Representative Bacillus sp. AM1 from Gut Microbiota Harbor Versatile Molecular Pathways for Bisphenol A Biodegradation
Abstract
1. Introduction
2. Results and Discussion
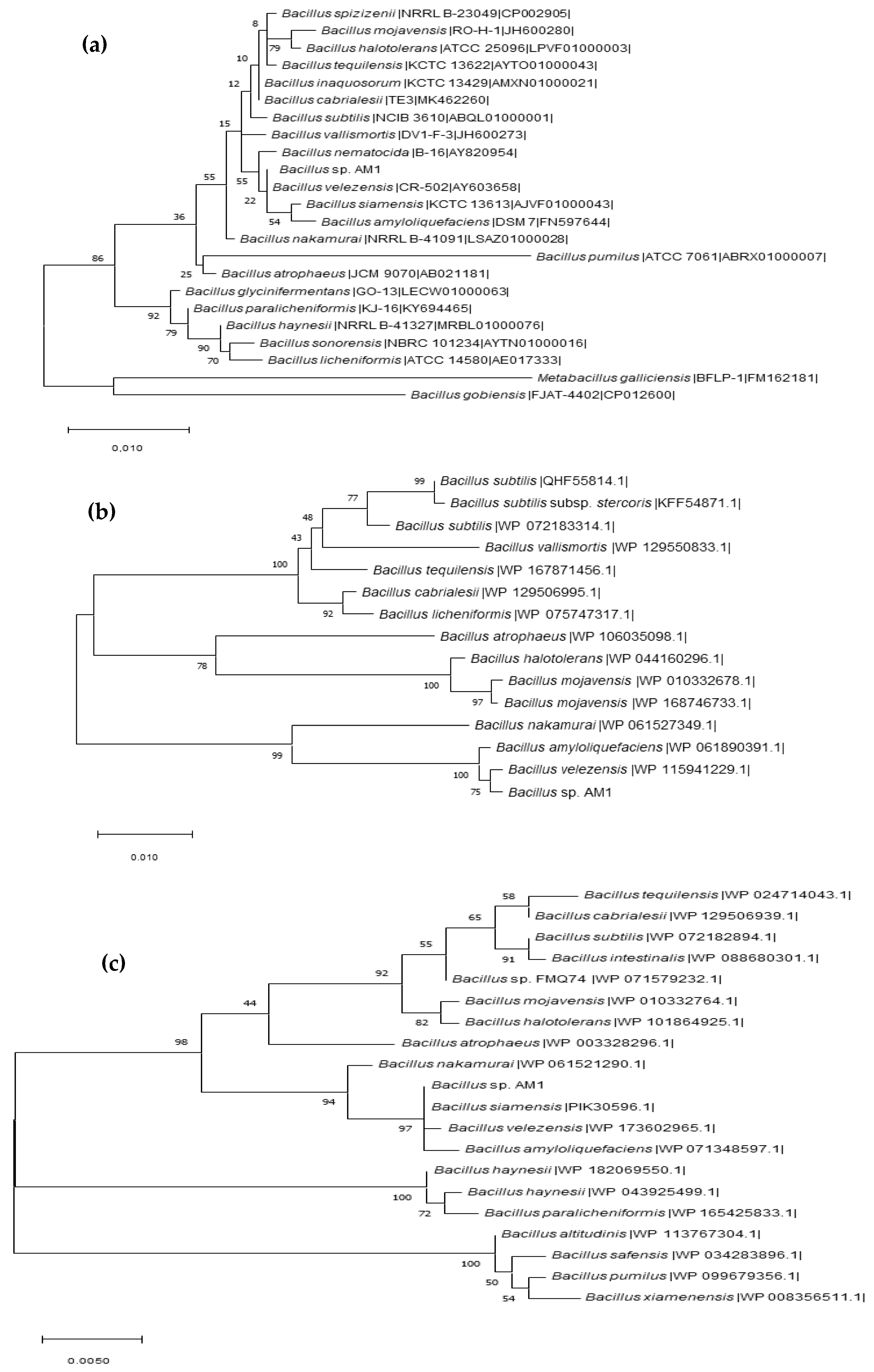
2.1. BPA-Tolerant Microorganisms Isolated from Human Microbiota: 16S RNA Partial Gene Identification and Phylogenetic Analysis
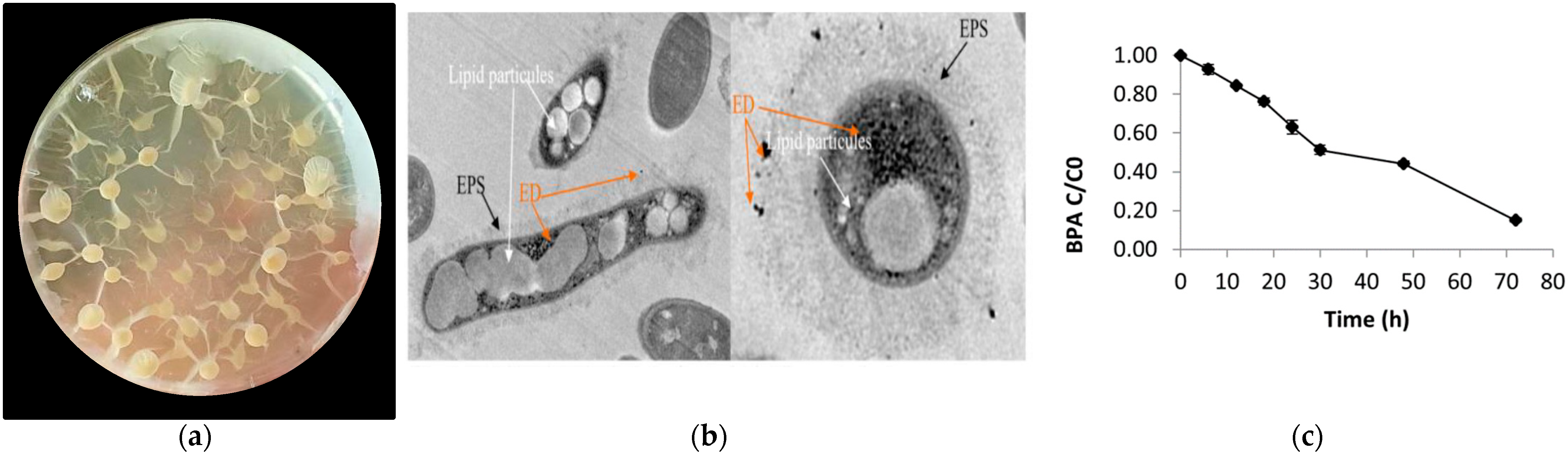
2.2. Identification of the Representative BPA-Tolerant Microbiota Strain and Morphological Features
2.3. Whole Genome Sequence of Bacillus sp. AM1 Analysis
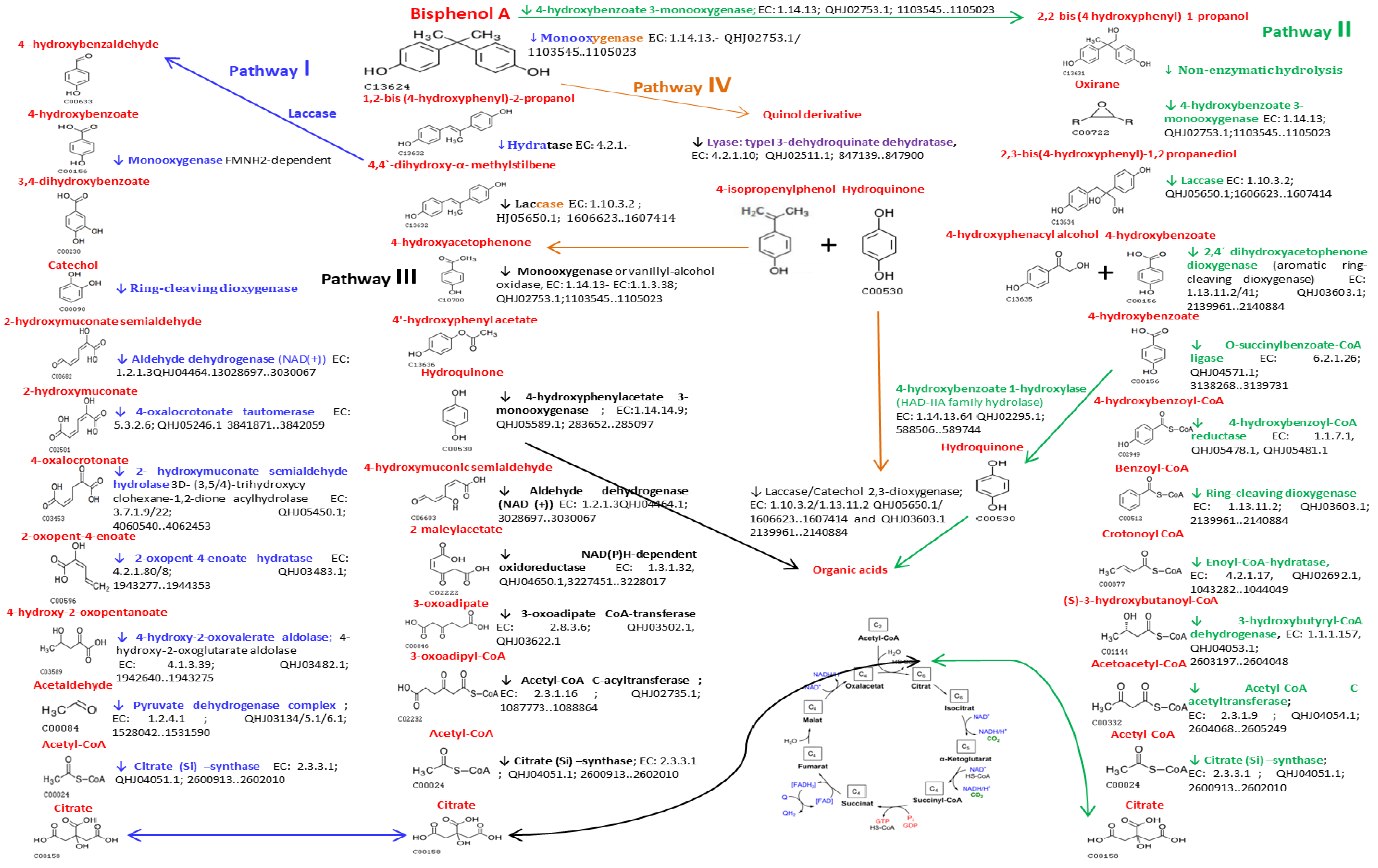
2.4. Description of Putative BPA Biodegradation Pathways by Microbiota BPA-Tolerant Isolate
3. Materials and Methods
3.1. Tolerant-BPA bacterial Isolation from Human Gut Microbiota with Biotechnological Interest
3.2. Genomic DNA from Tolerant-BPA Bacterial Isolates, Taxonomy Identification and Phylogenetic Analysis
3.3. Morphological Features and Identification of BPA-Tolerant Bacterial Strains
3.4. BPA Tolerance Testing
3.5. Whole Genome Sequence of BPA-Tolerant Bacterial Strain
3.6. Whole Genome Sequence Analyses
3.6.1. Phylogenomic Analysis
3.6.2. Molecular and Gene Prediction and Annotations
3.6.3. Identification of Microbial Enzymes Isolated from Bisphenol A Tolerant Microorganisms
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
Patents
References
- Gibson, M.K.; Crofts, T.S.; Dantas, G. Antibiotics and the developing infant gut microbiota and resistome. Curr. Opin. Microbiol. 2015, 27, 51–56. [Google Scholar] [CrossRef]
- Pagliari, D.; Gambassi, G.; Piccirillo, C.A.; Cianci, R. The intricate link among gut “Immunological Niche,” Microbiota, and Xenobiotics in intestinal pathology. Mediators Inflamm. 2017, 8390595. [Google Scholar] [CrossRef] [PubMed]
- Andújar, N.; Gálvez-Ontiveros, Y.; Zafra-Gómez, A.; Rodrigo, L.; Álvarez-Cubero, M.J.; Aguilera, M.; Monteagudo, C.; Rivas, A.A. Bisphenol A analogues in food and their hormonal and obesogenic effects: A Review. Nutrients 2019, 11, 2136. [Google Scholar] [CrossRef]
