Post-Translational Modifications of Retroviral HIV-1 Gag Precursors: An Overview of Their Biological Role
Abstract
1. Introduction
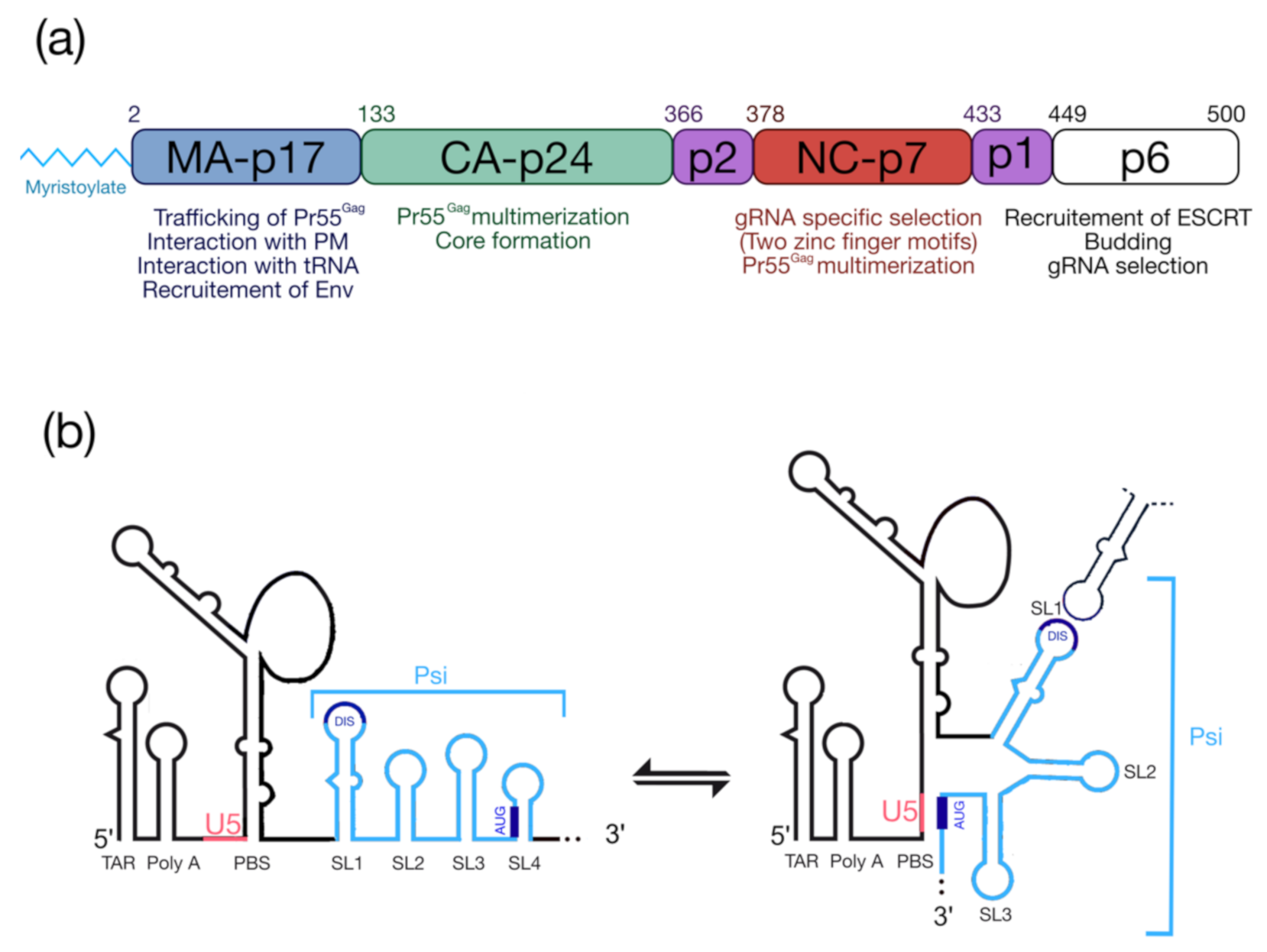
2. HIV-1 Pr55Gag
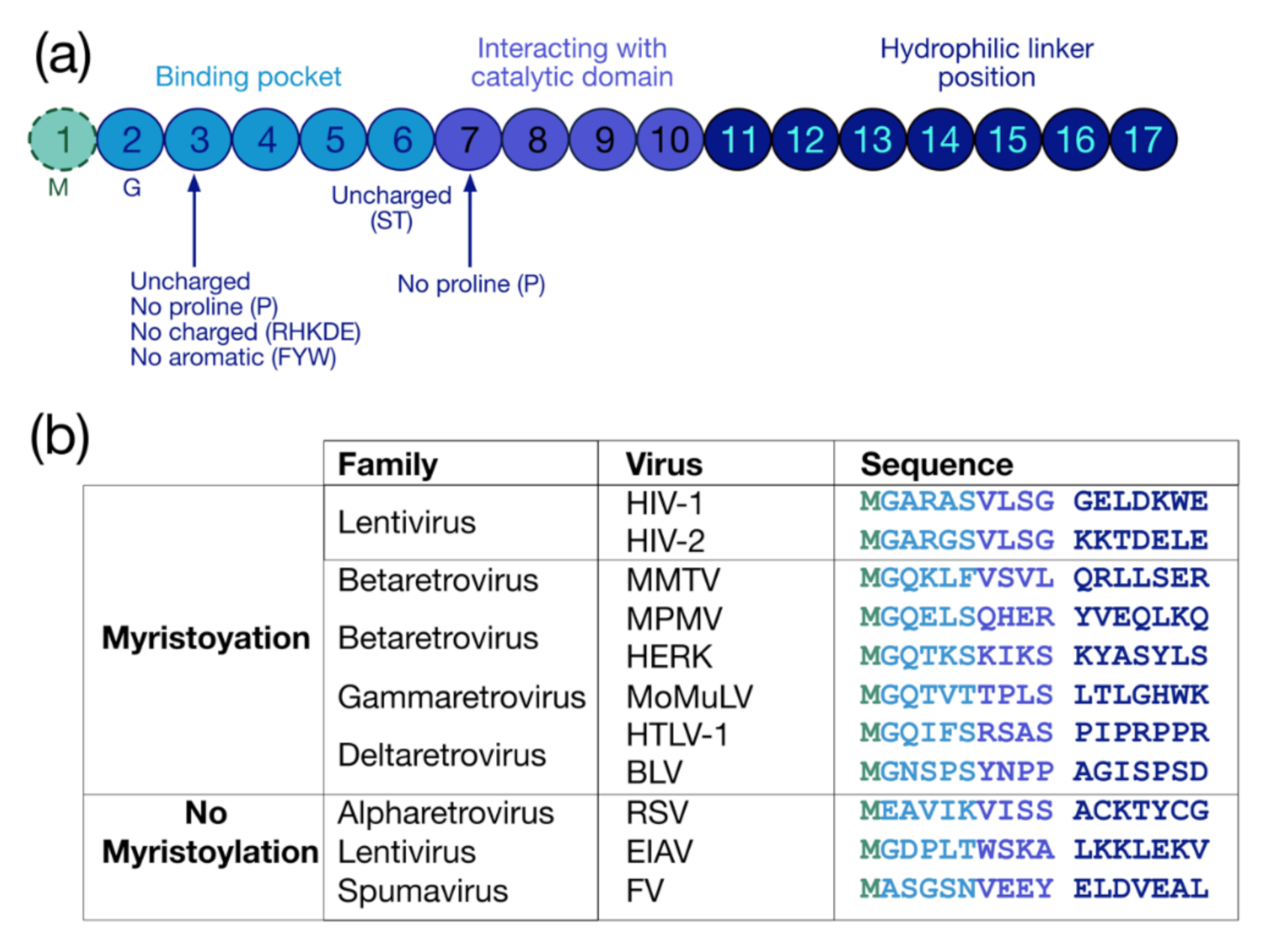
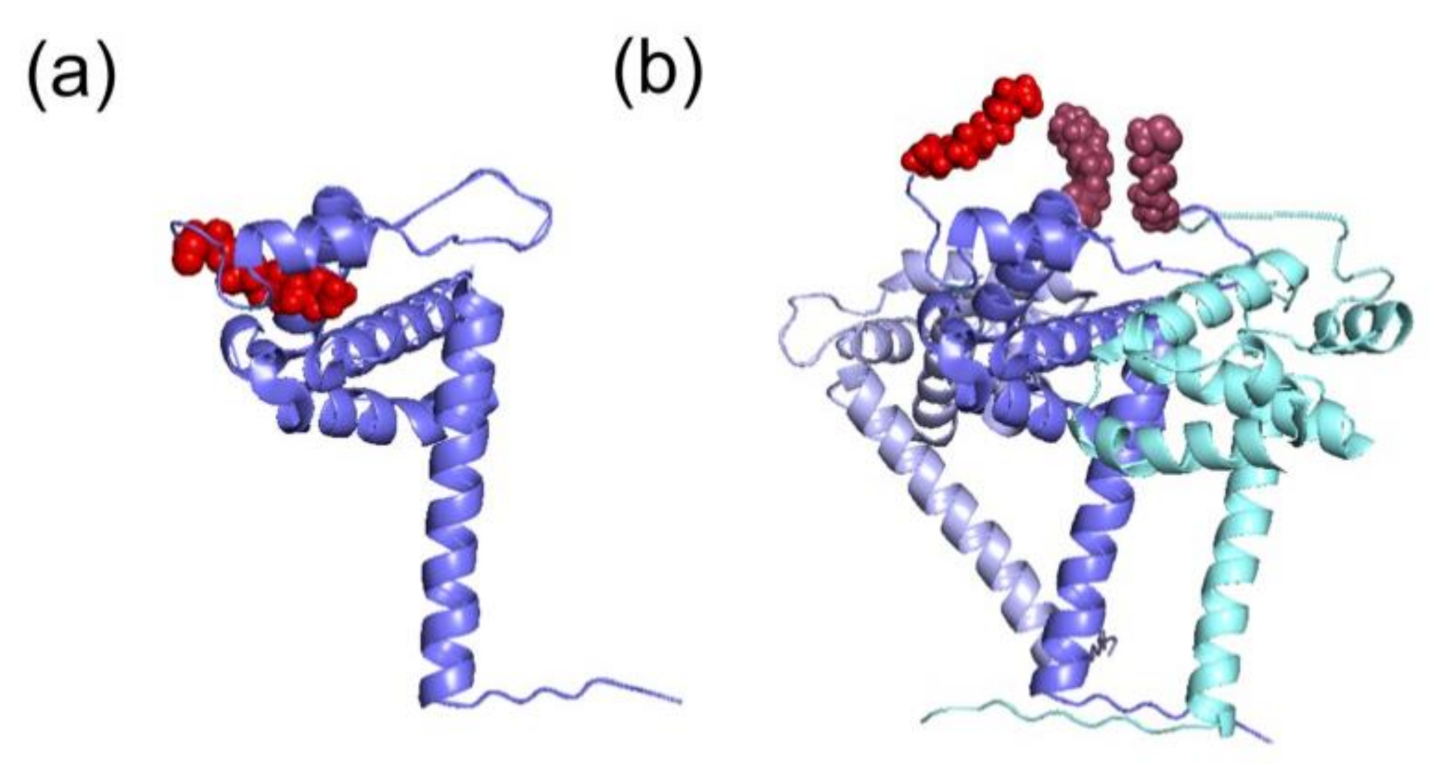
3. HIV-1 Pr55Gag Myristoylation
4. Gag Myristoylation in Other Retroviruses
5. HIV-1 Pr55Gag Phosphorylation
6. Gag Phosphorylation in Other Retroviruses
7. HIV-1 Pr55Gag Ubiquitination
8. Gag Ubiquitination in Other Retroviruses
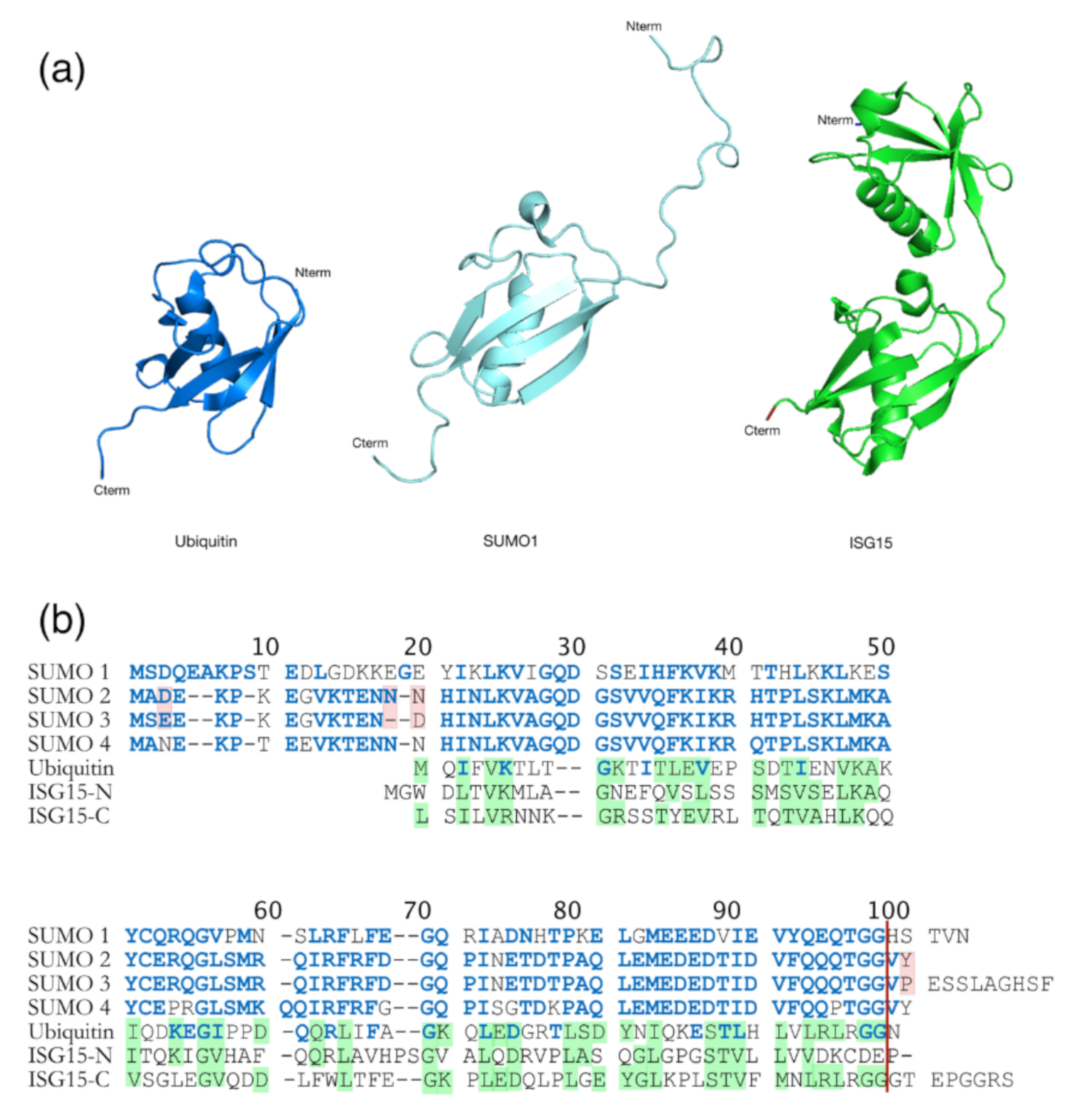
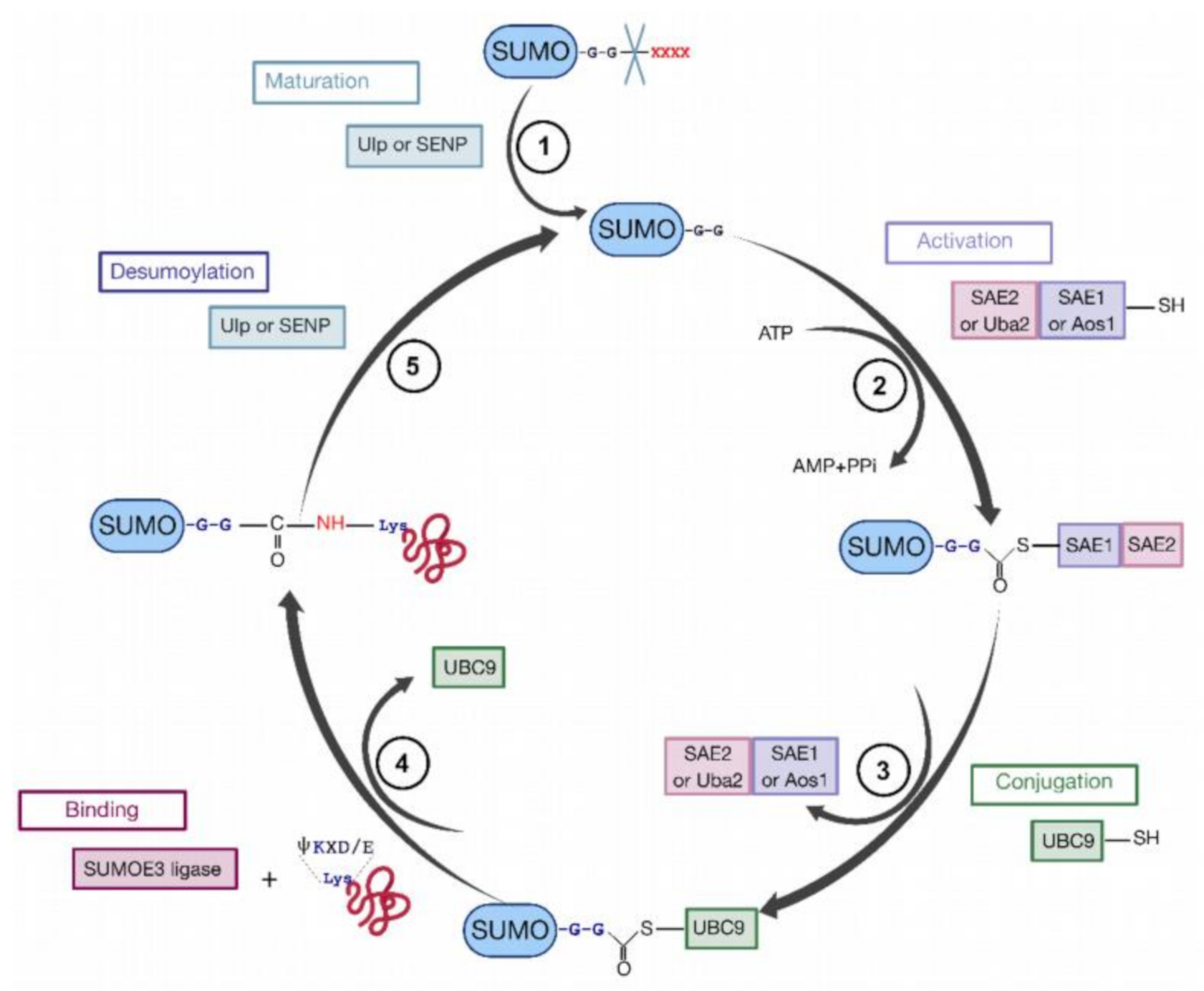
9. HIV-1 Pr55Gag Sumoylation
10. Gag Sumoylation in Other Retroviruses
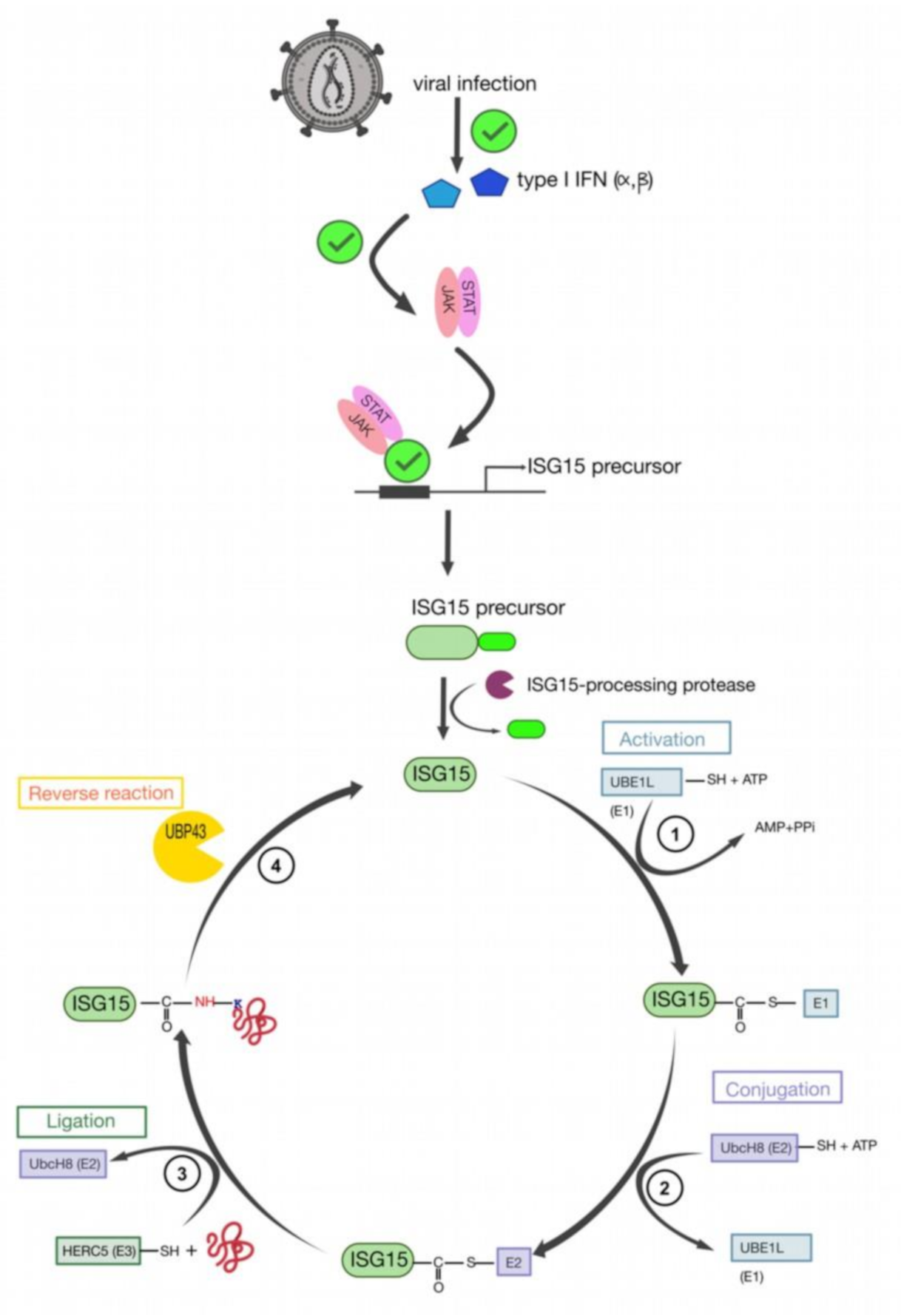
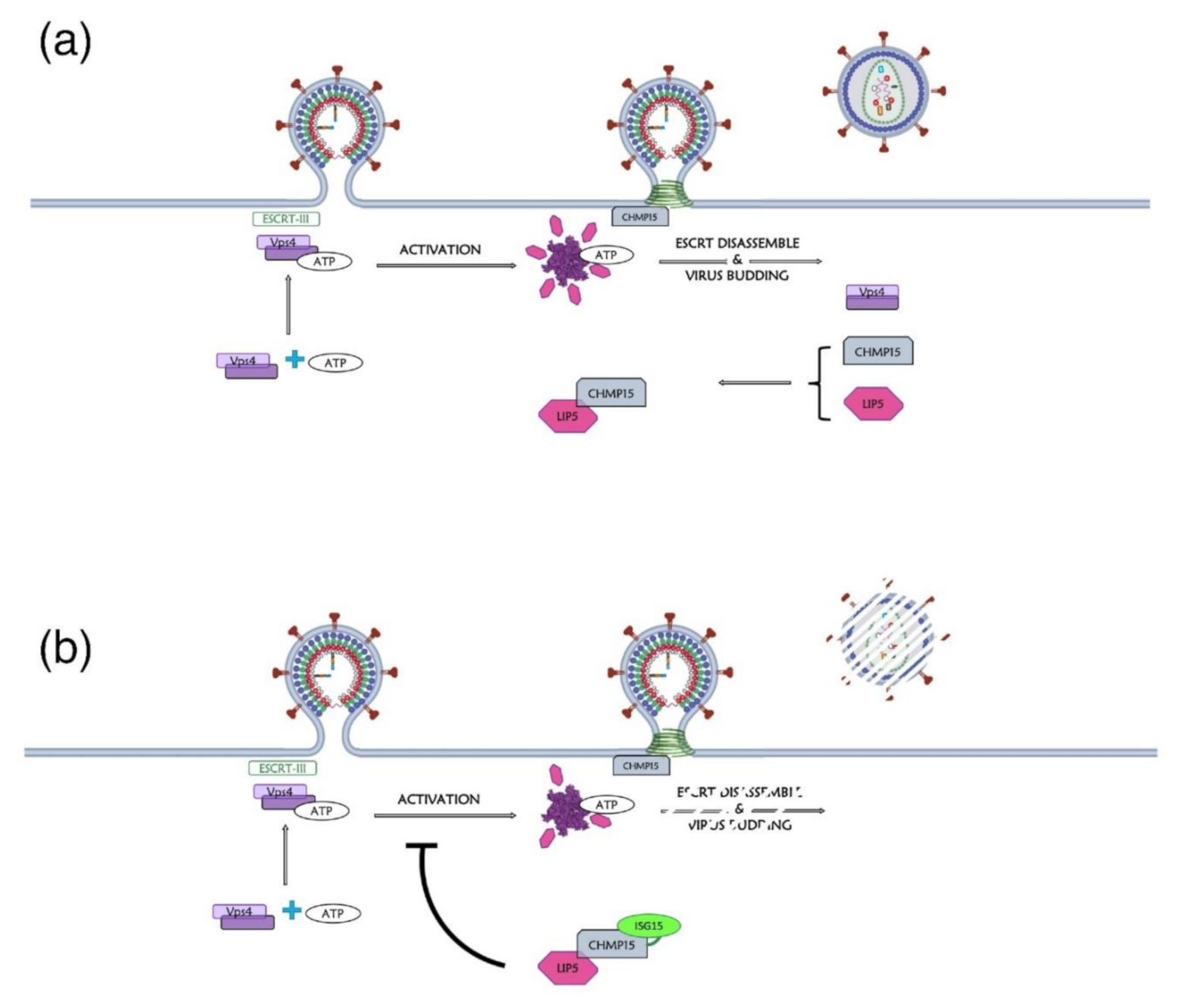
11. Retroviral Gag Protein ISGylation
12. Post-Translational Methylation of Retroviral Gag Proteins
13. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Audagnotto, M.; Peraro, M.D. Protein post-translational modifications: In silico prediction tools and molecular modeling. Comput. Struct. Biotechnol. J. 2017, 15, 307–319. [Google Scholar] [CrossRef]
- Spoel, S.H. Orchestrating the proteome with post-translational modifications. J. Exp. Bot. 2018, 69, 4499–4503. [Google Scholar] [CrossRef]
- Biard-Piechaczyk, M.; Borel, S.; Espert, L.; De Bettignies, G.; Coux, O. HIV-1, ubiquitin and ubiquitin-like proteins: The dialectic interactions of a virus with a sophisticated network of post-translational modifications. Biol. Cell 2012, 104, 165–187. [Google Scholar] [CrossRef]
- Chen, L.; Keppler, O.T.; Schölz, C. Post-translational Modification-Based Regulation of HIV Replication. Front. Microbiol. 2018, 9, 2131. [Google Scholar] [CrossRef] [PubMed]
- Lu, K.; Heng, X.; Garyu, L.; Monti, S.; Garcia, E.L.; Kharytonchyk, S.; Dorjsuren, B.; Kulandaivel, G.; Jones, S.; Hiremath, A.; et al. NMR Detection of Structures in the HIV-15′-Leader RNA That Regulate Genome Packaging. Science 2011, 334, 242–245. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, A.K.; Gorelick, R.J.; Vasa, S.M.; Guex, N.; Rein, A.; Mathews, D.H.; Giddings, M.C.; Weeks, K.M. High-Throughput SHAPE Analysis Reveals Structures in HIV-1 Genomic RNA Strongly Conserved across Distinct Biological States. PLoS Biol. 2008, 6, 96. [Google Scholar] [CrossRef] [PubMed]
- El-Wahab, E.W.A.; Smyth, R.P.; Mailler, E.; Bernacchi, S.; Vivet-Boudou, V.; Hijnen, M.; Jossinet, F.; Mak, J.; Paillart, J.-C.; Marquet, R. Specific recognition of the HIV-1 genomic RNA by the Gag precursor. Nat. Commun. 2014, 5, 4304. [Google Scholar] [CrossRef]
- Smyth, R.P.; Despons, L.; Huili, G.; Bernacchi, S.; Hijnen, M.; Mak, J.; Jossinet, F.; Weixi, L.; Paillart, J.-C.; Von Kleist, M.; et al. Mutational interference mapping experiment (MIME) for studying RNA structure and function. Nat. Methods 2015, 12, 866–872. [Google Scholar] [CrossRef]
- Bernacchi, S.; El-Wahab, E.W.A.; Dubois, N.; Hijnen, M.; Smyth, R.P.; Mak, J.; Marquet, R.; Paillart, J.-C. HIV-1 Pr55Gag binds genomic and spliced RNAs with different affinity and stoichiometry. RNA Biol. 2017, 14, 90–103. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, M.; Clerté, C.; Chamontin, C.; Basyuk, E.; Lainé, S.; Hottin, J.; Bertrand, E.; Margeat, E.; Mougel, M. Imaging HIV-1 RNA dimerization in cells by multicolor super-resolution and fluctuation microscopies. Nucleic Acids Res. 2016, 44, 7922–7934. [Google Scholar] [CrossRef]
- Jouvenet, N.; Simon, S.M.; Bieniasz, P.D. Imaging the interaction of HIV-1 genomes and Gag during assembly of individual viral particles. Proc. Natl. Acad. Sci. USA 2009, 106, 19114–19119. [Google Scholar] [CrossRef]
- Kutluay, S.B.; Bieniasz, P.D. Analysis of the Initiating Events in HIV-1 Particle Assembly and Genome Packaging. PLoS Pathog. 2010, 6, e1001200. [Google Scholar] [CrossRef]
- Mailler, E.; Bernacchi, S.; Marquet, R.; Paillart, J.-C.; Vivet-Boudou, V.; Smyth, R.P. The Life-Cycle of the HIV-1 Gag–RNA Complex. Viruses 2016, 8, 248. [Google Scholar] [CrossRef] [PubMed]
- Bieniasz, P.; Telesnitsky, A. Multiple, Switchable Protein:RNA Interactions Regulate Human Immunodeficiency Virus Type 1 Assembly. Annu. Rev. Virol. 2018, 5, 165–183. [Google Scholar] [CrossRef] [PubMed]
- Comas-Garcia, M.; Davis, S.R.; Rein, A. On the Selective Packaging of Genomic RNA by HIV-1. Viruses 2016, 8, 246. [Google Scholar] [CrossRef] [PubMed]
- Bell, N.M.; Lever, A.M. HIV Gag polyprotein: Processing and early viral particle assembly. Trends Microbiol. 2013, 21, 136–144. [Google Scholar] [CrossRef] [PubMed]
- Olety, B.; Ono, A. Roles played by acidic lipids in HIV-1 Gag membrane binding. Virus Res. 2014, 193, 108–115. [Google Scholar] [CrossRef]
- Kutluay, S.B.; Zang, T.; Blanco-Melo, D.; Powell, C.; Jannain, D.; Errando, M.; Bieniasz, P.D. Global Changes in the RNA Binding Specificity of HIV-1 Gag Regulate Virion Genesis. Cell 2014, 159, 1096–1109. [Google Scholar] [CrossRef] [PubMed]
- Gaines, C.R.; Tkacik, E.; Rivera-Oven, A.; Somani, P.; Achimovich, A.; Alabi, T.; Zhu, A.; Getachew, N.; Yang, A.L.; McDonough, M.; et al. HIV-1 Matrix Protein Interactions with tRNA: Implications for Membrane Targeting. J. Mol. Biol. 2018, 430, 2113–2127. [Google Scholar] [CrossRef]
- Thornhill, D.; Olety, B.; Ono, A. Relationships between MA-RNA Binding in Cells and Suppression of HIV-1 Gag Mislocalization to Intracellular Membranes. J. Virol. 2019, 93, 23. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Perilla, J.R.; Yufenyuy, E.L.; Meng, X.; Chen, B.; Ning, J.; Ahn, J.; Gronenborn, A.M.; Schulten, K.; Aiken, C.; et al. Mature HIV-1 capsid structure by cryo-electron microscopy and all-atom molecular dynamics. Nat. Cell Biol. 2013, 497, 643–646. [Google Scholar] [CrossRef] [PubMed]
- Schur, F.K.M.; Hagen, W.J.H.; Rumlová, M.; Ruml, T.; Müller, B.; Kräusslich, H.-G.; Briggs, J.A.G. Structure of the immature HIV-1 capsid in intact virus particles at 8.8 A resolution. Nat. Cell Biol. 2015, 517, 505–508. [Google Scholar] [CrossRef]
- Pornillos, O.; Ganser-Pornillos, B.K.; Kelly, B.N.; Hua, Y.; Whitby, F.G.; Stout, C.D.; Sundquist, W.I.; Hill, C.P.; Yeager, M. X-Ray Structures of the Hexameric Building Block of the HIV Capsid. Cell 2009, 137, 1282–1292. [Google Scholar] [CrossRef]
- Dannull, J.; Surovoy, A.; Jung, G.; Moelling, K. Specific binding of HIV-1 nucleocapsid protein to PSI RNA in vitro requires N-terminal zinc finger and flanking basic amino acid residues. EMBO J. 1994, 13, 1525–1533. [Google Scholar] [CrossRef]
- Webb, J.A.; Jones, C.P.; Parent, L.J.; Rouzina, I.; Musier-Forsyth, K. Distinct binding interactions of HIV-1 Gag to Psi and non-Psi RNAs: Implications for viral genomic RNA packaging. RNA 2013, 19, 1078–1088. [Google Scholar] [CrossRef] [PubMed]
