Integrated Metabolome and Transcriptome Analysis Unveils Novel Pathway Involved in the Formation of Yellow Peel in Cucumber
Abstract
1. Introduction
2. Results
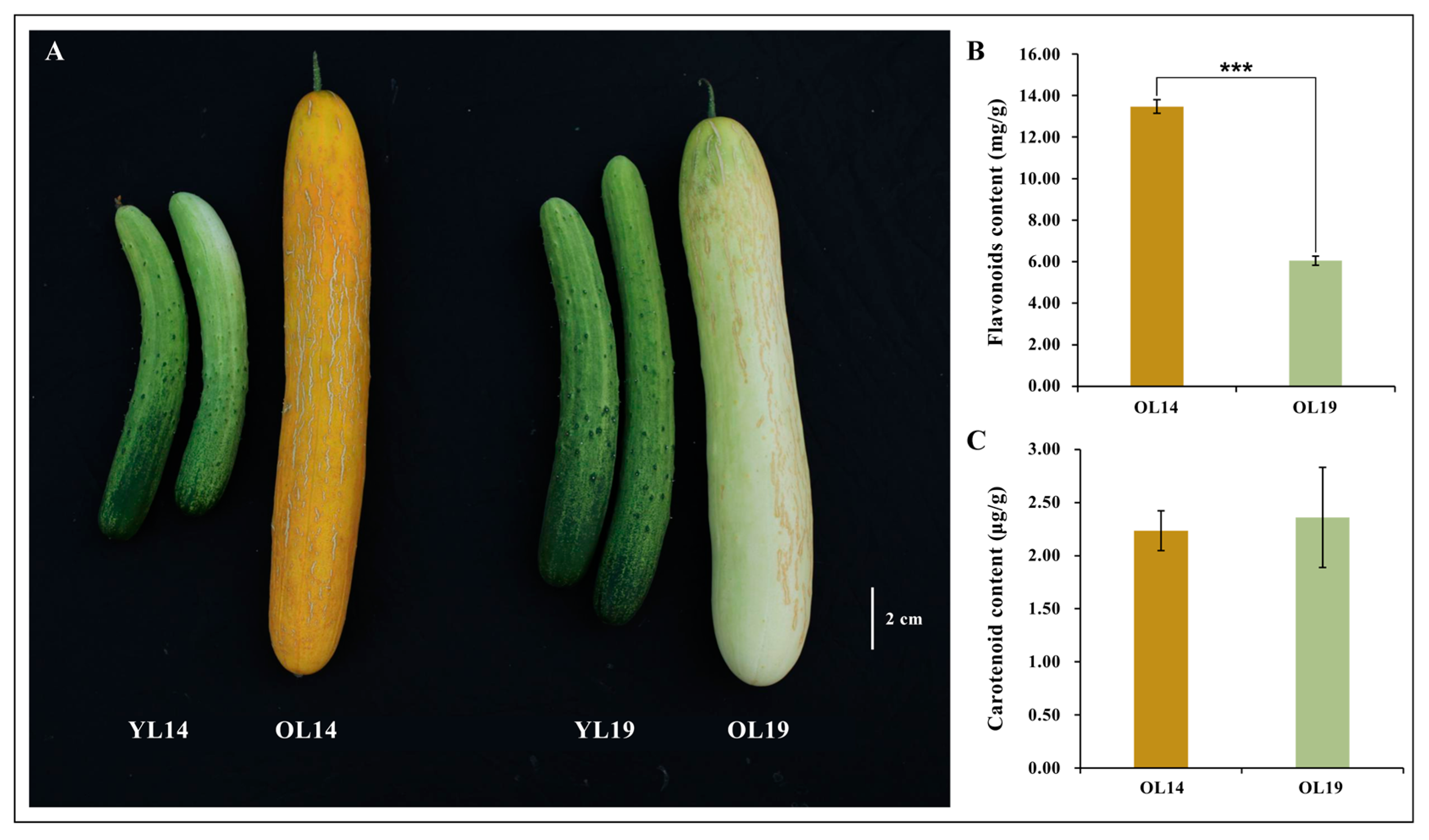
2.1. Phenotype of Two Near-Isogenic Cucumber Varieties
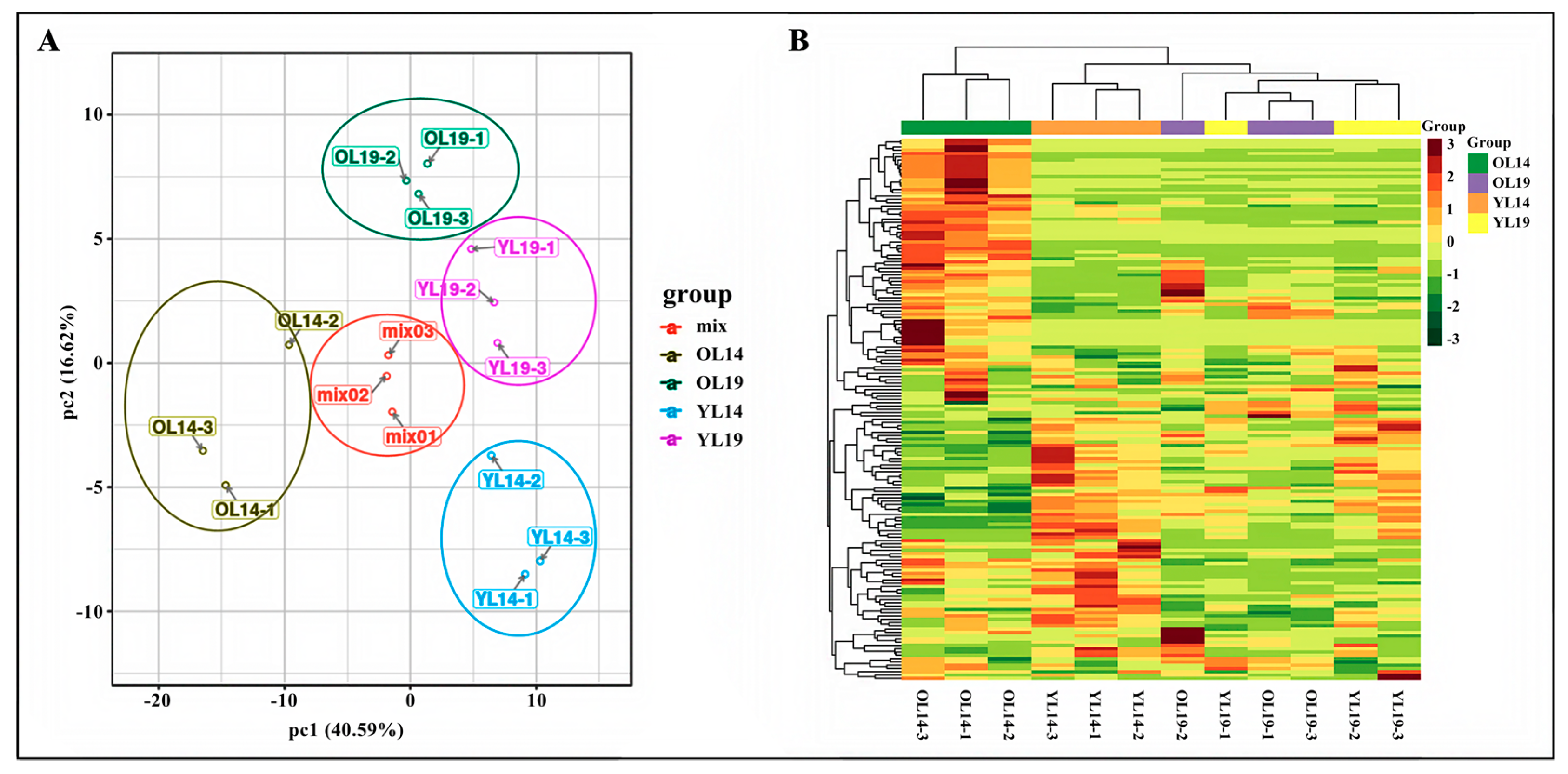
2.2. Differential Accumulation of Flavonoid Metabolites between L14 and L19
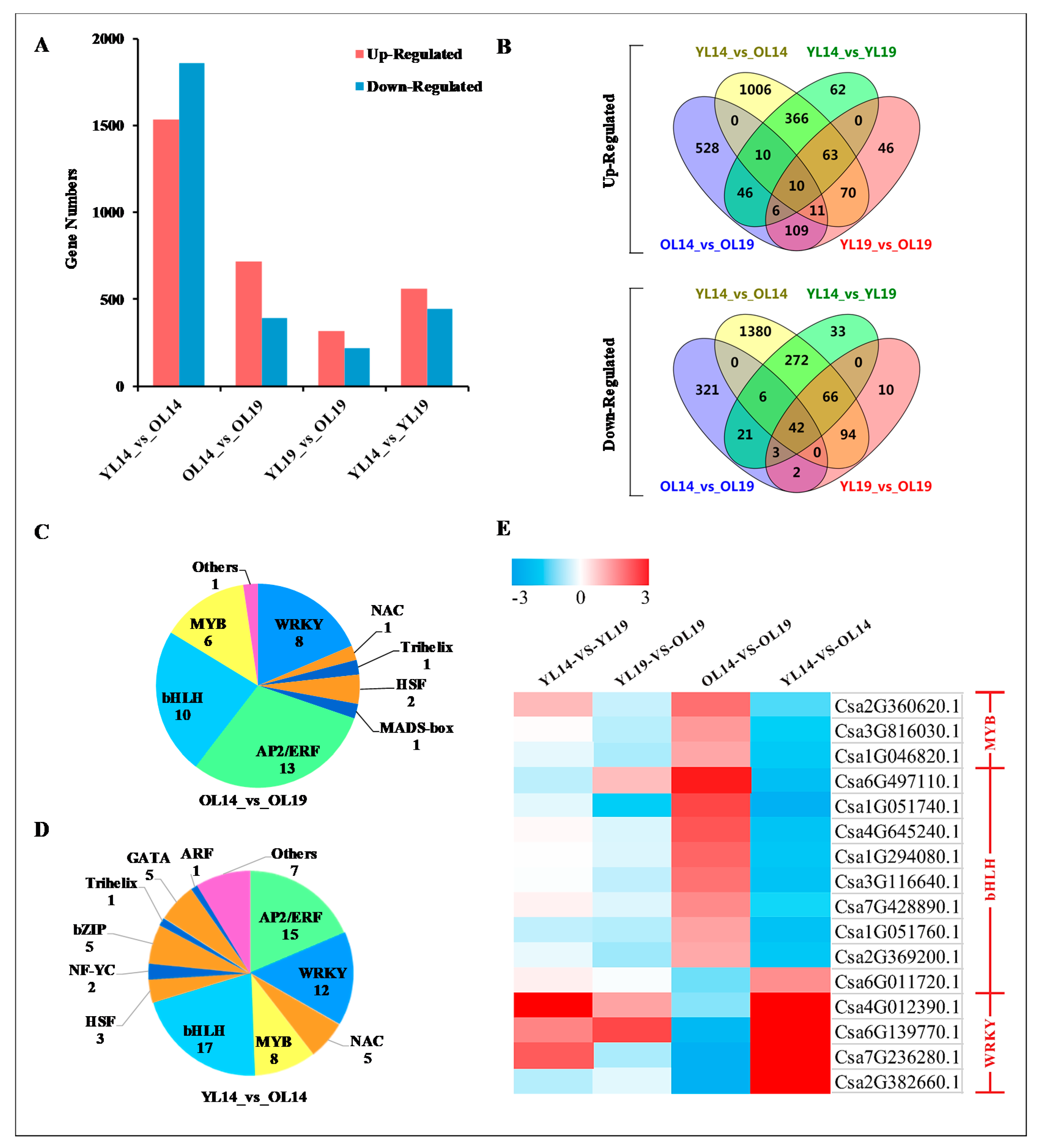
2.3. L14 and L19 Transcriptome Analysis
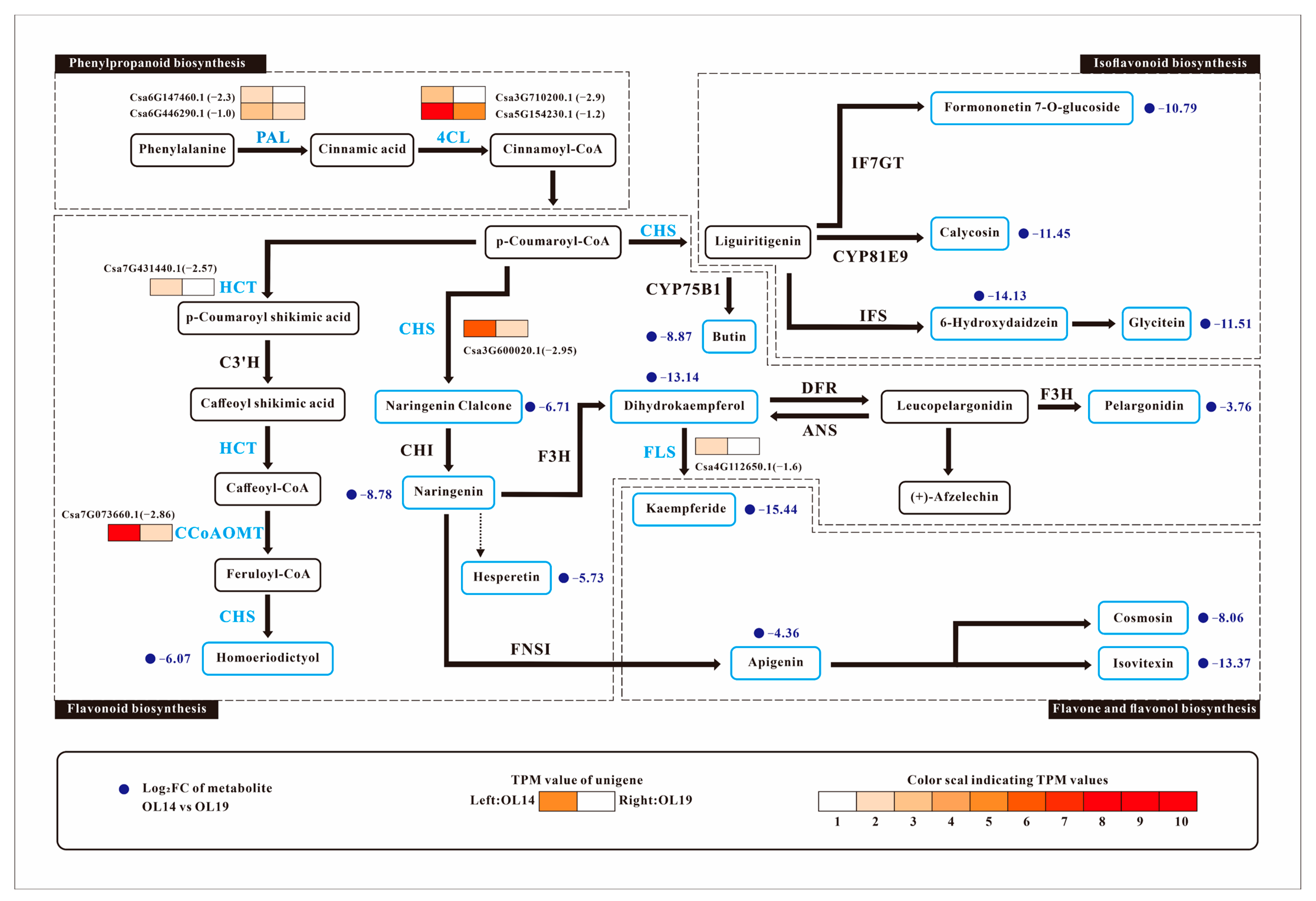
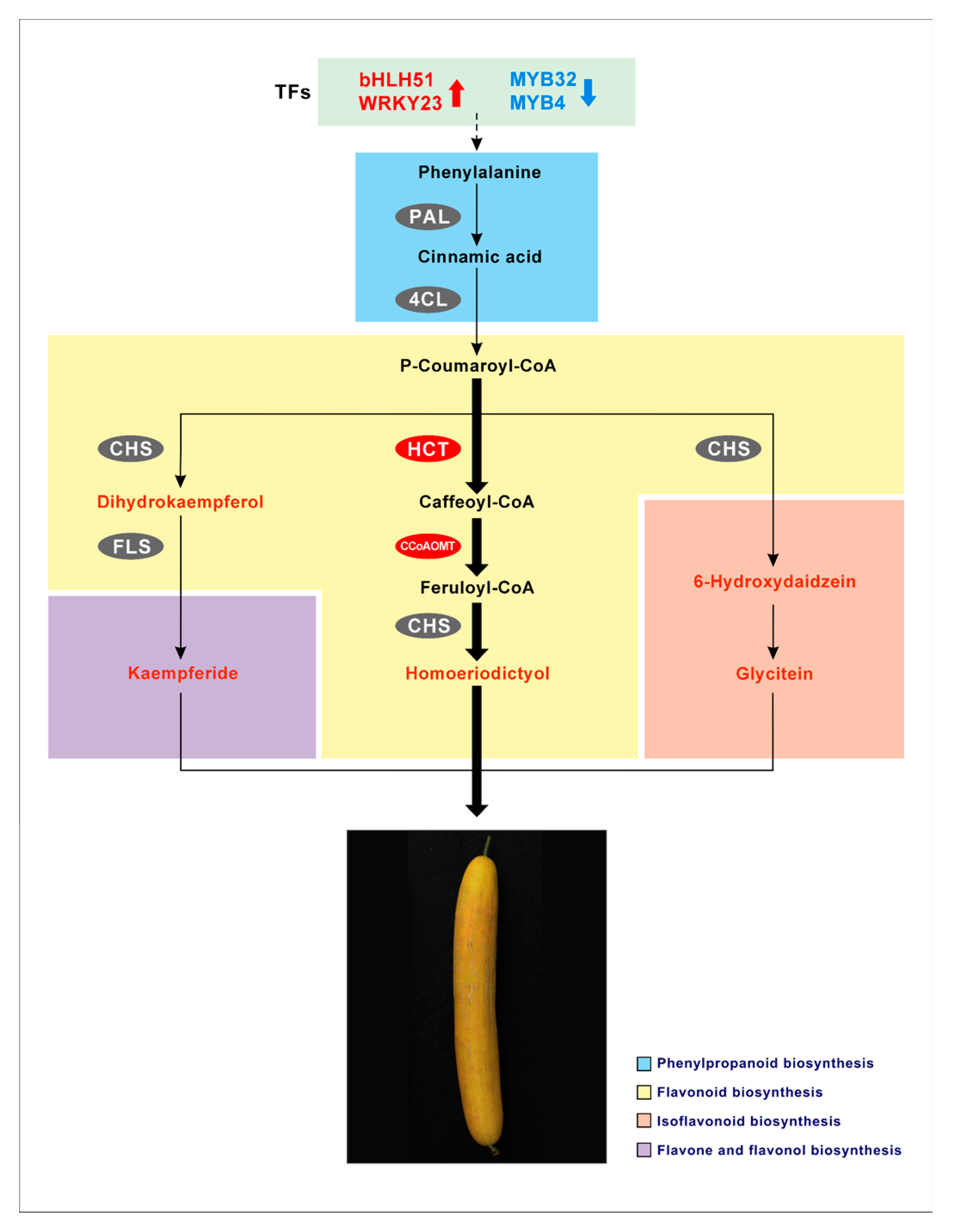
2.4. Association Analysis of Genes and Metabolites Related to Yellowing of Cucumber Peel
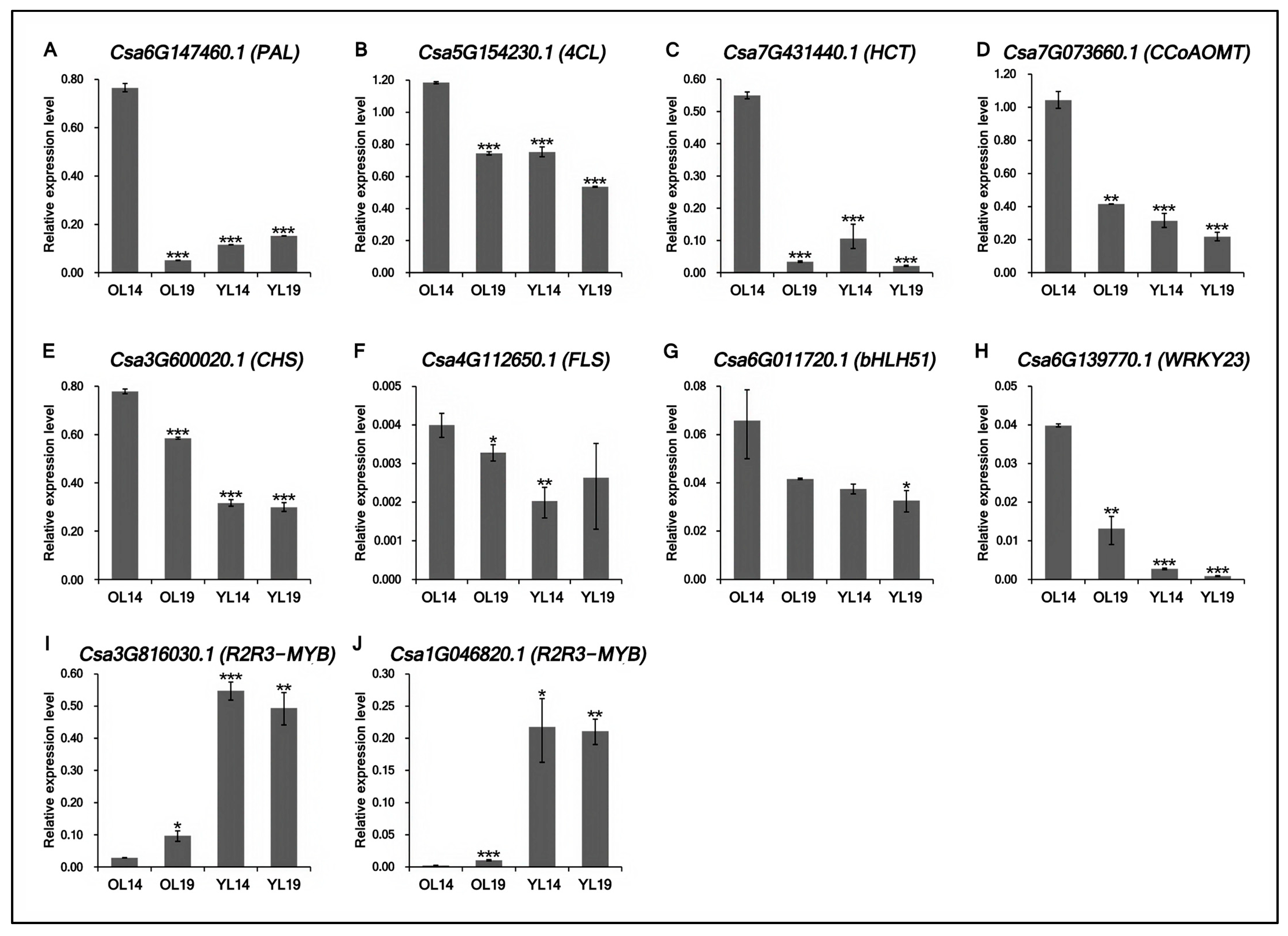
2.5. Validation of Transcriptomic Data Accuracy by qRT-PCR
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Determination of Total Flavonoids and Carotenoids
4.3. Sample Extraction and LC–MS/MS Analysis
4.4. RNA Extraction, cDNA Library Preparation and RNA Sequencing
4.5. Analysis and Functional Annotation of Differentially Expressed Genes
4.6. qRT-PCR Verification of Gene Expression
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Miao, L.; Di, Q.H.; Sun, T.S.; Li, Y.S.; Duan, Y.; Wang, J.; Yan, Y.; He, C.X.; Wang, C.L.; Yu, X.C. Integrated metabolome and transcriptome analysis provide insights into the effects of grafting on fruit flavor of cucumber with different rootstocks. Int. J. Mol. Sci. 2019, 20, 3592. [Google Scholar] [CrossRef]
- Sebastian, P.; Schaefer, H.; Telford, I.R.H.; Renner, S.S. Cucumber (Cucumis sativus) and melon (C. melo) have numerous wild relatives in Asia and Australia, and the sister species of melon is from Australia. Proc. Natl. Acad. Sci. USA 2010, 107, 14269–14273. [Google Scholar] [CrossRef]
- Yundaeng, C.; Somta, P.; Tangphatsornruang, S.; Chankaew, S.; Srinives, P. A single base substitution in BADH/AMADH is responsible for fragrance in cucumber (Cucumis sativus L.), and development of SNAP markers for the fragrance. Appl. Genet. 2015, 128, 1881–1892. [Google Scholar] [CrossRef]
- Wang, M.; Jiang, B.; Peng, Q.W.; Liu, W.R.; He, X.M.; Liang, Z.J.; Lin, Y.E. Transcriptome analyses in different cucumber cultivars provide novel insights into drought stress responses. Int. J. Mol. Sci. 2018, 19, 2067. [Google Scholar] [CrossRef]
- Che, G.; Zhang, X.L. Molecular basis of cucumber fruit domestication. Curr. Opin. Plant. Biol. 2019, 47, 38–46. [Google Scholar] [CrossRef]
- Li, P.M.; Ma, F.W.; Cheng, L.L. Primary and secondary metabolism in the sun-exposed peel and the shaded peel of apple fruit. Physiol. Plant. 2013, 148, 9–24. [Google Scholar] [CrossRef]
- Wang, Z.R.; Cui, Y.Y.; Vainstein, A.; Chen, S.W.; Ma, H.Q. Regulation of Fig (Ficus carica L.) Fruit Color: Metabolomic and Transcriptomic Analyses of the Flavonoid Biosynthetic Pathway. Front. Plant. Sci. 2017, 8, 1990. [Google Scholar] [CrossRef]
- Hao, N.; Du, Y.L.; Li, H.Y.; Wang, C.; Wang, C.; Gong, S.Y.; Zhou, S.G.; Wu, T. CsMYB36 is involved in the formation of yellow green peel in cucumber (Cucumis sativus L.). Appl. Genet. 2018, 131, 1659–1669. [Google Scholar] [CrossRef]
- Hutchins, A.E. Inheritance in the cucumber. J. Agric. Res. 1940, 60, 117–128. [Google Scholar]
