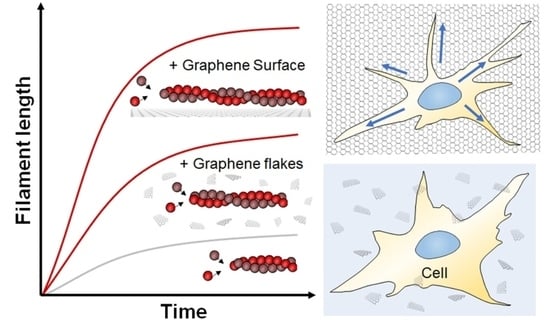
Graphene Enhances Actin Filament Assembly Kinetics and Modulates NIH-3T3 Fibroblast Cell Spreading
Abstract
:1. Introduction
2. Results and Discussions
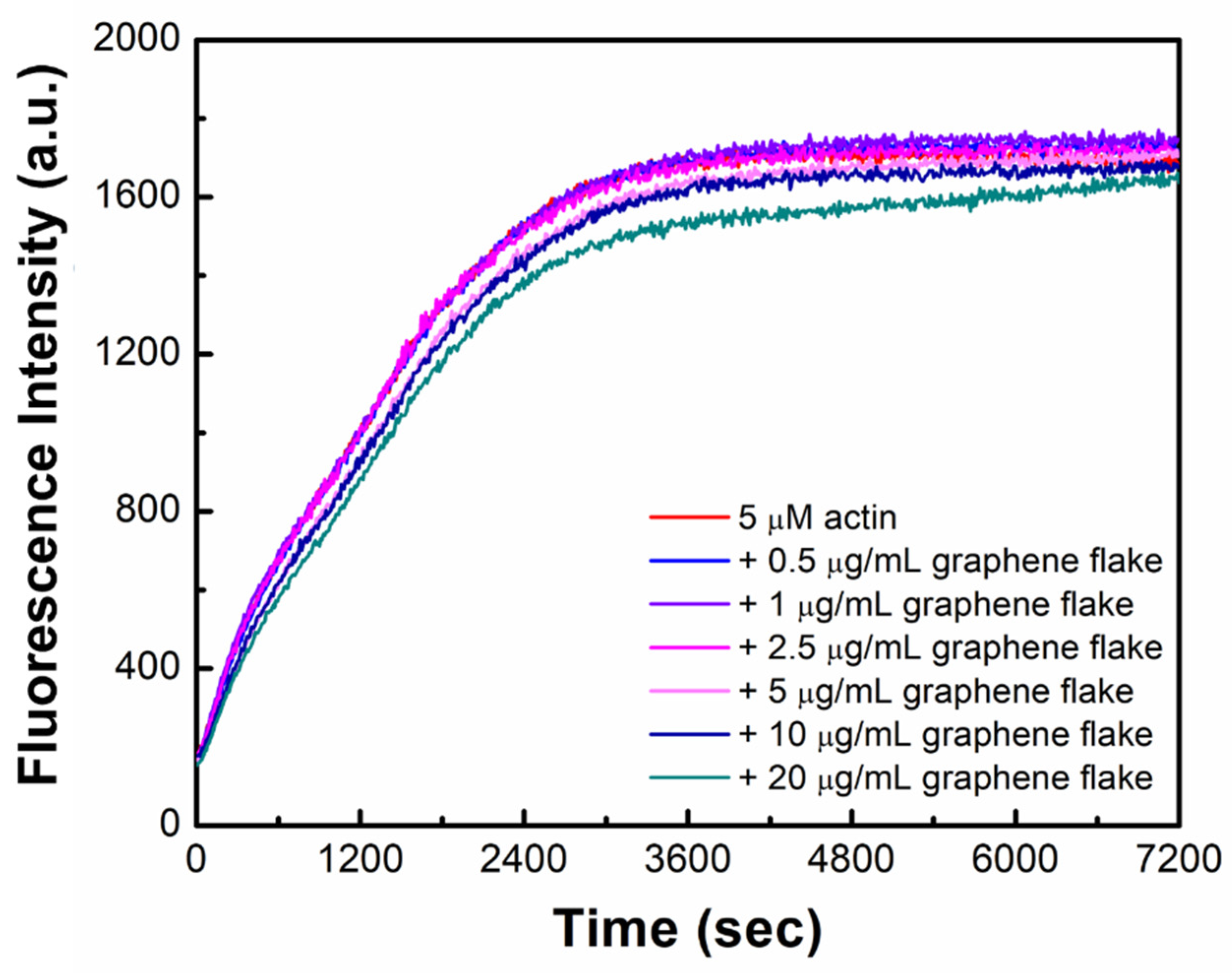
2.1. Graphene Flakes Affect Actin Filament Length without Hampering Polymerization
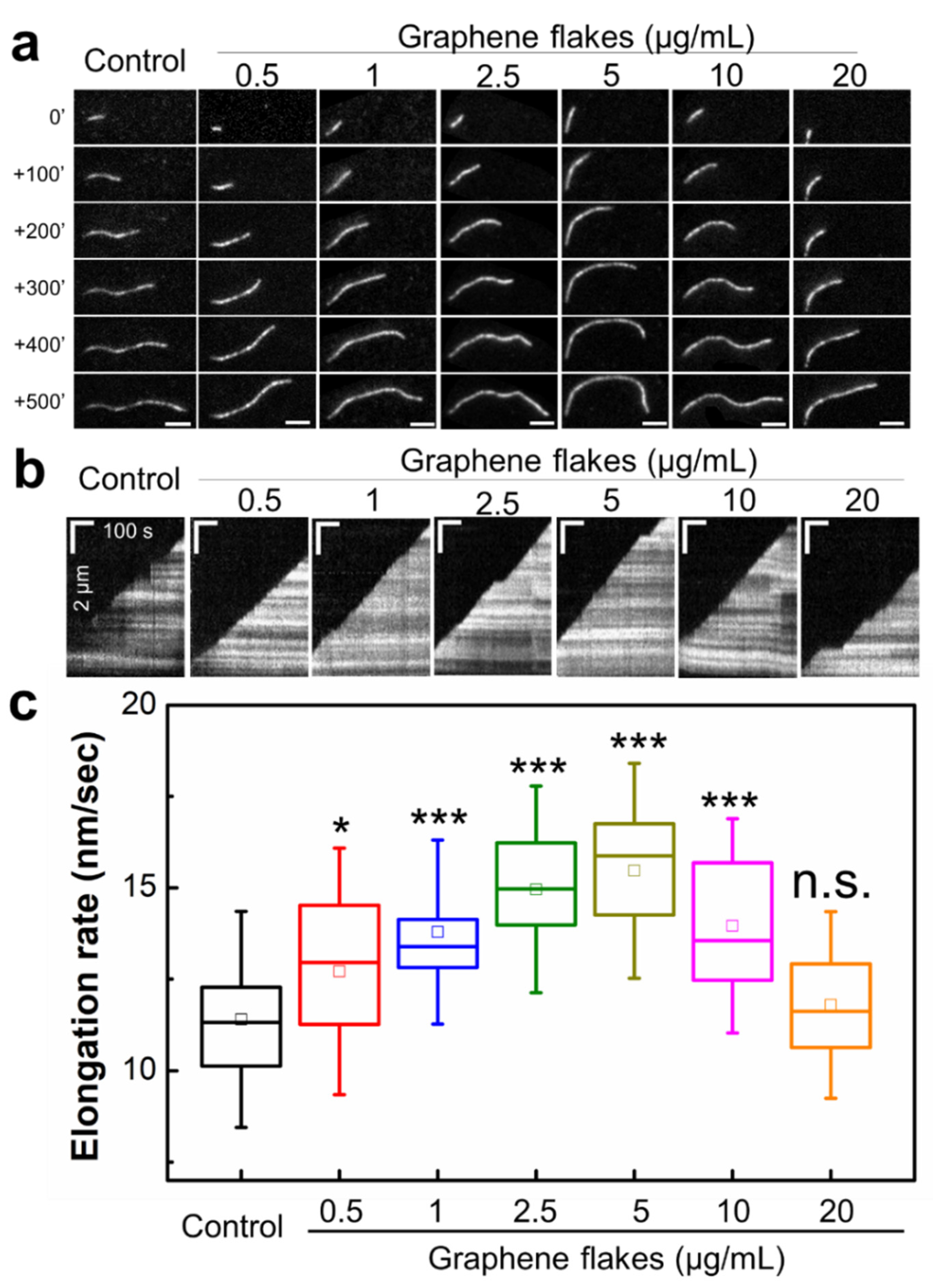
2.2. Graphene Flakes Modulate the Rates of Actin Filament Elongation
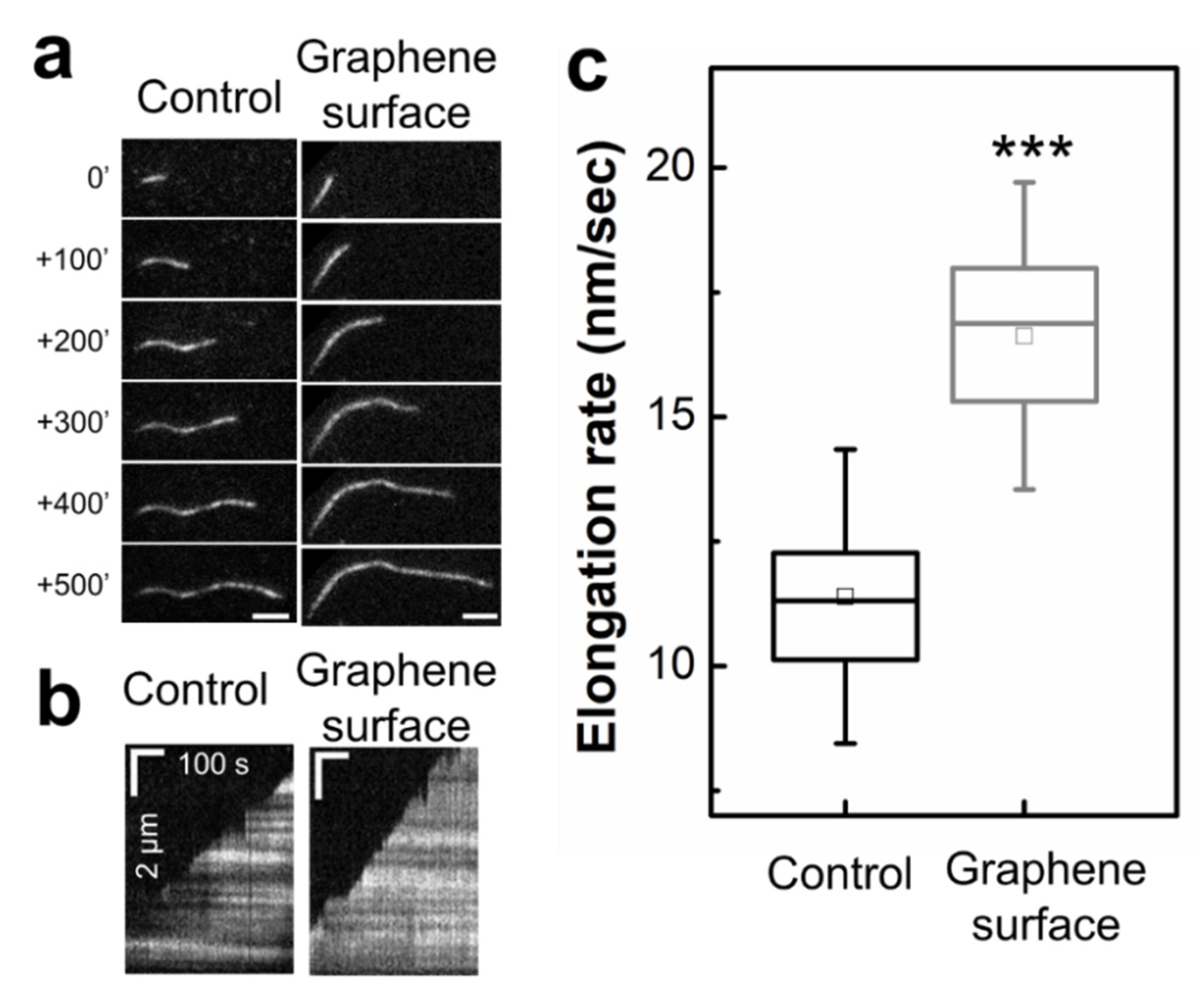
2.3. Graphene Surface Accelerates Actin Filament Elongation
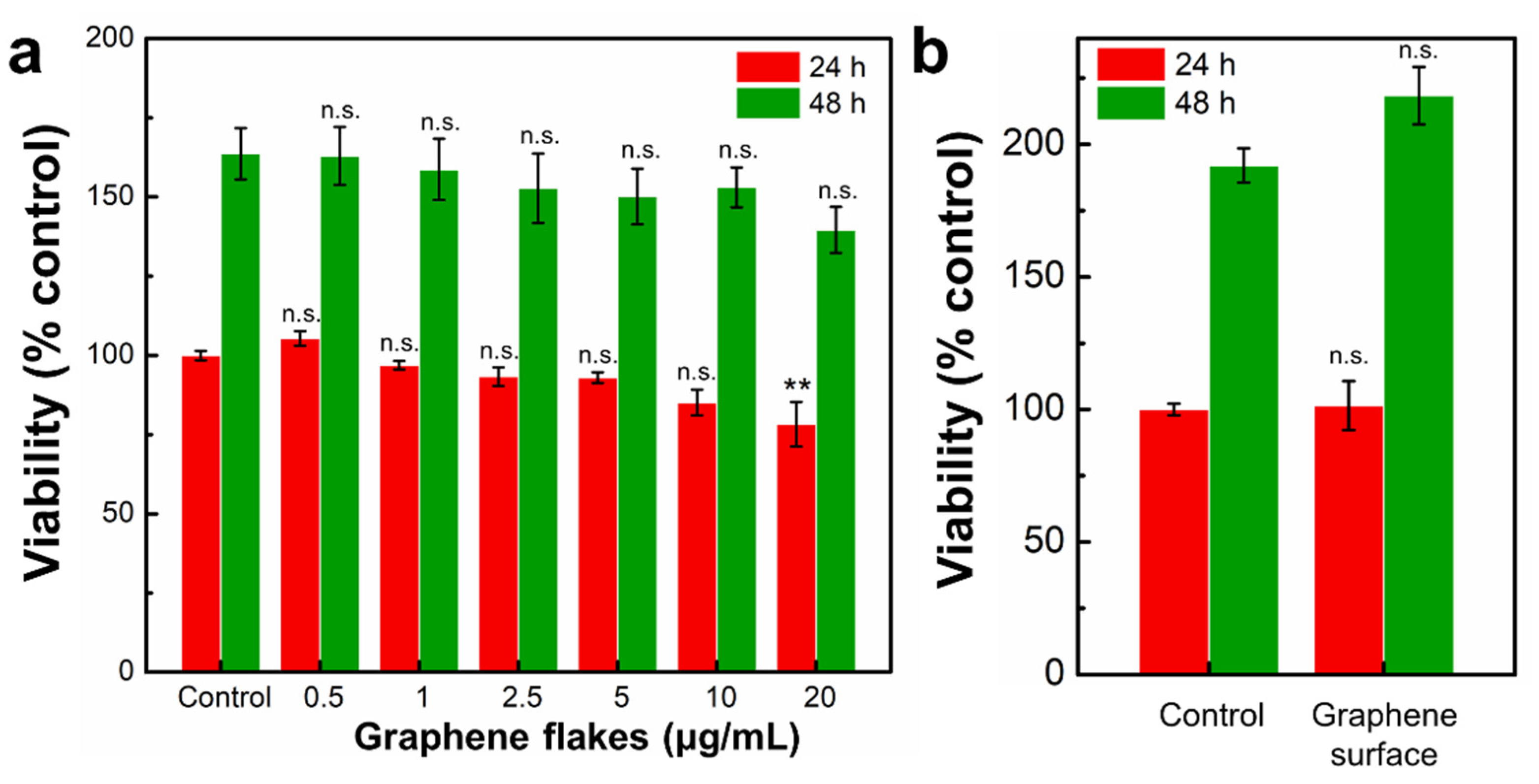
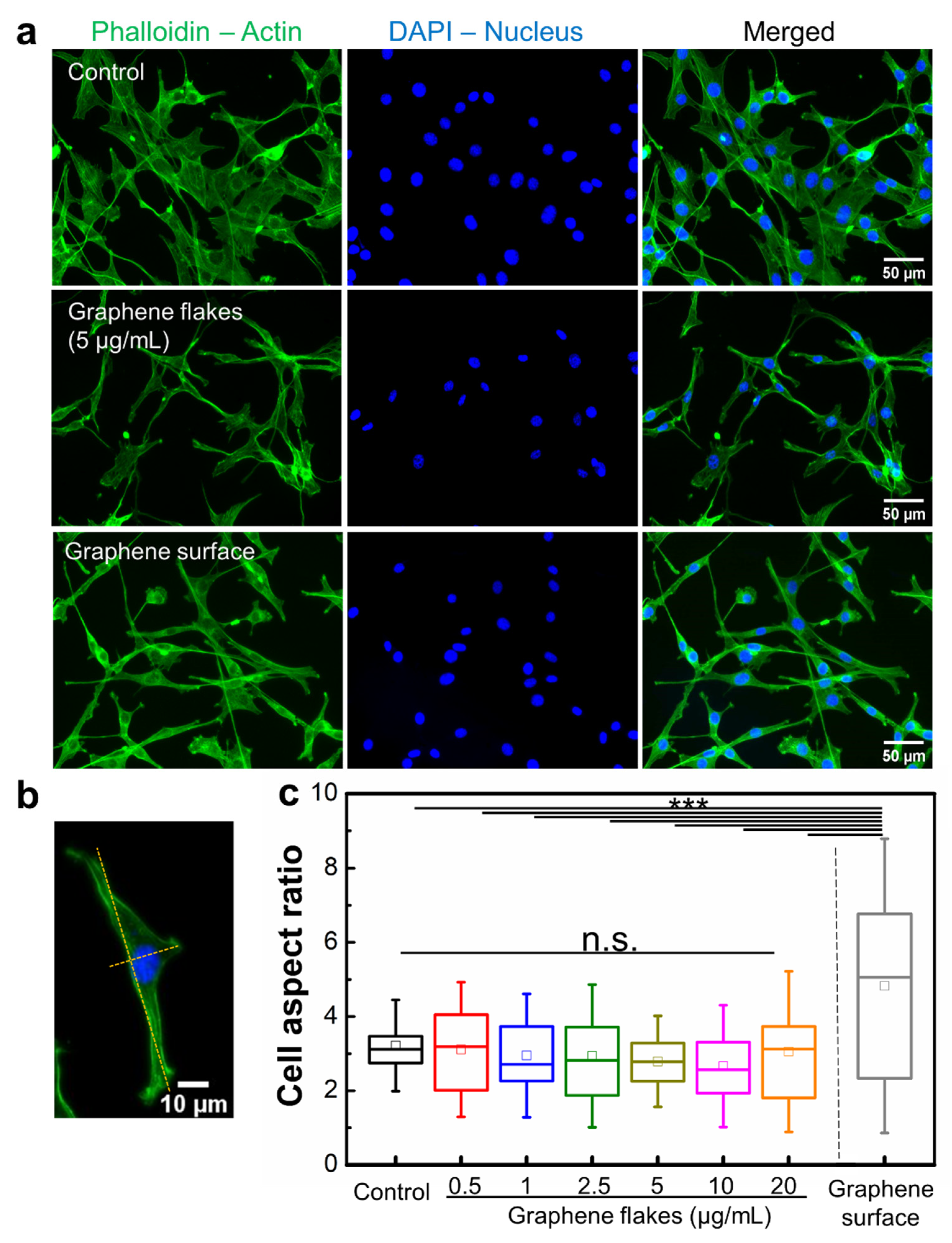
2.4. Non-Cytotoxic Graphene Surface Alters Morphology of Mouse Embryo Fibroblasts
3. Materials and Methods
3.1. Sample Preparation
3.2. Steady-State TIRF Microscopy Imaging and Data Analysis
3.3. Pyrene Assay
3.4. Flow Cell Preparation and Real-Time TIRF Microscopy Imaging
3.5. Cell Culture
3.6. Cell Viability Assay, Cell Imaging, and Morphology Analysis
3.7. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Blanchoin, L.; Boujemaa-Paterski, R.; Sykes, C.; Plastino, J. Actin dynamics, architecture, and mechanics in cell motility. Physiol. Rev. 2014, 94, 235–263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fletcher, D.A.; Mullins, R.D. Cell mechanics and the cytoskeleton. Nature 2010, 463, 485–492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pollard, T.D.; Cooper, J.A. Actin, a central player in cell shape and movement. Science 2009, 326, 1208–1212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scipion, C.P.M.; Ghoshdastider, U.; Ferrer, F.J.; Yuen, T.-Y.; Wongsantichon, J.; Robinson, R.C. Structural evidence for the roles of divalent cations in actin polymerization and activation of ATP hydrolysis. Proc. Natl. Acad. Sci. USA 2018, 115, 10345–10350. [Google Scholar] [CrossRef] [Green Version]
- Dominguez, R.; Holmes, K.C. Actin structure and function. Annu. Rev. Biophys. 2011, 40, 169–186. [Google Scholar] [CrossRef] [Green Version]
- Pollard, T.D. Actin and actin-binding proteins. Cold Spring Harb. Perspect. Biol. 2016, 8, a018226. [Google Scholar] [CrossRef] [Green Version]
- Kang, H.; Bradley, M.J.; McCullough, B.R.; Pierre, A.; Grintsevich, E.E.; Reisler, E.; de la Cruz, E.M. Identification of cation-binding sites on actin that drive polymerization and modulate bending stiffness. Proc. Natl. Acad. Sci. USA 2012, 109, 16923–16927. [Google Scholar] [CrossRef] [Green Version]