- Cimarelli, A.; Sandin, S.; Höglund, S.; Luban, J. Basic Residues in Human Immunodeficiency Virus Type 1 Nucleocapsid Promote Virion Assembly via Interaction with RNA. J. Virol. 2000, 74, 3046–3057. [Google Scholar] [CrossRef] [PubMed]
- Ott, D.E.; Coren, L.V.; Shatzer, T. The Nucleocapsid Region of Human Immunodeficiency Virus Type 1 Gag Assists in the Coordination of Assembly and Gag Processing: Role for RNA-Gag Binding in the Early Stages of Assembly. J. Virol. 2009, 83, 7718–7727. [Google Scholar] [CrossRef]
- El Meshri, S.E.; Dujardin, D.; Godet, J.; Richert, L.; Boudier, C.; Darlix, J.L.; Didier, P.; Mély, Y.; De Rocquigny, H. Role of the Nucleocapsid Domain in HIV-1 Gag Oligomerization and Trafficking to the Plasma Membrane: A Fluorescence Lifetime Imaging Microscopy Investigation. J. Mol. Biol. 2015, 427, 1480–1494. [Google Scholar] [CrossRef]
- Dubois, N.; Khoo, K.K.; Ghossein, S.; Seissler, T.; Wolff, P.; McKinstry, W.J.; Mak, J.; Paillart, J.-C.; Marquet, R.; Bernacchi, S. The C-terminal p6 domain of the HIV-1 Pr55Gag precursor is required for specific binding to the genomic RNA. RNA Biol. 2018, 15, 923–936. [Google Scholar] [CrossRef]
- Farazi, T.A.; Waksman, G.; Gordon, J.I. The Biology and Enzymology of ProteinN-Myristoylation. J. Biol. Chem. 2001, 276, 39501–39504. [Google Scholar] [CrossRef]
- Martin, D.D.; Beauchamp, E.; Berthiaume, L.G. Post-translational myristoylation: Fat matters in cellular life and death. Biochimie 2011, 93, 18–31. [Google Scholar] [CrossRef] [PubMed]
- Dyda, F.; Klein, D.C.; Hickman, A.B. GCN5-Related N-Acetyltransferases: A Structural Overview. Annu. Rev. Biophys. Biomol. Struct. 2000, 29, 81–103. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.R.; Bhatnagar, R.S.; Knoll, L.J.; Gordon, I.J. Genetic and Biochemical Studies of Protein N-Myristoylation. Annu. Rev. Biochem. 1994, 63, 869–914. [Google Scholar] [CrossRef]
- Cao, W.; Sumikoshi, K.; Nakamura, S.; Terada, T.; Shimizu, K. Prediction of N-myristoylation modification of proteins by SVM. Bioinformation 2011, 6, 204–206. [Google Scholar] [CrossRef] [PubMed]
- Maurer-Stroh, S.; Eisenhaber, B.; Eisenhaber, F. N-terminal N -myristoylation of proteins: Refinement of the sequence motif and its taxon-specific differences 1 1Edited by J. Thornton. J. Mol. Biol. 2002, 317, 523–540. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.; Loeliger, E.; Luncsford, P.; Kinde, I.; Beckett, D.; Summers, M.F. Entropic switch regulates myristate exposure in the HIV-1 matrix protein. Proc. Natl. Acad. Sci. USA 2003, 101, 517–522. [Google Scholar] [CrossRef] [PubMed]
- Hogue, I.B.; Llewellyn, G.N.; Ono, A. Dynamic Association between HIV-1 Gag and Membrane Domains. Mol. Biol. Int. 2012, 2012, 979765. [Google Scholar] [CrossRef]
- Inlora, J.; Collins, D.R.; Trubin, M.E.; Chung, J.Y.J.; Ono, A. Membrane Binding and Subcellular Localization of Retroviral Gag Proteins Are Differentially Regulated by MA Interactions with Phosphatidylinositol-(4,5)-Bisphosphate and RNA. mBio 2014, 5, e02202-14. [Google Scholar] [CrossRef]
- Resh, M.D. A myristoyl switch regulates membrane binding of HIV-1 Gag. Proc. Natl. Acad. Sci. USA 2004, 101, 417–418. [Google Scholar] [CrossRef] [PubMed]
- Ono, A. Relationships between plasma membrane microdomains and HIV-1 assembly. Biol. Cell 2010, 102, 335–350. [Google Scholar] [CrossRef] [PubMed]
- Saad, J.S.; Miller, J.; Tai, J.; Kim, A.; Ghanam, R.H.; Summers, M.F. Structural basis for targeting HIV-1 Gag proteins to the plasma membrane for virus assembly. Proc. Natl. Acad. Sci. USA 2006, 103, 11364–11369. [Google Scholar] [CrossRef]
- Saad, J.S.; Loeliger, E.; Luncsford, P.; Liriano, M.; Tai, J.; Kim, A.; Miller, J.; Joshi, A.; Freed, E.O.; Summers, M.F. Point Mutations in the HIV-1 Matrix Protein Turn Off the Myristyl Switch. J. Mol. Biol. 2007, 366, 574–585. [Google Scholar] [CrossRef] [PubMed]
- Dalton, A.K.; Ako-Adjei, D.; Murray, P.S.; Murray, D.; Vogt, V.M. Electrostatic Interactions Drive Membrane Association of the Human Immunodeficiency Virus Type 1 Gag MA Domain. J. Virol. 2007, 81, 6434–6445. [Google Scholar] [CrossRef]
- Paillart, J.-C.; Göttlinger, H.G. Opposing Effects of Human Immunodeficiency Virus Type 1 Matrix Mutations Support a Myristyl Switch Model of Gag Membrane Targeting. J. Virol. 1999, 73, 2604–2612. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Resh, M.D. Differential membrane binding of the human immunodeficiency virus type 1 matrix protein. J. Virol. 1996, 70, 8540–8548. [Google Scholar] [CrossRef] [PubMed]
- Murray, P.S.; Li, Z.; Wang, J.; Tang, C.L.; Honig, B.; Murray, D. Retroviral Matrix Domains Share Electrostatic Homology: Models for Membrane Binding Function throughout the Viral Life Cycle. Structure 2005, 13, 1521–1531. [Google Scholar] [CrossRef] [PubMed]
- Hamard-Peron, E.; Muriaux, D. Retroviral matrix and lipids, the intimate interaction. Retrovirology 2011, 8, 15. [Google Scholar] [CrossRef]
- Parent, L.J.; Gudleski, N. Beyond Plasma Membrane Targeting: Role of the MA domain of Gag in Retroviral Genome Encapsidation. J. Mol. Biol. 2011, 410, 553–564. [Google Scholar] [CrossRef] [PubMed]
- Murray, D.; Ben-Tal, N.; Honig, B.; McLaughlin, S. Electrostatic interaction of myristoylated proteins with membranes: Simple physics, complicated biology. Structure 1997, 5, 985–989. [Google Scholar] [CrossRef]
- Kozak, C.A. Origins of the Endogenous and Infectious Laboratory Mouse Gammaretroviruses. Viruses 2014, 7, 1–26. [Google Scholar] [CrossRef]
- Nitta, T.; Kuznetsov, Y.; McPherson, A.; Fan, H. Murine leukemia virus glycosylated Gag (gPr80gag) facilitates interferon-sensitive virus release through lipid rafts. Proc. Natl. Acad. Sci. USA 2009, 107, 1190–1195. [Google Scholar] [CrossRef]
- Houzet, L.; Gay, B.; Morichaud, Z.; Briant, L.; Mougel, M. Intracellular assembly and budding of the Murine Leukemia Virus in infected cells. Retrovirology 2006, 3, 12. [Google Scholar] [CrossRef]
- Hron, T.; Elleder, D.; Gifford, R.J. Deltaretroviruses have circulated since at least the Paleogene and infected a broad range of mammalian species. Retrovirology 2019, 16, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Egessain, A.; Ecassar, O. Epidemiological Aspects and World Distribution of HTLV-1 Infection. Front. Microbiol. 2012, 3, 388. [Google Scholar] [CrossRef]
- Barez, P.-Y.; De Brogniez, A.; Carpentier, A.; Gazon, H.; Gillet, N.; Gutiérrez, G.; Hamaidia, M.; Jacques, J.-R.; Perike, S.; Sriramareddy, S.N.; et al. Recent Advances in BLV Research. Viruses 2015, 7, 6080–6088. [Google Scholar] [CrossRef] [PubMed]