- Aguilera, M.; Gálvez-Ontiveros, Y.; Rivas, A. Endobolome, a new concept for determining the influence of microbiota disrupting chemicals (MDC) in relation to specific endocrine pathogenesis. Front. Microbiol. 2020, 11, 578007. [Google Scholar] [CrossRef] [PubMed]
- Gálvez-Ontiveros, Y.; Páez, S.; Monteagudo, C.; Rivas, A. Endocrine disruptors in food: Impact on gut microbiota and metabolic diseases. Nutrients 2020, 12, 1158. [Google Scholar] [CrossRef]
- Verbanck, M.; Canouil, M.; Leloire, A.; Dhennin, V.; Coumoul, X.; Yengo, L.; Froguel, P.; Poulain-Godefroy, O. Low-dose exposure to bisphenols A, F and S of human primary adipocyte impacts coding and non-coding RNA profiles. PLoS ONE 2017, 12, e0179583. [Google Scholar] [CrossRef] [PubMed]
- Gálvez-Ontiveros, Y.; Moscoso-Ruiz, I.; Rodrigo, L.; Aguilera, M.; Rivas, A.; Zafra-Gómez, A. Presence of parabens and bisphenols in food commonly consumed in Spain. Foods 2021, 10, 92. [Google Scholar] [CrossRef]
- Zhang, Y.; Dong, T.; Hu, W.; Wang, X.; Xu, B.; Lin, Z.; Hofer, T.; Stefanoff, P.; Chen, Y.; Wang, X.; et al. Association between exposure to a mixture of Phenols, Pesticides, and Phthalates and Obesity: Comparison of three statistical models. Environ. Int. 2019, 123, 325–336. [Google Scholar] [CrossRef]
- Milošević, N.; Milanović, M.; Sudji, J.; Bosić Živanović, D.; Stojanoski, S.; Vuković, B.; Milić, N.; Medić Stojanoska, M. Could Phthalates exposure contribute to the development of metabolic syndrome and liver disease in humans? Environ. Sci. Pollut. Res. Int. 2020, 27, 772–784. [Google Scholar] [CrossRef] [PubMed]
- Roman, P.; Cardona, D.; Sempere, L.; Carvajal, F. Microbiota and Organophosphates. Neurotoxicology 2019, 75, 200–208. [Google Scholar] [CrossRef] [PubMed]
- Yuan, X.; Pan, Z.; Jin, C.; Ni, Y.; Fu, Z.; Jin, Y. Gut Microbiota: An underestimated and unintended recipient for Pesticide-induced toxicity. Chemosphere 2019, 227, 425–434. [Google Scholar] [CrossRef] [PubMed]
- Woldemariam, K.; Wan, Z.; Yu, Q.; Li, H.; Wei, X.; Liu, Y.; Wang, J.; Sun, B. Prebiotic, probiotic, antimicrobial, and functional food applications of Bacillus amyloliquefaciens. J. Agric. Food Chem. 2020, 68, 14709–14727. [Google Scholar] [CrossRef] [PubMed]
