Conserved Conformational Hierarchy across Functionally Divergent Glycosyltransferases of the GT-B Structural Superfamily as Determined from Microsecond Molecular Dynamics
Abstract
1. Introduction
2. Results
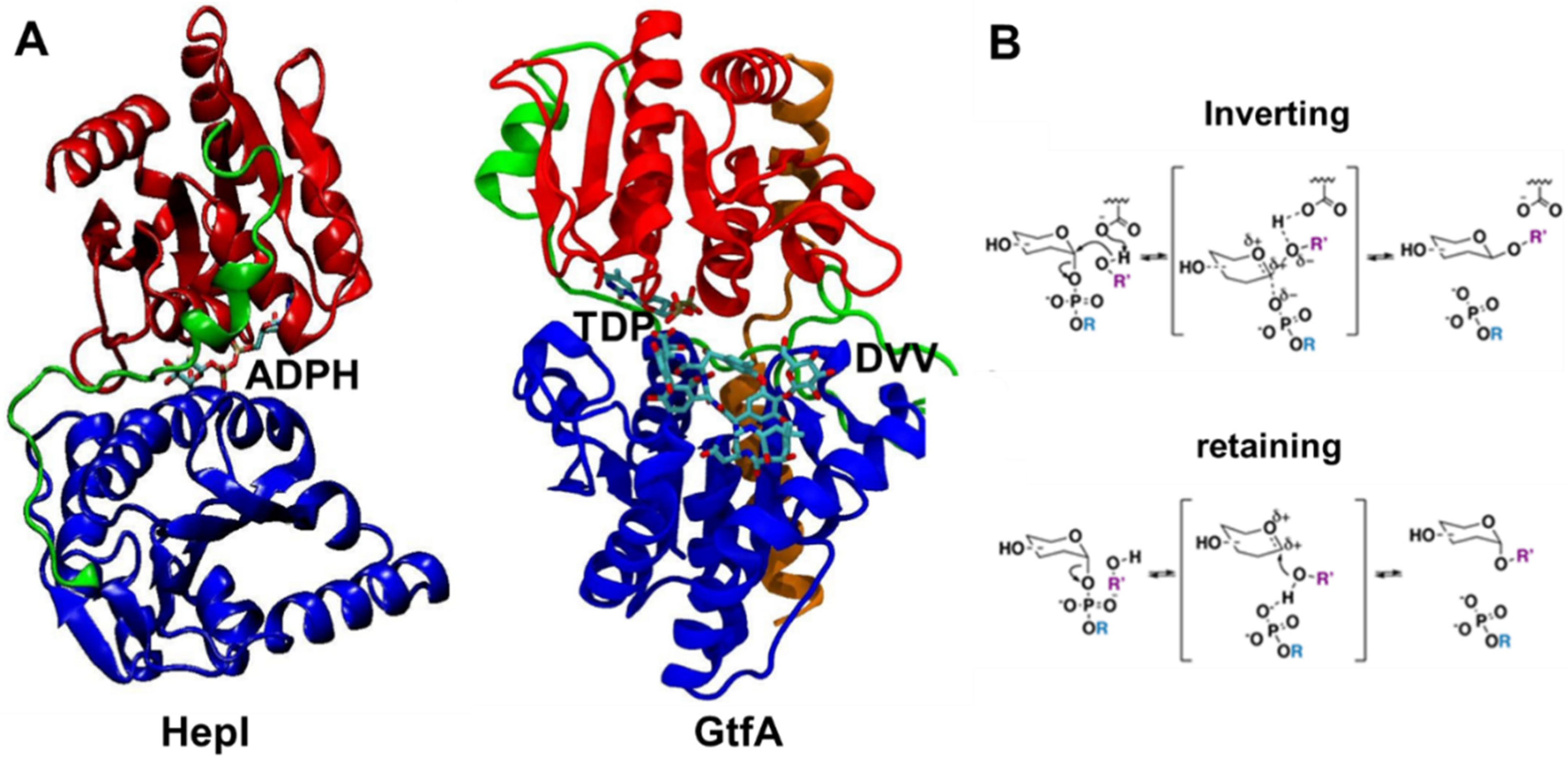
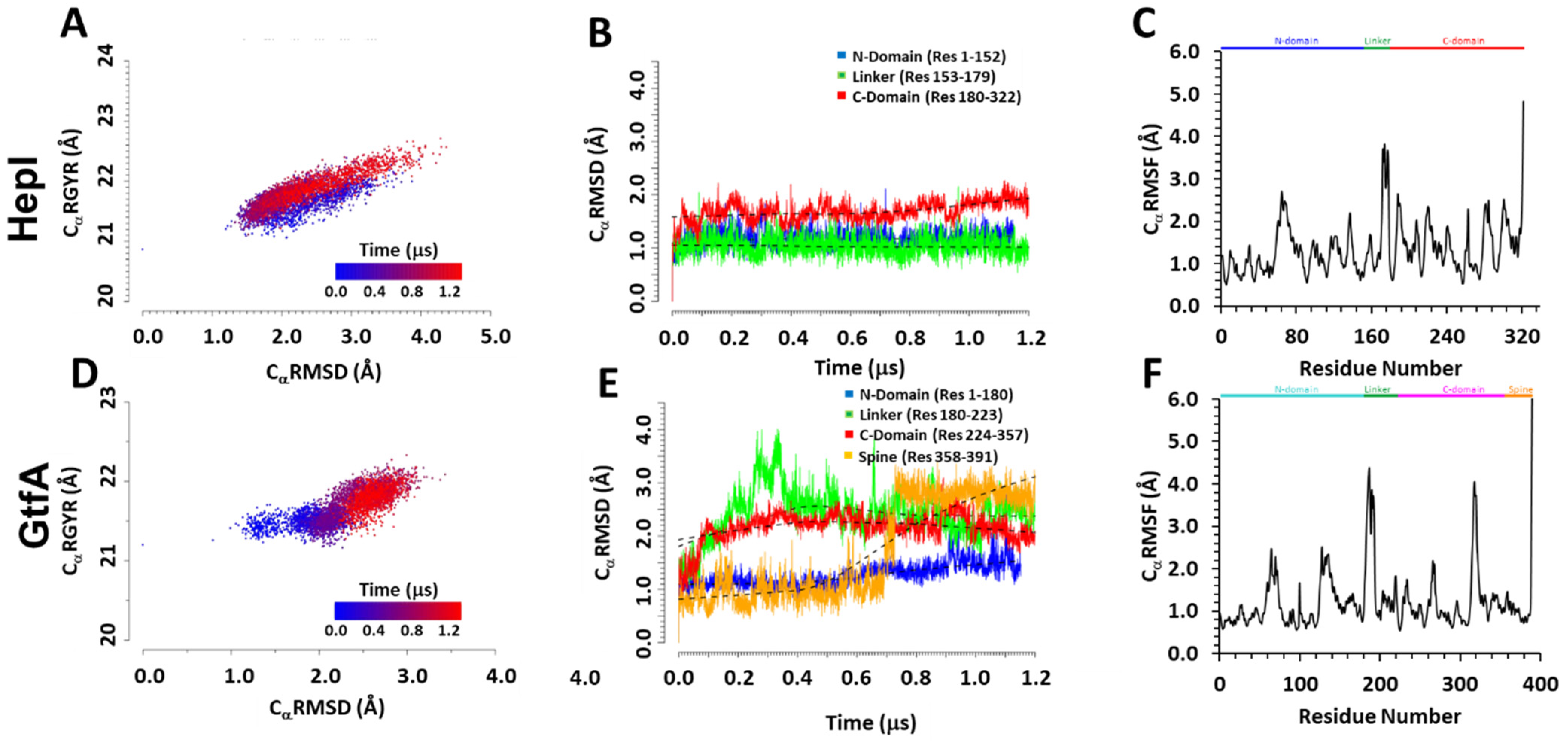
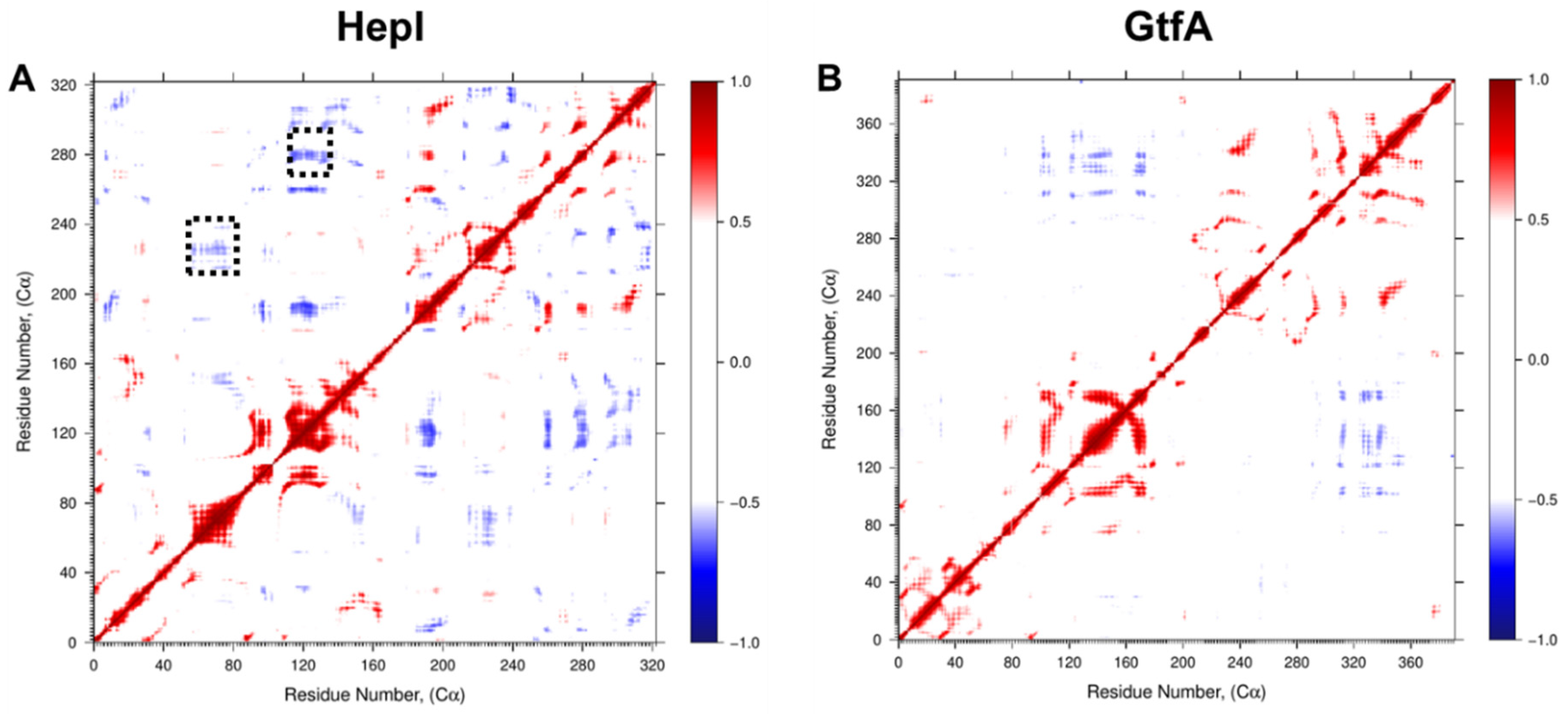
2.1. Differential Dynamic Flexibilities in Open and Closed Configurations in Hepl and GtfA
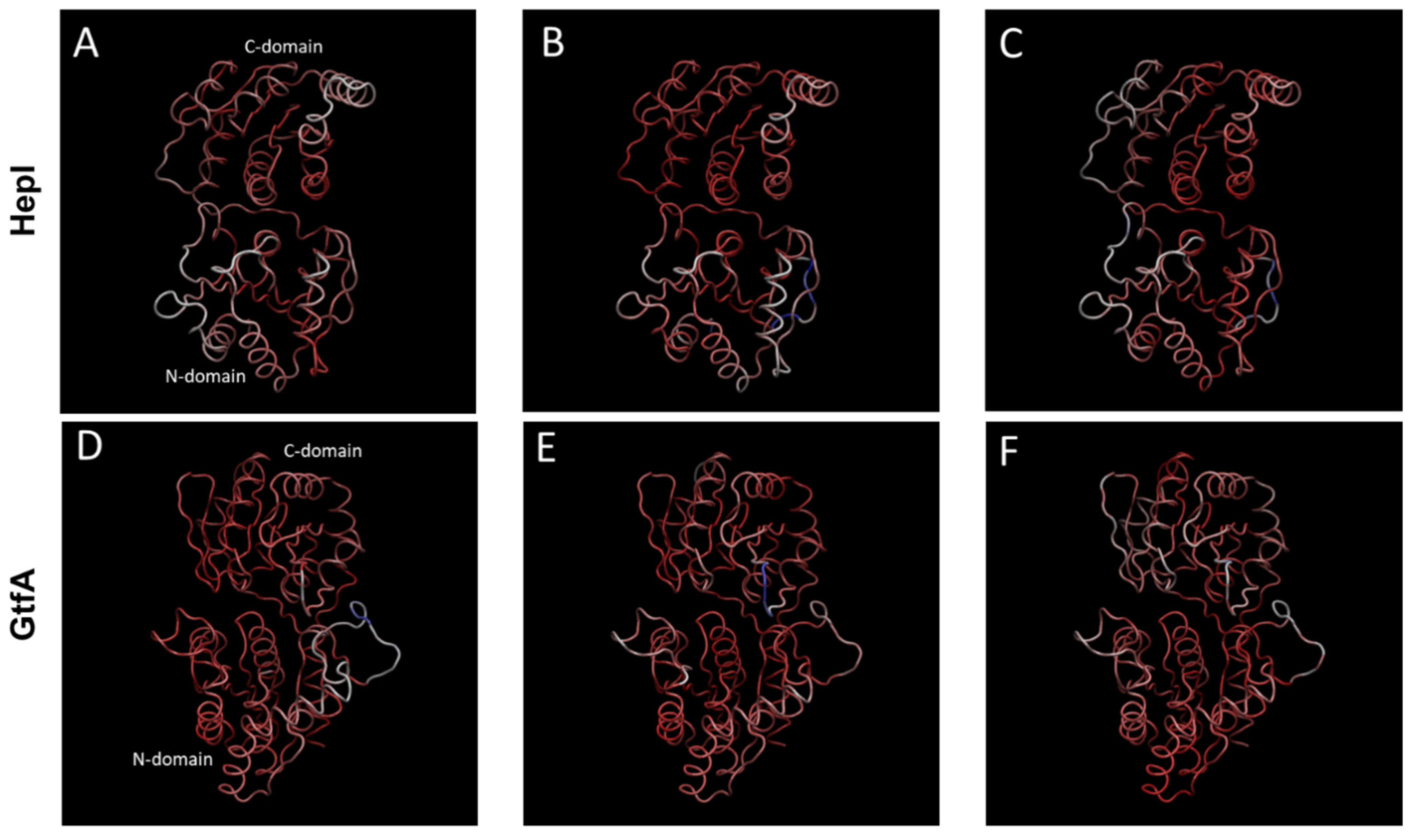
2.2. Distinguishing Characters in Dynamic Fluctuations from Different Secondary Structures
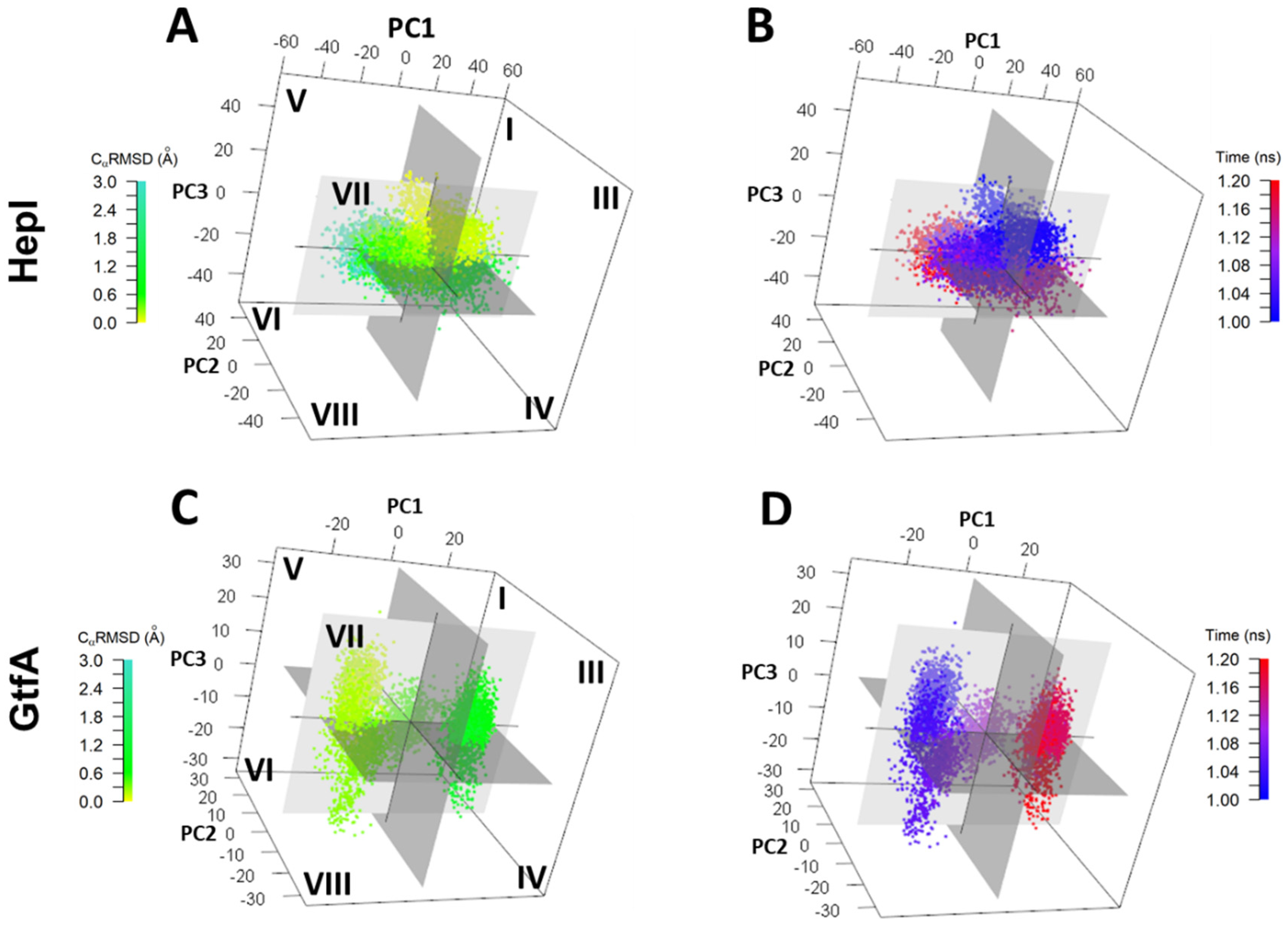
2.3. Conformation Distribution Due to Low-Frequency Large Amplitude Motions
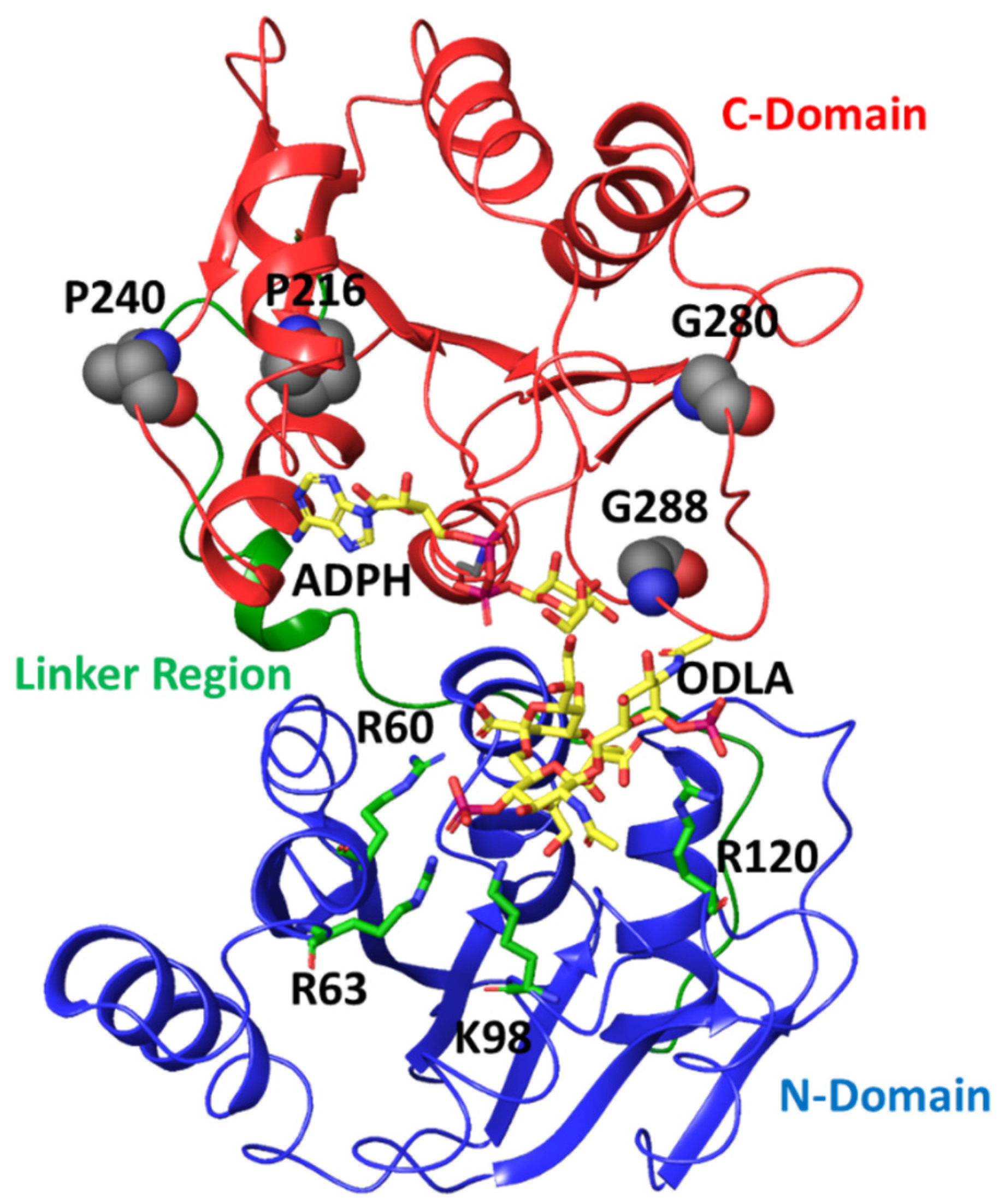
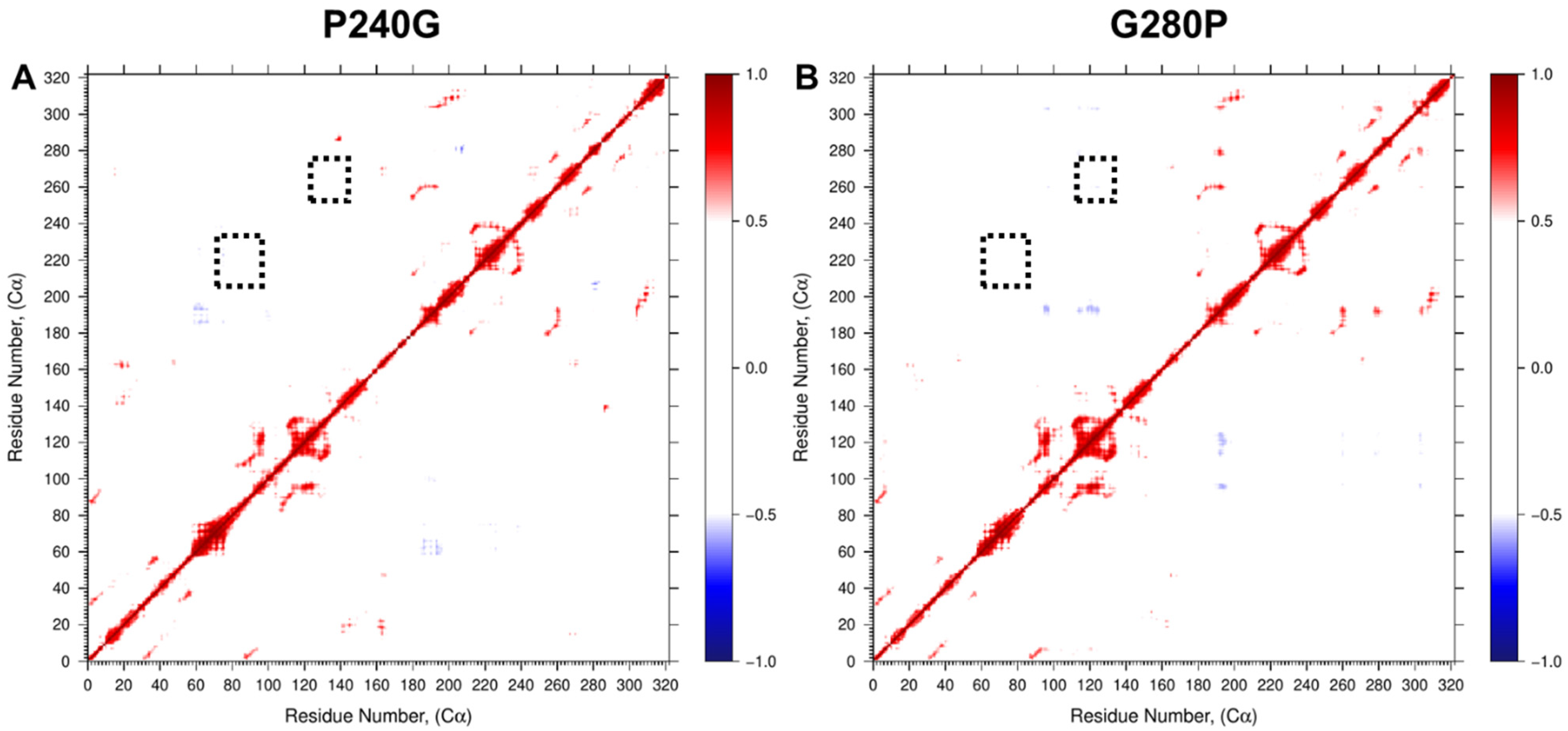
2.4. Dynamics Restriction Affecting Catalytic Efficiency from Site-Directed Mutations
3. Discussion
4. Material and Methods
4.1. Multiple Sequence Alignment (MSA)
4.2. Molecular Dynamics (MD) Simulations
4.3. Site-Directed Mutagenesis of HepI from E. coli K12
- P216G Forward Primer: gcgcgccccacccaagtttaatccgtattcctg
- P216G Reverse Primer: caggaatacggattaaacttgggtggggcgcgc
- P240G Forward Primer: gttgaagtattgggcaagatgagtctggaaggcgttg
- P240G Reverse Primer: caacgccttccagactcatcttgcccaatacttcaac
- G280P Forward Primer: ggatagacccaatatcacggtttatccgccaaccgatccg
- G280P Reverse Primer: cggatcggttggcggataaaccgtgatattgggtctatcc
- G288P Forward Primer: ccgggattaattcctgggtatgggaagaatcagatggtatgtagggctcc
- G288P Reverse Primer: ggagccctacataccatctgattcttcccatacccaggaattaatcccgg
4.4. Mutant Protein Expression and Purification
4.5. HepI Mutant Kinetics Characterization
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Henrissat, B.; Surolia, A.; Stanley, P. A Genomic View of Glycobiology. In Essentials of Glycobiology; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2009. [Google Scholar]
