OsCBE1, a Substrate Receptor of Cullin4-Based E3 Ubiquitin Ligase, Functions as a Regulator of Abiotic Stress Response and Productivity in Rice
Abstract
1. Introduction
2. Results
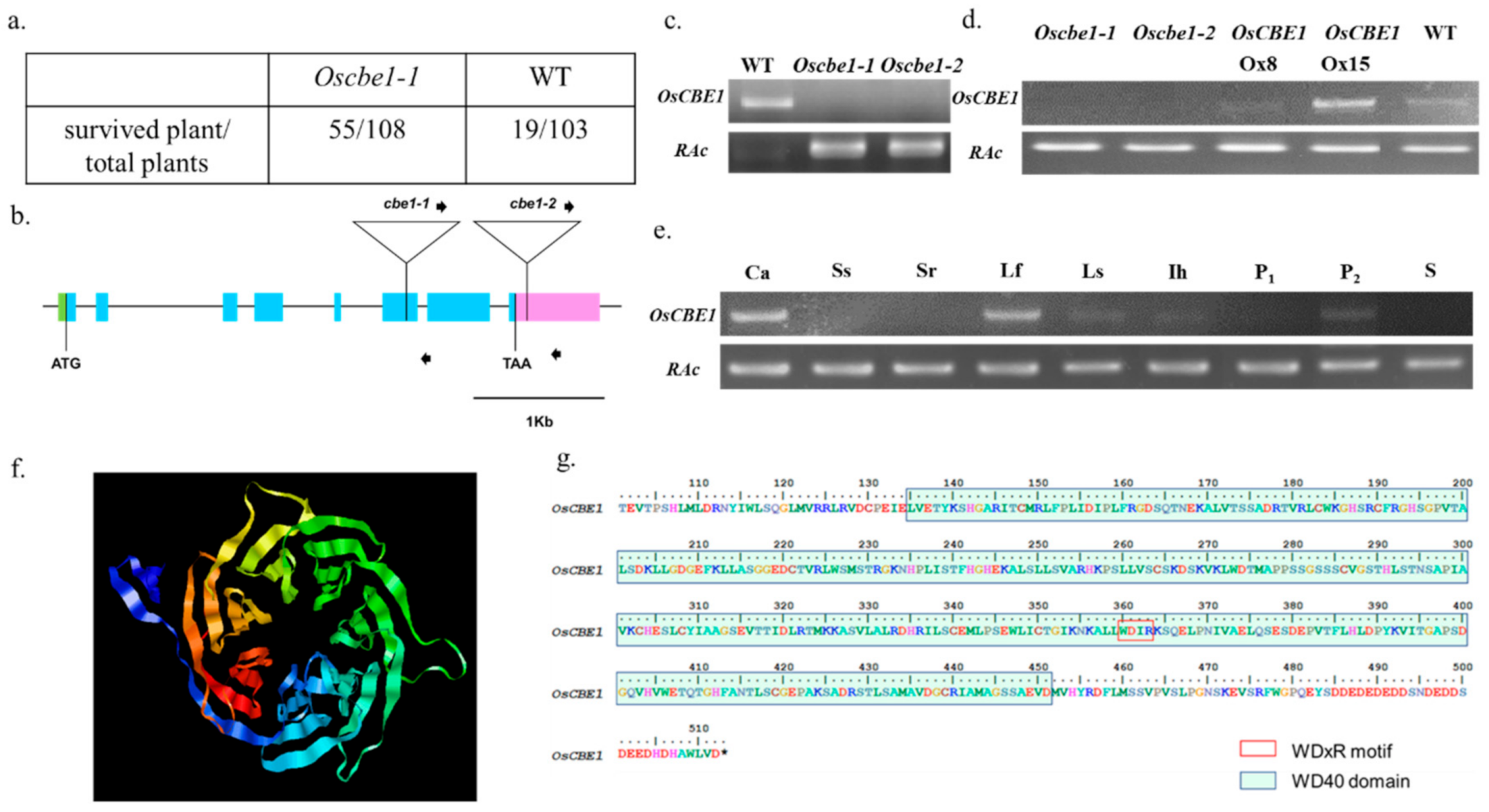
2.1. Identification of Cold Stress Mutant Oscbe1
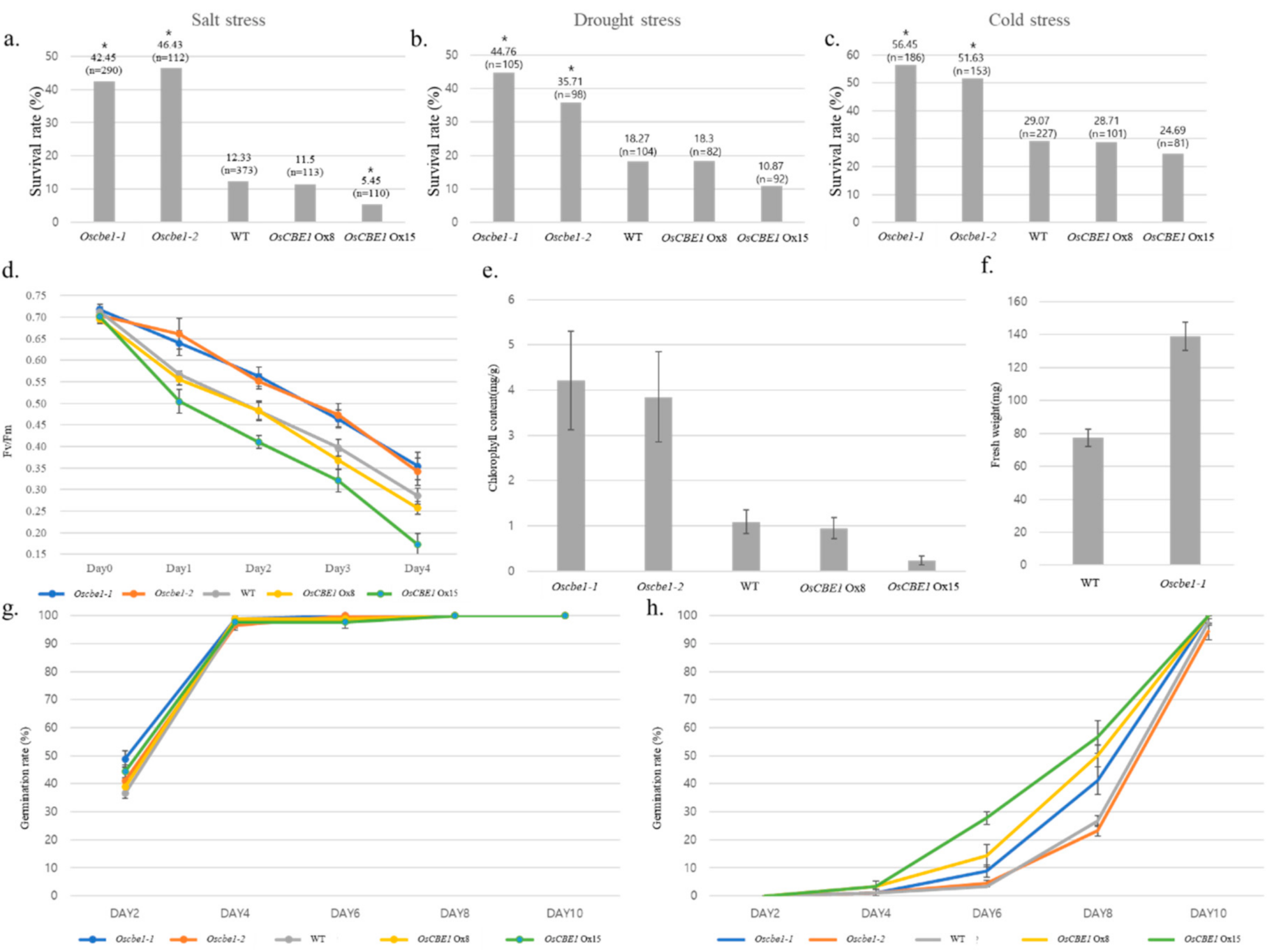
2.2. OsCBE1 Negatively Regulates Stress Response and ABA Signaling
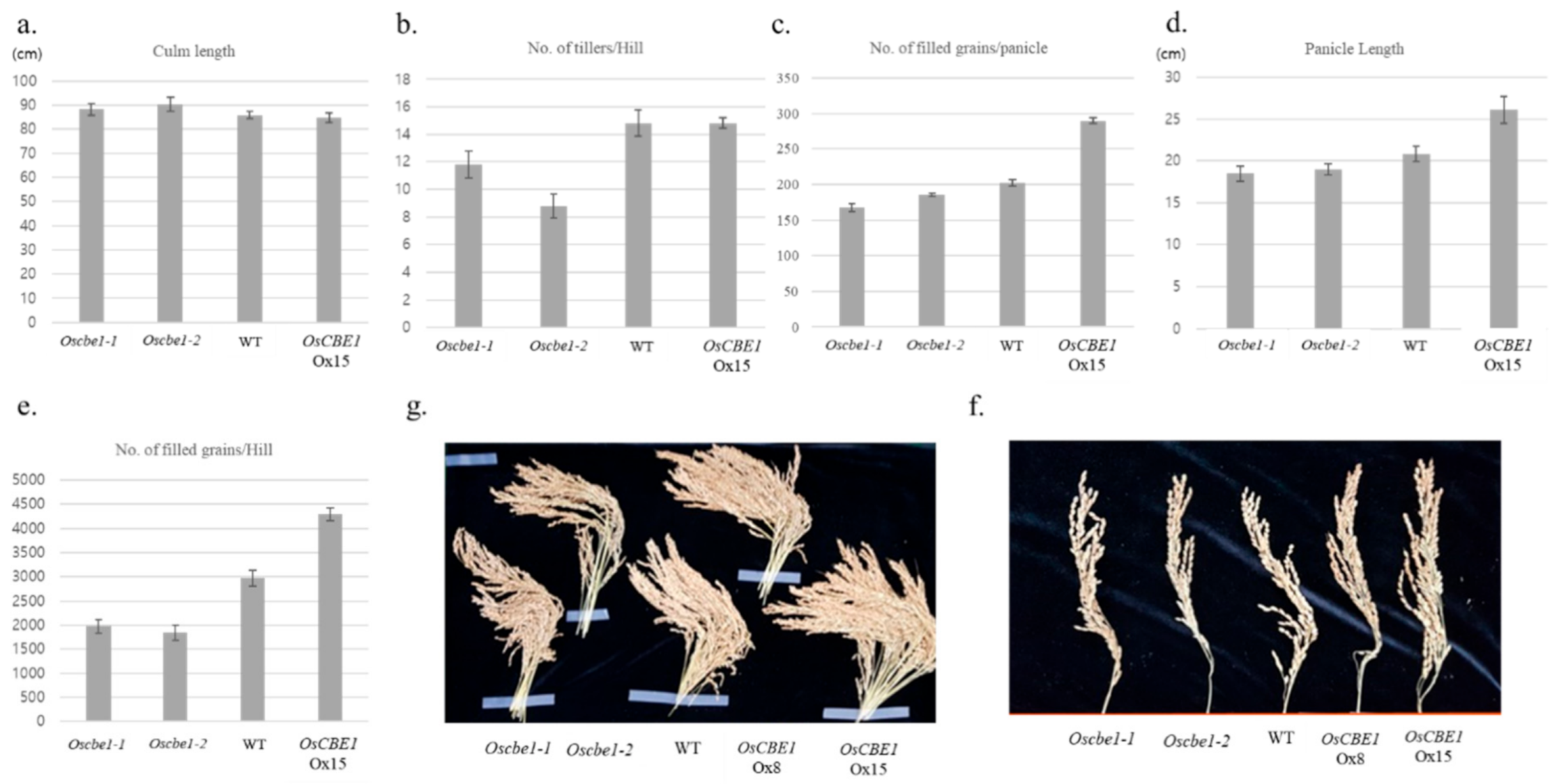
2.3. Expression of OsCBE1 Positively Related to Crop Productivity
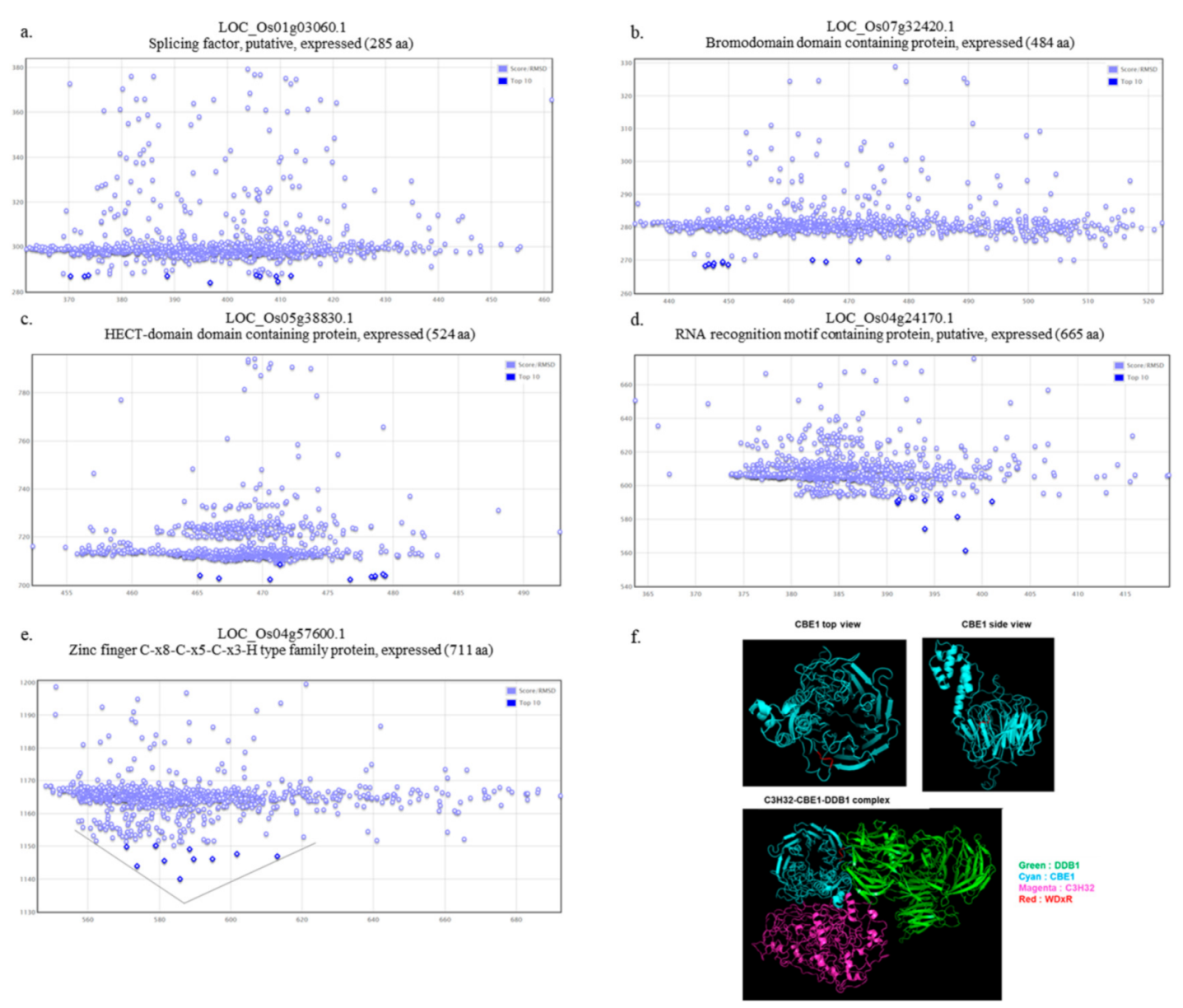
2.4. Functional Role of OsCBE1 and In Silico Identification of the OsCBE1 Binding Partners
2.5. Physical Interaction of OsCBE1 with OsDDB1 and OsC3H32
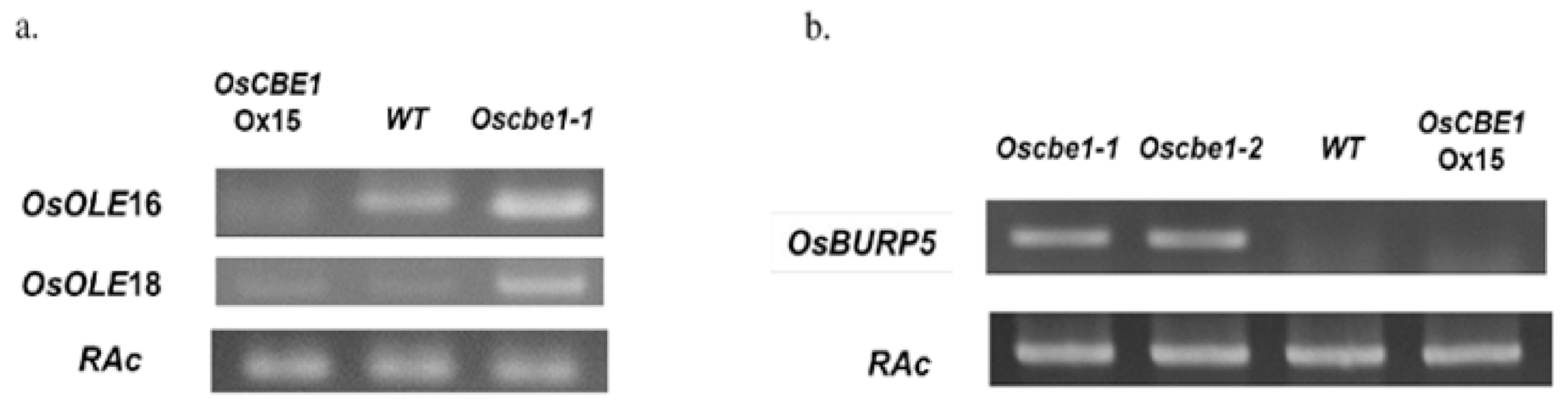
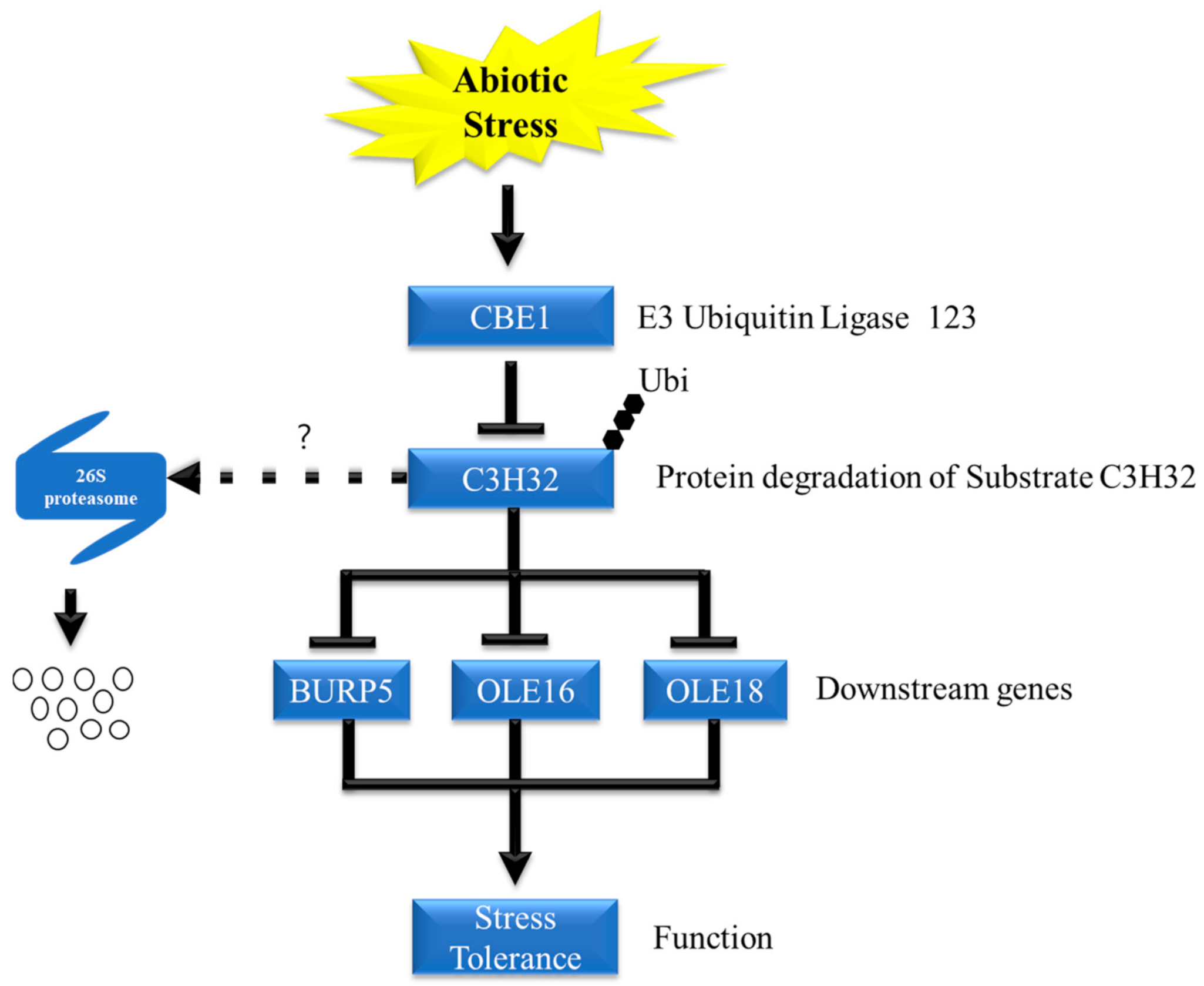
2.6. Oleosin Genes and an ABA Responsive Gene Are Negatively Regulated by OsCBE1
3. Discussion
4. Materials and Methods
4.1. Plant Growth and Stress Tolerance Test
4.2. Analysis of Chlorophyll Content and Fluorescence
4.3. PCR and RT-PCR Analysis
4.4. Vector Construction
4.5. Generation of OsCBE1 Overexpression Lines
4.6. Germination Test
4.7. Evaluation of Agronomic Traits
4.8. In Silico Analysis
4.9. Co-IP Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Van Velthuizen, H. Mapping Biophysical Factors that Influence Agricultural Production and Rural Vulnerability; Food & Agriculture Organization: Rome, Italy, 2007. [Google Scholar]
- Ashraf, M.; Ahmad, M.S.A.; Öztürk, M.; Aksoy, A. Crop improvement through different means: Challenges and prospects. In Crop Production for Agricultural Improvement; Springer: Berlin, Germany, 2012; pp. 1–15. [Google Scholar]
- Corlett, R.T.; Westcott, D.A. Will plant movements keep up with climate change? Trends Ecol. Evol. 2013, 28, 482–488. [Google Scholar] [CrossRef]
- Zhu, J.-K. Abiotic stress signaling and responses in plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef]
- Yuan, F.; Yang, H.; Xue, Y.; Kong, D.; Ye, R.; Li, C.; Zhang, J.; Theprungsirikul, L.; Shrift, T.; Krichilsky, B. OSCA1 mediates osmotic-stress-evoked Ca 2+ increases vital for osmosensing in Arabidopsis. Nature 2014, 514, 367. [Google Scholar] [CrossRef]
- Ma, Y.; Dai, X.; Xu, Y.; Luo, W.; Zheng, X.; Zeng, D.; Pan, Y.; Lin, X.; Liu, H.; Zhang, D. COLD1 confers chilling tolerance in rice. Cell 2015, 160, 1209–1221. [Google Scholar] [CrossRef] [PubMed]
- Park, S.-Y.; Fung, P.; Nishimura, N.; Jensen, D.R.; Fujii, H.; Zhao, Y.; Lumba, S.; Santiago, J.; Rodrigues, A.; Tsz-fung, F.C. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 2009, 324, 1068–1071. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Szostkiewicz, I.; Korte, A.; Moes, D.; Yang, Y.; Christmann, A.; Grill, E. Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 2009, 324, 1064–1068. [Google Scholar] [CrossRef] [PubMed]
- Boudsocq, M.; Barbier-Brygoo, H.; Laurière, C. Identification of nine sucrose nonfermenting 1-related protein kinases 2 activated by hyperosmotic and saline stresses in Arabidopsis thaliana. J. Biol. Chem. 2004, 279, 41758–41766. [Google Scholar] [CrossRef]
