Bioactive Lipids in MSCs Biology: State of the Art and Role in Inflammation
Abstract
1. Introduction: Lipidomics and Lipids Mediated Inflammation in Mesenchymal Stem Cells
1.1. Lipidomics
1.2. Involvement of MSC in Inflammatory Processes
1.3. Functional Role of Endogenous Bioactive Lipids in Inflammation
2. Lipids as Signaling Mediators in Inflammation
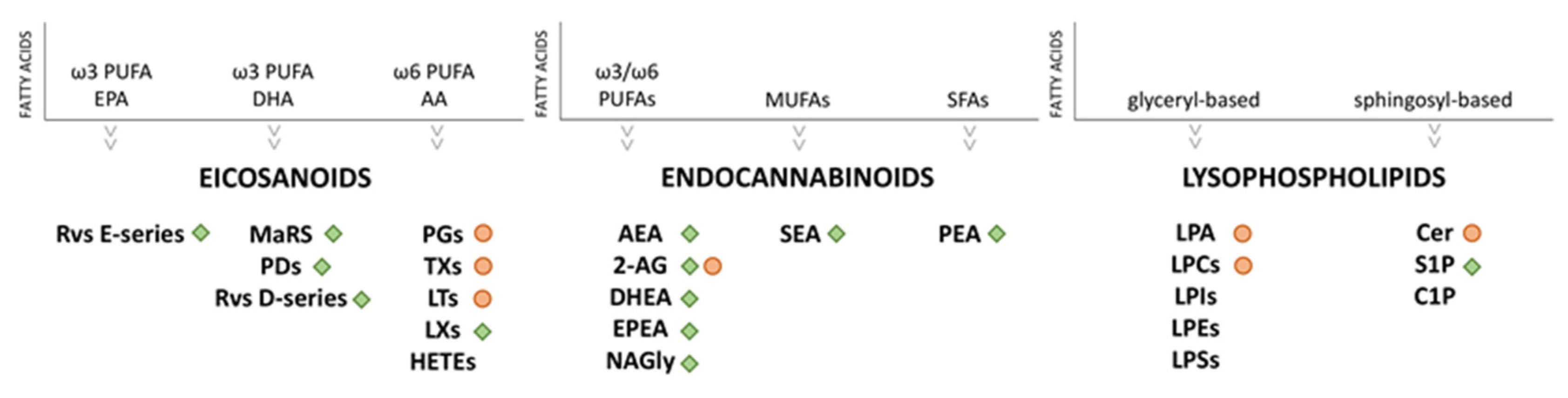
2.1. Eicosanoids
2.2. Endocannabinoids (eCBs) and Endocannabinoid-Like Compounds
2.3. Lysophospholipids (LPLs)
3. Bioactive Lipids in MSCs
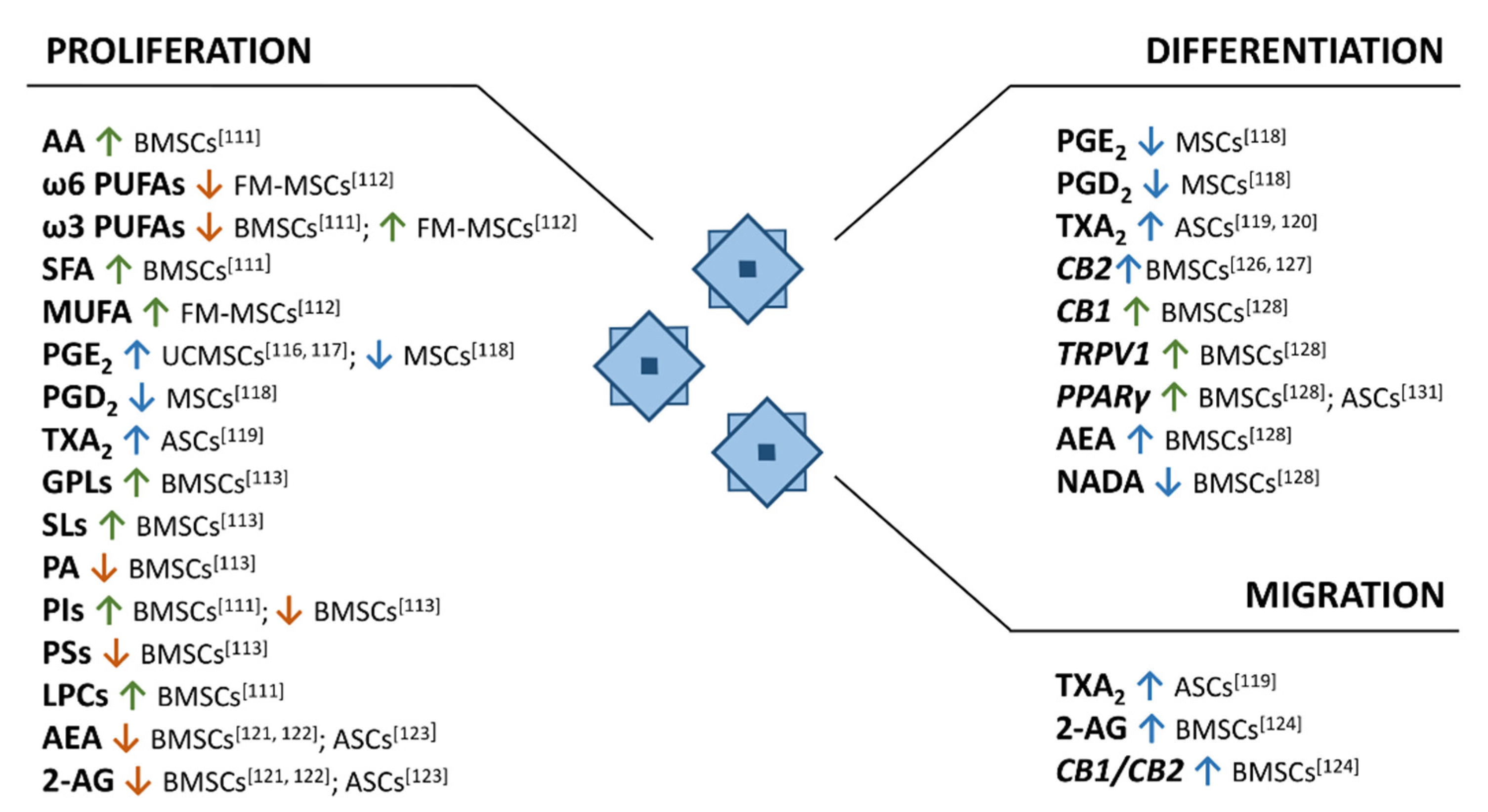
3.1. Lipid Metabolism in MSCs Maintenance and Differentiation
3.2. Pro and Anti-Inflammatory Properties of MSC-Derived Lipids
3.3. Effect of Exogenous Supplements of PUFAs on MSCs
3.4. MSCs as an Alternative Treatment of Inflammatory Diseases: The Example of Osteoarthritis
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Fahy, E.; Subramaniam, S.; Murphy, R.C.; Nishijima, M.; Raetz, C.R.H.; Shimizu, T.; Spener, F.; Van Meer, G.; Wakelam, M.J.O.; Dennis, E.A. Update of the LIPID MAPS comprehensive classification system for lipids. J. Lipid Res. 2009, 50, S9–S14. [Google Scholar] [CrossRef]
- Fahy, E.; Cotter, D.; Sud, M.; Subramaniam, S. Lipid classification, structures and tools. Biochim. Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2011, 1811, 637–647. [Google Scholar] [CrossRef]
- García-Cañaveras, J.C.; Peris-Díaz, M.D.; Alcoriza-Balaguer, M.I.; Cerdán-Calero, M.; Donato, M.T.; Lahoz, A. A lipidomic cell-based assay for studying drug-induced phospholipidosis and steatosis. Electrophoresis 2017, 38, 2331–2340. [Google Scholar] [CrossRef]
- Lydic, T.; Goo, Y.-H. Lipidomics unveils the complexity of the lipidome in metabolic diseases. Clin. Transl. Med. 2018, 7, 4. [Google Scholar] [CrossRef]
- Sethi, S.; Brietzke, E. Recent advances in lipidomics: Analytical and clinical perspectives. Prostaglandins Other Lipid Mediat. 2017, 128–129, 8–16. [Google Scholar] [CrossRef]
- Nguyen, V.L.; Haber, P.; Seth, D. Applications and Challenges for the Use of Phosphatidylethanol Testing in Liver Disease Patients (Mini Review). Alcohol. Clin. Exp. Res. 2018, 42, 238–243. [Google Scholar] [CrossRef]
- German, J.B.; Gillies, L.A.; Smilowitz, J.T.; Zivkovic, A.M.; Watkins, S.M. Lipidomics and lipid profiling in metabolomics. Curr. Opin. Lipidol. 2007, 18, 66–71. [Google Scholar]
- Wenk, M.R. The emerging field of lipidomics. Nat. Rev. Drug Discov. 2005, 4, 594–610. [Google Scholar] [CrossRef]
- Wenk, M.R. Lipidomics: New Tools and Applications. Cell 2010, 143, 888–895. [Google Scholar] [CrossRef]
- Schwudke, D.; Shevchenko, A.; Hoffmann, N.; Ahrends, R. Lipidomics informatics for life-science. J. Biotechnol. 2017, 261, 131–136. [Google Scholar] [CrossRef]
- Shevchenko, A.; Simons, K. Lipidomics: Coming to grips with lipid diversity. Nat. Rev. Mol. Cell Biol. 2010, 11, 593–598. [Google Scholar] [CrossRef]
- Li, L.; Han, J.; Wang, Z.; Liu, J.; Wei, J.; Xiong, S.; Zhao, Z. Mass Spectrometry Methodology in Lipid Analysis. Int. J. Mol. Sci. 2014, 15, 10492–10507. [Google Scholar] [CrossRef]
- Yang, K.; Han, X. Lipidomics: Techniques, Applications, and Outcomes Related to Biomedical Sciences. Trends Biochem. Sci. 2016, 41, 954–969. [Google Scholar] [CrossRef]
- Casati, S.; Giannasi, C.; Minoli, M.; Niada, S.; Ravelli, A.; Angeli, I.; Mergenthaler, V.; Ottria, R.; Ciu, P.; Orioli, M.; et al. Quantitative Lipidomic Analysis of Osteosarcoma Cell-Derived Products by UHPLC-MS/MS. Biomolecules 2020, 10, 1302. [Google Scholar] [CrossRef]
- Locke, M.; Windsor, J.; Dunbar, P.R. Human adipose-derived stem cells: Isolation, characterization and applications in surgery. ANZ J. Surg. 2009, 79, 235–244. [Google Scholar] [CrossRef]
- Strem, B.M.; Hicok, K.C.; Zhu, M.; Wulur, I.; Alfonso, Z.; Schreiber, R.E.; Fraser, J.K.; Hedrick, M.H. Multipotential differentiation of adipose tissue-derived stem cells. Keio J. Med. 2005, 54, 132–141. [Google Scholar] [CrossRef]
- Dominici, M.; Le Blanc, K.; Mueller, I.; Slaper-Cortenbach, I.; Marini, F.; Krause, D.; Deans, R.; Keating, A.; Prockop, D.; Horwitz, E. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 2006, 8, 315–317. [Google Scholar] [CrossRef]