- Kang, H.; Bradley, M.J.; Cao, W.; Zhou, K.; Grintsevich, E.E.; Michelot, A.; Sindelar, C.V.; Hochstrasser, M.; Enrique, M. Site-specific cation release drives actin filament severing by vertebrate cofilin. Proc. Natl. Acad. Sci. USA 2014, 111, 17821–17826. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuznetsova, I.M.; Turoverov, K.K.; Uversky, V.N. What macromolecular crowding can do to a protein. Int. J. Mol. Sci. 2014, 15, 23090–23140. [Google Scholar] [CrossRef] [Green Version]
- Frederick, K.B.; Sept, D.; de la Cruz, E.M. Effects of solution crowding on actin polymerization reveal the energetic basis for nucleotide-dependent filament stability. J. Mol. Biol. 2008, 378, 540–550. [Google Scholar] [CrossRef] [Green Version]
- Rosin, C.; Estel, K.; Halker, J.; Winter, R. Combined effects of temperature, pressure, and co-solvents on the polymerization kinetics of actin. ChemPhysChem 2015, 16, 1379–1385. [Google Scholar] [CrossRef]
- Pernier, J.; Shekhar, S.; Jegou, A.; Guichard, B.; Carlier, M.-F. Profilin interaction with actin filament barbed end controls dynamic instability, capping, branching, and motility. Dev. Cell 2016, 36, 201–214. [Google Scholar] [CrossRef] [Green Version]
- Ghosh, M.; Song, X.; Mouneimne, G.; Sidani, M.; Lawrence, D.S.; Condeelis, J.S. Cofilin promotes actin polymerization and defines the direction of cell motility. Science 2004, 304, 743–746. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, Y.; Sun, H.; Li, X.; Li, X.; Zhao, L. Impact of carbon nanomaterials on actin polymerization. J. Nanosci. Nanotechnol. 2016, 16, 2408–2417. [Google Scholar] [CrossRef]
- Holt, B.D.; Shams, H.; Horst, T.A.; Basu, S.; Rape, A.D.; Wang, Y.-L.; Rohde, G.K.; Mofrad, M.R.K.; Islam, M.F.; Dahl, K.N. Altered cell mechanics from the inside: Dispersed single wall carbon nanotubes integrate with and restructure actin. J. Funct. Biomater. 2012, 3, 398–417. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tian, X.; Yang, Z.; Duan, G.; Wu, A.; Gu, Z.; Zhang, L.; Chen, C.; Chai, Z.; Ge, C.; Zhou, R. Graphene oxide nanosheets retard cellular migration via disruption of actin cytoskeleton. Small 2017, 13, 1602133. [Google Scholar] [CrossRef]
