The Relationship between the Gut Microbiome and Metformin as a Key for Treating Type 2 Diabetes Mellitus
Abstract
1. Introduction
2. Gut Microbiome and T2DM
3. Potential Mechanisms of Metformin on the Gut Microbiome
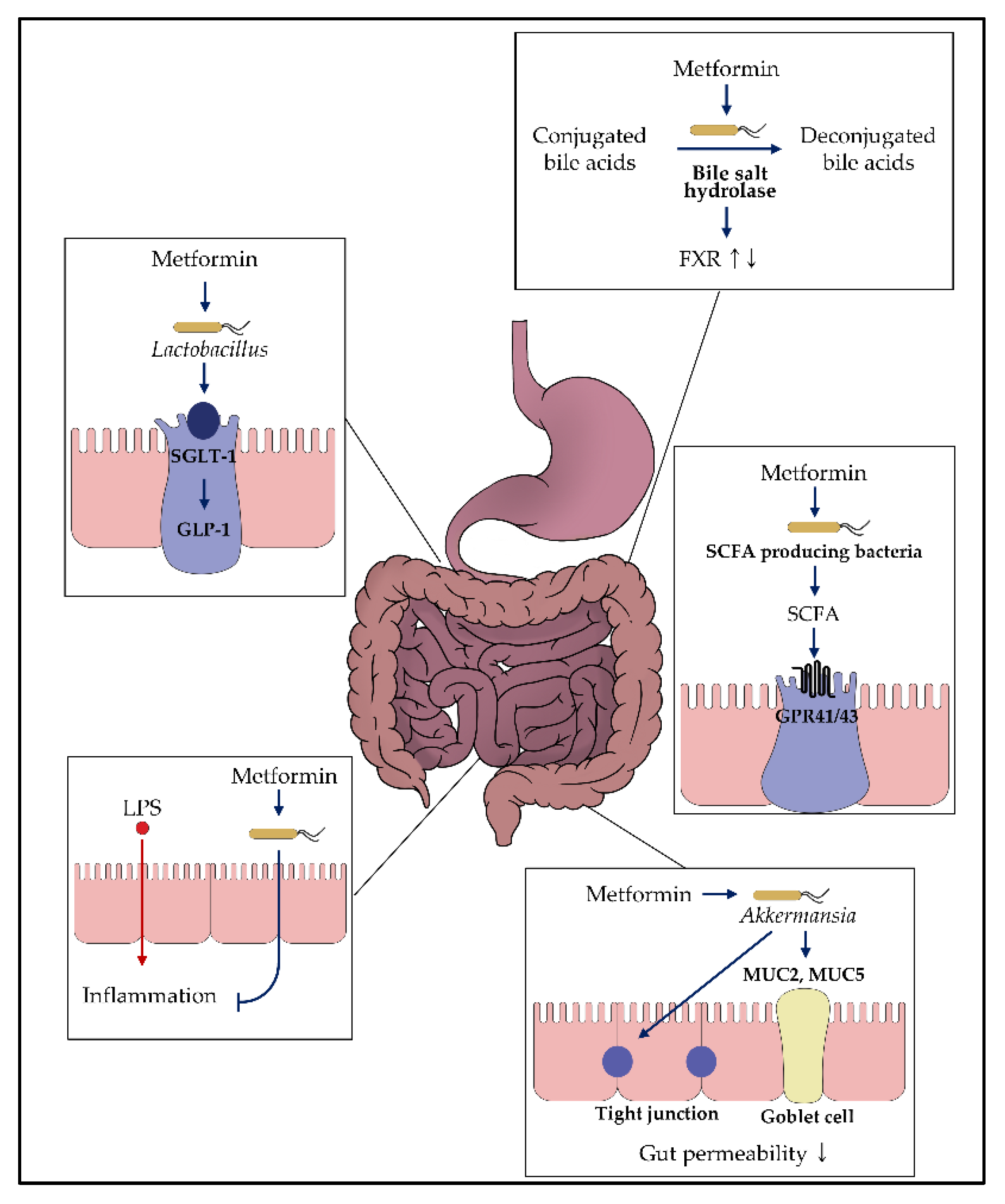
3.1. Regulation of Glucose Homeostasis
3.2. Effects on Bacteria Producing Short-Chain Fatty Acid
3.3. Enhancement of the Gut Permeability
| Ref * | Animal | Study Design | Gut Microbiota | Biochemical Alterations |
|---|---|---|---|---|
| [76] | Rats | Metformin treatment in high-fat diet | versus without metformin treatment in high fat diet Family: Lactobacillaceae ↑ Genus: Achromobacter –, Acinetobacter –, Azorhiziphilus –, Enterococcus –, Escherichia –, Klebsiella –, tobacillus ↑, Sarcina –, Stenotrophomnas – | NA |
| [88] | Mice | Metformin treatment in high-fat diet | versus without metformin treatment in high fat diet α-diversity (Shannon): ↓ Phylum: Bacteroidetes ↑, Verrucomicrobia ↑ Family: Bacteroidaceae ↑, Clostridiales familyXIII ↑, Incertae sedis ↑, Rikenellaceae ↑, Ruminococcaceae ↑, Verrucomicrobioaceae ↑ Species: Akkermansia muciniphila ↑, Clostridium cocleatum ↑ | Inflammation scores – |
| Metformin treatment in normal diet | versus without metformin treatment in normal diet α-diversity (Shannon): – Phylum: Bacteroidetes – Family: Rikenellaceae ↑, Ruminococcaceae ↑, Verrucomicrobioaceae ↑ Genus: Alistipes spp. ↑, Akkermansia spp. ↑, Clostridium spp. ↑ | NA | ||
| [89] | Mice | Metformin treatment for 16 weeks in high-fat diet | versus without metformin treatment in high fat diet α-diversity (observed OTU): – Phylum: Bacteroidetes ↑, Firmicutes ↓, Verrucomicrobia ↑ Genus: Akkermansia ↑, Bacteroides ↑, Butyricimonas ↑, Parabacteroides ↑ | IL-6 mRNA ↓ IL-1β mRNA ↓ |
| [90] | Mice | Metformin treatment for 24 weeks in high-fat diet | versus without metformin treatment in high fat diet α-diversity (Shannon): ↓ Phylum: Bacteroidetes ↑, Firmicutes ↓, Verrucomicrobia ↑ Family: Desulfovibrionaceae Genus: Akkermansia ↑, Bacteroides ↑, Christensenella ↑, Coprococcus↓, Dorea ↓, Lachnoclostridium ↓, Parabacteroides ↑, Papillibacter ↓, Oscillospira ↓, Ruminococcus ↓, Desulfovibrio ↓, Muribaculum ↓ | Plasma threonine ↓, methionine sulfoxide ↓, Tetradecanoylcarnitine ↓, Hexadecenoylcarnitine ↓ |
| [91] | Mice | Metformin treatment in obese mice (db/db mice) | versus without metformin treatment in obese mice (db/db mice) α-diversity (Shannon): ↑ Genus: Akkermansia ↑, Butyricimonas ↑, Clostridium ↓, Coprococcus ↑, Dehalobacterium ↑, Dorea ↑, Lactobacillus ↑, Oscillospira ↑, Parabacteroides ↓, Paraprevotella ↑, Prevotella ↓, Proteus ↓, Ruminococcus ↑ | Total SCFA concentration in feces ↑ Acetic acid ↑, Butyric acid ↑ LPS levels ↓ |
| [92] | Mice | Metformin treatment in high-fat diet | versus without metformin treatment in high fat diet α-diversity (Shannon, evenness): – Phylum: Bacteroidetes ↑ Family: Coriobacteriaceae ↓, Ruminococcaceae ↑, S24_7 ↑, Veilonellaceae ↓ Genus: Dorea ↓, Dehalobacterium ↓, Lactobacillus ↓, Lactococcus ↑, Roseburia ↓, SMB53 ↓ | IL-6 ↓, IL-1β ↓, TNF α ↓ Taurine ↑, Butyrate ↑, Total Bile acids ↑, Propionate ↑, Leucine ↑, Creatinine ↓, Sarcosine ↓, Glutamate ↓, Pyruvate ↓, Formate ↓ |
| [93] | Rats | Metformin treatment in high-fat diet combined with a low dose streptozocin | versus without metformin treatment in high fat diet α-diversity (Simpson, Shannon): ↑ Class: Coriobacteriia ↑ Family: S24_7 ↑ | Total SCFAs ↑, Butyric acid ↑, Isovaleric acid ↑ |
| [94] | Rats | Metformin treatment in high-fat diet combined with a low dose streptozocin | versus without metformin treatment in high fat diet α-diversity (Chao1): ↑ Family: S24_7 ↓ Genus: Anaerotruncus ↑, Escherichia-Shiegella ↓, Eubacterium xylanophilum ↑, Lachnospiraceae NK4A136 ↑, Lachnospiraceae-UCG_006 ↑, Roseburia ↑ | Serum LPS ↓, Serum CRP↓, Serum TNF α ↓, Serum IL-6 ↓ Propionate in cecum ↑, Butyrate in cecum ↑ |
| [99] | Mice (female) | Metformin treatment in fat, fructose and cholesterol rich diet | versus without metformin treatment in fructose and cholesterol rich diet Family: Alloprevotella ↓ Genus: Bacteroides –, Romboutsia ↓ Species: Akkermansia muciniphila –, Lactobacillus animalis ↓ | TNF α ↓ Endotoxin ↓ |
| [109] | Mice | Metformin treatment for 5 weeks in high-fat diet combined with a low dose streptozocin | versus without metformin treatment in high fat diet α-diversity (observed OTU): – Genus: Akkermansia ↑, Bacteroides spp. ↓ | NA |
| [110] | Mice | Metformin treatment in high-fat diet | versus without metformin treatment in high fat diet Phylum: Verrucomicrobia ↑, Genus: Akkermansia ↑, Alistipes ↑, Anaerotruncus ↓, Blautia ↓, Lactococcus ↓, Lactonifactor ↓, Lawsonia ↓, Odoribacter ↓, Parabacteroides ↓ | IL-6 mRNA ↓ IL-1β mRNA ↓ |
| [121] | Mice | Metformin treatment for 30 days | versus without metformin treatment in healthy mice α-diversity (Shannon): – Class: Lachnopiraceae ↓, Porphyromonadaceae ↑, Prevoltellaceae ↑, Rhodobacteraceae ↓, Rikenellaceae ↑, Verrucomicrobiaceae ↑ | NA |
| [123] | Rats | Metformin treatment in high-fat diet | versus without metformin treatment in high fat diet α-diversity (Shannon): ↓ Phylum: Bacteroidetes –, Firmicutes –, Proteobacteria ↑ Species: Akkermansia ↑, Allobaculum ↑, Bacteroides ↑, Blautia ↑, Butyricoccus ↑, Clostridium ↓, Klebsiella ↑, Lactobacillus ↑, Parasutterella ↑, Phascolarctobacterium ↑, Prevotella ↑, Roseburia ↓ | NA |
| [124] | Rats | Metformin treatment in high-fat diet combined with a low dose streptozocin | versus without metformin treatment in high fat diet α-diversity (Chao1, Shannon): ↑ Phylum: Bacteroidetes ↑, Firmicutes ↓, Proteobacteria ↓ Order: Clostridiales ↑, Enterobacteriales ↓, Lactobacillales ↑ Genus: Akkermansia ↑, Desulfovibrio ↓, Lachnospiraceae NK4A136 ↓, Lactobacillus ↑, Roseburia ↑ | NA |
| [125] | Mice | Metformin treatment for 3 weeks in high-fat diet | versus without metformin treatment in high fat diet α-diversity (Shannon, evenness): – Genus: Akkermansia ↑, Allobaculum ↓, Clostridium ↓, Enterococcus ↓, Lactococcus ↓, Leuconostoc ↓, Oscillospira ↑, Parabacteroides ↑, Prevotella ↑, Ruminococcus ↓, Streptococcus ↓ | NA |
| [126] | Mice with | Metformin treatment for 5 weeks in high-fat diet combined with a low dose streptozocin | versus without metformin treatment in high fat diet α-diversity (Chao1): ↓ Phylum: Bacteroidetes ↓, Firmicutes ↑, Proteobacteria ↓ Genus: Lactobacillus ↑ | NA |
| [127] | Mice | Metformin treatment in high-fat diet | versus without metformin treatment in high fat diet α-diversity (Shannon, evenness): – Species: Bacteriodetes fragilis ↓, Escherichia coli ↓ | Serum endotoxin ↓ IL-6 ↓, TLR4 ↓ |
| [128] | Rats | Metformin treatment in high-fat diet combined with a low dose streptozocin | versus without metformin treatment in high fat diet α-diversity (Chao1): ↑ Phylum: Bacteroidetes ↑, Proteobacteria ↓, Verrucomicrobia ↓ Family: Alcaligenaceae ↑, Peptococcaceae ↑, Prevotellaceae ↑, S24_7 ↑ Genus: Prevotella ↑, Sutterella ↑, 02d06 ↑, rc4 ↑ | IL-6 mRNA in pancrease ↓, TNF α mRNA in pancrease ↓, LPS ↓ |
| [129] | Rats | Metformin treatment in high-fat diet combined with a low dose streptozocin | versus without metformin treatment in high fat diet Genus: Bifidobacterium ↑, Lactobacillus ↑ Species: Clostridium perfringens ↓, Escherichia coli ↓ | Plasma endotoxin ↓, Total SCFAs in cecum ↑, Lactic acid in cecum ↑, Acetic acid in cecum ↑ |
| [130] | Rats | Metformin treatment in Otsuka Long-Evans Tokushima Fatty (OLETF) rats | versus without metformin treatment Genus: Akkermansia ↑, Prevotella ↓, Roseburia ↑ Species: Escherichia coli ↓ | Serum endotoxin ↓, Fecal endotoxin ↓, serum TNF α ↓, serum IL-6 ↓ |
| [131] | Rats | Metformin treatment in Zucker diabetic fatty rats | versus without metformin treatment α-diversity (Shannon): – Phylum: Bacteroidetes –, Firmicutes –↑, Proteobacteria ↓, Tenericutes –, Verrucomicrobia ↑ Genus: Lactobacillus ↑ Species: Lactobacillus intestinalis ↑, Lactobacillus johnsonii ↑ | NA |
3.4. Modulation of the Immune Response
3.5. Actions on the Circulation of the Bile Acids
4. Relationships between Metformin and Gut Microbiome in Human Studies
5. Perspective and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Foretz, M.; Guigas, B.; Bertrand, L.; Pollak, M.; Viollet, B. Metformin: From mechanisms of action to therapies. Cell Metab. 2014, 20, 953–966. [Google Scholar] [CrossRef]
- Andujar-Plata, P.; Pi-Sunyer, X.; Laferrere, B. Metformin effects revisited. Diabetes Res. Clin. Pract. 2012, 95, 1–9. [Google Scholar] [CrossRef]
- Rena, G.; Pearson, E.R.; Sakamoto, K. Molecular mechanism of action of metformin: Old or new insights? Diabetologia 2013, 56, 1898–1906. [Google Scholar] [CrossRef]
- Rojas, L.B.; Gomes, M.B. Metformin: An old but still the best treatment for type 2 diabetes. Diabetol. Metab. Syndr. 2013, 5, 6. [Google Scholar] [CrossRef]
- Viollet, B.; Guigas, B.; Sanz Garcia, N.; Leclerc, J.; Foretz, M.; Andreelli, F. Cellular and molecular mechanisms of metformin: An overview. Clin. Sci. 2012, 122, 253–270. [Google Scholar] [CrossRef]
