med. 2022, 2022, 4605329. [Google Scholar] [CrossRef]
- Schilb, A.L.; Ayat, N.R.; Vaidya, A.M.; Hertz, L.M.; Hall, R.C.; Scheidt, J.H.; Sun, D.; Sun, Z.; Gopalakrishnan, R.; Lu, Z.R. Efficacy of Targeted ECO/miR-200c Nanoparticles for Modulating Tumor Microenvironment and Treating Triple Negative Breast Cancer as Non-invasively Monitored by MR Molecular Imaging. Pharm. Res. 2021, 38, 1405–1418. [Google Scholar] [CrossRef]
- Reid, G.; Williams, M.; Kirschner, M.B.; Mugridge, N.; Weiss, J.; Brahmbhatt, H.; Macdiarmid, J.; Molloy, M.; Lin, R.; Van Zandwijk, N. Abstract 3976: Targeted delivery of a synthetic microRNA-based mimic as an approach to cancer therapy. Cancer Res. 2015, 75 (Suppl. 15). [Google Scholar] [CrossRef]
- Jiang, K.; Zou, H. microRNA-20b-5p overexpression combing Pembrolizumab potentiates cancer cells to radiation therapy via repressing programmed death-ligand 1. Bioengineered 2022, 13, 917–929. [Google Scholar] [CrossRef]
- Yan, J.; Yao, L.; Li, P.; Wu, G.; Lv, X. Long non-coding RNA MIR17HG sponges microRNA-21 to upregulate PTEN and regulate homoharringtonine-based chemoresistance of acute myeloid leukemia cells. Oncol. Lett. 2022, 23, 24. [Google Scholar] [CrossRef]
- Liang, A.L.; Zhang, T.T.; Zhou, N.; Wu, C.Y.; Lin, M.H.; Liu, Y.J. miRNA-10b sponge: An anti-breast cancer study in vitro. Oncol. Rep. 2016, 35, 1950–1958. [Google Scholar] [CrossRef]
- Durmaz, B.; Bagca, B.G.; Cogulu, O.; Susluer, S.Y.; Alpay, A.; Aksoylar, S.; Gunduz, C. Antileukemic Effects of Anti-miR-146a, Anti-miR-155, Anti-miR-181a, and Prednisolone on Childhood Acute Lymphoblastic Leukemia. BioMed Res. Int. 2021, 2021, 3207328. [Google Scholar] [CrossRef]
- Chen, H.; Wang, J.; Wang, H.; Liang, J.; Dong, J.; Bai, H.; Jiang, G. Advances in the application of Let-7 microRNAs in the diagnosis, treatment and prognosis of leukemia. Oncol. Lett. 2022, 23, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Fasihi-Ramandi, M.; Moridnia, A.; Najafi, A.; Sharifi, M. Inducing cell proliferative prevention in human acute promyelocytic leukemia by miR-182 inhibition through modulation of CASP9 expression. Biomed. Pharmacother. 2017, 89, 1152–1158. [Google Scholar] [CrossRef] [PubMed]
- Lai, T.H.; Ewald, B.; Zecevic, A.; Liu, C.; Sulda, M.; Papaioannou, D.; Garzon, R.; Blachly, J.S.; Plunkett, W.; Sampath, D. HDAC Inhibition Induces MicroRNA-182, which Targets RAD51 and Impairs HR Repair to Sensitize Cells to Sapacitabine in Acute Myelogenous Leukemia. Clin. Cancer Res. 2016, 22, 3537–3549. [Google Scholar] [CrossRef] [PubMed]
- Bamodu, O.A.; Kuo, K.T.; Yuan, L.P.; Cheng, W.H.; Lee, W.H.; Ho, Y.S.; Chao, T.Y.; Yeh, C.T. HDAC inhibitor suppresses proliferation and tumorigenicity of drug-resistant chronic myeloid leukemia stem cells through regulation of hsa-miR-196a targeting BCR/ABL1. Exp. Cell Res. 2018, 370, 519–530. [Google Scholar] [CrossRef]
