NGS-Based Application for Routine Non-Invasive Pre-Implantation Genetic Assessment in IVF
Abstract
1. Introduction
2. Results
2.1. Sample Collection
2.2. Next-Generation Sequencing and Primary Data Analysis
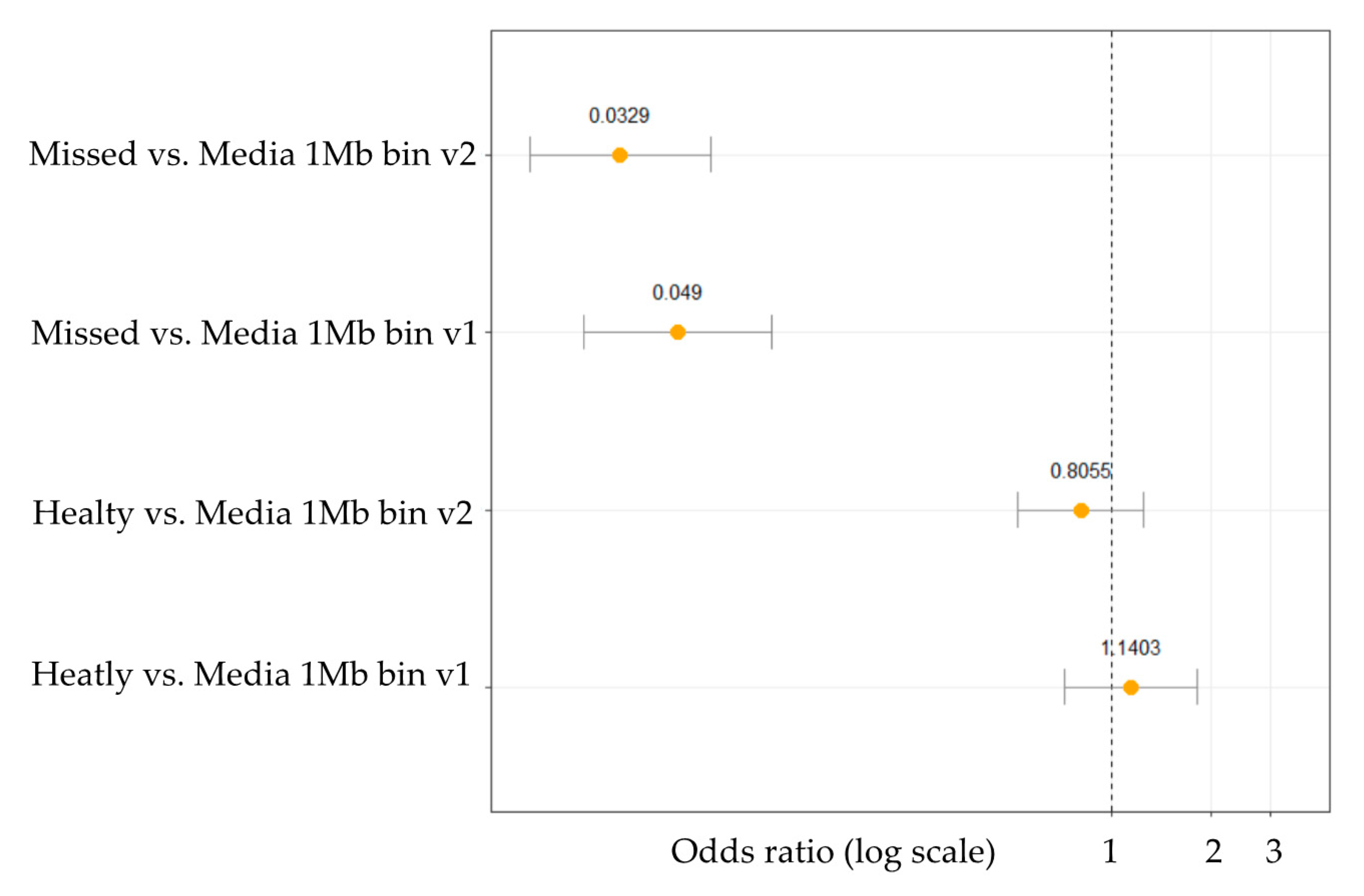
2.3. Identified CNVs and Statistical Testing
3. Discussion
4. Materials and Methods
4.1. Study Design and Workflow
4.2. Step 1: IVF Procedure and Sample Collection
4.3. Step 2: Whole-Genome Amplification
4.4. Step 3: Next-Generation Sequencing
4.5. Step 4: Bioinformatics Analysis
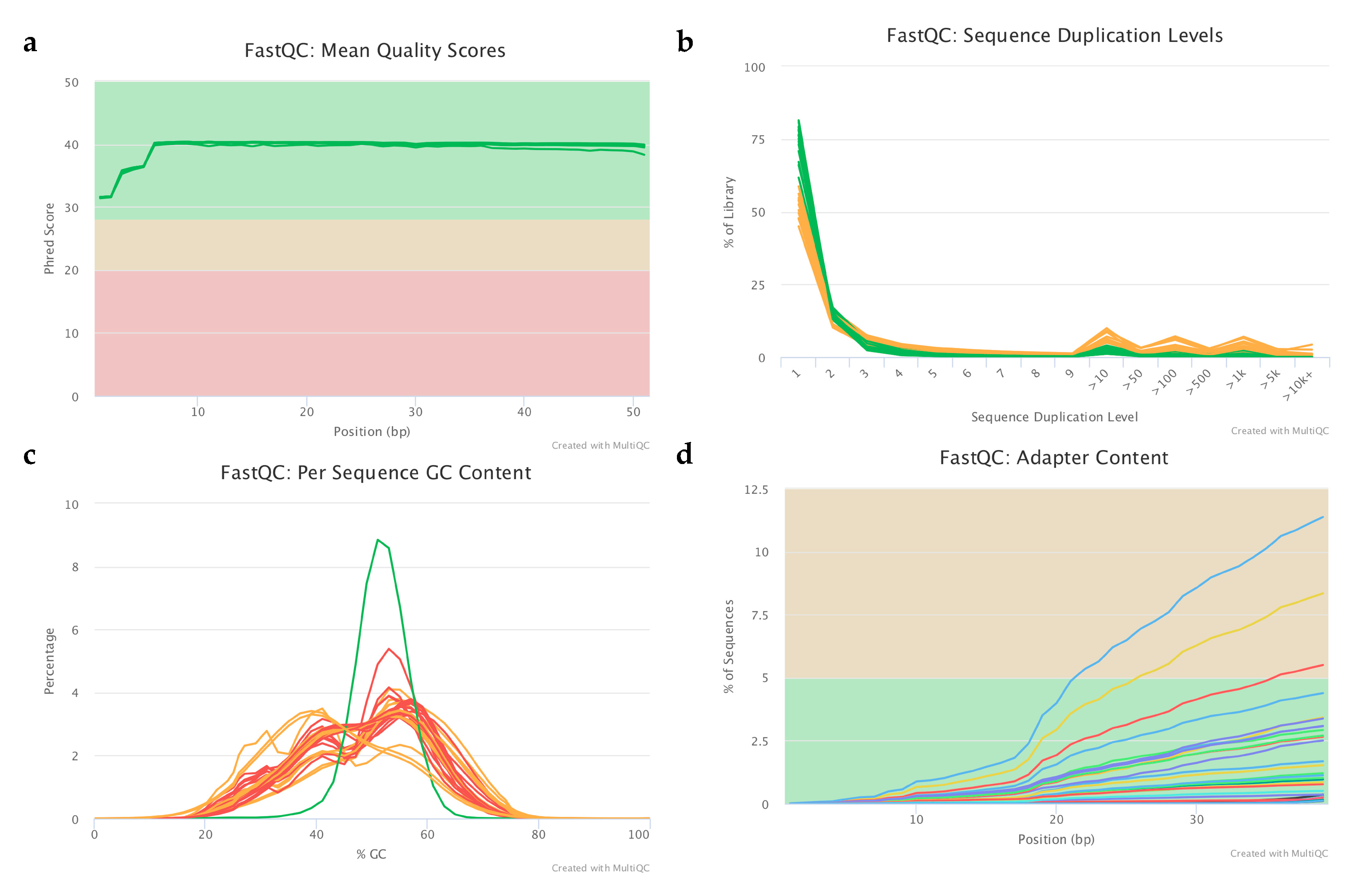
4.5.1. Raw Data Quality Control and Filtering
4.5.2. Sequence Alignment and Mapping Quality
4.5.3. CNV Identification
4.5.4. Statistics for CNV Analysis
4.5.5. Functional Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Alpha Scientists in Reproductive Medicine and ESHRE Special Interest Group of Embryology. Istanbul consensus workshop on embryo assessment: Proceedings of an expert meeting. Reprod. Biomed. Online 2011, 22, 632–646. [Google Scholar] [CrossRef]
- Paternot, G.; Debrock, S.; De Neubourg, D.; D’Hooghe, T.M.; Spiessens, C. Semi-automated morphometric analysis of human embryos can reveal correlations between total embryo volume and clinical pregnancy. Hum. Reprod. 2013, 28, 627–633. [Google Scholar] [CrossRef] [PubMed]
- Gardner, D.K.; Meseguer, M.; Rubio, C.; Treff, N.R. Diagnosis of human preimplantation embryo viability. Hum. Reprod. Update 2015, 21, 727–747. [Google Scholar] [CrossRef] [PubMed]
- Katz-Jaffe, M.G.; Gardner, D.K.; Schoolcraft, W.B. Proteomic analysis of individual human embryos to identify novel biomarkers of development and viability. Fertil. Steril. 2006, 85, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Mains, L.M.; Christenson, L.; Yang, B.; Sparks, A.E.; Mathur, S.; Van Voorhis, B.J. Identification of apolipoprotein AI in the human embryonic secretome. Fertil. Steril. 2011, 2, 422–427. [Google Scholar] [CrossRef] [PubMed]