- Casado, V.; Martín, D.; Torres, C.; Reglero, G. Phospholipases in food industry: A Review. Methods Mol. Biol. 2012, 861, 495–523. [Google Scholar] [CrossRef] [PubMed]
- Moran, K.; de Lange, C.F.M.; Ferket, P.; Fellner, V.; Wilcock, P.; van Heugten, E. Enzyme supplementation to improve the nutritional value of fibrous feed ingredients in swine diets fed in dry or liquid form. J. Anim. Sci. 2016, 94, 1031–1040. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Gallego, C.; Pohl, S.; Salminen, S.; De Vos, W.M.; Kneifel, W. Akkermansia muciniphila: A novel functional microbe with probiotic properties. Benef. Microbes 2016, 7, 571–584. [Google Scholar] [CrossRef]
- Brodmann, T.; Endo, A.; Gueimonde, M.; Vinderola, G.; Kneifel, W.; de Vos, W.M.; Salminen, S.; Gómez-Gallego, C. Safety of novel microbes for human consumption: Practical examples of assessment in the European Union. Front. Microbiol. 2017, 8, 1725. [Google Scholar] [CrossRef] [PubMed]
- EFSA Panel on Biological Hazards (BIOHAZ); Koutsoumanis, K.; Allende, A.; Alvarez-Ordóñez, A.; Bolton, D.; Bover-Cid, S.; Chemaly, M.; Davies, R.; De Cesare, A.; Hilbert, F.; et al. Scientific opinion on the update of the list of QPS-recommended biological agents intentionally added to food or feed as notified to EFSA (2017–2019). EFSA J. 2020, 18, e05966. [Google Scholar] [CrossRef]
- EFSA Panel on Additives and Products or Substances used in Animal Feed (FEEDAP); Bampidis, V.; Azimonti, G.; de Bastos, M.L.; Christensen, H.; Dusemund, B.; Kouba, M.; Fašmon Durjava, M.; López-Alonso, M.; López Puente, S.; et al. Safety and efficacy of Bacillus subtilis PB6 (Bacillus velezensis ATCC PTA-6737) as a feed additive for chickens for fattening, chickens reared for laying, minor poultry species (except for laying purposes), ornamental, sporting and game birds. EFSA J. 2020, 18, e06280. [Google Scholar] [CrossRef]
