Abstract
TaCKX gene family members (GFMs) play essential roles in the regulation of cytokinin during wheat development and significantly influence yield-related traits. However, detailed function of most of them is not known. To characterize the role of TaCKX2.2 genes we silenced all homoeologous copies of both TaCKX2.2.1 and TaCKX2.2.2 by RNAi technology and observed the effect of silencing in 7 DAP spikes of T1 and T2 generations. The levels of gene silencing of these developmentally regulated genes were different in both generations, which variously determined particular phenotypes. High silencing of TaCKX2.2.2 in T2 was accompanied by slight down-regulation of TaCKX2.2.1 and strong up-regulation of TaCKX5 and TaCKX11, and expression of TaCKX1, TaCKX2.1, and TaCKX9 was comparable to the non-silenced control. Co-ordinated expression of TaCKX2.2.2 with other TaCKX GFMs influenced phytohormonal homeostasis. Contents of isoprenoid, active cytokinins, their conjugates, and auxin in seven DAP spikes of silenced T2 plants increased from 1.27 to 2.51 times. However, benzyladenine (BA) and abscisic acid (ABA) contents were significantly reduced and GA3 was not detected. We documented a significant role of TaCKX2.2.2 in the regulation of thousand grain weight (TGW), grain number, and chlorophyll content, and demonstrated the formation of a homeostatic feedback loop between the transcription of tested genes and phytohormones. We also discuss the mechanism of regulation of yield-related traits.
1. Introduction
Bread wheat (Triticum aestivum L.) is one of the most widely grown and economically important cereal species in the world. This high-yielding species is comparable in production to rice and maize but contains higher protein level, and is rich in vitamins and dietary fibres [1,2]. A further increase of wheat yield is important to feed the growing world population [3,4]. The large and complex, allohexaploid genome of bread wheat, which is composed of three diploid genomes (AABBDD), is a great reservoir of homoeologous genes. Knowledge of their function can be applied directly or after genetic modifications in wheat breeding as reviewed previously [5]. Therefore, functional characterization of yield-related genes in this species is especially important. Transformation technology used in this type of research is still a bottle-neck in the case of species with a large and complex genome. This is a possible reason for the weak progress in research on bread wheat compared to other cereals.
Cytokinin (CK) is an important growth regulator of plant development influencing many yield-related processes as reviewed in Kieber and Schaller [6] and in Sakakibara [7]. This phytohormone is considered as ‘a key driver’ of seed yield in many crops [8] including barley [9,10] and wheat [11,12,13]. Its function is coordinated by various phytohormones: auxin, gibberellins (GAs), brassinosteroids and others [14]. Generally, cytokinin promotes cell division, and auxin cell expansion and the balance between these two processes controls root and shoot growth. Biosynthesis of gibberellins is supported by auxin transport. Therefore, GAs repress tillering but promote stem elongation. Tillering is inhibited by auxin, as well as by reducing CK content [12,15], reviewed in Guo et al. [14]. Moreover, CKs are negative regulators of resistance to abiotic stress and root growth [16,17] and reduction of their level increases ABA accumulation [18]. However, in wheat kernels an increased cis-zeatin (cZ) level was followed by increased ABA accumulation [13]. Hormone homeostasis is further regulated by co-expressed genes and other factors including miRNA and signalling proteins. Integrators of these various developmental signals are transcription factors, which might reprogram expression of the interacting genes [19]. Fine tuning the expression of key genes’ might be a promising strategy to improve yield-related traits [14].
Cytokinin levels in tissues and organs of plants is regulated by biosynthesis and metabolism as well as compartmentalization and translocation through the plant [8,20]. Cytokinin metabolites differ in their biological activity [21,22]. Among different forms of CKs, the most abundant are isoprenoid CKs: N6-(Δ2-isopentenyl)adenine (iP), trans-zeatin (tZ), cis-zeatin (cZ) and dihydrozeatin (DZ). Benzyladenine (BA) belongs to the second smaller group of aromatic CKs [23]. These free base CKs listed above, which are products of biosynthesis, are the most biologically active. They undergo a series of interconversions to nucleosides (CK ribosides) and nucleotides as summarized in Hosek et al. [24]. Nucleosides-tZ riboside (tZR), cZR, DZR and iPR-are believed to be the main transporter metabolites, although CK bases, tZ and iP were found in xylem sap as well [20]. Translocation of tZ and tZR via the xylem from roots to shoots controls various shoot traits [7]. Conjugation of free bases with glucose via O-glucosyltransferases leads to their rapid deactivation to O-glucosides, which can be converted back to free bases and ribosides by β-glucosidase. This enzyme was identified in maize [25], products were detected in 7 days after pollination (DAP) spikes of wheat [11] and alleles of TaZOGs and cisZOGs were localized [26] and updated (J. Song, personal communication). Another enzyme, N-glucosyltransferase glycosylate bases of all CK-types at the N7 and N9 positions, making them biologically inactive in Arabidopsis [27]. However, N9-glucosides of iP, tZ and cZ were reported to be substrates of cytokinin oxidase/dehydrogenase enzyme in maize [28] and tZ-N7G, tZ-N9G in Arabidopsis [24].
Cytokinin oxidase/dehydrogenase (CKX) catalyzes irreversible degradation of free bases iP, tZ, cZ and their ribosides iPR, tZR, cZR by cleaving the N6-side chain. DZs are resistant to this enzyme [23]. This step of CK metabolism plays an important role in the regulation of cytokinin level and regulation of yield-related traits in cereals, inter alia rice, barley and wheat, as was documented by RNAi silencing of selected genes [9,10,11,29,30].
CKX enzymes are encoded by CKX gene family members (GFMs). According to the latest revision by Chen et al. [12] there are 13 TaCKX GFMs basically numbered from one to 11, and 12 of them are located on the three homoeologous chromosomes of the A, B and D genome, giving the number of 35 GFMs. Exception are TaCKX2.2.2 and TaCKX2.2.3 allocated only to the D genome. This new numbering is based on the newly released wheat genome database (IWGSC RefSeq v1.0 & v2.0) and phylogenic cladogram with other cereals (barley and rice). The numbering of TaCKX2, which has undergone gene duplication [31], is the most expanded. The previous number of TaCKX2.2 [32] was revised for TaCKX2.2.1 located on A, B, and D genomes and mentioned above TaCKX2.2.2 and 2.2.3. Other genes revised according to previous publications [26,32], which are referred to this study, are TaCKX10 renamed as TaCKX9 and TaCKX3 renamed as TaCKX11. The TaCKX GFMs are specifically expressed in various tissues and organs [12,32,33], which might indicate their role in reproductive development [10]. According to the last assignments, TaCKX1, TaCKX2.2.1 and TaCKX2.2.2 are mainly expressed in developing spikes, TaCKX3 and TaCKX10 in seedling roots, TaCKX5, TaCKX9 and TaCKX8 in younger organs of developing plants from seedling roots to 0 days after pollination (DAP) spikes, and TaCKX11 and TaCKX4 in all organs [32,33]. Similar tissue specificity of expression of TaCKX GFMs was presented based on RNA-seq data [12], as well as for TaCKX1 and TaCKX2 during early kernel development [13].
Only some TaCKX GFMs are functionally characterized. Silencing of TaCKX1 by RNAi influenced levels of expression of the other TaCKX GFMs, and increased content of most CK forms and GA3, which caused an increase of yield parameters such as a higher spike number, grain number, and grain yield, but lower thousand grain weight [11]. The effect of TaCKX2.4 gene silencing by RNAi increased grain number per spike [34]. Another source of analysis of gene function and/or selection of desirable phenotypes in bread wheat, for which natural or induced mutants are not available, is natural variation among genotypes [33]. Haplotype variants of TaCKX6a02 and TaCKX6-D1 were associated with higher filling rate and grain size [35,36], and higher copy number of TaCKX4 was related to increased chlorophyll content and grain size [37]. The above-mentioned genes are now annotated: TaCKX6a02 as TaCKX2.2.1–3A, TaCKX6-D1 (JQ797673) as TaCKX2.2.2–3D and TaCKX2.4 as TaCKX2.2.1–3B [12].
Some functions of these genes might also be predicted by known function of their orthologues in other cereal species. Silencing of HvCKX1 orthologous to TaCKX1 resulted in elevated grain number in these two, closely related species [9,11]. However, the same effect of increased grain number in rice was obtained by silencing or knocking-out OsCKX2 [30]. OsCKX11, orthologous to wheat TaCKX11, which is expressed in all organs of the wheat plant [12,32], coordinated sink–source relationships and leaf senescence and grain number in rice [38].
Both methods of gene silencing by RNAi and knock-out mutation by CRISPR/Cas9 have been successfully used for analysis of gene function. The advantages of the first one are the possibility of silencing homoeologous copies of the genes and obtaining plants with different levels of silencing. This is especially important in the case of developmentally regulated genes, for which the level of silencing might lead to different phenotypes [11], or in the case of genes coding proteins of key importance for life [39]. The phenotypic differences between silencing of HvCKX1 [9] and knocking-out the same gene [40] are documented.
In this study we silenced homoeologues of both TaCKX2.2 genes, TaCKX2.2.1 and TaCKX2.2.2, by RNAi technology and analyzed the phenotypic effect of silencing them in T1. In addition to these two gene family members we monitored TaCKX2.1 and other TaCKX GFMs in the T2 generation. Similar study has already been performed in our lab with another GFM, TaCKX1, silenced in the same genotype [11] and the data are compared. To reveal mechanisms of regulation of yield-related traits cross-talk of silenced genes with other TaCKX GFMs as well as homeostatic feedback loop between their transcription and phytohormones was investigated.
2. Results
2.1. Expression Levels of Silenced TaCKX2.2.1, TaCKX2.2.2 and TaCKX2.1 in T1 and T2
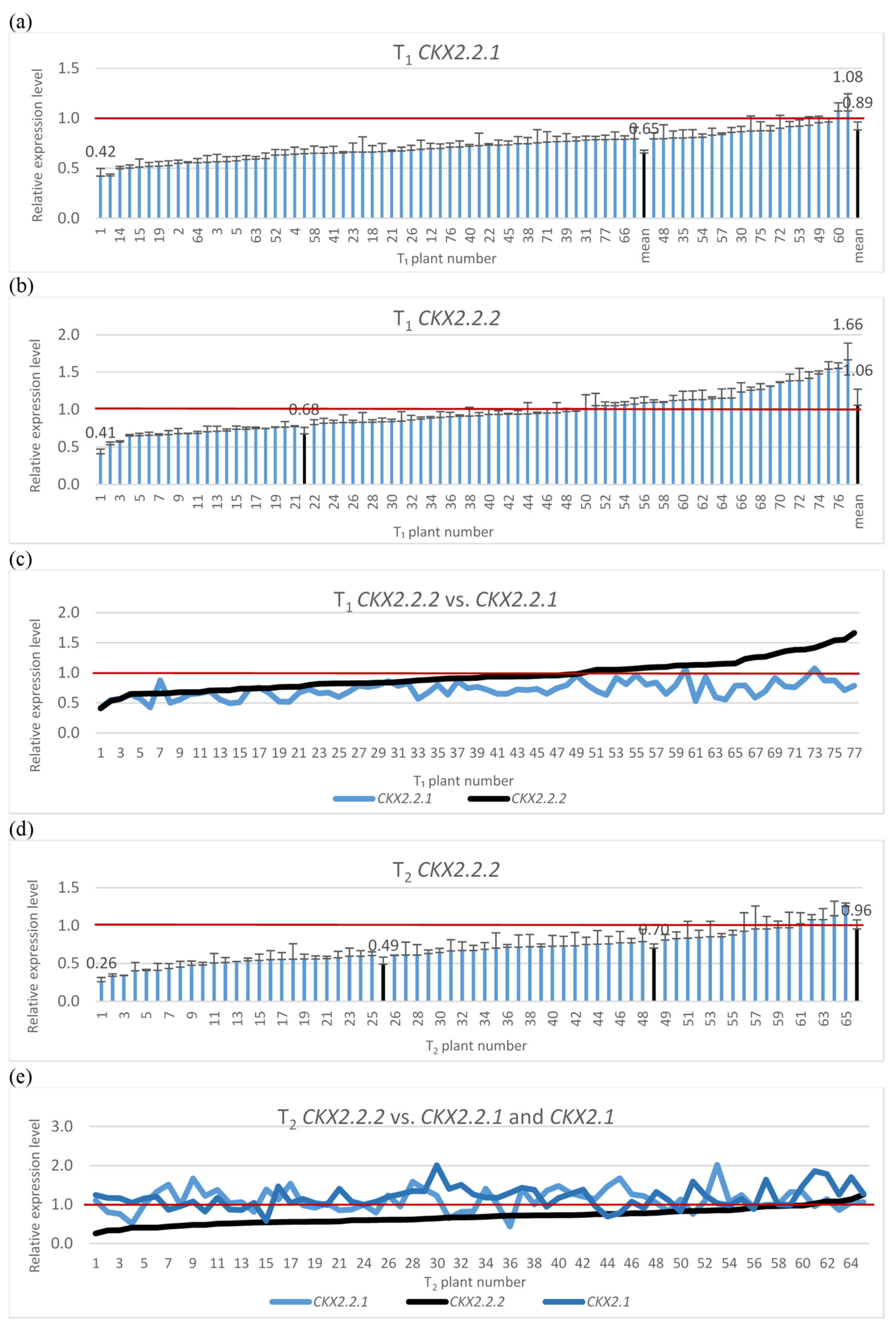
The silencing cassette contains homologous sequences for silencing of three TaCKX2 genes, TaCKX2.2.1, TaCKX2.2.2 and TaCKX2.1. Therefore, the silencing effect for the two was tested in T1 (Figure 1a,c) and a test for TaCKX2.1 was included in the T2 generation (Figure 1d,e). The range of relative expression levels for TaCKX2.2.1 in T1 was from 0.42 to 1.08, and the mean was 0.89. The mean for silenced plants (≤0.8) was 0.65. Relative expression levels for TaCKX2.2.2 ranged from 0.41 to 1.66 and the mean for silenced plants was 0.68. Silencing for both genes in tested T1 plants was similar for those with higher silencing but differed in the plants with higher expression levels. The correlation coefficient between the levels of expression of TaCKX2.2.1 and TaCKX2.2.2 was 0.53 for 77 plants (N 77).