- Choi, H.-I.; Hong, J.-H.; Ha, J.-O.; Kang, J.-Y.; Kim, S.Y. ABFs, a family of ABA-responsive element binding factors. J. Biol. Chem. 2000, 275, 1723–1730. [Google Scholar] [CrossRef]
- Abe, H.; Yamaguchi-Shinozaki, K.; Urao, T.; Iwasaki, T.; Hosokawa, D.; Shinozaki, K. Role of Arabidopsis MYC and MYB homologs in drought-and abscisic acid-regulated gene expression. Plant Cell 1997, 9, 1859–1868. [Google Scholar]
- Agarwal, P.K.; Agarwal, P.; Reddy, M.; Sopory, S.K. Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep. 2006, 25, 1263–1274. [Google Scholar] [CrossRef]
- Nakashima, K.; Takasaki, H.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. NAC transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta BBA Gene Regul. Mech. 2012, 1819, 97–103. [Google Scholar] [CrossRef]
- Dubouzet, J.G.; Sakuma, Y.; Ito, Y.; Kasuga, M.; Dubouzet, E.G.; Miura, S.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt-and cold-responsive gene expression. Plant J. 2003, 33, 751–763. [Google Scholar] [CrossRef] [PubMed]
- Gilmour, S.J.; Sebolt, A.M.; Salazar, M.P.; Everard, J.D.; Thomashow, M.F. Overexpression of the Arabidopsis CBF3transcriptional activator mimics multiple biochemical changes associated with cold acclimation. Plant Phys. 2000, 124, 1854–1865. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Zou, H.-F.; Wang, H.-W.; Zhang, W.-K.; Ma, B.; Zhang, J.-S.; Chen, S.-Y. Soybean GmMYB76, GmMYB92, and GmMYB177 genes confer stress tolerance in transgenic Arabidopsis plants. Cell Res. 2008, 18, 1047. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Gong, F.; Cao, D.; Hu, X.; Wang, W. Advances in crop proteomics: PTMs of proteins under abiotic stress. Proteomics 2016, 16, 847–865. [Google Scholar] [CrossRef] [PubMed]
- Komander, D.; Rape, M. The ubiquitin code. Annu. Rev. Biochem. 2012, 81, 203–229. [Google Scholar] [CrossRef] [PubMed]
- Ohtake, F.; Tsuchiya, H. The emerging complexity of ubiquitin architecture. J. Biochem. 2016, 161, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, Y.; de la Bastide, M.; Hamilton, J.P.; Kanamori, H.; McCombie, W.R.; Ouyang, S.; Schwartz, D.C.; Tanaka, T.; Wu, J.; Zhou, S. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 2013, 6, 4. [Google Scholar] [CrossRef]
- Lamesch, P.; Berardini, T.Z.; Li, D.; Swarbreck, D.; Wilks, C.; Sasidharan, R.; Muller, R.; Dreher, K.; Alexander, D.L.; Garcia-Hernandez, M. The Arabidopsis Information Resource (TAIR): Improved gene annotation and new tools. Nucleic Acids Res. 2011, 40, D1202–D1210. [Google Scholar] [CrossRef]
- Du, Z.; Zhou, X.; Li, L.; Su, Z. PlantsUPS: A database of plants’ Ubiquitin Proteasome System. BMC Gen. 2009, 10, 227. [Google Scholar] [CrossRef]
- Lee, J.-H.; Terzaghi, W.; Gusmaroli, G.; Charron, J.-B.F.; Yoon, H.-J.; Chen, H.; He, Y.J.; Xiong, Y.; Deng, X.W. Characterization of Arabidopsis and rice DWD proteins and their roles as substrate receptors for CUL4-RING E3 ubiquitin ligases. Plant Cell 2008, 20, 152–167. [Google Scholar] [CrossRef]
- Smalle, J.; Vierstra, R.D. The ubiquitin 26S proteasome proteolytic pathway. Annu. Rev. Plant Biol. 2004, 55, 555–590. [Google Scholar] [CrossRef]