- Ghorbani, M.; Soleymani, H.; Hashemzadeh, H.; Mortezazadeh, S.; Sedghi, M.; Shojaeilangari, S.; Allahverdi, A.; Naderi-Manesh, H. Microfluidic investigation of the effect of graphene oxide on mechanical properties of cell and actin cytoskeleton networks: Experimental and theoretical approaches. Sci. Rep. 2021, 11, 16216. [Google Scholar] [CrossRef]
- Holt, B.D.; Short, P.A.; Rape, A.D.; Wang, Y.-L.; Islam, M.F.; Dahl, K.N. Carbon nanotubes reorganize actin structures in cells and ex vivo. ACS Nano 2010, 4, 4872–4878. [Google Scholar] [CrossRef]
- Hamzah, A.A.; Selvarajan, R.S.; Majlis, B. Graphene for biomedical applications: A review. Sains Malays. 2017, 46, 1125–1139. [Google Scholar] [CrossRef]
- Zhu, Y.; Murali, S.; Cai, W.; Li, X.; Suk, J.W.; Potts, J.R.; Ruoff, R.S. Graphene and graphene oxide: Synthesis, properties, and applications. Adv. Mater. 2010, 22, 3906–3924. [Google Scholar] [CrossRef]
- Avouris, P. Graphene: Electronic and photonic properties and devices. Nano Lett. 2010, 10, 4285–4294. [Google Scholar] [CrossRef]
- Papageorgiou, D.G.; Kinloch, I.A.; Young, R.J. Mechanical properties of graphene and graphene-based nanocomposites. Prog. Mater. Sci. 2017, 90, 75–127. [Google Scholar] [CrossRef]
- Liao, C.; Li, Y.; Tjong, S.C. Graphene nanomaterials: Synthesis, biocompatibility, and cytotoxicity. Int. J. Mol. Sci. 2018, 19, 3564. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Syama, S.; Mohanan, P.V. Safety and biocompatibility of graphene: A new generation nanomaterial for biomedical application. Int. J. Biol. Macromol. 2016, 86, 546–555. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Parekh, S.H. Linking graphene-based material physicochemical properties with molecular adsorption, structure and cell fate. Commun. Chem. 2020, 3, 8. [Google Scholar] [CrossRef] [Green Version]
- Iannazzo, D.; Pistone, A.; Salamò, M.; Galvagno, S.; Romeo, R.; Giofré, S.V.; Branca, C.; Visalli, G.; di Pietro, A. Graphene quantum dots for cancer targeted drug delivery. Int. J. Pharm. 2017, 518, 185–192. [Google Scholar] [CrossRef]
- Bai, Y.; Xu, T.; Zhang, X. Graphene-based biosensors for detection of biomarkers. Micromachines 2020, 11, 60. [Google Scholar] [CrossRef] [Green Version]