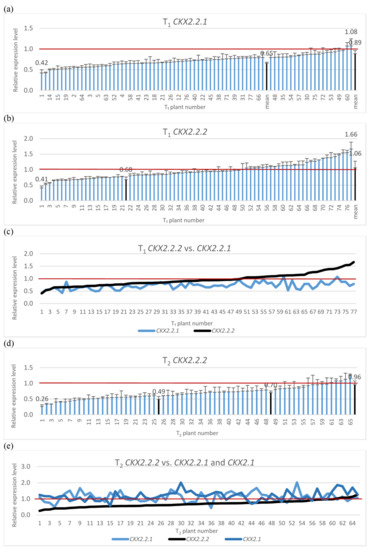
Figure 1.
Relative expression levels of silenced TaCKX2.2.1 (a), TaCKX2.2.2 (b), TaCKX2.2.2 vs. TaCKX2.2.1 (c) in T1 and TaCKX2.2.2 (d), and TaCKX2.2.2 vs. TaCKX2.2.1 and TaCKX2.1 (e). The error bars denote ± SD. The thresholds of classified plants are 0.6 and 0.8.
The most silenced gene in T2 was TaCKX2.2.2 (Figure 1d,e). Levels of silencing of this gene in tested lines ranged from 0.26 to 1.25 and the mean for the highly silenced one (≤0.60) was 0.49; for those with relative expression between 0.61 to 0.79 it was 0.7, and for the third group of non-silenced genes it ranged from 0.8 to 1.25 with a mean of 0.96 (see also Figure 2 and Supplementary Table S1). Levels of expression of TaCKX2.2.1 ranged from 0.44 to 2.01 with the mean of 1.14, but silencing below 0.79 was observed only in six out of 62 tested plants. Expression levels of TaCKX2.1 ranged from 0.69 to 2.01, the mean was 1.17 and only in three plants the expression level was below 0.79.
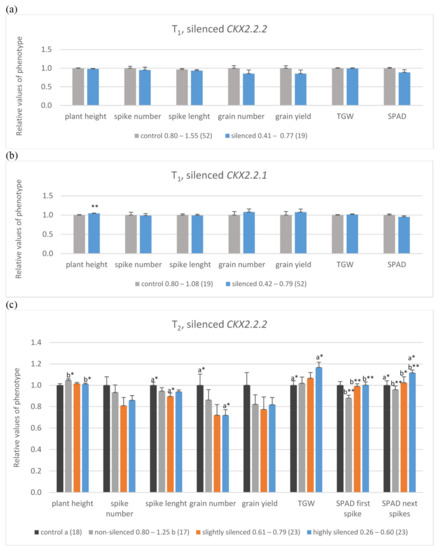
Figure 2.
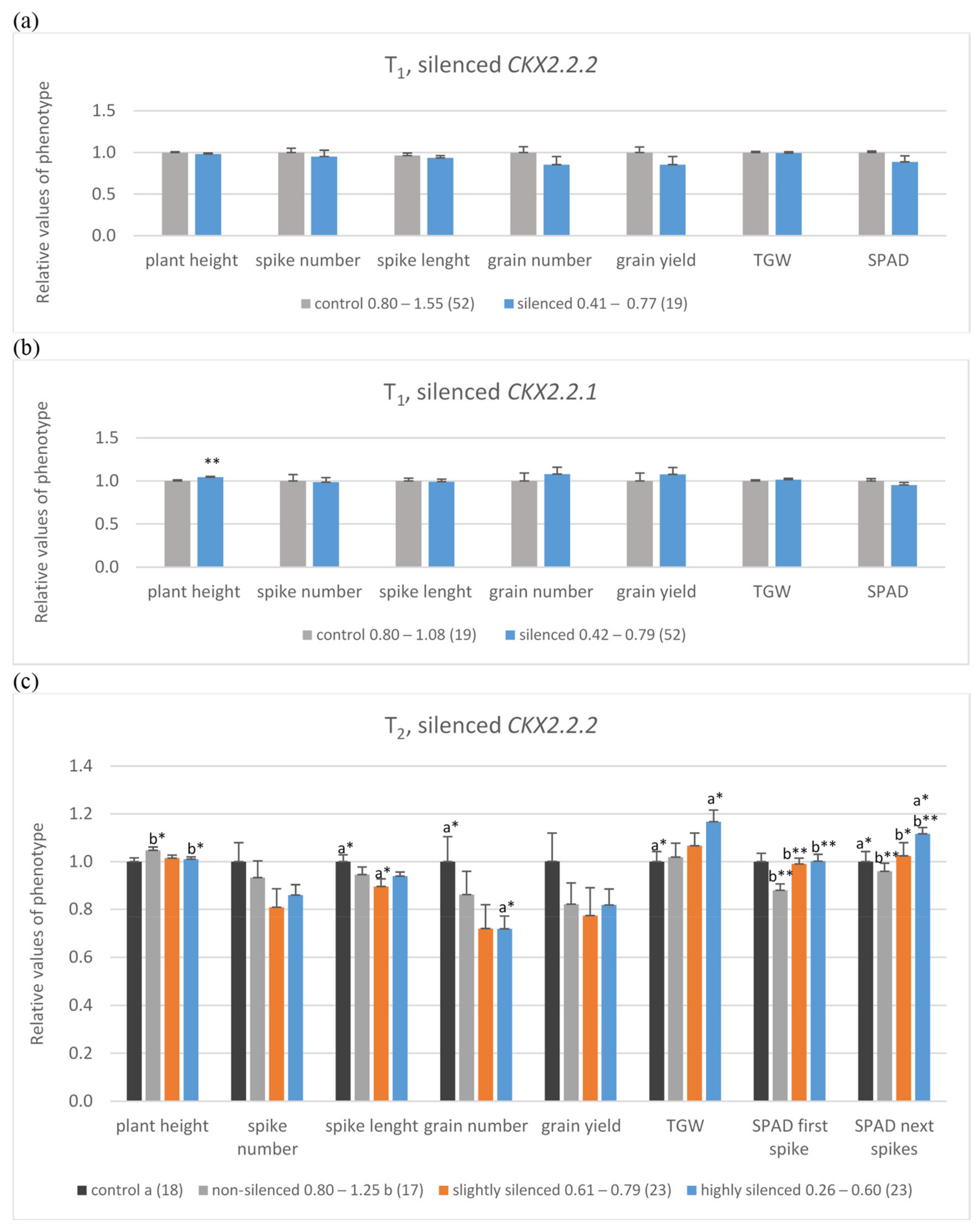
Mean, relative values of yield-related traits in control, non-silenced (grey bars), silenced (orange bars) or highly silenced (blue bars) plants of T1 with silenced TaCKX2.2.2 (a) and TaCKX2.2.1 genes (b) and T2 lines with silenced TaCKX2.2.2, including R2, control (black bars) (c). SPAD was measured on the flag leaves of the first (SPAD first spike) or four next spikes (SPAD next spikes). The error bars denote ± SE. Different letters mean significant differences: *—significant at p ≤ 0.05; **—significant at p ≤ 0.01.
2.2. Yield-Related Traits are Determined Depending on the Levels of Silencing of TaCKX2.2.1 and TaCKX2.2.2
The T1 silenced plants were divided into two groups. The first one included TaCKX2.2.2 silenced plants (level of expression 0.41 to 0.77; mean 0.68; Figure 2a, Supplemantary Table S1a), and the second contained TaCKX2.2.1 silenced plants (level of expression 0.42 to 0.79; mean 0.65; Figure 2b, Supplementary Table S1b). In both groups, expression of the second, respective TaCKX2.2 gene was silent at the level of 0.59 or slightly silenced (0.90) respectively. Data of yield-related traits in both groups of silenced plants were compared to those in non-silenced groups of plants with the expression level from 0.8 to above 1.00. In TaCKX2.2.2 silenced plants, spikes were shorter and data of plant height, spike number, spike length, grain number, grain yield, thousand grain weight (TGW), and chlorophyll content measured by SPAD in flag leaves of the 1st spike (SPAD 1st) were slightly lower compared to the non-transgenic control (Figure 2a, Supplementary Table S1a). Otherwise CKX activity was slightly higher (Supplementary Table S1a). Different data were obtained in TaCKX2.2.1 silenced plants (Figure 2b, Supplementary Table S1b). These plants were significantly taller and the values of spike length, grain number, grain yield and TGW were slightly higher compared to the non-transgenic control.
Among 65 T2 analyzed plants (Figure 2c, Supplementary Table S1c) the level of TaCKX2.2.2 expression in 25 of them was very low (0.26–0.60), in 23 of them was moderate (0.61–0.79) and in 17 of them it was from 0.80 to 1.25 (non-silenced plants). Additionally, 18 plants were not-silenced, R2 in-vitro control (second generation of in-vitro regenerated plants). Relative expression levels of TaCKX2.2.1 and TaCKX2.1 in these three groups of T2 plants were above 1.00 and relative CKX activity was slightly lower in the most silenced (0.94) and higher in non-silenced plants. Similarly, as in the case of T1 plants with silenced TaCKX2.2.2, the data of plant height, spike number, spike length and grain number were lower compared to the control group (Figure 2c, Supplementary Table S1c). However, the data of TGW and SPAD in the first and in the next spikes were significantly higher, especially in the first, most silenced group compared to the control.