- Gray, W.M.; del Pozo, J.C.; Walker, L.; Hobbie, L.; Risseeuw, E.; Banks, T.; Crosby, W.L.; Yang, M.; Ma, H.; Estelle, M. Identification of an SCF ubiquitin–ligase complex required for auxin response in Arabidopsis thaliana. Gen. Dev. 1999, 13, 1678–1691. [Google Scholar] [CrossRef] [PubMed]
- Marrocco, K.; Thomann, A.; Parmentier, Y.; Genschik, P.; Criqui, M.C. The APC/C E3 ligase remains active in most post-mitotic Arabidopsis cells and is required for proper vasculature development and organization. Development 2009, 136, 1475–1485. [Google Scholar] [CrossRef] [PubMed]
- Figueroa, P.; Gusmaroli, G.; Serino, G.; Habashi, J.; Ma, L.; Shen, Y.; Feng, S.; Bostick, M.; Callis, J.; Hellmann, H. Arabidopsis has two redundant Cullin3 proteins that are essential for embryo development and that interact with RBX1 and BTB proteins to form multisubunit E3 ubiquitin ligase complexes in vivo. Plant Cell 2005, 17, 1180–1195. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Shen, Y.; Tang, X.; Yu, L.; Wang, J.; Guo, L.; Zhang, Y.; Zhang, H.; Feng, S.; Strickland, E. Arabidopsis CULLIN4 forms an E3 ubiquitin ligase with RBX1 and the CDD complex in mediating light control of development. Plant Cell 2006, 18, 1991–2004. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Feng, S.; Chen, F.; Chen, H.; Wang, J.; McCall, C.; Xiong, Y.; Deng, X.W. Arabidopsis DDB1-CUL4 ASSOCIATED FACTOR1 forms a nuclear E3 ubiquitin ligase with DDB1 and CUL4 that is involved in multiple plant developmental processes. Plant Cell 2008, 20, 1437–1455. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Huang, X.; Gusmaroli, G.; Terzaghi, W.; Lau, O.S.; Yanagawa, Y.; Zhang, Y.; Li, J.; Lee, J.-H.; Zhu, D. Arabidopsis CULLIN4-damaged DNA binding protein 1 interacts with constitutively photomorphogenic1-suppressor of phya complexes to regulate photomorphogenesis and flowering time. Plant Cell 2010, 22, 108–123. [Google Scholar] [CrossRef] [PubMed]
- Maeda, K.; Houjyou, Y.; Komatsu, T.; Hori, H.; Kodaira, T.; Ishikawa, A. AGB1 and PMR5 contribute to PEN2-mediated preinvasion resistance to Magnaporthe oryzae in Arabidopsis thaliana. Mol. Plant-Microbe Interact. 2009, 22, 1331–1340. [Google Scholar] [CrossRef]
- Lee, J.-H.; Yoon, H.-J.; Terzaghi, W.; Martinez, C.; Dai, M.; Li, J.; Byun, M.-O.; Deng, X.W. DWA1 and DWA2, two Arabidopsis DWD protein components of CUL4-based E3 ligases, act together as negative regulators in ABA signal transduction. Plant Cell 2010, 22, 1716–1732. [Google Scholar] [CrossRef]
- Lee, J.; Zhou, P. DCAFs, the missing link of the CUL4-DDB1 ubiquitin ligase. Mol. Cell 2007, 26, 775–780. [Google Scholar] [CrossRef]
- Biedermann, S.; Hellmann, H. WD40 and CUL4-based E3 ligases: Lubricating all aspects of life. Trends Plant Sci. 2011, 16, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R. CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2016, 45, D200–D203. [Google Scholar] [CrossRef]
- Kim, D.E.; Chivian, D.; Baker, D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 2004, 32, W526–W531. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Franceschini, A.; Wyder, S.; Forslund, K.; Heller, D.; Huerta-Cepas, J.; Simonovic, M.; Roth, A.; Santos, A.; Tsafou, K.P. STRING v10: Protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2014, 43, D447–D452. [Google Scholar] [CrossRef]