- Rahman, M.; Ahmad, M.Z.; Ahmad, J.; Firdous, J.; Ahmad, F.J.; Mushtaq, G.; Kamal, M.A.; Akhter, S. Role of graphene nano-composites in cancer therapy: Theranostic applications, metabolic fate and toxicity issues. Curr. Drug Metab. 2015, 16, 397–409. [Google Scholar] [CrossRef]
- Feng, L.; Wu, L.; Qu, X. New horizons for diagnostics and therapeutic applications of graphene and graphene oxide. Adv. Mater. 2013, 25, 168–186. [Google Scholar] [CrossRef]
- Zhou, H.; Zhao, K.; Li, W.; Yang, N.; Liu, Y.; Chen, C.; Wei, T. The interactions between pristine graphene and macrophages and the production of cytokines/chemokines via TLR- and NF-κB-related signaling pathways. Biomaterials 2012, 33, 6933–6942. [Google Scholar] [CrossRef] [PubMed]
- Sasidharan, A.; Panchakarla, L.S.; Chandran, P.; Menon, D.; Nair, S.; Rao, C.N.R.; Koyakutty, M. Differential nano-bio interactions and toxicity effects of pristine versus functionalized graphene. Nanoscale 2011, 3, 2461–2464. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, A.; Basu, S.; Singha, S.; Patra, H.K. Inner-view of nanomaterial incited protein conformational changes: Insights into designable interaction. Research 2018, 2018, 9712832. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, H.; Zhang, B.; Zheng, J.; Yu, M.; Zhou, T.; Zhao, K.; Jia, Y.; Gao, X.; Chen, C.; Wei, T. The inhibition of migration and invasion of cancer cells by graphene via the impairment of mitochondrial respiration. Biomaterials 2014, 35, 1597–1607. [Google Scholar] [CrossRef]
- Li, T.; Oloyede, A.; Gu, Y. Adhesive characteristics of low dimensional carbon nanomaterial on actin. Appl. Phys. Lett. 2014, 104, 023702. [Google Scholar] [CrossRef] [Green Version]
- Kuziel, A.W.; Milowska, K.Z.; Chau, P.-L.; Boncel, S.; Koziol, K.K.; Yahya, N.; Payne, M.C. The true amphipathic nature of graphene flakes: A versatile 2D stabilizer. Adv. Mater. 2020, 32, 2000608. [Google Scholar] [CrossRef]
- Kuhn, J.R.; Pollard, T.D. Real-time measurements of actin filament polymerization by total internal reflection fluorescence microscopy. Biophys. J. 2005, 88, 1387–1402. [Google Scholar] [CrossRef] [Green Version]
- Korn, E.D.; Carlier, M.-F.; Pantaloni, D. Actin polymerization and ATP hydrolysis. Science 1987, 238, 638–644. [Google Scholar] [CrossRef]
- Pollard, T.D.; Blanchoin, L.; Mullins, R.D. Molecular mechanisms controlling actin filament dynamics in nonmuscle cells. Annu. Rev. Biophys. Biomol. Struct. 2000, 29, 545–576. [Google Scholar] [CrossRef] [Green Version]
