Nod-Like Receptors in Host Defence and Disease at the Epidermal Barrier
Abstract
1. Introduction
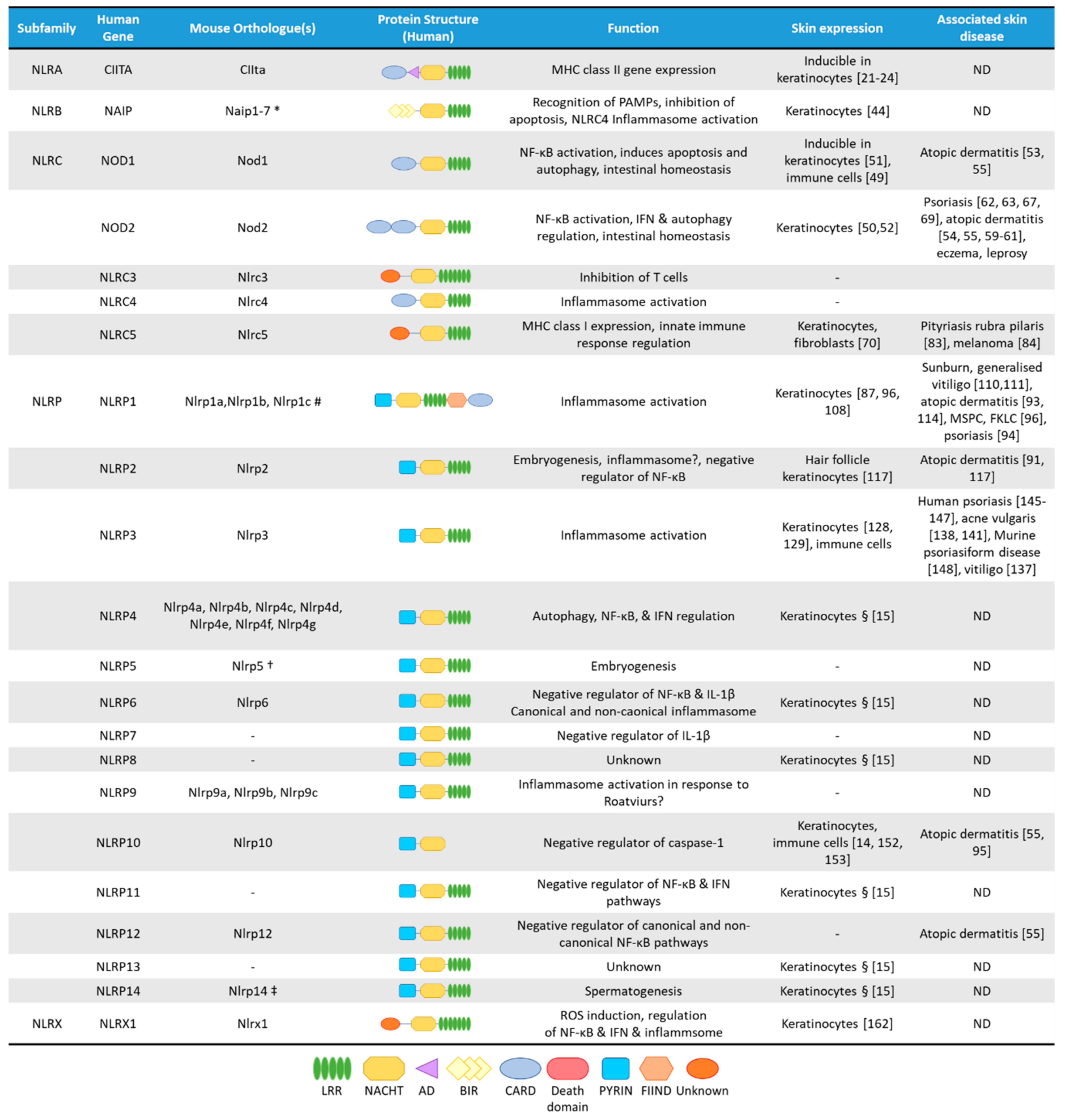
2. The NLRA Subfamily
3. The NLRB Subfamily
4. The NLRC Subfamily
4.1. NOD1 and NOD2
Skin Diseases Associated with NOD1 and NOD2 Functions
4.2. NLRC5
Skin Diseases Associated with NLRC5 Functions
5. The NLRP Subfamily
5.1. NLRP1
Skin Diseases Associated with Genetic Variants in NLRP1
5.2. NLRP2
Skin Diseases Associated with NLRP2 Functions
5.3. NLRP3
5.3.1. The Role of NLRP3 in Skin Diseases
5.3.2. Inflammasome Activation and Epidermal Differentiation Are Interconnected
5.4. NLRP10
6. The NLRX Subfamily
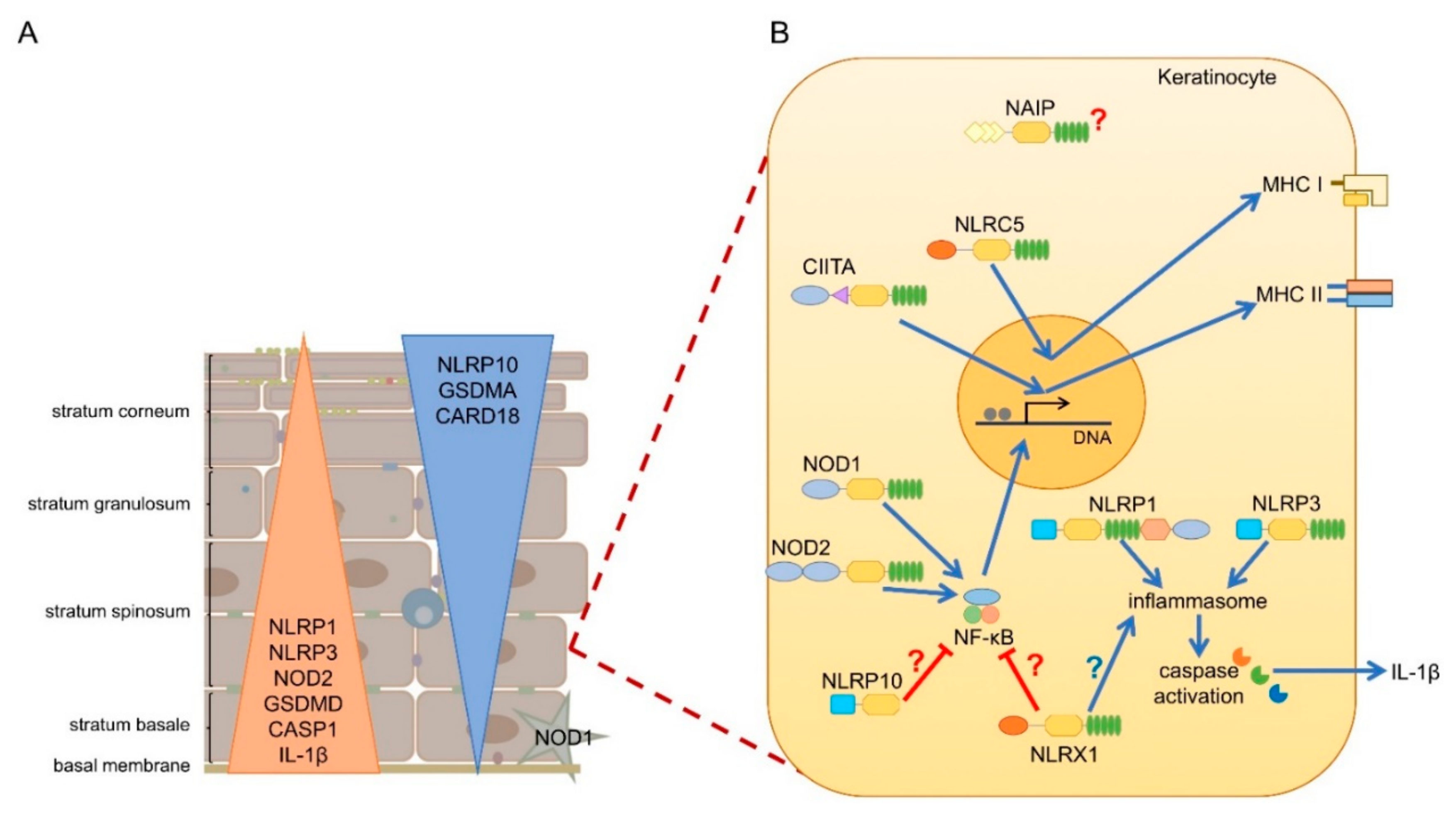
7. NLRs in Human and Mouse Skin
| Human Gene | Murine Gene | Function in Human Keratincoytes/Skin | Function in Murine Keratinocytes/Skin |
|---|---|---|---|
| CIITA | CIIta | Inducible in keratinocytes [21,22,24] Regulates MHC-II expression and presentation of intact proteins to T-cells [22,27,28], | Inducible in keratinocytes [23,30] |
| NOD2 | Nod2 | Gene variants associated with atopic dermatitis [53,54,55] Deficiency promotes Staphylococcus aureus colonisation [61]. Upregulated in psoriasis [67] | Nod2–/– are unable to clear Staphylococcus aureus infection [59,60] Nod2–/– mice are susceptible to imiquimod-induced psoriasiform disease [68] |
| NLRP1 | Nlrp1a, Nlrp1b, Nlrp1c # | Expressed in keratinocytes [87] dsRNA and UVB induced activation in keratinocytes [87,90,107] | Not expressed in keratinocytes [87] UVB induced activation in skin-homing immune cells [87] |
| NLRP3 | Nlrp3 | Expressed and activated in keratinocytes [128,129] Activated in sebocytes by Cutibacterium acnes [138] Activated in keratinocytes by Staphylococcus aureus [144] Upregulated in psoriasis [145] | Expressed mainly in skin-homing immune cells [132,133] Role in wound healing [132,133] Activated in keratinocytes by Staphylococcus aureus [15] Upregulated in psoriasiform disease [146,148,149] |
| NLRP10 | Nlrp10 | Expressed in differentiated keratinocytes [14,152] Gene variants associated with atopic dermatitis [55,95] Regulates inflammasome activation [152,154] | Expressed in differentiated keratinocytes [14,153] Plays a role in contact hypersensitivity [157] Nlrp10–/– mice exhibit normal inflammasome activation [155,156] |
8. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chandler, C.E.; Harberts, E.M.; Ernst, R.K. Pathogen Sensing: Toll-Like Receptors and NODs (Innate Immunity). In Encyclopedia of Microbiology (Fourth Edition); Elsevier: Amsterdam, The Netherlands, 2019; pp. 443–456. [Google Scholar]
- Zhang, Q.; Zmasek, C.M.; Godzik, A. Domain architecture evolution of pattern-recognition receptors. Immunogenetics 2010, 62, 263–272. [Google Scholar] [CrossRef]
- Jann, O.C.; King, A.; Lopez Corrales, N.; Anderson, S.I.; Jensen, K.; Ait-ali, T.; Tang, H.; Wu, C.; Cockett, N.E.; Archibald, A.L.; et al. Comparative genomics of Toll-like receptor signalling in five species. BMC Genom. 2009, 10, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Kuo, I.-H.; Yoshida, T.; De Benedetto, A.; Beck, L.A. The cutaneous innate immune response in patients with atopic dermatitis. J. Allergy Clin. Immunol. 2013, 131, 266–278. [Google Scholar] [CrossRef]
- Sun, L.; Liu, W.; Zhang, L.J. The Role of Toll-Like Receptors in Skin Host Defense, Psoriasis, and Atopic Dermatitis. J. Immunol. Res. 2019, 1824624. [Google Scholar] [CrossRef]
- Miller, L.S. Toll-like receptors in skin. Adv. Dermatol. 2008, 24, 71–87. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.; Zhou, F. Inflammasomes in Common Immune-Related Skin Diseases. Front. Immunol. 2020, 11, 1–15. [Google Scholar] [CrossRef]
- Shaw, M.H.; Reimer, T.; Kim, Y.-G.; Nuñez, G. NOD-like receptors (NLRs): Bona fide intracellular microbial sensors. Curr. Opin. Immunol. 2008, 20, 377–382. [Google Scholar] [CrossRef] [PubMed]
- Ting, J.P.Y.; Lovering, R.C.; Alnemri, E.S.P.D.; Bertin, J.; Boss, J.M.; Davis, B.; Flavell, R.A.; Girardin, S.E.; Godzik, A.; Harton, J.A.; et al. The NLR gene family: An official nomenclature. Immunity 2008, 28, 285–287. [Google Scholar] [CrossRef] [PubMed]
- Bentham, A.; Burdett, H.; Anderson, P.A.; Williams, S.J.; Kobe, B. Animal NLRs provide structural insights into plant NLR function. Ann. Bot. 2016, 119, 698–702. [Google Scholar] [CrossRef] [PubMed]