- Zhou, Q.; Wang, S.H.; Hu, B.W.; Chen, H.M.; Zhang, Z.H.; Huang, S.W. An ACCUMULATION AND REPLICATION OF CHLOROPLASTS 5 gene mutation confers light green peel in cucumber. J. Integr. Plant. Biol. 2015, 57, 936–942. [Google Scholar] [CrossRef]
- Lun, Y.Y.; Wang, X.; Zhang, C.Z.; Yang, L.; Gao, D.L.; Chen, H.M.; Huang, S.W. A CsYcf54 variant conferring light green coloration in cucumber. Euphytica 2016, 208, 509–517. [Google Scholar] [CrossRef]
- Liu, H.Q.; Jiao, J.Q.; Liang, X.J.; Liu, J.; Meng, H.W.; Chen, S.X.; Li, Y.H.; Cheng, Z.H. Map-based cloning, identification and characterization of the w gene controlling white immature fruit color in cucumber (Cucumis sativus L.). Appl. Genet. 2016, 129, 1247–1256. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.H.; Wen, C.L.; Weng, Y.Q. Fine mapping of the pleiotropic locus B for black spine and orange mature fruit color in cucumber identifies a 50 kb region containing a R2R3-MYB transcription factor. Appl. Genet. 2013, 126, 2187–2196. [Google Scholar] [CrossRef]
- Cho, K.; Cho, K.S.; Sohn, H.B.; Ha, I.J.; Hong, S.Y.; Lee, H.; Kim, Y.M.; Nam, M.H. Network analysis of the metabolome and transcriptome reveals novel regulation of potato pigmentation. J. Exp. Bot. 2016, 67, 1519–1533. [Google Scholar] [CrossRef]
- Gu, Z.Y.; Men, S.Q.; Zhu, J.; Hao, Q.; Tong, N.N.; Liu, Z.A.; Zhang, H.C.; Shu, Q.Y.; Wang, L.S. Chalcone synthase is ubiquitinated and degraded via interactions with a RING-H2 protein in petals of Paeonia ‘He Xie’. J. Exp. Bot. 2019, 70, 4749–4762. [Google Scholar] [CrossRef]
- Nabavi, S.M.; Šamec, D.; Tomczyk, M.; Milella, L.; Russo, D.; Habtemariam, S.; Suntar, I.; Rastrelli, L.; Daglia, M.; Xiao, J.B.; et al. Flavonoid biosynthetic pathways in plants: Versatile targets for metabolic engineering. Biotechnol. Adv. 2020, 38, 107316. [Google Scholar] [CrossRef]
- Xu, X.W.; Pan, J.W.; He, M.; Tian, H.L.; Qi, X.H.; Xu, Q.; Chen, X.H. Transcriptome profiling reveals key genes related to astringency during cucumber fruit development. 3 Biotech 2019, 9, 1–9. [Google Scholar] [CrossRef]
- Zhuang, H.M.; Lou, Q.; Liu, H.F.; Han, H.W.; Wang, Q.; Tang, Z.H.; Ma, Y.M.; Wang, H. Differential regulation of anthocyanins in green and purple turnips revealed by combined de novo transcriptome and metabolome analysis. Int. J. Mol. Sci. 2019, 20, 4387. [Google Scholar] [CrossRef]
- Lou, Q.; Liu, Y.L.; Qi, Y.Y.; Jiao, S.Z.; Tian, F.F.; Jiang, L.; Wang, Y.J. Transcriptome sequencing and metabolite analysis reveals the role of delphinidin metabolism in flower colour in grape hyacinth. J. Exp. Bot. 2014, 65, 3157–3164. [Google Scholar] [CrossRef]
- Matus, J.T. Transcriptomic and metabolomic networks in the grape berry illustrate that it takes more than flavonoids to fight against ultraviolet radiation. Front. Plant. Sci. 2016, 7, 1337. [Google Scholar] [CrossRef]
- Shen, Y.H.; Yang, F.Y.; Lu, B.G.; Zhao, W.W.; Jiang, T.; Feng, L.; Chen, X.J.; Ming, R. Exploring the differential mechanisms of carotenoid biosynthesis in the yellow peel and red flesh of papaya. BMC Genom. 2019, 20, 49. [Google Scholar] [CrossRef]