- Abe, F.; Kitadate, A.; Ikeda, S.; Yamashita, J.; Nakanishi, H.; Takahashi, N.; Asaka, C.; Teshima, K.; Miyagaki, T.; Sugaya, M.; et al. Histone deacetylase inhibitors inhibit metastasis by restoring a tumor suppressive microRNA-150 in advanced cutaneous T-cell lymphoma. Oncotarget 2017, 8, 7572–7585. [Google Scholar] [CrossRef]
- Zhang, X.; Tang, J.; Zhi, X.; Xie, K.; Wang, W.; Li, Z.; Zhu, Y.; Yang, L.; Xu, H.; Xu, Z. miR-874 functions as a tumor suppressor by inhibiting angiogenesis through STAT3/VEGF-A pathway in gastric cancer. Oncotarget 2015, 6, 1605–1617. [Google Scholar] [CrossRef]
- Moloudizargari, M.; Hekmatirad, S.; Mofarahe, Z.S.; Asghari, M.H. Exosomal microRNA panels as biomarkers for hematological malignancies. Curr. Probl. Cancer 2021, 45, 100726. [Google Scholar] [CrossRef]
- Hornick, N.I.; Huan, J.; Doron, B.; Goloviznina, N.A.; Lapidus, J.; Chang, B.H.; Kurre, P. Serum Exosome MicroRNA as a Minimally-Invasive Early Biomarker of AML. Sci. Rep. 2015, 5, 11295. [Google Scholar] [CrossRef]
- Jiang, D.; Wu, X.; Sun, X.; Tan, W.; Dai, X.; Xie, Y.; Du, A.; Zhao, Q. Bone mesenchymal stem cell-derived exosomal microRNA-7-5p inhibits progression of acute myeloid leukemia by targeting OSBPL11. J. Nanobiotechnol. 2022, 20, 29. [Google Scholar] [CrossRef]
- Zhang, L.; Khadka, B.; Wu, J.; Feng, Y.; Long, B.; Xiao, R.; Liu, J. Bone Marrow Mesenchymal Stem Cells-Derived Exosomal miR-425-5p Inhibits Acute Myeloid Leukemia Cell Proliferation, Apoptosis, Invasion and Migration by Targeting WTAP. OncoTargets Ther. 2021, 14, 4901–4914. [Google Scholar] [CrossRef]
- Zhang, L.; Zhou, S.; Zhou, T.; Li, X.; Tang, J. Potential of the tumor-derived extracellular vesicles carrying the miR-125b-5p target TNFAIP3 in reducing the sensitivity of diffuse large B cell lymphoma to rituximab. Int. J. Oncol. 2021, 58, 31. [Google Scholar] [CrossRef]
- Feng, Y.; Zhong, M.; Zeng, S.; Wang, L.; Liu, P.; Xiao, X.; Liu, Y. Exosome-derived miRNAs as predictive biomarkers for diffuse large B-cell lymphoma chemotherapy resistance. Epigenomics 2019, 11, 35–51. [Google Scholar] [CrossRef]
- Lee, J.Y.; Ryu, D.; Lim, S.W.; Ryu, K.J.; Choi, M.E.; Yoon, S.E.; Kim, K.; Park, C.; Kim, S.J. Exosomal miR-1305 in the oncogenic activity of hypoxic multiple myeloma cells: A biomarker for predicting prognosis. J. Cancer 2021, 12, 2825–2834. [Google Scholar] [CrossRef]
- Tian, F.; Wang, P.; Lin, D.; Dai, J.; Liu, Q.; Guan, Y.; Zhan, Y.; Yang, Y.; Wang, W.; Wang, J.; et al. Exosome-delivered miR-221/222 exacerbates tumor liver metastasis by targeting SPINT1 in colorectal cancer. Cancer Sci. 2021, 112, 3744–3755. [Google Scholar] [CrossRef]
- Si, W.; Shen, J.; Zheng, H.; Weimin, F. The role and mechanisms of action of microRNAs in cancer drug resistance. Clin. Epigenet. 2019, 11, 25. [Google Scholar] [CrossRef]
- Ye, J.; Zou, M.; Li, P.; Liu, H. MicroRNA Regulation of Energy Metabolism to Induce Chemoresistance in Cancers. Technol. Cancer Res. Treat. 2018, 17, 1533033818805997. [Google Scholar] [CrossRef]
- Mollaei, H.; Safaralizadeh, R.; Rostami, Z. MicroRNA replacement therapy in cancer. J. Cell. Physiol. 2019, 234, 12369–12384. [Google Scholar] [CrossRef]