- Horwitz, E.M.; Le Blanc, K.; Dominici, M.; Mueller, I.; Slaper-Cortenbach, I.; Marini, F.C.; Deans, R.J.; Krause, D.S.; Keating, A. Clarification of the nomenclature for MSC: The International Society for Cellular Therapy position statement. Cytotherapy 2005, 7, 393–395. [Google Scholar] [CrossRef]
- Glassberg, M.K.; Minkiewicz, J.; Toonkel, R.L.; Simonet, E.S.; Rubio, G.A.; DiFede, D.; Shafazand, S.; Khan, A.; Pujol, M.V.; LaRussa, V.F.; et al. Allogeneic Human Mesenchymal Stem Cells in Patients with Idiopathic Pulmonary Fibrosis via Intravenous Delivery (AETHER): A Phase I Safety Clinical Trial. Chest 2017, 151, 971–981. [Google Scholar] [CrossRef]
- Lindsay, J.O.; Allez, M.; Clark, M.; Labopin, M.; Ricart, E.; Rogler, G.; Rovira, M.; Satsangi, J.; Farge, D.; Hawkey, C.J.; et al. Autologous stem-cell transplantation in treatment-refractory Crohn’s disease: An analysis of pooled data from the ASTIC trial. Lancet Gastroenterol. Hepatol. 2017, 2, 399–406. [Google Scholar] [CrossRef]
- Shi, Y.; Wang, Y.; Li, Q.; Liu, K.; Hou, J.; Shao, C.; Wang, Y. Immunoregulatory mechanisms of mesenchymal stem and stromal cells in inflammatory diseases. Nat. Rev. Nephrol. 2018, 14, 493–507. [Google Scholar] [CrossRef] [PubMed]
- Mehta, P.; McAuley, D.F.; Brown, M.; Sanchez, E.; Tattersall, R.S.; Manson, J.J. COVID-19: Consider cytokine storm syndromes and immunosuppression. Lancet 2020, 395, 1033–1034. [Google Scholar] [CrossRef]
- Golchin, A.; Seyedjafari, E.; Ardeshirylajimi, A. Mesenchymal Stem Cell Therapy for COVID-19: Present or Future. Stem Cell Rev. Rep. 2020, 16, 427–433. [Google Scholar] [CrossRef] [PubMed]
- Gowen, A.; Shahjin, F.; Chand, S.; Odegaard, K.E.; Yelamanchili, S.V. Mesenchymal Stem Cell-Derived Extracellular Vesicles: Challenges in Clinical Applications. Front. Cell Dev. Biol. 2020, 8, 149. [Google Scholar] [CrossRef] [PubMed]
- Sagini, K.; Costanzi, E.; Emiliani, C.; Buratta, S.; Urbanelli, L. Extracellular Vesicles as Conveyors of Membrane-Derived Bioactive Lipids in Immune System. Int. J. Mol. Sci. 2018, 19, 1227. [Google Scholar] [CrossRef]
- Harrell, C.R.; Fellabaum, C.; Jovicic, N.; Djonov, V.; Arsenijevic, N.; Volarevic, V. Molecular Mechanisms Responsible for Therapeutic Potential of Mesenchymal Stem Cell-Derived Secretome. Cells 2019, 8, 467. [Google Scholar] [CrossRef]
- English, K. Mechanisms of mesenchymal stromal cell immunomodulation. Immunol. Cell Biol. 2013, 91, 19–26. [Google Scholar] [CrossRef]
- Aggarwal, S.; Pittenger, M.F. Human mesenchymal stem cells modulate allogeneic immune cell responses. Blood 2005, 105, 1815–1822. [Google Scholar] [CrossRef]
- Bernardo, M.E.; Locatelli, F.; Fibbe, W.E. Mesenchymal Stromal Cells. Ann. N. Y. Acad. Sci. 2009, 1176, 101–117. [Google Scholar] [CrossRef]
- Bernardo, M.E.; Cometa, A.M.; Pagliara, D.; Vinti, L.; Rossi, F.; Cristantielli, R.; Palumbo, G.; Locatelli, F. Ex vivo expansion of mesenchymal stromal cells. Best Pract. Res. Clin. Haematol. 2011, 24, 73–81. [Google Scholar] [CrossRef]
- Mushahary, D.; Spittler, A.; Kasper, C.; Weber, V.; Charwat, V. Isolation, cultivation, and characterization of human mesenchymal stem cells. Cytom. Part A 2018, 93, 19–31. [Google Scholar] [CrossRef]
- Baer, P.C.; Geiger, H. Adipose-Derived Mesenchymal Stromal/Stem Cells: Tissue Localization, Characterization, and Heterogeneity. Stem Cells Int. 2012, 2012, 1–11. [Google Scholar] [CrossRef]
- Bertozzi, N.; Simonacci, F.; Grieco, M.P.; Grignaffini, E.; Raposio, E. The biological and clinical basis for the use of adipose-derived stem cells in the field of wound healing. Ann. Med. Surg. 2017, 20, 41–48. [Google Scholar] [CrossRef]
- Kokai, L.E.; Marra, K.; Rubin, J.P. Adipose stem cells: Biology and clinical applications for tissue repair and regeneration. Transl. Res. 2014, 163, 399–408. [Google Scholar] [CrossRef]
- Muhammad, G.; Xu, J.; Bulte, J.; Jablonska, A.; Walczak, P.; Janowski, M. Transplanted adipose-derived stem cells can be short-lived yet accelerate healing of acid-burn skin wounds: A multimodal imaging study. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef]