- Lyskov, S.; Chou, F.-C.; Conchúir, S.Ó.; Der, B.S.; Drew, K.; Kuroda, D.; Xu, J.; Weitzner, B.D.; Renfrew, P.D.; Sripakdeevong, P. Serverification of molecular modeling applications: The Rosetta Online Server that Includes Everyone (ROSIE). PLoS ONE 2013, 8, e63906. [Google Scholar] [CrossRef]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 2015, 10, 845. [Google Scholar] [CrossRef]
- Seok, H.-Y.; Nguyen, L.V.; Park, H.-Y.; Tarte, V.N.; Ha, J.; Lee, S.-Y.; Moon, Y.-H. Arabidopsis non-TZF gene AtC3H17 functions as a positive regulator in salt stress response. Biochem. Biophys. Res. Commun. 2018, 498, 954–959. [Google Scholar] [CrossRef] [PubMed]
- Seok, H.-Y.; Woo, D.-H.; Park, H.-Y.; Lee, S.-Y.; Tran, H.T.; Lee, E.-H.; Vu Nguyen, L.; Moon, Y.-H. AtC3H17, a non-tandem CCCH zinc finger protein, functions as a nuclear transcriptional activator and has pleiotropic effects on vegetative development, flowering and seed development in Arabidopsis. Plant Cell Physiol. 2016, 57, 603–615. [Google Scholar] [CrossRef]
- Seo, K.-I.; Lee, J.-H.; Nezames, C.D.; Zhong, S.; Song, E.; Byun, M.-O.; Deng, X.W. ABD1 is an Arabidopsis DCAF substrate receptor for CUL4-DDB1–based E3 ligases that acts as a negative regulator of abscisic acid signaling. Plant Cell 2014, 26, 695–711. [Google Scholar] [CrossRef]
- Dutilleul, C.; Ribeiro, I.; Blanc, N.; Nezames, C.D.; Deng, X.W.; Zglobicki, P.; Palacio Barrera, A.M.; Atehortùa, L.; Courtois, M.; Labas, V. ASG2 is a farnesylated DWD protein that acts as ABA negative regulator in Arabidopsis. Plant Cell Environ. 2016, 39, 185–198. [Google Scholar] [CrossRef]
- Irigoyen, M.L.; Iniesto, E.; Rodriguez, L.; Puga, M.I.; Yanagawa, Y.; Pick, E.; Strickland, E.; Paz-Ares, J.; Wei, N.; De Jaeger, G. Targeted degradation of abscisic acid receptors is mediated by the ubiquitin ligase substrate adaptor DDA1 in Arabidopsis. Plant Cell 2014, 26, 712–728. [Google Scholar] [CrossRef]
- Fernando, V.D.; Schroeder, D.F. Arabidopsis DDB1-CUL4 E3 ligase complexes in det1 salt/osmotic stress resistant germination. Plant Signal. Behav. 2016, 11, e1223004. [Google Scholar] [CrossRef][Green Version]
- Lee, I.; Ambaru, B.; Thakkar, P.; Marcotte, E.M.; Rhee, S.Y. Rational association of genes with traits using a genome-scale gene network for Arabidopsis thaliana. Nat. Biotechnol. 2010, 28, 149. [Google Scholar] [CrossRef]
- Lee, J.-H.; Terzaghi, W.; Deng, X.W. DWA3, an Arabidopsis DWD protein, acts as a negative regulator in ABA signal transduction. Plant Sci. 2011, 180, 352–357. [Google Scholar] [CrossRef]
- Ali, A.; Yun, D.-J. Arabidopsis HOS15 is a multifunctional protein that negatively regulate ABA-signaling and drought stress. Plant Biotechnol. Rep. 2020, 14, 1–5. [Google Scholar] [CrossRef]
- Li, D.; Zhang, L.; Li, X.; Kong, X.; Wang, X.; Li, Y.; Liu, Z.; Wang, J.; Li, X.; Yang, Y. AtRAE1 is involved in degradation of ABA receptor RCAR1 and negatively regulates ABA signalling in Arabidopsis. Plant Cell Environ. 2018, 41, 231–244. [Google Scholar] [CrossRef] [PubMed]