2.3. Co-Expression of Silenced TaCKX2.2.2 with Other TaCKX GFMs in T2
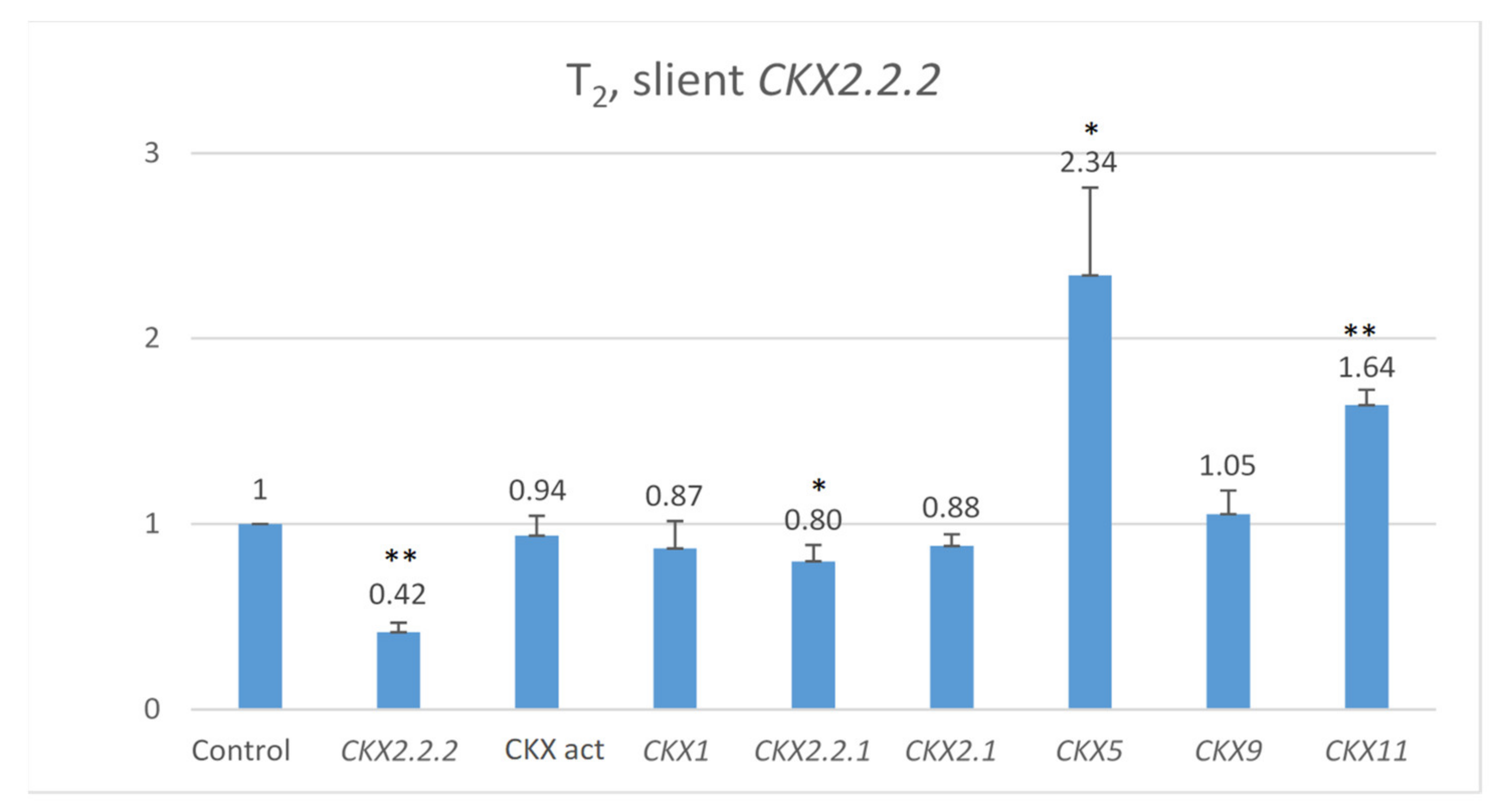
Co-expression of other genes was measured in 7 DAP spikes of seven T2 plants representing the highly silenced TaCKX2.2.2 (Figure 3). The mean level of silencing for TaCKX2.2.2 compared to the control (=1.00) was 0.42. Expression levels of TaCKX1, TaCKX2.1 and TaCKX9 were close to 1.00 (0.87, 0.88 and 1.05 respectively). For TaCKX2.2.1, the level was 0.80, and for two of them, TaCKX5 and TaCKX11, levels were much higher, reaching 2.34 and 1.64 respectively. CKX activity was slightly lower.
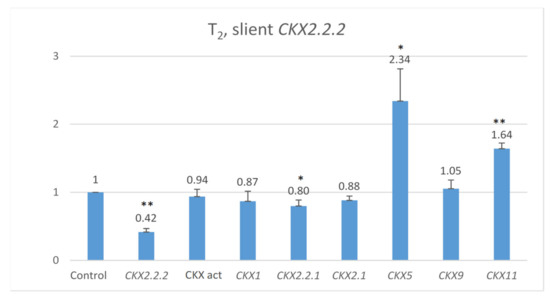
Figure 3.
Relative expression of other TaCKX GFMs in 7 DAP spikes, related to the control, non-silent TaCKX2.2.2 set as 1.00, and relative CKX enzyme activity (CKX act bar). The error bars denote ± SE. *—significant at p ≤ 0.05; **—significant at p ≤ 0.01.
2.4. Phytohormone Content in Highly Silenced TaCKX2.2.2 T2 Lines
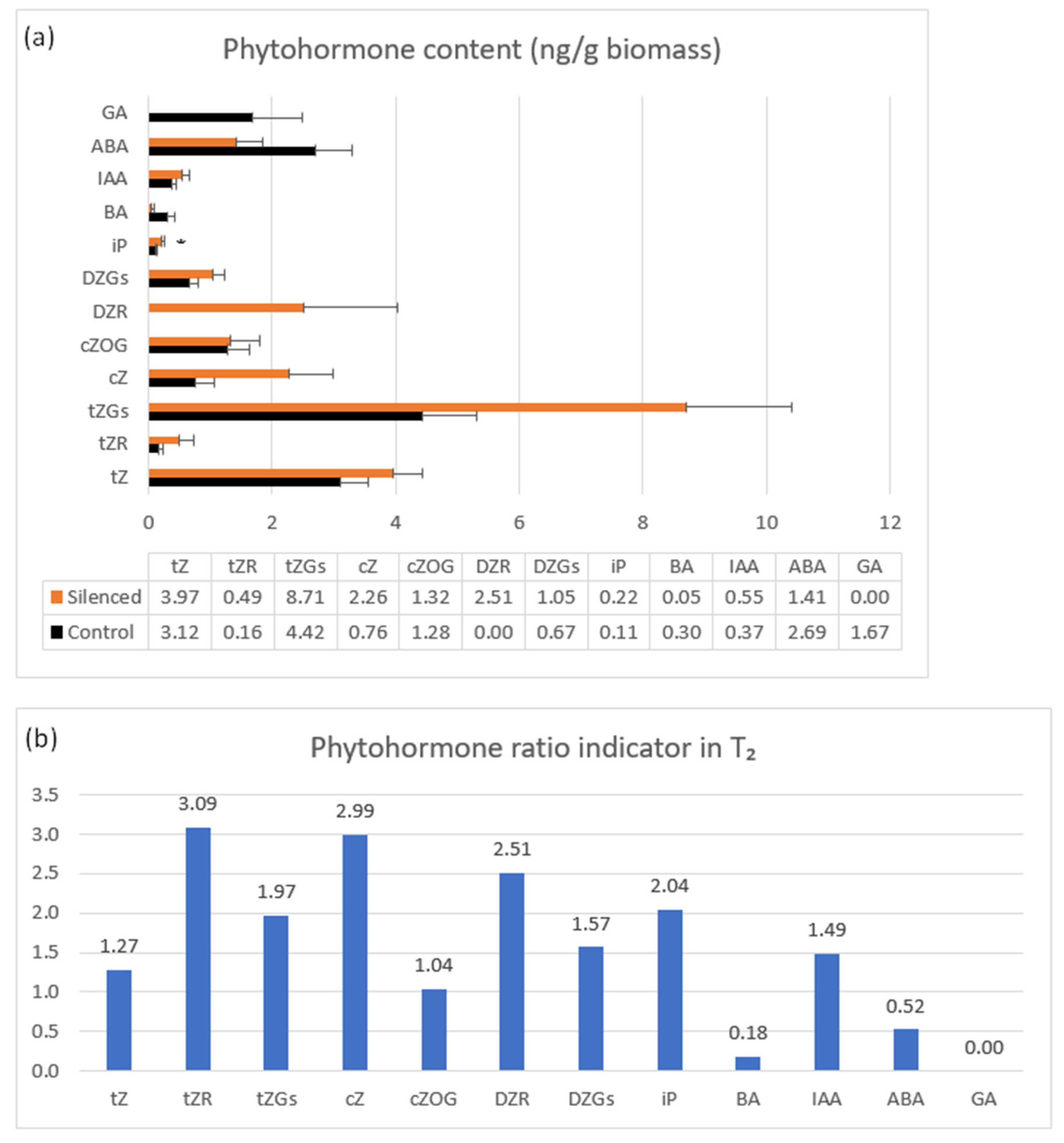
Phytohormone content was measured in 7 DAP spikes of seven T2 plants representing the highly silenced TaCKX2.2.2 gene with mean relative expression 0.42 (Figure 4a,b). The highest content in both silenced and non-silenced controls was observed for tZ and tZGs, cZ and cZOG, DZGs and ABA. These contents were even higher for almost all listed cytokinins in silenced plants, apart from BA (and ABA). A larger increase in content was noted for tZGs, cZ, and DZR. The latter was not present in the non-silenced control. Interestingly, BA and ABA contents in silenced plants were decreased and GA, detected only in non-silenced, control plants was reduced to 0.
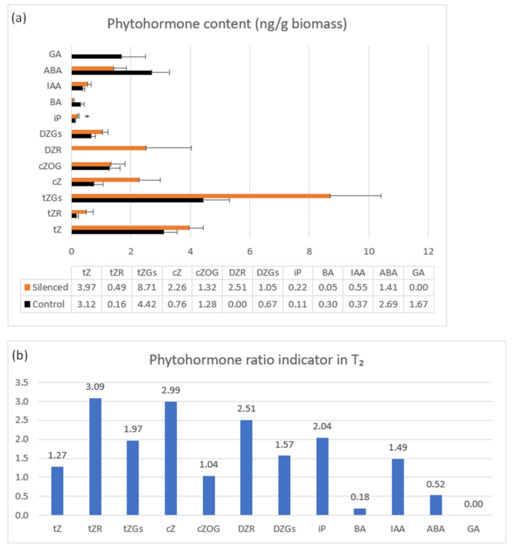
Figure 4.
Phytohormone content in control (black bars) and TaCKX2.2.2 silent T2 plants (a) and phytohormone ratio indicator (b). Not shown: small amounts (≤0.20 ng/g biomass): cZOGR, DZ7G, DZOGR, iP, iP7G, BA; trace amounts (≤0.02 ng/g biomass): cZR, cZ9G, DZ9G. Not detected: DZ, iPR, IBA, IPA, PAA. The error bars denote ± SE. * p ≤ 0.05.
Small amounts (≤0.20 ng/g biomass) were detected for DZOGR, DZ7G, cZOGR, IP7G; trace amounts (≤0.02 ng/g biomass) for: cZ9G, CZR DZ9G; and DZ, iPR, IBA, IPA, and PAA were not detected.
The phytohormone ratio indicator presented in Figure 4b is the mean value in silenced per mean value in non-silenced, control plants indicating a decrease or increase in content above the control level.