- Tenthorey, J.L.; Kofoed, E.M.; Daugherty, M.D.; Malik, H.S.; Vance, R.E. Molecular Basis for Specific Recognition of Bacterial Ligands by NAIP/NLRC4 Inflammasomes. Mol. Cell 2014, 54, 17–29. [Google Scholar] [CrossRef]
- Hu, Z.; Yan, C.; Liu, P.; Huang, Z.; Ma, R.; Zhang, C.; Wang, R.; Zhang, Y.; Martinon, F.; Miao, D.; et al. Crystal Structure of NLRC4 Reveals Its Autoinhibition Mechanism. Science 2013, 341, 172–175. [Google Scholar] [CrossRef] [PubMed]
- Meunier, E.; Broz, P. Evolutionary Convergence and Divergence in NLR Function and Structure. Trends Immunol. 2017, 38, 744–757. [Google Scholar] [CrossRef] [PubMed]
- Lachner, J.; Mlitz, V.; Tschachler, E.; Eckhart, L. Epidermal cornification is preceded by the expression of a keratinocyte-specific set of pyroptosis-related genes. Sci. Rep. 2017, 7, 17446. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, H.; Gaide, O.; Pétrilli, V.; Martinon, F.; Contassot, E.; Roques, S.; Kummer, J.A.; Tschopp, J.; French, L.E. Activation of the IL-1beta-processing inflammasome is involved in contact hypersensitivity. J. Investig. Dermatol. 2007, 127, 1956–1963. [Google Scholar] [CrossRef]
- León Machado, J.A.; Steimle, V. The MHC class II transactivator CIITA: Not (quite) the odd-one-out anymore among NLR proteins. Int. J. Mol. Sci. 2021, 22, 1074. [Google Scholar] [CrossRef] [PubMed]
- Steimle, V.; Otten, L.A.; Zufferey, M.; Mach, B. Complementation cloning of an MHC class II transactivator mutated in hereditary MHC class II deficiency (or bare lymphocyte syndrome). Cell 1993, 75, 135–146. [Google Scholar] [CrossRef]
- Chang, C.-H.; Guerder, S.; Hong, S.-C.; van Ewijk, W.; Flavell, R.A. Mice Lacking the MHC Class II Transactivator (CIITA) Show Tissue-Specific Impairment of MHC Class II Expression. Immunity 1996, 4, 167–178. [Google Scholar] [CrossRef]
- Reith, W.; LeibundGut-Landmann, S.; Waldburger, J.-M. Regulation of MHC class II gene expression by the class II transactivator. Nat. Rev. Immunol. 2005, 5, 793–806. [Google Scholar] [CrossRef] [PubMed]
- Choi, N.M.; Majumder, P.; Boss, J.M. Regulation of major histocompatibility complex class II genes. Curr. Opin. Immunol. 2011, 23, 81–87. [Google Scholar] [CrossRef]
- Albanesi Cavani, C.A.; Girolomoni, G. Interferon-γ-Stimulated Human Keratinocytes Express the Genes Necessary for the Production of Peptide-Loaded MHC Class II Molecules. J. Investig. Dermatol. 1998, 110, 138–142. [Google Scholar] [CrossRef]
- Takagi, A.; Nishiyama, C.; Kanada, S.; Niwa, Y.; Fukuyama, K.; Ikeda, S.; Okumura, K.; Ogawa, H. Prolonged MHC class II expression and CIITA transcription in human keratinocytes. Biochem. Biophys. Res. Commun. 2006, 347, 388–393. [Google Scholar] [CrossRef]
- Tamoutounour, S.; Han, S.-J.; Deckers, J.; Constantinides, M.G.; Hurabielle, C.; Harrison, O.J.; Bouladoux, N.; Linehan, J.L.; Link, V.M.; Vujkovic-Cvijin, I.; et al. Keratinocyte-intrinsic MHCII expression controls microbiota-induced Th1 cell responses. Proc. Natl. Acad. Sci. USA 2019, 116, 23643–23652. [Google Scholar] [CrossRef]
- Czernielewski, J.M.; Bagot, M. Class II MHC antigen expression by human keratinocytes results from lympho-epidermal interactions and gamma-interferon production. Clin. Exp. Immunol. 1986, 66, 295–302. [Google Scholar]
- Aubock, J.; Romani, N.; Grubauer, G.; Fritsch, P. HLA-DR expression on keratinocytes is a common feature of diseased skin. Br. J. Dermatol. 1986, 114, 465–472. [Google Scholar] [CrossRef] [PubMed]
- Krueger, J.G.; Krane, J.F.; Carter, D.M.; Gottlieb, A.B. Role of Growth Factors, Cytokines, and Their Receptors in the Pathogenesis of Psoriasis. J. Investig. Dermatol. 1990, 94, s135–s140. [Google Scholar] [CrossRef] [PubMed]
- Wittmann, M.; Purwar, R.; Hartmann, C.; Gutzmer, R.; Werfel, T. Human Keratinocytes Respond to Interleukin-18: Implication for the Course of Chronic Inflammatory Skin Diseases. J. Investig. Dermatol. 2005, 124, 1225–1233. [Google Scholar] [CrossRef] [PubMed]
- Mutis, T.; Bueger, M.; Bakker, A.; Ottenhoff, T.H.M. HLA Class II+ Human Keratinocytes present Mycobacterium leprae Antigens to CD4+ Thl-Like Cells. Scand. J. Immunol. 1993, 37, 43–51. [Google Scholar] [CrossRef] [PubMed]
- Nickoloff, B.J.; Basham, T.Y.; Merigan, T.C.; Torseth, J.W.; Morhenn, V.B. Human Keratinocyte-Lymphocyte Reactions In Vitro. J. Investig. Dermatol. 1986, 87, 11–18. [Google Scholar] [CrossRef]
- Gaspari, A.A.; Katz, S.I. Induction and functional characterization of class II MHC (Ia) antigens on murine keratinocytes. J. Immunol. 1988, 140, 2956–2963. [Google Scholar]