- Blot, V.; Perugi, F.; Gay, B.; Prévost, M.-C.; Briant, L.; Tangy, F.; Abriel, H.; Staub, O.; Dokhélar, M.-C.; Pique, C. Nedd4.1-mediated ubiquitination and subsequent recruitment of Tsg101 ensure HTLV-1 Gag trafficking towards the multivesicular body pathway prior to virus budding. J. Cell Sci. 2004, 117, 2357–2367. [Google Scholar] [CrossRef]
- Inlora, J.; Chukkapalli, V.; Derse, D.; Ono, A. Gag Localization and Virus-Like Particle Release Mediated by the Matrix Domain of Human T-Lymphotropic Virus Type 1 Gag Are Less Dependent on Phosphatidylinositol-(4,5)-Bisphosphate than Those Mediated by the Matrix Domain of HIV-1 Gag. J. Virol. 2011, 85, 3802–3810. [Google Scholar] [CrossRef] [PubMed]
- Le Blanc, I.; Rosenberg, A.R.; Dokhélar, M.-C. Multiple Functions for the Basic Amino Acids of the Human T-Cell Leukemia Virus Type 1 Matrix Protein in Viral Transmission. J. Virol. 1999, 73, 1860–1867. [Google Scholar] [CrossRef]
- Fogarty, K.H.; Zhang, W.; Grigsby, I.F.; Johnson, J.L.; Chen, Y.; Mueller, J.D.; Mansky, L.M. New Insights into HTLV-1 Particle Structure, Assembly, and Gag-Gag Interactions in Living Cells. Viruses 2011, 3, 770–793. [Google Scholar] [CrossRef] [PubMed]
- Yeung, T.; Gilbert, G.E.; Shi, J.; Silvius, J.; Kapus, A.; Grinstein, S. Membrane Phosphatidylserine Regulates Surface Charge and Protein Localization. Science 2008, 319, 210–213. [Google Scholar] [CrossRef]
- Chukkapalli, V.; Oh, S.J.; Ono, A. Opposing mechanisms involving RNA and lipids regulate HIV-1 Gag membrane binding through the highly basic region of the matrix domain. Proc. Natl. Acad. Sci. USA 2010, 107, 1600–1605. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Norris, K.M.; Mansky, L.M. Involvement of the Matrix and Nucleocapsid Domains of the Bovine Leukemia Virus Gag Polyprotein Precursor in Viral RNA Packaging. J. Virol. 2003, 77, 9431–9438. [Google Scholar] [CrossRef]
- Prchal, J.; Srb, P.; Hunter, E.; Ruml, T.; Hrabal, R. The Structure of Myristoylated Mason-Pfizer Monkey Virus Matrix Protein and the Role of Phosphatidylinositol-(4,5)-Bisphosphate in Its Membrane Binding. J. Mol. Biol. 2012, 423, 427–438. [Google Scholar] [CrossRef]
- Ross, S.R. Mouse Mammary Tumor Virus Molecular Biology and Oncogenesis. Viruses 2010, 2, 2000–2012. [Google Scholar] [CrossRef] [PubMed]
- Hayward, A.J.; Tachedjian, M.; Cui, J.; Field, H.; Holmes, E.C.; Wang, L.-F.; Tachedjian, G. Identification of diverse full-length endogenous betaretroviruses in megabats and microbats. Retrovirology 2013, 10, 35. [Google Scholar] [CrossRef] [PubMed]
- Monde, K.; Contreras-Galindo, R.; Kaplan, M.H.; Markovitz, D.M.; Ono, A. Human Endogenous Retrovirus K Gag Coassembles with HIV-1 Gag and Reduces the Release Efficiency and Infectivity of HIV-1. J. Virol. 2012, 86, 11194–11208. [Google Scholar] [CrossRef]
- Doležal, M.; Zábranský, A.; Dostál, J.; Vaněk, O.; Brynda, J.; Lepšík, M.; Hadravová, R.; Pichová, I. Myristoylation drives dimerization of matrix protein from mouse mammary tumor virus. Retrovirology 2016, 13, 1–15. [Google Scholar] [CrossRef]
- Dalton, A.K.; Murray, P.S.; Murray, D.; Vogt, V.M. Biochemical Characterization of Rous Sarcoma Virus MA Protein Interaction with Membranes. J. Virol. 2005, 79, 6227–6238. [Google Scholar] [CrossRef][Green Version]
- Nadaraia-Hoke, S.; Bann, D.V.; Lochmann, T.L.; Gudleski-O’Regan, N.; Parent, L.J. Alterations in the MA and NC Domains Modulate Phosphoinositide-Dependent Plasma Membrane Localization of the Rous Sarcoma Virus Gag Protein. J. Virol. 2013, 87, 3609–3615. [Google Scholar] [CrossRef] [PubMed]
- Callahan, E.M.; Wills, J.W. Repositioning Basic Residues in the M Domain of the Rous Sarcoma Virus Gag Protein. J. Virol. 2000, 74, 11222–11229. [Google Scholar] [CrossRef]
- Dick, R.A.; Vogt, V.M. Membrane interaction of retroviral Gag proteins. Front. Microbiol. 2014, 5, 187. [Google Scholar] [CrossRef]
- Hatanaka, H.; Iourin, O.; Rao, Z.; Fry, E.; Kingsman, A.; Stuart, D.I. Structure of Equine Infectious Anemia Virus Matrix Protein. J. Virol. 2002, 76, 1876–1883. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Ma, J.; Zhang, X.; Su, C.; Yao, Q.-C.; Wang, X. Equine Infectious Anemia Virus Gag Assembly and Export Are Directed by Matrix Protein throughtrans-Golgi Networks and Cellular Vesicles. J. Virol. 2015, 90, 1824–1838. [Google Scholar] [CrossRef]
- Fernandes, F.; Chen, K.; Ehrlich, L.S.; Jin, J.; Chen, M.H.; Medina, G.N.; Symons, M.; Montelaro, R.; Donaldson, J.; Tjandra, N.; et al. Phosphoinositides Direct Equine Infectious Anemia Virus Gag Trafficking and Release. Traffic 2010, 12, 438–451. [Google Scholar] [CrossRef]
- Goldstone, D.C.; Flower, T.G.; Ball, N.J.; Sanz-Ramos, M.; Yap, M.W.; Ogrodowicz, R.W.; Stanke, N.; Reh, J.; Lindemann, D.; Stoye, J.P.; et al. A Unique Spumavirus Gag N-terminal Domain with Functional Properties of Orthoretroviral Matrix and Capsid. PLoS Pathog. 2013, 9, e1003376. [Google Scholar] [CrossRef]
- Heneine, W.; Schweizer, M.; Sandstrom, P.; Folks, T. Human Infection with Foamy Viruses. Curr. Top. Microbiol. Immunol. 2003, 277, 181–196. [Google Scholar] [CrossRef] [PubMed]
- Shaw, K.L.; Lindemann, D.; Mulligan, M.J.; Goepfert, P.A. Foamy Virus Envelope Glycoprotein Is Sufficient for Particle Budding and Release. J. Virol. 2003, 77, 2338–2348. [Google Scholar] [CrossRef] [PubMed]
- Nishi, H.; Shaytan, A.; Panchenko, A.R. Physicochemical mechanisms of protein regulation by phosphorylation. Front. Genet. 2014, 5, 270. [Google Scholar] [CrossRef] [PubMed]
- Mandell, D.J.; Chorny, I.; Groban, E.S.; Wong, S.E.; Levine, E.; Rapp, A.C.S.; Jacobson, M.P. Strengths of Hydrogen Bonds Involving Phosphorylated Amino Acid Side Chains. J. Am. Chem. Soc. 2007, 129, 820–827. [Google Scholar] [CrossRef] [PubMed]
- Burnett, G.; Kennedy, E.P. The Enzymatic Phosphorylation of Proteins. J. Biol. Chem. 1954, 211, 969–980. [Google Scholar] [CrossRef]
- Francis, A.C.; Di Primio, C.; Allouch, A.; Cereseto, A. Role of phosphorylation in the nuclear biology of HIV-1. Curr. Med. Chem. 2011, 18, 2904–2912. [Google Scholar] [CrossRef] [PubMed]
- Burnette, B.; Yu, G.; Felsted, R. Phosphorylation of HIV-1 gag proteins by protein kinase C. J. Biol. Chem. 1993, 268, 8698–8703. [Google Scholar] [CrossRef]
- Yu, G.; Shen, F.S.; Sturch, S.; Aquino, A.; Glazer, R.I.; Felsted, R.L. Regulation of HIV-1 gag Protein Subcellular Targeting by Protein Kinase C. J. Biol. Chem. 1995, 270, 4792–4796. [Google Scholar] [CrossRef] [PubMed]
- Kräusslich, H.G. Morphogenesis and Maturation of Retroviruses—Google Livres, 1st ed.; Springer Science & Business Media: Berlin/Heidelberg, Germany, 1996. [Google Scholar]