- Albesa-Jové, D.; Romero-García, J.; Sancho-Vaello, E.; Contreras, F.-X.; Rodrigo-Unzueta, A.; Comino, N.; Carreras-González, A.; Arrasate, P.; Urresti, S.; Biarnés, X.; et al. Structural Snapshots and Loop Dynamics along the Catalytic Cycle of Glycosyltransferase GpgS. Structure 2017, 25, 1034–1044.e3. [Google Scholar] [CrossRef]
- Grizot, S.; Salem, M.; Vongsouthi, V.; Durand, L.; Moreau, F.; Dohi, H.; Vincent, S.; Escaich, S.; Ducruix, A. Structure of the Escherichia coli Heptosyltransferase WaaC: Binary Complexes with ADP AND ADP-2-deoxy-2-fluoro Heptose. J. Mol. Biol. 2006, 363, 383–394. [Google Scholar] [CrossRef]
- Vetting, M.W.; Frantom, P.A.; Blanchard, J.S. Structural and Enzymatic Analysis of MshA from Corynebacterium glutamicum: SUBSTRATE-ASSISTED CATALYSIS. J. Biol. Chem. 2008, 283, 15834–15844. [Google Scholar] [CrossRef] [PubMed]
- Mulichak, A.M.; Losey, H.C.; Lu, W.; Wawrzak, Z.; Walsh, C.T.; Garavito, R.M. Structure of the TDP-epi-vancosaminyltransferase GtfA from the chloroeremomycin biosynthetic pathway. Proc. Natl. Acad. Sci. USA 2003, 100, 9238–9243. [Google Scholar] [CrossRef] [PubMed]
- Hurtado-Guerrero, R.; Zusman, T.; Pathak, S.; Ibrahim, A.F.M.; Shepherd, S.; Prescott, A.; Segal, G.; Van Aalten, D.M.F. Molecular mechanism of elongation factor 1A inhibition by a Legionella pneumophila glycosyltransferase. Biochem. J. 2010, 426, 281–292. [Google Scholar] [CrossRef] [PubMed]
- Koç, C.; Gerlach, D.; Beck, S.; Peschel, A.; Xia, G.; Stehle, T. Structural and Enzymatic Analysis of TarM Glycosyltransferase from Staphylococcus aureus Reveals an Oligomeric Protein Specific for the Glycosylation of Wall Teichoic Acid. J. Biol. Chem. 2015, 290, 9874–9885. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, H.; Hansen, G.; Singh, S.; Hanuszkiewicz, A.; Lindner, B.; Fukase, K.; Woodard, R.W.; Holst, O.; Hilgenfeld, R.; Mamat, U.; et al. Structural and mechanistic analysis of the membrane-embedded glycosyltransferase WaaA required for lipopolysaccharide synthesis. Proc. Natl. Acad. Sci. USA 2012, 109, 6253–6258. [Google Scholar] [CrossRef] [PubMed]
