Acceptive Immunity: The Role of Fucosylated Glycans in Human Host–Microbiome Interactions
Abstract
1. Introduction
2. Recognition of Bacterial Surface Antigens
2.1. Toll-Like Receptors (TLR)
2.2. DC-SIGN
3. Stages of Immune System Development in the First Year of Life
4. Commensal Gut Microbiota and Changes in Short-Chain Fatty Acid Composition
5. Control over the Intestinal Colonization of Infants through Breastfeeding
6. Degradation of HMOs by Microorganisms
7. Lactobacilli and Bifidobacteria Cell Wall Components and Adhesins and Immune Response Modulation
8. Bacteroid Cell Wall
9. Helicobacter Pylori LPS
10. Fucosylation of IECs
11. Intestinal Immunoglobulins
12. Immunoglobulin Coating of Bacteria
13. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vakhitov, T.I.; Sitkin, S.I. The superorganism concept in biology and medicine. Exp. Clin. Gastroenterol. 2014, 7, 72–85. [Google Scholar]
- Xu, X.; Xu, P.; Ma, C.; Tang, J.; Zhang, X. Gut microbiota, host health, and polysaccharides. Biotechnol. Adv. 2013, 31, 318–337. [Google Scholar] [CrossRef]
- Tremaroli, V.; Bäckhed, F. Functional interactions between the gut microbiota and host metabolism. Nature 2012, 489, 242–249. [Google Scholar] [CrossRef]
- Levy, M.; Blacher, E.; Elinav, E. Microbiome, metabolites and host immunity. Curr. Opin. Microbiol. 2017, 35, 8–15. [Google Scholar] [CrossRef]
- Klimovich, V.B. Actual problems of evolutional immunology. Z. Evol. Biokhim. Fiziol. 2002, 38, 442–451. [Google Scholar]
- Byndloss, M.X.; Litvak, Y.; Bäumler, A.J. Microbiota-nourishing Immunity and Its Relevance for Ulcerative Colitis. Inflamm. Bowel Dis. 2019, 25, 811–815. [Google Scholar] [CrossRef]
- Halim, A.; Anonsen, J.H. Microbial glycoproteomics. Curr. Opin. Struct. Biol. 2017, 44, 143–150. [Google Scholar] [CrossRef]
- Wilson, I.B.; Zeleny, R.; Kolarich, D.; Staudacher, E.; Stroop, C.J.; Kamerling, J.P.; Altmann, F. Analysis of Asn-linked glycans from vegetable foodstuffs: Widespread occurrence of Lewis a, core α1, 3-linked fucose and xylose substitutions. Glycobiology 2001, 11, 261–274. [Google Scholar] [CrossRef]
- Khasbiullina, N.R.; Bovin, N.V. Hypotheses of the origin of natural antibodies: A glycobiologist’s opinion. Biochemistry 2015, 80, 820–835. [Google Scholar] [CrossRef]
- Li, J.; Hsu, H.C.; Mountz, J.D.; Allen, J.G. Unmasking Fucosylation: From Cell Adhesion to Immune System Regulation and Diseases. Cell Chem. Biol. 2018, 25, 499–512. [Google Scholar] [CrossRef]
- Kononova, S.V. How Fucose of Blood Group Glycotopes Programs Human Gut Microbiota. Biochemistry 2017, 82, 973–989. [Google Scholar] [CrossRef]
- Agace, W.W.; McCoy, K.D. Regionalized Development and Maintenance of the Intestinal Adaptive Immune Landscape. Immunity 2017, 46, 532–548. [Google Scholar] [CrossRef] [PubMed]
- Hug, H.; Mohajeri, M.H.; La Fata, G. Toll-Like Receptors: Regulators of the Immune Response in the Human Gut. Nutrients 2018, 10, 203. [Google Scholar] [CrossRef] [PubMed]
- Fitzgerald, K.A.; McWhirter, S.M.; Faia, K.L.; Rowe, D.C.; Latz, E.; Golenbock, D.T.; Coyle, A.J.; Liao, S.M.; Maniatis, T. IKKε and TBK1 are essential components of the IRF3 signaling pathway. Nat. Immunol. 2003, 4, 491–496. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, L.A.; Bowie, A.G. The family of five: TIR-domain-containing adaptors in Toll-like receptor signaling. Nat. Rev. Immunol. 2007, 7, 353–364. [Google Scholar] [CrossRef] [PubMed]
- Naiki, Y.; Michelsen, K.S.; Zhang, W.; Chen, S.; Doherty, T.M.; Arditi, M. Transforming growth factor-beta differentially inhibits MyD88-dependent, but not TRAM- and TRIF-dependent, lipopolysaccharide-induced TLR4 signaling. J. Biol. Chem. 2005, 280, 5491–5495. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Kunkel, S.L.; Chang, C.H. Negative regulation of MyD88-dependent signaling by IL-10 in dendritic cells. Proc. Natl. Acad. Sci. USA 2009, 106, 18327–18332. [Google Scholar] [CrossRef]
- Dziarski, R.; Gupta, D. Role of MD-2 in TLR2- and TLR4-mediated recognition of Gram-negative and Gram-positive bacteria and activation of chemokine genes. J. Endotoxin Res. 2000, 6, 401–405. [Google Scholar] [CrossRef]
- Tundup, S.; Srivastava, L.; Norberg, T.; Watford, W.; Harn, D. A neoglycoconjugate containing the human milk sugar LNFPIII drives anti-inflammatory activation of antigen presenting cells in a CD14 dependent pathway. PLoS ONE 2015, 10, e0137495. [Google Scholar] [CrossRef]
- Cummings, R.D.; McEver, R.P. C-type lectins. In Essentials of Glycobiology, 3rd ed.; Varki, A., Cummings, R.D., Esko, J.D., Stanley, P., Hart, G.W., Aebi, M., Darvill, A.G., Kinoshita, T., Packer, N.H., Prestegard, J.H., et al., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2017; pp. 435–452. [Google Scholar] [CrossRef]
- Manabe, Y.; Marchetti, R.; Takakura, Y.; Nagasaki, M.; Nihei, W.; Takebe, T.; Tanaka, K.; Kabayama, K.; Chiodo, F.; Hanashima, S.; et al. The core fucose on an IgG antibody is an endogenous ligand of Dectin-1. Angew. Chem. Int. Ed. Engl. 2019, 131, 18870–18875. [Google Scholar] [CrossRef]
- Cummings, R.D.; Schnaar, R.L. R-Type Lectins. In Essentials of Glycobiology, 3rd ed.; Varki, A., Cummings, R.D., Esko, J.D., Stanley, P., Hart, G.W., Aebi, M., Darvill, A.G., Kinoshita, T., Packer, N.H., Prestegard, J.H., et al., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2017; pp. 401–412. [Google Scholar] [CrossRef]
- Švajger, U.; Anderluh, M.; Jeras, M.; Obermajer, N. C-type lectin DC-SIGN: An adhesion, signalling and antigen-uptake molecule that guides dendritic cells in immunity. Cell. Signal. 2010, 22, 1397–1405. [Google Scholar] [CrossRef]
- Mitchell, D.A.; Fadden, A.J.; Drickamer, K. A novel mechanism of carbohydrate recognition by the C-type lectins DC-SIGN and DC-SIGNR subunit organization and binding to multivalent ligands. J. Biol. Chem. 2001, 276, 28939–28945. [Google Scholar] [CrossRef]
- Soilleux, E.J.; Morris, L.S.; Leslie, G.; Chehimi, J.; Luo, Q.; Levroney, E.; Trowsdale, J.; Montaner, L.J.; Doms, R.W.; Weissman, D.; et al. Constitutive and induced expression of DC-SIGN on dendritic cell and macrophage subpopulations in situ and in vitro. J. Leukoc. Biol. 2002, 71, 445–457. [Google Scholar] [PubMed]
- Chen, W.; Ma, L.; Li, R.; Huang, S.; Xie, R.; Chen, Y.; Zhao, B.; Fei, J.; Qu, H.; Chen, H.; et al. DC-SIGN Expression in Intestinal Epithelial Cells Regulates Sepsis-Associated Acute Intestinal Injury Via Activating ERK1/2-NF-κB/P65 Signaling. Shock 2019, 52, 434–442. [Google Scholar] [CrossRef] [PubMed]
- Konieczna, P.; Schiavi, E.; Ziegler, M.; Groeger, D.; Healy, S.; Grant, R.; O’Mahony, L. Human dendritic cell DC-SIGN and TLR-2 mediate complementary immune regulatory activities in response to Lactobacillus rhamnosus JB-1. PLoS ONE 2015, 10, e0120261. [Google Scholar] [CrossRef]
- Konieczna, P.; Groeger, D.; Ziegler, M.; Frei, R.; Ferstl, R.; Shanahan, F.; Quigley, E.M.; Kiely, B.; Akdis, C.A.; O’Mahony, L. Bifidobacterium infantis 35624 administration induces Foxp3 T regulatory cells in human peripheral blood: Potential role for myeloid and plasmacytoid dendritic cells. Gut 2012, 61, 354–366. [Google Scholar] [CrossRef] [PubMed]
- Bry, L.; Falk, P.G.; Gordon, J.L. Genetic engineering of carbohydrate biosynthetic pathways in transgenic mice demonstrates cell cycle-associated regulation of glycoconjugate production in small intestinal epithelial cells. Proc. Natl. Acad. Sci. USA 1996, 93, 1161–1166. [Google Scholar] [CrossRef]
- Miyake, M.; Zenita, K.; Tanaka, O.; Okada, Y.; Kannagi, R. Stage-specific expression of SSEA-1-related antigens in the developing lung of human embryos and its relation to the distribution of these antigens in lung cancers. Cancer Res. 1988, 48, 7150–7158. [Google Scholar] [PubMed]
- Gadhoum, S.Z.; Sackstein, R. CD15 expression in human myeloid cell differentiation is regulated by sialidase activity. Nat. Chem. Biol. 2008, 4, 751–757. [Google Scholar] [CrossRef]
- Sabatte, J.; Faigle, W.; Ceballos, A.; Morelle, W.; Rodrígues, C.R.; Lenicov, F.R.; Thépaut, M.; Fieschi, F.; Malchiodi, E.; Fernández, M.; et al. Semen clusterin is a novel DC-SIGN ligand. J. Immunol. 2011, 187, 5299–5309. [Google Scholar] [CrossRef]