- Kudoh, A.; Takahama, S.; Sawasaki, T.; Ode, H.; Yokoyama, M.; Okayama, A.; Ishikawa, A.; Miyakawa, K.; Matsunaga, S.; Kimura, H.; et al. The phosphorylation of HIV-1 Gag by atypical protein kinase C facilitates viral infectivity by promoting Vpr incorporation into virions. Retrovirology 2014, 11, 9. [Google Scholar] [CrossRef] [PubMed]
- Radestock, B.; Morales, I.; Rahman, S.A.; Radau, S.; Glass, B.; Zahedi, R.P.; Müller, B.; Kräusslich, H.-G. Comprehensive Mutational Analysis Reveals p6Gag Phosphorylation to Be Dispensable for HIV-1 Morphogenesis and Replication. J. Virol. 2012, 87, 724–734. [Google Scholar] [CrossRef]
- Votteler, J.; Neumann, L.; Hahn, S.; Hahn, F.; Rauch, P.; Schmidt, K.; Studtrucker, N.; Solbak, S.M.; Fossen, T.; Henklein, P.; et al. Highly conserved serine residue 40 in HIV-1 p6 regulates capsid processing and virus core assembly. Retrovirology 2011, 8, 11. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.M.; Chen, M.-H.; Khan, M.; Ehrlich, L.; Kemal, K.S.; Weiser, B.; Shi, B.; Chen, C.; Powell, M.; Anastos, K.; et al. The S40 residue in HIV-1 Gag p6 impacts local and distal budding determinants, revealing additional late domain activities. Retrovirology 2013, 10, 143. [Google Scholar] [CrossRef]
- Radestock, B.; Burk, R.; Müller, B.; Kräusslich, H.-G. Re-visiting the functional Relevance of the highly conserved Serine 40 Residue within HIV-1 p6Gag. Retrovirology 2014, 11, 1–5. [Google Scholar] [CrossRef]
- Müller, B.; Patschinsky, T.; Kräusslich, H.-G. The Late-Domain-Containing Protein p6 Is the Predominant Phosphoprotein of Human Immunodeficiency Virus Type 1 Particles. J. Virol. 2002, 76, 1015–1024. [Google Scholar] [CrossRef]
- Hemonnot, B.; Cartier, C.; Gay, B.; Rebuffat, S.; Bardy, M.; Devaux, C.; Boyer, V.; Briant, L. The Host Cell MAP Kinase ERK-2 Regulates Viral Assembly and Release by Phosphorylating the p6 Protein of HIV-1. J. Biol. Chem. 2004, 279, 32426–32434. [Google Scholar] [CrossRef]
- Wang, Z.; Canagarajah, B.J.; Boehm, J.C.; Kassisà, S.; Cobb, M.H.; Young, P.R.; Abdel-Meguid, S.; Adams, J.L.; Goldsmith, E.J. Structural basis of inhibitor selectivity in MAP kinases. Structure 1998, 6, 1117–1128. [Google Scholar] [CrossRef]
- Cartier, C.; Deckert, M.; Grangeasse, C.; Trauger, R.; Jensen, F.; Bernard, A.; Cozzone, A.; Desgranges, C.; Boyer, V. Association of ERK2 mitogen-activated protein kinase with human immunodeficiency virus particles. J. Virol. 1997, 71, 4832–4837. [Google Scholar] [CrossRef]
- Cartier, C.; Hemonnot, B.; Gay, B.; Bardy, M.; Sanchiz, C.; Devaux, C.; Briant, L. Active cAMP-dependent Protein Kinase Incorporated within Highly Purified HIV-1 Particles Is Required for Viral Infectivity and Interacts with Viral Capsid Protein. J. Biol. Chem. 2003, 278, 35211–35219. [Google Scholar] [CrossRef]
- Jacqué, J.; Mann, A.; Enslen, H.; Sharova, N.; Brichacek, B.; Davis, R.J.; Stevenson, M. Modulation of HIV-1 infectivity by MAPK, a virion-associated kinase. EMBO J. 1998, 17, 2607–2618. [Google Scholar] [CrossRef]
- Dochi, T.; Nakano, T.; Inoue, M.; Takamune, N.; Shoji, S.; Sano, K.; Misumi, S. Phosphorylation of human immunodeficiency virus type 1 capsid protein at serine 16, required for peptidyl-prolyl isomerase-dependent uncoating, is mediated by virion-incorporated extracellular signal-regulated kinase 2. J. Gen. Virol. 2014, 95, 1156–1166. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Gabuzda, D. Regulation of Human Immunodeficiency Virus Type 1 Infectivity by the ERK Mitogen-Activated Protein Kinase Signaling Pathway. J. Virol. 1999, 73, 3460–3466. [Google Scholar] [CrossRef]
- Greenway, A.; Azad, A.; Mills, J.; McPhee, D. Human immunodeficiency virus type 1 Nef binds directly to Lck and mitogen-activated protein kinase, inhibiting kinase activity. J. Virol. 1996, 70, 6701–6708. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Goncalves, J.; Gabuzda, D. Phosphorylation of Vif and Its Role in HIV-1 Replication. J. Biol. Chem. 1996, 271, 10121–10129. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Gabuzda, D. Mitogen-activated Protein Kinase Phosphorylates and Regulates the HIV-1 Vif Protein. J. Biol. Chem. 1998, 273, 29879–29887. [Google Scholar] [CrossRef]
- Kaushik, R.; Ratner, L. Role of Human Immunodeficiency Virus Type 1 Matrix Phosphorylation in an Early Postentry Step of Virus Replication. J. Virol. 2004, 78, 2319–2326. [Google Scholar] [CrossRef] [PubMed]
- Strasner, A.B.; Natarajan, M.; Doman, T.; Key, D.; August, A.; Henderson, A.J. The Src Kinase Lck Facilitates Assembly of HIV-1 at the Plasma Membrane. J. Immunol. 2008, 181, 3706–3713. [Google Scholar] [CrossRef]
- Gallay, P.; Swingler, S.; Song, J.; Bushman, F.; Trono, D. HIV nuclear import is governed by the phosphotyrosine-mediated binding of matrix to the core domain of integrase. Cell 1995, 83, 569–576. [Google Scholar] [CrossRef]
- Hahn, F.; Setz, C.; Friedrich, M.; Rauch, P.; Solbak, S.M.; Frøystein, N.Å.; Henklein, P.; Votteler, J.; Fossen, T.; Schubert, U. Mutation of the Highly Conserved Ser-40 of the HIV-1 p6 Gag Protein to Phe Causes the Formation of a Hydrophobic Patch, Enhances Membrane Association, and Polyubiquitination of Gag. Viruses 2014, 6, 3738–3765. [Google Scholar] [CrossRef] [PubMed]
- Leis, J.; Phillips, N.; Fu, X.; Tuazon, P.T.; Traugh, J.A. Phosphorylation of avian retrovirus matrix protein by Ca2+/phospholipid-dependent protein kinase. JBIC J. Biol. Inorg. Chem. 1989, 179, 415–422. [Google Scholar] [CrossRef] [PubMed]
- Nelle, T.D.; Verderame, M.F.; Leis, J.; Wills, J.W. The Major Site of Phosphorylation within the Rous Sarcoma Virus MA Protein Is Not Required for Replication. J. Virol. 1998, 72, 1103–1107. [Google Scholar] [CrossRef] [PubMed]
- Pepinsky, R.B.; Papayannopoulos, A.I.; Campbell, S.; Vogt, V.M. Analysis of Rous sarcoma virus Gag protein by mass spectrometry indicates trimming by host exopeptidase. J. Virol. 1996, 70, 3313–3318. [Google Scholar] [CrossRef]
- Fu, X.; Tuazon, P.T.; Traugh, J.A.; Leis, J. Site-Directed Mutagenesis of the Avian Retrovirus Nucleocapsid Protein, Pp12, at Serine 40, the Primary Site of Phosphorylation in Vivo. J. Biol. Chem. 1988, 268, 2134–2139. [Google Scholar] [CrossRef]
- Wang, H.; Machesky, N.J.; Mansky, L.M. Both the PPPY and PTAP Motifs Are Involved in Human T-Cell Leukemia Virus Type 1 Particle Release. J. Virol. 2004, 78, 1503–1512. [Google Scholar] [CrossRef] [PubMed]
- Hemonnot, B.; Molle, D.; Bardy, M.; Gay, B.; Laune, D.; Devaux, C.; Briant, L. Phosphorylation of the HTLV-1 matrix L-domain-containing protein by virus-associated ERK-2 kinase. Virology 2006, 349, 430–439. [Google Scholar] [CrossRef]