- Yao, Q.; Lu, Q.; Wan, X.; Song, F.; Xu, Y.; Hu, M.; Zamyatina, A.; Liu, X.; Huang, N.; Zhu, P.; et al. A structural mechanism for bacterial autotransporter glycosylation by a dodecameric heptosyltransferase family. eLife 2014, 3, e03714. [Google Scholar] [CrossRef] [PubMed]
- Lairson, L.L.; Henrissat, B.; Davies, G.J.; Withers, S.G. Glycosyltransferases: Structures, Functions, and Mechanisms. Annu. Rev. Biochem. 2008, 77, 521–555. [Google Scholar] [CrossRef]
- Mulichak, A.M.; Losey, H.C.; Walsh, C.T.; Garavito, R.M. Structure of the UDP-Glucosyltransferase GtfB That Modifies the Heptapeptide Aglycone in the Biosynthesis of Vancomycin Group Antibiotics. Structure 2001, 9, 547–557. [Google Scholar] [CrossRef]
- Coutinho, P.M.; Deleury, E.; Davies, G.J.; Henrissat, B. An Evolving Hierarchical Family Classification for Glycosyltransferases. J. Mol. Biol. 2003, 328, 307–317. [Google Scholar] [CrossRef]
- Bourne, Y.; Henrissat, B. Glycoside hydrolases and glycosyltransferases: Families and functional modules. Curr. Opin. Struct. Biol. 2001, 11, 593–600. [Google Scholar] [CrossRef]
- Breton, C.; Fournel-Gigleux, S.; Palcic, M.M. Recent structures, evolution and mechanisms of glycosyltransferases. Curr. Opin. Struct. Biol. 2012, 22, 540–549. [Google Scholar] [CrossRef]
- Breton, C.; Šnajdrová, L.; Jeanneau, C.; Koča, J.; Imberty, A. Structures and mechanisms of glycosyltransferases. Glycobiology 2005, 16, 29R–37R. [Google Scholar] [CrossRef]
- Vrielink, A.; Rüger, W.; Driessen, H.P.; Freemont, P.S. Crystal structure of the DNA modifying enzyme beta-glucosyltransferase in the presence and absence of the substrate uridine diphosphoglucose. EMBO J. 1994, 13, 3413–3422. [Google Scholar] [CrossRef]
- Cote, J.M.; Ramirez-Mondragon, C.A.; Siegel, Z.S.; Czyzyk, D.J.; Gao, J.; Sham, Y.Y.; Mukerji, I.; Taylor, E.A. The Stories Tryptophans Tell: Exploring Protein Dynamics of Heptosyltransferase I from Escherichia coli. Biochemistry 2017, 56, 886–895. [Google Scholar] [CrossRef] [PubMed]