- Graham, G.G.; Punt, J.; Arora, M.; Day, R.O.; Doogue, M.P.; Duong, J.K.; Furlong, T.J.; Greenfield, J.R.; Greenup, L.C.; Kirkpatrick, C.M.; et al. Clinical pharmacokinetics of metformin. Clin. Pharmacokinet. 2011, 50, 81–98. [Google Scholar] [CrossRef]
- Beckmann, R. [Absorption, distribution in the organism and elimination of metformin]. Diabetologia 1969, 5, 318–324. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wilcock, C.; Wyre, N.D.; Bailey, C.J. Subcellular distribution of metformin in rat liver. J. Pharm. Pharmacol. 1991, 43, 442–444. [Google Scholar] [CrossRef] [PubMed]
- Rena, G.; Hardie, D.G.; Pearson, E.R. The mechanisms of action of metformin. Diabetologia 2017, 60, 1577–1585. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.S.; Jonker, J.W.; Kato, Y.; Kusuhara, H.; Schinkel, A.H.; Sugiyama, Y. Involvement of organic cation transporter 1 in hepatic and intestinal distribution of metformin. J. Pharmacol. Exp. Ther. 2002, 302, 510–515. [Google Scholar] [CrossRef] [PubMed]
- Shu, Y.; Sheardown, S.A.; Brown, C.; Owen, R.P.; Zhang, S.; Castro, R.A.; Ianculescu, A.G.; Yue, L.; Lo, J.C.; Burchard, E.G.; et al. Effect of genetic variation in the organic cation transporter 1 (oct1) on metformin action. J. Clin. Investig. 2007, 117, 1422–1431. [Google Scholar] [CrossRef] [PubMed]
- Natali, A.; Ferrannini, E. Effects of metformin and thiazolidinediones on suppression of hepatic glucose production and stimulation of glucose uptake in type 2 diabetes: A systematic review. Diabetologia 2006, 49, 434–441. [Google Scholar] [CrossRef] [PubMed]
- Foretz, M.; Hebrard, S.; Leclerc, J.; Zarrinpashneh, E.; Soty, M.; Mithieux, G.; Sakamoto, K.; Andreelli, F.; Viollet, B. Metformin inhibits hepatic gluconeogenesis in mice independently of the lkb1/ampk pathway via a decrease in hepatic energy state. J. Clin. Investig. 2010, 120, 2355–2369. [Google Scholar] [CrossRef]
- Madiraju, A.K.; Erion, D.M.; Rahimi, Y.; Zhang, X.M.; Braddock, D.T.; Albright, R.A.; Prigaro, B.J.; Wood, J.L.; Bhanot, S.; MacDonald, M.J.; et al. Metformin suppresses gluconeogenesis by inhibiting mitochondrial glycerophosphate dehydrogenase. Nature 2014, 510, 542–546. [Google Scholar] [CrossRef] [PubMed]
- Bonora, E.; Cigolini, M.; Bosello, O.; Zancanaro, C.; Capretti, L.; Zavaroni, I.; Coscelli, C.; Butturini, U. Lack of effect of intravenous metformin on plasma concentrations of glucose, insulin, c-peptide, glucagon and growth hormone in non-diabetic subjects. Curr. Med. Res. Opin. 1984, 9, 47–51. [Google Scholar] [CrossRef]
- Stepensky, D.; Friedman, M.; Raz, I.; Hoffman, A. Pharmacokinetic-pharmacodynamic analysis of the glucose-lowering effect of metformin in diabetic rats reveals first-pass pharmacodynamic effect. Drug Metab. Dispos. 2002, 30, 861–868. [Google Scholar] [CrossRef]
- Bailey, C.J.; Mynett, K.J.; Page, T. Importance of the intestine as a site of metformin-stimulated glucose utilization. Br. J. Pharmacol. 1994, 112, 671–675. [Google Scholar] [CrossRef]
- Bailey, C.J.; Wilcock, C.; Scarpello, J.H. Metformin and the intestine. Diabetologia 2008, 51, 1552–1553. [Google Scholar] [CrossRef]
- Tucker, G.T.; Casey, C.; Phillips, P.J.; Connor, H.; Ward, J.D.; Woods, H.F. Metformin kinetics in healthy subjects and in patients with diabetes mellitus. Br. J. Clin. Pharmacol. 1981, 12, 235–246. [Google Scholar] [CrossRef]
- Wilcock, C.; Bailey, C.J. Accumulation of metformin by tissues of the normal and diabetic mouse. Xenobiotica 1994, 24, 49–57. [Google Scholar] [CrossRef]
- Jensen, J.B.; Sundelin, E.I.; Jakobsen, S.; Gormsen, L.C.; Munk, O.L.; Frokiaer, J.; Jessen, N. [11c]-labeled metformin distribution in the liver and small intestine using dynamic positron emission tomography in mice demonstrates tissue-specific transporter dependency. Diabetes 2016, 65, 1724–1730. [Google Scholar] [CrossRef]
- Dujic, T.; Zhou, K.; Donnelly, L.A.; Tavendale, R.; Palmer, C.N.; Pearson, E.R. Association of organic cation transporter 1 with intolerance to metformin in type 2 diabetes: A godarts study. Diabetes 2015, 64, 1786–1793. [Google Scholar] [CrossRef]
- Buse, J.B.; DeFronzo, R.A.; Rosenstock, J.; Kim, T.; Burns, C.; Skare, S.; Baron, A.; Fineman, M. The primary glucose-lowering effect of metformin resides in the gut, not the circulation: Results from short-term pharmacokinetic and 12-week dose-ranging studies. Diabetes Care 2016, 39, 198–205. [Google Scholar] [CrossRef]
- Kho, Z.Y.; Lal, S.K. The human gut microbiome—A potential controller of wellness and disease. Front. Microbiol. 2018, 9, 1835. [Google Scholar] [CrossRef]
- Hooper, L.V.; Gordon, J.I. Commensal host-bacterial relationships in the gut. Science 2001, 292, 1115–1118. [Google Scholar] [CrossRef]
- Le Chatelier, E.; Nielsen, T.; Qin, J.; Prifti, E.; Hildebrand, F.; Falony, G.; Almeida, M.; Arumugam, M.; Batto, J.M.; Kennedy, S.; et al. Richness of human gut microbiome correlates with metabolic markers. Nature 2013, 500, 541–546. [Google Scholar] [CrossRef]
- Qin, J.; Li, Y.; Cai, Z.; Li, S.; Zhu, J.; Zhang, F.; Liang, S.; Zhang, W.; Guan, Y.; Shen, D.; et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 2012, 490, 55–60. [Google Scholar] [CrossRef]
- Sato, J.; Kanazawa, A.; Ikeda, F.; Yoshihara, T.; Goto, H.; Abe, H.; Komiya, K.; Kawaguchi, M.; Shimizu, T.; Ogihara, T.; et al. Gut dysbiosis and detection of “live gut bacteria” in blood of japanese patients with type 2 diabetes. Diabetes Care 2014, 37, 2343–2350. [Google Scholar] [CrossRef]
- Karlsson, F.H.; Tremaroli, V.; Nookaew, I.; Bergstrom, G.; Behre, C.J.; Fagerberg, B.; Nielsen, J.; Backhed, F. Gut metagenome in european women with normal, impaired and diabetic glucose control. Nature 2013, 498, 99–103. [Google Scholar] [CrossRef]
- Larsen, N.; Vogensen, F.K.; van den Berg, F.W.; Nielsen, D.S.; Andreasen, A.S.; Pedersen, B.K.; Al-Soud, W.A.; Sorensen, S.J.; Hansen, L.H.; Jakobsen, M. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS ONE 2010, 5, e9085. [Google Scholar] [CrossRef]
- Forslund, K.; Hildebrand, F.; Nielsen, T.; Falony, G.; Le Chatelier, E.; Sunagawa, S.; Prifti, E.; Vieira-Silva, S.; Gudmundsdottir, V.; Pedersen, H.K.; et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 2015, 528, 262–266. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Shen, D.; Fang, Z.; Jie, Z.; Qiu, X.; Zhang, C.; Chen, Y.; Ji, L. Human gut microbiota changes reveal the progression of glucose intolerance. PLoS ONE 2013, 8, e71108. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Wang, M.; Yang, J.; Xu, Q.; Liang, C.; Chen, B.; Zhang, J.; Yang, Y.; Wang, H.; Shang, Y.; et al. Response of gut microbiota in type 2 diabetes to hypoglycemic agents. Endocrine 2019, 66, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, A.; Yang, W.; Chen, G.; Shafiq, M.; Javed, S.; Ali Zaidi, S.S.; Shahid, R.; Liu, C.; Bokhari, H. Analysis of gut microbiota of obese individuals with type 2 diabetes and healthy individuals. PLoS ONE 2019, 14, e0226372. [Google Scholar] [CrossRef] [PubMed]
- Chavez-Carbajal, A.; Pizano-Zarate, M.L.; Hernandez-Quiroz, F.; Ortiz-Luna, G.F.; Morales-Hernandez, R.M.; De Sales-Millan, A.; Hernandez-Trejo, M.; Garcia-Vite, A.; Beltran-Lagunes, L.; Hoyo-Vadillo, C.; et al. Characterization of the gut microbiota of individuals at different t2d stages reveals a complex relationship with the host. Microorganisms 2020, 8, 94. [Google Scholar] [CrossRef]
- Gurung, M.; Li, Z.; You, H.; Rodrigues, R.; Jump, D.B.; Morgun, A.; Shulzhenko, N. Role of gut microbiota in type 2 diabetes pathophysiology. EBioMedicine 2020, 51, 102590. [Google Scholar] [CrossRef]
- Rodriguez, J.; Hiel, S.; Delzenne, N.M. Metformin: Old friend, new ways of action-implication of the gut microbiome? Curr. Opin. Clin. Nutr. Metab. Care 2018, 21, 294–301. [Google Scholar] [CrossRef]
- Wu, T.; Horowitz, M.; Rayner, C.K. New insights into the anti-diabetic actions of metformin: From the liver to the gut. Expert Rev. Gastroenterol. Hepatol. 2017, 11, 157–166. [Google Scholar] [CrossRef]
- Hur, K.Y.; Lee, M.S. New mechanisms of metformin action: Focusing on mitochondria and the gut. J. Diabetes Investig. 2015, 6, 600–609. [Google Scholar] [CrossRef]
- Vallianou, N.G.; Stratigou, T.; Tsagarakis, S. Metformin and gut microbiota: Their interactions and their impact on diabetes. Hormones 2019, 18, 141–144. [Google Scholar] [CrossRef]
- Zhang, Q.; Hu, N. Effects of metformin on the gut microbiota in obesity and type 2 diabetes mellitus. Diabetes Metab. Syndr. Obes. 2020, 13, 5003–5014. [Google Scholar] [CrossRef]
- Morgan, X.C.; Tickle, T.L.; Sokol, H.; Gevers, D.; Devaney, K.L.; Ward, D.V.; Reyes, J.A.; Shah, S.A.; LeLeiko, N.; Snapper, S.B.; et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 2012, 13, R79. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Hamady, M.; Yatsunenko, T.; Cantarel, B.L.; Duncan, A.; Ley, R.E.; Sogin, M.L.; Jones, W.J.; Roe, B.A.; Affourtit, J.P.; et al. A core gut microbiome in obese and lean twins. Nature 2009, 457, 480–484. [Google Scholar] [CrossRef]