- Minton, A.P. Implications of macromolecular crowding for protein assembly. Curr. Opin. Struct. Biol. 2000, 10, 34–39. [Google Scholar] [CrossRef]
- Minton, A.P. The influence of macromolecular crowding and macromolecular confinement on biochemical reactions in physiological media. J. Biol. Chem. 2001, 276, 10577–10580. [Google Scholar] [CrossRef] [Green Version]
- Rashid, R.; Chee, S.M.L.; Raghunath, M.; Wohland, T. Macromolecular crowding gives rise to microviscosity, anomalous diffusion and accelerated actin polymerization. Phys. Biol. 2015, 12, 034001. [Google Scholar] [CrossRef]
- Geigenfeind, T.; Heras, D.D.L. Principal component analysis of the excluded area of two-dimensional hard particles. J. Chem. Phys. 2019, 150, 184906. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, V.C.; Jachak, A.; Hurt, R.H.; Kane, A.B. Biological interactions of graphene-family nanomaterials: An interdisciplinary review. Chem. Res. Toxicol. 2012, 25, 15–34. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Whitesides, G.M.; Mathias, J.P.; Seto, C.T. Molecular self-assembly and nanochemistry: A chemical strategy for the synthesis of nanostructures. Science 1991, 254, 1312–1319. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Adams, D.J.; Marlow, M.; Zelzer, M. Surface-mediated supramolecular self-assembly of protein, peptide, and nucleoside derivatives: From surface design to the underlying mechanism and tailored functions. Langmuir 2018, 34, 15109–15125. [Google Scholar] [CrossRef] [Green Version]
- Nayak, A.; Dutta, A.K.; Belfort, G. Surface-enhanced nucleation of insulin amyloid fibrillation. Biochem. Biophys. Res. Commun. 2008, 369, 303–307. [Google Scholar] [CrossRef]
- Ryoo, S.R.; Kim, Y.K.; Kim, M.H.; Min, D.H. Behaviors of NIH-3T3 fibroblasts on graphene/carbon nanotubes: Proliferation, focal adhesion, and gene transfection studies. ACS Nano 2010, 4, 6587–6598. [Google Scholar] [CrossRef]
- Lin, F.; Du, F.; Huang, J.; Chau, A.; Zhou, Y.; Duan, H.; Wang, J.; Xiong, C. Substrate effect modulates adhesion and proliferation of fibroblast on graphene layer. Colloids Surf. B Biointerfaces 2016, 146, 785–793. [Google Scholar] [CrossRef]