- Butler, S.A.; Luttoo, J.; Freire, M.O.; Abban, T.K.; Borrelli, P.T.; Iles, R.K. Human chorionic gonadotropin (hCG) in the secretome of cultured embryos: Hyperglycosylated hCG and hCG free beta subunit are potential markers for infertility management and treatment. Reprod. Sci. 2013, 9, 1038–1045. [Google Scholar] [CrossRef] [PubMed]
- Montskó, G.; Zrínyi, Z.; Janáky, T.; Szabó, Z.; Várnagy, Á.; Kovács, G.L.; Bódis, J. Noninvasive embryo viability assessment by quantitation of human haptoglobin alpha-1 fragment in the in vitro fertilization culture medium: An additional tool to increase success rate. Fertil. Steril. 2015, 103, 687–693. [Google Scholar] [CrossRef]
- Montskó, G.; Gödöny, K.; Herczeg, R.; Várnagy, Á.; Bódis, J.; Kovács, G.L. Alpha-1 chain of human haptoglobin as viability marker of in vitro fertilized human embryos: Information beyond morphology. Syst. Biol. Reprod. Med. 2019, 65, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Devreker, F.; Hardy, K.; Van den Bergh, M.; Winston, J.; Biramane, J.; Englert, Y. Noninvasive assessment of glucose and pyruvate uptake by human embryos after intracytoplasmic sperm injection and during the formation of pronuclei. Fertil. Steril. 2000, 73, 947–954. [Google Scholar] [CrossRef]
- Gardner, D.K.; Lane, M.; Stevens, J.; Schoolcraft, W.B. Noninvasive assessment of human embryo nutrient consumption as a measure of developmental potential. Fertil. Steril. 2001, 76, 1175–1180. [Google Scholar] [CrossRef]
- Hernandez-Vargas, P.; Munoz, M.; Domínguez, F. Identifying biomarkers for predicting successful embryo implantation: Applying single to multi-OMICSs to improve reproductive outcomes. Hum. Reprod. Update 2020, 26, 264–301. [Google Scholar] [CrossRef] [PubMed]
- Cortezzi, S.S.; Cabral, E.C.; Trevisan, M.G.; Ferreira, C.R.; Setti, A.S.; Braga, D.P.; Figueiredo, R.C.; van Jaconelli, A.; Eberlin, M.N.; Van Borges, E. Prediction of embryo implantation potential by mass spectrometry fingerprinting of the culture medium. Reproduction 2013, 5, 453–462. [Google Scholar] [CrossRef] [PubMed]