- Okano, M.; Satoskar, A.R.; Nishizaki, K.; Harn, D.A. Lacto-N-fucopentaose III found on Schitosoma mansoni egg antigens functions as adjuvant for proteins by inducing Th2-type response. J. Immunol. 2001, 167, 442–450. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Ge, Z.; Rasko, D.A.; Taylor, D.E. Lewis antigens in Helicobacter pylori: Biosynthesis and phase variation. Mol. Microbiol. 2000, 36, 1187–1196. [Google Scholar] [CrossRef]
- Lee, J.H.; Pandey, R.P.; Kim, D.; Sohng, J.K. Cloning and functional characterization of an α-1,3-fucosyltransferase from Bacteroides fragilis. Biotechnol. Bioproc. Eng. 2013, 18, 843–849. [Google Scholar] [CrossRef]
- Lee, R.T.; Hsu, T.L.; Huang, S.K.; Hsieh, S.L.; Wong, C.H.; Lee, Y.C. Survey of immune-related, mannose/fucose-binding C-type lectin receptors reveals widely divergent sugar-binding specificities. Glycobiology 2011, 21, 512–520. [Google Scholar] [CrossRef]
- Feinberg, H.; Mitchell, D.A.; Drickamer, K.; Weis, W.I. Structural basis for selective recognition of oligosaccharides by DC-SIGN and DC-SIGNR. Science 2001, 294, 2163–2166. [Google Scholar] [CrossRef]
- Hsu, S.C.; Tsai, T.H.; Kawasaki, H.; Chen, C.H.; Plunkett, B.; Lee, R.T.; Lee, Y.C.; Huang, S.K. Antigen coupled with Lewis-x trisaccharides elicits potent immune responses in mice. J. Allergy Clin. Immunol. 2007, 119, 1522–1528. [Google Scholar] [CrossRef]
- van Liempt, E.; Bank, C.M.; Mehta, P.; Garcı’a-Vallejo, J.J.; Kawar, Z.S.; Geyer, R.; Alvarez, R.A.; Cummings, R.D.; Kooyk, Y.; van Die, I. Specificity of DC-SIGN for mannose-and fucose-containing glycans. FEBS Lett. 2006, 580, 6123–6131. [Google Scholar] [CrossRef]
- Holla, A.; Skerra, A. Comparative analysis reveals selective recognition of glycans by the dendritic cell receptors DC-SIGN and Langerin. Protein Eng. Des. Sel. 2011, 24, 659–669. [Google Scholar] [CrossRef] [PubMed]
- Noll, A.J.; Yu, Y.; Lasanajak, Y.; Duska-McEwen, G.; Buck, R.; Smith, D.F.; Cummings, R.D. Human DC-SIGN binds specific human milk glycans. Biochem. J. 2016, 473, 1343–1353. [Google Scholar] [CrossRef]
- Gringhuis, S.I.; den Dunnen, J.; Litjens, M.; van Het Hof, B.; van Kooyk, Y.; Geijtenbeek, T.B. C-type lectin DC-SIGN modulates Toll-like receptor signaling via Raf-1 kinase-dependent acetylation of transcription factor NF-κB. Immunity 2007, 26, 605–616. [Google Scholar] [CrossRef]
- Geijtenbeek, T.B.; Gringhuis, S.I. C-type lectin receptors in the control of T helper cell differentiation. Nat. Rev. Immunol. 2016, 16, 433–448. [Google Scholar] [CrossRef]
- van Gisbergen, K.P.; Sanchez-Hernandez, M.; Geijtenbeek, T.B.; van Kooyk, Y. Neutrophils mediate immune modulation of dendritic cells through glycosylation-dependent interactions between Mac-1 and DC-SIGN. J. Exp. Med. 2005, 201, 1281–1292. [Google Scholar] [CrossRef] [PubMed]
- Hornef, M.W.; Torow, N. ‘Layered immunity’ and the ‘neonatal window of opportunity’–timed succession of non-redundant phases to establish mucosal host–microbial homeostasis after birth. Immunology 2020, 159, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Houghteling, P.D.; Walker, W.A. Why is initial bacterial colonization of the intestine important to the infant’s and child’s health? J. Pediatr. Gastroenterol. Nutr. 2015, 60, 294–307. [Google Scholar] [CrossRef] [PubMed]
- Sudo, N.; Sawamura, S.; Tanaka, K.; Aiba, Y.; Kubo, C.; Koga, Y. The requirement of intestinal bacterial flora for the development of an IgE production system fully susceptible to oral tolerance induction. J. Immunol. 1997, 159, 1739–1745. [Google Scholar] [PubMed]
- Gensollen, T.; Iyer, S.S.; Kasper, D.L.; Blumberg, R.S. How colonization by microbiota in early life shapes the immune system. Science 2016, 352, 539–544. [Google Scholar] [CrossRef]
- Feehley, T.; Stefka, A.T.; Cao, S.; Nagler, C.R. Microbial regulation of allergic responses to food. Semin. Immunopathol. 2012, 34, 671–688. [Google Scholar] [CrossRef][Green Version]
- Yasmin, F.; Tun, H.M.; Konya, T.B.; Guttman, D.S.; Chari, R.S.; Field, C.J.; Becker, A.B.; Mandhane, P.J.; Turvey, S.E.; Subbarao, P.; et al. Cesarean section, formula feeding, and infant antibiotic exposure: Separate and combined impacts on gut microbial changes in later infancy. Front. Pediatr. 2017, 5, 200. [Google Scholar] [CrossRef]
- Molloy, J.; Allen, K.; Collier, F.; Tang, M.L.; Ward, A.C.; Vuillermin, P. The potential link between gut microbiota and IgE-mediated food allergy in early life. Int. J. Environ. Res. Public Health 2013, 10, 7235–7256. [Google Scholar] [CrossRef]
- Olszak, T.; An, D.; Zeissig, S.; Vera, M.P.; Richter, J.; Franke, A.; Glickman, J.N.; Siebert, R.; Baron, R.M.; Kasper, D.L.; et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 2012, 336, 489–493. [Google Scholar] [CrossRef]
- An, D.; Oh, S.F.; Olszak, T.; Neves, J.F.; Avci, F.Y.; Erturk-Hasdemir, D.; Lu, X.; Zeissig, S.; Blumberg, R.S.; Kasper, D.L. Sphingolipids from a symbiotic microbe regulate homeostasis of host intestinal natural killer T cells. Cell 2014, 156, 123–133. [Google Scholar] [CrossRef]
- Xiao, L.; van De Worp, W.R.; Stassen, R.; van Maastrigt, C.; Kettelarij, N.; Stahl, B.; Blijenberg, B.; Overbeek, S.A.; Folkerts, G.; Garssen, J.; et al. Human milk oligosaccharides promote immune tolerance via direct interactions with human dendritic cells. Eur. J. Immunol. 2019, 49, 1001–1014. [Google Scholar] [CrossRef] [PubMed]
- Buddington, R.K.; Weiher, E. The application of ecological principles and fermentable fibers to manage the gastrointestinal tract ecosystem. J. Nutr. 1999, 129, 1446S–1450S. [Google Scholar] [CrossRef]
- Fisher, C.K.; Mehta, P. Identifying keystone species in the human gut microbiome from metagenomic timeseries using sparse linear regression. PLoS ONE 2014, 9, e102451. [Google Scholar] [CrossRef] [PubMed]
- Trosvik, P.; de Muinck, E.J. Ecology of bacteria in the human gastrointestinal tract—identification of keystone and foundation taxa. Microbiome 2015, 3, 1–12. [Google Scholar] [CrossRef]
- Cockburn, D.W.; Koropatkin, N.M. Polysaccharide degradation by the intestinal microbiota and its influence on human health and disease. J. Mol. Biol. 2016, 428, 3230–3252. [Google Scholar] [CrossRef]
- Tuncil, Y.E.; Xiao, Y.; Porter, N.T.; Reuhs, B.L.; Martens, E.C.; Hamaker, B.R. Reciprocal prioritization to dietary glycans by gut bacteria in a competitive environment promotes stable coexistence. mBio 2017, 8, e01068-17. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.P. Glycan utilisation system in Bacteroides and Bifidobacteria and their roles in gut stability and health. Appl. Environ. Biotechnol. 2019, 103, 7287–7315. [Google Scholar] [CrossRef]
- La Reau, A.J.; Suen, G. The Ruminococci: Key symbionts of the gut ecosystem. J. Microbiol. 2018, 56, 199–208. [Google Scholar] [CrossRef]
- Ze, X.; Duncan, S.H.; Louis, P.; Flint, H.J. Ruminococcus bromii is a keystone species for the degradation of resistant starch in the human colon. ISME J. 2012, 6, 1535–1543. [Google Scholar] [CrossRef]
- Salazar, N.; Gueimonde, M.; De Los Reyes-Gavilán, C.G.; Ruas-Madiedo, P. Exopolysaccharides produced by lactic acid bacteria and bifidobacteria as fermentable substrates by the intestinal microbiota. Crit. Rev. Food Sci. Nutr. 2016, 56, 1440–1453. [Google Scholar] [CrossRef] [PubMed]
- Rios-Covian, D.; Arboleya, S.; Hernandez-Barranco, A.M.; Alvarez-Buylla, J.R.; Ruas-Madiedo, P.; Gueimonde, M.; Clara, G. Interactions between Bifidobacterium and Bacteroides species in cofermentations are affected by carbon sources, including exopolysaccharides produced by bifidobacteria. Appl. Environ. Microbiol. 2013, 79, 7518–7524. [Google Scholar] [CrossRef] [PubMed]
- Nilsen, M.; Madelen Saunders, C.; Leena Angell, I.; Arntzen, M.Ø.; Lødrup Carlsen, K.C.; Carlsen, K.-H.; Haugen, G.; Hagen, L.H.; Carlsen, M.H.; Hedlin, G.; et al. Butyrate Levels in the Transition from an Infant- to an Adult-Like Gut Microbiota Correlate with Bacterial Networks Associated with Eubacterium Rectale and Ruminococcus Gnavus. Genes 2020, 11, 1245. [Google Scholar] [CrossRef]
- Differding, M.K.; Benjamin-Neelon, S.E.; Hoyo, C.; Østbye, T.; Mueller, N.T. Timing of complementary feeding is associated with gut microbiota diversity and composition and short chain fatty acid concentrations over the first year of life. BMC Microbiol. 2020, 20, 1–13. [Google Scholar] [CrossRef]