- Wang, F.; Chen, L.; Chen, H.P.; Chen, S.W.; Liu, Y.P. Analysis of flavonoid metabolites in citrus peels (Citrus reticulata “Dahongpao”) using UPLC-ESI-MS/MS. Molecules 2019, 24, 2680. [Google Scholar] [CrossRef]
- Chen, Y.H.; Zhang, R.P.; Song, Y.M.; He, J.M.; Sun, J.H.; Bai, J.F.; An, Z.L.; Dong, L.J.; Zhan, Q.M.; Abliz, Z. RRLC-MS/MS-based metabonomics combined with in-depth analysis of metabolic correlation network: Finding potential biomarkers for breast cancer. Analyst 2009, 134, 2003. [Google Scholar] [CrossRef]
- Latchman, D.S. Transcription factors: An overview. Int. J. Exp. Pathol. 1993, 74, 417–422. [Google Scholar] [CrossRef]
- Schwechheimer, C.; Bevan, M. The regulation of transcription factor activity in plants. Trends Plant. Sci. 1998, 3, 378–383. [Google Scholar] [CrossRef]
- Burbulis, I.E.; Winkel-Shirley, B. Interactions among enzymes of the Arabidopsis flavonoid biosynthetic pathway. Proc. Natl. Acad. Sci. USA 1999, 96, 12929–12934. [Google Scholar] [CrossRef]
- Ferrer, J.L.; Austin, M.B.; Stewart, C.; Noel, J.P. Structure and function of enzymes involved in the biosynthesis of phenylpropanoids. Plant. Physiol. Bioch. 2008, 46, 356–370. [Google Scholar] [CrossRef]
- Jun, S.Y.; Sattler, S.A.; Cortez, G.S.; Vermerris, W.; Sattler, S.E.; Kang, C.H. Biochemical and structural analysis of substrate specificity of a phenylalanine ammonia-lyase. Plant. Physiol. 2018, 176, 1452–1468. [Google Scholar] [CrossRef]
- González-Aguilar, G.A.; Zavaleta-Gatica, R.; Tiznado-Hernández, M.E. Improving postharvest quality of mango ‘Haden’ by UV-C treatment. Postharvest Biol. Tec. 2007, 45, 108–116. [Google Scholar] [CrossRef]
- Boerjan, W.; Ralph, J.; Baucher, M. Lignin biosynthesis. Annu. Rev. Plant. Biol. 2003, 54, 519–546. [Google Scholar] [CrossRef]
- Alberstein, M.; Eisenstein, M.; Abeliovich, H. Removing allosteric feedback inhibition of tomato 4-coumarate:CoA ligase by directed evolution. Plant. J. 2012, 69, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Martin, C.R. Structure, function, and regulation of the chalcone synthase. In International Review of Cytology; Jeon, K.W., Jarvik, J., Eds.; Academic Press: Cambridge, MA, USA, 1993; Volume 147, pp. 233–284. [Google Scholar]
- Abe, I.; Watanabe, T.; Morita, H.; Kohno, T.; Noguchi, H. Engineered biosynthesis of plant polyketides: Manipulation of chalcone synthase. Org. Lett. 2006, 8, 499–502. [Google Scholar] [CrossRef]
- Xie, S.X.; Cheng, L.; Zeng, W.; Zhang, W.; Yuan, Y.; Yuan, H.Y. Molecular cloning and expression of HCT gene from camellia sinensis. J. Northeast For. Univ. 2013, 41, 19–22. [Google Scholar]
- Hoffmann, L.; Maury, S.P.; Bergdoll, M.; Thion, L.; Erard, M.; Legrand, M. Identification of the enzymatic active site of tobacco caffeoyl-coenzyme A O-methyltransferase by site-directed mutagenesis. J. Biol. Chem. 2001, 276, 36831. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.F.; Balint-Kurti, P.J. Maize Homologs of CCoAOMT and HCT, two key enzymes in lignin biosynthesis, form complexes with the NLR Rp1 protein to modulate the defense response. Plant. Physiol. 2016, 171, 2166–2177. [Google Scholar] [CrossRef]