2.5. Co-Ordinated Regulation of CKX2.2.2 Silencing with Expression of Other TaCKX GFMs, Phytohormone Content and with Yield-Related Traits
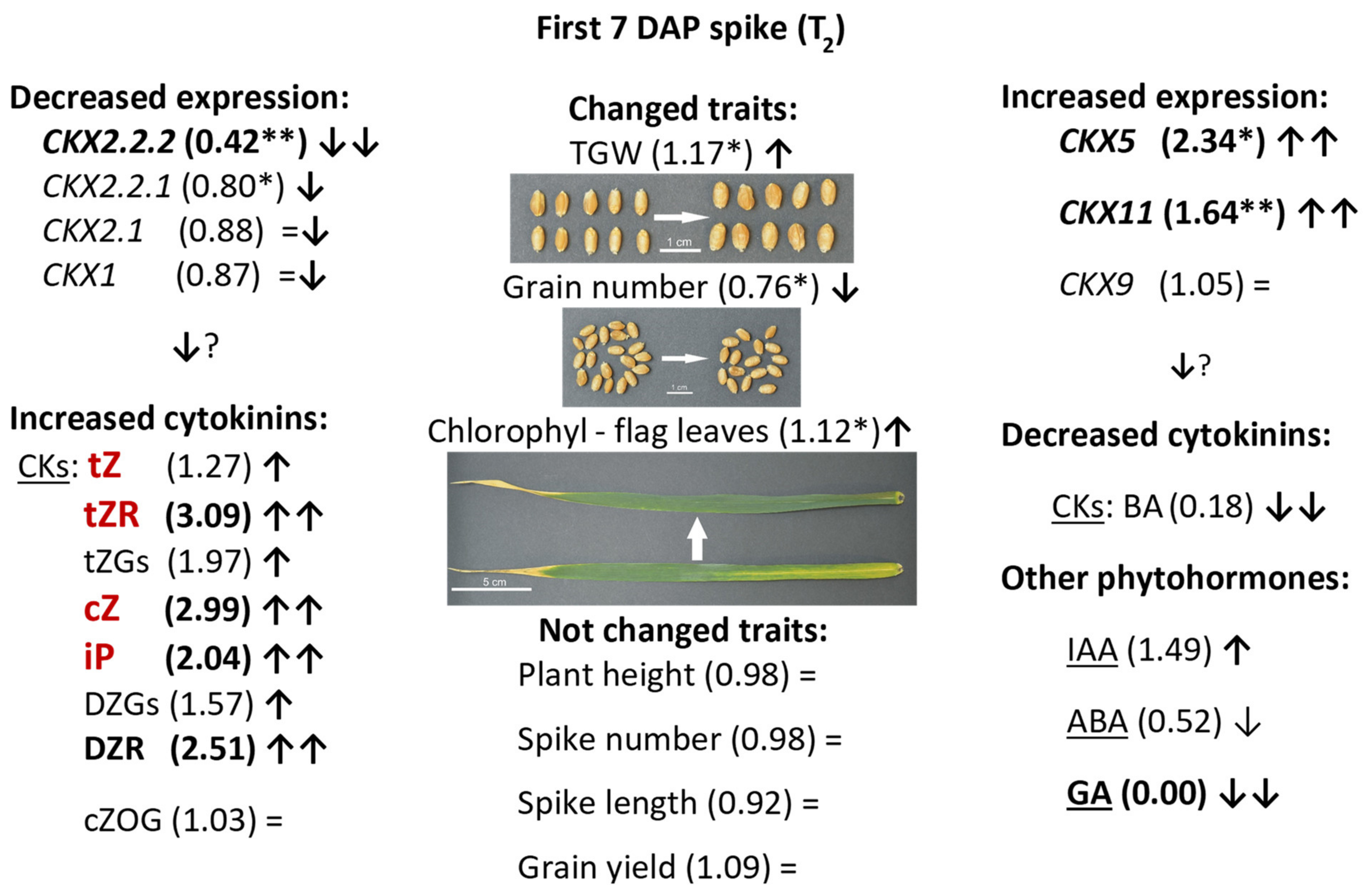
A schematic presentation of co-ordinated regulation of expression of TaCKX GFMs, phytohormone content and yield-related traits is presented in Figure 5. Significantly decreased expression of TaCKX2.2.2 (58% less than in control) in 7 DAP spikes of silenced T2 plants was co-ordinated with slightly decreased expression of TaCKX2.2.1 (20%) and highly increased expression of TaCKX5 and TaCKX11 (234% and 164% respectively). Relative expression levels of TaCKX1, TaCKX2.1, and TaCKX9 were close to the control (±5−13%). Most of isoprenoid cytokinin levels measured in seven DAP spikes of the same plants were increased. There were tZ (127%), tZGs (173%), tZR (203%), cZ (214%), DZGs (162%), DZR (251%), and iP (1.73%). cZOG content was not changed (103%). Content of aromatic cytokinin, BA was significantly reduced to 17% of control plants. The contents of other phytohormones were also different in silenced compared to control plants; IAA content was increased to 232% of the control, GA was not detected and ABA content reduced to 52% of control plants. All these changes significantly influenced three out of eight yield-related traits. TGW in silenced plants was significantly increased to 147%, grain number was reduced by 24% and flag leaf senescence was increased to 126%. Plant height, spike number and length, grain yield, and root weight were not changed compared to the control plants.
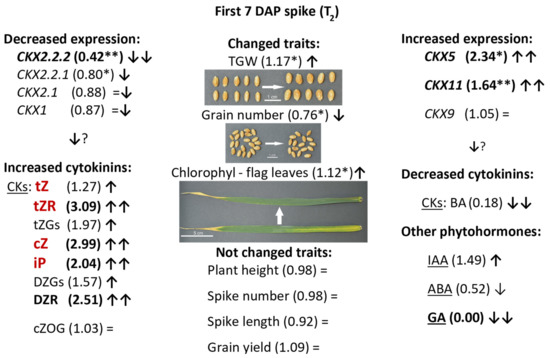
Figure 5.
Schematic presentation of up (↑) or down (↓) regulation of expression of TaCKX GFMs, phytohormone content and yield-related traits in 7 DAP spikes of silenced, T2 lines. Relative to the control = 1.00 values are in brackets. Highly decreased (↓↓) or highly increased (↑↑) values are in bold. *—significant at p ≤ 0.05; **—significant at p ≤ 0.01. CKs in red are substrates for CKX enzyme. =—not changed; ?—expected results.
Correlation coefficients between TaCKX GFMs and phytohormones are presented in Table 1. Down-regulated TaCKX1 and TaCKX2.2.1, unchanged TaCKX9 and up-regulated TaCKX5 expression are negatively correlated with tZR and IAA. Decreased expression of TaCKX2.1 is strongly and negatively correlated with DZR as well as with tZGs and iP. Down-regulated TaCKX2.2.2 is strongly, positively correlated with ABA, but up-regulated TaCKX11 is negatively correlated with ABA.

Table 1.
Correlation coefficients between TaCKX GFMs and phytohormones. Values in bold are significant at p < 0.05. = not changed; =↓ slightly silenced; ↓ silenced or down-regulated; ↓↓ highly silenced, significant; ↑ increased expression or up-regulated phytohormone.
2.6. Effect of TaCKX GFMs on Regulation of Phytohormone Content and Yield-Related Traits
The effect of TaCKX GFMs expression levels on regulation of phytohormone content and yield-related traits is based on correlation coefficients (cc; all collected in Supplementary Table S2) and presented in Figure 6.
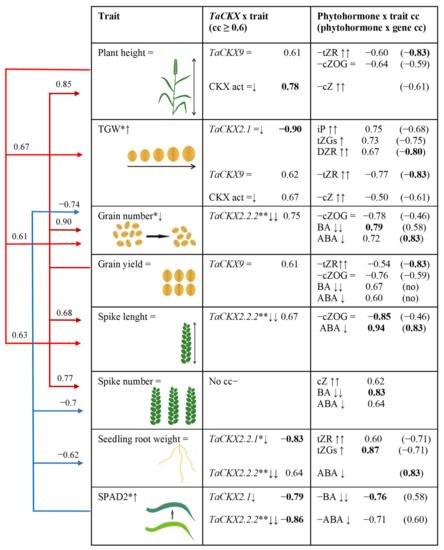
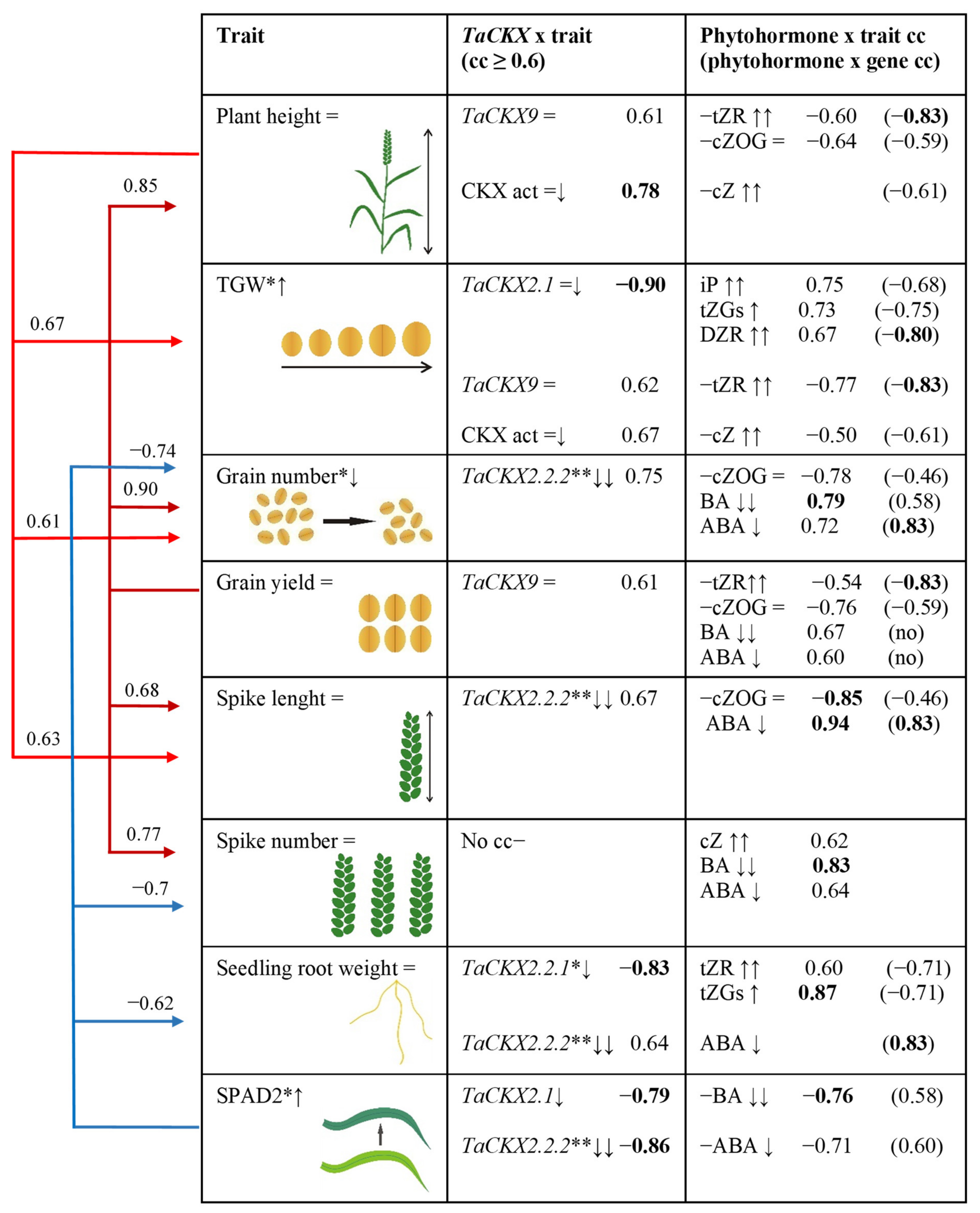
Figure 6.
Effect of increased (↑), decreased (↓) or unchanged (=) expression of TaCKX GFMs as well as phytohormones in 7 DAP spikes on yield-related traits based on correlation coefficients (cc). Red arrows—positive cc; blue arrows—negative cc. Values of cc in bold are significant at p < 0.05; no—no correlation; =↓ slightly silenced; ↓↓ highly silenced. *—significant at p ≤ 0.05; **—significant at p ≤ 0.01.
Plant height is in positive correlation with grain yield, TGW, spike length and grain number positively correlated with TaCKX9 in seven DAP spikes of T2 plants silenced for TaCKX2.2.2, and negatively with growing content of tZR and unchanged cZOG. Expression level of TaCKX9 negatively correlated with tZR and cZOG as well. Moreover, plant height was also correlated with CKX activity, which is negatively correlated with cZ content.
An increase in silenced plants’ TGW is positively correlated with plant height as well as SPAD2. The trait is strongly, negatively correlated with slightly decreased expression of TaCKX2.1 and positively with increased content of iP, tZGs and DZR. However, expression level of TaCKX2.1 is negatively correlated with these phytohormones. In contrast, TGW is positively correlated with TaCKX9 and CKX activity and negatively with tZR and cZ. The gene was positively correlated with tZR and CKX activity negatively correlated with cZ.
Grain number reduced in silenced plants is positively correlated with grain yield, plant height, spike number, and spike length (cc 0.90, 0.85, 0.77, and 0.68 respectively). The trait is positively correlated with strongly decreased expression of TaCKX2.2.2 and negatively with cZOG but positively with decreased contents of BA and ABA. TaCKX2.2.2 is also negatively correlated with cZOG but positively with ABA (cc 0.83) and BA (cc 0.58).