- Corrêa-Oliveira, R.; Fachi, J.L.; Vieira, A.; Sato, F.T.; Vinolo, M.A.R. Regulation of immune cell function by short-chain fatty acids. Clin. Transl. Immunol. 2016, 5, e73. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Gurav, A.; Sivaprakasam, S.; Brady, E.; Padia, R.; Shi, H.; Thangaraju, M.; Prasad, P.D.; Manicassamy, S.; Munn, D.H.; et al. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 2014, 40, 128–139. [Google Scholar] [CrossRef]
- Nastasi, C.; Fredholm, S.; Willerslev-Olsen, A.; Hansen, M.; Bonefeld, C.M.; Geisler, C.; Andersen, M.H.; Ødum, N.; Woetmann, A. Butyrate and propionate inhibit antigen-specific CD8+T cell activation by suppressing IL-12 production by antigen-presenting cells. Sci. Rep. 2017, 7, 14516. [Google Scholar] [CrossRef]
- Zúñiga, M.; Monedero, V.; Yebra, M.J. Utilization of host-derived glycans by intestinal Lactobacillus and Bifidobacterium species. Front. Microbiol. 2018, 9, 1917. [Google Scholar] [CrossRef]
- Sakanaka, M.; Gotoh, A.; Yoshida, K.; Odamaki, T.; Koguchi, H.; Xiao, J.Z.; Kitaoka, M.; Katayama, T. Varied pathways of infant gut-associated Bifidobacterium to assimilate human milk oligosaccharides: Prevalence of the gene set and its correlation with bifidobacteria-rich microbiota formation. Nutrients 2020, 12, 71. [Google Scholar] [CrossRef]
- Marcobal, A.; Barboza, M.; Sonnenburg, E.D.; Pudlo, N.; Martens, E.C.; Desai, P.; Lebrilla, C.B.; Weimer, B.C.; Mills, D.A.; German, J.B.; et al. Bacteroides in the infant gut consume milk oligosaccharides via mucus-utilization pathways. Cell Host Microbe 2011, 10, 507–514. [Google Scholar] [CrossRef]
- Crost, E.H.; Tailford, L.E.; Le Gall, G.; Fons, M.; Henrissat, B.; Juge, N. Utilisation of mucin glycans by the human gut symbiont Ruminococcus gnavus is strain-dependent. PLoS ONE 2013, 8, e76341. [Google Scholar] [CrossRef] [PubMed]
- Audy, J.; Labrie, S.; Roy, D.; LaPointe, G. Sugar source modulates exopolysaccharide biosynthesis in Bifidobacterium longum subsp. longum CRC 002. Microbiology 2010, 156, 653–664. [Google Scholar] [CrossRef] [PubMed]
- Bessman, N.J.; Sonnenberg, G.F. Emerging roles for antigen presentation in establishing host–microbiome symbiosis. Immunol. Rev. 2016, 272, 139–150. [Google Scholar] [CrossRef]
- Cabrera-Rubio, R.; Collado, M.C.; Laitinen, K.; Salminen, S.; Isolauri, E.; Mira, A. The human milk microbiome changes over lactation and is shaped by maternal weight and mode of delivery. Am. J. Clin. Nutr. 2012, 96, 544–551. [Google Scholar] [CrossRef]
- Le Doare, K.; Holder, B.; Bassett, A.; Pannaraj, P.S. Mother’s milk: A purposeful contribution to the development of the infant microbiota and immunity. Front. Immunol. 2018, 9, 361. [Google Scholar] [CrossRef]
- Collado, M.C.; Delgado, S.; Maldonado, A.; Rodríguez, J.M. Assessment of the bacterial diversity of breast milk of healthy women by quantitative real-time PCR. Lett. Appl. Microbiol. 2009, 48, 523–528. [Google Scholar] [CrossRef]
- Goldsmith, F.; O’sullivan, A.; Smilowitz, J.T.; Freeman, S.L. Lactation and intestinal microbiota: How early diet shapes the infant gut. J. Mammary Gland Biol. Neoplasia 2015, 20, 149–158. [Google Scholar] [CrossRef]
- Karav, S.; Le Parc, A.; de Moura, J.M.L.N.; Frese, S.A.; Kirmiz, N.; Block, D.E.; Barile, D.; Mills, D.A. Oligosaccharides released from milk glycoproteins are selective growth substrates for infant-associated bifidobacteria. Appl. Environ. Microbiol. 2016, 82, 3622–3630. [Google Scholar] [CrossRef]
- Aakko, J.; Kumar, H.; Rautava, S.; Wise, A.; Autran, C.; Bode, L.; Isolauri, E.; Salminen, S. Human milk oligosaccharide categories define the microbiota composition in human colostrum. Benef. Microbes 2017, 8, 563–567. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Li, M.; Wu, S.; Lebrilla, C.B.; Chapkin, R.S.; Ivanov, I.; Donovan, S.M. Fecal microbiota composition of breast-fed infants is correlated with human milk oligosaccharides consumed. J. Pediatr. Gastroenterol. Nutr. 2015, 60, 825–833. [Google Scholar] [CrossRef] [PubMed]
- Thurl, S.; Munzert, M.; Boehm, G.; Matthews, C.; Stahl, B. Systematic review of the concentrations of oligosaccharides in human milk. Nutr. Rev. 2017, 75, 920–933. [Google Scholar] [CrossRef] [PubMed]
- Thurl, S.; Henker, J.; Siegel, M.; Tovar, K.; Sawatzki, G. Detection of four human milk groups with respect to Lewis blood group dependent oligosaccharides. Glycoconj. J. 1997, 14, 795–799. [Google Scholar] [CrossRef]
- Hall, A.B.; Tolonen, A.C.; Xavier, R.J. Human genetic variation and the gut microbiome in disease. Nat. Rev. Genet. 2017, 18, 690–699. [Google Scholar] [CrossRef]
- Plaza-Díaz, J.; Fontana, L.; Gil, A. Human milk oligosaccharides and immune system development. Nutrients 2018, 10, 1038. [Google Scholar] [CrossRef] [PubMed]
- McGuire, M.K.; Meehan, C.L.; McGuire, M.A.; Williams, J.E.; Foster, J.; Sellen, D.W.; Kamau-Mbuthia, E.W.; Kamundia, E.W.; Mbugua, S.; Moore, S.E.; et al. What’s normal? Oligosaccharide concentrations and profiles in milk produced by healthy women vary geographically. Am. J. Clin. Nutr. 2017, 105, 1086–1100. [Google Scholar] [CrossRef] [PubMed]
- Lackey, K.A.; Williams, J.E.; Meehan, C.L.; Zachek, J.A.; Benda, E.D.; Price, W.J.; Foster, J.A.; Sellen, D.W.; Kamau-Mbuthia, E.W.; Kamundia, E.W.; et al. What’s normal? Microbiomes in human milk and infant feces are related to each other but vary geographically: The INSPIRE study. Front. Nutr. 2019, 6, 45. [Google Scholar] [CrossRef]
- Rausch, P.; Künzel, S.; Suwandi, A.; Grassl, G.A.; Rosenstiel, P.; Baines, J.F. Multigenerational influences of the Fut2 gene on the dynamics of the gut microbiota in mice. Front. Microbiol. 2017, 8, 991. [Google Scholar] [CrossRef]
- Davenport, E.R.; Goodrich, J.K.; Bell, J.T.; Spector, T.D.; Ley, R.E.; Clark, A.G. ABO antigen and secretor statuses are not associated with gut microbiota composition in 1,500 twins. BMC Genom. 2016, 17, 1–14. [Google Scholar] [CrossRef]
- Urashima, T.; Asakuma, S.; Leo, F.; Fukuda, K.; Messer, M.; Oftedal, O.T. The Predominance of Type I Oligosaccharides Is a Feature Specific to Human Breast Milk. Adv. Nutr. 2012, 3, 473S–482S. [Google Scholar] [CrossRef]
- Becerra, J.E.; Rodríguez-Díaz, J.; Gozalbo-Rovira, R.; Palomino-Schätzlein, M.; Zúñiga, M.; Monedero, V.; Yebra, M.J. Unique Microbial Catabolic Pathway for the Human Core N-Glycan Constituent Fucosyl-α-1,6-N-Acetylglucosamine-Asparagine. mBio 2020, 11, e02804-19. [Google Scholar] [CrossRef] [PubMed]
- Carbohydrate-Active enZYmes Database. Available online: http://www.cazy.org/ (accessed on 4 April 2021).
- Bunesova, V.; Lacroix, C.; Schwab, C. Fucosyllactose and L-fucose utilization of infant Bifidobacterium longum and Bifidobacterium kashiwanohense. BMC Microbiol. 2016, 16, 248. [Google Scholar] [CrossRef] [PubMed]
- Matsuki, T.; Yahagi, K.; Mori, H.; Matsumoto, H.; Hara, T.; Tajima, S.; Ogawa, E.; Kodama, H.; Yamamoto, K.; Yamada, T.; et al. A key genetic factor for fucosyllactose utilization affects infant gut microbiota development. Nat. Commun. 2016, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Lewis, Z.T.; Totten, S.M.; Smilowitz, J.T.; Popovic, M.; Parker, E.; Lemay, D.G.; Van Tassell, M.L.; Miller, M.J.; Jin, Y.S.; German, J.B.; et al. Maternal fucosyltransferase 2 status affects the gut bifidobacterial communities of breastfed infants. Microbiome 2015, 3, 13. [Google Scholar] [CrossRef] [PubMed]
- Becerra, J.E.; Coll-Marqués, J.M.; Rodríguez-Díaz, J.; Monedero, V.; Yebra, M.J. Preparative scale purification of fucosyl-N-acetylglucosamine disaccharides and their evaluation as potential prebiotics and antiadhesins. Appl. Environ. Biotechnol. 2015, 99, 7165–7176. [Google Scholar] [CrossRef]
- Garrido, D.; Ruiz-Moyano, S.; Kirmiz, N.; Davis, J.C.; Totten, S.M.; Lemay, D.G.; Ugalde, J.A.; German, J.B.; Lebrilla, C.B.; Mills, D.A. A novel gene cluster allows preferential utilization of fucosylated milk oligosaccharides in Bifidobacterium longum subsp. longum SC596. Sci. Rep. 2016, 6, 35045. [Google Scholar] [CrossRef]
- Altermann, E.; Russell, W.M.; Azcarate-Peril, M.A.; Barrangou, R.; Buck, B.L.; McAuliffe, O.; Souther, N.; Dobson, A.; Duong, T.; Callanan, M.; et al. Complete genome sequence of the probiotic lactic acid bacterium Lactobacillus acidophilus NCFM. Proc. Natl. Acad. Sci. USA 2005, 102, 3906–3912. [Google Scholar] [CrossRef]