- Czyzyk, D.J.; Sawant, S.S.; Ramirez-Mondragon, C.A.; Hingorani, M.M.; Taylor, E.A. Escherichia coli Heptosyltransferase I: Investigation of Protein Dynamics of a GT-B Structural Enzyme. Biochemistry 2013, 52, 5158–5160. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Coleman, W.G., Jr. Cloning and characterization of the Escherichia coli K-12 rfa-2 (rfaC) gene, a gene required for lipopolysaccharide inner core synthesis. J. Bacteriol. 1993, 175, 2534–2540. [Google Scholar] [CrossRef] [PubMed][Green Version]
- De Kievit, T.R.; Lam, J.S. Isolation and characterization of two genes, waaC (rfaC) and waaF (rfaF), involved in Pseudomonas aeruginosa serotype O5 inner-core biosynthesis. J. Bacteriol. 1997, 179, 3451–3457. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Schnaitman, C.A.; Klena, J.D. Genetics of lipopolysaccharide biosynthesis in enteric bacteria. Microbiol. Rev. 1993, 57, 655–682. [Google Scholar] [CrossRef]
- Sirisena, D.M.; Brozek, K.A.; MacLachlan, P.R.; Sanderson, K.E.; Raetz, C.R. The rfaC gene of Salmonella typhimurium. Cloning, sequencing, and enzymatic function in heptose transfer to lipopolysaccharide. J. Biol. Chem. 1992, 267, 18874–18884. [Google Scholar] [CrossRef]
- Romero-García, J.; Francisco, C.; Biarnés, X.; Planas, A. Structure-Function Features of a Mycoplasma Glycolipid Synthase Derived from Structural Data Integration, Molecular Simulations, and Mutational Analysis. PLoS ONE 2013, 8, e81990. [Google Scholar] [CrossRef]
- Righino, B.; Bozzi, M.; Pirolli, D.; Sciandra, F.; Bigotti, M.G.; Brancaccio, A.; De Rosa, M.C. Identification and Modeling of a GT-A Fold in the α-Dystroglycan Glycosylating Enzyme LARGE1. J. Chem. Inf. Model. 2020, 60, 3145–3156. [Google Scholar] [CrossRef]
- Janos, P.; Kozmon, S.; Tvaroska, I.; Koca, J. Three-dimensional homology model of GlcNAc-TV glycosyltransferase. Glycobiology 2016, 26, 757–771. [Google Scholar] [CrossRef]