- Wells, D. Embryo aneuploidy and the role of morphological and genetic screening. Reprod. Biomed. Online 2010, 21, 274–277. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Bogale, B.; Tang, Y.; Lu, S.; Xie, X.S.; Racowsky, C. Noninvasive preimplantation genetic testing for aneuploidy in spent medium may be more reliable than trophectoderm biopsy. Proc. Natl. Acad. Sci. USA 2019, 116, 14105–14112. [Google Scholar] [CrossRef] [PubMed]
- Shitara, A.; Takahasi, K.; Goto, M.; Takahashi, H.; Iwasawa, T.; Onodera, Y.; Makino, M.; Miura, H.; Shirasawa, H.; Sato, W.; et al. Cell-free DNA in spent culture medium effectively reflects the chromosomal status of embryos following culturing beyond implantation compared to trophectoderm biopsy. PLoS ONE 2021, 16, e0246438. [Google Scholar] [CrossRef]
- Rubio, C.; Navarro-Sánchez, L.; García-Pascual, C.M.; Ocali, O.; Cimadomo, D.; Venier, W.; Barroso, G.; Kopcow, L.; Bahçeci, M.; Kulmann, M.I.R.; et al. Multicenter prospective study of concordance between embryonic cell-free DNA and trophectoderm biopsies from 1301 human blastocysts. Am. J. Obstet. Gynecol 2020, 223, 751. [Google Scholar] [CrossRef]
- Vera-Rodriguez, M.; Diez-Juan, A.; Jimenez-Almazan, J.; Martinez, S.; Navarro, R.; Peinado, V.; Simon, C. Origin and composition of cell-free DNA in spent medium from human embryo culture during preimplantation development. Hum. Reprod. 2018, 33, 745–756. [Google Scholar] [CrossRef]
- Handyside, A.H. Noninvasive preimplantation genetic testing: Dream or reality? Fertil. Steril. 2016, 106, 1324–1325. [Google Scholar] [CrossRef]
- Kuznyetsov, V.; Madjunkova, S.; Antes, R.; Abramov, R.; Motamedi, G.; Ibarrientos, Z.; Librach, C. Evaluation of a novel non-invasive preimplantation genetic screening approach. PLoS ONE 2018, 13, e0197262. [Google Scholar] [CrossRef]
- Fang, R.; Yang, W.; Zhao, X.; Xiong, F.; Guo, C.; Xiao, J.; Cai, L.Y. Chromosome screening using culture medium of embryos fertilised in vitro: A pilot clinical study. J. Transl. Med. 2019, 17, 73. [Google Scholar] [CrossRef] [PubMed]