- Pridmore, R.D.; Berger, B.; Desiere, F.; Vilanova, D.; Barretto, C.; Pittet, A.C.; Zwahlen, M.C.; Rouvet, M.; Altermann, E.; Barrangou, R.; et al. The genome sequence of the probiotic intestinal bacterium Lactobacillus johnsonii NCC 533. Proc. Natl. Acad. Sci. USA 2004, 101, 2512–2517. [Google Scholar] [CrossRef]
- Azcarate-Peril, M.A.; Altermann, E.; Goh, Y.J.; Tallon, R.; Sanozky-Dawes, R.B.; Pfeiler, E.A.; O’Flaherty, S.; Buck, B.L.; Dobson, A.; Duong, T.; et al. Analysis of the genome sequence of Lactobacillus gasseri ATCC 33323 reveals the molecular basis of an autochthonous intestinal organism. Appl. Environ. Microbiol. 2008, 74, 4610–4625. [Google Scholar] [CrossRef]
- Ceapa, C.; Davids, M.; Ritari, J.; Lambert, J.; Wels, M.; Douillard, F.P.; Smokvina, T.; de Vos, W.M.; Knol, J.; Kleerebezem, M. The variable regions of Lactobacillus rhamnosus genomes reveal the dynamic evolution of metabolic and host-adaptation repertoires. Genome Biol. Evol. 2016, 8, 1889–1905. [Google Scholar] [CrossRef]
- Rodríguez-Díaz, J.; Rubio-del-Campo, A.; Yebra, M.J. Lactobacillus casei ferments the N-acetylglucosamine moiety of fucosyl-α-1,3-N-acetylglucosamine and excretes l-fucose. Appl. Environ. Microbiol. 2012, 78, 4613–4619. [Google Scholar] [CrossRef]
- Rodríguez-Díaz, J.; Monedero, V.; Yebra, M.J. Utilization of natural fucosylated oligosaccharides by three novel α-l-fucosidases from a probiotic Lactobacillus casei strain. Appl. Environ. Microbiol. 2011, 77, 703–705. [Google Scholar] [CrossRef] [PubMed]
- Becerra, J.E.; Yebra, M.J.; Monedero, V. An l-fucose operon in the probiotic Lactobacillus rhamnosus GG is involved in adaptation to gastrointestinal conditions. Appl. Environ. Microbiol. 2015, 81, 3880–3888. [Google Scholar] [CrossRef] [PubMed]
- Crouzet, L.; Derrien, M.; Cherbuy, C.; Plancade, S.; Foulon, M.; Chalin, B.; van Hylckama Vlieg, J.E.T.; Grompone, G.; Rigottier-Gois, L.; Serror, P. Lactobacillus paracasei CNCM I-3689 reduces vancomycin-resistant Enterococcus persistence and promotes Bacteroidetes resilience in the gut following antibiotic challenge. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef]
- Manley, K.J.; Fraenkel, M.B.; Mayall, B.C.; Power, D.A. Probiotic treatment of vancomycin-resistant enterococci: A randomised controlled trial. Med. J. Aust. 2007, 186, 454–457. [Google Scholar] [CrossRef]
- Szachta, P.; Ignys, I.; Cichy, W. An evaluation of the ability of the probiotic strain Lactobacillus rhamnosus GG to eliminate the gastrointestinal carrier state of vancomycin-resistant enterococci in colonized children. J. Clin. Gastroenterol. 2011, 45, 872–877. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Shen, Y.; Jiang, Y.; He, L.; Sun, Z. Bioinformatic Survey of S-Layer Proteins in Bifidobacteria. Comput. Mol. Biosci. 2018, 8, 68–79. [Google Scholar] [CrossRef]
- Johnson, B.R.; Hymes, J.; Sanozky-Dawes, R.; Henriksen, E.D.; Barrangou, R.; Klaenhammer, T.R. Conserved S-layer-associated proteins revealed by exoproteomic survey of S-layer-forming lactobacilli. Appl. Environ. Microbiol. 2016, 82, 134–145. [Google Scholar] [CrossRef]
- Pyclik, M.; Srutkova, D.; Schwarzer, M.; Górska, S. Bifidobacteria cell wall-derived exo-polysaccharides, lipoteichoic acids, peptidoglycans, polar lipids and proteins-their chemical structure and biological attributes. Int. J. Biol. Macromol. 2020, 147, 333–349. [Google Scholar] [CrossRef]
- González-Rodríguez, I.; Ruiz, L.; Gueimonde, M.; Margolles, A.; Sánchez, B. Factors involved in the colonization and survival of bifidobacteria in the gastrointestinal tract. FEMS Microbiol. Lett. 2013, 340, 1–10. [Google Scholar] [CrossRef]
- Kinoshita, H.; Imoto, S.; Suda, Y.; Ishida, M.; Watanabe, M.; Kawai, Y.; Kitazawa, H.; Miura, K.; Horii, A.; Saito, T. Proposal of screening method for intestinal mucus adhesive lactobacilli using the enzymatic activity of glyceraldehyde-3-phosphate dehydrogenase (GAPDH). Anim. Sci. J. 2013, 84, 150–158. [Google Scholar] [CrossRef]
- Houeix, B.; Synowsky, S.; Cairns, M.T.; Kane, M.; Kilcoyne, M.; Joshi, L. Identification of putative adhesins and carbohydrate ligands of Lactobacillus paracasei using a combinatorial in silico and glycomics microarray profiling approach. Integr. Biol. 2019, 11, 315–329. [Google Scholar] [CrossRef] [PubMed]
- Hymes, J.P.; Klaenhammer, T.R. Stuck in the middle: Fibronectin-binding proteins in Gram-positive bacteria. Front. Microbiol. 2016, 7, 1504. [Google Scholar] [CrossRef]
- Guerin, J.; Burgain, J.; Francius, G.; El-Kirat-Chatel, S.; Beaussart, A.; Scher, J.; Gaiani, C. Adhesion of Lactobacillus rhamnosus GG surface biomolecules to milk proteins. Food Hydrocoll. 2018, 82, 296–303. [Google Scholar] [CrossRef]
- Ruiz, L.; Delgado, S.; Ruas-Madiedo, P.; Margolles, A.; Sánchez, B. Proteinaceous Molecules Mediating Bifidobacterium-Host Interactions. Front. Microbiol. 2016, 7, 1193. [Google Scholar] [CrossRef] [PubMed]
- Fanning, S.; Hall, L.J.; Cronin, M.; Zomer, A.; MacSharry, J.; Goulding, D.; Motherway, M.O.; Shanahan, F.; Nally, K.; Dougan, G.; et al. Bifidobacterial surface-exopolysaccharide facilitates commensal-host interaction through immune modulation and pathogen protection. Proc. Natl. Acad. Sci. USA 2012, 109, 2108–2113. [Google Scholar] [CrossRef]
- Oleksy, M.; Klewicka, E. Exopolysaccharides produced by Lactobacillus sp.: Biosynthesis and applications. Crit. Rev. Food Sci. Nutr. 2018, 58, 450–462. [Google Scholar] [CrossRef] [PubMed]
- Salazar, N.; Prieto, A.; Leal, J.A.; Mayo, B.; Bada-Gancedo, J.C.; de Los Reyes-Gavilán, C.G.; Ruas-Madiedo, P. Production of exopolysaccharides by Lactobacillus and Bifidobacterium strains of human origin, and metabolic activity of the producing bacteria in milk. J. Dairy Sci. 2009, 92, 4158–4168. [Google Scholar] [CrossRef]
- Hidalgo-Cantabrana, C.; López, P.; Gueimonde, M.; Clara, G.; Suárez, A.; Margolles, A.; Ruas-Madiedo, P. Immune modulation capability of exopolysaccharides synthesised by lactic acid bacteria and bifidobacteria. Probiotics Antimicrob. Proteins 2012, 4, 227–237. [Google Scholar] [CrossRef]
- Speciale, I.; Verma, R.; Di Lorenzo, F.; Molinaro, A.; Im, S.H.; De Castro, C. Bifidobacterium bifidum presents on the cell surface a complex mixture of glucans and galactans with different immunological properties. Carbohydr. Polym. 2019, 218, 269–278. [Google Scholar] [CrossRef]
- Chen, Y.C.; Wu, Y.J.; Hu, C.Y. Monosaccharide composition influence and immunomodulatory effects of probiotic exopolysaccharides. Int. J. Biol. Macromol. 2019, 133, 575–582. [Google Scholar] [CrossRef]
- Ding, Y.H.; Qian, L.Y.; Pang, J.; Lin, J.Y.; Xu, Q.; Wang, L.H.; Huang, D.S.; Zou, H. The regulation of immune cells by Lactobacilli: A potential therapeutic target for anti-atherosclerosis therapy. Oncotarget 2017, 8, 59915. [Google Scholar] [CrossRef] [PubMed]
- Foligne, B.; Zoumpopoulou, G.; Dewulf, J.; Younes, A.B.; Chareyre, F.; Sirard, J.C.; Pot, B.; Grangette, C. A key role of dendritic cells in probiotic functionality. PLoS ONE 2007, 2, e313. [Google Scholar] [CrossRef]
- Mirpuri, J.; Sotnikov, I.; Myers, L.; Denning, T.L.; Yarovinsky, F.; Parkos, C.A.; Denning, P.W.; Louis, N.A. Lactobacillus rhamnosus (LGG) regulates IL-10 signaling in the developing murine colon through upregulation of the IL-10R2 receptor subunit. PLoS ONE 2012, 7, e51955. [Google Scholar] [CrossRef]
- Liu, J.; Gu, Z.; Song, F.; Zhang, H.; Zhao, J.; Chen, W. Lactobacillus plantarum ZS2058 and Lactobacillus rhamnosus GG use different mechanisms to prevent Salmonella infection in vivo. Front. Microbiol. 2019, 10, 299. [Google Scholar] [CrossRef] [PubMed]
- Tytgat, H.L.; van Teijlingen, N.H.; Sullan, R.M.A.; Douillard, F.P.; Rasinkangas, P.; Messing, M.; Reunanen, J.; Satokari, R.; Vanderleyden, J.; Dufrêne, Y.F.; et al. Probiotic gut microbiota isolate interacts with dendritic cells via glycosylated heterotrimeric pili. PLoS ONE 2016, 11, e0151824. [Google Scholar] [CrossRef]
- García, C.E.V.; Petrova, M.; Claes, I.J.; De Boeck, I.; Verhoeven, T.L.; Dilissen, E.; von Ossowski, I.; Palva, A.; Bullens, D.M.; Vanderleyden, J.; et al. Piliation of Lactobacillus rhamnosus GG promotes adhesion, phagocytosis, and cytokine modulation in macrophages. Appl. Environ. Microbiol. 2015, 81, 2050–2062. [Google Scholar] [CrossRef] [PubMed]
- Weiss, G.; Christensen, H.R.; Zeuthen, L.H.; Vogensen, F.K.; Jakobsen, M.; Frøkiær, H. Lactobacilli and bifidobacteria induce differential interferon-β profiles in dendritic cells. Cytokine 2011, 56, 520–530. [Google Scholar] [CrossRef] [PubMed]