- Panda, S.K.; Saxena, S.; Guruprasad, L. Homology modeling, docking and structure-based virtual screening for new inhibitor identification of Klebsiella pneumoniae heptosyltransferase-III. J. Biomol. Struct. Dyn. 2019, 38, 1887–1902. [Google Scholar] [CrossRef]
- Ge, C.; Gómez-Llobregat, J.; Skwark, M.J.; Ruysschaert, J.-M.; Wieslander, Å.; Lindén, M. Membrane remodeling capacity of a vesicle-inducing glycosyltransferase. FEBS J. 2014, 281, 3667–3684. [Google Scholar] [CrossRef] [PubMed]
- Ramírez, A.S.; Boilevin, J.; Mehdipour, A.R.; Hummer, G.; Darbre, T.; Reymond, J.-L.; Locher, K.P. Structural basis of the molecular ruler mechanism of a bacterial glycosyltransferase. Nat. Commun. 2018, 9, 445. [Google Scholar] [CrossRef] [PubMed]
- Salinas, S.; Petruk, A.; Brukman, N.; Bianco, M.; Jacobs, M.; Marti, M.; Ielpi, L. Binding of the substrate UDP-glucuronic acid induces conformational changes in the xanthan gum glucuronosyltransferase. Protein Eng. Des. Sel. 2016, 29, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Šnajdrová, L.; Kulhánek, P.; Imberty, A.; Koča, J. Molecular dynamics simulations of glycosyltransferase LgtC. Carbohydr. Res. 2004, 339, 995–1006. [Google Scholar] [CrossRef]
- Shaw, D.E.; Dror, R.O.; Salmon, J.K.; Grossman, J.P.; Mackenzie, K.M.; Bank, J.A.; Young, C.; Deneroff, M.M.; Batson, B.; Bowers, K.J.; et al. Millisecond-scale molecular dynamics simulations on Anton. In Proceedings of the Conference on High Performance Computing Networking, Storage and Analysis, Portland, OR, USA, 14–20 November 2009; ACM: New York, NY, USA, 2009; pp. 1–11. [Google Scholar]
- Gaieb, Z.; Morikis, D. Detection of Side Chain Rearrangements Mediating the Motions of Transmembrane Helices in Molecular Dynamics Simulations of G Protein-Coupled Receptors. Comput. Struct. Biotechnol. J. 2017, 15, 131–137. [Google Scholar] [CrossRef]