- Niederwieser, D.; Auböck, J.; Troppmair, J.; Herold, M.; Schuler, G.; Boeck, G.; Lotz, J.; Fritsch, P.; Huber, C. IFN-mediated induction of MHC antigen expression on human keratinocytes and its influence on in vitro alloimmune responses. J. Immunol. 1988, 140, 2556–2564. [Google Scholar] [PubMed]
- Ortiz-Sanchez, E.; Chávez-Olmos, P.; Pina-Sanchez, P.; Salcedo, M.; Garrigo, E. Expression of the costimulatory molecule CD86, but not CD80, in keratinocytes of normal cervical epithelium and human papillomavirus-16 positive low squamous intraepithelial lesions. Int. J. Gynecol. Cancer 2007, 17, 571–580. [Google Scholar] [CrossRef]
- Nickoloff, B.J.; Mitra, R.S.; Lee, K.; Turka, L.A.; Green, J.; Thompson, C.; Shimizu, Y. Discordant expression of CD28 ligands, BB-1, and B7 on keratinocytes in vitro and psoriatic cells in vivo. Am. J. Pathol. 1993, 142, 1029–1040. [Google Scholar]
- Gaspari, A.A.; Ferbel, B.; Chen, Z.; Razvi, F.; Polakowska, R. Accessory and Alloantigen-Presenting Cell Functions of A431 Keratinocytes That Stably Express the B7 Antigen. Cell. Immunol. 1993, 149, 291–302. [Google Scholar] [CrossRef]
- Nickoloff, B.J.; Turka, L.A. Immunological functions of non-professional antigen-presenting cells: New insights from studies of T-cell interactions with keratinocytes. Immunol. Today 1994, 15, 464–469. [Google Scholar] [CrossRef]
- Kofoed, E.M.; Vance, R.E. Innate immune recognition of bacterial ligands by NAIPs determines inflammasome specificity. Nature 2011, 477, 592–595. [Google Scholar] [CrossRef]
- Maier, J.K.X.; Lahoua, Z.; Gendron, N.H.; Fetni, R.; Johnston, A.; Davoodi, J.; Rasper, D.; Roy, S.; Slack, R.S.; Nicholson, D.W.; et al. The Neuronal Apoptosis Inhibitory Protein Is a Direct Inhibitor of Caspases 3 and 7. J. Neurosci. 2002, 22, 2035–2043. [Google Scholar] [CrossRef] [PubMed]
- Davoodi, J.; Ghahremani, M.-H.; Es-haghi, A.; Mohammad-gholi, A.; MacKenzie, A. Neuronal apoptosis inhibitory protein, NAIP, is an inhibitor of procaspase-9. Int. J. Biochem. Cell Biol. 2010, 42, 958–964. [Google Scholar] [CrossRef]
- Sanna, M.G.; da Correia, J.S.; Ducrey, O.; Lee, J.; Nomoto, K.; Schrantz, N.; Deveraux, Q.L.; Ulevitch, R.J. IAP Suppression of Apoptosis Involves Distinct Mechanisms: The TAK1/JNK1 Signaling Cascade and Caspase Inhibition. Mol. Cell. Biol. 2002, 22, 1754–1766. [Google Scholar] [CrossRef] [PubMed]
- Vance, R.E. The NAIP/NLRC4 inflammasomes. Curr. Opin. Immunol. 2015, 32, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zhao, Y.; Shi, J.; Shao, F. Human NAIP and mouse NAIP1 recognize bacterial type III secretion needle protein for inflammasome activation. Proc. Natl. Acad. Sci. USA 2013, 110, 14408–14413. [Google Scholar] [CrossRef]
- Mariathasan, S.; Newton, K.; Monack, D.M.; Vucic, D.; French, D.M.; Lee, W.P.; Roose-Girma, M.; Erickson, S.; Dixit, V.M. Differential activation of the inflammasome by caspase-1 adaptors ASC and Ipaf. Nature 2004, 430, 213–218. [Google Scholar] [CrossRef]
- Zamboni, D.S.; Kobayashi, K.S.; Kohlsdorf, T.; Ogura, Y.; Long, E.M.; Vance, R.E.; Kuida, K.; Mariathasan, S.; Dixit, V.M.; Flavell, R.A.; et al. The Birc1e cytosolic pattern-recognition receptor contributes to the detection and control of Legionella pneumophila infection. Nat. Immunol. 2006, 7, 318–325. [Google Scholar] [CrossRef]
- Human Protein Atlas NAIP. Available online: https://www.proteinatlas.org/ENSG00000249437-NAIP (accessed on 19 February 2021).
- Velloso, F.J.; Trombetta-Lima, M.; Anschau, V.; Sogayar, M.C.; Correa, R.G. NOD-like receptors: Major players (and targets) in the interface between innate immunity and cancer. Biosci. Rep. 2019, 39. [Google Scholar] [CrossRef]
- Human Protein Atlas. Available online: http://www.proteinatlas.org (accessed on 31 January 2021).
- Correa, R.G.; Milutinovic, S.; Reed, J.C. Roles of NOD1 (NLRC1) and NOD2 (NLRC2) in innate immunity and inflammatory diseases. Biosci. Rep. 2012, 32, 597–608. [Google Scholar] [CrossRef] [PubMed]
- Travassos, L.H.; Carneiro, L.A.M.; Girardin, S.; Philpott, D.J. Nod proteins link bacterial sensing and autophagy. Autophagy 2010, 6, 409–411. [Google Scholar] [CrossRef]
- Human Protein Atlas NOD1. Available online: https://www.proteinatlas.org/ENSG00000106100-NOD1/celltype (accessed on 19 February 2021).
- Human Protein Atlas NOD2. Available online: https://www.proteinatlas.org/ENSG00000167207-NOD2/celltype (accessed on 19 February 2021).