- Stigliani, S.; Anserini, P.; Venturini, P.L.; Scaruffi, P. Mitochondrial DNA content in embryo culture medium is significantly associated with human embryo fragmentation. Hum. Reprod. 2013, 28, 2652–2660. [Google Scholar] [CrossRef]
- Farra, C.; Choucair, F.; Awwad, J. Non-invasive pre-implantation genetic testing of human embryos: An emerging concept. Hum. Reprod. 2018, 33, 2162–2167. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Ding, C.; Shen, X.; Wang, J.; Li, R.; Cai, B.; Zhou, C. Medium-based noninvasive preimplantation genetic diagnosis for human α-thalassemias-SEA. Medicine 2015, 94, e669. [Google Scholar] [CrossRef] [PubMed]
- Hammond, E.R.; McGillivray, B.C.; Wicker, S.M.; Peek, J.C.; Shelling, A.N.; Stone, P.; Cree, L.M. Characterizing nuclear and mitochondrial DNA in spent embryo culture media: Genetic contamination identified. Fertil. Steril. 2017, 107, 220–228. [Google Scholar] [CrossRef] [PubMed]
- Hammond, E.R.; Shelling, A.N.; Cree, L.M. Nuclear and mitochondrial DNA in blastocoele fluid and embryo culture medium: Evidence and potential clinical use. Hum. Reprod. 2016, 31, 1653–1661. [Google Scholar] [CrossRef]
- Palini, S.; Galluzzi, L.; De Stefani, S.; Bianchi, M.; Wells, D.; Magnani, M.; Bulletti, C. Genomic DNA in human blastocoele fluid. Reprod. Biomed. Online 2013, 26, 603–610. [Google Scholar] [CrossRef] [PubMed]
- Gianaroli, L.; Magli, M.C.; Pomante, A.; Crivello, A.M.; Cafueri, G.; Valerio, M.; Ferraretti, A.P. Blastocentesis: A source of DNA for preimplantation genetic testing. Results from a pilot study. Fertil. Steril. 2014, 102, 1692–1699. [Google Scholar] [CrossRef]
- Yeung, Q.S.Y.; Zhang, Y.X.; Chung, J.P.W.; Lui, W.T.; Kwok, Y.K.Y.; Gui, B.; Choy, K.W. A prospective study of non-invasive preimplantation genetic testing for aneuploidies (NiPGT-A) using next-generation sequencing (NGS) on spent culture media (SCM). J. Assist. Reprod. Genet. 2019, 36, 1609–1621. [Google Scholar] [CrossRef] [PubMed]