Grain yield is not changed in silenced compared to non-silenced plants. The trait is positively correlated with grain number (0.90), plant height (0.85), spike number (0.77), and spike length (0.68), and with TaCKX9, but negatively with tZR and cZOG. Correlation coefficients between TaCKX9 expression and both cytokinins were negative (−0.83 and −0.59). Also grain yield is positively correlated with reduced contents of BA and ABA.
Spike length is in positive correlation with grain number and yield, plant height and spike number. The trait is positively correlated with TaCKX2.2.2 and strongly, negatively with cZOG (cc-0.85) but strongly, positively with ABA (cc 0.94). Interestingly the gene is also negatively correlated with cZOG but strongly, positively correlated with ABA (cc 0.83).
Spike number, positively correlated with other traits such as grain number and yield, spike length and negatively with chlorophyll content measured in flag leaves (SPAD2). The trait was not correlated with expression of any gene, but was positively correlated with BA (cc 0.83), ABA (cc 0.64), and cZ (cc 0.62).
Seedling root weight, which is negatively correlated with SPAD2, was strongly, negatively regulated by TaCKX2.2.1 (cc −0.83) and positively with tZR and tZGs measured in seven DAP spikes. The gene is negatively correlated with both tZ metabolites. Root weight is also positively regulated by TaCKX2.2.2.
SPAD2 negatively correlated with spike number, grain number and root weight but positively with TGW. The trait is strongly, negatively correlated with TaCKX2.1 and TaCKX2.2.2 as well as with BA and ABA. Moreover, both genes positively correlated with the contents of both phytohormones.
3. Discussion
3.1. Phenotype Is Dependent on the Level of TaCKX2.2.2 and TaCKX2.2.1 Silencing
Silencing of particular genes essentially influenced the phenotype of wheat plants. Decreased expression of both, TaCKX2.2.2 and TaCKX2.2.1 by 41% in the T1 group of plants resulted in obtaining shorter plants and slightly lower parameters of yield-related traits. The opposite phenotype was obtained for plants, in which TaCKX2.2.1 expression was decreased and TaCKX2.2.2 expression was comparable to the control. These results indicate different functions of the two genes. Different phenotypes were also obtained with slightly silent (reduced by 30%) and highly silent (reduced by 51%) expression of TaCKX2.2.2 in T2. A similar dependence of yield-related traits on the level of silencing of another GFM, TaCKX1, has already been documented in our earlier work with the same genotype [11]. Different levels of silencing of particular gene resulted in up or down-regulation of expression of other TaCKX genes as well as appropriate phytohormone homeostasis (discussed below).
3.2. TaCKX2.2.2 Silencing Co-Ordinates Expression Levels of Other TaCKX GFMs and Phytohormone Homeostasis, Influencing Yield
A high level of TaCKX2.2.2 silencing (in T2) was coordinated with a slight decrease of TaCKX2.2.1 expression but a large increase of TaCKX5 and TaCKX11 expression. Otherwise, the effect of silencing of TaCKX1 coordinated expression of the same genes in the opposite way [11]. Silencing of both TaCKX2.2 GFMs, which are specifically expressed in developing spikes, caused slight silencing of TaCKX2.1 and TaCKX1 expressed in the same organ, as well. Increased expression was observed for TaCKX5, which is expressed in younger organs (from seedling roots to 0 DAP spikes) as well as TaCKX11 expressed in all organs up to 14 DAP [32]. Growth of expression of these genes in 7 DAP spikes of plants representing decreased expression of TaCKX2.2 genes specific for developing spikes might be a mechanism of maintaining CKX isozyme homeostasis, since total CKX activity was not changed. A similar result was observed in TaCKX1 silenced plants [11]. In these plants down-regulation of expression of TaCKX1 was coordinated with down-regulation of expression of TaCKX11 and up-regulation of TaCKX2.1 and TaCKX9, but total CKX activity was as in non-silenced plants.
As expected, in both cases of TaCKX2.2.2 and TaCKX1 silencing, most cytokinins and their conjugates in seven DAP spikes increased. However, there are significant differences in the contents of various forms. The main differences include highly increased content of tZ and DZ ribosides in TaCKX2.2.2 silenced compared to TaCKX1 silenced plants, in which contents of these ribosides were very small or not detected [11]. Other differences concerning CKs are a larger increase of cZ and glucosides of tZ and DZ in TaCKX2.2.2 silenced compared to TaCKX1 silenced plants, which in contrast were very rich in tZ7G and tZ9GOG. Both riboside forms and free base cZ are transporter metabolites, which when transported from roots to shoots control shoot traits [7,20]. Moreover, tZ riboside and cZ are degradation substrates of CKX [23,24]. Therefore, their growth might be the joint effect of transport as well as decreased CKX activity of specific isozyme encoded by silent CKX2.2.2. Glycosylation conjugates are deactivation products of free bases CKs, and might be converted back to free bases and ribosides during reactivation [24,25]. Their growth of content in seven DAP spikes of TaCKX2.2.2 silenced plants is possibly a result of maintaining/regulation of cytokinin homeostasis, to decrease intensive growth of free base tZ, cZ and their ribosides. Some of the TaGlu genes encoding β-glucosidases, the enzymes catalyzing the process of reactivation of O-glucosides to free bases and ribosides, were highly expressed in developing spikes of wheat, with the highest expression of TaGlu4 in 7 DAP spikes [26].
Other differences in the effect of silencing of TaCKX2.2.2 versus TaCKX1 were associated with changes in IAA and GA3 contents as well as phenotype. IAA in silenced TaCKX2.2.2 was 1.45 times higher, but GA content was reduced to zero. Conversely, IAA in silenced TaCKX1 plants was not changed and GA was over 10 times higher [11]. The opposite effect of coordinated expression in the case of both TaCKX2.2.2 and TaCKX1 as well as CKs, auxin, ABA and GA resulted in opposite phenotype changes. The CKX2.2.2 silenced plants were characterized by higher TGW, lower grain number, and higher chlorophyll content in flag leaves. The TaCKX1 silencing resulted in a higher spike number, grain number, and grain yield, but lower TGW [11]. The possible mechanisms of regulation of these phenotypic traits are discussed below.
3.3. Homeostatic Feedback Loop Between TaCKX Genes Transcription and Phytohormones
Expression of Arabidopsis AtCKX4 [41] activated by cytokinins was found to be down-regulated by IAA [42], underlining its importance in auxin-cytokinin crosstalk [43]. A similar feedback loop was observed in TaCKX2.2.2 silent wheat plants. Down-regulation of this gene together with TaCKX2.2.1 and TaCKX1 in developing spikes influenced growing content of active forms of CKs, mainly cZ and iP as well as ribosides of tZ and DZ. Such a high CKX gene specific affinity to cZ was also shown in Arabidopsis by AtCKX1 and AtCKX7 [44]. Moreover, iP and tZ were found as the main substrates of AtCKX4 [44], but it was associated with abiotic stress tolerance [42]. Activated by these cytokinins, highly expressed TaCKX5 showed a strong, negative correlation with IAA. As in the case of AtCKX4, TaCKX5 was highly expressed in younger organs (from seedling roots to 0 DAP spikes) but both TaCKX2.2 genes are expressed in developing spikes [32]. Therefore, these three TaCKX genes take part in maintaining a homeostatic loop between CKs and IAA in developing wheat plants. As reviewed by Cao et al. [45], auxin was proven to be a key regulator of different processes involved in seed development, seed size and weight. Seed development has been influenced by auxin level, their transport and signalling pathway and mediated by auxin homeostasis. IAA was also found to stimulate transport of photoassymilates to and within developing grains of wheat [46].
ABA level was positively correlated with TaCKX2.2.2 expression and negatively with cZOG. Therefore, the down-regulation of this gene, which is specifically expressed in developing spikes, was associated with decreased ABA and unchanged cZOG level. Moreover, high level of DZ riboside negatively correlated with slightly down-regulated TaCKX2.1 but positively with high level of tZ glucosides. High tZ riboside level was negatively correlated with TaCKX9 expressed through the younger organs of developing wheat plants.
Homeostatic balance of phytohormones in a selected organ is also regulated by many other processes. According to earlier reports, iP-type cytokinins might be converted to tZ-type cytokinins by cytochrome P450 mono-oxygenase proteins [43,47]. Both tZ and iP are also main, long-range transport forms. The tZ and its riboside are transported from root to shoot in the xylem, and iP is transported in the opposite direction through the phloem [48,49,50]. Possible consequences of the crosstalk between TaCKX genes and phytohormones in determination of yield-related traits are discussed below.
3.4. Regulation of Developmental Processes and Yield-Related Traits
Growing active CK content in 7 DAP spikes of TaCKX2.2.2 silenced plants, which are in the middle of the cell division/cell expansion stage [51,52], means that both processes of cell differentiation and cell division in the embryo promoted by CKs [53] are active. In contrast, content of one kind of gibberellin measured, GA3 was reduced to 0. GA and ABA are considered to be the main regulators of seed germination and dormancy, playing antagonistic roles during this process. GA enhances germination while ABA positively regulates the induction and maintenance of dormancy [54,55]. The induction of seed dormancy by ABA takes place in the third phase of wheat seed development [56]. The seven DAP spikes of TaCKX2.2.2 silenced plants showed decreased ABA content and GA3 content was not detected compared to non-silenced, control plants. These data might indicate a delay in seed germination and dormancy of silenced plants.
Silencing of TaCKX2.2.2 influenced three out of eight yield-related traits testedgrowth of TGW, reduction of grain number per plant and increase of chlorophyll content in flag leaves. Higher TGW and chlorophyll content has been negatively regulated by TaCKX2.1 expression as well, which in the case of TGW up-regulated contents of active cytokinins, iP and DZR, and glucosides of tZ. Moreover, TGW is positively regulated by TaCKX9, leading to increased tZR content. Interestingly, such an opposite effect to TaCKX2.1 in TaCKX1 silenced plants was maintained by TaCKX11, expressed in more developed spikes as well. Likewise, in the case of TaCKX1 silencing [11], total CKX activity was not changed, but the proportions of specific isozymes are expected to be modified. These modifications of pulled isozymes might be responsible for highly increased cZ content. The opposite effect of TaCKX2.1 on TGW and chlorophyll content in flag leaves was documented in our earlier report, in which silencing of TaCKX1 resulted in lower TGW and SPAD values, which was regulated by increased expression of TaCKX2.1 [11]. Therefore, we confirmed our previous results that TaCKX2.1 in co-operation with other TaCKX GFMs downregulates TGW and with TaCKX2.2.2 chlorophyll content in flag leaves. Similar correlations with grain size, weight and grain filling rate were reported for the allele of TaCKX6a02 [35], which is TaCKX2.1–3D [12]. Opposed data were reported for other TaCKX2 GFM, TaCKX6 [36], renamed as TaCKX2.2.1–3D [12]. An 18-bp deletion in intron 2 of the gene resulted in its decreased expression, which was associated with a greater TGW, compared to its wild-type haplotype. However, these and our data are not well comparable because TaCKX2.2.1 in our research is only slightly decreased and nature of these genetic modifications in both genomes are different. The results of Zhang et al. [36] refer specifically to the mutation of one homoeologous gene, and in our case all three homoeologous genes were slightly downregulated.
The changes observed by us in phytohormonal homeostasis are consistent with the obtained phenotype of higher TGW. Lack of GA and decreased ABA content in 7 DAP spikes of TaCKX2.2.2 silenced plants should have resulted in delay of grain germination and effect on grain filling, which in consequence led to higher TGW and chlorophyll content in flag leaves. This result was proved by the opposite effect in TaCKX1 silenced plants, showing ten times increased GA content. Moreover, growth of TGW in TaCKX2.2.2 silenced plants was positively correlated with iP content, which was higher than in TaCKX1 silenced plants. As reported, [56] decrease in the level of this phytohormone was associated with seed maturation in wheat. Therefore, increased iP is expected to delay this process in TaCKX2.2.2 silenced plants, which in turn resulted in higher TGW. It was also documented that tZ and other CKs are implicated in the control of seed dormancy induction and maintenance, and IAA level is decreased during seed maturation [56,57]. As suggested by Fahy et al. [58], final grain weight is largely determined by developmental processes prior to grain filling.