- Park, S.-R.; Hwang, J.; Kim, M. The Arabidopsis WDR55 is positively involved in ABA-mediated drought tolerance response. Plant Biotechnol. Rep. 2020, 14, 407–418. [Google Scholar] [CrossRef]
- Bae, Y.; Kim, H.; Kim, S.-H.; Lee, J.-H. Expression study on the Scaffold gene of CRL4 complex in rice (Oryza sativa L.). J. Life Sci. 2018, 28, 1132–1139. [Google Scholar]
- Shabek, N.; Ruble, J.; Waston, C.J.; Garbutt, K.C.; Hinds, T.R.; Li, T.; Zheng, N. Structural insights into DDA1 function as a core component of the CRL4-DDB1 ubiquitin ligase. Cell Discov. 2018, 4, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.S.; Zhao, P.; Jung, C.; Chua, N.-H. PLANT U-BOX PROTEIN 10 negatively regulates abscisic acid response in Arabidopsis. Appl. Biol. Chem. 2019, 62, 39. [Google Scholar] [CrossRef]
- Wang, N.; Liu, Y.; Cai, Y.; Tang, J.; Li, Y.; Gai, J. The soybean U-box gene GmPUB6 regulates drought tolerance in Arabidopsis thaliana. Plant Physiol. Biochem. 2020, 155, 284–296. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Liu, Y.; Cong, Y.; Wang, T.; Zhong, X.; Yang, S.; Li, Y.; Gai, J. Genome-wide identification of soybean U-box E3 ubiquitin ligases and roles of GmPUB8 in negative regulation of drought stress response in Arabidopsis. Plant Cell Physiol. 2016, 57, 1189–1209. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.K.; Ryu, M.Y.; Kim, J.H.; Hong, J.S.; Oh, T.R.; Kim, W.T.; Yang, S.W. RING E3 ligases: Key regulatory elements are involved in abiotic stress responses in plants. BMB Rep. 2017, 50, 393. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Zhao, J.; Li, L.; Gao, Y.; Zhao, L.; Patil, S.B.; Fang, J.; Zhang, W.; Yang, Y.; Li, M. The Arabidopsis U-box E3 ubiquitin ligase PUB30 negatively regulates salt tolerance by facilitating BRI1 kinase inhibitor 1 (BKI1) degradation. Plant Cell Environ. 2017, 40, 2831–2843. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhao, L.; Zhang, M.; Zafar, S.A.; Fang, J.; Li, M.; Zhang, W.; Li, X. Arabidopsis E3 ubiquitin ligases PUB22 and PUB23 negatively regulate drought tolerance by targeting ABA receptor PYL9 for degradation. Int. J. Mol. Sci. 2017, 18, 1841. [Google Scholar] [CrossRef]
- Seo, D.H.; Ahn, M.Y.; Park, K.Y.; Kim, E.Y.; Kim, W.T. The N-terminal UND motif of the Arabidopsis U-box E3 ligase PUB18 is critical for the negative regulation of ABA-mediated stomatal movement and determines its ubiquitination specificity for exocyst subunit Exo70B1. Plant Cell 2016, 28, 2952–2973. [Google Scholar] [CrossRef]
- Seo, D.H.; Ryu, M.Y.; Jammes, F.; Hwang, J.H.; Turek, M.; Kang, B.G.; Kwak, J.M.; Kim, W.T. Roles of four Arabidopsis U-box E3 ubiquitin ligases in negative regulation of abscisic acid-mediated drought stress responses. Plant Physiol. 2012, 160, 556–568. [Google Scholar] [CrossRef] [PubMed]
- Makino, A. Photosynthesis, grain yield, and nitrogen utilization in rice and wheat. Plant Physiol. 2011, 155, 125–129. [Google Scholar] [CrossRef]
- Yoshida, S. Fundamentals of Rice Crop Science; The International Rice Research Institute: Los Baños, Philippines, 1981. [Google Scholar]