- Harder, J.; Núñez, G. Functional Expression of the Intracellular Pattern Recognition Receptor NOD1 in Human Keratinocytes. J. Investig. Dermatol. 2009, 129, 1299–1302. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Voss, E.; Wehkamp, J.; Wehkamp, K.; Stange, E.F.; Schröder, J.M.; Harder, J. NOD2/CARD15 mediates induction of the antimicrobial peptide human beta-defensin-2. J. Biol. Chem. 2006, 281, 2005–2011. [Google Scholar] [CrossRef]
- Weidinger, S.; Klopp, N.; Rummler, L.; Wagenpfeil, S.; Novak, N.; Baurecht, H.-J.; Groer, W.; Darsow, U.; Heinrich, J.; Gauger, A. Association of NOD1 polymorphisms with atopic eczema and related phenotypes. J. Allergy Clin. Immunol. 2005, 116, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Kabesch, M.; Peters, W.; Carr, D.; Leupold, W.; Weiland, S.K.; von Mutius, E. Association between polymorphisms in caspase recruitment domain containing protein 15 and allergy in two German populations. J. Allergy Clin. Immunol. 2003, 111, 813–817. [Google Scholar] [CrossRef]
- Macaluso, F.; Nothnagel, M.; Parwez, Q.; Petrasch-Parwez, E.; Bechara, F.G.; Epplen, J.T.; Hoffjan, S. Polymorphisms in NACHT-LRR (NLR) genes in atopic dermatitis. Exp. Dermatol. 2007, 16, 692–698. [Google Scholar] [CrossRef] [PubMed]
- Bylund, S.; Von Kobyletzki, L.B.; Svalstedt, M.; Svensson, Å. Prevalence and incidence of atopic dermatitis: A systematic review. Acta Derm. Venereol. 2020, 100, 320–329. [Google Scholar] [CrossRef]
- Tsang, M.S.M.; Hou, T.; Chan, B.C.L.; Wong, C.K. Immunological roles of NLR in allergic diseases and its underlying mechanisms. Int. J. Mol. Sci. 2021, 22, 1507. [Google Scholar] [CrossRef]
- De Benedetto, A.; Kubo, A.; Beck, L.A. Skin Barrier Disruption: A Requirement for Allergen Sensitization? J. Investig. Dermatol. 2012, 132, 949–963. [Google Scholar] [CrossRef]
- Deshmukh, H.S.; Hamburger, J.B.; Ahn, S.H.; McCafferty, D.G.; Yang, S.R.; Fowler, V.G. Critical Role of NOD2 in Regulating the Immune Response to Staphylococcus aureus. Infect. Immun. 2009, 77, 1376–1382. [Google Scholar] [CrossRef]
- Hruz, P.; Zinkernagel, A.S.; Jenikova, G.; Botwin, G.J.; Hugot, J.-P.; Karin, M.; Nizet, V.; Eckmann, L. NOD2 contributes to cutaneous defense against Staphylococcus aureus through -toxin-dependent innate immune activation. Proc. Natl. Acad. Sci. USA 2009, 106, 12873–12878. [Google Scholar] [CrossRef] [PubMed]
- Roth, S.A.; Simanski, M.; Rademacher, F.; Schröder, L.; Harder, J. The Pattern Recognition Receptor NOD2 Mediates Staphylococcus aureus –Induced IL-17C Expression in Keratinocytes. J. Investig. Dermatol. 2014, 134, 374–380. [Google Scholar] [CrossRef]
- Rahman, P.; Bartlett, S.; Siannis, F.; Pellett, F.J.; Farewell, V.T.; Peddle, L.; Schentag, C.T.; Alderdice, C.A.; Hamilton, S.; Khraishi, M.; et al. CARD15: A Pleiotropic Autoimmune Gene That Confers Susceptibility to Psoriatic Arthritis. Am. J. Hum. Genet. 2003, 73, 677–681. [Google Scholar] [CrossRef]
- Zhu, K.; Yin, X.; Tang, X.; Zhang, F.; Yang, S.; Zhang, X. Meta-analysis of NOD2/CARD15 polymorphisms with psoriasis and psoriatic arthritis. Rheumatol. Int. 2012, 32, 1893–1900. [Google Scholar] [CrossRef]
- Christophers, E.; Metzler, G.; Röcken, M. Bimodal immune activation in psoriasis. Br. J. Dermatol. 2014, 170, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Rendon, A.; Schäkel, K. Psoriasis Pathogenesis and Treatment. Int. J. Mol. Sci. 2019, 20, 1475. [Google Scholar] [CrossRef] [PubMed]
- Bozó, R.; Flink, L.B.; Belső, N.; Gubán, B.; Széll, M.; Kemény, L.; Bata-Csörgő, Z. Could basement membrane alterations, resembling micro-wounds at the dermo-epidermal junction in psoriatic non-lesional skin, make the skin susceptible to lesion formation? Exp. Dermatol. 2021, 1–8. [Google Scholar] [CrossRef]
- Tervaniemi, M.; Katayama, S.; Skoog, T.; Siitonen, H.; Vuola, J.; Nuutila, K.; Sormunen, R.; Johnsson, A.; Linnarsson, S.; Suomela, S.; et al. NOD-like receptor signaling and inflammasome-related pathways are highlighted in psoriatic epidermis. Sci. Rep. 2016, 6, 22745. [Google Scholar] [CrossRef]
- Hedl, M.; Li, J.; Cho, J.H.; Abraham, C. Chronic stimulation of Nod2 mediates tolerance to bacterial products. Proc. Natl. Acad. Sci. USA 2007, 104, 19440–19445. [Google Scholar] [CrossRef]
- Guryanova, S.; Udzhukhu, V.; Kubylinsky, A. Pathogenetic Therapy of Psoriasis by Muramyl Peptide. Front. Immunol. 2019, 10. [Google Scholar] [CrossRef]
- Human Protein Atlas NLRC5. Available online: https://www.proteinatlas.org/ENSG00000140853-NLRC5/celltype (accessed on 19 February 2021).
- Mótyán, J.; Bagossi, P.; Benkő, S.; Tőzsér, J. A molecular model of the full-length human NOD-like receptor family CARD domain containing 5 (NLRC5) protein. BMC Bioinform. 2013, 14, 275. [Google Scholar] [CrossRef]
- Downs, I.; Vijayan, S.; Sidiq, T.; Kobayashi, K.S. CITA/NLRC5: A critical transcriptional regulator of MHC class I gene expression. BioFactors 2016, 42, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Neerincx, A.; Castro, W.; Guarda, G.; Kufer, T.A. NLRC5, at the Heart of Antigen Presentation. Front. Immunol. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, K.S.; van den Elsen, P.J. NLRC5: A key regulator of MHC class I-dependent immune responses. Nat. Rev. Immunol. 2012, 12, 813–820. [Google Scholar] [CrossRef] [PubMed]
- Neerincx, A.; Jakobshagen, K.; Utermöhlen, O.; Büning, H.; Steimle, V.; Kufer, T.A. The N-Terminal Domain of NLRC5 Confers Transcriptional Activity for MHC Class I and II Gene Expression. J. Immunol. 2014, 193, 3090–3100. [Google Scholar] [CrossRef]
- Benkő, S.; Kovács, E.G.; Hezel, F.; Kufer, T.A. NLRC5 Functions beyond MHC I Regulation—What Do We Know So Far? Front. Immunol. 2017, 8. [Google Scholar] [CrossRef]
- Cui, J.; Zhu, L.; Xia, X.; Wang, H.Y.; Legras, X.; Hong, J.; Ji, J.; Shen, P.; Zheng, S.; Chen, Z.J.; et al. NLRC5 Negatively Regulates the NF-κB and Type I Interferon Signaling Pathways. Cell 2010, 141, 483–496. [Google Scholar] [CrossRef] [PubMed]
- Benko, S.; Magalhaes, J.G.; Philpott, D.J.; Girardin, S.E. NLRC5 Limits the Activation of Inflammatory Pathways. J. Immunol. 2010, 185, 1681–1691. [Google Scholar] [CrossRef]