- Pavyde, E.; Maciulaitis, R.; Mauricas, M.; Sudzius, G.; Didziokiene, E.I.; Laurinavicius, A.; Sutkeviciene, N.; Stankevicius, E.; Maciulaitis, J.; Usas, A. Skeletal Muscle-Derived Stem/Progenitor Cells: A Potential Strategy for the Treatment of Acute Kidney Injury. Stem Cells Int. 2016, 2016. [Google Scholar] [CrossRef]
- Jankowski, R.J.; Deasy, B.M.; Huard, J. Muscle-derived stem cells. Gene Ther. 2002, 9, 642–647. [Google Scholar] [CrossRef]
- Ceusters, J.; Lejeune, J.P.J.; Sandersen, C.C.; Niesten, A.; Lagneaux, L.; Serteyn, D.D. From skeletal muscle to stem cells: An innovative and minimally-invasive process for multiple species. Sci. Rep. 2017, 7, 1–9. [Google Scholar] [CrossRef]
- Shimizu, T. Lipid Mediators in Health and Disease: Enzymes and Receptors as Therapeutic Targets for the Regulation of Immunity and Inflammation. Annu. Rev. Pharmacol. Toxicol. 2009, 49, 123–150. [Google Scholar] [CrossRef]
- Leuti, A.; Fazio, D.; Fava, M.; Piccoli, A.; Oddi, S.; Maccarrone, M. Bioactive lipids, inflammation and chronic diseases. Adv. Drug Deliv. Rev. 2020, 159, 133–169. [Google Scholar] [CrossRef]
- Chiurchiù, V.; Leuti, A.; Maccarrone, M. Bioactive Lipids and Chronic Inflammation: Managing the Fire Within. Front. Immunol. 2018, 9, 38. [Google Scholar] [CrossRef]
- Chiurchiù, V.; Battistini, L.; Maccarrone, M. Endocannabinoid signalling in innate and adaptive immunity. Immunology 2015, 144, 352–364. [Google Scholar] [CrossRef]
- Chiurchiù, V.; Maccarrone, M. Bioactive lipids as modulators of immunity, inflammation and emotions. Curr. Opin. Pharmacol. 2016, 29, 54–62. [Google Scholar] [CrossRef]
- Maceyka, M.; Spiegel, S. Sphingolipid metabolites in inflammatory disease. Nat. Cell Biol. 2014, 510, 58–67. [Google Scholar] [CrossRef]
- Smith, W.L.; DeWitt, D.L.; Garavito, R.M. Cyclooxygenases: Structural, Cellular, and Molecular Biology. Annu. Rev. Biochem. 2000, 69, 145–182. [Google Scholar] [CrossRef]
- Stitham, J.; Midgett, C.R.; Martin, K.A.; Hwa, J. Prostacyclin: An Inflammatory Paradox. Front. Pharmacol. 2011, 2, 24. [Google Scholar] [CrossRef]
- Mandal, A.K.; Jones, P.B.; Bair, A.M.; Christmas, P.; Miller, D.; Yamin, T.-T.D.; Wisniewski, D.; Menke, J.; Evans, J.F.; Hyman, B.T.; et al. The nuclear membrane organization of leukotriene synthesis. Proc. Natl. Acad. Sci. USA 2008, 105, 20434–20439. [Google Scholar] [CrossRef]
- Murphy, R.; Gijón, M.A. Biosynthesis and metabolism of leukotrienes. Biochem. J. 2007, 405, 379–395. [Google Scholar] [CrossRef]
- Serhan, C.N. Pro-resolving lipid mediators are leads for resolution physiology. Nature 2014, 510, 92–101. [Google Scholar] [CrossRef]
- Dennis, E.A.; Norris, P.C. Eicosanoid storm in infection and inflammation. Nat. Rev. Immunol. 2015, 15, 511–523. [Google Scholar] [CrossRef]
- Aoki, T.; Narumiya, S. Prostaglandins and chronic inflammation. Trends Pharmacol. Sci. 2012, 33, 304–311. [Google Scholar] [CrossRef]
- Narumiya, S.; Furuyashiki, T. Fever, inflammation, pain and beyond: Prostanoid receptor research during these 25 years. FASEB J. 2010, 25, 813–818. [Google Scholar] [CrossRef]
- Honda, T.; Segi-Nishida, E.; Miyachi, Y.; Narumiya, S. Prostacyclin-IP signaling and prostaglandin E2-EP2/EP4 signaling both mediate joint inflammation in mouse collagen-induced arthritis. J. Exp. Med. 2006, 203, 325–335. [Google Scholar] [CrossRef]
- Hirata, T.; Narumiya, S. Chapter Five—Prostanoids as Regulators of Innate and Adaptive Immunity; Elsevier: Amsterdam, The Netherlands, 2012; Volume 116. [Google Scholar]
- Yao, C.; Sakata, D.; Esaki, Y.; Li, Y.; Matsuoka, T.; Kuroiwa, K.; Sugimoto, Y.; Narumiya, S. Prostaglandin E2–EP4 signaling promotes immune inflammation through TH1 cell differentiation and TH17 cell expansion. Nat. Med. 2009, 15, 633–640. [Google Scholar] [CrossRef]
- Chen, Q.; Muramoto, K.; Masaaki, N.; Ding, Y.; Yang, H.; Mackey, M.F.; Li, W.; Inoue, Y.; Ackermann, K.; Shirota, H.; et al. A novel antagonist of the prostaglandin E2 EP4 receptor inhibits Th1 differentiation and Th17 expansion and is orally active in arthritis models. Br. J. Pharmacol. 2010, 160, 292–310. [Google Scholar] [CrossRef]