- De Oliveira, J.C.; Molinari Roberto, G.; Baroni, M.; Bezerra Salomão, K.; Alejandra Pezuk, J.; Sol Brassesco, M. miRNA Dysregulation in Childhood Hematological Cancer. Int. J. Mol. Sci. 2018, 19, 2688. [Google Scholar] [CrossRef]
| Differential Diagnosis | Prognostic Stratification | Chemoresistance | |
|---|---|---|---|
| ALL | miR-128a, miR-128b, let-7b, miR-223 | miR-146a, miR-429, miR-124 hypermethylation, miR-99a, miR-100, miR-16 | miR-206, miR-34a, miR-142-3p, miR-17-92 cluster |
| Pediatric ALL | miR-29c-5p, miR-326, miR-200c | miR-10a, miR-134, miR-214, miR-708, miR-99a, miR-151-5p, miR-451, miR-1290, miR-155, miR-181a, miR-143, miR-182, miR-152 promoter methylation, miR-101-3p, miR-4774-5p, miR-1324, miR-631, miR-4699-5p, miR-922 | miR-125b, miR-99a, miR-100, miR-324-3p, miR-508-5p, miR-18a, miR-532, miR-218, miR-625, miR-193a, miR-638, miR-550, miR-633, miR-21, miR-326, miR-221 |
| AML | miR-128a, miR-128b, let-7b, miR-223, miR-92a/miR-638 ratio | miR-181 family, miR-504-3p, miR-191, miR-199a, miR-195, miR-363 | miR-874-3p, miR-15a-5p, miR-21-5p, miR-9, miR-217, miR-143, miR-204 |
| Pediatric AML | miR-196b, miR-155, miR-25, miR-370 | miR-193b-3p, miR-370, miR-29a, miR-509, miR-542, miR-146a, miR-3667, miR-199a | miR-29a, miR-100, miR-125b |
| CLL | miR-192, miR-32-5p, miR-98-5p, miR-374b-5p, miR-145-5p, miR-185-5p | miR-181b, miR-650, miR-708, miR-29b, miR-29c, miR-18a, miR-19b-1, miR-92a-1, miR-17 | miR-148a, miR-222, miR-21, miR-181a, miR-155 |
| CML | miR-451, miR-222, miR-126, miR-155 | miR-486-5p, miR-320a, miR-150 | miR-217, miR-199b, miR-221, miR-577, miR-451, miR-146a, miR-9, miR-142-5p, miR-365a-3p |
| Lymphomas | let-7f, miR-9, miR-27a, miR-142-3p, miR-155, miR-203, miR-30c | miR-130a, miR-199a, miR-497, miR-34a-5p, miR-22, miR-129-5p, miR-27b, miR-7, miR-223 | miR-125b-5p, miR-155, miR-1244, miR-193b-5p, miR-223-3p |
| MM | miR-34a, let-7e, miR-4254 | miR-223-3p, miR-744, let-7e, miR-720, miR-1246 | miR-145-3p, miR-155, miR-221, miR-222 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gaál, Z. Implication of microRNAs in Carcinogenesis with Emphasis on Hematological Malignancies and Clinical Translation. Int. J. Mol. Sci. 2022, 23, 5838. https://doi.org/10.3390/ijms23105838
Gaál Z. Implication of microRNAs in Carcinogenesis with Emphasis on Hematological Malignancies and Clinical Translation. International Journal of Molecular Sciences. 2022; 23(10):5838. https://doi.org/10.3390/ijms23105838
Chicago/Turabian StyleGaál, Zsuzsanna. 2022. "Implication of microRNAs in Carcinogenesis with Emphasis on Hematological Malignancies and Clinical Translation" International Journal of Molecular Sciences 23, no. 10: 5838. https://doi.org/10.3390/ijms23105838
APA StyleGaál, Z. (2022). Implication of microRNAs in Carcinogenesis with Emphasis on Hematological Malignancies and Clinical Translation. International Journal of Molecular Sciences, 23(10), 5838. https://doi.org/10.3390/ijms23105838