- Duan, G.; Zhang, Y.; Luan, B.; Weber, J.K.; Zhou, R.W.; Yang, Z.; Zhao, L.; Xu, J.; Luo, J.; Zhou, R. Graphene-Induced pore formation on cell membranes. Sci. Rep. 2017, 7, 42767. [Google Scholar] [CrossRef] [Green Version]
- Liao, K.-H.; Lin, Y.-S.; Macosko, C.W.; Haynes, C.L. Cytotoxicity of graphene oxide and graphene in human erythrocytes and skin fibroblasts. ACS Appl. Mater. Interfaces 2011, 3, 2607–2615. [Google Scholar] [CrossRef]
- Lasocka, I.; Szulc-Dąbrowska, L.; Skibniewski, M.; Skibniewska, E.; Gregorczyk-Zboroch, K.; Pasternak, I.; Hubalek Kalbacova, M. Cytocompatibility of graphene monolayer and its impact on focal cell adhesion, mitochondrial morphology and activity in BALB/3T3 fibroblasts. Materials 2021, 14, 643. [Google Scholar] [CrossRef] [PubMed]
- Lasocka, I.; Szulc-Dabrowska, L.; Skibniewski, M.; Skibniewska, E.; Strupinski, W.; Pasternak, I.; Kmiec, H.; Kowalczyk, P. Biocompatibility of pristine graphene monolayer: Scaffold for fibroblasts. Toxicol. In Vitro 2018, 48, 276–285. [Google Scholar] [CrossRef]
- Zhang, K.; Arranja, A.; Chen, H.; Mytnyk, S.; Wang, Y.; Oldenhof, S.; van Esch, J.H.; Mendes, E. A nano-fibrous platform of copolymer patterned surfaces for controlled cell alignment. RSC Adv. 2018, 8, 21777–21785. [Google Scholar] [CrossRef] [Green Version]
- Mancia, A.; Elliott, J.T.; Halter, M.; Bhadriraju, K.; Tona, A.; Spurlin, T.A.; Middlebrooks, B.L.; Baatz, J.E.; Warr, G.W.; Plant, A.L. Quantitative methods to characterize morphological properties of cell lines. Biotechnol. Prog. 2012, 28, 1069–1078. [Google Scholar] [CrossRef]
- Jeong, J.-T.; Choi, M.-K.; Sim, Y.; Lim, J.-T.; Kim, G.-S.; Seong, M.-J.; Hyung, J.-H.; Kim, K.S.; Umar, A.; Lee, S.-K. Effect of graphene oxide ratio on the cell adhesion and growth behavior on a graphene oxide-coated silicon substrate. Sci. Rep. 2016, 6, 33835. [Google Scholar] [CrossRef]
- Kim, D.-H.; Han, K.; Gupta, K.; Kwon, K.W.; Suh, K.-Y.; Levchenko, A. Mechanosensitivity of fibroblast cell shape and movement to anisotropic substratum topography gradients. Biomaterials 2009, 30, 5433–5444. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, T.-J.; Park, S.; Bhang, S.H.; Yoon, J.-K.; Jo, I.; Jeong, G.-J.; Hong, B.H.; Kim, B.-S. Graphene enhances the cardiomyogenic differentiation of human embryonic stem cells. Biochem. Biophys. Res. Commun. 2014, 452, 174–180. [Google Scholar] [CrossRef]