- Koliada, A.; Syzenko, G.; Moseiko, V.; Budovska, L.; Puchkov, K.; Perederiy, V.; Gavalko, Y.; Dorofeyev, A.; Romanenko, M.; Tkach, S.; et al. Association between body mass index and firmicutes/bacteroidetes ratio in an adult ukrainian population. BMC Microbiol. 2017, 17, 120. [Google Scholar] [CrossRef]
- Vrieze, A.; Van Nood, E.; Holleman, F.; Salojarvi, J.; Kootte, R.S.; Bartelsman, J.F.; Dallinga-Thie, G.M.; Ackermans, M.T.; Serlie, M.J.; Oozeer, R.; et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology 2012, 143, 913–916.e7. [Google Scholar] [CrossRef]
- Halawa, M.R.; El-Salam, M.A.; Mostafa, B.M.; Sallout, S.S. The gut microbiome, lactobacillus acidophilus; relation with type 2 diabetes mellitus. Curr. Diabetes Rev. 2019, 15, 480–485. [Google Scholar] [CrossRef]
- Zeuthen, L.H.; Christensen, H.R.; Frokiaer, H. Lactic acid bacteria inducing a weak interleukin-12 and tumor necrosis factor alpha response in human dendritic cells inhibit strongly stimulating lactic acid bacteria but act synergistically with gram-negative bacteria. Clin. Vaccine Immunol. 2006, 13, 365–375. [Google Scholar] [CrossRef][Green Version]
- Cani, P.D.; Amar, J.; Iglesias, M.A.; Poggi, M.; Knauf, C.; Bastelica, D.; Neyrinck, A.M.; Fava, F.; Tuohy, K.M.; Chabo, C.; et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 2007, 56, 1761–1772. [Google Scholar] [CrossRef]
- Delzenne, N.M.; Cani, P.D.; Everard, A.; Neyrinck, A.M.; Bindels, L.B. Gut microorganisms as promising targets for the management of type 2 diabetes. Diabetologia 2015, 58, 2206–2217. [Google Scholar] [CrossRef]
- Carvalho, B.M.; Guadagnini, D.; Tsukumo, D.M.L.; Schenka, A.A.; Latuf-Filho, P.; Vassallo, J.; Dias, J.C.; Kubota, L.T.; Carvalheira, J.B.C.; Saad, M.J.A. Modulation of gut microbiota by antibiotics improves insulin signalling in high-fat fed mice. Diabetologia 2012, 55, 2823–2834. [Google Scholar] [CrossRef]
- Song, M.J.; Kim, K.H.; Yoon, J.M.; Kim, J.B. Activation of toll-like receptor 4 is associated with insulin resistance in adipocytes. Biochem. Biophys. Res. Commun. 2006, 346, 739–745. [Google Scholar] [CrossRef] [PubMed]
- Balakumar, M.; Prabhu, D.; Sathishkumar, C.; Prabu, P.; Rokana, N.; Kumar, R.; Raghavan, S.; Soundarajan, A.; Grover, S.; Batish, V.K.; et al. Improvement in glucose tolerance and insulin sensitivity by probiotic strains of indian gut origin in high-fat diet-fed c57bl/6j mice. Eur. J. Nutr. 2018, 57, 279–295. [Google Scholar] [CrossRef] [PubMed]
- Panwar, H.; Rashmi, H.M.; Batish, V.K.; Grover, S. Probiotics as potential biotherapeutics in the management of type 2 diabetes—Prospects and perspectives. Diabetes Metab. Res. Rev. 2013, 29, 103–112. [Google Scholar] [CrossRef]
- Napolitano, A.; Miller, S.; Nicholls, A.W.; Baker, D.; Van Horn, S.; Thomas, E.; Rajpal, D.; Spivak, A.; Brown, J.R.; Nunez, D.J. Novel gut-based pharmacology of metformin in patients with type 2 diabetes mellitus. PLoS ONE 2014, 9, e100778. [Google Scholar] [CrossRef]
- Wu, H.; Esteve, E.; Tremaroli, V.; Khan, M.T.; Caesar, R.; Manneras-Holm, L.; Stahlman, M.; Olsson, L.M.; Serino, M.; Planas-Felix, M.; et al. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat. Med. 2017, 23, 850–858. [Google Scholar] [CrossRef]
- De la Cuesta-Zuluaga, J.; Mueller, N.T.; Corrales-Agudelo, V.; Velasquez-Mejia, E.P.; Carmona, J.A.; Abad, J.M.; Escobar, J.S. Metformin is associated with higher relative abundance of mucin-degrading akkermansia muciniphila and several short-chain fatty acid-producing microbiota in the gut. Diabetes Care 2017, 40, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.; Nilholm, C.; Roth, B.; Linninge, C.; Hoglund, P.; Nyman, M.; Ohlsson, B. Anthropometric and metabolic improvements in human type 2 diabetes after introduction of an okinawan-based nordic diet are not associated with changes in microbial diversity or scfa concentrations. Int. J. Food Sci. Nutr. 2018, 69, 729–740. [Google Scholar] [CrossRef]
- Sun, L.; Xie, C.; Wang, G.; Wu, Y.; Wu, Q.; Wang, X.; Liu, J.; Deng, Y.; Xia, J.; Chen, B.; et al. Gut microbiota and intestinal fxr mediate the clinical benefits of metformin. Nat. Med. 2018, 24, 1919–1929. [Google Scholar] [CrossRef]
- Elbere, I.; Kalnina, I.; Silamikelis, I.; Konrade, I.; Zaharenko, L.; Sekace, K.; Radovica-Spalvina, I.; Fridmanis, D.; Gudra, D.; Pirags, V.; et al. Association of metformin administration with gut microbiome dysbiosis in healthy volunteers. PLoS ONE 2018, 13, e0204317. [Google Scholar] [CrossRef]
- Bryrup, T.; Thomsen, C.W.; Kern, T.; Allin, K.H.; Brandslund, I.; Jorgensen, N.R.; Vestergaard, H.; Hansen, T.; Hansen, T.H.; Pedersen, O.; et al. Metformin-induced changes of the gut microbiota in healthy young men: Results of a non-blinded, one-armed intervention study. Diabetologia 2019, 62, 1024–1035. [Google Scholar] [CrossRef] [PubMed]
- Lam, T.K. Neuronal regulation of homeostasis by nutrient sensing. Nat. Med. 2010, 16, 392–395. [Google Scholar] [CrossRef]
- Duca, F.A.; Bauer, P.V.; Hamr, S.C.; Lam, T.K. Glucoregulatory relevance of small intestinal nutrient sensing in physiology, bariatric surgery, and pharmacology. Cell Metab. 2015, 22, 367–380. [Google Scholar] [CrossRef] [PubMed]
- Gorboulev, V.; Schurmann, A.; Vallon, V.; Kipp, H.; Jaschke, A.; Klessen, D.; Friedrich, A.; Scherneck, S.; Rieg, T.; Cunard, R.; et al. Na(+)-d-glucose cotransporter sglt1 is pivotal for intestinal glucose absorption and glucose-dependent incretin secretion. Diabetes 2012, 61, 187–196. [Google Scholar] [CrossRef] [PubMed]
- Gribble, F.M.; Williams, L.; Simpson, A.K.; Reimann, F. A novel glucose-sensing mechanism contributing to glucagon-like peptide-1 secretion from the glutag cell line. Diabetes 2003, 52, 1147–1154. [Google Scholar] [CrossRef]
- Kuhre, R.E.; Frost, C.R.; Svendsen, B.; Holst, J.J. Molecular mechanisms of glucose-stimulated glp-1 secretion from perfused rat small intestine. Diabetes 2015, 64, 370–382. [Google Scholar] [CrossRef]
- Moriya, R.; Shirakura, T.; Ito, J.; Mashiko, S.; Seo, T. Activation of sodium-glucose cotransporter 1 ameliorates hyperglycemia by mediating incretin secretion in mice. Am. J. Physiol. Endocrinol. Metab. 2009, 297, E1358–E1365. [Google Scholar] [CrossRef]
- Parker, H.E.; Adriaenssens, A.; Rogers, G.; Richards, P.; Koepsell, H.; Reimann, F.; Gribble, F.M. Predominant role of active versus facilitative glucose transport for glucagon-like peptide-1 secretion. Diabetologia 2012, 55, 2445–2455. [Google Scholar] [CrossRef]
- Maida, A.; Lamont, B.J.; Cao, X.; Drucker, D.J. Metformin regulates the incretin receptor axis via a pathway dependent on peroxisome proliferator-activated receptor-alpha in mice. Diabetologia 2011, 54, 339–349. [Google Scholar] [CrossRef]
- Vardarli, I.; Arndt, E.; Deacon, C.F.; Holst, J.J.; Nauck, M.A. Effects of sitagliptin and metformin treatment on incretin hormone and insulin secretory responses to oral and “isoglycemic” intravenous glucose. Diabetes 2014, 63, 663–674. [Google Scholar] [CrossRef]
- Duca, F.A.; Cote, C.D.; Rasmussen, B.A.; Zadeh-Tahmasebi, M.; Rutter, G.A.; Filippi, B.M.; Lam, T.K. Metformin activates a duodenal ampk-dependent pathway to lower hepatic glucose production in rats. Nat. Med. 2015, 21, 506–511. [Google Scholar] [CrossRef]
- Lenzen, S.; Lortz, S.; Tiedge, M. Effect of metformin on sglt1, glut2, and glut5 hexose transporter gene expression in small intestine from rats. Biochem. Pharmacol. 1996, 51, 893–896. [Google Scholar] [CrossRef]
- El Aidy, S.; Merrifield, C.A.; Derrien, M.; van Baarlen, P.; Hooiveld, G.; Levenez, F.; Dore, J.; Dekker, J.; Holmes, E.; Claus, S.P.; et al. The gut microbiota elicits a profound metabolic reorientation in the mouse jejunal mucosa during conventionalisation. Gut 2013, 62, 1306–1314. [Google Scholar] [CrossRef] [PubMed]
- Ejtahed, H.S.; Mohtadi-Nia, J.; Homayouni-Rad, A.; Niafar, M.; Asghari-Jafarabadi, M.; Mofid, V. Probiotic yogurt improves antioxidant status in type 2 diabetic patients. Nutrition 2012, 28, 539–543. [Google Scholar] [CrossRef] [PubMed]
- Yadav, H.; Jain, S.; Sinha, P.R. Antidiabetic effect of probiotic dahi containing lactobacillus acidophilus and lactobacillus casei in high fructose fed rats. Nutrition 2007, 23, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Bauer, P.V.; Duca, F.A.; Waise, T.M.Z.; Rasmussen, B.A.; Abraham, M.A.; Dranse, H.J.; Puri, A.; O’Brien, C.A.; Lam, T.K.T. Metformin alters upper small intestinal microbiota that impact a glucose-sglt1-sensing glucoregulatory pathway. Cell Metab. 2018, 27, 101–117.e105. [Google Scholar] [CrossRef]
- Rooj, A.K.; Kimura, Y.; Buddington, R.K. Metabolites produced by probiotic lactobacilli rapidly increase glucose uptake by caco-2 cells. BMC Microbiol. 2010, 10, 16. [Google Scholar] [CrossRef]
- Fredborg, M.; Theil, P.K.; Jensen, B.B.; Purup, S. G protein-coupled receptor120 (gpr120) transcription in intestinal epithelial cells is significantly affected by bacteria belonging to the bacteroides, proteobacteria, and firmicutes phyla. J. Anim. Sci. 2012, 90 (Suppl. 4), 10–12. [Google Scholar] [CrossRef]