- Day, A.; Neutelings, G.; Nolin, F.; Grec, S.; Habrant, A.; Crônier, D.; Maher, B.; Rolando, C.; David, H.; Chabbert, B.; et al. Caffeoyl coenzyme A O-methyltransferase down-regulation is associated with modifications in lignin and cell-wall architecture in flax secondary xylem. Plant. Physiol. Bioch. 2009, 47, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Widiez, T.; Hartman, T.G.; Dudai, N.; Yan, Q.; Lawton, M.; Havkin-Frenkel, D.; Belanger, F.C. Functional characterization of two new members of the caffeoyl CoA O-methyltransferase-like gene family from Vanilla planifolia reveals a new class of plastid-localized O-methyltransferases. Plant. Mol. Biol. 2011, 76, 475–488. [Google Scholar] [CrossRef]
- Lado, J.; Gambetta, G.; Zacarias, L. Key determinants of citrus fruit quality: Metabolites and main changes during maturation. Sci. Hortic. 2018, 233, 238–248. [Google Scholar] [CrossRef]
- Brand, A.; Borovsky, Y.; Hill, T.; Rahman, K.A.A.; Bellalou, A.; Van Deynze, A.; Paran, I. CaGLK2 regulates natural variation of chlorophyll content and fruit color in pepper fruit. Appl. Genet. 2014, 127, 2139–2148. [Google Scholar] [CrossRef]
- Nguyen, C.V.; Vrebalov, J.T.; Gapper, N.E.; Zheng, Y.; Zhong, S.L.; Fei, Z.J.; Giovannoni, J.J. Tomato GOLDEN2-LIKE transcription factors reveal molecular gradients that function during fruit development and ripening. Plant. Cell 2014, 26, 585–601. [Google Scholar] [CrossRef]
- Ni, F.R.; Wu, L.; Wang, Q.; Hong, J.; Qi, Y.J.; Zhou, X.P. Turnip yellow mosaic virus P69 Interacts with and suppresses GLK transcription factors to cause pale-green symptoms in Arabidopsis. Mol. Plant. 2017, 10, 764–766. [Google Scholar] [CrossRef] [PubMed]
- Hwang, I.; Kim, Y.; Han, J.; Nou, I.S. Orange color is associated with CYC-B expression in tomato fleshy fruit. Mol. Breed. 2016, 36, 42. [Google Scholar] [CrossRef]
- Ballester, A.R.; Molthoff, J.; de Vos, R.; Hekkert, B.T.L.; Orzaez, D.; Fernández-Moreno, J.; Tripodi, P.; Grandillo, S.; Martin, C.; Heldens, J.; et al. Biochemical and molecular analysis of pink tomatoes: Deregulated expression of the gene encoding transcription factor SlMYB12 leads to pink tomato fruit color. Plant. Physiol. 2010, 152, 71–84. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Moreno, J.; Tzfadia, O.; Forment, J.; Presa, S.; Rogachev, I.; Meir, S.; Orzaez, D.; Aharoni, A.; Granell, A. Characterization of a New pink-fruited tomato mutant results in the identification of a null allele of the SlMYB12 transcription factor. Plant. Physiol. 2016, 171, 1821–1836. [Google Scholar] [CrossRef]
- Xu, W.J.; Dubos, C.; Lepiniec, L. Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant. Sci. 2015, 20, 176–185. [Google Scholar] [CrossRef]
- Wang, Z.B.; Yu, Q.B.; Shen, W.X.; El Mohtar, C.A.; Zhao, X.C.; Gmitter, F.G., Jr. Functional study of CHS gene family members in citrus revealed a novel CHS gene affecting the production of flavonoids. BMC Plant. Biol 2018, 18, 189. [Google Scholar] [CrossRef]
- Goławska, S.; Goławska, S.; Sprawka, I.; Sprawka, I.; Łukasik, I.; Łukasik, I.; Goławski, A.; Goławski, A. Are naringenin and quercetin useful chemicals in pest-management strategies? J. Pest. Sci. 2014, 87, 173–180. [Google Scholar] [CrossRef]