- Ho, J.R.; Arrach, N.; Rhodes-Long, K.; Ahmady, A.; Ingles, S.; Chung, K.; McGinnis, L.K. Pushing the limits of detection: Investigation of cell-free DNA for aneuploidy screening in embryos. Fertil. Steril. 2018, 110, 467–475. [Google Scholar] [CrossRef]
- Xu, J.; Fang, R.; Chen, L.; Chen, D.; Xiao, J.P.; Yang, W.; Lu, S. Noninvasive chromosome screening of human embryos by genome sequencing of embryo culture medium for in vitro fertilization. Proc. Natl. Acad. Sci. USA 2016, 113, 11907–11912. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 15 September 2020).
- Marcel, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Andrews, S. Trim Galore: A Wrapper Tool Around Cutadapt and FastQC to Consistently Apply QUALITY and Adapter Trimming to FastQ Files, with Some Extra Functionality for MspI-Digested RRBS-Type (Reduced Representation Bisufite-Seq) Libraries. 2012. Available online: https://www.bioinformatics.babraham.ac.uk/projects/trim_galore (accessed on 5 September 2020).
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Durbin, R. The sequence alignment/map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Okonechnikov, K.; Conesa, A.; García-Alcalde, F. Qualimap 2: Advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 2016, 32, 292–294. [Google Scholar] [CrossRef] [PubMed]
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: Summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef]
- Klambauer, G.; Schwarzbauer, K.; Mayr, A.; Clevert, D.A.; Mitterecker, A.; Bodenhofer, U.; Hochreiter, S. cn.MOPS: Mixture of Poissons for discovering copy number variations in next-generation sequencing data with a low false discovery rate. Nucleic Acids Res. 2012, 40, e69. [Google Scholar] [CrossRef] [PubMed]
- Stevenson, M.; Nunes, T.; Sanchez, J.; Thornton, R.; Reiczigel, J.; Robison-Cox, J.; Sebastiani, P.; EpiR: An R Package for the Analysis of Epidemiological Data. R Pakage Version 1.0-10. 2013. Available online: https://CRAN.R-project.org/package=epiR (accessed on 15 September 2020).