- Bouamr, F.; Melillo, J.A.; Wang, M.Q.; Nagashima, K.; Santos, M.D.L.; Rein, A.; Goff, S.P. PPPYEPTAP Motif Is the Late Domain of Human T-Cell Leukemia Virus Type 1 Gag and Mediates Its Functional Interaction with Cellular Proteins Nedd4 and Tsg101. J. Virol. 2003, 77, 11882–11895. [Google Scholar] [CrossRef]
- Yasuda, J.; Hunter, E. A Proline-Rich Motif (PPPY) in the Gag Polyprotein of Mason-Pfizer Monkey Virus Plays a Maturation-Independent Role in Virion Release. J. Virol. 1998, 72, 4095–4103. [Google Scholar] [CrossRef]
- Henderson, E.L.; Sowder, R.; Smythers, G.; Benveniste, R.E.; Oroszlan, S. Purification and N-terminal amino acid sequence comparisons of structural proteins from retrovirus-D/Washington and Mason-Pfizer monkey virus. J. Virol. 1985, 55, 778–787. [Google Scholar] [CrossRef] [PubMed]
- Uckert, W.; Wunderlich, V.; Fiebach, H.; Hertling, I.; Stein, U.; Kraft, R.; Desrosiers, R. Biochemical and immunological characterization of structural proteins from retrovirus-D/New England and comparison to Mason-Pfizer monkey virus and permanent human fibroblast virus. Arch. Virol. 1987, 94, 267–282. [Google Scholar] [CrossRef]
- Enssle, J.; Fischer, N.; Moebes, A.; Mauer, B.; Smola, U.; Rethwilm, A. Carboxy-terminal cleavage of the human foamy virus Gag precursor molecule is an essential step in the viral life cycle. J. Virol. 1997, 71, 7312–7317. [Google Scholar] [CrossRef]
- Yuan, B.; Li, X.; Goff, S.P. Mutations altering the moloney murine leukemia virus p12 Gag protein affect virion production and early events of the virus life cycle. EMBO J. 1999, 18, 4700–4710. [Google Scholar] [CrossRef] [PubMed]
- Yueh, A.; Goff, S.P. Phosphorylated Serine Residues and an Arginine-Rich Domain of the Moloney Murine Leukemia Virus p12 Protein Are Required for Early Events of Viral Infection. J. Virol. 2003, 77, 1820–1829. [Google Scholar] [CrossRef][Green Version]
- Brzezinski, J.D.; Felkner, R.; Modi, A.; Liu, M.; Roth, M.J. Phosphorylation Requirement of Murine Leukemia Virus p12. J. Virol. 2016, 90, 11208–11219. [Google Scholar] [CrossRef] [PubMed]
- Callis, J. The Ubiquitination Machinery of the Ubiquitin System. Arab. Book 2014, 12, e0174. [Google Scholar] [CrossRef]
- Vijay-Kumar, S.; Bugg, C.; Wilkinson, K.; Vierstra, R.; Hatfield, P.; Cook, W. Comparison of the three-dimensional structures of human, yeast, and oat ubiquitin. J. Biol. Chem. 1987, 262, 6396–6399. [Google Scholar] [CrossRef]
- Komander, D. The emerging complexity of protein ubiquitination. Biochem. Soc. Trans. 2009, 37, 937–953. [Google Scholar] [CrossRef]
- Shields, S.B.; Piper, R.C. How Ubiquitin Functions with ESCRTs. Traffic 2011, 12, 1306–1317. [Google Scholar] [CrossRef]
- Ohtake, F.; Saeki, Y.; Ishido, S.; Kanno, J.; Tanaka, K. The K48-K63 Branched Ubiquitin Chain Regulates NF-κB Signaling. Mol. Cell 2016, 64, 251–266. [Google Scholar] [CrossRef] [PubMed]
- Suresh, B.; Lee, J.; Kim, K.-S.; Ramakrishna, S. The Importance of Ubiquitination and Deubiquitination in Cellular Reprogramming. Stem Cells Int. 2016, 2016, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Gottwein, E.; Kräusslich, H.-G. Analysis of Human Immunodeficiency Virus Type 1 Gag Ubiquitination. J. Virol. 2005, 79, 9134–9144. [Google Scholar] [CrossRef]
- Gottwein, E.; Jäger, S.; Habermann, A.; Kräusslich, H.-G. Cumulative Mutations of Ubiquitin Acceptor Sites in Human Immunodeficiency Virus Type 1 Gag Cause a Late Budding Defect. J. Virol. 2006, 80, 6267–6275. [Google Scholar] [CrossRef]
- Ott, D.E.; Coren, L.V.; Ii, R.C.S.; Adams, J.; Schubert, U. Retroviruses Have Differing Requirements for Proteasome Function in the Budding Process. J. Virol. 2003, 77, 3384–3393. [Google Scholar] [CrossRef] [PubMed]
- Sette, P.; Jadwin, J.A.; Dussupt, V.; Bello, N.F.; Bouamr, F. The ESCRT-Associated Protein Alix Recruits the Ubiquitin Ligase Nedd4-1 To Facilitate HIV-1 Release through the LYPXnL L Domain Motif. J. Virol. 2010, 84, 8181–8192. [Google Scholar] [CrossRef]
- Sette, P.; Nagashima, K.; Piper, R.C.; Bouamr, F. Ubiquitin conjugation to Gag is essential for ESCRT-mediated HIV-1 budding. Retrovirology 2013, 10, 79. [Google Scholar] [CrossRef] [PubMed]
- Ott, D.E.; Coren, L.V.; Copeland, T.D.; Kane, B.P.; Johnson, D.G.; Sowder, R.C.; Yoshinaka, Y.; Oroszlan, S.; Arthur, L.O.; Henderson, L.E. Ubiquitin Is Covalently Attached to the p6GagProteins of Human Immunodeficiency Virus Type 1 and Simian Immunodeficiency Virus and to the p12Gag Protein of Moloney Murine Leukemia Virus. J. Virol. 1998, 72, 2962–2968. [Google Scholar] [CrossRef]
- Schubert, U.; Ott, D.E.; Chertova, E.N.; Welker, R.; Tessmer, U.; Princiotta, M.F.; Bennink, J.R.; Kräusslich, H.-G.; Yewdell, J.W. Proteasome inhibition interferes with Gag polyprotein processing, release, and maturation of HIV-1 and HIV-2. Proc. Natl. Acad. Sci. USA 2000, 97, 13057–13062. [Google Scholar] [CrossRef] [PubMed]
- Ott, D.E.; Coren, L.V.; Chertova, E.N.; Gagliardi, T.D.; Schubert, U. Ubiquitination of HIV-1 and MuLV Gag. Virology 2000, 278, 111–121. [Google Scholar] [CrossRef]
- Patnaik, A.; Chau, V.; Wills, J.W. Ubiquitin is part of the retrovirus budding machinery. Proc. Natl. Acad. Sci. USA 2000, 97, 13069–13074. [Google Scholar] [CrossRef]
- Jäger, S.; Gottwein, E.; Kräusslich, H.-G. Ubiquitination of Human Immunodeficiency Virus Type 1 Gag Is Highly Dependent on Gag Membrane Association. J. Virol. 2007, 81, 9193–9201. [Google Scholar] [CrossRef] [PubMed]
- Von Schwedler, U.K.; Stuchell, M.; Müller, B.; Ward, D.M.; Chung, H.Y.; Morita, E.; Wang, H.E.; Davis, T.; He, G.P.; Cimbora, D.M.; et al. The protein network of HIV budding. Cell 2003, 114, 701–713. [Google Scholar] [CrossRef]
- Pornillos, O.; Alam, S.L.; Rich, R.L.; Myszka, D.G.; Davis, D.R.; Sundquist, W.I. Structure and functional interactions of the Tsg101 UEV domain. EMBO J. 2002, 21, 2397–2406. [Google Scholar] [CrossRef]
- Watanabe, S.M.; Strickland, M.; Tjandra, N.; Carter, C.A. RNA Binding Suppresses Tsg101 Recognition of Ub-Modified Gag and Facilitates Recruitment to the Plasma Membrane. Viruses 2020, 12, 447. [Google Scholar] [CrossRef]
- Garrus, J.E.; von Schwedler, U.K.; Pornillos, O.W.; Morham, S.G.; Zavitz, K.H.; Wang, H.E.; Wettstein, D.A.; Stray, K.M.; Côté, M.; Rich, R.L.; et al. Tsg101 and the Vacuolar Protein Sorting Pathway Are Essential for HIV-1 Budding. Cell 2001, 107, 55–65. [Google Scholar] [CrossRef]