- Yoshida, H.; Horie, T.; Shiraiwa, T. A model explaining genotypic and environmental variation of rice spikelet number per unit area measured by cross-locational experiments in Asia. Field Crops Res. 2006, 97, 337–343. [Google Scholar] [CrossRef]
- Kim, H.; Lee, K.; Hwang, H.; Bhatnagar, N.; Kim, D.-Y.; Yoon, I.S.; Byun, M.-O.; Kim, S.T.; Jung, K.-H.; Kim, B.-G. Overexpression of PYL5 in rice enhances drought tolerance, inhibits growth, and modulates gene expression. J. Exp. Bot. 2014, 65, 453–464. [Google Scholar] [CrossRef] [PubMed]
- Miao, C.; Xiao, L.; Hua, K.; Zou, C.; Zhao, Y.; Bressan, R.A.; Zhu, J.-K. Mutations in a subfamily of abscisic acid receptor genes promote rice growth and productivity. Proc. Natl. Acad. Sci. USA 2018, 115, 6058–6063. [Google Scholar] [CrossRef]
- Benanti, J.A.; Cheung, S.K.; Brady, M.C.; Toczyski, D.P. A proteomic screen reveals SCF Grr1 targets that regulate the glycolytic–gluconeogenic switch. Nat. Cell Biol. 2007, 9, 1184. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.; Wang, X.; Li, H.; Gao, Y. Screening E3 substrates using a live phage display library. PLoS ONE 2013, 8, e76622. [Google Scholar] [CrossRef] [PubMed]
- Yen, H.-C.S.; Xu, Q.; Chou, D.M.; Zhao, Z.; Elledge, S.J. Global protein stability profiling in mammalian cells. Science 2008, 322, 918–923. [Google Scholar] [CrossRef]
- Kim, T.Y.; Siesser, P.F.; Rossman, K.L.; Goldfarb, D.; Mackinnon, K.; Yan, F.; Yi, X.; MacCoss, M.J.; Moon, R.T.; Der, C.J. Substrate trapping proteomics reveals targets of the βTrCP2/FBXW11 ubiquitin ligase. Mol. Cell. Biol. 2015, 35, 167–181. [Google Scholar] [CrossRef]
- Lee, K.A.; Hammerle, L.P.; Andrews, P.S.; Stokes, M.P.; Mustelin, T.; Silva, J.C.; Black, R.A.; Doedens, J.R. Ubiquitin ligase substrate identification through quantitative proteomics at both the protein and peptide levels. J. Biol. Chem. 2011, 286, 41530–41538. [Google Scholar] [CrossRef]
- Andrews, P.S.; Schneider, S.; Yang, E.; Michaels, M.; Chen, H.; Tang, J.; Emkey, R. Identification of substrates of SMURF1 ubiquitin ligase activity utilizing protein microarrays. Assay Drug Dev. Technol. 2010, 8, 471–487. [Google Scholar] [CrossRef]
- Maier, A.; Schrader, A.; Kokkelink, L.; Falke, C.; Welter, B.; Iniesto, E.; Rubio, V.; Uhrig, J.F.; Hülskamp, M.; Hoecker, U. Light and the E3 ubiquitin ligase COP 1/SPA control the protein stability of the MYB transcription factors PAP 1 and PAP 2 involved in anthocyanin accumulation in Arabidopsis. Plant J. 2013, 74, 638–651. [Google Scholar] [CrossRef] [PubMed]
- Iconomou, M.; Saunders, D.N. Systematic approaches to identify E3 ligase substrates. Biochem. J. 2016, 473, 4083–4101. [Google Scholar] [CrossRef]
- Li, Y.; Xie, P.; Lu, L.; Wang, J.; Diao, L.; Liu, Z.; Guo, F.; He, Y.; Liu, Y.; Huang, Q. An integrated bioinformatics platform for investigating the human E3 ubiquitin ligase-substrate interaction network. Nat. Commun. 2017, 8, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Sato, Y.; Takehisa, H.; Kamatsuki, K.; Minami, H.; Namiki, N.; Ikawa, H.; Ohyanagi, H.; Sugimoto, K.; Antonio, B.A.; Nagamura, Y. RiceXPro version 3.0: Expanding the informatics resource for rice transcriptome. Nucleic Acids Res. 2013, 41, D1206–D1213. [Google Scholar] [CrossRef] [PubMed]