- Randle, A.M.V.; Zhuo, J.C. ggplot2: Elegant graphics for data analysis (2nd ed.). Meas. Interdiscip. Res. Perspect. 2019, 17, 160–167. [Google Scholar] [CrossRef]



| Sample Name | Group | Avg. GC | ≥1X | ≥5X | Median Coverage | % Aligned |
|---|---|---|---|---|---|---|
| G1_plus_HSA1 | c | 49% | 0.9% | 0.3% | 0.0X | 40.9% |
| G1_plus_HSA2 | c | 49% | 0.5% | 0.2% | 0.0X | 39.7% |
| G1_plus_HSA4 | c | 47% | 0.8% | 0.2% | 0.0X | 35.1% |
| G1_plus_HSA5 | c | 48% | 0.7% | 0.3% | 0.0X | 36.2% |
| G1_plus_HSA6 | c | 48% | 0.8% | 0.3% | 0.0X | 44.2% |
| 7567_1A | 0 | 50% | 10.6% | 0.4% | 0.0X | 91.9% |
| 7567_1B | 0 | 48% | 4.0% | 1.1% | 0.0X | 76.0% |
| 7010_1A | 0 | 49% | 1.4% | 0.4% | 0.0X | 41.7% |
| 7010_1B | 0 | 50% | 12.6% | 0.4% | 0.0X | 96.2% |
| 7301_1A | 0 | 50% | 11.9% | 0.4% | 0.0X | 95.2% |
| 7301_1B | 0 | 50% | 5.0% | 1.0% | 0.0X | 84.7% |
| 7316_1A | 0 | 49% | 6.1% | 1.0% | 0.0X | 86.4% |
| 7316_1B | 0 | 50% | 9.9% | 0.3% | 0.0X | 95.7% |
| 7370_1B | 0 | 50% | 7.5% | 0.9% | 0.0X | 87.4% |
| 6341_4B | 1 | 49% | 1.5% | 0.4% | 0.0X | 40.4% |
| 6341_4C | 1 | 50% | 2.4% | 0.5% | 0.0X | 47.9% |
| 7793_1A | 1 | 49% | 5.7% | 1.3% | 0.0X | 83.1% |
| 7793_1B | 1 | 50% | 7.8% | 1.1% | 0.0X | 87.8% |
| 7938_1A | 1 | 50% | 8.1% | 1.1% | 0.0X | 88.8% |
| 7938_1C | 1 | 49% | 9.9% | 1.1% | 0.0X | 92.2% |
| A7Down | 2 | 44% | 17.8% | 0.0% | 0.0X | 98.0% |
| A8Down | 2 | 44% | 21.1% | 0.0% | 0.0X | 98.0% |
| Chromosomal Location | Type of Alteration | Function |
|---|---|---|
| 2q35 | deletion | XRCC5 gene inactivation- defect in DNA repair function |
| 2q37 | 2.3-2.4 mb deletion | IGFBP2 inactivation |
| 3p25.3-p25.1 | deletion | miR-885 inactivation, impaired differentiation |
| 4p16.3-p16.1 | duplication | CNV identified with chromosomal microarray in individuals with developmental disabilities or congenital anomalies (ISCA) |
| 8q24.3 | duplication | myc proto-oncogene gene desert in GWAS studies |
| 9p12-p11.2 | deletion | ANKRD20A3 gene inactivation syndromic hydrocephalus due to diffuse hyperplasia of choroid plexus, glioma |
| 10q22.1 | duplication | COLl13A1 frameshift with pathogenic interpretation (ClinVar) |
| 11q23.1-23.3 | duplication | Beckwith-Wiedemann syndrome |
| 14q31.1-q31.3 | deletion | autosomal dominant disorder (HPPD)involving hypertelorism and deafness |
| 14q32.2-q32.33 | deletion | FOXG1 inactivation, impaired development and structural brain abnormalities |
| 15q13.3 | deletion | MTMR10, FAN1 frameshift associated with karyomegalic interstitial nephritis |
| 16q23.3-24.3 | duplication | APRT, FOXC2 indel, adenine phosphoribosyl transferase deficiency, disichtiasis lymphoedema syndrome |
| 17q22-p23.2 | deletion | Ateleiotic dwarfism, isolated growth hormone deficiency |
| 20p12.2-p12.1 | deletion | JAG1 related Alagille syndrome |
| 20q13.31-q13.33 | duplication | PKC1, phosphoenolpyruvate carboxikinase deficiency |