- Staub, O.; Dho, S.; Henry, P.; Correa, J.; Ishikawa, T.; McGlade, J.; Rotin, D. WW domains of Nedd4 bind to the proline-rich PY motifs in the epithelial Na+ channel deleted in Liddle’s syndrome. EMBO J. 1996, 15, 2371–2380. [Google Scholar] [CrossRef] [PubMed]
- Babst, M.; Katzmann, D.J.; Estepa-Sabal, E.J.; Meerloo, T.; Emr, S.D. Escrt-III. Dev. Cell 2002, 3, 271–282. [Google Scholar] [CrossRef]
- Dunn, R.; Klos, D.A.; Adler, A.S.; Hicke, L. The C2 domain of the Rsp5 ubiquitin ligase binds membrane phosphoinositides and directs ubiquitination of endosomal cargo. J. Cell Biol. 2004, 165, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Spidel, J.L.; Craven, R.C.; Wilson, C.B.; Patnaik, A.; Wang, H.; Mansky, L.M.; Wills, J.W. Lysines Close to the Rous Sarcoma Virus Late Domain Critical for Budding. J. Virol. 2004, 78, 10606–10616. [Google Scholar] [CrossRef] [PubMed]
- Ott, D.E.; Coren, L.V.; Sowder, R.C.; Adams, J.; Nagashima, K.; Schubert, U. Equine Infectious Anemia Virus and the Ubiquitin-Proteasome System. J. Virol. 2002, 76, 3038–3044. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Li, F.; Montelaro, R.C. Functional Roles of Equine Infectious Anemia Virus Gag p9 in Viral Budding and Infection. J. Virol. 2001, 75, 9762–9770. [Google Scholar] [CrossRef][Green Version]
- Heidecker, G.; Lloyd, P.A.; Fox, K.; Nagashima, K.; Derse, D. Late Assembly Motifs of Human T-Cell Leukemia Virus Type 1 and Their Relative Roles in Particle Release. J. Virol. 2004, 78, 6636–6648. [Google Scholar] [CrossRef]
- Heidecker, G.; Lloyd, P.A.; Soheilian, F.; Nagashima, K.; Derse, D. The Role of WWP1-Gag Interaction and Gag Ubiquitination in Assembly and Release of Human T-Cell Leukemia Virus Type 1. J. Virol. 2007, 81, 9769–9777. [Google Scholar] [CrossRef] [PubMed]
- Medina, G.; Pincetic, A.; Ehrlich, L.S.; Zhang, Y.; Tang, Y.; Leis, J.; Carter, C.A. Tsg101 can replace Nedd4 function in ASV Gag release but not membrane targeting. Virology 2008, 377, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Bartusch, C.; Prange, R. ESCRT Requirements for Murine Leukemia Virus Release. Viruses 2016, 8, 103. [Google Scholar] [CrossRef] [PubMed]
- Vana, M.L.; Tang, Y.; Chen, A.; Medina, G.; Carter, C.; Leis, J. Role of Nedd4 and Ubiquitination of Rous Sarcoma Virus Gag in Budding of Virus-Like Particles from Cells. J. Virol. 2004, 78, 13943–13953. [Google Scholar] [CrossRef]
- Stanke, N.; Stange, A.; Lüftenegger, D.; Zentgraf, H.; Lindemann, D. Ubiquitination of the Prototype Foamy Virus Envelope Glycoprotein Leader Peptide Regulates Subviral Particle Release. J. Virol. 2005, 79, 15074–15083. [Google Scholar] [CrossRef]
- Yang, Y.; He, Y.; Wang, X.; Liang, Z.; He, G.; Zhang, P.; Zhu, H.; Xu, N.; Liang, S. Protein SUMOylation modification and its associations with disease. Open Biol. 2017, 7, 170167. [Google Scholar] [CrossRef]
- Han, Z.-J.; Feng, Y.-H.; Gu, B.-H.; Li, Y.-M.; Chen, H. The post-translational modification, SUMOylation, and cancer (Review). Int. J. Oncol. 2018, 52, 1081–1094. [Google Scholar] [CrossRef]
- Matunis, M.J.; Coutavas, E.; Blobel, G. A novel ubiquitin-like modification modulates the partitioning of the Ran-GTPase-activating protein RanGAP1 between the cytosol and the nuclear pore complex. J. Cell Biol. 1996, 135, 1457–1470. [Google Scholar] [CrossRef] [PubMed]
- Owerbach, D.; McKay, E.M.; Yeh, E.T.; Gabbay, K.H.; Bohren, K.M. A proline-90 residue unique to SUMO-4 prevents maturation and sumoylation. Biochem. Biophys. Res. Commun. 2005, 337, 517–520. [Google Scholar] [CrossRef] [PubMed]
- Mahajan, R.; Gerace, L.; Melchior, F. Molecular Characterization of the SUMO-1 Modification of RanGAP1 and Its Role in Nuclear Envelope Association. J. Cell Biol. 1998, 140, 259–270. [Google Scholar] [CrossRef]
- Saitoh, H.; Hinchey, J. Functional Heterogeneity of Small Ubiquitin-related Protein Modifiers SUMO-1 versus SUMO-2/3. J. Biol. Chem. 2000, 275, 6252–6258. [Google Scholar] [CrossRef]
- Desterro, J.M.; Rodriguez, M.S.; Hay, R.T. SUMO-1 Modification of IκBα Inhibits NF-κB Activation. Mol. Cell 1998, 2, 233–239. [Google Scholar] [CrossRef]
- Nayak, A.; Müller, S. SUMO-specific proteases/isopeptidases: SENPs and beyond. Genome Biol. 2014, 15, 422. [Google Scholar] [CrossRef] [PubMed]
- Tatham, M.H.; Kim, S.; Jaffray, E.; Song, J.; Chen, Y.; Hay, R.T. Unique binding interactions among Ubc9, SUMO and RanBP2 reveal a mechanism for SUMO paralog selection. Nat. Struct. Mol. Biol. 2004, 12, 67–74. [Google Scholar] [CrossRef]
- Eisenhardt, N.; Ilic, D.; Nagamalleswari, E.; Pichler, A. Biochemical characterization of SUMO-conjugating enzymes by in vitro sumoylation assays. Methods Enzym. 2019, 618, 167–185. [Google Scholar] [CrossRef]
- Jeon, Y.J.; Yoo, H.M.; Chung, C.H. ISG15 and immune diseases. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2010, 1802, 485–496. [Google Scholar] [CrossRef]
- Gurer, C.; Berthoux, L.; Luban, J. Covalent Modification of Human Immunodeficiency Virus Type 1 p6 by SUMO-1. J. Virol. 2005, 79, 910–917. [Google Scholar] [CrossRef]
- Jaber, T.; Bohl, C.R.; Lewis, G.L.; Wood, C.; West, J.T.; Weldon, R.A. Human Ubc9 Contributes to Production of Fully Infectious Human Immunodeficiency Virus Type 1 Virions. J. Virol. 2009, 83, 10448–10459. [Google Scholar] [CrossRef]
- Martinez, N.W.; Xue, X.; Berro, R.G.; Kreitzer, G.; Resh, M.D. Kinesin KIF4 Regulates Intracellular Trafficking and Stability of the Human Immunodeficiency Virus Type 1 Gag Polyprotein. J. Virol. 2008, 82, 9937–9950. [Google Scholar] [CrossRef] [PubMed]
- Lamoliatte, F.; McManus, F.P.; Maarifi, G.; Chelbi-Alix, M.K.; Thibault, P. Uncovering the SUMOylation and ubiquitylation crosstalk in human cells using sequential peptide immunopurification. Nat. Commun. 2017, 8, 14109. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.; Tang, Y.; Okada, Y.; Torrey, T.A.; Chattopadhyay, S.K.; Pfleiderer, M.; Falkner, F.G.; Dorner, F.; Choi, W.; Hirokawa, N.; et al. Binding of Murine Leukemia Virus Gag Polyproteins to KIF4, a Microtubule-Based Motor Protein. J. Virol. 1998, 72, 6898–6901. [Google Scholar] [CrossRef]
- Weldon, A.R.; Sarkar, P.; Brown, S.M.; Weldon, S.K. Mason–Pfizer monkey virus Gag proteins interact with the human sumo conjugating enzyme, hUbc9. Virology 2003, 314, 62–73. [Google Scholar] [CrossRef]
- Yueh, A.; Leung, J.; Bhattacharyya, S.; Perrone, L.A.; Santos, K.D.L.; Pu, S.-Y.; Goff, S.P. Interaction of Moloney Murine Leukemia Virus Capsid with Ubc9 and PIASy Mediates SUMO-1 Addition Required Early in Infection. J. Virol. 2006, 80, 342–352. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wen, S.; Zhao, R.; Qi, J.; Liu, Z.; Li, W.; An, J.; Wood, C.; Wang, Y. Covalent conjugation of the equine infectious anemia virus Gag with SUMO. Biochem. Biophys. Res. Commun. 2017, 486, 712–719. [Google Scholar] [CrossRef]