- Gur, M.; Zomot, E.; Bahar, I. Global motions exhibited by proteins in micro- to milliseconds simulations concur with anisotropic network model predictions. J. Chem. Phys. 2013, 139, 121912. [Google Scholar] [CrossRef]
- Hu, X.; Hong, L.; Smith, M.D.; Neusius, T.; Cheng, X.; Smith, J.C. The dynamics of single protein molecules is non-equilibrium and self-similar over thirteen decades in time. Nat. Phys. 2016, 12, 171–174. [Google Scholar] [CrossRef]
- Shaw, D.E.; Maragakis, P.; Lindorff-Larsen, K.; Piana, S.; Dror, R.O.; Eastwood, M.P.; Bank, J.A.; Jumper, J.M.; Salmon, J.K.; Shan, Y.; et al. Atomic-Level Characterization of the Structural Dynamics of Proteins. Science 2010, 330, 341–346. [Google Scholar] [CrossRef]
- Cote, J.M.; Hecht, C.J.S.; Patel, K.R.; Ramirez-Mondragon, C.A.; Sham, Y.Y.; Taylor, E.A. Opposites Attract: Escherichia coli Heptosyltransferase I Conformational Changes Induced by Interactions between the Substrate and Positively Charged Residues. Biochemistry 2020, 59, 3135–3147. [Google Scholar] [CrossRef]
- Bourgeois, D.; Vallone, B.; Schotte, F.; Arcovito, A.; Miele, A.E.; Sciara, G.; Wulff, M.; Anfinrud, P.; Brunori, M. Complex Landscape of Protein Structural Dynamics Unveiled by Nanosecond Laue Crystallography. Proc. Natl. Acad. Sci. USA 2003, 100, 8704–8709. [Google Scholar] [CrossRef]
- Frauenfelder, H.; Sligar, S.G.; Wolynes, P.G. The energy landscapes and motions of proteins. Science 1991, 254, 1598–1603. [Google Scholar] [CrossRef] [PubMed]
- Henzler-Wildman, K.A.; Thai, V.; Lei, M.; Ott, M.; Wolf-Watz, M.; Fenn, T.; Pozharski, E.; Wilson, M.A.; Petsko, G.A.; Karplus, M.; et al. Intrinsic motions along an enzymatic reaction trajectory. Nature 2007, 450, 838–844. [Google Scholar] [CrossRef] [PubMed]
- Blaukopf, M.; Worrall, L.; Kosma, P.; Strynadka, N.C.J.; Withers, S.G. Insights into Heptosyltransferase I Catalysis and Inhibition through the Structure of Its Ternary Complex. Structure 2018, 26, 1399–1407.e5. [Google Scholar] [CrossRef] [PubMed]