- Gilroy, D.W.; Bishop-Bailey, D. Lipid mediators in immune regulation and resolution. Br. J. Pharmacol. 2019, 176, 1009–1023. [Google Scholar] [CrossRef]
- Chiurchiù, V.; Leuti, A.; Dalli, J.; Jacobsson, A.; Battistini, L.; Maccarrone, M.; Serhan, C.N. Proresolving lipid mediators resolvin D1, resolvin D2, and maresin 1 are critical in modulating T cell responses. Sci. Transl. Med. 2016, 8, 353ra111. [Google Scholar] [CrossRef]
- Schwab, J.M.; Chiang, N.; Arita, M.; Serhan, C.N. Resolvin E1 and protectin D1 activate inflammation-resolution programmes. Nat. Cell Biol. 2007, 447, 869–874. [Google Scholar] [CrossRef]
- Serhan, C.N.; Chiang, N. Resolution phase lipid mediators of inflammation: Agonists of resolution. Curr. Opin. Pharmacol. 2013, 13, 632–640. [Google Scholar] [CrossRef]
- Serhan, C.N.; Chiang, N.; Dalli, J. The resolution code of acute inflammation: Novel pro-resolving lipid mediators in resolution. Semin. Immunol. 2015, 27, 200–215. [Google Scholar] [CrossRef]
- Miyahara, T.; Runge, S.; Chatterjee, A.; Chen, M.; Mottola, G.; Fitzgerald, J.M.; Serhan, C.N.; Conte, M.S. D-series resolvin attenuates vascular smooth muscle cell activation and neointimal hyperplasia following vascular injury. FASEB J. 2013, 27, 2220–2232. [Google Scholar] [CrossRef]
- Basil, M.C.; Levy, B.D. Specialized pro-resolving mediators: Endogenous regulators of infection and inflammation. Nat. Rev. Immunol. 2016, 16, 51–67. [Google Scholar] [CrossRef]
- Stables, M.J.; Gilroy, D.W. Old and new generation lipid mediators in acute inflammation and resolution. Prog. Lipid Res. 2011, 50, 35–51. [Google Scholar] [CrossRef]
- Herlong, J.; Scott, T. Positioning prostanoids of the D and J series in the immunopathogenic scheme. Immunol. Lett. 2006, 102, 121–131. [Google Scholar] [CrossRef]
- Bannenberga, G.; Serhanb, C.N. Specialized Pro-Resolving Lipid Mediators in the Inflammatory Response: An Update. Biochim. Biophys. Acta (BBA) 2010, 1801, 1260–1273. [Google Scholar] [CrossRef]
- Spector, A.A.; Fang, X.; Snyder, G.D.; Weintraub, N.L. Epoxyeicosatrienoic acids (EETs): Metabolism and biochemical function. Prog. Lipid Res. 2004, 43, 55–90. [Google Scholar] [CrossRef]
- Ligresti, A.; De Petrocellis, L.; Di Marzo, V. From Phytocannabinoids to Cannabinoid Receptors and Endocannabinoids: Pleiotropic Physiological and Pathological Roles Through Complex Pharmacology. Physiol. Rev. 2016, 96, 1593–1659. [Google Scholar] [CrossRef]
- Pertwee, R.G. Cannabidiol as a potential medicine. In Cannabinoids as Therapeutics; Birkhäuser: Basel, Switzerland, 2005. [Google Scholar] [CrossRef]
- Di Marzo, V. Targeting the endocannabinoid system: To enhance or reduce? Nat. Rev. Drug Discov. 2008, 7, 438–455. [Google Scholar] [CrossRef]
- Devane, W.A.; Hanus, L.; Breuer, A.; Pertwee, R.G.; Stevenson, L.A.; Griffin, G.; Gibson, D.; Mandelbaum, A.; Etinger, A.; Mechoulam, R. Isolation and structure of a brain constituent that binds to the cannabinoid receptor. Science 1992, 258, 1946–1949. [Google Scholar] [CrossRef]
- Sugiura, T.; Kondo, S.; Sukagawa, A.; Nakane, S.; Shinoda, A.; Itoh, K.; Yamashita, A.; Waku, K. 2-Arachidonoylgylcerol: A Possible Endogenous Cannabinoid Receptor Ligand in Brain. Biochem. Biophys. Res. Commun. 1995, 215, 89–97. [Google Scholar] [CrossRef]
- Mechoulam, R.; Ben-Shabat, S.; Hanus, L.; Ligumsky, M.; Kaminski, N.E.; Schatz, A.R.; Gopher, A.; Almog, S.; Martin, B.R.; Compton, D.R.; et al. Identification of an endogenous 2-monoglyceride, present in canine gut, that binds to cannabinoid receptors. Biochem. Pharmacol. 1995, 50, 83–90. [Google Scholar] [CrossRef]
- Ribeiro, A.; Pontis, S.; Mengatto, L.; Armirotti, A.; Chiurchiù, V.; Capurro, V.; Fiasella, A.; Nuzzi, A.; Romeo, E.; Moreno-Sanz, G.; et al. A Potent Systemically Active N-Acylethanolamine Acid Amidase Inhibitor that Suppresses Inflammation and Human Macrophage Activation. ACS Chem. Biol. 2015, 10, 1838–1846. [Google Scholar] [CrossRef]
- Maestroni, G.J.M. The endogenous cannabinoid 2-arachidonoyl glycerol as in vivo chemoattractant for dendritic cells and adjuvant for Th1 response to a soluble protein. FASEB J. 2004, 18, 1914–1916. [Google Scholar] [CrossRef]