The role of CKs in maintaining chlorophyll level in cereals has already been reported [8,59,60]. Down-regulation of TaCKX2.1 and TaCKX2.2.2 in 7 DAP spikes of TaCKX2.2.2 silenced plants resulted in down-regulation of BA and ABA, lack of GA3 and up-regulation of free base isoprenoid cytokinins, cZ, tZ, iP. These changes positively influenced chlorophyll content in flag leaves of silenced plants. In contrast, up-regulation of TaCKX2.1 in TaCKX1 silenced plants was correlated with a slight increase of tZ, iP and cZ content and a very large increase of GA3, which resulted in lower chlorophyll content/early senescence of flag leaves [11]. It has already been documented that premature leaf senescence resulted in lower grain yield [59] and increased CK during seed development might improve chlorophyll content and determine grain number [61,62]. Moreover, ABA was reported to play an important role in leaf senescence as well [63,64]. The longer stay-green characteristic of flag leaves is expected to influence longer grain filling and higher TGW, since during further seed development nutrient compounds and phytohormones are transported from flag leaves to the seeds [38,60]. Similar results as for TaCKX2.2.2 and TaCKX1 silent plants of wheat were reported for knock-out OsCKX11 in rice, which was significantly induced by ABA [38]. Increase of CK levels, mainly tZ and cZ was correlated with reduction in ABA levels determining delayed leaf senescence and increased branch, tiller, and grain number. Therefore, both TaCKX genes and OsCKX11 function antagonistically between CKs and ABA in leaf senescence of wheat and rice.
Decreased TaCKX2.2.2 expression resulted in higher TGW but reduced grain number. The same gene also participates in regulation of spike length and root weight and, in the opposite way, in chlorophyll content of flag leaves. The gene is strongly, positively correlated with ABA, involved in regulation of all these traits. As would be expected, grain number, grain yield, and spike number are oppositely regulated to chlorophyll content by BA and ABA. Moreover, cZOG, negatively correlated with TaCKX2.2.2, is involved in strong, negative regulation of grain number and spike length but positively with spike number. The latter trait is also importantly regulated by BA and ABA as well as cZ. Therefore, the spike number is up-regulated by cZ and down-regulated by BA, which is co-ordinated by ABA. The content of ABA in seven DAP spikes, which is the milk/early dough stage of wild-type wheat plants was low, about 2.6 ng/g biomass [11] and decreased by 50% in TaCKX2.2.2 silenced plants. Since the highest concentrations of ABA were detected in the late dough stage [13], we may suppose that the lower level of ABA in seven DAP spikes resulted in extending chlorophyll content in flag leaves and also the maturation time of the spikes, which led to higher TGW. High content of cZ was observed during early embryo development of barley, underlying its important role in this stage of development [57]. cZ has also been detected in other cereal crop species such as rice [65] and maize [66], but its role in spike development was not known. Similarly, as in maize, the cZ of wheat was preferentially glycosylated to O-glucoside. The content of this conjugate was negatively correlated with grain number, grain yield, spike length and plant height. Since the content of cZOG was not changed, and only grain number was reduced, it could be positively affected by decreasing BA. A positive correlation of BA with grain number, grain yield, and spike number and negative with chlorophyll content underline its role in posttranscriptional or posttranslational regulation of protein abundance involved in ABA biosynthesis and response [67]. Again, we are not able to confirm that silencing by RNAi of expression of TaCKX2.2.1–3A (originally TaCKX2.4) determines grain number per spike [34]. Such a differences might be dependent on individual genetic background, which modulates co-operation of other TaCKX GFMs or other regulatory genes, existing in that background, which take part in establishing hormonal homeostasis.
TaCKX9 is one more gene which coordinates development of TGW as well as plant height and grain yield in TaCKX2.2.2 silenced plants. The gene is expressed in seedling roots as well as younger developing organs, showing very high expression in leaves [32], and is co-expressed positively with similarly expressed TaCKX5 [33]. We revealed a positive correlation of TaCKX9 expression with the riboside of tZ. Since tZR belongs to transporter metabolites [20], and the gene is expressed through the developing plant, we might predict its role in coordination of other genes involved in transport of tZR from roots to developing spikes. Indeed, there is a positive correlation between root weight and tZR and a negative correlation between plant height/TGW/grain yield and tZR measured in seven DAP spikes.
4. Materials and Methods
4.1. Plant Material
The spring wheat (Triticum aestivum L.) cultivar Kontesa, which is susceptible to Agrobacterium-mediated transformation and in vitro regeneration, was used in all experiments.
4.2. Vector Construction
The binary vector pBract207 (https://www.jic.ac.uk/technologies/crop-transformation-bract/, accessed on 10 March 2021) containing the hpRNA type of silencing cassette was assembled via Gateway cloning. In the first step, the 385 bp fragment of a coding sequence of TaCKX2.2.2–3D (FJ648070, 1259., 1643; TraesCS3D02G143300) was amplified using the primers: forward 5′- CTGGCTCAACCTGTTCCTC-3′, reverse 5′- ATACTTCCTCTTCCGATCCACG-3′ and cloned into the entry vector pCR8/GW/TOPO (Invitrogen, Karlsbad, CA, USA). This fragment has 93%–94.3% identity with the same fragments of TaCKX2.2.1, located on 3A, 3B, and 3D chromosomes and 87.9%–92.3% identity with TaCKX2.1, located on 3A, 3B, and 3D chromosomes as well (Supplementary Table S3). Next, the TaCKX2.2.2 insert from the entry vector was cloned to the pBract207 destination vector in a sense and antisense orientation using LR Clonase (Invitrogen) according to the manufacturer’s protocol. The correct assembly of the silencing cassette in the pBract207 vector was verified by restriction enzyme analysis and Sanger sequencing. The vector was then electroporated into the AGL1 strain of Agrobacterium tumefaciens and used for transformation.
4.3. Plant Transformation and Selection of Transgenic Lines
Donor plants for transformation experiments were grown in growth chambers under controlled environmental conditions at 20 °C day and 18 °C night temperatures, and a 16 h light/8 h dark photoperiod. Light intensity of 350 μmol/m2/s was provided by fluorescent lamps. Agrobacterium-mediated transformation of immature embryos and in vitro plant regeneration were performed according to our previously developed protocols for wheat [68,69]. Putative transgenic events as well as T1 plants were verified by PCR screening for the presence of a T-DNA fragment using primers specific to the hpt gene. PCR screening was performed for T0 plants as well as 14-day old T1 seedlings using the KAPA3G Plant PCR Kit (Kapa Biosystems, Wilmington, MA, USA). The following primers were used for the amplification of a 405 bp fragment of the hpt gene: forward 5′-ATGACGCACAATCCCACTATCCT-3′ and reverse 5′-AGTTCGGTTTCAGGCAGGTCTT-3′. The reaction was carried out in a final volume of 50 µl containing 1X KAPA Plant PCR Buffer, 0.3 μM of each primer, and 1 U of KAPA3G Plant DNA Polymerase. A 0.5 × 0.5 mm fragment of leaf tissue was used as a template. The PCR was run with the following temperature profile: an initial denaturation step at 95 °C for 3 min.; 40 cycles of 95 °C for 20 s, 68 °C for 30 s, 72 °C for 30 s, and the final extension step at 72 °C for 2 min. The PCR products were separated and visualized on agarose gels. Respective generation of control, in-vitro plants was applied in verification by PCR screening. Non-transgenic null segregants were used as control samples in all experiments.
4.4. Quantitative RT-qPCR
Total RNA from was extracted from the middle part of 7 DAP (days after pollination) spikes using TRI Reagent (Sigma-Aldrich, St. Louis, MO, USA) according to the manufacturer’s protocol. Purification of RNA and cDNA synthesis was performed as described by Ogonowska et al. [32]. RT-qPCR analysis was done for seven target genes: TaCKX1 (JN128583), TaCKX2.1 (JF293079), TaCKX2.2.1 (FJ648070), TaCKX2.2.2 (GU084177), TaCKX11 (former TaCKX3) (JN128587), TaCKX5 [32] and TaCKX9 (former TaCKX10) (JN128591). Sequences of specific primers designed for the genes tested are shown in Supplementary Table S4. ADP-ribosylation factor (Ref2) (AB050957) was used as a reference gene in all qPCR reactions. qPCR was carried out in a 10-µl mixture containing 1x HOT FIREPol EvaGreen qPCR Mix Plus (no ROX) (Solis BioDyne, Tartu, Estonia), 0.2 µM of each primer and 4 µL of 20 x diluted cDNA. The following temperature profile was used: an initial denaturation/polymerase activation step of 95 °C for 12 min; 45 cycles of amplification at 95 °C for 20 s, 62 °C for 20 s, 72 °C for 20 s; final denaturation step at 72 °C for 5 min and the melting curve profile from 72 °C to 99 °C, with the temperature rising 1 °C at each step and continuous fluorescence measurements. The expression levels of TaCKX GFMs were calculated from three technical replicates (repeated samples in RT-qPCR reaction) according to the standard curve method using the Ref2 gene as a normalizer [70]. The relative expression levels in transgenic/silenced plants were calculated as x-fold of 1 based on mean expression value in null segregants, which was set to 1. The analysis was performed for three biological replicates.
Experimental plants were divided into three groups depending on the level of silencing: ≤0.60 as highly silenced, 0.61–0.79 as silenced and ≥0.80 as non-silenced. This three-group scale is used to find if there was dependence between expression level of silenced genes and phenotype.
4.5. CKX Activity Assays
Cytokinin oxidase/dehydrogenase activity was measured in 7 DAP spikes of T1/T2 plants as described previously [9,33].
4.6. Quantification of ABA, Auxins, Cytokinins and GA3
For quantification, the standards of phytohormones were used: ABA, five auxins: indole-3-acetic acid (IAA), indole-3-butyric acid (IBA), indole-3-propionic acid (IPA), 1-naphthaleneacetic acid (NAA), and 2-phenylacetic acid (PAA); twenty-seven standards of CKs: trans-zeatin (tZ), trans-zeatin riboside (tZR), trans-zeatin-9-glucoside (tZ9G), trans-zeatin-7-glucoside (tZ7G), trans-zeatin-O-glucoside (tZOG), trans-zeatin riboside-O-glucoside (tZROG), trans-zeatin-9-glucoside-O-glucoside (tZ9GOG), trans-zeatin-9-glucoside riboside (tZ9GR), cZ, cis-zeatin-riboside (cZR), cis-zeatin O-glucoside (cZOG), cis-zeatin 9-glucoside (cZ9G), cis-zeatin-O-glucoside-riboside (cZROG), dihydrozeatin (DZ), dihydrozeatin-riboside (DZR), dihydrozeatin-9-glucoside (DZ9G), dihydrozeatin-7-glucoside (DZ7G), dihydrozeatin-O-glucoside (DZOG), dihydrozeatin riboside-O-glucoside (DZROG), N6-(Δ2-isopentenyl)adenine (iP), N6-isopentenyladenosine (iPR), N6-isopentenyladenosine-7-glucoside (iP7G), para-topolin (pT), meta-topolin (mT), ortho-topolin (oT), 6-benzylaminopurine (6-BAP), and GA3.
The procedure of phytohormone quantification is described in Jablonski et al. [11].
4.7. Analysis of Phenotypic Traits
The following parameters were determined after harvesting mature plants: plant height, spike length, number of spikes per plant, number of grains per plant, weight of grains per plant and 1000 grain weight. Additionally, the weight of fresh mass of roots from 5-day old seedlings was determined in T2 lines. Seeds were germinated on wet glass beads in Petri dishes for 5 days in the dark, at room temperature. The excess of water in the roots was removed by placing them on the filter paper and then, the roots were immediately cut from the seedlings and weighed. Chlorophyll content was measured in the flag leaves from the first five tillers with spikes, periodically from 0 DAP up to leaf desiccation using a SPAD chlorophyll meter (Konica Minolta SPAD-502Plus). All measurements were performed for six biological replicates.