- EFSA Panel on Additives and Products or Substances used in Animal Feed (FEEDAP); Rychen, G.; Aquilina, G.; Azimonti, G.; Bampidis, V.; de Bastos, M.L.; Bories, G.; Chesson, A.; Cocconcelli, P.S.; Flachowsky, G.; et al. Guidance on the characterisation of microorganisms used as feed additives or as production organisms. EFSA J. 2018, 16, e05206. [Google Scholar] [CrossRef]
- Fan, B.; Blom, J.; Klenk, H.-P.; Borriss, R. Bacillus amyloliquefaciens, Bacillus velezensis, and Bacillus siamensis Form an “Operational group B. amyloliquefaciens” within the B. subtilis species complex. Front. Microbiol. 2017, 8, 22. [Google Scholar] [CrossRef]
- Yamanaka, H.; Moriyoshi, K.; Ohmoto, T.; Ohe, T.; Sakai, K. Degradation of Bisphenol A by Bacillus pumilus isolated from Kimchi, a traditionally fermented food. Appl. Biochem. Biotechnol. 2007, 136, 39–51. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Zu, L.; Wong, P.-K.; Hui, X.; Lu, Y.; Xiong, J.; An, T. Biodegradation and detoxification of Bisphenol A with one newly-isolated strain Bacillus sp. GZB: Kinetics, mechanism and estrogenic transition. Bioresour. Technol. 2012, 114, 224–230. [Google Scholar] [CrossRef] [PubMed]
- Das, R.; Liang, Z.; Li, G.; Mai, B.; An, T. Genome sequence of a spore-laccase forming, BPA-degrading Bacillus sp. GZB isolated from an electronic-waste recycling site reveals insights into BPA degradation pathways. Arch. Microbiol. 2019, 201, 623–638. [Google Scholar] [CrossRef] [PubMed]
- Jandhyala, S.M.; Talukdar, R.; Subramanyam, C.; Vuyyuru, H.; Sasikala, M.; Nageshwar Reddy, D. Role of the normal gut microbiota. World J. Gastroenterol. 2015, 21, 8787–8803. [Google Scholar] [CrossRef] [PubMed]
- Indiani, C.; Rizzardi, K.F.; Castelo, P.M.; Ferraz, L.F.C.; Darrieux, M.; Parisotto, T.M. Childhood Obesity and Firmicutes/Bacteroidetes ratio in the gut microbiota: A systematic review. Child Obes. 2018, 14, 501–509. [Google Scholar] [CrossRef] [PubMed]
- Andra, S.S.; Austin, C.; Yang, J.; Patel, D.; Arora, M. Recent advances in simultaneous analysis of Bisphenol A and its conjugates in human matrices: Exposure biomarker perspectives. Sci. Total Environ. 2016, 572, 770–781. [Google Scholar] [CrossRef]