- Perng, Y.-C.; Lenschow, D.J. ISG15 in antiviral immunity and beyond. Nat. Rev. Genet. 2018, 16, 423–439. [Google Scholar] [CrossRef] [PubMed]
- Potter, J.L.; Narasimhan, J.; Mende-Mueller, L.; Haas, A.L. Precursor Processing of Pro-ISG15/UCRP, an Interferon-β-induced Ubiquitin-like Protein. J. Biol. Chem. 1999, 274, 25061–25068. [Google Scholar] [CrossRef] [PubMed]
- Villarroya-Beltri, C.; Guerra, S.; Sánchez-Madrid, F. ISGylation—A key to lock the cell gates for preventing the spread of threats. J. Cell Sci. 2017, 130, 2961–2969. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Collins, M.N.; Hsiang, T.-Y.; Krug, R.M. Interferon-induced ISG15 pathway: An ongoing virus–host battle. Trends Microbiol. 2013, 21, 181–186. [Google Scholar] [CrossRef]
- Hou, S.X.; Zheng, Z.; Chen, X.; Perrimon, N. The JAK/STAT Pathway in Model Organisms. Dev. Cell 2002, 3, 765–778. [Google Scholar] [CrossRef]
- Malakhov, M.P.; Malakhova, O.A.; Kim, K.I.; Ritchie, K.J.; Zhang, D.-E. UBP43 (USP18) Specifically Removes ISG15 from Conjugated Proteins. J. Biol. Chem. 2002, 277, 9976–9981. [Google Scholar] [CrossRef] [PubMed]
- Levy, A.J.; Scott, I.; Mackewicz, C. Protection from HIV/AIDS: The importance of innate immunity. Clin. Immunol. 2003, 108, 167–174. [Google Scholar] [CrossRef]
- Shirazi, Y.; Pitha, P.M. Alpha interferon inhibits early stages of the human immunodeficiency virus type 1 replication cycle. J. Virol. 1992, 66, 1321–1328. [Google Scholar] [CrossRef]
- Cheney, K.M.; Áine, M. Interferon-Alpha Mediates Restriction of Human Immunodeficiency Virus Type-1 Replication in Primary Human Macrophages at an Early Stage of Replication. PLoS ONE 2010, 5, e13521. [Google Scholar] [CrossRef] [PubMed]
- Pincetic, A.; Kuang, Z.; Seo, E.J.; Leis, J. The Interferon-Induced Gene ISG15 Blocks Retrovirus Release from Cells Late in the Budding Process. J. Virol. 2010, 84, 4725–4736. [Google Scholar] [CrossRef]
- Woods, M.W.; Kelly, J.N.; Hattlmann, C.J.; Tong, J.G.K.; Xu, L.S.; Coleman, M.D.; Quest, G.R.; Smiley, J.R.; Barr, S.D. Human HERC5 restricts an early stage of HIV-1 assembly by a mechanism correlating with the ISGylation of Gag. Retrovirology 2011, 8, 95. [Google Scholar] [CrossRef]
- Okumura, A.; Lu, G.; Pitha-Rowe, I.; Pitha, P.M. Innate antiviral response targets HIV-1 release by the induction of ubiquitin-like protein ISG15. Proc. Natl. Acad. Sci. USA 2006, 103, 1440–1445. [Google Scholar] [CrossRef] [PubMed]
- Morales, D.J.; Lenschow, D.J. The antiviral activities of ISG15. J. Mol. Biol. 2013, 425, 4995–5008. [Google Scholar] [CrossRef]
- Zou, W.; Papov, V.; Malakhova, O.; Kim, K.I.; Dao, C.; Li, J.; Zhang, D.-E. ISG15 modification of ubiquitin E2 Ubc13 disrupts its ability to form thioester bond with ubiquitin. Biochem. Biophys. Res. Commun. 2005, 336, 61–68. [Google Scholar] [CrossRef]
- Ambler, R.P.; Rees, M.W. ɛ-N-Methyl-lysine in Bacterial Flagellar Protein. Nat. Cell Biol. 1959, 184, 56–57. [Google Scholar] [CrossRef]
- Levy, D. Lysine methylation signaling of non-histone proteins in the nucleus. Cell. Mol. Life Sci. 2019, 76, 2873–2883. [Google Scholar] [CrossRef]
- Lanouette, S.; Mongeon, V.; Figeys, D.; Couture, J. The functional diversity of protein lysine methylation. Mol. Syst. Biol. 2014, 10, 724. [Google Scholar] [CrossRef]
- Blanchet, F.; Cardona, A.; Letimier, F.A.; Hershfield, M.S.; Acuto, O. CD28 costimulatory signal induces protein arginine methylation in T cells. J. Exp. Med. 2005, 202, 371–377. [Google Scholar] [CrossRef] [PubMed]
- Smith, W.A.; Schurter, B.T.; Wong-Staal, F.; David, M. Arginine Methylation of RNA Helicase A Determines Its Subcellular Localization. J. Biol. Chem. 2004, 279, 22795–22798. [Google Scholar] [CrossRef] [PubMed]
- Willemsen, N.M.; Hitchen, E.M.; Bodetti, T.J.; Apolloni, A.; Warrilow, D.; Piller, S.C.; Harrich, D. Protein methylation is required to maintain optimal HIV-1 infectivity. Retrovirology 2006, 3, 92. [Google Scholar] [CrossRef]
- Lochmann, T.L.; Bann, D.V.; Ryan, E.P.; Beyer, A.R.; Mao, A.; Cochrane, A.; Parent, L.J. NC-mediated nucleolar localization of retroviral gag proteins. Virus Res. 2013, 171, 304–318. [Google Scholar] [CrossRef] [PubMed]
- Invernizzi, C.D.F.; Xie, B.; Frankel, F.A.; Feldhammer, M.; Roy, B.B.; Richard, S.; Wainberg, M.A. Arginine methylation of the HIV-1 nucleocapsid protein results in its diminished function. AIDS 2007, 21, 795–805. [Google Scholar] [CrossRef] [PubMed]
- Paris, J.; Tobaly-Tapiero, J.; Giron, M.-L.; Burlaud-Gaillard, J.; Buseyne, F.; Roingeard, P.; Lesage, P.; Zamborlini, A.; Saïb, A. The invariant arginine within the chromatin-binding motif regulates both nucleolar localization and chromatin binding of Foamy virus Gag. Retrovirology 2018, 15, 48. [Google Scholar] [CrossRef] [PubMed]



| Domain | Residue | Enzyme | Observations and Associated (or Proposed) Roles | References |
|---|---|---|---|---|
| MA | S9 | ERK2 | Involved in the viral replication Phosphorylation of the mature form of MA | [95,101] |
| S67 | ||||
| S72 | ||||
| S77 | ||||
| S111 | PKC could be involved in membrane binding by regulating the exposure of the myristoyl group | [82,83,84] | ||
| Y132 | Src | In MA mature 1% of Y132 is phosphorylated Src overexpression was found to promote the localization of Pr55Gag at the PM | [102,103] | |
| CA | S148 | ERK2 | Belongs to S-P motif involved in recruitment of ERK-2 | [93,94,95,96] |
| p6 | T456 | Belongs to the PTAP late domain Potential role in viral infectivity and assembly | [86] | |
| T470 | Redundancy with T471, S473, S488, S491, and S499 | [91] | ||
| T471 | ERK-2 | Belongs to T-P motif involved in the recruitment of ERK-2 | [91] | |
| Its substitution induces the accumulation of immature viral particles incompletely separated from PM | [91] | |||
| Redundancy with T470, S473, S488, S491, and S499 | [91] | |||
| Effects on assembly or on viral release is not due to phosphorylation | [86] | |||
| S473 | Redundancy with T470, S471, S491, and S499 | [91] | ||
| S488 | ERK2 | Viral particles without active ERK2 were found to be poorly infectious due to a defect in reverse transcription | [93,95] | |
| Involved in the phosphorylation of other viral proteins: Rev, Nef, Vif, mature MA | [97,98,99,100] | |||
| PKC | The p6 domain of Pr55Gag is a target for PKC | [85,86,90] | ||
| The inhibition of PKC activity reduced Vpr level in virions | [85,87,88] | |||
| Its mutation with F perturbs: - Viral morphology, maturation and infectivity | [87,88,89] | |||