- Ayala, R.; Zhang, C.; Yang, D.; Hwang, Y.; Aung, A.; Shroff, S.S.; Arce, F.T.; Lal, R.; Arya, G.; Varghese, S. Engineering the cell–material interface for controlling stem cell adhesion, migration, and differentiation. Biomaterials 2011, 32, 3700–3711. [Google Scholar] [CrossRef] [PubMed]
- Yilbas, B.S.; Ibrahim, A.; Ali, H.; Khaled, M.; Laoui, T. Hydrophobic and optical characteristics of graphene and graphene oxide films transferred onto functionalized silica particles deposited glass surface. Appl. Surf. Sci. 2018, 442, 213–223. [Google Scholar] [CrossRef]
- Tamada, Y.; Ikada, Y. Effect of preadsorbed proteins on cell adhesion to polymer surfaces. J. Colloid Interface Sci. 1993, 155, 334–339. [Google Scholar] [CrossRef] [Green Version]
- Dowling, D.P.; Miller, I.S.; Ardhaoui, M.; Gallagher, W.M. Effect of surface wettability and topography on the adhesion of osteosarcoma cells on plasma-modified polystyrene. J. Biomater. Appl. 2011, 26, 327–347. [Google Scholar] [CrossRef]
- McGrath, J.L. Cell spreading: The power to simplify. Curr. Biol. 2007, 17, R357–R358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Spudich, J.A.; Watt, S. The regulation of rabbit skeletal muscle contraction. I. Biochemical studies of the interaction of the tropomyosin-troponin complex with actin and the proteolytic fragments of myosin. J. Biol. Chem. 1971, 246, 4866–4871. [Google Scholar] [CrossRef]
- Castaneda, N.; Lee, M.; Rivera-Jacquez, H.J.; Marracino, R.R.; Merlino, T.R.; Kang, H. Actin filament mechanics and structure in crowded environments. J. Phys. Chem. B 2019, 123, 2770–2779. [Google Scholar] [CrossRef]
- Chan, J.; Venugopal, A.; Pirkle, A.; McDonnell, S.; Hinojos, D.; Magnuson, C.W.; Ruoff, R.S.; Colombo, L.; Wallace, R.M.; Vogel, E.M. Reducing extrinsic performance-limiting factors in graphene grown by chemical vapor deposition. ACS Nano 2012, 6, 3224–3229. [Google Scholar] [CrossRef]
- Castaneda, N.; Zheng, T.; Rivera-Jacquez, H.J.; Lee, H.J.; Hyun, J.; Balaeff, A.; Huo, Q.; Kang, H. Cations modulate actin bundle mechanics, assembly dynamics, and structure. J. Phys. Chem. B 2018, 122, 3826–3835. [Google Scholar] [CrossRef] [PubMed]
- Graham, J.S.; McCullough, B.R.; Kang, H.; Elam, W.A.; Cao, W.; de la Cruz, E.M. Multi-platform compatible software for analysis of polymer bending mechanics. PLoS ONE 2014, 9, e94766. [Google Scholar] [CrossRef] [PubMed]