- Cao, G.T.; Dai, B.; Wang, K.L.; Yan, Y.; Xu, Y.L.; Wang, Y.X.; Yang, C.M. Bacillus licheniformis, a potential probiotic, inhibits Obesity by modulating colonic microflora in C57BL/6J mice model. J. Appl. Microbiol. 2019, 127, 880–888. [Google Scholar] [CrossRef]
- Wang, Y.; Wu, Y.; Wang, B.; Xu, H.; Mei, X.; Xu, X.; Zhang, X.; Ni, J.; Li, W. Bacillus amyloliquefaciens SC06 protects mice against High-Fat Diet-Induced Obesity and liver injury via regulating host metabolism and gut microbiota. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- López-Moreno, A.; Suárez, A.; Avanzi, C.; Monteoliva-Sánchez, M.; Aguilera, M. Probiotic strains and intervention total doses for modulating Obesity-related microbiota dysbiosis: A systematic review and meta-analysis. Nutrients 2020, 12, 1912. [Google Scholar] [CrossRef]
- Pariza, M.W.; Gillies, K.O.; Kraak-Ripple, S.F.; Leyer, G.; Smith, A.B. Determining the safety of microbial cultures for consumption by humans and animals. Regul. Toxicol. Pharmacol. 2015, 73, 164–171. [Google Scholar] [CrossRef]
- Thomson, N.; Summers, D.; Sivaniah, E. Synthesis, properties and uses of bacterial storage lipid granules as naturally occurring nanoparticles. Soft Matter 2010, 6, 4045–4057. [Google Scholar] [CrossRef]
- Zhang, K.; Zhou, C.; Zhou, Z.; Li, X.; Li, Z.; Zhang, M.; Zhang, X.; Zhang, C.; Wei, T.; Zhang, S.; et al. Identification of lipid droplets in gut microbiota. bioRxiv 2020. [Google Scholar] [CrossRef]
- Devaraj, K.; Aathika, S.; Periyasamy, K.; Manickam Periyaraman, P.; Palaniyandi, S.; Subramanian, S. Production of thermostable multiple enzymes from Bacillus amyloliquefaciens KUB29. Nat. Prod. Res. 2019, 33, 1674–1677. [Google Scholar] [CrossRef]
- Deb, P.; Talukdar, S.A.; Mohsina, K.; Sarker, P.K.; Sayem, S.A. Production and partial characterization of extracellular amylase enzyme from Bacillus amyloliquefaciens P-001. Springerplus 2013, 2, 154. [Google Scholar] [CrossRef] [PubMed]