| Effects on assembly or on viral release could be not due to phosphorylation | [86] | |||
| S491 | Redundancy with T470, S471, S473, and S499 | [91] | ||
| S499 | Redundancy with T470, S471, S473, and S491 | [91] |
| Retrovirus | Protein | Residues | Enzyme | Observation and Associated (or Proposed) Roles | References |
|---|---|---|---|---|---|
| RSV | MA | Y15 | PKC | No effect on the viral cycle | [105,106] |
| Y46 | |||||
| S68 | |||||
| S106 | PKC | Major site of phosphorylation Involved in the recruitment of factors which promote NC phosphorylation | [105,106] | ||
| Y155 | PKC | No effect on the viral cycle | [105,106] | ||
| NC | S529 | Role for the specific interaction with the gRNA | [106,107] | ||
| HTLV-1 | MA | S105 | ERK2 | Close to late domains (PPPY et PTAP) Involved in viral release and budding efficiency | [110] |
| MoMuLV | p12 | S137 | - Redundancy - Modulation of early and late functions and the RNA-binding activity of p12 | [117] | |
| S148 | |||||
| S150 | |||||
| S173 | |||||
| S192 | - S192 mainly contributes to p12 phosphorylation and its substitution by A impairs viral assembly and infectivity | [117] | |||
| S209 | |||||
| MPMV | p18 | Y205 | Belongs to proline-rich motif (PPPY) Necessary for the viral release | [112] | |
| S167 | Redundancy | [113,114] | |||
| S176 | |||||
| S211 | |||||
| FV | p4 | S116 | Redundancy | [115] | |
| S119 | |||||
| S120 | |||||
| S124 | |||||
| Domain | Residues | Observation and Associated (or Proposed) Roles | References |
|---|---|---|---|
| MA | Mono-ubiquitination | [125,126] | |
| CA | K157 K162 K202 K263 K272 K290 K302 K314 K331 K335 K359 | Mono-ubiquitination Observed redundancy | [125,126] |
| NC | K388 K391 K397 K403 K410 K411 K415 K424 | Mono-ubiquitination | [125,126] |
| p2 | K436 K442 | Mono or di-ubiquitination | [125,126] |
| p6 | Mono or di-ubiquitination. Most ubiquitinated domain in Pr55Gag | [125,126] | |
| K475 | Major target for mono-ubiquitination No effect in the viral release and infectivity Involved in global Pr55Gag ubiquitination | [125,132] | |
| K481 | Major targets for mono-ubiquitination No effect on virus release and infectivity Involved in the global Pr55Gag ubiquitination | [125,132] | |
| S488F | Conformal changes: formation of a hydrophobic patch in a-helix at the C-terminus of p6 Leads to strong interaction of Pr55Gag with the PM This structure promotes L48 linked polyubiquitination | [104] |
| Retrovirus | Domain | Residues | Associated (or Proposed) Roles | References |
|---|---|---|---|---|
| HIV | About 100 free Ubs are incorporated into viral particles 2–5% mono-ubiquitinated | [130,132,133,134] | ||
| Pr55Gag ubiquitination promotes the virus release K475 and K481 in p6 domain are major targets for ubiquitinations Pr55Gag ubiquitination is correlated with the ability of the precursor to bind the PM | [132,134] | |||
| MLV | Increases viral release and infectivity | [127] | ||
| p12 | PPPY late domain is involved in the recruitment of NEDD4 | [148] | ||
| HTLV-1 | MA | Ubiquitination of this domain has a crucial role in release | [111,145,146] | |
| 40% of MA are ubiquitinated MA can be mono- or di-ubiquitinated | [111,145,146] | |||
| K74 | Substrate for Pr53Gag ubiquitination | [146] | ||
| MPMV | PPPY late domain is involved in the recruitment of NEDD4 | [112] | ||
| RSV | - Mono-ubiquitination is crucial for viral release - Ubiquitylation is required for the recruitment of ESCRT machinery and for the budding | [133,143,149] | ||
| - Contains free Ubs into mature particles - Pr76Gag mono-ubiquitination is necessary for budding and to recruit the ESCRT machinery | [142,149] | |||
| EIAV | 10–15% of the molar level of the Gag protein of free Ub | [143] | ||
| Proteasome inhibition: does not impair the release | [143] | |||
| p9 | Ub-like motif (NVKEKD) | |||
| Mono-ubiquitinated domain Contains YPDL late domain | [143,144] | |||
| MMTV | Proteasome inhibition: does not decrease the release | [143] | ||
| MA (p10) | YXXL Late domain | [127] | ||
| pp21 | YXXL Late domain | [127] | ||
| p8 | Mono-ubiquitinated | [127] | ||
| CA (p27) | Mono-ubiquitinated | [127] | ||
| PSAP domain | [127] | |||
| NC (p14) | Di-ubiquitinated | [127] | ||
| PFV | Encodes for a very restricted number of K residues | [150] |
| Retrovirus | Domains | Residues | Associated (or Proposed) Roles | References |
|---|---|---|---|---|
| HIV-1 | Sumoylation and ubiquitination co-regulate each other | [165] | ||
| p6 | More than one domain should be involved in Ubc9 recruitment | [162,163] | ||
| SUMO-Ubc9 could be involved in intracellular trafficking of Pr55Gag - in the early phase: perinuclear region or - in the late stages of replication: potential role in Env incorporation | [163,164] | |||
| K 475 | Sumoylation could be then involved in the negative regulation of viral replication | [162] | ||
| Belongs to QKQE consensus sequence Sumoylation and mono-ubiquitination of p6 can both occur on K475 | [162] | |||
| MoMuLV | CA | CA domain of MLV Gag interacts with Ubc9 and with PIASy | [168] | |
| MPMV | Recruitment of Ubc9 involved in the active transport of MPMV Pr78Gag to the PM | [167] | ||
| RSV | CA | K 244 | Its substitution with non sumoylable R reduces the overall viral infectivity | [142] |
| EIAV | MA | K 13 K 86 K 107 | Targets of sumoylation | [144,169] |
| CA | K 282 K 297 | |||
| NC | K 368 K 373 K 388 K 420 K 423 | |||
| p9 | K 465 | Constitutes the main target for sumoylation | [144,169] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bussienne, C.; Marquet, R.; Paillart, J.-C.; Bernacchi, S. Post-Translational Modifications of Retroviral HIV-1 Gag Precursors: An Overview of Their Biological Role. Int. J. Mol. Sci. 2021, 22, 2871. https://doi.org/10.3390/ijms22062871
Bussienne C, Marquet R, Paillart J-C, Bernacchi S. Post-Translational Modifications of Retroviral HIV-1 Gag Precursors: An Overview of Their Biological Role. International Journal of Molecular Sciences. 2021; 22(6):2871. https://doi.org/10.3390/ijms22062871
Chicago/Turabian StyleBussienne, Charlotte, Roland Marquet, Jean-Christophe Paillart, and Serena Bernacchi. 2021. "Post-Translational Modifications of Retroviral HIV-1 Gag Precursors: An Overview of Their Biological Role" International Journal of Molecular Sciences 22, no. 6: 2871. https://doi.org/10.3390/ijms22062871
APA StyleBussienne, C., Marquet, R., Paillart, J.-C., & Bernacchi, S. (2021). Post-Translational Modifications of Retroviral HIV-1 Gag Precursors: An Overview of Their Biological Role. International Journal of Molecular Sciences, 22(6), 2871. https://doi.org/10.3390/ijms22062871