- Gallily, R.; Breuer, A.; Mechoulam, R. 2-Arachidonylglycerol, an endogenous cannabinoid, inhibits tumor necrosis factor-α production in murine macrophages, and in mice. Eur. J. Pharmacol. 2000, 406, R5–R7. [Google Scholar] [CrossRef]
- Barrie, N.; Manolios, N. The endocannabinoid system in pain and inflammation: Its relevance to rheumatic disease. Eur. J. Rheumatol. 2017, 4, 210–218. [Google Scholar] [CrossRef]
- Artmann, A.; Petersen, G.; Hellgren, L.I.; Boberg, J.; Skonberg, C.; Nellemann, C.L.; Hansen, S.H.; Hansen, H.S. Influence of dietary fatty acids on endocannabinoid and N-acylethanolamine levels in rat brain, liver and small intestine. Biochim. Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2008, 1781, 200–212. [Google Scholar] [CrossRef]
- Lucanic, M.; Held, J.M.; Vantipalli, M.C.; Klang, I.M.; Graham, J.B.; Gibson, B.W.; Lithgow, G.J.; Gill, M.S. N-acylethanolamine signalling mediates the effect of diet on lifespan in Caenorhabditis elegans. Nat. Cell Biol. 2011, 473, 226–229. [Google Scholar] [CrossRef]
- Brown, I.; Cascio, M.G.; Wahle, K.W.; Smoum, R.; Mechoulam, R.; Ross, R.A.; Pertwee, R.G.; Heys, S.D. Cannabinoid receptor-dependent and -independent anti-proliferative effects of omega-3 ethanolamides in androgen receptor-positive and -negative prostate cancer cell lines. Carcinogenesis 2010, 31, 1584–1591. [Google Scholar] [CrossRef]
- Meijerink, J.; Plastina, P.; Vincken, J.-P.; Poland, M.; Attya, M.; Balvers, M.G.J.; Gruppen, H.; Gabriele, B.; Witkamp, R.F. The ethanolamide metabolite of DHA, docosahexaenoylethanolamine, shows immunomodulating effects in mouse peritoneal and RAW264.7 macrophages: Evidence for a new link between fish oil and inflammation. Br. J. Nutr. 2011, 105, 1798–1807. [Google Scholar] [CrossRef]
- Piomelli, D. More surprises lying ahead. The endocannabinoids keep us guessing. Neuropharmacology 2014, 76, 228–234. [Google Scholar] [CrossRef]
- Payahoo, L.; Khajebishak, Y.; Ostadrahimi, A. Oleoylethanolamide: A novel pharmaceutical agent in the management of obesity-an updated review. J. Cell. Physiol. 2019, 234, 7893–7902. [Google Scholar] [CrossRef]
- TuTunchi, H.; Saghafi-Asl, M.; Ostadrahimi, A. A systematic review of the effects of oleoylethanolamide, a high-affinity endogenous ligand of PPAR-α, on the management and prevention of obesity. Clin. Exp. Pharmacol. Physiol. 2020, 47, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Raso, G.M.; Russo, R.; Calignano, A.; Meli, R. Palmitoylethanolamide in CNS health and disease. Pharmacol. Res. 2014, 86, 32–41. [Google Scholar] [CrossRef] [PubMed]
- Petrosino, S.; Di Marzo, V. The pharmacology of palmitoylethanolamide and first data on the therapeutic efficacy of some of its new formulations. Br. J. Pharmacol. 2017, 174, 1349–1365. [Google Scholar] [CrossRef] [PubMed]
- Ryberg, E.; Larsson, N.; Sjögren, S.; Hjorth, S.; Hermansson, N.-O.; Leonova, J.; Elebring, T.; Nilsson, K.; Drmota, T.; Greasley, P.J. The orphan receptor GPR55 is a novel cannabinoid receptor. Br. J. Pharmacol. 2007, 152, 1092–1101. [Google Scholar] [CrossRef]
- Hansen, H.S. Palmitoylethanolamide and other anandamide congeners. Proposed role in the diseased brain. Exp. Neurol. 2010, 224, 48–55. [Google Scholar] [CrossRef]
- Maccarrone, M.; Cartoni, A.; Parolaro, D.; Margonelli, A.; Massi, P.; Bari, M.; Battista, N.; Finazzi-Agrò, A. Cannabimimetic activity, binding, and degradation of stearoylethanolamide within the mouse central nervous system. Mol. Cell. Neurosci. 2002, 21, 126–140. [Google Scholar] [CrossRef]
- Carbonare, M.D.; Del Giudice, E.; Stecca, A.; Colavito, D.; Fabris, M.; D’Arrigo, A.; Bernardini, D.; Dam, M.; León, A. A Saturated N-Acylethanolamine Other than N-Palmitoyl Ethanolamine with Anti-inflammatory Properties: A Neglected Story. J. Neuroendocr. 2008, 20, 26–34. [Google Scholar] [CrossRef]
- Balvers, M.G.J.; Verhoeckx, K.C.; Witkamp, R.F. Development and validation of a quantitative method for the determination of 12 endocannabinoids and related compounds in human plasma using liquid chromatography–tandem mass spectrometry. J. Chromatogr. B 2009, 877, 1583–1590. [Google Scholar] [CrossRef]