- Tanaka, T.; Katsuma, S.; Adachi, T.; Koshimizu, T.A.; Hirasawa, A.; Tsujimoto, G. Free fatty acids induce cholecystokinin secretion through gpr120. Naunyn-Schmiedeberg’s Arch. Pharmacol. 2008, 377, 523–527. [Google Scholar] [CrossRef]
- Bauer, P.V.; Duca, F.A.; Waise, T.M.Z.; Dranse, H.J.; Rasmussen, B.A.; Puri, A.; Rasti, M.; O’Brien, C.A.; Lam, T.K.T. Lactobacillus gasseri in the upper small intestine impacts an acsl3-dependent fatty acid-sensing pathway regulating whole-body glucose homeostasis. Cell Metab. 2018, 27, 572–587.e6. [Google Scholar] [CrossRef]
- Den Besten, G.; van Eunen, K.; Groen, A.K.; Venema, K.; Reijngoud, D.J.; Bakker, B.M. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 2013, 54, 2325–2340. [Google Scholar] [CrossRef]
- Karaki, S.; Tazoe, H.; Hayashi, H.; Kashiwabara, H.; Tooyama, K.; Suzuki, Y.; Kuwahara, A. Expression of the short-chain fatty acid receptor, gpr43, in the human colon. J. Mol. Histol. 2008, 39, 135–142. [Google Scholar] [CrossRef]
- Karaki, S.; Mitsui, R.; Hayashi, H.; Kato, I.; Sugiya, H.; Iwanaga, T.; Furness, J.B.; Kuwahara, A. Short-chain fatty acid receptor, gpr43, is expressed by enteroendocrine cells and mucosal mast cells in rat intestine. Cell Tissue Res. 2006, 324, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Tolhurst, G.; Heffron, H.; Lam, Y.S.; Parker, H.E.; Habib, A.M.; Diakogiannaki, E.; Cameron, J.; Grosse, J.; Reimann, F.; Gribble, F.M. Short-chain fatty acids stimulate glucagon-like peptide-1 secretion via the g-protein-coupled receptor ffar2. Diabetes 2012, 61, 364–371. [Google Scholar] [CrossRef]
- Cherbut, C.; Ferrier, L.; Roze, C.; Anini, Y.; Blottiere, H.; Lecannu, G.; Galmiche, J.P. Short-chain fatty acids modify colonic motility through nerves and polypeptide yy release in the rat. Am. J. Physiol. 1998, 275, G1415–G1422. [Google Scholar] [CrossRef] [PubMed]
- Holz, G.G., IV; Kuhtreiber, W.M.; Habener, J.F. Pancreatic beta-cells are rendered glucose-competent by the insulinotropic hormone glucagon-like peptide-1(7-37). Nature 1993, 361, 362–365. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 2021, 19, 55–71. [Google Scholar] [CrossRef]
- Lee, H.; Ko, G. Effect of metformin on metabolic improvement and gut microbiota. Appl. Environ. Microbiol. 2014, 80, 5935–5943. [Google Scholar] [CrossRef]
- Lee, H.; Lee, Y.; Kim, J.; An, J.; Lee, S.; Kong, H.; Song, Y.; Lee, C.K.; Kim, K. Modulation of the gut microbiota by metformin improves metabolic profiles in aged obese mice. Gut Microbes 2018, 9, 155–165. [Google Scholar] [CrossRef]
- Ryan, P.M.; Patterson, E.; Carafa, I.; Mandal, R.; Wishart, D.S.; Dinan, T.G.; Cryan, J.F.; Tuohy, K.M.; Stanton, C.; Ross, R.P. Metformin and dipeptidyl peptidase-4 inhibitor differentially modulate the intestinal microbiota and plasma metabolome of metabolically dysfunctional mice. Can. J. Diabetes 2020, 44, 146–155.e2. [Google Scholar] [CrossRef]
- Zhang, W.; Xu, J.H.; Yu, T.; Chen, Q.K. Effects of berberine and metformin on intestinal inflammation and gut microbiome composition in db/db mice. Biomed. Pharmacother. 2019, 118, 109131. [Google Scholar] [CrossRef] [PubMed]
- Ahmadi, S.; Razazan, A.; Nagpal, R.; Jain, S.; Wang, B.; Mishra, S.P.; Wang, S.; Justice, J.; Ding, J.; McClain, D.A.; et al. Metformin reduces aging-related leaky gut and improves cognitive function by beneficially modulating gut microbiome/goblet cell/mucin axis. J. Gerontol. A Biol. Sci. Med. Sci. 2020, 75, e9–e21. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Liang, L.; Yu, G.; Li, Q. Pumpkin polysaccharide modifies the gut microbiota during alleviation of type 2 diabetes in rats. Int. J. Biol. Macromol. 2018, 115, 711–717. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, C.; Li, J.; Li, T.; Zhang, Y.; Liang, Y.; Mei, Y. Phellinus linteus polysaccharide extract improves insulin resistance by regulating gut microbiota composition. FASEB J. 2020, 34, 1065–1078. [Google Scholar] [CrossRef] [PubMed]
- Macfarlane, S.; Macfarlane, G.T. Regulation of short-chain fatty acid production. Proc. Nutr. Soc. 2003, 62, 67–72. [Google Scholar] [CrossRef]
- Rios-Covian, D.; Arboleya, S.; Hernandez-Barranco, A.M.; Alvarez-Buylla, J.R.; Ruas-Madiedo, P.; Gueimonde, M.; de los Reyes-Gavilan, C.G. Interactions between bifidobacterium and bacteroides species in cofermentations are affected by carbon sources, including exopolysaccharides produced by bifidobacteria. Appl. Environ. Microbiol. 2013, 79, 7518–7524. [Google Scholar] [CrossRef]
- Hao, Z.; Li, L.; Ning, Z.; Zhang, X.; Mayne, J.; Cheng, K.; Walker, K.; Liu, H.; Figeys, D. Metaproteomics reveals growth phase-dependent responses of an in vitro gut microbiota to metformin. J. Am. Soc. Mass Spectrom. 2020, 31, 1448–1458. [Google Scholar] [CrossRef]
- Pryor, R.; Norvaisas, P.; Marinos, G.; Best, L.; Thingholm, L.B.; Quintaneiro, L.M.; De Haes, W.; Esser, D.; Waschina, S.; Lujan, C.; et al. Host-microbe-drug-nutrient screen identifies bacterial effectors of metformin therapy. Cell 2019, 178, 1299–1312.e29. [Google Scholar] [CrossRef]
- Brandt, A.; Hernandez-Arriaga, A.; Kehm, R.; Sanchez, V.; Jin, C.J.; Nier, A.; Baumann, A.; Camarinha-Silva, A.; Bergheim, I. Metformin attenuates the onset of non-alcoholic fatty liver disease and affects intestinal microbiota and barrier in small intestine. Sci. Rep. 2019, 9, 6668. [Google Scholar] [CrossRef]
- Wankhade, U.D.; Zhong, Y.; Lazarenko, O.P.; Chintapalli, S.V.; Piccolo, B.D.; Chen, J.R.; Shankar, K. Sex-specific changes in gut microbiome composition following blueberry consumption in c57bl/6j mice. Nutrients 2019, 11, 313. [Google Scholar] [CrossRef]
- Caslin, B.; Maguire, C.; Karmakar, A.; Mohler, K.; Wylie, D.; Melamed, E. Alcohol shifts gut microbial networks and ameliorates a murine model of neuroinflammation in a sex-specific pattern. Proc. Natl. Acad. Sci. USA 2019, 116, 25808–25815. [Google Scholar] [CrossRef]
- Geer, E.B.; Shen, W. Gender differences in insulin resistance, body composition, and energy balance. Gend. Med. 2009, 6 (Suppl. 1), 60–75. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.N.; O’Sullivan, A.J. Sex differences in energy metabolism need to be considered with lifestyle modifications in humans. J. Nutr. Metab. 2011, 2011, 391809. [Google Scholar] [CrossRef] [PubMed]
- Kornman, K.S.; Loesche, W.J. Effects of estradiol and progesterone on bacteroides melaninogenicus and bacteroides gingivalis. Infect. Immun. 1982, 35, 256–263. [Google Scholar] [CrossRef]
- Gao, Z.; Yin, J.; Zhang, J.; Ward, R.E.; Martin, R.J.; Lefevre, M.; Cefalu, W.T.; Ye, J. Butyrate improves insulin sensitivity and increases energy expenditure in mice. Diabetes 2009, 58, 1509–1517. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.V.; Frassetto, A.; Kowalik, E.J., Jr.; Nawrocki, A.R.; Lu, M.M.; Kosinski, J.R.; Hubert, J.A.; Szeto, D.; Yao, X.; Forrest, G.; et al. Butyrate and propionate protect against diet-induced obesity and regulate gut hormones via free fatty acid receptor 3-independent mechanisms. PLoS ONE 2012, 7, e35240. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, E.; Yin, B.; Fang, D.; Chen, P.; Wang, G.; Zhao, J.; Zhang, H.; Chen, W. Effects of lactobacillus casei ccfm419 on insulin resistance and gut microbiota in type 2 diabetic mice. Benef. Microbes 2017, 8, 421–432. [Google Scholar] [CrossRef]
- Wang, K.; Liao, M.; Zhou, N.; Bao, L.; Ma, K.; Zheng, Z.; Wang, Y.; Liu, C.; Wang, W.; Wang, J.; et al. Parabacteroides distasonis alleviates obesity and metabolic dysfunctions via production of succinate and secondary bile acids. Cell Rep. 2019, 26, 222–235.e5. [Google Scholar] [CrossRef]
- Zheng, J.; Li, H.; Zhang, X.; Jiang, M.; Luo, C.; Lu, Z.; Xu, Z.; Shi, J. Prebiotic mannan-oligosaccharides augment the hypoglycemic effects of metformin in correlation with modulating gut microbiota. J. Agric. Food Chem. 2018, 66, 5821–5831. [Google Scholar] [CrossRef]
- Shin, N.R.; Lee, J.C.; Lee, H.Y.; Kim, M.S.; Whon, T.W.; Lee, M.S.; Bae, J.W. An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 2014, 63, 727–735. [Google Scholar] [CrossRef]
- Cani, P.D.; Bibiloni, R.; Knauf, C.; Waget, A.; Neyrinck, A.M.; Delzenne, N.M.; Burcelin, R. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 2008, 57, 1470–1481. [Google Scholar] [CrossRef]
- Cani, P.D.; Neyrinck, A.M.; Fava, F.; Knauf, C.; Burcelin, R.G.; Tuohy, K.M.; Gibson, G.R.; Delzenne, N.M. Selective increases of bifidobacteria in gut microflora improve high-fat-diet-induced diabetes in mice through a mechanism associated with endotoxaemia. Diabetologia 2007, 50, 2374–2383. [Google Scholar] [CrossRef]
- Tanti, J.F.; Gual, P.; Gremeaux, T.; Gonzalez, T.; Barres, R.; Le Marchand-Brustel, Y. Alteration in insulin action: Role of irs-1 serine phosphorylation in the retroregulation of insulin signalling. Ann. Endocrinol. 2004, 65, 43–48. [Google Scholar] [CrossRef]
- Macchione, I.G.; Lopetuso, L.R.; Ianiro, G.; Napoli, M.; Gibiino, G.; Rizzatti, G.; Petito, V.; Gasbarrini, A.; Scaldaferri, F. Akkermansia muciniphila: Key player in metabolic and gastrointestinal disorders. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 8075–8083. [Google Scholar] [PubMed]