- Kuenzel, S.; Till, A.; Winkler, M.; Häsler, R.; Lipinski, S.; Jung, S.; Grötzinger, J.; Fickenscher, H.; Schreiber, S.; Rosenstiel, P. The Nucleotide-Binding Oligomerization Domain-Like Receptor NLRC5 Is Involved in IFN-Dependent Antiviral Immune Responses. J. Immunol. 2010, 184, 1990–2000. [Google Scholar] [CrossRef]
- Neerincx, A.; Lautz, K.; Menning, M.; Kremmer, E.; Zigrino, P.; Hösel, M.; Büning, H.; Schwarzenbacher, R.; Kufer, T.A. A Role for the Human Nucleotide-binding Domain, Leucine-rich Repeat-containing Family Member NLRC5 in Antiviral Responses*. J. Biol. Chem. 2010, 285, 26223–26232. [Google Scholar] [CrossRef] [PubMed]
- Ranjan, P.; Singh, N.; Kumar, A.; Neerincx, A.; Kremmer, E.; Cao, W.; Davis, W.G.; Katz, J.M.; Gangappa, S.; Lin, R.; et al. NLRC5 interacts with RIG-I to induce a robust antiviral response against influenza virus infection. Eur. J. Immunol. 2015, 45, 758–772. [Google Scholar] [CrossRef]
- Davis, B.K.; Roberts, R.A.; Huang, M.T.; Willingham, S.B.; Conti, B.J.; Brickey, W.J.; Barker, B.R.; Kwan, M.; Taxman, D.J.; Accavitti-Loper, M.-A.; et al. Cutting Edge: NLRC5-Dependent Activation of the Inflammasome. J. Immunol. 2011, 186, 1333–1337. [Google Scholar] [CrossRef]
- Spoerri, I.; Herms, S.; Eytan, O.; Sair, O.; Heinimann, K.; Sprecher, E.; Itin, P.H.; Burger, B. Immune-regulatory genes as possible modifiers of familial pityriasis rubra pilaris—Lessons from a family with PRP and psoriasis. J. Eur. Acad. Dermatol. Venereol. 2018. [Google Scholar] [CrossRef]
- Kim, H.; Kim, H.; Feng, Y.; Li, Y.; Tamiya, H.; Tocci, S.; Ronai, Z.A. PRMT5 control of cGAS/STING and NLRC5 pathways defines melanoma response to antitumor immunity. Sci. Transl. Med. 2020, 12, eaaz5683. [Google Scholar] [CrossRef]
- Ma, H.-L.; Zhao, X.-F.; Chen, G.-Z.; Fang, R.-H.; Zhang, F.-R. Silencing NLRC5 inhibits extracellular matrix expression in keloid fibroblasts via inhibition of transforming growth factor-β1/Smad signaling pathway. Biomed. Pharmacother. 2016, 83, 1016–1021. [Google Scholar] [CrossRef] [PubMed]
- Corridoni, D.; Arseneau, K.O.; Cifone, M.G.; Cominelli, F. The Dual Role of Nod-Like Receptors in Mucosal Innate Immunity and Chronic Intestinal Inflammation. Front. Immunol. 2014, 5, 317. [Google Scholar] [CrossRef]
- Sand, J.; Haertel, E.; Biedermann, T.; Contassot, E.; Reichmann, E.; French, L.E.; Werner, S.; Beer, H.-D. Expression of inflammasome proteins and inflammasome activation occurs in human, but not in murine keratinocytes. Cell Death Dis. 2018, 9, 24. [Google Scholar] [CrossRef]
- Niebuhr, M.; Baumert, K.; Heratizadeh, A.; Satzger, I.; Werfel, T. Impaired NLRP3 inflammasome expression and function in atopic dermatitis due to Th2 milieu. Allergy 2014. [Google Scholar] [CrossRef]
- Schroder, K.; Tschopp, J. The inflammasomes. Cell 2010, 140, 821–832. [Google Scholar] [CrossRef] [PubMed]
- Feldmeyer, L.; Keller, M.; Niklaus, G.; Hohl, D.; Werner, S.; Beer, H.-D. The inflammasome mediates UVB-induced activation and secretion of interleukin-1beta by keratinocytes. Curr. Biol. 2007, 17, 1140–1145. [Google Scholar] [CrossRef]
- Thürmann, L.; Grützmann, K.; Klös, M.; Bieg, M.; Winter, M.; Polte, T.; Bauer, T.; Schick, M.; Bewerunge-Hudler, M.; Röder, S.; et al. Early-onset childhood atopic dermatitis is related to NLRP2 repression. J. Allergy Clin. Immunol. 2018, 141, 1482–1485.e16. [Google Scholar] [CrossRef] [PubMed]
- Levandowski, C.B.; Mailloux, C.M.; Ferrara, T.M.; Gowan, K.; Ben, S.; Jin, Y.; McFann, K.K.; Holland, P.J.; Fain, P.R.; Dinarello, C.A.; et al. NLRP1 haplotypes associated with vitiligo and autoimmunity increase interleukin-1 processing via the NLRP1 inflammasome. Proc. Natl. Acad. Sci. USA 2013, 110, 2952–2956. [Google Scholar] [CrossRef]
- Bivik, C.; Verma, D.; Winge, M.C.; Lieden, A.; Bradley, M.; Rosdahl, I.; Söderkvist, P. Genetic Variation in the Inflammasome and Atopic Dermatitis Susceptibility. J. Investig. Dermatol. 2013, 133, 2486–2489. [Google Scholar] [CrossRef]
- Ekman, A.-K.; Verma, D.; Fredrikson, M.; Bivik, C.; Enerbäck, C. Genetic variations of NLRP1: Susceptibility in psoriasis. Br. J. Dermatol. 2014, 171, 1517–1520. [Google Scholar] [CrossRef] [PubMed]
- Hirota, T.; Takahashi, A.; Kubo, M.; Tsunoda, T.; Tomita, K.; Sakashita, M.; Yamada, T.; Fujieda, S.; Tanaka, S.; Doi, S.; et al. Genome-wide association study identifies eight new susceptibility loci for atopic dermatitis in the Japanese population. Nat. Genet. 2012, 44, 1222–1226. [Google Scholar] [CrossRef]
- Zhong, F.L.; Mamaï, O.; Sborgi, L.; Boussofara, L.; Hopkins, R.; Robinson, K.; Szeverényi, I.; Takeichi, T.; Balaji, R.; Lau, A.; et al. Germline NLRP1 Mutations Cause Skin Inflammatory and Cancer Susceptibility Syndromes via Inflammasome Activation. Cell 2016, 167, 187–202.e17. [Google Scholar] [CrossRef]
- Martinon, F.; Burns, K.; Tschopp, J. The Inflammasome. Mol. Cell 2002, 10, 417–426. [Google Scholar] [CrossRef]
- Zhong, Y.; Kinio, A.; Saleh, M. Functions of NOD-Like Receptors in Human Diseases. Front. Immunol. 2013, 4, 1–18. [Google Scholar] [CrossRef]
- Faustin, B.; Lartigue, L.; Bruey, J.-M.; Luciano, F.; Sergienko, E.; Bailly-Maitre, B.; Volkmann, N.; Hanein, D.; Rouiller, I.; Reed, J.C. Reconstituted NALP1 Inflammasome Reveals Two-Step Mechanism of Caspase-1 Activation. Mol. Cell 2007, 25, 713–724. [Google Scholar] [CrossRef]
- Boyden, E.D.; Dietrich, W.F. Nalp1b controls mouse macrophage susceptibility to anthrax lethal toxin. Nat. Genet. 2006, 38, 240–244. [Google Scholar] [CrossRef]
- Mitchell, P.S.; Sandstrom, A.; Vance, R.E. The NLRP1 inflammasome: New mechanistic insights and unresolved mysteries. Curr. Opin. Immunol. 2019, 60, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Sandstrom, A.; Mitchell, P.S.; Goers, L.; Mu, E.W.; Lesser, C.F.; Vance, R.E. Functional degradation: A mechanism of NLRP1 inflammasome activation by diverse pathogen enzymes. Science 2019, 364, eaau1330. [Google Scholar] [CrossRef] [PubMed]
- Chavarría-Smith, J.; Vance, R.E. Direct Proteolytic Cleavage of NLRP1B Is Necessary and Sufficient for Inflammasome Activation by Anthrax Lethal Factor. PLoS Pathog. 2013, 9, e1003452. [Google Scholar] [CrossRef]