- Han, B.; Wright, R.; Kirchhoff, A.M.; Chester, J.A.; Cooper, B.R.; Davisson, V.J.; Barker, E.L. Quantitative LC–MS/MS analysis of arachidonoyl amino acids in mouse brain with treatment of FAAH inhibitor. Anal. Biochem. 2013, 432, 74–81. [Google Scholar] [CrossRef]
- Burstein, S.; McQuain, C.A.; Ross, A.H.; Salmonsen, R.A.; Zurier, R.E. Resolution of inflammation by N-arachidonoylglycine. J. Cell. Biochem. 2011, 112, 3227–3233. [Google Scholar] [CrossRef] [PubMed]
- Burstein, S.; Adams, J.K.; Bradshaw, H.B.; Fraioli, C.; Rossetti, R.G.; Salmonsen, R.A.; Shaw, J.W.; Walker, J.M.; Zipkin, R.E.; Zurier, R.B. Potential anti-inflammatory actions of the elmiric (lipoamino) acids. Bioorg. Med. Chem. 2007, 15, 3345–3355. [Google Scholar] [CrossRef] [PubMed]
- Parmar, N.; Ho, W.S.V. N-arachidonoyl glycine, an endogenous lipid that acts as a vasorelaxant via nitric oxide and large conductance calcium-activated potassium channels. Br. J. Pharmacol. 2010, 160, 594–603. [Google Scholar] [CrossRef] [PubMed]
- McHugh, D.; Hu, S.S.-J.; Rimmerman, N.; Juknat, A.; Vogel, Z.; Walker, J.M.; Bradshaw, H. N-arachidonoyl glycine, an abundant endogenous lipid, potently drives directed cellular migration through GPR18, the putative abnormal cannabidiol receptor. BMC Neurosci. 2010, 11, 44. [Google Scholar] [CrossRef]
- Cascio, M.G.; Minassi, A.; Ligresti, A.; Appendino, G.; Burstein, S.; Di Marzo, V. A structure-activity relationship study on N-arachidonoyl-amino acids as possible endogenous inhibitors of fatty acid amide hydrolase. Biochem. Biophys. Res. Commun. 2004, 314, 192–196. [Google Scholar] [CrossRef]
- Goetzl, E.J.; Graeler, M.; Huang, M.-C.; Shankar, G. Lysophospholipid Growth Factors and Their G Protein-Coupled Receptors in Immunity, Coronary Artery Disease, and Cancer. Sci. World J. 2002, 2, 324–338. [Google Scholar] [CrossRef][Green Version]
- Huang, M.-C.; Graeler, M.; Shankar, G.; Spencer, J.; Goetzl, E.J. Lysophospholipid mediators of immunity and neoplasia. Biochim. Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2002, 1582, 161–167. [Google Scholar] [CrossRef]
- Makide, K.; Kitamura, H.; Sato, Y.; Okutani, M.; Aoki, J. Emerging lysophospholipid mediators, lysophosphatidylserine, lysophosphatidylthreonine, lysophosphatidylethanolamine and lysophosphatidylglycerol. Prostaglandins Other Lipid Mediat. 2009, 89, 135–139. [Google Scholar] [CrossRef]
- Sevastou, I.; Kaffe, E.; Mouratis, M.; Aidinis, V. Lysoglycerophospholipids in chronic inflammatory disorders: The PLA2/LPC and ATX/LPA axes. Biochim. Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2013, 1831, 42–60. [Google Scholar] [CrossRef]
- Gault, C.R.; Obeid, L.M.; Hannun, Y.A. An Overview of Sphingolipid Metabolism: From Synthesis to Breakdown. In Sphingolipids as Signaling and Regulatory Molecules; Springer: New York, NY, USA, 2010. [Google Scholar] [CrossRef]
- McArthur, S.; Gobbetti, T.; Kusters, D.H.M.; Reutelingsperger, C.P.; Flower, R.J.; Perretti, M. Definition of a Novel Pathway Centered on Lysophosphatidic Acid to Recruit Monocytes during the Resolution Phase of Tissue Inflammation. J. Immunol. 2015, 195, 1139–1151. [Google Scholar] [CrossRef]
- El Alwani, M.; Wu, B.X.; Obeid, L.M.; Hannun, Y.A. Bioactive sphingolipids in the modulation of the inflammatory response. Pharmacol. Ther. 2006, 112, 171–183. [Google Scholar] [CrossRef]
- Gomez-Muñoz, A.; Presa, N.; Gomez-Larrauri, A.; Rivera, I.-G.; Trueba, M.; Ordoñez, M. Control of inflammatory responses by ceramide, sphingosine 1-phosphate and ceramide 1-phosphate. Prog. Lipid Res. 2016, 61, 51–62. [Google Scholar] [CrossRef]
- Bartke, N.; Hannun, Y.A. Bioactive sphingolipids: Metabolism and function. J. Lipid Res. 2009, 50, S91–S96. [Google Scholar] [CrossRef]
- Weichand, B.; Weis, N.; Weigert, A.; Grossmann, N.; Levkau, B.; Brüne, B. Apoptotic cells enhance sphingosine-1-phosphate receptor 1 dependent macrophage migration. Eur. J. Immunol. 2013, 43, 3306–3313. [Google Scholar] [CrossRef]
- Yanes, O.; Clark, J.; Wong, D.M.; Patti, G.J.; Sánchez-Ruiz, A.; Benton, H.P.; Trauger, S.A.; Desponts, C.; Ding, S.; Siuzdak, G. Metabolic oxidation regulates embryonic stem cell differentiation. Nat. Chem. Biol. 2010, 6, 411–417. [Google Scholar] [CrossRef]