- Latorre, J.D.; Hernandez-Velasco, X.; Wolfenden, R.E.; Vicente, J.L.; Wolfenden, A.D.; Menconi, A.; Bielke, L.R.; Hargis, B.M.; Tellez, G. Evaluation and selection of Bacillus species based on enzyme production, antimicrobial activity, and biofilm synthesis as direct-fed microbial candidates for poultry. Front. Vet. Sci. 2016, 3. [Google Scholar] [CrossRef] [PubMed]
- Ohore, O.E.; Zhang, S. Endocrine Disrupting Effects of Bisphenol A exposure and recent advances on its removal by water treatment systems: A Review. Sci. Afr. 2019, 5, e00135. [Google Scholar] [CrossRef]
- Sasaki, M.; Tsuchido, T.; Matsumura, Y. Molecular cloning and characterization of cytochrome P450 and ferredoxin genes involved in Bisphenol A degradation in Sphingomonas bisphenolicum strain AO1. J. Appl. Microbiol. 2008, 105, 1158–1169. [Google Scholar] [CrossRef]
- Nakamura, S.; Tezuka, Y.; Ushiyama, A.; Kawashima, C.; Kitagawara, Y.; Takahashi, K.; Ohta, S.; Mashino, T. Ipso Substitution of Bisphenol A catalyzed by microsomal cytochrome P450 and enhancement of estrogenic activity. Toxicol. Lett. 2011, 203, 92–95. [Google Scholar] [CrossRef] [PubMed]
- Lobos, J.H.; Leib, T.K.; Su, T.M. Biodegradation of Bisphenol A and other bisphenols by a gram-negative aerobic bacterium. Appl. Environ. Microbiol. 1992, 58, 1823–1831. [Google Scholar] [CrossRef]
- Sasaki, M.; Akahira, A.; Oshiman, K.; Tsuchido, T.; Matsumura, Y. Purification of Cytochrome P450 and ferredoxin, involved in Bisphenol A degradation, from Sphingomonas sp. strain AO1. Appl. Environ. Microbiol. 2005, 71, 8024–8030. [Google Scholar] [CrossRef]
- Menasria, T.; Aguilera, M.; Hocine, H.; Benammar, L.; Ayachi, A.; Si Bachir, A.; Dekak, A.; Monteoliva-Sánchez, M. Diversity and bioprospecting of extremely halophilic archaea isolated from Algerian arid and semi-arid wetland ecosystems for halophilic-active hydrolytic enzymes. Microbiol. Res. 2018, 207, 289–298. [Google Scholar] [CrossRef]
- Menasria, T.; Monteoliva-Sánchez, M.; Benammar, L.; Benhadj, M.; Ayachi, A.; Hacène, H.; Gonzalez-Paredes, A.; Aguilera, M. Culturable halophilic bacteria inhabiting algerian saline ecosystems: A source of promising features and potentialities. World J. Microbiol. Biotechnol. 2019, 35, 132. [Google Scholar] [CrossRef] [PubMed]