- Kim, Y.S.; Ho, S.B. Intestinal goblet cells and mucins in health and disease: Recent insights and progress. Curr. Gastroenterol. Rep. 2010, 12, 319–330. [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.E.; Larsson, J.M.; Hansson, G.C. The two mucus layers of colon are organized by the muc2 mucin, whereas the outer layer is a legislator of host-microbial interactions. Proc. Natl. Acad. Sci. USA 2011, 108 (Suppl. 1), 4659–4665. [Google Scholar] [CrossRef] [PubMed]
- Amar, J.; Chabo, C.; Waget, A.; Klopp, P.; Vachoux, C.; Bermudez-Humaran, L.G.; Smirnova, N.; Berge, M.; Sulpice, T.; Lahtinen, S.; et al. Intestinal mucosal adherence and translocation of commensal bacteria at the early onset of type 2 diabetes: Molecular mechanisms and probiotic treatment. EMBO Mol. Med. 2011, 3, 559–572. [Google Scholar] [CrossRef]
- Everard, A.; Belzer, C.; Geurts, L.; Ouwerkerk, J.P.; Druart, C.; Bindels, L.B.; Guiot, Y.; Derrien, M.; Muccioli, G.G.; Delzenne, N.M.; et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl. Acad. Sci. USA 2013, 110, 9066–9071. [Google Scholar] [CrossRef]
- Belzer, C.; de Vos, W.M. Microbes inside—From diversity to function: The case of Akkermansia. ISME J. 2012, 6, 1449–1458. [Google Scholar] [CrossRef]
- Van Passel, M.W.; Kant, R.; Zoetendal, E.G.; Plugge, C.M.; Derrien, M.; Malfatti, S.A.; Chain, P.S.; Woyke, T.; Palva, A.; de Vos, W.M.; et al. The genome of akkermansia muciniphila, a dedicated intestinal mucin degrader, and its use in exploring intestinal metagenomes. PLoS ONE 2011, 6, e16876. [Google Scholar] [CrossRef]
- Ma, W.; Chen, J.; Meng, Y.; Yang, J.; Cui, Q.; Zhou, Y. Metformin alters gut microbiota of healthy mice: Implication for its potential role in gut microbiota homeostasis. Front. Microbiol. 2018, 9, 1336. [Google Scholar] [CrossRef]
- Rosario, D.; Benfeitas, R.; Bidkhori, G.; Zhang, C.; Uhlen, M.; Shoaie, S.; Mardinoglu, A. Understanding the representative gut microbiota dysbiosis in metformin-treated type 2 diabetes patients using genome-scale metabolic modeling. Front. Physiol. 2018, 9, 775. [Google Scholar] [CrossRef]
- Zhang, X.; Zhao, Y.; Xu, J.; Xue, Z.; Zhang, M.; Pang, X.; Zhang, X.; Zhao, L. Modulation of gut microbiota by berberine and metformin during the treatment of high-fat diet-induced obesity in rats. Sci. Rep. 2015, 5, 14405. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.X.; Zhang, L.S.; Luo, Y.; Yuan, K.; Huang, Z.Y.; Guo, Y. A purified anthraquinone-glycoside preparation from rhubarb ameliorates type 2 diabetes mellitus by modulating the gut microbiota and reducing inflammation. Front. Microbiol. 2019, 10, 1423. [Google Scholar] [CrossRef] [PubMed]
- Ji, S.; Wang, L.; Li, L. Effect of metformin on short-term high-fat diet-induced weight gain and anxiety-like behavior and the gut microbiota. Front. Endocrinol. 2019, 10, 704. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.C.; Fan, Z.Y.; Wang, H.Y.; Wen, D.C.; Zhang, S.Y. Effect of polysaccharides from adlay seed on anti-diabetic and gut microbiota. Food Funct. 2019, 10, 4372–4380. [Google Scholar] [CrossRef]
- Wang, J.H.; Bose, S.; Shin, N.R.; Chin, Y.W.; Choi, Y.H.; Kim, H. Pharmaceutical impact of houttuynia cordata and metformin combination on high-fat-diet-induced metabolic disorders: Link to intestinal microbiota and metabolic endotoxemia. Front. Endocrinol. 2018, 9, 620. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Bei, J.; Liang, L.; Yu, G.; Li, L.; Li, Q. Stachyose improves inflammation through modulating gut microbiota of high-fat diet/streptozotocin-induced type 2 diabetes in rats. Mol. Nutr. Food Res. 2018, 62, e1700954. [Google Scholar] [CrossRef]
- Khat-Udomkiri, N.; Toejing, P.; Sirilun, S.; Chaiyasut, C.; Lailerd, N. Antihyperglycemic effect of rice husk derived xylooligosaccharides in high-fat diet and low-dose streptozotocin-induced type 2 diabetic rat model. Food Sci. Nutr. 2020, 8, 428–444. [Google Scholar] [CrossRef]
- Wang, J.H.; Bose, S.; Lim, S.K.; Ansari, A.; Chin, Y.W.; Choi, H.S.; Kim, H. Houttuynia cordata facilitates metformin on ameliorating insulin resistance associated with gut microbiota alteration in oletf rats. Genes 2017, 8, 239. [Google Scholar] [CrossRef]
- Zhang, M.; Feng, R.; Yang, M.; Qian, C.; Wang, Z.; Liu, W.; Ma, J. Effects of metformin, acarbose, and sitagliptin monotherapy on gut microbiota in zucker diabetic fatty rats. BMJ Open Diabetes Res. Care 2019, 7, e000717. [Google Scholar] [CrossRef]
- Chelakkot, C.; Choi, Y.; Kim, D.K.; Park, H.T.; Ghim, J.; Kwon, Y.; Jeon, J.; Kim, M.S.; Jee, Y.K.; Gho, Y.S.; et al. Akkermansia muciniphila-derived extracellular vesicles influence gut permeability through the regulation of tight junctions. Exp. Mol. Med. 2018, 50, e450. [Google Scholar] [CrossRef]
- Derrien, M.; Collado, M.C.; Ben-Amor, K.; Salminen, S.; de Vos, W.M. The mucin degrader Akkermansia muciniphila is an abundant resident of the human intestinal tract. Appl. Environ. Microbiol. 2008, 74, 1646–1648. [Google Scholar] [CrossRef] [PubMed]
- Plovier, H.; Everard, A.; Druart, C.; Depommier, C.; Van Hul, M.; Geurts, L.; Chilloux, J.; Ottman, N.; Duparc, T.; Lichtenstein, L.; et al. A purified membrane protein from Akkermansia muciniphila or the pasteurized bacterium improves metabolism in obese and diabetic mice. Nat. Med. 2017, 23, 107–113. [Google Scholar] [CrossRef] [PubMed]
- Artis, D.; Wang, M.L.; Keilbaugh, S.A.; He, W.; Brenes, M.; Swain, G.P.; Knight, P.A.; Donaldson, D.D.; Lazar, M.A.; Miller, H.R.; et al. Relmbeta/fizz2 is a goblet cell-specific immune-effector molecule in the gastrointestinal tract. Proc. Natl. Acad. Sci. USA 2004, 101, 13596–13600. [Google Scholar] [CrossRef] [PubMed]
- Suemori, S.; Lynch-Devaney, K.; Podolsky, D.K. Identification and characterization of rat intestinal trefoil factor: Tissue- and cell-specific member of the trefoil protein family. Proc. Natl. Acad. Sci. USA 1991, 88, 11017–11021. [Google Scholar] [CrossRef]
- Heilbronn, L.K.; Campbell, L.V. Adipose tissue macrophages, low grade inflammation and insulin resistance in human obesity. Curr. Pharm. Des. 2008, 14, 1225–1230. [Google Scholar] [CrossRef]
- Lumeng, C.N.; Saltiel, A.R. Inflammatory links between obesity and metabolic disease. J. Clin. Investig. 2011, 121, 2111–2117. [Google Scholar] [CrossRef]
- Osborn, O.; Olefsky, J.M. The cellular and signaling networks linking the immune system and metabolism in disease. Nat. Med. 2012, 18, 363–374. [Google Scholar] [CrossRef]
- Brestoff, J.R.; Artis, D. Immune regulation of metabolic homeostasis in health and disease. Cell 2015, 161, 146–160. [Google Scholar] [CrossRef]
- Lackey, D.E.; Olefsky, J.M. Regulation of metabolism by the innate immune system. Nat. Rev. Endocrinol. 2016, 12, 15–28. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.Y.; Lee, S.H.; Yang, E.J.; Kim, E.K.; Kim, J.K.; Shin, D.Y.; Cho, M.L. Metformin ameliorates inflammatory bowel disease by suppression of the stat3 signaling pathway and regulation of the between th17/treg balance. PLoS ONE 2015, 10, e0135858. [Google Scholar] [CrossRef]
- Li, S.N.; Wang, X.; Zeng, Q.T.; Feng, Y.B.; Cheng, X.; Mao, X.B.; Wang, T.H.; Deng, H.P. Metformin inhibits nuclear factor kappab activation and decreases serum high-sensitivity c-reactive protein level in experimental atherogenesis of rabbits. Heart Vessel. 2009, 24, 446–453. [Google Scholar] [CrossRef] [PubMed]
- Huang, N.L.; Chiang, S.H.; Hsueh, C.H.; Liang, Y.J.; Chen, Y.J.; Lai, L.P. Metformin inhibits tnf-alpha-induced ikappab kinase phosphorylation, ikappab-alpha degradation and il-6 production in endothelial cells through pi3k-dependent ampk phosphorylation. Int. J. Cardiol. 2009, 134, 169–175. [Google Scholar] [CrossRef]
- Png, C.W.; Linden, S.K.; Gilshenan, K.S.; Zoetendal, E.G.; McSweeney, C.S.; Sly, L.I.; McGuckin, M.A.; Florin, T.H. Mucolytic bacteria with increased prevalence in ibd mucosa augment in vitro utilization of mucin by other bacteria. Am. J. Gastroenterol. 2010, 105, 2420–2428. [Google Scholar] [CrossRef] [PubMed]
- Hansen, C.H.; Holm, T.L.; Krych, L.; Andresen, L.; Nielsen, D.S.; Rune, I.; Hansen, A.K.; Skov, S. Gut microbiota regulates nkg2d ligand expression on intestinal epithelial cells. Eur. J. Immunol. 2013, 43, 447–457. [Google Scholar] [CrossRef] [PubMed]
- Depommier, C.; Everard, A.; Druart, C.; Plovier, H.; Van Hul, M.; Vieira-Silva, S.; Falony, G.; Raes, J.; Maiter, D.; Delzenne, N.M.; et al. Supplementation with akkermansia muciniphila in overweight and obese human volunteers: A proof-of-concept exploratory study. Nat. Med. 2019, 25, 1096–1103. [Google Scholar] [CrossRef] [PubMed]