- Knobloch, M.; Braun, S.M.G.; Zurkirchen, L.; Von Schoultz, C.; Zamboni, N.; Araúzo-Bravo, M.J.; Kovacs, W.J.; Karalay, Ö.; Suter, U.; Machado, R.A.C.; et al. Metabolic control of adult neural stem cell activity by Fasn-dependent lipogenesis. Nat. Cell Biol. 2013, 493, 226–230. [Google Scholar] [CrossRef]
- Clémot, M.; Demarco, R.S.; Jones, D.L. Lipid Mediated Regulation of Adult Stem Cell Behavior. Front. Cell Dev. Biol. 2020, 8, 115. [Google Scholar] [CrossRef]
- Kilpinen, L.; Tigistu-Sahle, F.; Oja, S.; Greco, D.; Parmar, A.; Saavalainen, P.; Nikkilä, J.; Korhonen, M.; Lehenkari, P.; Käkelä, R.; et al. Aging bone marrow mesenchymal stromal cells have altered membrane glycerophospholipid composition and functionality. J. Lipid Res. 2013, 54, 622–635. [Google Scholar] [CrossRef]
- Chatgilialoglu, A.; Rossi, M.; Alviano, F.; Poggi, P.; Zannini, C.; Marchionni, C.; Ricci, F.; Tazzari, P.L.; Taglioli, V.; Calder, P.C.; et al. Restored in vivo-like membrane lipidomics positively influence in vitro features of cultured mesenchymal stromal/stem cells derived from human placenta. Stem Cell Res. Ther. 2017, 8, 1–11. [Google Scholar] [CrossRef]
- Lu, X.; Chen, Y.; Wang, H.; Bai, Y.; Zhao, J.; Zhang, X.; Liang, L.; Chen, Y.; Ye, C.; Li, Y.; et al. Integrated Lipidomics and Transcriptomics Characterization upon Aging-Related Changes of Lipid Species and Pathways in Human Bone Marrow Mesenchymal Stem Cells. J. Proteome Res. 2019, 18, 2065–2077. [Google Scholar] [CrossRef]
- Wagner, W.; Horn, P.; Castoldi, M.; Diehlmann, A.; Bork, S.; Saffrich, R.; Benes, V.; Blake, J.; Pfister, S.; Eckstein, V.; et al. Replicative Senescence of Mesenchymal Stem Cells: A Continuous and Organized Process. PLoS ONE 2008, 3, e2213. [Google Scholar] [CrossRef]
- Gharibi, B.; Hughes, F.J. Effects of Medium Supplements on Proliferation, Differentiation Potential, and In Vitro Expansion of Mesenchymal Stem Cells. Stem Cells Transl. Med. 2012, 1, 771–782. [Google Scholar] [CrossRef] [PubMed]
- Jang, M.W.; Yun, S.P.; Park, J.H.; Ryu, J.M.; Lee, J.H.; Han, H.J. Cooperation of Epac1/Rap1/Akt and PKA in prostaglandin E2-induced proliferation of human umbilical cord blood derived mesenchymal stem cells: Involvement of c-Myc and VEGF expression. J. Cell. Physiol. 2012, 227, 3756–3767. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.P.; Ryu, J.M.; Jang, M.W.; Han, H.J. Interaction of profilin-1 and F-actin via a β-arrestin-1/JNK signaling pathway involved in prostaglandin E2-induced human mesenchymal stem cells migration and proliferation. J. Cell. Physiol. 2010, 226, 559–571. [Google Scholar] [CrossRef] [PubMed]
- Ern, C.; Frasheri, I.; Berger, T.; Kirchner, H.; Heym, R.; Hickel, R.; Folwaczny, M. Effects of prostaglandin E2 and D2 on cell proliferation and osteogenic capacity of human mesenchymal stem cells. Prostaglandins Leukot. Essent. Fat. Acids 2019, 151, 1–7. [Google Scholar] [CrossRef]
- Yun, D.H.; Song, H.Y.; Lee, M.J.; Kim, M.R.; Kim, M.Y.; Lee, J.S.; Kim, J.H. Thromboxane A2modulates migration, proliferation, and differentiation of adipose tissue-derived mesenchymal stem cells. Exp. Mol. Med. 2009, 41, 17–24. [Google Scholar] [CrossRef][Green Version]
- Kim, M.R.; Jeon, E.S.; Kim, Y.M.; Lee, J.S.; Kim, J.H. Thromboxane A2Induces Differentiation of Human Mesenchymal Stem Cells to Smooth Muscle-Like Cells. Stem Cells 2009, 27, 191–199. [Google Scholar] [CrossRef]
- Rossi, F.; Bernardo, M.E.; Bellini, G.; Luongo, L.; Conforti, A.; Manzo, I.; Guida, F.; Cristino, L.; Imperatore, R.; Petrosino, S.; et al. The Cannabinoid Receptor Type 2 as Mediator of Mesenchymal Stromal Cell Immunosuppressive Properties. PLoS ONE 2013, 8, e80022. [Google Scholar] [CrossRef]
- Köse, S.; Aerts-Kaya, F.S.F.; Köprü, Ç.Z.; Nemutlu, E.; Kuşkonmaz, B.; Karaosmanoğlu, B.; Taşkıran, E.Z.; Altun, B.; Çetinkaya, D.U.; Korkusuz, P. Human bone marrow mesenchymal stem cells secrete endocannabinoids that stimulate in vitro hematopoietic stem cell migration effectively comparable to beta-adrenergic stimulation. Exp. Hematol. 2018, 57, 30–41.e1. [Google Scholar] [CrossRef]