- Deng, Y.X.; Lu, S.F. Biosynthesis and regulation of phenylpropanoids in plants. Crit. Rev. Plant. Sci 2017, 36, 1–34. [Google Scholar] [CrossRef]
- Deng, Y.X.; Li, C.L.; Li, H.Q.; Lu, S.F. Identification and characterization of flavonoid biosynthetic enzyme genes in Salvia miltiorrhiza (Lamiaceae). Molecules 2018, 23, 1467. [Google Scholar] [CrossRef]
- Muhlemann, J.K.; Younts, T.L.B.; Muday, G.K. Flavonols control pollen tube growth and integrity by regulating ROS homeostasis during high-temperature stress. Proc. Natl. Acad. Sci. 2018, 115, E11188–E11197. [Google Scholar] [CrossRef]
- Brodowska, K.M. Natural flavonoids: Classification, potential role, and application of flavonoid analogues. Eur. J. Biol. Res. 2017, 7, 108–123. [Google Scholar]
- Liu, M.Y.; Zhang, C.J.; Duan, L.X.; Luan, Q.Q.; Li, J.L.; Yang, A.G.; Qi, X.Q.; Ren, Z.H. CsMYB60 is a key regulator of flavonols and proanthocyanidans that determine the colour of fruit spines in cucumber. J. Exp. Bot. 2019, 70, 69–84. [Google Scholar] [CrossRef] [PubMed]
- Peterson, J.J.; Dwyer, J.T.; Beecher, G.R.; Bhagwat, S.A.; Gebhardt, S.E.; Haytowitz, D.B.; Holden, J.M. Flavanones in oranges, tangerines (mandarins), tangors, and tangelos: A compilation and review of the data from the analytical literature. J. Food Compos. Anal. 2006, 19, S66–S73. [Google Scholar] [CrossRef]
- Goulas, V.; Manganaris, G.A. Exploring the phytochemical content and the antioxidant potential of Citrus fruits grown in Cyprus. Food Chem. 2012, 131, 39–47. [Google Scholar] [CrossRef]
- Zhao, D.Q.; Tang, W.H.; Hao, Z.J.; Tao, J. Identification of flavonoids and expression of flavonoid biosynthetic genes in two coloured tree peony flowers. Biochem. Bioph. Res. Commun. 2015, 459, 450–456. [Google Scholar] [CrossRef]
- Wellmann, F.; Lukačin, R.; Moriguchi, T.; Britsch, L.; Schiltz, E.; Matern, U. Functional expression and mutational analysis of flavonol synthase from Citrus unshiu. Eur. J. Biochem. 2002, 269, 4134–4142. [Google Scholar] [CrossRef]
- Bajpai, A.; Khan, K.; Muthukumar, M.; Rajan, S.; Singh, N.K. Molecular analysis of anthocyanin biosynthesis pathway genes and their differential expression in mango peel. Genome 2018, 61, 157–166. [Google Scholar] [CrossRef]
- Singh, R.S.; Kesari, R.; Kumar, U.; Jha, V.K.; Kumar, A.; Kumar, T.; Pal, A.K.; Singh, P.K. Candidate genes of flavonoid biosynthesis in Selaginella bryopteris (L.) Baker identified by RNA-Seq. Funct. Integr. Genom. 2018, 18, 505–517. [Google Scholar] [CrossRef]
- Park, S.; Kim, D.H.; Lee, J.Y.; Ha, S.H.; Lim, S.H. Comparative Analysis of Two Flavonol Synthases from Different-Colored Onions Provides Insight into Flavonoid Biosynthesis. J. Agr. Food Chem. 2017, 65, 5287–5298. [Google Scholar] [CrossRef]
- Yan, J.; Yu, L.; He, L.Z.; Zhu, L.; Xu, S.; Wan, Y.H.; Wang, H.; Wang, Y.; Zhu, W.M. Comparative Transcriptome Analysis of Celery Leaf Blades Identified an R2R3-MYB Transcription Factor that Regulates Apigenin Metabolism. J. Agr. Food Chem. 2019, 67, 5265–5277. [Google Scholar] [CrossRef]