- Jacob, J.; Duclohier, H.; Cafiso, D.S. The Role of Proline and Glycine in Determining the Backbone Flexibility of a Channel-Forming Peptide. Biochem. J. 1999, 76, 1367–1376. [Google Scholar] [CrossRef]
- Krieger, F.; Möglich, A.; Kiefhaber, T. Effect of Proline and Glycine Residues on Dynamics and Barriers of Loop Formation in Polypeptide Chains. J. Am. Chem. Soc. 2005, 127, 3346–3352. [Google Scholar] [CrossRef]
- Warshel, A. Electrostatic Origin of the Catalytic Power of Enzymes and the Role of Preorganized Active Sites. J. Biol. Chem. 1998, 273, 27035–27038. [Google Scholar] [CrossRef]
- Olsson, M.H.M.; Parson, W.W.; Warshel, A. Dynamical Contributions to Enzyme Catalysis: Critical Tests of A Popular Hypothesis. Chem. Rev. 2006, 106, 1737–1756. [Google Scholar] [CrossRef]
- Ryckaert, J.-P.; Ciccotti, G.; Berendsen, H.J.C. Numerical integration of the cartesian equations of motion of a system with constraints: Molecular dynamics of n-alkanes. J. Comput. Phys. 1977, 23, 327–341. [Google Scholar] [CrossRef]
- Elber, R.; Ruymgaart, A.P.; Hess, B. SHAKE parallelization. Eur. Phys. J. Spéc. Top. 2011, 200, 211–223. [Google Scholar] [CrossRef] [PubMed]
- Phillips, J.C.; Hardy, D.J.; Maia, J.D.C.; Stone, J.E.; Ribeiro, J.V.; Bernardi, R.C.; Buch, R.; Fiorin, G.; Hénin, J.; Jiang, W.; et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 2020, 153, 044130. [Google Scholar] [CrossRef] [PubMed]
- Grant, B.J.; Rodrigues, A.P.; ElSawy, K.M.; McCammon, J.A.; Caves, L.S. Bio3d: An R package for the comparative analysis of protein structures. Bioinformatics 2006, 22, 2695–2696. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wen, B.; Peng, J.; Zuo, X.; Gong, Q.; Zhang, Z. Determining structural ensembles of flexible multi-domain proteins using small-angle X-ray scattering and molecular dynamics simulations. Protein Cell 2015, 6, 619–623. [Google Scholar] [CrossRef] [PubMed]
| HepI | kcat (s−1) | Fold Change | KM (M) | Fold Change | kcat/kM |
|---|---|---|---|---|---|
| Wild type | 0.59 ± 0.09 | - | 8.5 ± 3.6 | - | 6.8 × 104 |