- EFSA Panel on additives and products or substances used in animal feed (FEEDAP) 2012; guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance (question no EFSA-Q-2011-01108, adopted: 23 May 2012 by European Food Safety Authority). EFSA J. 2012, 10, 10. [CrossRef]
- Humphries, R.M.; Ambler, J.; Mitchell, S.L.; Castanheira, M.; Dingle, T.; Hindler, J.A.; Koeth, L.; Sei, K. CLSI Methods development and standardization working group best practices for evaluation of antimicrobial susceptibility tests. J. Clin. Microbiol. 2018, 56. [Google Scholar] [CrossRef] [PubMed]
- García-Córcoles, M.T.; Cipa, M.; Rodríguez-Gómez, R.; Rivas, A.; Olea-Serrano, F.; Vílchez, J.L.; Zafra-Gómez, A. Determination of bisphenols with estrogenic activity in plastic packaged baby food samples using solid-liquid extraction and clean-up with dispersive sorbents followed by gas chromatography tandem mass spectrometry analysis. Talanta 2018, 178, 441–448. [Google Scholar] [CrossRef]
- Wick, R.R.; Judd, L.M.; Gorrie, C.L.; Holt, K.E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 2017, 13. [Google Scholar] [CrossRef] [PubMed]
- Jolley, K.A.; Bliss, C.M.; Bennett, J.S.; Bratcher, H.B.; Brehony, C.; Colles, F.M.; Wimalarathna, H.; Harrison, O.B.; Sheppard, S.K.; Cody, A.J.; et al. Ribosomal multilocus sequence typing: Universal characterization of bacteria from domain to strain. Microbiology (Reading) 2012, 158 (Pt 4), 1005–1015. [Google Scholar] [CrossRef]
- Yoon, S.-H.; Ha, S.-M.; Lim, J.; Kwon, S.; Chun, J. A Large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 2017, 110, 1281–1286. [Google Scholar] [CrossRef]
- Meier-Kolthoff, J.P.; Auch, A.F.; Klenk, H.-P.; Göker, M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 2013, 14, 60. [Google Scholar] [CrossRef]

| Enzyme Test | Microorganisms | |||||
|---|---|---|---|---|---|---|
| Bacillus sp. AM1 | B. amyloliquefaciens (DSM7) | B. velezensis (CR-502) | B. siamensis (KCTC13613) | B.cereus (B7) | B.mycoides (B12) | |
| Starch | + | + | + | ++ | ++ | ++ |