- Nie, X.-F.; Maclean, K.N.; Kumar, V.; McKay, I.A.; Bustin, S.A. ERF-2, the human homologue of the murine Tis11d early response gene. Gene 1995, 152, 285–286. [Google Scholar] [CrossRef]
- Mello, C.C.; Schubert, C.; Draper, B.; Zhang, W.; Lobel, R.; Priess, J.R. The PIE-1 protein and germline specification in C. elegans embryos. Nature 1996, 382, 710–712. [Google Scholar] [CrossRef]
- Ma, Q.; Wadleigh, D.; Chi, T.; Herschman, H. The Drosophila TIS11 homologue encodes a developmentally controlled gene. Oncogene 1994, 9, 3329–3334. [Google Scholar] [PubMed]
- Thompson, M.J.; Lai, W.S.; Taylor, G.A.; Blackshear, P.J. Cloning and characterization of two yeast genes encoding members of the CCCH class of zinc finger proteins: Zinc finger-mediated impairment of cell growth. Gene 1996, 174, 225–233. [Google Scholar] [CrossRef]
- Wang, D.; Guo, Y.; Wu, C.; Yang, G.; Li, Y.; Zheng, C. Genome-wide analysis of CCCH zinc finger family in Arabidopsis and rice. BMC Genom. 2008, 9, 44. [Google Scholar] [CrossRef]
- Peng, X.; Zhao, Y.; Cao, J.; Zhang, W.; Jiang, H.; Li, X.; Ma, Q.; Zhu, S.; Cheng, B. CCCH-type zinc finger family in maize: Genome-wide identification, classification and expression profiling under abscisic acid and drought treatments. PLoS ONE 2012, 7, e40120. [Google Scholar] [CrossRef]
- Pi, B.; He, X.; Ruan, Y.; Jang, J.-C.; Huang, Y. Genome-wide analysis and stress-responsive expression of CCCH zinc finger family genes in Brassica rapa. BMC Plant Biol. 2018, 18, 1–15. [Google Scholar] [CrossRef]
- Wang, W.; Liu, B.; Xu, M.; Jamil, M.; Wang, G. ABA-induced CCCH tandem zinc finger protein OsC3H47 decreases ABA sensitivity and promotes drought tolerance in Oryza sativa. Biochem. Biophys. Res. Commun. 2015, 464, 33–37. [Google Scholar] [CrossRef]
- Kumar, M.; Choi, J.; An, G.; Kim, S.-R. Ectopic expression of OsSta2 enhances salt stress tolerance in rice. Front. Plant Sci. 2017, 8, 316. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-H.; Choi, H.-S.; Cho, Y.-C.; Kim, S.-R. Cold-responsive regulation of a flower-preferential class III peroxidase gene, OsPOX1, in rice (Oryza sativa L.). J. Plants Biol. 2012, 55, 123–131. [Google Scholar] [CrossRef]
- Lee, S.-C.; Kim, S.-H.; Kim, S.-R. Drought inducible OsDhn1 promoter is activated by OsDREB1A and OsDREB1D. J. Plants Biol. 2013, 56, 115–121. [Google Scholar] [CrossRef]
- Ni, Z.; Kim, E.-D.; Chen, Z.J. Chlorophyll and starch assays. Protoc. Exch. 2009, 10, 1038. [Google Scholar] [CrossRef]
- Chen, D.-H.; Ronald, P. A rapid DNA minipreparation method suitable for AFLP and other PCR applications. Plant. Mol. Biol. Rep. 1999, 17, 53–57. [Google Scholar] [CrossRef]
- Koh, S.; Lee, S.-C.; Kim, M.-K.; Koh, J.H.; Lee, S.; An, G.; Choe, S.; Kim, S.-R. T-DNA tagged knockout mutation of rice OsGSK1, an orthologue of Arabidopsis BIN2, with enhanced tolerance to various abiotic stresses. Plant. Mol. Biol. 2007, 65, 453–466. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Kim, S.-R.; Lee, D.-Y.; Yang, J.-I.; Moon, S.; An, G. Cloning vectors for rice. J. Plants Biol. 2009, 52, 73. [Google Scholar] [CrossRef]