- Rotter, V.; Nagaev, I.; Smith, U. Interleukin-6 (il-6) induces insulin resistance in 3t3-l1 adipocytes and is, like il-8 and tumor necrosis factor-alpha, overexpressed in human fat cells from insulin-resistant subjects. J. Biol. Chem. 2003, 278, 45777–45784. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.S.; Ban, M.; Choi, E.J.; Moon, H.G.; Jeon, J.S.; Kim, D.K.; Park, S.K.; Jeon, S.G.; Roh, T.Y.; Myung, S.J.; et al. Extracellular vesicles derived from gut microbiota, especially Akkermansia muciniphila, protect the progression of dextran sulfate sodium-induced colitis. PLoS ONE 2013, 8, e76520. [Google Scholar] [CrossRef] [PubMed]
- Weigert, C.; Hennige, A.M.; Brodbeck, K.; Haring, H.U.; Schleicher, E.D. Interleukin-6 acts as insulin sensitizer on glycogen synthesis in human skeletal muscle cells by phosphorylation of ser473 of akt. Am. J. Physiol. Endocrinol. Metab. 2005, 289, E251–E257. [Google Scholar] [CrossRef]
- Bharti, A.C.; Aggarwal, B.B. Nuclear factor-kappa b and cancer: Its role in prevention and therapy. Biochem. Pharmacol. 2002, 64, 883–888. [Google Scholar] [CrossRef]
- Inan, M.S.; Rasoulpour, R.J.; Yin, L.; Hubbard, A.K.; Rosenberg, D.W.; Giardina, C. The luminal short-chain fatty acid butyrate modulates nf-kappab activity in a human colonic epithelial cell line. Gastroenterology 2000, 118, 724–734. [Google Scholar] [CrossRef]
- Kinoshita, M.; Suzuki, Y.; Saito, Y. Butyrate reduces colonic paracellular permeability by enhancing ppargamma activation. Biochem. Biophys. Res. Commun. 2002, 293, 827–831. [Google Scholar] [CrossRef]
- Li, X.; Wang, N.; Yin, B.; Fang, D.; Jiang, T.; Fang, S.; Zhao, J.; Zhang, H.; Wang, G.; Chen, W. Effects of lactobacillus plantarum ccfm0236 on hyperglycaemia and insulin resistance in high-fat and streptozotocin-induced type 2 diabetic mice. J. Appl. Microbiol. 2016, 121, 1727–1736. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Zhang, Q.; Dang, H.; Liu, X.; Tian, F.; Zhao, J.; Chen, Y.; Zhang, H.; Chen, W. Antidiabetic effect of lactobacillus casei ccfm0412 on mice with type 2 diabetes induced by a high-fat diet and streptozotocin. Nutrition 2014, 30, 1061–1068. [Google Scholar] [CrossRef] [PubMed]
- Tian, P.; Li, B.; He, C.; Song, W.; Hou, A.; Tian, S.; Meng, X.; Li, K.; Shan, Y. Antidiabetic (type 2) effects of lactobacillus g15 and q14 in rats through regulation of intestinal permeability and microbiota. Food Funct. 2016, 7, 3789–3797. [Google Scholar] [CrossRef]
- Sun, K.Y.; Xu, D.H.; Xie, C.; Plummer, S.; Tang, J.; Yang, X.F.; Ji, X.H. Lactobacillus paracasei modulates lps-induced inflammatory cytokine release by monocyte-macrophages via the up-regulation of negative regulators of nf-kappab signaling in a tlr2-dependent manner. Cytokine 2017, 92, 1–11. [Google Scholar] [CrossRef]
- Xue, J.; Li, X.; Liu, P.; Li, K.; Sha, L.; Yang, X.; Zhu, L.; Wang, Z.; Dong, Y.; Zhang, L.; et al. Inulin and metformin ameliorate polycystic ovary syndrome via anti-inflammation and modulating gut microbiota in mice. Endocr. J. 2019, 66, 859–870. [Google Scholar] [CrossRef]
- Li, T.; Chiang, J.Y. Bile acid signaling in metabolic disease and drug therapy. Pharmacol. Rev. 2014, 66, 948–983. [Google Scholar] [CrossRef]
- Sansome, D.J.; Xie, C.; Veedfald, S.; Horowitz, M.; Rayner, C.K.; Wu, T. Mechanism of glucose-lowering by metformin in type 2 diabetes: Role of bile acids. Diabetes Obes. Metab. 2020, 22, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Caspary, W.F.; Creutzfeldt, W. Inhibition of bile salt absorption by blood-sugar lowering biguanides. Diabetologia 1975, 11, 113–117. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Scarpello, J.H.; Hodgson, E.; Howlett, H.C. Effect of metformin on bile salt circulation and intestinal motility in type 2 diabetes mellitus. Diabet. Med. 1998, 15, 651–656. [Google Scholar] [CrossRef]
- Meng, X.M.; Ma, X.X.; Tian, Y.L.; Jiang, Q.; Wang, L.L.; Shi, R.; Ding, L.; Pang, S.G. Metformin improves the glucose and lipid metabolism via influencing the level of serum total bile acids in rats with streptozotocin-induced type 2 diabetes mellitus. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 2232–2237. [Google Scholar]
- Trabelsi, M.S.; Daoudi, M.; Prawitt, J.; Ducastel, S.; Touche, V.; Sayin, S.I.; Perino, A.; Brighton, C.A.; Sebti, Y.; Kluza, J.; et al. Farnesoid x receptor inhibits glucagon-like peptide-1 production by enteroendocrine l cells. Nat. Commun. 2015, 6, 7629. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Jiang, C.; Krausz, K.W.; Li, Y.; Albert, I.; Hao, H.; Fabre, K.M.; Mitchell, J.B.; Patterson, A.D.; Gonzalez, F.J. Microbiome remodelling leads to inhibition of intestinal farnesoid x receptor signalling and decreased obesity. Nat. Commun 2013, 4, 2384. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, M.; Horai, Y.; Houten, S.M.; Morimoto, K.; Sugizaki, T.; Arita, E.; Mataki, C.; Sato, H.; Tanigawara, Y.; Schoonjans, K.; et al. Lowering bile acid pool size with a synthetic farnesoid x receptor (fxr) agonist induces obesity and diabetes through reduced energy expenditure. J. Biol. Chem. 2011, 286, 26913–26920. [Google Scholar] [CrossRef]
- Sayin, S.I.; Wahlstrom, A.; Felin, J.; Jantti, S.; Marschall, H.U.; Bamberg, K.; Angelin, B.; Hyotylainen, T.; Oresic, M.; Backhed, F. Gut microbiota regulates bile acid metabolism by reducing the levels of tauro-beta-muricholic acid, a naturally occurring fxr antagonist. Cell Metab. 2013, 17, 225–235. [Google Scholar] [CrossRef]
- Pathak, P.; Xie, C.; Nichols, R.G.; Ferrell, J.M.; Boehme, S.; Krausz, K.W.; Patterson, A.D.; Gonzalez, F.J.; Chiang, J.Y.L. Intestine farnesoid x receptor agonist and the gut microbiota activate g-protein bile acid receptor-1 signaling to improve metabolism. Hepatology 2018, 68, 1574–1588. [Google Scholar] [CrossRef]
- Ma, K.; Saha, P.K.; Chan, L.; Moore, D.D. Farnesoid x receptor is essential for normal glucose homeostasis. J. Clin. Investig. 2006, 116, 1102–1109. [Google Scholar] [CrossRef]
- Cariou, B.; van Harmelen, K.; Duran-Sandoval, D.; van Dijk, T.H.; Grefhorst, A.; Abdelkarim, M.; Caron, S.; Torpier, G.; Fruchart, J.C.; Gonzalez, F.J.; et al. The farnesoid x receptor modulates adiposity and peripheral insulin sensitivity in mice. J. Biol. Chem. 2006, 281, 11039–11049. [Google Scholar] [CrossRef]
- Zhang, Y.; Lee, F.Y.; Barrera, G.; Lee, H.; Vales, C.; Gonzalez, F.J.; Willson, T.M.; Edwards, P.A. Activation of the nuclear receptor fxr improves hyperglycemia and hyperlipidemia in diabetic mice. Proc. Natl. Acad. Sci. USA 2006, 103, 1006–1011. [Google Scholar] [CrossRef]
- Clarke, S.F.; Murphy, E.F.; Nilaweera, K.; Ross, P.R.; Shanahan, F.; O’Toole, P.W.; Cotter, P.D. The gut microbiota and its relationship to diet and obesity: New insights. Gut Microbes 2012, 3, 186–202. [Google Scholar] [CrossRef]
- Louis, S.; Tappu, R.M.; Damms-Machado, A.; Huson, D.H.; Bischoff, S.C. Characterization of the gut microbial community of obese patients following a weight-loss intervention using whole metagenome shotgun sequencing. PLoS ONE 2016, 11, e0149564. [Google Scholar] [CrossRef]
- Hollister, E.B.; Gao, C.; Versalovic, J. Compositional and functional features of the gastrointestinal microbiome and their effects on human health. Gastroenterology 2014, 146, 1449–1458. [Google Scholar] [CrossRef]
- Hu, S.; Wang, Y.; Lichtenstein, L.; Tao, Y.; Musch, M.W.; Jabri, B.; Antonopoulos, D.; Claud, E.C.; Chang, E.B. Regional differences in colonic mucosa-associated microbiota determine the physiological expression of host heat shock proteins. Am. J. Physiol. Gastrointest. Liver Physiol. 2010, 299, G1266–G1275. [Google Scholar] [CrossRef]
- Harrell, L.; Wang, Y.; Antonopoulos, D.; Young, V.; Lichtenstein, L.; Huang, Y.; Hanauer, S.; Chang, E. Standard colonic lavage alters the natural state of mucosal-associated microbiota in the human colon. PLoS ONE 2012, 7, e32545. [Google Scholar] [CrossRef] [PubMed]
- Winter, S.E.; Winter, M.G.; Xavier, M.N.; Thiennimitr, P.; Poon, V.; Keestra, A.M.; Laughlin, R.C.; Gomez, G.; Wu, J.; Lawhon, S.D.; et al. Host-derived nitrate boosts growth of e. Coli in the inflamed gut. Science 2013, 339, 708–711. [Google Scholar] [CrossRef] [PubMed]
- Rhee, S.H. Lipopolysaccharide: Basic biochemistry, intracellular signaling, and physiological impacts in the gut. Intest. Res. 2014, 12, 90–95. [Google Scholar] [CrossRef] [PubMed]
- Florez, H.; Pan, Q.; Ackermann, R.T.; Marrero, D.G.; Barrett-Connor, E.; Delahanty, L.; Kriska, A.; Saudek, C.D.; Goldberg, R.B.; Rubin, R.R.; et al. Impact of lifestyle intervention and metformin on health-related quality of life: The diabetes prevention program randomized trial. J. Gen. Intern. Med. 2012, 27, 1594–1601. [Google Scholar] [CrossRef]
- Van Eldere, J.; Robben, J.; De Pauw, G.; Merckx, R.; Eyssen, H. Isolation and identification of intestinal steroid-desulfating bacteria from rats and humans. Appl. Environ. Microbiol. 1988, 54, 2112–2117. [Google Scholar] [CrossRef] [PubMed]
- Manichanh, C.; Eck, A.; Varela, E.; Roca, J.; Clemente, J.C.; Gonzalez, A.; Knights, D.; Knight, R.; Estrella, S.; Hernandez, C.; et al. Anal gas evacuation and colonic microbiota in patients with flatulence: Effect of diet. Gut 2014, 63, 401–408. [Google Scholar] [CrossRef]