- Xu, F.; Li, L.L.; Zhang, W.W.; Cheng, H.; Sun, N.N.; Cheng, S.Y.; Wang, Y. Isolation, characterization, and function analysis of a flavonol synthase gene from Ginkgo biloba. Mol. Biol. Rep. 2012, 39, 2285–2296. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.Q.; Guo, J.; Zhou, Q.; Xin, Y.; Wang, G.B.; Xu, L.A. De novo transcriptome analysis revealed genes involved in flavonoid biosynthesis, transport and regulation in Ginkgo biloba. Ind. Crop. Prod. 2018, 124, 226–235. [Google Scholar] [CrossRef]
- Wang, C.H.; Zhi, S.; Liu, C.Y.; Xu, F.X.; Zhao, A.C.; Wang, X.L.; Tang, X.; Li, Z.G.; Huang, P.; Yu, M.D. Isolation and characterization of a novel chalcone synthase gene family from mulberry. Plant. Physiol. Biochem. 2017, 115, 107–118. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Tang, X.H.; Ren, C.X.; Wei, B.; Wu, Y.Y.; Wu, Q.H.; Pei, J. Full-length transcriptome sequences and the identification of putative genes for flavonoid biosynthesis in safflower. BMC Genom. 2018, 19, 1–13. [Google Scholar] [CrossRef]
- He, X.J.; Zhao, X.C.; Gao, L.P.; Shi, X.X.; Dai, X.L.; Liu, Y.J.; Xia, T.; Wang, Y.S. Isolation and characterization of key genes that promote flavonoid accumulation in purple-leaf tea (Camellia sinensis L.). Sci. Rep. 2018, 8, 1–13. [Google Scholar] [CrossRef]
- Liu, S.; Liu, L.; Tang, Y.; Xiong, S.; Long, J.; Liu, Z.; Tian, N. Comparative transcriptomic analysis of key genes involved in flavonoid biosynthetic pathway and identification of a flavonol synthase from Artemisia annua L. Plant. Biol. 2017, 19, 618–629. [Google Scholar] [CrossRef]
- Baldi, P.; Moser, M.; Brilli, M.; Vrhovsek, U.; Pindo, M.; Si-Ammour, A. Fine-tuning of the flavonoid and monolignol pathways during apple early fruit development. Planta 2017, 245, 1021–1035. [Google Scholar] [CrossRef]
- Besseau, S.; Hoffmann, L.; Geoffroy, P.; Lapierre, C.; Pollet, B.; Legrand, M. Flavonoid accumulation in Arabidopsis repressed in lignin synthesis affects auxin transport and plant growth. Plant. Cell 2007, 19, 148–162. [Google Scholar] [CrossRef]
- Kreuzaler, F.; Hahlbrock, K. Enzymatic synthesis of aromatic compounds in higher plants: Formation of naringenin (5, 7, 4′-trihydroxyflavanone) from p-coumaroyl coenzyme A and malonyl coenzyme A. FEBS Lett. 1972, 28, 69–72. [Google Scholar] [CrossRef]
- Awasthi, P.; Mahajan, V.; Jamwal, V.L.; Kapoor, N.; Rasool, S.; Bedi, Y.S.; Gandhi, S.G. Cloning and expression analysis of chalcone synthase gene from Coleus forskohlii. J. Genet. 2016, 95, 647–657. [Google Scholar] [CrossRef]
- Winkel-Shirley, B. Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant. Physiol. 2001, 126, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Chaudhary, P.R.; Bang, H.; Jayaprakasha, G.K.; Patil, B.S. Variation in key flavonoid biosynthetic enzymes and phytochemicals in ‘Rio Red’ Grapefruit (Citrus paradisi Macf.) during fruit development. J. Agr. Food Chem. 2016, 64, 9022–9032. [Google Scholar] [CrossRef] [PubMed]
- Shaipulah, N.F.M.; Muhlemann, J.K.; Woodworth, B.D.; Van Moerkercke, A.; Verdonk, J.C.; Ramirez, A.A.; Haring, M.A.; Dudareva, N.; Schuurink, R.C. CCoAOMT Down-regulation activates anthocyanin biosynthesis in petunia1. Plant. Physiol. 2016, 170, 717–731. [Google Scholar] [CrossRef] [PubMed]