- Hellmich, K.A.; Levinsohn, J.L.; Fattah, R.; Newman, Z.L.; Maier, N.; Sastalla, I.; Liu, S.; Leppla, S.H.; Moayeri, M. Anthrax Lethal Factor Cleaves Mouse Nlrp1b in Both Toxin-Sensitive and Toxin-Resistant Macrophages. PLoS ONE 2012, 7, e49741. [Google Scholar] [CrossRef] [PubMed]
- Levinsohn, J.L.; Newman, Z.L.; Hellmich, K.A.; Fattah, R.; Getz, M.A.; Liu, S.; Sastalla, I.; Leppla, S.H.; Moayeri, M. Anthrax Lethal Factor Cleavage of Nlrp1 Is Required for Activation of the Inflammasome. PLoS Pathog. 2012, 8, e1002638. [Google Scholar] [CrossRef] [PubMed]
- Ewald, S.E.; Chavarria-Smith, J.; Boothroyd, J.C. NLRP1 Is an Inflammasome Sensor for Toxoplasma gondii. Infect. Immun. 2014, 82, 460–468. [Google Scholar] [CrossRef]
- Bauernfried, S.; Scherr, M.J.; Pichlmair, A.; Duderstadt, K.E.; Hornung, V. Human NLRP1 is a sensor for double-stranded RNA. Science 2021, 371, eabd0811. [Google Scholar] [CrossRef] [PubMed]
- Fenini, G.; Karakaya, T.; Hennig, P.; Di Filippo, M.; Beer, H.D. The NLRP1 inflammasome in human skin and beyond. Int. J. Mol. Sci. 2020, 21, 4788. [Google Scholar] [CrossRef]
- Bachovchin, D.A. NLRP1: A jack of all trades, or a master of one? Mol. Cell 2021, 81, 423–425. [Google Scholar] [CrossRef]
- Cavailles, P.; Flori, P.; Papapietro, O.; Bisanz, C.; Lagrange, D.; Pilloux, L.; Massera, C.; Cristinelli, S.; Jublot, D.; Bastien, O.; et al. A Highly Conserved Toxo1 Haplotype Directs Resistance to Toxoplasmosis and Its Associated Caspase-1 Dependent Killing of Parasite and Host Macrophage. PLoS Pathog. 2014, 10, e1004005. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Mailloux, C.M.; Gowan, K.; Riccardi, S.L.; LaBerge, G.; Bennett, D.C.; Fain, P.R.; Spritz, R.A. NALP1 in Vitiligo-Associated Multiple Autoimmune Disease. N. Engl. J. Med. 2007, 356, 1216–1225. [Google Scholar] [CrossRef] [PubMed]
- Boniface, K.; Jacquemin, C.; Darrigade, A.-S.; Dessarthe, B.; Martins, C.; Boukhedouni, N.; Vernisse, C.; Grasseau, A.; Thiolat, D.; Rambert, J.; et al. Vitiligo Skin Is Imprinted with Resident Memory CD8 T Cells Expressing CXCR3. J. Investig. Dermatol. 2018, 138, 355–364. [Google Scholar] [CrossRef]
- Ciążyńska, M.; Bednarski, I.A.; Wódz, K.; Narbutt, J.; Lesiak, A. NLRP1 and NLRP3 inflammasomes as a new approach to skin carcinogenesis (Review). Oncol. Lett. 2020, 19, 1649–1656. [Google Scholar] [CrossRef]
- Grigoryev, D.N.; Howell, M.D.; Watkins, T.N.; Chen, Y.-C.; Cheadle, C.; Boguniewicz, M.; Barnes, K.C.; Leung, D.Y.M. Vaccinia virus–specific molecular signature in atopic dermatitis skin. J. Allergy Clin. Immunol. 2010, 125, 153–159.e28. [Google Scholar] [CrossRef] [PubMed]
- Bruey, J.M.; Bruey-Sedano, N.; Newman, R.; Chandler, S.; Stehlik, C.; Reed, J.C. PAN1/NALP2/PYPAF2, an Inducible Inflammatory Mediator That Regulates NF-κB and Caspase-1 Activation in Macrophages. J. Biol. Chem. 2004, 279, 51897–51907. [Google Scholar] [CrossRef]
- Ji, S.; Shin, J.E.; Kim, Y.S.; Oh, J.-E.; Min, B.-M.; Choi, Y. Toll-Like Receptor 2 and NALP2 Mediate Induction of Human Beta-Defensins by Fusobacterium nucleatum in Gingival Epithelial Cells. Infect. Immun. 2009, 77, 1044–1052. [Google Scholar] [CrossRef]
- Yoshikawa, Y.; Sasahara, Y.; Takeuchi, K.; Tsujimoto, Y.; Hashida-Okado, T.; Kitano, Y.; Hashimoto-Tamaoki, T. Transcriptional Analysis of Hair Follicle-Derived Keratinocytes from Donors with Atopic Dermatitis Reveals Enhanced Induction of IL32 Gene by IFN-γ. Int. J. Mol. Sci. 2013, 14, 3215–3227. [Google Scholar] [CrossRef]
- Netea, M.G.; Nold-Petry, C.A.; Nold, M.F.; Joosten, L.A.B.; Opitz, B.; van der Meer, J.H.M.; van de Veerdonk, F.L.; Ferwerda, G.; Heinhuis, B.; Devesa, I.; et al. a Differential requirement for the activation of the inflammasome for processing and release of IL-1beta in monocytes and macrophages. Blood 2009, 113, 2324–2335. [Google Scholar] [CrossRef]
- Pétrilli, V.; Papin, S.; Dostert, C.; Mayor, A.; Martinon, F.; Tschopp, J. Activation of the NALP3 inflammasome is triggered by low intracellular potassium concentration. Cell Death Differ. 2007, 14, 1583–1589. [Google Scholar] [CrossRef] [PubMed]
- Martinon, F.; Pétrilli, V.; Mayor, A.; Tardivel, A.; Tschopp, J. Gout-associated uric acid crystals activate the NALP3 inflammasome. Nature 2006, 440, 237–241. [Google Scholar] [CrossRef]
- Martinon, F.; Agostini, L.; Meylan, E.; Tschopp, J. Identification of Bacterial Muramyl Dipeptide as Activator of the NALP3/Cryopyrin Inflammasome. Curr. Biol. 2004, 14, 1929–1934. [Google Scholar] [CrossRef]
- Kanneganti, T.-D.; Body-Malapel, M.; Amer, A.; Park, J.-H.; Whitfield, J.; Franchi, L.; Taraporewala, Z.F.; Miller, D.; Patton, J.T.; Inohara, N.; et al. Critical Role for Cryopyrin/Nalp3 in Activation of Caspase-1 in Response to Viral Infection and Double-stranded RNA. J. Biol. Chem. 2006, 281, 36560–36568. [Google Scholar] [CrossRef]
- Broz, P.; Dixit, V.M. Inflammasomes: Mechanism of assembly, regulation and signalling. Nat. Rev. Immunol. 2016, 16, 407–420. [Google Scholar] [CrossRef]
- He, W.; Wan, H.; Hu, L.; Chen, P.; Wang, X.; Huang, Z.; Yang, Z.-H.; Zhong, C.-Q.; Han, J. Gasdermin D is an executor of pyroptosis and required for interleukin-1β secretion. Cell Res. 2015, 25, 1285–1298. [Google Scholar] [CrossRef] [PubMed]
- Rühl, S.; Broz, P. Caspase-11 activates a canonical NLRP3 inflammasome by promoting K + efflux. Eur. J. Immunol. 2015, 45, 2927–2936. [Google Scholar] [CrossRef]
- Shi, J.; Zhao, Y.; Wang, K.; Shi, X.; Wang, Y.; Huang, H.; Zhuang, Y.; Cai, T.; Wang, F.; Shao, F. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 2015, 526, 660–665. [Google Scholar] [CrossRef] [PubMed]
- Grossi, S.; Fenini, G.; Kockmann, T.; Hennig, P.; Di Filippo, M.; Beer, H.-D. Inactivation of the Cytoprotective Major Vault Protein by Caspase-1 and -9 in Epithelial Cells during Apoptosis. J. Investig. Dermatol. 2020, 140, 1335–1345.e10. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, T.; Li, S.; Yi, X.; Guo, S.; Wang, Y.; Chen, J.; Liu, L.; Jian, Z.; Gao, T.; Kang, P.; et al. Tranilast Directly Targets NLRP3 to Protect Melanocytes from Keratinocyte-Derived IL-1β Under Oxidative Stress. Front. Cell Dev. Biol. 2020, 8, 588. [Google Scholar] [CrossRef]