| Carboxymethylcelulose | − | − | − | − | − | − |
| Gelatine | − | − | − | − | − | − |
| Inuline | + | − | − | − | + | + |
| Tween 80 | − | + | + | − | − | − |
| DNase | ++ | + | + | − | − | − |
| Antibiotic | Concentration (μg) | Resistance (R) Sensitive (S) | Zone inhibition (mm) |
|---|---|---|---|
| Ampicillin | 10 | R | - |
| Amikacin | 30 | S | 14 |
| Ceftazidime | 30 | R | - |
| Ciprofloxacin | 5 | S | 40 |
| Chloramphenicol | 30 | S | 42 |
| Clindamycin | 2 | S | 22 |
| Erythromycin | 15 | S | 25 |
| Gentamicin | 10 | S | 22 |
| Imipenem | 10 | S | 50 |
| Penicillin | 10 | R | - |
| Rifampicin | 30 | S | 29 |
| Vancomycin | 30 | S | 22 |
| Kanamycin | 30 | S | 19 |
| Streptomycin | 10 | R | - |
| Tetracycline | 30 | S | 17 |
| Co-Trimoxazole | 25 | S | 39 |
| Amoxicillin/ Clavulanic acid | 20/10 | R | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
López-Moreno, A.; Torres-Sánchez, A.; Acuña, I.; Suárez, A.; Aguilera, M. Representative Bacillus sp. AM1 from Gut Microbiota Harbor Versatile Molecular Pathways for Bisphenol A Biodegradation. Int. J. Mol. Sci. 2021, 22, 4952. https://doi.org/10.3390/ijms22094952
López-Moreno A, Torres-Sánchez A, Acuña I, Suárez A, Aguilera M. Representative Bacillus sp. AM1 from Gut Microbiota Harbor Versatile Molecular Pathways for Bisphenol A Biodegradation. International Journal of Molecular Sciences. 2021; 22(9):4952. https://doi.org/10.3390/ijms22094952
Chicago/Turabian StyleLópez-Moreno, Ana, Alfonso Torres-Sánchez, Inmaculada Acuña, Antonio Suárez, and Margarita Aguilera. 2021. "Representative Bacillus sp. AM1 from Gut Microbiota Harbor Versatile Molecular Pathways for Bisphenol A Biodegradation" International Journal of Molecular Sciences 22, no. 9: 4952. https://doi.org/10.3390/ijms22094952
APA StyleLópez-Moreno, A., Torres-Sánchez, A., Acuña, I., Suárez, A., & Aguilera, M. (2021). Representative Bacillus sp. AM1 from Gut Microbiota Harbor Versatile Molecular Pathways for Bisphenol A Biodegradation. International Journal of Molecular Sciences, 22(9), 4952. https://doi.org/10.3390/ijms22094952