- Feller, A.; Machemer, K.; Braun, E.L.; Grotewold, E. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant. J. 2011, 66, 94–116. [Google Scholar] [CrossRef] [PubMed]
- Grunewald, W.; De Smet, I.; Lewis, D.R.; Lofke, C.; Jansen, L.; Goeminne, G.; Vanden Bossche, R.; Karimi, M.; De Rybel, B.; Vanholme, B.; et al. Transcription factor WRKY23 assists auxin distribution patterns during Arabidopsis root development through local control on flavonol biosynthesis. Proc. Natl. Acad. Sci. USA 2012, 109, 1554–1559. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yang, Z.; Zeng, Q.W.; Wang, S.B.; Luo, Y.W.; Huang, Y.; Xin, Y.C.; He, N.J. Abnormal expression of bHLH3 disrupts a flavonoid homeostasis network, causing differences in pigment composition among mulberry fruits. Hortic. Res. 2020, 7, 1–19. [Google Scholar] [CrossRef]
- An, X.H.; Tian, Y.; Chen, K.Q.; Wang, X.F.; Hao, Y.J. The apple WD40 protein MdTTG1 interacts with bHLH but not MYB proteins to regulate anthocyanin accumulation. J. Plant. Physiol. 2012, 169, 710–717. [Google Scholar] [CrossRef] [PubMed]
- Dubos, C.; Stracke, R.; Grotewold, E.; Weisshaar, B.; Martin, C.; Lepiniec, L. MYB transcription factors in Arabidopsis. Trends Plant. Sci. 2010, 15, 573–581. [Google Scholar] [CrossRef]
- Stracke, R.; Werber, M.; Weisshaar, B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin Plant. Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef]
- Zimmermann, I.M.; Heim, M.A.; Weisshaar, B.; Uhrig, J.F. Comprehensive identification of Arabidopsis thaliana MYB transcription factors interacting with R/B-like BHLH proteins. Plant. J. 2004, 40, 22–34. [Google Scholar] [CrossRef]
- Jin, H.L.; Cominelli, E.; Bailey, P.; Parr, A.; Mehrtens, F.; Jones, J.; Tonelli, C.; Weisshaar, B.; Martin, C. Transcriptional repression by AtMYB4 controls production of UV-protecting sunscreens in Arabidopsis. EMBO J. 2000, 19, 6150–6161. [Google Scholar] [CrossRef] [PubMed]
- Bray, N.L.; Pimentel, H.; Melsted, P.; Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 2016, 34, 525–527. [Google Scholar] [CrossRef] [PubMed]

| Component Name | Compounds | OL14 vs. OL19 LogFC | YL14 vs. OL14 LogFC | Pathway ID | Index |
|---|---|---|---|---|---|
| Isoflavone | |||||
| 6-Hydroxydaidzein | −14.12976 | 14.12976 | c | pme3261 | |
| Glycitein | −11.50735 | 11.50735 | c | pme3251 | |
| Calycosin | −11.45121 | 11.45121 | c | pme3230 | |
| Formononetin 7-O-glucoside (Ononin) | −10.78560 | 10.78560 | c | pme3502 | |
| Genistein 7-O-Glucoside (Genistin) | −8.44094 | 9.46536 | c | pme3210 | |
| Flavanone | |||||
| Naringenin O-malonylhexoside | −17.18100 | 17.18100 | - | pma0791 | |
| Naringenin | −8.77594 | 10.47588 | a, c | pme0376 | |
| Naringenin chalcone | −6.71026 | 6.83489 | a | pme2957 | |
| Homoeriodictyol | −6.07142 | 6.00144 | a | pme3461 | |
| Hesperetin | −5.73054 | 6.56258 | a | pme2319 | |
| Isosakuranetin (4’-Methylnaringenin) | −3.78556 | 8.45274 | - | pme3464 | |
| Flavone C-glycosides | |||||
| Apigenin C-glucoside | −16.90395 | 5.52477 | - | pma1108 | |
| Isovitexin | −13.37429 | 4.78364 | d | pme0374 | |
| Naringenin C-hexoside | −12.92101 | 3.95331 | - | pma0724 | |