- Dai, X.; Tohyama, M.; Murakami, M.; Sayama, K. Epidermal keratinocytes sense dsRNA via the NLRP3 inflammasome, mediating interleukin (IL)-1β and IL-18 release. Exp. Dermatol. 2017, 26, 904–911. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Xiao, C.; Dang, E.; Cao, J.; Zhu, Z.; Fu, M.; Yao, X.; Liu, Y.; Jin, B.; Wang, G.; et al. CD100–Plexin-B2 Promotes the Inflammation in Psoriasis by Activating NF-κB and the Inflammasome in Keratinocytes. J. Investig. Dermatol. 2018, 138, 375–383. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, T.; Nakashima, M.; Suzuki, Y. Nuclear DNA damage-triggered NLRP3 inflammasome activation promotes UVB-induced inflammatory responses in human keratinocytes. Biochem. Biophys. Res. Commun. 2016, 477, 329–335. [Google Scholar] [CrossRef]
- Ito, H.; Kanbe, A.; Sakai, H.; Seishima, M. Activation of NLRP3 signalling accelerates skin wound healing. Exp. Dermatol. 2018, 27, 80–86. [Google Scholar] [CrossRef]
- Weinheimer-Haus, E.M.; Mirza, R.E.; Koh, T.J. Nod-Like Receptor Protein-3 Inflammasome Plays an Important Role during Early Stages of Wound Healing. PLoS ONE 2015, 10, e0119106. [Google Scholar] [CrossRef]
- Wang, T.; Zhao, J.; Zhang, J.; Mei, J.; Shao, M.; Pan, Y.; Yang, W.; Jiang, Y.; Liu, F.; Jia, W. Heparan sulfate inhibits inflammation and improves wound healing by downregulating the NLR family pyrin domain containing 3 (NLRP3) inflammasome in diabetic rats. J. Diabetes 2018, 10, 556–563. [Google Scholar] [CrossRef] [PubMed]
- Qing, L.; Fu, J.; Wu, P.; Zhou, Z.; Yu, F.; Tang, J. Metformin induces the m2 macrophage polarization to accelerate the wound healing via regulating ampk/mtor/nlrp3 inflammasome singling pathway. Am. J. Transl. Res. 2019, 11, 655–668. [Google Scholar]
- Wang, D.; Duncan, B.; Li, X.; Shi, J. The role of NLRP3 inflammasome in infection-related, immune-mediated and autoimmune skin diseases. J. Dermatol. Sci. 2020, 98, 146–151. [Google Scholar] [CrossRef]
- Li, S.; Kang, P.; Zhang, W.; Jian, Z.; Zhang, Q.; Yi, X.; Guo, S.; Guo, W.; Shi, Q.; Li, B.; et al. Activated NLR family pyrin domain containing 3 (NLRP3) inflammasome in keratinocytes promotes cutaneous T-cell response in patients with vitiligo. J. Allergy Clin. Immunol. 2020, 145, 632–645. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.J.; Choi, D.K.; Sohn, K.C.; Seo, M.S.; Lee, H.E.; Lee, Y.; Seo, Y.J.; Lee, Y.H.; Shi, G.; Zouboulis, C.C.; et al. Propionibacterium acnes Activates the NLRP3 Inflammasome in Human Sebocytes. J. Investig. Dermatol. 2014, 134, 2747–2756. [Google Scholar] [CrossRef]
- Szabó, K.; Erdei, L.; Bolla, B.S.; Tax, G.; Bíró, T.; Kemény, L. Factors shaping the composition of the cutaneous microbiota. Br. J. Dermatol. 2017, 176, 344–351. [Google Scholar] [CrossRef] [PubMed]
- McLaughlin, J.; Watterson, S.; Layton, A.M.; Bjourson, A.J.; Barnard, E.; McDowell, A. Propionibacterium acnes and Acne Vulgaris: New Insights from the Integration of Population Genetic, Multi-Omic, Biochemical and Host-Microbe Studies. Microorganisms 2019, 7, 128. [Google Scholar] [CrossRef]
- Shen, C.; Wang, Q.Z.; Shen, Z.Y.; Yuan, H.Y.; Yu, W.J.; Chen, X.D.; Xu, H. Genetic association between the NLRP3 gene and acne vulgaris in a Chinese population. Clin. Exp. Dermatol. 2019, 44, 184–189. [Google Scholar] [CrossRef]
- Muñoz-Planillo, R.; Franchi, L.; Miller, L.S.; Núñez, G. A Critical Role for Hemolysins and Bacterial Lipoproteins in Staphylococcus aureus-Induced Activation of the Nlrp3 Inflammasome. J. Immunol. 2009, 183, 3942–3948. [Google Scholar] [CrossRef] [PubMed]
- Craven, R.R.; Gao, X.; Allen, I.C.; Gris, D.; Wardenburg, J.B.; McElvania-TeKippe, E.; Ting, J.P.; Duncan, J.A. Staphylococcus aureus α-Hemolysin Activates the NLRP3-Inflammasome in Human and Mouse Monocytic Cells. PLoS ONE 2009, 4, e7446. [Google Scholar] [CrossRef]
- Dai, X.; Sayama, K.; Tohyama, M.; Shirakata, Y.; Hanakawa, Y.; Tokumaru, S.; Yang, L.; Hirakawa, S.; Hashimoto, K. Mite allergen is a danger signal for the skin via activation of inflammasome in keratinocytes. J. Allergy Clin. Immunol. 2011, 127, 806–814.e4. [Google Scholar] [CrossRef]
- Su, F.; Xia, Y.; Huang, M.; Zhang, L.; Chen, L. Expression of NLPR3 in Psoriasis Is Associated with Enhancement of Interleukin-1β and Caspase-1. Med. Sci. Monit. 2018, 24, 7909–7913. [Google Scholar] [CrossRef] [PubMed]
- Irrera, N.; Vaccaro, M.; Bitto, A.; Pallio, G.; Pizzino, G.; Lentini, M.; Arcoraci, V.; Minutoli, L.; Scuruchi, M.; Cutroneo, G.; et al. BAY 11-7082 inhibits the NF-κB and NLRP3 inflammasome pathways and protects against IMQ-induced psoriasis. Clin. Sci. 2017, 131, 487–498. [Google Scholar] [CrossRef] [PubMed]
- Yu, P.; Hao, S.; Zheng, H.; Zhao, X.; Li, Y. Association of NLRP1 and NLRP3 Polymorphisms with Psoriasis Vulgaris Risk in the Chinese Han Population. Biomed. Res. Int. 2018, 2018, 1–8. [Google Scholar] [CrossRef]
- Primiano, M.J.; Lefker, B.A.; Bowman, M.R.; Bree, A.G.; Hubeau, C.; Bonin, P.D.; Mangan, M.; Dower, K.; Monks, B.G.; Cushing, L.; et al. Efficacy and Pharmacology of the NLRP3 Inflammasome Inhibitor CP-456,773 (CRID3) in Murine Models of Dermal and Pulmonary Inflammation. J. Immunol. 2016, 197, 2421–2433. [Google Scholar] [CrossRef]
- Deng, G.; Chen, W.; Wang, P.; Zhan, T.; Zheng, W.; Gu, Z.; Wang, X.; Ji, X.; Sun, Y. Inhibition of NLRP3 inflammasome-mediated pyroptosis in macrophage by cycloastragenol contributes to amelioration of imiquimod-induced psoriasis-like skin inflammation in mice. Int. Immunopharmacol. 2019, 74, 105682. [Google Scholar] [CrossRef]
- Edqvist, P.D.; Fagerberg, L.; Hallström, B.M.; Danielsson, A.; Edlund, K.; Uhlén, M.; Pontén, F. Expression of Human Skin-Specific Genes Defined by Transcriptomics and Antibody-Based Profiling. J. Histochem. Cytochem. 2015, 63. [Google Scholar] [CrossRef]
- Tanaka, S.; Mizushina, Y.; Kato, Y.; Tamura, M.; Shiroishi, T. Functional Conservation of Gsdma Cluster Genes Speci fi cally Duplicated in the Mouse Genome. G3 Genes Genomes Genet. 2013, 3, 1843–1850. [Google Scholar] [CrossRef]
- Lautz, K.; Damm, A.; Menning, M.; Wenger, J.; Adam, A.C.; Zigrino, P.; Kremmer, E.; Kufer, T.A. NLRP10 enhances shigella-induced pro-inflammatory responses. Cell. Microbiol. 2012, 14, 1568–1583. [Google Scholar] [CrossRef] [PubMed]