- Jeon, S.G.; Kayama, H.; Ueda, Y.; Takahashi, T.; Asahara, T.; Tsuji, H.; Kiyono, H.; Ma, J.S.; Kusu, T.; Okumura, R.; et al. Probiotic Bifidobacterium breve induces IL-10-producing Tr1 cells in the colon. PLoS Pathog. 2012, 8, e1002714. [Google Scholar] [CrossRef]
- Ashida, N.; Yanagihara, S.; Shinoda, T.; Yamamoto, N. Characterization of adhesive molecule with affinity to Caco-2 cells in Lactobacillus acidophilus by proteome analysis. J. Biosci. Bioeng. 2011, 112, 333–337. [Google Scholar] [CrossRef] [PubMed]
- Bene, K.P.; Kavanaugh, D.W.; Leclaire, C.; Gunning, A.P.; MacKenzie, D.A.; Wittmann, A.; Young, I.D.; Kawasaki, N.; Rajnavolgyi, E.; Juge, N. Lactobacillus reuteri Surface Mucus Adhesins Upregulate Inflammatory Responses Through Interactions With Innate C-Type Lectin Receptors. Front. Microbiol. 2017, 8, 321. [Google Scholar] [CrossRef]
- Altmann, F.; Kosma, P.; O’Callaghan, A.; Leahy, S.; Bottacini, F.; Molloy, E.; Plattner, S.; Schiavi, E.; Gleinser, M.; Groeger, D.; et al. Genome analysis and characterisation of the exopolysaccharide produced by Bifidobacterium longum subsp. longum 35624™. PLoS ONE 2016, 11, e0162983. [Google Scholar] [CrossRef] [PubMed]
- Schiavi, E.; Gleinser, M.; Molloy, E.; Groeger, D.; Frei, R.; Ferstl, R.; Rodriguez-Perez, N.; Ziegler, M.; Grant, R.; Moriarty, T.F.; et al. The surface-associated exopolysaccharide of Bifidobacterium longum 35624 plays an essential role in dampening host proinflammatory responses and repressing local TH17 responses. Appl. Environ. Microbiol. 2016, 82, 7185–7196. [Google Scholar] [CrossRef] [PubMed]
- Coyne, M.J.; Reinap, B.; Lee, M.M.; Comstock, L.E. Human symbionts use a host-like pathway for surface fucosylation. Science 2005, 307, 1778–1781. [Google Scholar] [CrossRef] [PubMed]
- Coyne, M.J.; Chatzidaki-Livanis, M.; Paoletti, L.C.; Comstock, L.E. Role of glycan synthesis in colonization of the mammalian gut by the bacterial symbiont Bacteroides fragilis. Proc. Natl. Acad. Sci. USA 2008, 105, 13099–13104. [Google Scholar] [CrossRef]
- Fletcher, C.M.; Coyne, M.J.; Bentley, D.L.; Villa, O.F.; Comstock, L.E. Phase-variable expression of a family of glycoproteins imparts a dynamic surface to a symbiont in its human intestinal ecosystem. Proc. Natl. Acad. Sci. USA 2007, 104, 2413–2418. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, C.M.; Coyne, M.J.; Villa, O.F.; Chatzidaki-Livanis, M.; Comstock, L.E. A general O-glycosylation system important to the physiology of a major human intestinal symbiont. Cell 2009, 137, 321–331. [Google Scholar] [CrossRef] [PubMed]
- TerAvest, M.A.; He, Z.; Rosenbaum, M.A.; Martens, E.C.; Cotta, M.A.; Gordon, J.I.; Angenent, L.T. Regulated expression of polysaccharide utilization and capsular biosynthesis loci in biofilm and planktonic Bacteroides thetaiotaomicron during growth in chemostats. Biotechnol. Bioeng. 2014, 111, 165–173. [Google Scholar] [CrossRef]
- Bechon, N.; Mihajlovic, J.; Vendrell-Fernández, S.; Chain, F.; Langella, P.; Beloin, C.; Ghigo, J.M. Capsular polysaccharides cross-regulation modulates Bacteroides thetaiotaomicron biofilm formation. bioRxiv 2020, 11, e00729-20. [Google Scholar] [CrossRef]
- Coyne, M.J.; Fletcher, C.M.; Chatzidaki-Livanis, M.; Posch, G.; Schaffer, C.; Comstock, L.E. Phylum-wide general protein O-glycosylation system of the Bacteroidetes. Mol. Microbiol. 2013, 88, 772–783. [Google Scholar] [CrossRef]
- Liu, C.H.; Lee, S.M.; VanLare, J.M.; Kasper, D.L.; Mazmanian, S.K. Regulation of surface architecture by symbiotic bacteria mediates host colonization. Proc. Natl. Acad. Sci. USA 2008, 105, 3951–3956. [Google Scholar] [CrossRef]
- Tzianabos, A.O.; Pantosti, A.; Baumann, H.; Brisson, J.R.; Jennings, H.J.; Kasper, D.L. The capsular polysaccharide of Bacteroides fragilis comprises two ionically linked polysaccharides. J. Biol. Chem. 1992, 267, 18230–18235. [Google Scholar] [CrossRef]
- Donaldson, G.P.; Ladinsky, M.S.; Yu, K.B.; Sanders, J.G.; Yoo, B.B.; Chou, W.C.; Conner, M.E.; Earl, A.M.; Knight, R.; Bjorkman, P.J.; et al. Gut microbiota utilize immunoglobulin A for mucosal colonization. Science 2018, 360, 795–800. [Google Scholar] [CrossRef]
- Arnolds, K.L.; Moreno-Huizar, N.; Stanislawski, M.A.; Palmer, B.; Lozupone, C. Hemagglutination by B. fragilis is mediated by capsular polysaccharides and is influenced by host ABO blood type. bioRxiv 2020. [Google Scholar] [CrossRef]
- Kalka-Moll, W.M.; Wang, Y.; Comstock, L.E.; Gonzalez, S.E.; Tzianabos, A.O.; Kasper, D.L. Immunochemical and biological characterization of three capsular polysaccharides from a single Bacteroides fragilis strain. Infect. Immun. 2001, 69, 2339–2344. [Google Scholar] [CrossRef]
- Mazmanian, S.K.; Liu, C.H.; Tzianabos, A.O.; Kasper, D.L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 2005, 122, 107–118. [Google Scholar] [CrossRef]
- Ennamorati, M.; Vasudevan, C.; Clerkin, K.; Halvorsen, S.; Verma, S.; Ibrahim, S.; Prosper, S.; Porter, C.; Yeliseyev, V.; Kim, M.; et al. Intestinal microbes influence development of thymic lymphocytes in early life. Proc. Natl. Acad. Sci. USA 2020, 117, 2570–2578. [Google Scholar] [CrossRef] [PubMed]
- Bloem, K.; García-Vallejo, J.J.; Vuist, I.M.; Cobb, B.; Van Vliet, S.J.; Van Kooyk, Y. Interaction of the capsular polysaccharide A from Bacteroides fragilis with DC-SIGN on human dendritic cells is necessary for its processing and presentation to T cells. Front. Immunol. 2013, 4, 103. [Google Scholar] [CrossRef] [PubMed]
- Erridge, C.; Pridmore, A.; Eley, A.; Stewart, J.; Poxton, I.R. Lipopolysaccharides of Bacteroides fragilis, Chlamydia trachomatis and Pseudomonas aeruginosa signal via toll-like receptor 2. J. Med. Microbiol. 2004, 53, 735–740. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, S.; Erturk-Hasdemir, D.; Ochoa-Reparaz, J.; Reinecker, H.C.; Kasper, D.L. Plasmacytoid dendritic cells mediate anti-inflammatory responses to a gut commensal molecule via both innate and adaptive mechanisms. Cell Host Microbe 2014, 15, 413–423. [Google Scholar] [CrossRef]
- Erturk-Hasdemir, D.; Oh, S.F.; Okan, N.A.; Stefanetti, G.; Gazzaniga, F.S.; Seeberger, P.H.; Plevy, S.E.; Kasper, D.L. Symbionts exploit complex signaling to educate the immune system. Proc. Natl. Acad. Sci. USA 2019, 116, 26157–26166. [Google Scholar] [CrossRef]
- Mazmanian, S.K.; Round, J.L.; Kasper, D.L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 2008, 453, 620–625. [Google Scholar] [CrossRef] [PubMed]
- Telesford, K.M.; Yan, W.; Ochoa-Reparaz, J.; Pant, A.; Kircher, C.; Christy, M.A.; Begum-Haque, S.; Kasper, D.L.; Kasper, L.H. A commensal symbiotic factor derived from Bacteroides fragilis promotes human CD39+ Foxp3+ T cells and Treg function. Gut Microbes 2015, 6, 234–242. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Bjursell, M.K.; Himrod, J.; Deng, S.; Carmichael, L.K.; Chiang, H.C.; Hooper, L.V.; Gordon, J.I. A genomic view of the human-Bacteroides thetaiotaomicron symbiosis. Science 2003, 299, 2074–2076. [Google Scholar] [CrossRef] [PubMed]
- Martens, E.C.; Roth, R.; Heuser, J.E.; Gordon, J.I. Coordinate regulation of glycan degradation and polysaccharide capsule biosynthesis by a prominent human gut symbiont. J. Biol. Chem. 2009, 284, 18445–18457. [Google Scholar] [CrossRef]
- Porter, N.T.; Canales, P.; Peterson, D.A.; Martens, E.C. A subset of polysaccharide capsules in the human symbiont Bacteroides thetaiotaomicron promote increased competitive fitness in the mouse gut. Cell Host Microbe 2017, 22, 494–506. [Google Scholar] [CrossRef]
- Hsieh, S.; Porter, N.T.; Donermeyer, D.L.; Horvath, S.; Strout, G.; Saunders, B.T.; Zhang, N.; Zinselmeyer, B.; Martens, E.C.; Stappenbeck, T.S.; et al. Polysaccharide Capsules Equip the Human Symbiont Bacteroides thetaiotaomicron to Modulate Immune Responses to a Dominant Antigen in the Intestine. J. Immunol. 2020, 204, 1035–1046. [Google Scholar] [CrossRef]
- Peterson, D.A.; McNulty, N.P.; Guruge, J.L.; Gordon, J.I. IgA response to symbiotic bacteria as a mediator of gut homeostasis. Cell Host Microbe 2007, 2, 328–339. [Google Scholar] [CrossRef]
- Bjursell, M.K.; Martens, E.C.; Gordon, J.I. Functional genomic and metabolic studies of the adaptations of a prominent adult human gut symbiont, Bacteroides thetaiotaomicron, to the suckling period. J. Biol. Chem. 2006, 281, 36269–36279. [Google Scholar] [CrossRef]