| P216G | 0.21 ± 0.02 | 2.8 ↓ | 1.6 ± 0.6 | 5.6 ↓ | 1.3 × 105 |
| P240G | 0.19 ± 0.01 | 3.1 ↓ | 0.9 ± 0.3 | 10 ↓ | 2.0 × 105 |
| G280P | 0.21 ± 0.02 | 2.8 ↓ | 1.5 ± 0.5 | 6.0 ↓ | 1.3 × 105 |
| G288P | 0.48 ± 0.07 | 1.2 ↓ | 10.2 ± 0.4 | 1.1 ↑ | 4.7 × 104 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramirez-Mondragon, C.A.; Nguyen, M.E.; Milicaj, J.; Hassan, B.A.; Tucci, F.J.; Muthyala, R.; Gao, J.; Taylor, E.A.; Sham, Y.Y. Conserved Conformational Hierarchy across Functionally Divergent Glycosyltransferases of the GT-B Structural Superfamily as Determined from Microsecond Molecular Dynamics. Int. J. Mol. Sci. 2021, 22, 4619. https://doi.org/10.3390/ijms22094619
Ramirez-Mondragon CA, Nguyen ME, Milicaj J, Hassan BA, Tucci FJ, Muthyala R, Gao J, Taylor EA, Sham YY. Conserved Conformational Hierarchy across Functionally Divergent Glycosyltransferases of the GT-B Structural Superfamily as Determined from Microsecond Molecular Dynamics. International Journal of Molecular Sciences. 2021; 22(9):4619. https://doi.org/10.3390/ijms22094619
Chicago/Turabian StyleRamirez-Mondragon, Carlos A., Megin E. Nguyen, Jozafina Milicaj, Bakar A. Hassan, Frank J. Tucci, Ramaiah Muthyala, Jiali Gao, Erika A. Taylor, and Yuk Y. Sham. 2021. "Conserved Conformational Hierarchy across Functionally Divergent Glycosyltransferases of the GT-B Structural Superfamily as Determined from Microsecond Molecular Dynamics" International Journal of Molecular Sciences 22, no. 9: 4619. https://doi.org/10.3390/ijms22094619
APA StyleRamirez-Mondragon, C. A., Nguyen, M. E., Milicaj, J., Hassan, B. A., Tucci, F. J., Muthyala, R., Gao, J., Taylor, E. A., & Sham, Y. Y. (2021). Conserved Conformational Hierarchy across Functionally Divergent Glycosyltransferases of the GT-B Structural Superfamily as Determined from Microsecond Molecular Dynamics. International Journal of Molecular Sciences, 22(9), 4619. https://doi.org/10.3390/ijms22094619