- Imamura, R.; Wang, Y.; Kinoshita, T.; Suzuki, M.; Noda, T.; Sagara, J.; Taniguchi, S.; Okamoto, H.; Suda, T. Anti-Inflammatory Activity of PYNOD and Its Mechanism in Humans and Mice. J. Immunol. 2010, 184, 5874–5884. [Google Scholar] [CrossRef]
- Wang, Y.; Hasegawa, M.; Imamura, R.; Kinoshita, T.; Kondo, C.; Konaka, K.; Suda, T. PYNOD, a novel Apaf-1/CED4-like protein is an inhibitor of ASC and caspase-1. Int. Immunol. 2004, 16, 777–786. [Google Scholar] [CrossRef] [PubMed]
- Joly, S.; Eisenbarth, S.C.; Olivier, A.K.; Williams, A.; Kaplan, D.H.; Cassel, S.L.; Flavell, R.A.; Sutterwala, F.S. Cutting Edge: Nlrp10 Is Essential for Protective Antifungal Adaptive Immunity against Candida albicans. J. Immunol. 2012, 189, 4713–4717. [Google Scholar] [CrossRef]
- Eisenbarth, S.C.; Williams, A.; Colegio, O.R.; Meng, H.; Strowig, T.; Rongvaux, A.; Henao-Mejia, J.; Thaiss, C.A.; Joly, S.; Gonzalez, D.G.; et al. NLRP10 is a NOD-like receptor essential to initiate adaptive immunity by dendritic cells. Nature 2012, 484, 510–513. [Google Scholar] [CrossRef]
- Damm, A.; Giebeler, N.; Zamek, J.; Zigrino, P.; Kufer, T.A. Epidermal NLRP10 contributes to contact hypersensitivity responses in mice. Eur. J. Immunol. 2016, 46, 1959–1969. [Google Scholar] [CrossRef] [PubMed]
- Suárez-Fariñas, M.; Tintle, S.J.; Shemer, A.; Chiricozzi, A.; Nograles, K.; Cardinale, I.; Duan, S.; Bowcock, A.M.; Krueger, J.G.; Guttman-Yassky, E. Nonlesional atopic dermatitis skin is characterized by broad terminal differentiation defects and variable immune abnormalities. J. Allergy Clin. Immunol. 2011, 127, 954–964.e4. [Google Scholar] [CrossRef]
- Reubold, T.F.; Hahne, G.; Wohlgemuth, S.; Eschenburg, S. Crystal structure of the leucine-rich repeat domain of the NOD-like receptor NLRP1: Implications for binding of muramyl dipeptide. FEBS Lett. 2014, 588, 3327–3332. [Google Scholar] [CrossRef] [PubMed]
- Arnoult, D.; Soares, F.; Tattoli, I.; Castanier, C.; Philpott, D.J.; Girardin, S.E. An N-terminal addressing sequence targets NLRX1 to the mitochondrial matrix. J. Cell Sci. 2009, 122, 3161–3168. [Google Scholar] [CrossRef]
- Moore, C.B.; Bergstralh, D.T.; Duncan, J.A.; Lei, Y.; Morrison, T.E.; Zimmermann, A.G.; Accavitti-Loper, M.A.; Madden, V.J.; Sun, L.; Ye, Z.; et al. NLRX1 is a regulator of mitochondrial antiviral immunity. Nature 2008, 451, 573–577. [Google Scholar] [CrossRef]
- Tattoli, I.; Carneiro, L.A.; Jéhanno, M.; Magalhaes, J.G.; Shu, Y.; Philpott, D.J.; Arnoult, D.; Girardin, S.E. NLRX1 is a mitochondrial NOD-like receptor that amplifies NF-κB and JNK pathways by inducing reactive oxygen species production. EMBO Rep. 2008, 9, 293–300. [Google Scholar] [CrossRef]
- Hung, S.-C.; Huang, P.-R.; Almeida-da-Silva, C.L.C.; Atanasova, K.R.; Yilmaz, O.; Ojcius, D.M. NLRX1 modulates differentially NLRP3 inflammasome activation and NF-κB signaling during Fusobacterium nucleatum infection. Microbes Infect. 2018, 20, 615–625. [Google Scholar] [CrossRef]
- Fekete, T.; Bencze, D.; Bíró, E.; Benkő, S.; Pázmándi, K. Focusing on the Cell Type Specific Regulatory Actions of NLRX1. Int. J. Mol. Sci. 2021, 22, 1316. [Google Scholar] [CrossRef]
- Mestas, J.; Hughes, C.C.W. Of Mice and Not Men: Differences between Mouse and Human Immunology. J. Immunol. 2004, 172, 2731–2738. [Google Scholar] [CrossRef]
- Zomer, H.D.; Trentin, A.G. Skin wound healing in humans and mice: Challenges in translational research. J. Dermatol. Sci. 2018, 90, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Vaure, C.; Liu, Y. A Comparative Review of Toll-Like Receptor 4 Expression and Functionality in Different Animal Species. Front. Immunol. 2014, 5, 316. [Google Scholar] [CrossRef]
- Chen, L.; Guo, S.; Ranzer, M.J.; DiPietro, L.A. Toll-Like Receptor 4 Has an Essential Role in Early Skin Wound Healing. J. Investig. Dermatol. 2013, 133, 258–267. [Google Scholar] [CrossRef] [PubMed]
- Iram, N.; Mildner, M.; Prior, M.; Petzelbauer, P.; Fiala, C.; Hacker, S.; Schoppl, A.; Tschachler, E.; Elbe-Burger, A. Age-related changes in expression and function of Toll-like receptors in human skin. Development 2012, 139, 4210–4219. [Google Scholar] [CrossRef]
- Applequist, S.E.; Wallin, R.P.A.; Ljunggren, H. Variable expression of Toll-like receptor in murine innate and adaptive immune cell lines. Int. Immunol. 2002, 14, 1065–1074. [Google Scholar] [CrossRef]
- Drexler, S.K.; Bonsignore, L.; Masin, M.; Tardivel, A.; Jackstadt, R.; Hermeking, H.; Schneider, P.; Gross, O.; Tschopp, J.; Yazdi, A.S. Tissue-specific opposing functions of the inflammasome adaptor ASC in the regulation of epithelial skin carcinogenesis. Proc. Natl. Acad. Sci. USA 2012, 109, 18384–18389. [Google Scholar] [CrossRef] [PubMed]
- Göblös, A.; Danis, J.; Vas, K.; Bata-Csörgő, Z.; Kemény, L.; Széll, M. Keratinocytes express functional CARD18, a negative regulator of inflammasome activation, and its altered expression in psoriasis may contribute to disease pathogenesis. Mol. Immunol. 2016, 73, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Qin, H.; Jin, J.; Fischer, H.; Mildner, M.; Gschwandtner, M.; Mlitz, V.; Eckhart, L.; Tschachler, E. The caspase-1 inhibitor CARD18 is specifically expressed during late differentiation of keratinocytes and its expression is lost in lichen planus. J. Dermatol. Sci. 2017, 87, 176–182. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Danis, J.; Mellett, M. Nod-Like Receptors in Host Defence and Disease at the Epidermal Barrier. Int. J. Mol. Sci. 2021, 22, 4677. https://doi.org/10.3390/ijms22094677
Danis J, Mellett M. Nod-Like Receptors in Host Defence and Disease at the Epidermal Barrier. International Journal of Molecular Sciences. 2021; 22(9):4677. https://doi.org/10.3390/ijms22094677
Chicago/Turabian StyleDanis, Judit, and Mark Mellett. 2021. "Nod-Like Receptors in Host Defence and Disease at the Epidermal Barrier" International Journal of Molecular Sciences 22, no. 9: 4677. https://doi.org/10.3390/ijms22094677
APA StyleDanis, J., & Mellett, M. (2021). Nod-Like Receptors in Host Defence and Disease at the Epidermal Barrier. International Journal of Molecular Sciences, 22(9), 4677. https://doi.org/10.3390/ijms22094677