4.8. Statistical Analysis
All statistical analyses were performed using Statistica v13.3 software (StatSoft, Kraków, Poland) and the mean ± SE was presented. The normality of the tested samples was verified by the Shapiro–Wilk test. Differences between tested and control samples in phytohormone content, phenotypic traits and expression levels were verified by Student’s t-test or the Mann–Whitney test at p < 0.01 and p < 0.05 confidence levels. Correlation coefficients of analyzed traits were verified by either the parametric Pearson test or the nonparametric Spearman test.
5. Conclusions
We documented that silencing of one of the TaCKX GFMs co-ordinates the expression of other genes from the family. This leads to a homeostatic feedback loop between the transcription of TaCKX genes and phytohormones and establishing phytohormonal homeostasis. Tested TaCKX genes showed specific correlations to the forms/metabolites of CKs and are involved in the regulation of other phytohormones. The TaCKX2.2 genes strongly modulate yield-related traits of wheat by cross-talk with other TaCKX GFMs and establishing homeostasis of phytohormones in developing wheat plants.
Supplementary Materials
The Supplementary Materials are available online at https://www.mdpi.com/article/10.3390/ijms22084142/s1.
Author Contributions
Conceptualization, A.N.-O. and W.O.; methodology, B.J., K.S., M.P., S.G., and A.B.; software, K.S.; validation, B.J., A.N.-O., and K.S.; formal analysis, B.J. and A.N.-O.; investigation, B.J., K.S., M.P., and M.C.; data curation, B.J. and A.N.-O.; writing—original draft preparation, A.N.-O. and W.O.; writing—review and editing, A.N.-O.; visualization, A.N.-O. and B.J.; supervision, A.N.-O. and W.O.; project administration, A.N.-O.; funding acquisition, A.N.-O. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Science Centre, Poland grant No. UMO-2014/13/B/NZ9/02376, grant No. UMO-2020/37/B/NZ9/00744 and a statutory grant of PBAI-NRI.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Acknowledgments
We thank Malgorzata Wojciechowska, Izabela Skuza, Agnieszka Glowacka, Maja Boczkowska and Agnieszka Onysk for excellent technical assistance.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Shewry, P.R.; Hey, S.J. The Contribution of Wheat to Human Diet and Health. Food Energy Secur. 2015, 4, 178–202. [Google Scholar] [CrossRef]
- Kurek, M.A.; Wyrwisz, J.; Karp, S.; Wierzbicka, A. Particle Size of Dietary Fiber Preparation Affects the Bioaccessibility of Selected Vitamin B in Fortified Wheat Bread. J. Cereal Sci. 2017, 77, 166–171. [Google Scholar] [CrossRef]
- Venske, E.; dos Santos, R.S.; Busanello, C.; Gustafson, P.; de Oliveira, A.C. Bread Wheat: A Role Model for Plant Domestication and Breeding. Hereditas 2019, 156, 16. [Google Scholar] [CrossRef]
- Foley, J.A.; Ramankutty, N.; Brauman, K.A.; Cassidy, E.S.; Gerber, J.S.; Johnston, M.; Mueller, N.D.; O’Connell, C.; Ray, D.K.; West, P.C.; et al. Solutions for a Cultivated Planet. Nature 2011, 478, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Nadolska-Orczyk, A.; Rajchel, I.K.; Orczyk, W.; Gasparis, S. Major Genes Determining Yield-related Traits in Wheat and Barley. Theor. Appl. Genet. 2017, 130, 1081–1098. [Google Scholar] [CrossRef]
- Kieber, J.J.; Schaller, G.E. Cytokinin Signaling in Plant Development. Development 2018, 145, 7. [Google Scholar] [CrossRef] [PubMed]
- Sakakibara, H. Cytokinin Biosynthesis and Transport for Systemic Nitrogen Signaling. Plant J. 2020, 105, 421–430. [Google Scholar] [CrossRef]
- Jameson, P.E.; Song, J.C. Cytokinin: A Key Driver of Seed Yield. J. Exp. Bot. 2016, 67, 593–606. [Google Scholar] [CrossRef]
- Zalewski, W.; Galuszka, P.; Gasparis, S.; Orczyk, W.; Nadolska-Orczyk, A. Silencing of the HvCKX1 Gene Decreases the Cytokinin Oxidase/Dehydrogenase Level in Barley and Leads to Higher Plant Productivity. J. Exp. Bot. 2010, 61, 1839–1851. [Google Scholar] [CrossRef]
- Zalewski, W.; Gasparis, S.; Boczkowska, M.; Rajchel, I.K.; Kala, M.; Orczyk, W.; Nadolska-Orczyk, A. Expression Patterns of HvCKX Genes Indicate Their Role in Growth and Reproductive Development of Barley. PLoS ONE 2014, 9, e115729. [Google Scholar] [CrossRef]
- Jablonski, B.; Ogonowska, H.; Szala, K.; Bajguz, A.; Orczyk, W.; Nadolska-Orczyk, A. Silencing of TaCKX1 Mediates Expression of Other TaCKX Genes to Increase Yield Parameters in Wheat. Int. J. Mol. Sci. 2020, 21, 4809. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Zhao, J.Q.; Song, J.C.; Jameson, P.E. Cytokinin Dehydrogenase: A Genetic Target for Yield Improvement in Wheat. Plant Biotechnol. J. 2020, 18, 614–630. [Google Scholar] [CrossRef]
- Nguyen, H.N.; Perry, L.; Kisiala, A.; Olechowski, H.; Emery, R.J.N. Cytokinin Activity during Early Kernel Development Corresponds Positively with Yield Potential and Later Stage ABA Accumulation in Field-grown Wheat (Triticum aestivum L.). Planta 2020, 252, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Chen, L.; Herrera-Estrella, L.; Cao, D.; Tran, L.P. Altering Plant Architecture to Improve Performance and Resistance. Trends Plant Sci. 2020, 25, 1154–1170. [Google Scholar] [CrossRef] [PubMed]
- Waldie, T.; Leyser, O. Cytokinin Targets Auxin Transport to Promote Shoot Branching. Plant Physiol. 2018, 177, 803–818. [Google Scholar] [CrossRef]
- Li, W.Q.; Herrera-Estrella, L.; Tran, L.S.P. The Yin-Yang of Cytokinin Homeostasis and Drought Acclimation/Adaptation. Trends Plant Sci. 2016, 21, 548–550. [Google Scholar] [CrossRef]
- Werner, T.; Nehnevajova, E.; Kollmer, I.; Novak, O.; Strnad, M.; Kramer, U.; Schmulling, T. Root-Specific Reduction of Cytokinin Causes Enhanced Root Growth, Drought Tolerance, and Leaf Mineral Enrichment in Arabidopsis and Tobacco. Plant Cell 2010, 22, 3905–3920. [Google Scholar] [CrossRef]
- Nishiyama, R.; Watanabe, Y.; Fujita, Y.; Le, D.T.; Kojima, M.; Werner, T.; Vankova, R.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Kakimoto, T.; et al. Analysis of Cytokinin Mutants and Regulation of Cytokinin Metabolic Genes Reveals Important Regulatory Roles of Cytokinins in Drought, Salt and Abscisic Acid Responses, and Abscisic Acid Biosynthesis. Plant Cell 2011, 23, 2169–2183. [Google Scholar] [CrossRef] [PubMed]
- Ko, D.K.; Brandizzi, F. Network-based Approaches for Understanding Gene Regulation and Function in Plants. Plant J. 2020, 104, 302–317. [Google Scholar] [CrossRef]
- Hirose, N.; Takei, K.; Kuroha, T.; Kamada-Nobusada, T.; Hayashi, H.; Sakakibara, H. Regulation of Cytokinin Biosynthesis, Compartmentalization and Translocation. J. Exp. Bot. 2008, 59, 75–83. [Google Scholar] [CrossRef]
- Spichal, L. Cytokinins–Recent News and Views of Evolutionally Old Molecules. Funct. Plant Biol. 2012, 39, 267–284. [Google Scholar] [CrossRef]
- Bajguz, A.; Piotrowska, A. Conjugates of Auxin and Cytokinin. Phytochemistry 2009, 70, 957–969. [Google Scholar] [CrossRef] [PubMed]
- Sakakibara, H. Cytokinins: Activity, Biosynthesis, and Translocation. Annu. Rev. Plant Biol. 2006, 57, 431–449. [Google Scholar] [CrossRef] [PubMed]
- Hosek, P.; Hoyerova, K.; Kiran, N.S.; Dobrev, P.I.; Zahajska, L.; Filepova, R.; Motyka, V.; Muller, K.; Kaminek, M. Distinct Metabolism of N-glucosides of Isopentenyladenine and Trans-zeatin Determines Cytokinin Metabolic Spectrum in Arabidopsis. New Phytol. 2020, 225, 2423–2438. [Google Scholar] [CrossRef]
- Brzobohaty, B.; Moore, I.; Kristoffersen, P.; Bako, L.; Campos, N.; Schell, J.; Palme, K. Release of Active Cytokinin by a Beta-Glucosidase Localized to the Maize Root-Meristem. Science 1993, 262, 1051–1054. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Jiang, L.; Jameson, P.E. Co-ordinate Regulation of Cytokinin Gene Family Members during Flag Leaf and Reproductive Development in Wheat. BMC Plant Biol. 2012, 12, 78. [Google Scholar] [CrossRef] [PubMed]
- Hou, B.K.; Lim, E.K.; Higgins, G.S.; Bowles, D.J. N-glucosylation of Cytokinins by Glycosyltransferases of Arabidopsis thaliana. J. Biol. Chem. 2004, 279, 47822–47832. [Google Scholar] [CrossRef] [PubMed]
- Zalabak, D.; Galuszka, P.; Mrizova, K.; Podlesakova, K.; Gu, R.L.; Frebortova, J. Biochemical Characterization of the Maize Cytokinin Dehydrogenase Family and Cytokinin Profiling in Developing Maize Plantlets in Relation to the Expression of Cytokinin Dehydrogenase Genes. Plant Physiol. Biochem. 2014, 74, 283–293. [Google Scholar] [CrossRef]
- Zalewski, W.; Orczyk, W.; Gasparis, S.; Nadolska-Orczyk, A. HvCKX2 Gene Silencing by Biolistic or Agrobacterium-mediated Transformation in Barley Leads to Different Phenotypes. BMC Plant Biol. 2012, 12, 206. [Google Scholar] [CrossRef]
- Ashikari, M.; Sakakibara, H.; Lin, S.Y.; Yamamoto, T.; Takashi, T.; Nishimura, A.; Angeles, E.R.; Qian, Q.; Kitano, H.; Matsuoka, M. Cytokinin Oxidase Regulates Rice Grain Production. Science 2005, 309, 741–745. [Google Scholar] [CrossRef]
- Mameaux, S.; Cockram, J.; Thiel, T.; Steuernagel, B.; Stein, N.; Taudien, S.; Jack, P.; Werner, P.; Gray, J.C.; Greenland, A.J.; et al. Molecular, Phylogenetic and Comparative Genomic Analysis of the Cytokinin Oxidase/Dehydrogenase Gene Family in the Poaceae. Plant Biotechnol. J. 2012, 10, 67–82. [Google Scholar] [CrossRef] [PubMed]
- Ogonowska, H.; Barchacka, K.; Gasparis, S.; Jablonski, B.; Orczyk, W.; Dmochowska-Boguta, M.; Nadolska-Orczyk, A. Specificity of Expression of TaCKX Family Genes in Developing Plants of Wheat and Their Co-operation within and among Organs. PLoS ONE 2019, 14, e0214239. [Google Scholar] [CrossRef] [PubMed]
- Szala, K.; Ogonowska, H.; Lugowska, B.; Zmijewska, B.; Wyszynska, R.; Dmochowska-Boguta, M.; Orczyk, W.; Nadolska-Orczyk, A. Different Sets of TaCKX Genes Affect Yield-related Traits in Wheat Plants Grown in a Controlled Environment and in Field Conditions. BMC Plant Biol. 2020, 20, 496. [Google Scholar] [CrossRef]