- Benítez-Páez, A.; Gómez del Pulgar, E.M.; Sanz, Y. The glycolytic versatility of Bacteroides uniformis IECT 7771 and its genome response to oligo and polysaccharides. Front. Cell. Infect. Microbiol. 2017, 7, 383. [Google Scholar] [CrossRef]
- Wexler, H.M. Bacteroides: The good, the bad, and the nitty-gritty. Clin. Microbiol. Rev. 2007, 20, 593–621. [Google Scholar] [CrossRef]
- Rios-Covian, D.; Salazar, N.; Gueimonde, M.; de los Reyes-Gavilan, C.G. Shaping the metabolism of intestinal Bacteroides population through diet to improve human health. Front. Microbiol. 2017, 8, 376. [Google Scholar] [CrossRef]
- Chmiela, M.; Gonciarz, W. Molecular mimicry in Helicobacter pylori infections. World J. Gastroenterol. 2017, 23, 3964–3977. [Google Scholar] [CrossRef] [PubMed]
- Blaser, M.J. Pathogenicity and symbiosis: Human gastric colonization by Helicobacter pylori as model system of amphibiosis. In Ending the War Metaphor: The Changing Agenda for Unraveling the Host-Microbe Relationship; National Academies Press: Washington, DC, USA, 2006; p. 115. [Google Scholar]
- Mishra, S. Is Helicobacter pylori good or bad? Eur. J. Clin. Microbiol. Infect. Dis. 2013, 32, 301–304. [Google Scholar] [CrossRef]
- Müller, A.; Arnold, I.; Hitzler, I. The immunomodulatory properties of Helicobacter pylori confer protection against allergic and chronic inflammatory disorders. Front. Cell. Infect. Microbiol. 2012, 2, 10. [Google Scholar] [CrossRef]
- Bergman, M.; Del Prete, G.; van Kooyk, Y.; Appelmelk, B. Helicobacter pylori phase variation, immune modulation and gastric autoimmunity. Nat. Rev. Microbiol. 2006, 4, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Bergman, M.P.; Engering, A.; Smits, H.H.; Van Vliet, S.J.; Van Bodegraven, A.A.; Wirth, H.P.; Kapsenberg, M.L.; Vandenbroucke-Grauls, C.M.; van Kooyk, Y.; Appelmelk, B.J. Helicobacter pylori modulates the T helper cell 1/T helper cell 2 balance through phase-variable interaction between lipopolysaccharide and DC-SIGN. J. Exp. Med. 2004, 200, 979–990. [Google Scholar] [CrossRef] [PubMed]
- Rad, R.; Gerhard, M.; Lang, R.; Schöniger, M.; Rösch, T.; Schepp, W.; Becker, I.; Wagner, H.; Prinz, C. The Helicobacter pylori blood group antigen-binding adhesin facilitates bacterial colonization and augments a nonspecific immune response. J. Immunol. 2002, 168, 3033–3041. [Google Scholar] [CrossRef]
- Chmiela, M.; Miszczyk, E.; Rudnicka, K. Structural modifications of Helicobacter pylori lipopolysaccharide: An idea for how to live in peace. World J. Gastroenterol. 2014, 20, 9882–9897. [Google Scholar] [CrossRef]
- Gil, J.H.; Seo, J.W.; Cho, M.S.; Ahn, J.H.; Sung, H.Y. Role of Treg and TH17 cells of the gastric mucosa in children with Helicobacter pylori gastritis. J. Pediatr. Gastroenterol. Nutr. 2014, 58, 245–251. [Google Scholar] [CrossRef]
- Horvath, D.J., Jr.; Washington, M.K.; Cope, V.A.; Algood, H.M. IL-23 Contributes to Control of Chronic Helicobacter Pylori Infection and the Development of T Helper Responses in a Mouse Model. Front. Immunol. 2012, 3, 56. [Google Scholar] [CrossRef]
- Razavi, A.; Bagheri, N.; Azadegan-Dehkordi, F.; Shirzad, M.; Rahimian, G.; Rafieian-Kopaei, M.; Shirzad, H. Comparative Immune Response in Children and Adults with H. pylori Infection. J. Immunol. Res. 2015, 2015, 315957. [Google Scholar] [CrossRef]
- Magalhães, A.; Reis, C.A. Helicobacter pylori adhesion to gastric epithelial cells is mediated by glycan receptors. Braz. J. Med. Biol. Res. 2010, 43, 611–618. [Google Scholar] [CrossRef] [PubMed]
- Bry, L.; Falk, P.G.; Midtvedt, T.; Gordon, J.I. A model of host-microbial interactions in an open mammalian ecosystem. Science 1996, 273, 1380–1383. [Google Scholar] [CrossRef]
- Kashyap, P.C.; Marcobal, A.; Ursell, L.K.; Smits, S.A.; Sonnenburg, E.D.; Costello, E.K.; Higginbottom, S.K.; Domino, S.E.; Holmes, S.P.; Relman, D.A.; et al. Genetically dictated change in host mucus carbohydrate landscape exerts a diet-dependent effect on the gut microbiota. Proc. Natl. Acad. Sci. USA 2013, 110, 17059–17064. [Google Scholar] [CrossRef]
- Pickard, J.M.; Chervonsky, A.V. Intestinal fucose as a mediator of host–microbe symbiosis. J. Immunol. 2015, 194, 5588–5593. [Google Scholar] [CrossRef] [PubMed]
- Biol-N’garagba, M.C.; Louisot, P. Regulation of the intestinal glycoprotein glycosylation during postnatal development: Role of hormonal and nutritional factors. Biochimie 2003, 85, 331–352. [Google Scholar] [CrossRef]
- Yoneshige, A.; Sasaki, A.; Miyazaki, M.; Kojima, N.; Suzuki, A.; Matsuda, J. Developmental changes in glycolipids and synchronized expression of nutrient transporters in the mouse small intestine. J. Nutr. Biochem. 2010, 21, 214–226. [Google Scholar] [CrossRef] [PubMed]
- Terahara, K.; Nochi, T.; Yoshida, M.; Takahashi, Y.; Goto, Y.; Hatai, H.; Kurokawa, S.; Jang, M.H.; Kweon, M.N.; Domino, S.E.; et al. Distinct fucosylation of M cells and epithelial cells by Fut1 and Fut2, respectively, in response to intestinal environmental stress. Biochem. Biophys. Res. Commun. 2011, 404, 822–828. [Google Scholar] [CrossRef]
- Pickard, J.M.; Maurice, C.F.; Kinnebrew, M.A.; Abt, M.C.; Schenten, D.; Golovkina, T.V.; Bogatyrev, S.R.; Ismagilov, R.F.; Pamer, E.G.; Turnbaugh, P.J.; et al. Rapid fucosylation of intestinal epithelium sustains host–commensal symbiosis in sickness. Nature 2014, 514, 638–641. [Google Scholar] [CrossRef]
- Goto, Y.; Obata, T.; Kunisawa, J.; Sato, S.; Ivanov, I.I.; Lamichhane, A.; Takeyama, N.; Kamioka, M.; Sakamoto, M.; Matsuki, T.; et al. Innate lymphoid cells regulate intestinal epithelial cell glycosylation. Science 2014, 345, 1254009. [Google Scholar] [CrossRef]
- Pham, T.A.N.; Clare, S.; Goulding, D.; Arasteh, J.M.; Stares, M.D.; Browne, H.P.; Keane, A.J.; Page, J.A.; Kumasaka, N.; Kane, L.; et al. Epithelial IL-22RA1-mediated fucosylation promotes intestinal colonization resistance to an opportunistic pathogen. Cell Host Microbe 2014, 16, 504–516. [Google Scholar] [CrossRef] [PubMed]
- Ng, K.M.; Ferreyra, J.A.; Higginbottom, S.K.; Lynch, J.B.; Kashyap, P.C.; Gopinath, S.; Naidu, N.; Choudhury, B.; Weimer, B.C.; Monack, D.M.; et al. Microbiota-liberated host sugars facilitate post-antibiotic expansion of enteric pathogens. Nature 2013, 502, 96–99. [Google Scholar] [CrossRef] [PubMed]
- Suwandi, A.; Galeev, A.; Riedel, R.; Sharma, S.; Seeger, K.; Sterzenbach, T.; García Pastor, L.; Boyle, E.C.; Gal-Mor, O.; Hensel, M.; et al. Std fimbriae-fucose interaction increases Salmonella-induced intestinal inflammation and prolongs colonization. PLoS Pathog. 2019, 15, e1007915. [Google Scholar] [CrossRef] [PubMed]
- Pacheco, A.R.; Curtis, M.M.; Ritchie, J.M.; Munera, D.; Waldor, M.K.; Moreira, C.G.; Sperandio, V. Fucose sensing regulates bacterial intestinal colonization. Nature 2012, 492, 113–117. [Google Scholar] [CrossRef]
- Nanthakumar, N.N.; Meng, D.; Newburg, D.S. Glucocorticoids and microbiota regulate ontogeny of intestinal fucosyltransferase 2 requisite for gut homeostasis. Glycobiology 2013, 23, 1131–1141. [Google Scholar] [CrossRef]
- Muñoz-Esparza, N.C.; Latorre-Moratalla, M.L.; Comas-Basté, O.; Toro-Funes, N.; Veciana-Nogués, M.T.; Vidal-Carou, M.C. Polyamines in food. Front. Nutr. 2019, 6, 108. [Google Scholar] [CrossRef]
- Hooper, L.V.; Midtvedt, T.; Gordon, J.I. How host-microbial interactions shape the nutrient environment of the mammalian intestine. Annu. Rev. Nutr. 2002, 22, 283–307. [Google Scholar] [CrossRef]
- Umesaki, Y.; Okada, Y.; Matsumoto, S.; Imaoka, A.; Setoyama, H. Segmented filamentous bacteria are indigenous intestinal bacteria that activate intraepithelial lymphocytes and induce MHC class II molecules and fucosyl asialo GM1 glycolipids on the small intestinal epithelial cells in the ex-germ-free mouse. Microbiol. Immunol. 1995, 39, 555–562. [Google Scholar] [CrossRef]
- Biol, M.C.; Lenoir, D.; Greco, S.; Galvain, D.; Hugueny, I.; Louisot, P. Role of insulin and nutritional factors in intestinal glycoprotein fucosylation during postnatal development. Am. J. Physiol. Gastrointest. Liver Physiol. 1998, 275, G936–G942. [Google Scholar] [CrossRef]