- Overbeek, R.; Olson, R.; Pusch, G.D.; Olsen, G.J.; Davis, J.J.; Disz, T.; Edwards, R.A.; Gerdes, S.; Parrello, B.; Shukla, M.; et al. The seed and the rapid annotation of microbial genomes using subsystems technology (rast). Nucleic Acids Res. 2014, 42, D206–D214. [Google Scholar] [CrossRef] [PubMed]
- Desai, M.S.; Seekatz, A.M.; Koropatkin, N.M.; Kamada, N.; Hickey, C.A.; Wolter, M.; Pudlo, N.A.; Kitamoto, S.; Terrapon, N.; Muller, A.; et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 2016, 167, 1339–1353.e21. [Google Scholar] [CrossRef] [PubMed]
- Roopchand, D.E.; Carmody, R.N.; Kuhn, P.; Moskal, K.; Rojas-Silva, P.; Turnbaugh, P.J.; Raskin, I. Dietary polyphenols promote growth of the gut bacterium akkermansia muciniphila and attenuate high-fat diet-induced metabolic syndrome. Diabetes 2015, 64, 2847–2858. [Google Scholar] [CrossRef] [PubMed]
- Anhe, F.F.; Roy, D.; Pilon, G.; Dudonne, S.; Matamoros, S.; Varin, T.V.; Garofalo, C.; Moine, Q.; Desjardins, Y.; Levy, E.; et al. A polyphenol-rich cranberry extract protects from diet-induced obesity, insulin resistance and intestinal inflammation in association with increased Akkermansia spp. population in the gut microbiota of mice. Gut 2015, 64, 872–883. [Google Scholar] [CrossRef] [PubMed]
- Greer, R.L.; Dong, X.; Moraes, A.C.; Zielke, R.A.; Fernandes, G.R.; Peremyslova, E.; Vasquez-Perez, S.; Schoenborn, A.A.; Gomes, E.P.; Pereira, A.C.; et al. Akkermansia muciniphila mediates negative effects of ifngamma on glucose metabolism. Nat. Commun. 2016, 7, 13329. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Sparks, J.B.; Karyala, S.V.; Settlage, R.; Luo, X.M. Host adaptive immunity alters gut microbiota. ISME J. 2015, 9, 770–781. [Google Scholar] [CrossRef]
- Collado, M.C.; Derrien, M.; Isolauri, E.; de Vos, W.M.; Salminen, S. Intestinal integrity and Akkermansia muciniphila, a mucin-degrading member of the intestinal microbiota present in infants, adults, and the elderly. Appl. Environ. Microbiol. 2007, 73, 7767–7770. [Google Scholar] [CrossRef]
- Kong, F.; Hua, Y.; Zeng, B.; Ning, R.; Li, Y.; Zhao, J. Gut microbiota signatures of longevity. Curr. Biol. 2016, 26, R832–R833. [Google Scholar] [CrossRef]
- Gupta, V.K.; Kim, M.; Bakshi, U.; Cunningham, K.Y.; Davis, J.M., 3rd; Lazaridis, K.N.; Nelson, H.; Chia, N.; Sung, J. A predictive index for health status using species-level gut microbiome profiling. Nat. Commun. 2020, 11, 4635. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.S.; Spakowicz, D.J.; Hong, B.Y.; Petersen, L.M.; Demkowicz, P.; Chen, L.; Leopold, S.R.; Hanson, B.M.; Agresta, H.O.; Gerstein, M.; et al. Evaluation of 16s rrna gene sequencing for species and strain-level microbiome analysis. Nat. Commun. 2019, 10, 5029. [Google Scholar] [CrossRef]
- Yang, J.; Pu, J.; Lu, S.; Bai, X.; Wu, Y.; Jin, D.; Cheng, Y.; Zhang, G.; Zhu, W.; Luo, X.; et al. Species-level analysis of human gut microbiota with metataxonomics. Front. Microbiol. 2020, 11, 2029. [Google Scholar] [CrossRef]
- Schmidt, T.S.B.; Raes, J.; Bork, P. The human gut microbiome: From association to modulation. Cell 2018, 172, 1198–1215. [Google Scholar] [CrossRef]
- Rodriguez, J.M.; Murphy, K.; Stanton, C.; Ross, R.P.; Kober, O.I.; Juge, N.; Avershina, E.; Rudi, K.; Narbad, A.; Jenmalm, M.C.; et al. The composition of the gut microbiota throughout life, with an emphasis on early life. Microb. Ecol. Health Dis. 2015, 26, 26050. [Google Scholar] [CrossRef] [PubMed]
- Wen, L.; Duffy, A. Factors influencing the gut microbiota, inflammation, and type 2 diabetes. J. Nutr. 2017, 147, 1468S–1475S. [Google Scholar] [CrossRef]
- Lundberg, R.; Toft, M.F.; August, B.; Hansen, A.K.; Hansen, C.H. Antibiotic-treated versus germ-free rodents for microbiota transplantation studies. Gut Microbes 2016, 7, 68–74. [Google Scholar] [CrossRef]
- Sen, P.; Oresic, M. Metabolic modeling of human gut microbiota on a genome scale: An overview. Metabolites 2019, 9, 22. [Google Scholar] [CrossRef] [PubMed]
- Prakash, T.; Taylor, T.D. Functional assignment of metagenomic data: Challenges and applications. Brief. Bioinform. 2012, 13, 711–727. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, J.A.; Field, D.; Swift, P.; Thomas, S.; Cummings, D.; Temperton, B.; Weynberg, K.; Huse, S.; Hughes, M.; Joint, I.; et al. The taxonomic and functional diversity of microbes at a temperate coastal site: A ’multi-omic’ study of seasonal and diel temporal variation. PLoS ONE 2010, 5, e15545. [Google Scholar] [CrossRef]
- Edwards, J.S.; Ibarra, R.U.; Palsson, B.O. In silico predictions of Escherichia coli metabolic capabilities are consistent with experimental data. Nat. Biotechnol. 2001, 19, 125–130. [Google Scholar] [CrossRef]
- Feist, A.M.; Henry, C.S.; Reed, J.L.; Krummenacker, M.; Joyce, A.R.; Karp, P.D.; Broadbelt, L.J.; Hatzimanikatis, V.; Palsson, B.O. A genome-scale metabolic reconstruction for escherichia coli k-12 mg1655 that accounts for 1260 orfs and thermodynamic information. Mol. Syst. Biol. 2007, 3, 121. [Google Scholar] [CrossRef] [PubMed]
- Zomorrodi, A.R.; Maranas, C.D. Optcom: A multi-level optimization framework for the metabolic modeling and analysis of microbial communities. PLoS Comput. Biol. 2012, 8, e1002363. [Google Scholar] [CrossRef] [PubMed]
- Harcombe, W.R.; Riehl, W.J.; Dukovski, I.; Granger, B.R.; Betts, A.; Lang, A.H.; Bonilla, G.; Kar, A.; Leiby, N.; Mehta, P.; et al. Metabolic resource allocation in individual microbes determines ecosystem interactions and spatial dynamics. Cell Rep. 2014, 7, 1104–1115. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.B.; Conner, C.; Nichols, G.A. Secondary failure of metformin monotherapy in clinical practice. Diabetes Care 2010, 33, 501–506. [Google Scholar] [CrossRef]
| Ref * | Population | Study Design | Gut Microbiota | Biochemical Alterations |
|---|---|---|---|---|
| [27] | Chinese | T2DM patients (n = 170) | versus Healthy subjects (n = 174) Family: Lachnospiraceae ↑, Erysipelotrichaceae ↓ Genus: Alistipes ↑, Clostridium ↑, Eubacterium ↓, Faecalibacterium ↓, Subdoligranulum ↑, Parabacteroides ↑ Species: Akkermansia muciniphila ↑, Bacteroides intestinalis ↑, Clostridium bolteae ↑, Clostridium hatheway ↑, Clostridium ramosum ↑, Clostridium symbiosum ↑, Eggerthella lenta ↑, Escherichia coli ↑, Eubacterium rectale ↓, Faecalibacterium prausnitzii ↓, Haemophilus parainfluenzae ↓, Roseburia intestinalis ↓, Roseburia inulinivorans ↓ | NA |
| [30] | Danish | T2DM patients (n = 18) | versus Healthy subjects (n = 18) α-diversity (Chao 1): – Phylum: Bacteroidetes ↑, Firmicutes ↓, Proteobacteria ↑ Class: Bacilli ↑, Bacteroidetes ↑, Betaproteobacteria ↑, Clostridia ↓ Genus: Akkermansia ↑, Alistipes ↑, Bacteroides ↓, Bifidobacterium ↓, Bilophila ↑, Catenibacterium ↓, Dialister ↑, Dorea ↑, Erysipelotrichaceae IS ↑, Faecalibacterium ↓, Lachnospiraceae IS ↓, Lactobacillus ↑, Parabacteroides ↑, Prevotella ↑, Roseburia ↓, Ruminococcus ↓, Sporobacter ↑, Subdoligranulum ↓, Succinivibrio ↑, Sutterella ↑ Species: Dorea longicatena ↓ | NA |
| [32] | Chinese | T2DM patients (n = 13) | versus Healthy subjects (n = 44) α-diversity (Chao 1, Shannon index): ↓ Class: Clostridia ↑, Clostridiales ↑ Family: Lachnospiraceae ↑ Genus: Abiotrophia ↑, Bacteroides ↓, Collinsella ↑, Dorea ↑, Eubacterium ↑, Haemophilus ↓, Megamonas ↓, Peptostreptococcus ↑, Prevotella ↑, Roseburia ↓, Ruminococcus ↑, Sporobacter ↑, Subdoligranulum ↑ | NA |
| [34] | Pakistani | Obese-T2DM patients (n = 40) | versus Healthy subjects (n = 20) α-diversity (Shannon index): ↓ Phylum: Bacteroidetes ↓, Elusimicrobia ↓, Firmicutes ↓, Proteobacteria ↓, Verrucomicrobioa ↓ Class: Bacilli ↓, Bacteroidia ↓, Clostridia ↑, Coriobacteriia ↑, Deltaproteobacteria ↓, Elusimicrobia ↓, Gammaproteobacteria ↓, Negativicutes ↑, Genus: Allisonella ↑, Bacillus ↓, Christensenellaceae_R_7 ↑, Dialister ↑, Escherichia_Shigella ↓, Eubacterium coprostanoligenes groups ↑, Lactobacillus ↑, Prevotella_9 ↓, Ruminococcus_2 ↓, Subdoligranulum ↑ | NA |
| Metformin Treatment Effects in T2DM Patients | ||||
| [28] | Japanese | T2DM patients (n = 50) | versus normal subjects (n = 50) Genus: Atopobium cluster ↓, Lactobacillus ↑, Prevotella ↓ Species: Clostridium coccoides ↓, Lactobacillus plantarum ↑, Lactobacillus reuteri ↑ | Fecal organic acids ↓ Acetic acid ↓ Propionic acid ↓ Fecal isovaleric acid ↑ CRP ↑, IL-6 ↑ |
| Metformin treated T2DM (n = 17) | versus non treated T2DM (n = 33) Family: Enterobacteriaceae ↑ Genus: Staphylococcus ↑ Species: Clostridium coccoides ↓, Lactobacillus plantarum ↑, Lactobacillus reuteri ↑ | NA | ||
| [29] | European old woman | T2DM patients (n = 53) | versus normal glucose tolerance (n = 43) Class: Clostridiales ↓ Family: Coriobacteriaceae ↓ Genus: Alistipes ↓, Clostridium ↓, Roseburia ↓, Species: Bacteroides intestinalis ↓, Eubacterium eligens ↓, Lactobacillus gasseri ↑, Streptococcus mutans ↑ | C-peptide ↑ |
| Metformin treated T2DM (n = 20) | versus non treated T2DM (n = 33) Genus: Clostridium ↓, Escherichia ↑, Eubacterium ↓, Klebsiella ↑, Salmonella ↑, Shigella ↑ Species: Escherichia coli ↑ | NA | ||