- Yoon, J.; Cho, L.-H.; Antt, H.W.; Koh, H.-J.; An, G. KNOX protein OSH15 induces grain shattering by repressing lignin biosynthesis genes. Plants Physiol. 2017, 174, 312–325. [Google Scholar] [CrossRef]
- Liu, C.; Mao, B.; Ou, S.; Wang, W.; Liu, L.; Wu, Y.; Chu, C.; Wang, X. OsbZIP71, a bZIP transcription factor, confers salinity and drought tolerance in rice. Plants Mol. Biol. 2014, 84, 19–36. [Google Scholar] [CrossRef]
- Schenkelberg, C.D.; Bystroff, C. InteractiveROSETTA: A graphical user interface for the PyRosetta protein modeling suite. Bioinformatics 2015, 31, 4023–4025. [Google Scholar] [CrossRef] [PubMed]
- Leaver-Fay, A.; Tyka, M.; Lewis, S.M.; Lange, O.F.; Thompson, J.; Jacak, R.; Kaufman, K.W.; Renfrew, P.D.; Smith, C.A.; Sheffler, W. ROSETTA3: An object-oriented software suite for the simulation and design of macromolecules. In Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 2011; Volume 487, pp. 545–574. [Google Scholar]
- Basu, S.; Wallner, B. DockQ: A quality measure for protein-protein docking models. PLoS ONE 2016, 11, e0161879. [Google Scholar] [CrossRef]
- Pierce, B.G.; Wiehe, K.; Hwang, H.; Kim, B.-H.; Vreven, T.; Weng, Z. ZDOCK server: Interactive docking prediction of protein–protein complexes and symmetric multimers. Bioinformatics 2014, 30, 1771–1773. [Google Scholar] [CrossRef] [PubMed]
- Schrodinger, L.L.C. The PyMOL Molecular Graphics System, Version 1.8; Schrodinger, L.L.C.: New York, NY, USA, 2015. [Google Scholar]
- Cho, L.-H.; Yoon, J.; Pasriga, R.; An, G. Homodimerization of Ehd1 is required to induce flowering in rice. Plants Physiol. 2016, 170, 2159–2171. [Google Scholar] [CrossRef] [PubMed]
- Ryu, C.H.; Lee, S.; Cho, L.H.; Kim, S.L.; Lee, Y.S.; Choi, S.C.; Jeong, H.J.; Yi, J.; Park, S.J.; Han, C.D. OsMADS50 and OsMADS56 function antagonistically in regulating long day (LD)-dependent flowering in rice. Plants Cell Environ. 2009, 32, 1412–1427. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Choi, J.; Lee, W.; An, G.; Kim, S.-R. OsCBE1, a Substrate Receptor of Cullin4-Based E3 Ubiquitin Ligase, Functions as a Regulator of Abiotic Stress Response and Productivity in Rice. Int. J. Mol. Sci. 2021, 22, 2487. https://doi.org/10.3390/ijms22052487
Choi J, Lee W, An G, Kim S-R. OsCBE1, a Substrate Receptor of Cullin4-Based E3 Ubiquitin Ligase, Functions as a Regulator of Abiotic Stress Response and Productivity in Rice. International Journal of Molecular Sciences. 2021; 22(5):2487. https://doi.org/10.3390/ijms22052487
Chicago/Turabian StyleChoi, Juyoung, Wonkyung Lee, Gynheung An, and Seong-Ryong Kim. 2021. "OsCBE1, a Substrate Receptor of Cullin4-Based E3 Ubiquitin Ligase, Functions as a Regulator of Abiotic Stress Response and Productivity in Rice" International Journal of Molecular Sciences 22, no. 5: 2487. https://doi.org/10.3390/ijms22052487
APA StyleChoi, J., Lee, W., An, G., & Kim, S.-R. (2021). OsCBE1, a Substrate Receptor of Cullin4-Based E3 Ubiquitin Ligase, Functions as a Regulator of Abiotic Stress Response and Productivity in Rice. International Journal of Molecular Sciences, 22(5), 2487. https://doi.org/10.3390/ijms22052487