- Li, Y.L.; Song, G.Q.; Gao, J.; Zhang, S.J.; Zhang, R.Z.; Li, W.; Chen, M.L.; Liu, M.; Xia, X.C.; Risacher, T.; et al. Enhancement of Grain Number per Spike by RNA Interference of Cytokinin Oxidase 2 Gene in Bread Wheat. Hereditas 2018, 155, 33. [Google Scholar] [CrossRef]
- Lu, J.; Chang, C.; Zhang, H.P.; Wang, S.X.; Sun, G.; Xiao, S.H.; Ma, C.X. Identification of a Novel Allele of TaCKX6a02 Associated with Grain Size, Filling Rate and Weight of Common Wheat. PLoS ONE 2015, 10, e0144765. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhao, Y.L.; Gao, L.F.; Zhao, G.Y.; Zhou, R.H.; Zhang, B.S.; Jia, J.Z. TaCKX6-D1, the Ortholog of Rice OsCKX2, is Associated with Grain Weight in Hexaploid Wheat. New Phytol. 2012, 195, 574–584. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.; Lu, J.; Zhang, H.P.; Ma, C.X.; Sun, G.L. Copy Number Variation of Cytokinin Oxidase Gene Tackx4 Associated with Grain Weight and Chlorophyll Content of Flag Leaf in Common Wheat. PLoS ONE 2015, 10, 15. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Peng, K.X.; Cui, F.B.; Wang, D.L.; Zhao, J.Z.; Zhang, Y.J.; Yu, N.N.; Wang, Y.Y.; Zeng, D.L.; Wang, Y.H.; et al. Cytokinin Oxidase/Dehydrogenase OsCKX11 Coordinates Source and Sink Relationship in Rice by Simultaneous Regulation of Leaf Senescence and Grain Number. Plant Biotechnol. J. 2020, 19, 335–350. [Google Scholar] [CrossRef]
- Travella, S.; Klimm, T.E.; Keller, B. RNA Interference-based Gene Silencing as an Efficient Tool for Functional Genomics in Hexaploid Bread Wheat. Plant Physiol. 2006, 142, 6–20. [Google Scholar] [CrossRef]
- Gasparis, S.; Przyborowski, M.; Kala, M.; Nadolska-Orczyk, A. Knockout of the HvCKX1 or HvCKX3 Gene in Barley (Hordeum vulgare L.) by RNA-Guided Cas9 Nuclease Affects the Regulation of Cytokinin Metabolism and Root Morphology. Cells 2019, 8, 782. [Google Scholar] [CrossRef] [PubMed]
- Brenner, W.G.; Romanov, G.A.; Kollmer, I.; Burkle, L.; Schmulling, T. Immediate-early and Delayed Cytokinin Response Genes of Arabidopsis thaliana Identified by Genome-wide Expression Profiling Reveal Novel Cytokinin-sensitive Processes and Suggest Cytokinin Action through Transcriptional Cascades. Plant J. 2005, 44, 314–333. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.J.; Lin, S.S.; Liu, D.C.; Gan, L.J.; McAvoy, R.; Ding, J.; Li, Y. Evolution and Roles of Cytokinin Genes in Angiosperms 1: Do Ancient IPTs Play Housekeeping While Non-ancient IPTs Play Regulatory Roles? Hortic. Res. 2020, 7, 1–15. [Google Scholar] [CrossRef]
- Kroll, C.K.; Brenner, W.G. Cytokinin Signaling Downstream of the His-Asp Phosphorelay Network: Cytokinin-Regulated Genes and Their Functions. Front. Plant Sci. 2020, 11, 604489. [Google Scholar] [CrossRef] [PubMed]
- Gajdosova, S.; Spichal, L.; Kaminek, M.; Hoyerova, K.; Novak, O.; Dobrev, P.I.; Galuszka, P.; Klima, P.; Gaudinova, A.; Zizkova, E.; et al. Distribution, Biological Activities, Metabolism, and the Conceivable Function of Cis-zeatin-type Cytokinins in Plants. J. Exp. Bot. 2011, 62, 2827–2840. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.S.; Li, G.J.; Qu, D.J.; Li, X.; Wang, Y.N. Into the Seed: Auxin Controls Seed Development and Grain Yield. Int. J. Mol. Sci. 2020, 21, 1662. [Google Scholar] [CrossRef]
- Darussalam; Cole, M.A.; Patrick, J.W. Auxin Control of Photoassimilate Transport to and within Developing Grains of Wheat. Aust. J. Plant Physiol. 1998, 25, 69–77. [Google Scholar] [CrossRef]
- Kiba, T.; Takei, K.; Kojima, M.; Sakakibara, H. Side-Chain Modification of Cytokinins Controls Shoot Growth in Arabidopsis. Dev. Cell 2013, 27, 452–461. [Google Scholar] [CrossRef]
- Kudo, T.; Kiba, T.; Sakakibara, H. Metabolism and Long-distance Translocation of Cytokinins. J. Integr. Plant Biol. 2010, 52, 53–60. [Google Scholar] [CrossRef]
- Bishopp, A.; Lehesranta, S.; Vaten, A.; Help, H.; El-Showk, S.; Scheres, B.; Helariutta, K.; Mahonen, A.P.; Sakakibara, H.; Helariutta, Y. Phloem-Transported Cytokinin Regulates Polar Auxin Transport and Maintains Vascular Pattern in the Root Meristem. Curr. Biol. 2011, 21, 927–932. [Google Scholar] [CrossRef]
- Takei, K.; Sakakibara, H.; Taniguchi, M.; Sugiyama, T. Nitrogen-dependent Accumulation of Cytokinins in Root and the Translocation to Leaf: Implication of Cytokinin Species That Induces Gene Expression of Maize Response Regulator. Plant Cell Physiol. 2001, 42, 85–93. [Google Scholar] [CrossRef]
- Gao, X.P.; Francis, D.; Ormrod, J.C.; Bennett, M.D. Changes in Cell Number and Cell-Division Activity during Endosperm Development in Allohexaploid Wheat, Triticum aestivum L. J. Exp. Bot. 1992, 43, 1603–1609. [Google Scholar] [CrossRef]
- Hess, J.R.; Carman, J.G.; Banowetz, G.M. Hormones in Wheat Kernels during Embryony. J. Plant Physiol. 2002, 159, 379–386. [Google Scholar] [CrossRef]
- Yang, J.C.; Zhang, J.H.; Huang, Z.L.; Wang, Z.Q.; Zhu, Q.S.; Liu, L.J. Correlation of Cytokinin Levels in the Endosperms and Roots with Cell Number and Cell Division Activity during Endosperm Development in Rice. Ann. Bot. 2002, 90, 369–377. [Google Scholar] [CrossRef]
- Finkelstein, R.; Reeves, W.; Ariizumi, T.; Steber, C. Molecular Aspects of Seed Dormancy. Annu. Rev. Plant Biol. 2008, 59, 387–415. [Google Scholar] [CrossRef] [PubMed]
- Tuan, P.A.; Kumar, R.; Rehal, P.K.; Toora, P.K.; Ayele, B.T. Molecular Mechanisms Underlying Abscisic Acid/Gibberellin Balance in the Control of Seed Dormancy and Germination in Cereals. Front. Plant Sci. 2018, 9, 668. [Google Scholar] [CrossRef]
- Tuan, P.A.; Yamasaki, Y.; Kanno, Y.; Seo, M.; Ayele, B.T. Transcriptomics of Cytokinin and Auxin Metabolism and Signaling Genes during Seed Maturation in Dormant and Non-dormant Wheat Genotypes. Sci. Rep. 2019, 9, 1–16. [Google Scholar] [CrossRef]
- Powell, A.E.; Paleczny, A.R.; Olechowski, H.; Emery, R.J.N. Changes in Cytokinin Form and Concentration in Developing Kernels Correspond with Variation in Yield among Field-grown Barley Cultivars. Plant Physiol. Biochem. 2013, 64, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Fahy, B.; Siddiqui, H.; David, L.C.; Powers, S.J.; Borrill, P.; Uauy, C.; Smith, A.M. Final Grain Weight is Not Limited by the Activity of Key Starch-synthesising Enzymes during Grain Filling in Wheat. J. Exp. Bot. 2018, 69, 5461–5475. [Google Scholar] [CrossRef]
- Wang, W.Q.; Hao, Q.Q.; Tian, F.X.; Li, Q.X.; Wang, W. The Stay-green Phenotype of Wheat Mutant Tasg1 is Associated with Altered Cytokinin Metabolism. Plant Cell Rep. 2016, 35, 585–599. [Google Scholar] [CrossRef]
- Wang, W.Q.; Hao, Q.Q.; Wang, W.L.; Li, Q.X.; Chen, F.J.; Ni, F.; Wang, Y.; Fu, D.L.; Wu, J.J.; Wang, W. The Involvement of Cytokinin and Nitrogen Metabolism in Delayed Flag Leaf Senescence in a Wheat Stay-green Mutant, Tasg1. Plant Sci. 2019, 278, 70–79. [Google Scholar] [CrossRef]
- Yang, D.Q.; Li, Y.; Shi, Y.H.; Cui, Z.Y.; Luo, Y.L.; Zheng, M.J.; Chen, J.; Li, Y.X.; Yin, Y.P.; Wang, Z.L. Exogenous Cytokinins Increase Grain Yield of Winter Wheat Cultivars by Improving Stay-Green Characteristics under Heat Stress. PLoS ONE 2016, 11, e0155437. [Google Scholar] [CrossRef]
- Luo, Y.L.; Tang, Y.H.; Zhang, X.; Li, W.Q.; Chang, Y.L.; Pang, D.W.; Xu, X.; Li, Y.; Wang, Z.L. Interactions between Cytokinin and Nitrogen Contribute to Grain Mass in Wheat Cultivars by Regulating the Flag Leaf Senescence Process. Crop J. 2018, 6, 538–551. [Google Scholar] [CrossRef]
- Mao, C.J.; Lu, S.C.; Lv, B.; Zhang, B.; Shen, J.B.; He, J.M.; Luo, L.Q.; Xi, D.D.; Chen, X.; Ming, F. A Rice NAC Transcription Factor Promotes Leaf Senescence via ABA Biosynthesis. Plant Physiol. 2017, 174, 1747–1763. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.W.; Gan, S.S. An Abscisic Acid-AtNAP Transcription Factor-SAG113 Protein Phosphatase 2C Regulatory Chain for Controlling Dehydration in Senescing Arabidopsis Leaves. Plant Physiol. 2012, 158, 961–969. [Google Scholar] [CrossRef]
- Takagi, M.; Yokota, T.; Murofushi, N.; Saka, H.; Takahashi, N. Quantitative Changes of Free-Base, Riboside, Ribotide and Glucoside Cytokinins in Developing Rice Grains. Plant Growth Regul. 1989, 8, 349–364. [Google Scholar] [CrossRef]
- Hluska, T.; Dobrev, P.I.; Tarkowska, D.; Frebortova, J.; Zalabak, D.; Kopecny, D.; Plihal, O.; Kokas, F.; Briozzo, P.; Zatloukal, M.; et al. Cytokinin Metabolism in Maize: Novel Evidence of Cytokinin Abundance, Interconversions and Formation of a New Trans-zeatin Metabolic Product with a Weak Anticytokinin Activity. Plant Sci. 2016, 247, 127–137. [Google Scholar] [CrossRef]
- Zd’arska, M.; Zatloukalova, P.; Benitez, M.; Sedo, O.; Potesil, D.; Novak, O.; Svacinova, J.; Pesek, B.; Malbeck, J.; Vasickova, J.; et al. Proteome Analysis in Arabidopsis Reveals Shoot- and Root-Specific Targets of Cytokinin Action and Differential Regulation of Hormonal Homeostasis. Plant Physiol. 2013, 161, 918–930. [Google Scholar] [CrossRef]
- Przetakiewicz, A.; Karas, A.; Orczyk, W.; Nadolska-Orczyk, A. Agrobacterium-mediated Transformation of Polyploid Cereals. The Efficiency of Selection and Transgene Expression in Wheat. Cell. Mol. Biol. Lett. 2004, 9, 903–917. [Google Scholar] [PubMed]
- Przetakiewicz, A.; Orczyk, W.; Nadolska-Orczyk, A. The Effect of Auxin on Plant Regeneration of Wheat, Barley and Triticale. Plant Cell Tissue Organ. Cult. 2003, 73, 245–256. [Google Scholar] [CrossRef]
- Paolacci, A.R.; Tanzarella, O.A.; Porceddu, E.; Ciaffi, M. Identification and Validation of Reference Genes for Quantitative RT-PCR Normalization in Wheat. BMC Mol. Biol. 2009, 10, 1–27. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).