| 21q22.3 | duplication | RIPK4, PCNT popliteal pterygum syndrome, lethal type |
| 21p13-p11.2 | deletion | short arm loss monosomy |
| 22q13.2-13.31 | duplication | SCO2 cardioencephalomyopathy due to cytochrome c oxidase deficiency, fatal |
| Performance | a (n = 116) | b (n = 48) | c (n = 42) | d (n = 71) | e (n = 41) | f (n = 20) | g (n = 1301) |
|---|---|---|---|---|---|---|---|
| sensitivity | 81.6% | 100% | 88.2% | 46.5% | 80% | 100% | 76.5–91.3% |
| specificity | 48.3% | 80% | 84% | 75% | 61% | 87.5% | 64.7–93.3% |
| positive predictive value | 82. 6% | 91.7% | 79% | 90.9% | 47% | 88.9% | 65.1–92% |
| negative predictive value | 46.7% | 100% | 91.3% | 20.7% | 88% | 100% | 65.2–94% |
| concordance for embryo ploidy | - | 93.8% | - | - | 56.3% | 100% | 70.1–77.3% |
| concordance for chromosome CN | - | 83.3% | - | - | - | 56.3% | 67.7% |
| concordance for TE result | 73.3% | - | 83.4% | 34.4% | - | 66.7% | 72.5–86.3% |
| Healthy Neonate (Group 1) | Miscarriage (Group 0) | |
|---|---|---|
| Number of embryonic culture media samples sequenced | 20 | 20 |
| ICCS Scoring parameters of D3 embryos | group 1 | group 0 |
| average blastomere number | 8.2 | 8.6 |
| fragmentation by volume | <10% | <10% |
| blastomere symmetry | full | full |
| Clinical characteristics | group 1 | group 0 |
| female average age | 35.18 | 34.74 |
| cause of infertility -tubal factor | 27.27 | 22.5 |
| cause of infertility male factor | 45.45 | 42.5 |
| cause of infertility -other | 27.27 | 25 |
| basal FSH cc (IU/µL) | 7.63 | 7.2 |
| previous miscarriage | 0 | 0 |
| oocyte collected | 9.3 | 8.6 |
| available embryos for culture | 2.5 | 2.5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gombos, K.; Gálik, B.; Kalács, K.I.; Gödöny, K.; Várnagy, Á.; Alpár, D.; Bódis, J.; Gyenesei, A.; Kovács, G.L. NGS-Based Application for Routine Non-Invasive Pre-Implantation Genetic Assessment in IVF. Int. J. Mol. Sci. 2021, 22, 2443. https://doi.org/10.3390/ijms22052443
Gombos K, Gálik B, Kalács KI, Gödöny K, Várnagy Á, Alpár D, Bódis J, Gyenesei A, Kovács GL. NGS-Based Application for Routine Non-Invasive Pre-Implantation Genetic Assessment in IVF. International Journal of Molecular Sciences. 2021; 22(5):2443. https://doi.org/10.3390/ijms22052443
Chicago/Turabian StyleGombos, Katalin, Bence Gálik, Krisztina Ildikó Kalács, Krisztina Gödöny, Ákos Várnagy, Donát Alpár, József Bódis, Attila Gyenesei, and Gábor L. Kovács. 2021. "NGS-Based Application for Routine Non-Invasive Pre-Implantation Genetic Assessment in IVF" International Journal of Molecular Sciences 22, no. 5: 2443. https://doi.org/10.3390/ijms22052443
APA StyleGombos, K., Gálik, B., Kalács, K. I., Gödöny, K., Várnagy, Á., Alpár, D., Bódis, J., Gyenesei, A., & Kovács, G. L. (2021). NGS-Based Application for Routine Non-Invasive Pre-Implantation Genetic Assessment in IVF. International Journal of Molecular Sciences, 22(5), 2443. https://doi.org/10.3390/ijms22052443