- Heidings, J.B.; Demosthene, B.; Merlino, T.R.; Castaneda, N.; Kang, E.H. Gelsolin-mediated actin filament severing in crowded environments. Biochem. Biophys. Res. Commun. 2020, 532, 548–554. [Google Scholar] [CrossRef]
- Winterhoff, M.; Brühmann, S.; Franke, C.; Breitsprecher, D.; Faix, J. Visualization of Actin Assembly and Filament Turnover by In Vitro Multicolor TIRF Microscopy. In Chemotaxis: Methods and Protocols; Jin, T., Hereld, D., Eds.; Springer: New York, NY, USA, 2016; pp. 287–306. [Google Scholar]
- Jiang, Z.; Feng, B.; Xu, J.; Qing, T.; Zhang, P.; Qing, Z. Graphene biosensors for bacterial and viral pathogens. Biosens. Bioelectron. 2020, 166, 112471. [Google Scholar] [CrossRef]
- Li, D.; Zhang, W.; Yu, X.; Wang, Z.; Su, Z.; Wei, G. When biomolecules meet graphene: From molecular level interactions to material design and applications. Nanoscale 2016, 8, 19491–19509. [Google Scholar] [CrossRef]
- Piguet, F.; Ouldali, H.; Discala, F.; Breton, M.-F.; Behrends, J.C.; Pelta, J.; Oukhaled, A. High temperature extends the range of size discrimination of nonionic polymers by a biological nanopore. Sci. Rep. 2016, 6, 38675. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Park, J.; Kravchuk, P.; Krishnaprasad, A.; Roy, T.; Kang, E.H. Graphene Enhances Actin Filament Assembly Kinetics and Modulates NIH-3T3 Fibroblast Cell Spreading. Int. J. Mol. Sci. 2022, 23, 509. https://doi.org/10.3390/ijms23010509
Park J, Kravchuk P, Krishnaprasad A, Roy T, Kang EH. Graphene Enhances Actin Filament Assembly Kinetics and Modulates NIH-3T3 Fibroblast Cell Spreading. International Journal of Molecular Sciences. 2022; 23(1):509. https://doi.org/10.3390/ijms23010509
Chicago/Turabian StylePark, Jinho, Pavlo Kravchuk, Adithi Krishnaprasad, Tania Roy, and Ellen Hyeran Kang. 2022. "Graphene Enhances Actin Filament Assembly Kinetics and Modulates NIH-3T3 Fibroblast Cell Spreading" International Journal of Molecular Sciences 23, no. 1: 509. https://doi.org/10.3390/ijms23010509
APA StylePark, J., Kravchuk, P., Krishnaprasad, A., Roy, T., & Kang, E. H. (2022). Graphene Enhances Actin Filament Assembly Kinetics and Modulates NIH-3T3 Fibroblast Cell Spreading. International Journal of Molecular Sciences, 23(1), 509. https://doi.org/10.3390/ijms23010509