| [31] | Danish | T2DM patients (n = 75) | versus normal subjects (n = 277) Family: bp Clostridiales ↓, Peptostreptococcaceae ↓ Genus: Akkermansia ↓, Acidaminococcus ↑, Bilophila ↑, Collinsella ↑, Coprococcus ↓, Escherichia ↑, Holdemania ↑, Lactobacillus ↑, Parabacteroides ↑, Roseburia ↓, Veillonella ↓ | NA |
| Metformin treated T2DM (n = 58) | versus non treated T2DM (n = 17) Family: Peptostreptococcaceae ↓ Genus: Akkermansia ↑, Bilophila ↓, Escherichia ↑, Holdemania ↑, Roseburia ↑, Veillonella ↓ | NA | ||
| Swedish female | T2DM patients (n = 53) | versus normal subjects (n = 92) Family: Peptostreptococcaceae ↓ Genus: Lactobacillus ↑ | NA | |
| Metformin treated T2DM (n = 20) | versus non treated T2DM (n = 33) Family: bp Clostridiales ↓, Peptostreptococcaceae ↓ Genus: Bilophila ↑, Escherichia ↑, Holdemania ↓, Lactobacillus ↑, Roseburia ↓, Veillonella ↓ | NA | ||
| Chinese | T2DM patients (n = 71) | versus normal subjects (n = 185) Family: bp Clostridiales ↓, Peptostreptococcaceae ↓, Genus: Acidaminococcus ↑, Bilophila ↑, Collinsella ↑, Coprococcus ↓, Escherichia ↑, Haemophilus ↓, Holdemania ↑, Lactobacillus ↑, Oscillibacter ↑, Roseburia ↓, Veillonella ↓ | NA | |
| Metformin treated T2DM (n = 15) | versus non treated T2DM (n = 56) Family: bp Clostridiales ↑, Peptostreptococcaceae ↓ Genus: Bilophila ↑, Collinsella ↑, Escherichia ↓, Holdemania ↑, Parabacteroides ↑, Roseburia ↑, Subdoligranulum ↑, Veillonella ↓ | NA | ||
| [33] | Chinese | T2DM patients (n = 26) | versus normal subjects (n = 50) α-diversity (Shannon index): ↓ Phylum: Firmicutes ↓ Class: Fusobacteriia ↑ Family: Enterobacteriaceae ↓, Erysipelotrichaceae ↑, Erysipelotrichaceae ↑, Porphyromonadaceae ↑ Genus: Faecalibacterium ↓, Fusobacterium ↑, Lactobacillus ↑, Ruminococcus ↓ | NA |
| Metformin treated T2DM (n = 51) | versus non treated T2DM (n = 26) α-diversity (Shannon index): – Phylum: Actinobacteria ↓ Family: Enterobacteriaceae ↓, Spirochaetaceae ↑, Turicibacteraceae ↑ Genus: Fusobacterium ↑, Turicibacter ↑ | NA | ||
| [55] | British | On metformin T2DM (visit 1 and 4, n = 12) | versus off metformin T2DM (visit 2 and 3, n = 12) Genus: SMB53 ↓, Adlercreutzia ↓, Eubacterium ↑ | Serum bile acids ↓ Fecal bile acids ↑ GLP-1 ↑ |
| [56] | Spanish | Metformin treated T2DM for 4 months (n = 22) | versus before metformin treatment in T2DM (n = 22) Phylum: Proteobacteria ↑, Firmicutes ↑ Genus: Actinetobacter ↑, Alkaliphilus ↓, Citrobacter ↑, Cronobacter ↑, Dermcoccus ↑, Desulfurispirillum ↑, Dickeya ↑, Edwardsiella ↑, Enterobacter ↑, Erwinia ↑, Escherichia ↑, Holdemania ↓, Intestinibacter ↓, Klebsiella ↓, Methylobaciilus ↑, Pantoea ↑, Pectobacterium ↑, Photorhabdus ↑, Providencia ↑, Pseudomonas ↑, Rahnella ↑, Rheinheimera ↑, Salmonella ↑, Subdoligranulum ↓, Xanthomonas ↑, Xenohabdus ↑, Yersinia ↑ Species: Akkermansia muciniphila ↑, Bifidobacterium adolescentis ↑ | Fecal propionate, butyrate, lactate and succinate ↑ Plasma bile acids ↑ |
| [57] | Colombian | T2DM patients (n = 28) | versus normal subjects (n = 84) Genus: Enterococcus casseliflavus ↓, Clostridiaceae 02d06 ↑, Prevotella ↑ | NA |
| Metformin treated T2DM (n = 14) | versus non treated T2DM (n = 14) Genus: Bacnesiellaceae ↓, Butyrivibrio ↑, Clostridiaceae 02d06 ↓, Megasphaera ↑, Oscillospira ↓, Prevotella ↑ | NA | ||
| [58] | Scandinavian | Metformin treated T2DM (n = 23) | versus non treated T2DM (n = 7) Family: Enterobacteriaceae ↑, Genus: Bacnesiellaceae ↓, Butyrivibrio ↑, Clostridiaceae 02d06 ↓, Megasphaera ↑, Oscillospira ↓, Prevotella ↑ | SCFA concentration – |
| [59] | Chinese | Metformin treated for 3 days in T2DM (n = 22) | versus before metformin treatment in T2DM (n = 22) Genus: Bacteroides ↓ Species: Bacteroides fragilis ↓, Bacteroides finegoldii ↓, Bacteroides thetaiotaomicron ↓, Bacteroides uniformis ↓, Bacteroides ovatus ↓, Bacteroides intestinalis ↓, Bacteroides stercoris ↓, Bacteroides eggerthii ↓, Bacteroides fluxus ↓, Bacteroides caccae ↓, Bacteroides dorei ↓ | GUDCA, Tauroursodeoxycholic acid, Conjugated Secondary bile acids ↑ Total bile acids – |
| Metformin Treatment Effects in Healthy Subjects | ||||
| [60] | Caucasian | Metformin treated for 7 days in healthy subjects (n = 18) | versus before metformin treatment in healthy subjects (n = 18) α-diversity (Shannon index): ↓ Class: Bacilli ↑, Enterobacteriales ↑, Episilonproteobacteria ↑, Gammaproteobacteria ↑, Negativicutes ↓ Order: Clostridiaceae_1 ↓, Lactobacillales ↑, Peptostreptococcaceae ↓, Selenomonadales ↓ Family: Asaccharospora ↓, Enterobacteriaceae ↑, Romboutsia ↓ Genus: Blautia ↑, Ruminiclostridium_6 ↓, Streptococcus ↑ | NA |
| [61] | Danish | Metformin treated for 6 weeks in healthy subjects (n = 22) | versus before metformin treatment in healthy subjects (n = 18) Genus: Bilophila ↑, Caproiciproducens ↑, Clostridium_sensu_stricto_1 ↓, Escherichia-Shigella ↑, Intestinibacter ↓, Prevotella ↑, Terrisporobacter ↓ Species: Alistipes finegoldii ↑, Bilophila wadsworthia ↑, Intestinibacter bartlettii ↓ | NA |
| Clinical Trials.gov Identifier | Study Title | Country | Study Population | Interventions |
|---|---|---|---|---|
| NCT04194515 | Gut Microbiota and Bile Acids in Type 2 Diabetes Mellitus | Taiwan | Outpatients and treatment-naïve male patients with type 2 diabetes | Drug: YH1 Drug: metformin |
| NCT04287387 | Response of Gut Microbiota in Type 2 Diabetes to Hypoglycemic Agents | China | Type 2 diabetes patients (18–65 years) | Drug: Glucophage 500 mg Tablet Drug: Acarbose Tablets Drug: Sitagliptin tablet Drug: Dapagliflozin Tablet Drug: Pioglitazone Tablets Drug: Glimepiride Tablets |
| NCT04639492 | Postbiotic MBS and Metformin Combination in Patients With T2DM | Taiwan | Type 2 diabetes patients (20–70 years) | Dietary Supplement: MBS oral solution Oral BIDAC, twice a day before breakfast and dinner times |
| NCT02960659 | Title: Therapeutic Targets in African-American Youth With Type 2 Diabetes | United States | African-American (12–25 years) | Drug: Metformin and Liraglutide Drug: Metformin |
| NCT03558867 | Personalized Medicine in Pre-diabetes and Early Type 2 Diabetes | Australia | Pre-diabetes or newly-diagnosed with type 2 diabetes (in the last 6 months) | Drug: Metformin + Healthy diet Drug: Metformin + Personalized diet |
| NCT03732690 | The Interaction Between Protein Intake, Gut Microbiota and Type 2 Diabetes in Subjects With Different Ethnic Backgrounds | France | T2DM patients: Caucasian (n = 80), Caribbean (n = 40) stable dose of metformin and do not use insulin or proton-pump inhibitors. | Other: Diet HP Other: Diet LP |
| NCT04089280 | Probiotics in Metformin Intolerant Patients With Type 2 Diabetes | Poland | T2DM patients (18–75 years) with metformin treatment in the last 3 months (<1500 mg/d) | Dietary Supplement: Sanprobi Barrier-multispecies probiotic Other: Placebo Comparator |
| NCT03718715 | The Interaction Between Metformin and Microbiota—The MEMO Study. (MEMO) | Sweden | Newly diagnosed patients with type 2 diabetes without previous treatment with metformin (40–80 years). | Drug: Metformin |
| NCT03489317 | Gut Microbiomes in Patients With Metabolic Syndrome | Hongkong | Residents in Hongkong (no metabolic syndrome, metabolic syndrome-partial, metabolic syndrome-full) | Drug: Metformin Behavioral: lifestyle modification Drug: Simvastatin 10 mg Drug: Amlodipine 5 mg |
| NCT02609815 | Initial Combination of Gemigliptin and Metformin on Microbiota Change | Republic of Korea | Type 2 patients with drug naive for 6 weeks | Drug: gemigliptin/metformin Drug: glimepiride/metformin |
| NCT04341571 | Effect of Probiotics Versus Metformin on Glycemic Control, Insulin Sensitivity and Insulin Secretion in Prediabetes. | Mexico | Pre-diabetes | Dietary Supplement: Probiotics Drug: Metformin |
| NCT04209075 | Prebiotics and Metformin Improve Gut and Hormones in Type 2 Diabetes in Youth (MIGHTY-fiber) | United States | Type 2 patients (10–25 years) | Dietary Supplement: Biomebliss Drug: Metformin Dietary Supplement: Placebo |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, C.B.; Chae, S.U.; Jo, S.J.; Jerng, U.M.; Bae, S.K. The Relationship between the Gut Microbiome and Metformin as a Key for Treating Type 2 Diabetes Mellitus. Int. J. Mol. Sci. 2021, 22, 3566. https://doi.org/10.3390/ijms22073566
Lee CB, Chae SU, Jo SJ, Jerng UM, Bae SK. The Relationship between the Gut Microbiome and Metformin as a Key for Treating Type 2 Diabetes Mellitus. International Journal of Molecular Sciences. 2021; 22(7):3566. https://doi.org/10.3390/ijms22073566
Chicago/Turabian StyleLee, Chae Bin, Soon Uk Chae, Seong Jun Jo, Ui Min Jerng, and Soo Kyung Bae. 2021. "The Relationship between the Gut Microbiome and Metformin as a Key for Treating Type 2 Diabetes Mellitus" International Journal of Molecular Sciences 22, no. 7: 3566. https://doi.org/10.3390/ijms22073566
APA StyleLee, C. B., Chae, S. U., Jo, S. J., Jerng, U. M., & Bae, S. K. (2021). The Relationship between the Gut Microbiome and Metformin as a Key for Treating Type 2 Diabetes Mellitus. International Journal of Molecular Sciences, 22(7), 3566. https://doi.org/10.3390/ijms22073566