- Jakobsson, H.E.; Abrahamsson, T.R.; Jenmalm, M.C.; Harris, K.; Quince, C.; Jernberg, C.; Björkstén, B.; Engstrand, L.; Andersson, A.F. Decreased gut microbiota diversity, delayed Bacteroidetes colonisation and reduced Th1 responses in infants delivered by caesarean section. Gut 2014, 63, 559–566. [Google Scholar] [CrossRef]
- Gregory, K.E.; LaPlante, R.D.; Shan, G.; Kumar, D.V.; Gregas, M. Mode of birth influences preterm infant intestinal colonization with bacteroides over the early neonatal period. Adv. Neonatal Care 2015, 15, 386–393. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, E.; De Palma, G.; Capilla, A.; Nova, E.; Pozo, T.; Castillejo, G.; Varea, V.; Marcos, A.; Garrote, J.A.; Polanco, I.; et al. Influence of environmental and genetic factors linked to celiac disease risk on infant gut colonization by Bacteroides species. Appl. Environ. Microbiol. 2011, 77, 5316–5323. [Google Scholar] [CrossRef]
- Hooper, L.V.; Xu, J.; Falk, P.G.; Midtvedt, T.; Gordon, J.I. A molecular sensor that allows a gut commensal to control its nutrient foundation in a competitive ecosystem. Proc. Natl. Acad. Sci. USA 1999, 96, 9833–9838. [Google Scholar] [CrossRef]
- Belenguer, A.; Duncan, S.H.; Holtrop, G.; Anderson, S.E.; Lobley, G.E.; Flint, H.J. Impact of pH on lactate formation and utilization by human fecal microbial communities. Appl. Environ. Microbiol. 2007, 73, 6526–6533. [Google Scholar] [CrossRef] [PubMed]
- Reichardt, N.; Duncan, S.H.; Young, P.; Belenguer, A.; Leitch, C.M.; Scott, K.P.; Flint, H.J.; Louis, P. Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J. 2014, 8, 1323–1335. [Google Scholar] [CrossRef] [PubMed]
- Park, D.; Xu, G.; Barboza, M.; Shah, I.M.; Wong, M.; Raybould, H.; Mills, D.A.; Lebrilla, C.B. Enterocyte glycosylation is responsive to changes in extracellular conditions: Implications for membrane functions. Glycobiology 2017, 27, 847–860. [Google Scholar] [CrossRef] [PubMed]
- Meng, D.; Newburg, D.S.; Young, C.; Baker, A.; Tonkonogy, S.L.; Sartor, R.B.; Walker, W.A.; Nanthakumar, N.N. Bacterial symbionts induce a FUT2-dependent fucosylated niche on colonic epithelium via ERK and JNK signaling. Am. J. Physiol. Gastrointest. Liver Physiol. 2007, 293, 780–787. [Google Scholar] [CrossRef]
- Sczesnak, A.; Segata, N.; Qin, X.; Gevers, D.; Petrosino, J.F.; Huttenhower, C.; Littman, D.R.; Ivanov, I.I. The genome of th17 cell-inducing segmented filamentous bacteria reveals extensive auxotrophy and adaptations to the intestinal environment. Cell Host Microbe 2011, 10, 260–272. [Google Scholar] [CrossRef]
- Jonsson, H.; Hugerth, L.W.; Sundh, J.; Andersson, A.F. Genome Sequence of Segmented Filamentous Bacteria Present in the Human Intestine. Commun. Biol. 2020, 3, 485. [Google Scholar] [CrossRef]
- Hedblom, G.A.; Reiland, H.A.; Sylte, M.J.; Johnson, T.J.; Baumler, D.J. Segmented Filamentous Bacteria-Metabolism Meets Immunity. Front Microbiol. 2018, 9, 1991. [Google Scholar] [CrossRef]
- Wang, Y.; Xiang, C.; Chen, H.; Yin, Y.; Chen, X.; Zhao, Y.; Wu, Y.; Li, Y.; Wang, X.; Chen, H.; et al. Induction of intestinal Th17 cells by flagellins from segmented filamentous bacteria. Front. Immunol. 2019, 10, 2750. [Google Scholar] [CrossRef] [PubMed]
- Ivanov, I.I.; Atarashi, K.; Manel, N.; Brodie, E.L.; Shima, T.; Karaoz, U.; Wei, D.; Goldfarb, K.C.; Santee, C.A.; Lynch, S.V.; et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 2009, 139, 485–498. [Google Scholar] [CrossRef] [PubMed]
- Goto, Y.; Panea, C.; Nakato, G.; Cebula, A.; Lee, C.; Diez, M.G.; Laufer, T.M.; Ignatowicz, L.; Ivanov, I.I. Segmented filamentous bacteria antigens presented by intestinal dendritic cells drive mucosal Th17 cell differentiation. Immunity 2014, 40, 594–607. [Google Scholar] [CrossRef]
- Yi, J.; Jung, J.; Han, D.; Surh, C.D.; Lee, Y.J. Segmented Filamentous Bacteria Induce Divergent Populations of Antigen-Specific CD4 T Cells in the Small Intestine. Mol. Cells 2019, 42, 228–236. [Google Scholar] [CrossRef]
- Parks, O.B.; Pociask, D.A.; Hodzic, Z.; Kolls, J.K.; Good, M. Interleukin-22 signaling in the regulation of intestinal health and disease. Front. Cell Dev. Biol. 2016, 3, 85. [Google Scholar] [CrossRef] [PubMed]
- Lejeune, D.; Dumoutier, L.; Constantinescu, S.; Kruijer, W.; Schuringa, J.J.; Renauld, J.C. Interleukin-22 (IL-22) activates the JAK/STAT, ERK, JNK, and p38 MAP kinase pathways in a rat hepatoma cell line. Pathways that are shared with and distinct from IL-10. J. Biol. Chem. 2002, 277, 33676–33682. [Google Scholar] [CrossRef] [PubMed]
- Sonnenberg, G.F.; Fouser, L.A.; Artis, D. Functional biology of the IL-22-IL-22R pathway in regulating immunity and inflammation at barrier surfaces. Adv. Immunol. 2010, 107, 1–29. [Google Scholar] [CrossRef]
- Sepahi, A.; Liu, Q.; Friesen, L.; Kim, C.H. Dietary fiber metabolites regulate innate lymphoid cell responses. Mucosal Immunol. 2020, 10. [Google Scholar] [CrossRef]
- Chun, E.; Lavoie, S.; Fonseca-Pereira, D.; Bae, S.; Michaud, M.; Hoveyda, H.R.; Fraser, G.L.; Gallini Comeau, C.A.; Glickman, J.N.; Fuller, M.H.; et al. Metabolite-sensing receptor Ffar2 regulates colonic group 3 innate lymphoid cells and gut immunity. Immunity 2019, 51, 871–884. [Google Scholar] [CrossRef]
- Berndt, B.E.; Zhang, M.; Owyang, S.Y.; Cole, T.S.; Wang, T.W.; Luther, J.; Veniaminova, N.A.; Merchant, J.L.; Chen, C.C.; Huffnagle, G.B.; et al. Butyrate increases IL-23 production by stimulated dendritic cells. Am. J. Physiol. Gastrointest. Liver Physiol. 2012, 303, G1384–G1392. [Google Scholar] [CrossRef] [PubMed]
- Goto, Y.; Lamichhane, A.; Kamioka, M.; Sato, S.; Honda, K.; Kunisawa, J.; Kiyono, H. IL-10-producing CD4+ T cells negatively regulate fucosylation of epithelial cells in the gut. Sci. Rep. 2015, 5, 15918. [Google Scholar] [CrossRef] [PubMed]
- Shouval, D.S.; Ouahed, J.; Biswas, A.; Goettel, J.A.; Horwitz, B.H.; Klein, C.; Muise, A.M.; Snapper, S.B. Interleukin 10 receptor signaling: Master regulator of intestinal mucosal homeostasis in mice and humans. Adv. Immunol. 2014, 122, 177–210. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Kelly, C.J.; Battista, K.D.; Schaefer, R.; Lanis, J.M.; Alexeev, E.E.; Wang, R.X.; Onyiah, J.C.; Kominsky, D.J.; Colgan, S.P. Microbial-derived butyrate promotes epithelial barrier function through IL-10 receptor–dependent repression of claudin-2. J. Immunol. 2017, 199, 2976–2984. [Google Scholar] [CrossRef]
- Kappler, K.; Hennet, T. Emergence and significance of carbohydrate-specific antibodies. Genes Immun. 2020, 21, 224–239. [Google Scholar] [CrossRef] [PubMed]
- Bunker, J.J.; Erickson, S.A.; Flynn, T.M.; Henry, C.; Koval, J.C.; Meisel, M.; Jabri, B.; Antonopoulos, D.A.; Wilson, P.C.; Bendelac, A. Natural polyreactive IgA antibodies coat the intestinal microbiota. Science 2017, 358, eaan6619. [Google Scholar] [CrossRef]
- Bovin, N.V. Natural antibodies to glycans. Biochemistry 2013, 78, 786–797. [Google Scholar] [CrossRef]
- Gayet, R.; Michaud, E.; Nicoli, F.; Chanut, B.; Paul, M.; Rochereau, N.; Guillon, C.; He, Z.; Papagno, L.; Bioley, G.; et al. Impact of IgA isoforms on their ability to activate dendritic cells and to prime T cells. Eur. J. Immunol. 2020, 50, 1295–1306. [Google Scholar] [CrossRef]
- Rochereau, N.; Drocourt, D.; Perouzel, E.; Pavot, V.; Redelinghuys, P.; Brown, G.D.; Tiraby, G.; Roblin, X.; Verrier, B.; Genin, C.; et al. Dectin-1 is essential for reverse transcytosis of glycosylated SIgA-antigen complexes by intestinal M cells. PLoS Biol. 2013, 11, e1001658. [Google Scholar] [CrossRef]
- Li, Y.; Jin, L.; Chen, T. The Effects of Secretory IgA in the Mucosal Immune System. BioMed Res. Int. 2020, 2020, 2032057. [Google Scholar] [CrossRef]
- Janzon, A.; Goodrich, J.K.; Koren, O.; TEDDY Study Group; Waters, J.L.; Ley, R.E. Interactions between the Gut Microbiome and Mucosal Immunoglobulins A, M, and G in the Developing Infant Gut. Msystems 2019, 4, e00612-19. [Google Scholar] [CrossRef]
- Misevic, G.; Garbarino, E. Glycan-to-Glycan Binding: Molecular Recognition through Polyvalent Interactions Mediates Specific Cell Adhesion. Molecules 2021, 26, 397. [Google Scholar] [CrossRef] [PubMed]
- Joglekar, P.; Ding, H.; Canales-Herrerias, P.; Pasricha, P.J.; Sonnenburg, J.L.; Peterson, D.A. Intestinal IgA regulates expression of a fructan polysaccharide utilization locus in colonizing gut commensal Bacteroides thetaiotaomicron. mBio 2019, 10, e02324-19. [Google Scholar] [CrossRef]