| C-hexosyl-apigenin O-caffeoylhexoside | −11.33398 | 2.92648 | - | pma6254 | |
| C-hexosyl-apigenin O-p-coumaroylhexoside | −7.01972 | 4.84490 | - | pmb0680 | |
| Apigenin 6-C-hexosyl-8-C-hexosyl-O-hexoside | −5.59166 | 3.92600 | - | pmb0613 | |
| 8-C-hexosyl-apigenin O-hexosyl-O-hexoside | −5.43549 | 3.87321 | - | pmb0639 | |
| Flavonol | |||||
| Kaempferide | −15.43731 | 15.43731 | d | pma1116 | |
| Aromadedrin (Dihydrokaempferol) | −13.13512 | 13.13512 | a | pme2963 | |
| Flavone | |||||
| Acacetin O-acetyl hexoside | −12.48981 | 12.48981 | - | pmb2987 | |
| sakuranetin | −11.85475 | 11.85475 | - | pme1662 | |
| Butin | −8.86582 | 10.58564 | a | pme3473 | |
| Apigenin 5-O-glucoside | −8.47617 | 19.03466 | - | pme0359 | |
| Apigenin 7-O-glucoside (Cosmosiin) | −8.05708 | 8.32675 | d | pmb0605 | |
| Apigenin | −4.35888 | 4.98833 | a, c, d | pme0379 | |
| Anthocyanins | |||||
| Pelargonidin 3-O-beta-D-glucoside (Callistephin chloride) | −14.15048 | 5.65299 | - | pme3392 | |
| Pelargonidin | −3.76095 | 14.92857 | a, b | pme1397 | |
| Catechin derivatives | |||||
| Epicatechin gallate (ECG) | −10.38881 | 10.38881 | - | pme1562 | |
| Epigallate catechin gallate (EGCG) | −9.86568 | 1.90675 | - | pme1486 | |
| GeneID | OL14_TPM | OL19_TPM | Log2FC | Description |
|---|---|---|---|---|
| Csa6G147460.1 | 14.38 | 2.82 | −2.35 | Phenylalanine ammonia-lyase, PAL |
| Csa6G446290.1 | 29.73 | 14.41 | −1.05 | Phenylalanine ammonia-lyase, PAL |
| Csa3G710200.1 | 27.32 | 3.75 | −2.86 | 4-coumarate-CoA ligase, 4CL |
| Csa5G154230.1 | 149.94 | 65.11 | −1.20 | 4-coumarate-CoA ligase, 4CL |
| Csa7G431440.1 | 11.08 | 1.86 | −2.57 | Shikimate O-hydroxycinnamoyltransferase, HCT |
| Csa7G073660.1 | 131.04 | 18.06 | −2.86 | Caffeoyl-CoA O-methyltransferase, CCoAOMT |
| Csa3G600020.1 | 78.22 | 10.12 | −2.95 | Chalcone synthase, CHS |
| Csa4G112650.1 | 15.13 | 4.97 | −1.60 | Flavonol synthase, FLS |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, C.; Zhou, G.; Chen, J.; Liu, X.; Lu, X.; Chen, H.; Tian, Y. Integrated Metabolome and Transcriptome Analysis Unveils Novel Pathway Involved in the Formation of Yellow Peel in Cucumber. Int. J. Mol. Sci. 2021, 22, 1494. https://doi.org/10.3390/ijms22031494
Chen C, Zhou G, Chen J, Liu X, Lu X, Chen H, Tian Y. Integrated Metabolome and Transcriptome Analysis Unveils Novel Pathway Involved in the Formation of Yellow Peel in Cucumber. International Journal of Molecular Sciences. 2021; 22(3):1494. https://doi.org/10.3390/ijms22031494
Chicago/Turabian StyleChen, Chen, Geng Zhou, Juan Chen, Xiaohong Liu, Xiangyang Lu, Huiming Chen, and Yun Tian. 2021. "Integrated Metabolome and Transcriptome Analysis Unveils Novel Pathway Involved in the Formation of Yellow Peel in Cucumber" International Journal of Molecular Sciences 22, no. 3: 1494. https://doi.org/10.3390/ijms22031494
APA StyleChen, C., Zhou, G., Chen, J., Liu, X., Lu, X., Chen, H., & Tian, Y. (2021). Integrated Metabolome and Transcriptome Analysis Unveils Novel Pathway Involved in the Formation of Yellow Peel in Cucumber. International Journal of Molecular Sciences, 22(3), 1494. https://doi.org/10.3390/ijms22031494