- Pabst, O.; Slack, E. IgA and the intestinal microbiota: The importance of being specific. Mucosal Immunol. 2019, 13, 12–21. [Google Scholar] [CrossRef]
- Sterlin, D.; Fadlallah, J.; Adams, O.; Fieschi, C.; Parizot, C.; Dorgham, K.; Rajkumar, A.; Autaa, G.; El-Kafsi, H.; Charuel, J.L.; et al. Human IgA binds a diverse array of commensal bacteria. J. Exp. Med. 2020, 217, e20181635. [Google Scholar] [CrossRef] [PubMed]
- Palm, N.W.; De Zoete, M.R.; Cullen, T.W.; Barry, N.A.; Stefanowski, J.; Hao, L.; Degnan, P.H.; Hu, J.; Peter, I.; Zhang, W.; et al. Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 2014, 158, 1000–1010. [Google Scholar] [CrossRef] [PubMed]
- Briliūtė, J.; Urbanowicz, P.A.; Luis, A.S.; Baslé, A.; Paterson, N.; Rebello, O.; Hendel, J.; Ndeh, D.A.; Lowe, E.C.; Martens, E.C.; et al. Complex N-glycan breakdown by gut Bacteroides involves an extensive enzymatic apparatus encoded by multiple co-regulated genetic loci. Nat. Microbiol. 2019, 4, 1571–1581. [Google Scholar] [CrossRef]
- Mathias, A.; Corthésy, B. N-Glycans on secretory component: Mediators of the interaction between secretory IgA and gram-positive commensals sustaining intestinal homeostasis. Gut Microbes 2011, 2, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Royle, L.; Roos, A.; Harvey, D.J.; Wormald, M.R.; Van Gijlswijk-Janssen, D.; Redwan, E.R.M.; Wilson, I.A.; Daha, M.R.; Dwek, R.A.; Rudd, P.M. Secretory IgA N- and O-glycans provide a link between the innate and adaptive immune systems. J. Biol. Chem. 2003, 278, 20140–20153. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Guerrero, A.; Parker, E.; Strum, J.S.; Smilowitz, J.T.; German, J.B.; Lebrilla, C.B. Site-specific glycosylation of secretory immunoglobulin A from human colostrum. J. Proteome Res. 2015, 14, 1335–1349. [Google Scholar] [CrossRef]
- Bunker, J.J.; Flynn, T.M.; Koval, J.C.; Shaw, D.G.; Meisel, M.; McDonald, B.D.; Ishizuka, I.E.; Dent, A.L.; Wilson, P.C.; Jabri, B.; et al. Innate and adaptive humoral responses coat distinct commensal bacteria with immunoglobulin A. Immunity 2015, 43, 541–553. [Google Scholar] [CrossRef]
- Planer, J.D.; Peng, Y.; Kau, A.L.; Blanton, L.V.; Ndao, I.M.; Tarr, P.I.; Warner, B.B.; Gordon, J.I. Development of the gut microbiota and mucosal IgA responses in twins and gnotobiotic mice. Nature 2016, 534, 263–266. [Google Scholar] [CrossRef]
- Magri, G.; Comerma, L.; Pybus, M.; Sintes, J.; Lligé, D.; Segura-Garzón, D.; Bascones, S.; Yeste, A.; Grasset, E.K.; Gutzeit, C.; et al. Human secretory IgM emerges from plasma cells clonally related to gut memory B cells and targets highly diverse commensals. Immunity 2017, 47, 118–134. [Google Scholar] [CrossRef] [PubMed]
- Mathias, A.; Duc, M.; Favre, L.; Benyacoub, J.; Blum, S.; Corthésy, B. Potentiation of polarized intestinal Caco-2 cell responsiveness to probiotics complexed with secretory IgA. J. Biol. Chem. 2010, 285, 33906–33913. [Google Scholar] [CrossRef]
- Huus, K.E.; Bauer, K.C.; Brown, E.M.; Bozorgmehr, T.; Woodward, S.E.; Serapio-Palacios, A.; Boutin, R.C.T.; Petersen, C.; Finlay, B.B. Commensal Bacteria Modulate Immunoglobulin A Binding in Response to Host Nutrition. Cell Host Microbe 2020, 27, 909–921. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, A.; Vogelzang, A.; Maruya, M.; Miyajima, M.; Murata, M.; Son, A.; Kuwahara, T.; Tsuruyama, T.; Yamada, S.; Matsuura, M.; et al. IgA regulates the composition and metabolic function of gut microbiota by promoting symbiosis between bacteria. J. Exp. Med. 2018, 215, 2019–2034. [Google Scholar] [CrossRef] [PubMed]
- Grosserichter-Wagener, C.; Radjabzadeh, D.; van der Weide, H.; Smit, K.N.; Kraaij, R.; Hays, J.P.; van Zelm, M.C. Differences in systemic IgA reactivity and circulating Th subsets in healthy volunteers with specific microbiota enterotypes. Front. Immunol. 2019, 10, 341. [Google Scholar] [CrossRef]
- Perruzza, L.; Strati, F.; Gargari, G.; D’Erchia, A.M.; Fosso, B.; Pesole, G.; Guglielmetti, S.; Grassi, F. Enrichment of intestinal Lactobacillus by enhanced secretory IgA coating alters glucose homeostasis in P2rx7−/− mice. Sci. Rep. 2019, 9, 9315. [Google Scholar] [CrossRef] [PubMed]
- Bunker, J.J.; Bendelac, A. IgA responses to microbiota. Immunity 2018, 49, 211–224. [Google Scholar] [CrossRef] [PubMed]
- Hino, S.; Mizushima, T.; Kaneko, K.; Kawai, E.; Kondo, T.; Genda, T.; Yamada, T.; Hase, K.; Nishimura, N.; Morita, T. Mucin-Derived O-Glycans Act as Endogenous Fiber and Sustain Mucosal Immune Homeostasis via Short-Chain Fatty Acid Production in Rat IECum. J. Nutr. 2020, 150, 2656–2665. [Google Scholar] [CrossRef]
- Bollinger, R.R.; Everett, M.L.; Palestrant, D.; Love, S.D.; Lin, S.S.; Parker, W. Human secretory immunoglobulin A may contribute to biofilm formation in the gut. Immunology 2003, 109, 580–587. [Google Scholar] [CrossRef]
- Kappler, K.; Restin, T.; Lasanajak, Y.; Smith, D.F.; Bassler, D.; Hennet, T. Limited Neonatal Carbohydrate-Specific Antibody Repertoire Consecutive to Partial Prenatal Transfer of Maternal Antibodies. Front. Microbiol. 2020, 11, 2513. [Google Scholar] [CrossRef] [PubMed]
- Kappler, K.; Lasanajak, Y.; Smith, D.F.; Opitz, L.; Hennet, T. Increased Antibody Response to Fucosylated Oligosaccharides and Fucose-Carrying Bacteroides Species in Crohn’s Disease. Front. Microbiol. 2020, 11, 1553. [Google Scholar] [CrossRef] [PubMed]
- Mikulic, J.; Longet, S.; Favre, L.; Benyacoub, J.; Corthesy, B. Secretory IgA in complex with Lactobacillus rhamnosus potentiates mucosal dendritic cell-mediated Treg cell differentiation via TLR regulatory proteins, RALDH2 and secretion of IL-10 and TGF-β. Cell. Mol. Immunol. 2017, 14, 546–556. [Google Scholar] [CrossRef] [PubMed]
- Sitkin, S.; Vakhitov, T.; Kononova, S.; Skalinskaya, M.; Pokrotnieks, J. Gut Microbiota-Mediated Pleiotropic Effects of Fucose Can Improve Inflammatory Bowel Disease by Modulating Bile Acid Metabolism and Enhancing Propionate Production. Inflamm. Bowel Dis. 2021, 27, e10–e11. [Google Scholar] [CrossRef]
- Goto, Y.; Uematsu, S.; Kiyono, H. Epithelial glycosylation in gut homeostasis and inflammation. Nat. Immunol. 2016, 17, 1244–1251. [Google Scholar] [CrossRef]
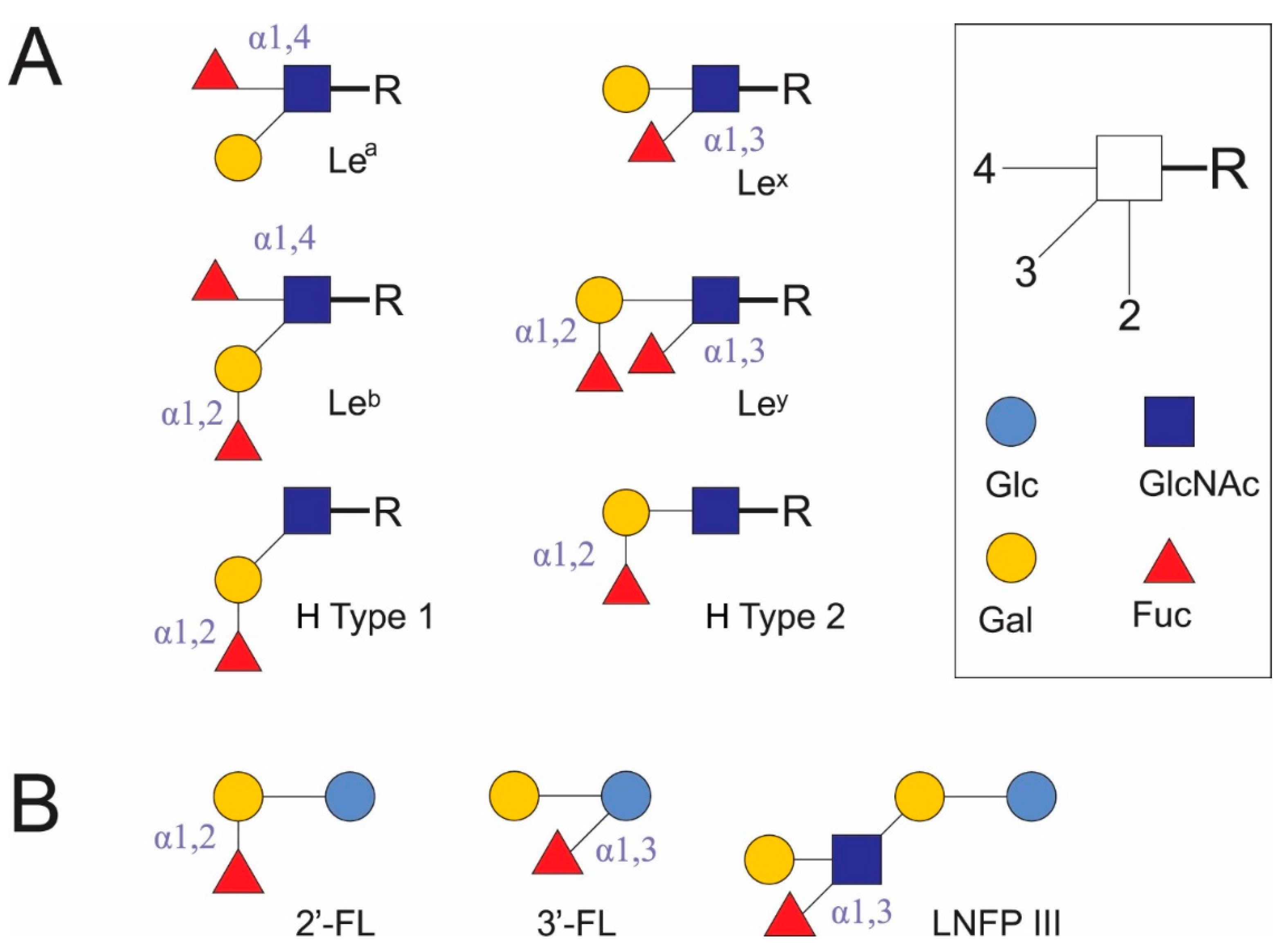
| HMO | Positive Correlations, Bacteria | Negative Correlations, Bacteria |
|---|---|---|
| LNFP III | Bifidobacterium breve > Staphylococcus aureus > Enterococcus spp. > other | Lactobacillus group > Faecalibacterium prausnitzii > Clostridium cluster XIVa |
| LNFP I | Akkermansia muciniphila > Bifidobacterium spp. > Enterococcus spp. > Bifidobacterium breve > Bifidobacterium longum group and Staphylococcus aureus > Streptococcus group > Staphylococcus spp. > other | Total bacteria > Faecalibacterium prausnitzii |
| 2′-FL | Akkermansia muciniphila > Bifidobacterium spp. > Clostridium cluster XIVa > Lactobacillus group | Staphylococcus aureus > Enterococcus spp. > total bacteria > Staphylococcus spp. > Bacteroides–Prevotella group > other |
| 3′-FL | Total bacteria > Lactobacillus group > Bifidobacterium bifidum | Enterococcus spp. > Akkermansia muciniphila > Staphylococcus aureus > Bifidobacterium longum group > Staphylococcus spp. > Bifidobacterium breve > Bifidobacterium spp. > Faecalibacterium prausnitzii > other |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kononova, S.; Litvinova, E.; Vakhitov, T.; Skalinskaya, M.; Sitkin, S. Acceptive Immunity: The Role of Fucosylated Glycans in Human Host–Microbiome Interactions. Int. J. Mol. Sci. 2021, 22, 3854. https://doi.org/10.3390/ijms22083854
Kononova S, Litvinova E, Vakhitov T, Skalinskaya M, Sitkin S. Acceptive Immunity: The Role of Fucosylated Glycans in Human Host–Microbiome Interactions. International Journal of Molecular Sciences. 2021; 22(8):3854. https://doi.org/10.3390/ijms22083854
Chicago/Turabian StyleKononova, Svetlana, Ekaterina Litvinova, Timur Vakhitov, Maria Skalinskaya, and Stanislav Sitkin. 2021. "Acceptive Immunity: The Role of Fucosylated Glycans in Human Host–Microbiome Interactions" International Journal of Molecular Sciences 22, no. 8: 3854. https://doi.org/10.3390/ijms22083854
APA StyleKononova, S., Litvinova, E., Vakhitov, T., Skalinskaya, M., & Sitkin, S. (2021). Acceptive Immunity: The Role of Fucosylated Glycans in Human Host–Microbiome Interactions. International Journal of Molecular Sciences, 22(8), 3854. https://doi.org/10.3390/ijms22083854