- Ivanov, I.; Borchert, P.; Hinz, B. A simple method for simultaneous determination of N-arachidonoylethanolamine, N-oleoylethanolamine, N-palmitoylethanolamine and 2-arachidonoylglycerol in human cells. Anal. Bioanal. Chem. 2014, 407, 1781–1787. [Google Scholar] [CrossRef] [PubMed]
- Scutt, A.; Williamson, E.M. Cannabinoids Stimulate Fibroblastic Colony Formation by Bone Marrow Cells Indirectly via CB2 Receptors. Calcif. Tissue Int. 2007, 80, 50–59. [Google Scholar] [CrossRef]
- Galve-Roperh, I.; Chiurchiù, V.; Díaz-Alonso, J.; Bari, M.; Guzmán, M.; Maccarrone, M. Cannabinoid receptor signaling in progenitor/stem cell proliferation and differentiation. Prog. Lipid Res. 2013, 52, 633–650. [Google Scholar] [CrossRef]
- Sun, Y.-X.; Xu, A.-H.; Yang, Y.; Zhang, J.-X.; Yu, A.-W. Activation of Cannabinoid Receptor 2 Enhances Osteogenic Differentiation of Bone Marrow Derived Mesenchymal Stem Cells. BioMed Res. Int. 2015, 2015, 1–8. [Google Scholar] [CrossRef]
- Xu, A.; Yang, Y.; Shao, Y.; Wu, M.; Sun, Y. Activation of cannabinoid receptor type 2-induced osteogenic differentiation involves autophagy induction and p62-mediated Nrf2 deactivation. Cell Commun. Signal. 2020, 18, 9–11. [Google Scholar] [CrossRef]
- Ahn, S.; Yi, S.; Seo, W.J.; Lee, M.J.; Song, Y.K.; Baek, S.Y.; Yu, J.; Hong, S.H.; Lee, J.; Shin, D.W.; et al. A Cannabinoid Receptor Agonist N-Arachidonoyl Dopamine Inhibits Adipocyte Differentiation in Human Mesenchymal Stem Cells. Biomol. Ther. 2015, 23, 218–224. [Google Scholar] [CrossRef]
- Bellocchio, L.; Cervino, C.; Vicennati, V.; Pasquali, R.; Pagotto, U. Cannabinoid Type 1 Receptor: Another Arrow in the Adipocytes’ Bow. J. Neuroendocr. 2008, 20, 130–138. [Google Scholar] [CrossRef]
- Karaliota, S.; Siafaka-Kapadai, A.; Gontinou, C.; Psarra, K.; Mavri-Vavayanni, M. Anandamide Increases the Differentiation of Rat Adipocytes and Causes PPARγ and CB1 Receptor Upregulation. Obesity 2009, 17, 1830–1838. [Google Scholar] [CrossRef]
- Pagano, C.; Pilon, C.; Calcagno, A.; Urbanet, R.; Rossato, M.; Milan, G.; Bianchi, K.; Rizzuto, R.; Bernante, P.; Federspil, G.; et al. The Endogenous Cannabinoid System Stimulates Glucose Uptake in Human Fat Cells via Phosphatidylinositol 3-Kinase and Calcium-Dependent Mechanisms. J. Clin. Endocrinol. Metab. 2007, 92, 4810–4819. [Google Scholar] [CrossRef]
- Da Silva, C.G.; Barretto, L.S.D.S.; Turco, E.G.L.; Santos, A.D.L.; Lessio, C.; Júnior, H.A.M.; De Almeida, F.G. Lipidomics of mesenchymal stem cell differentiation. Chem. Phys. Lipids 2020, 232, 232. [Google Scholar] [CrossRef]
- Griffin, M.; Elliman, S.; Cahill, E.; English, K.; Ceredig, R.; Ritter, T. Concise Review: Adult Mesenchymal Stromal Cell Therapy for Inflammatory Diseases: How Well Are We Joining the Dots? Stem Cells Regen. Med. 2013, 31, 2033–2041. [Google Scholar] [CrossRef]
- Campos, A.M.; Maciel, E.; Moreira, A.S.P.; Sousa, B.; Melo, T.; Domingues, P.; Curado, L.; Antunes, B.; Domingues, M.D.R.; Dos Santos, F. Lipidomics of Mesenchymal Stromal Cells: Understanding the Adaptation of Phospholipid Profile in Response to Pro-Inflammatory Cytokines. J. Cell. Physiol. 2016, 231, 1024–1032. [Google Scholar] [CrossRef]
- Cunningham, T.J.; Yao, L.; Lucena, A. Product inhibition of secreted phospholipase A2 may explain lysophosphatidylcholines’ unexpected therapeutic properties. J. Inflamm. 2008, 5, 1–10. [Google Scholar] [CrossRef]
- Chen, G.; Li, J.; Qiang, X.; Czura, C.J.; Ochani, M.; Ochani, K.; Ulloa, L.; Yang, H.; Tracey, K.J.; Wang, P.; et al. Suppression of HMGB1 release by stearoyl lysophosphatidylcholine:an additional mechanism for its therapeutic effects in experimental sepsis. J. Lipid Res. 2005, 46, 623–627. [Google Scholar] [CrossRef]
- Olivera, A.; Rivera, J. Sphingolipids and the Balancing of Immune Cell Function: Lessons from the Mast Cell. J. Immunol. 2005, 174, 1153–1158. [Google Scholar] [CrossRef]
- Masoodi, M.; Nicolaou, A. Lipidomic analysis of twenty-seven prostanoids and isoprostanes by liquid chromatography/electrospray tandem mass spectrometry. Rapid Commun. Mass Spectrom. 2006, 20, 3023–3029. [Google Scholar] [CrossRef]
- English, K.; Barry, F.P.; Field-Corbett, C.P.; Mahon, B.P. IFN-γ and TNF-α differentially regulate immunomodulation by murine mesenchymal stem cells. Immunol. Lett. 2007, 110, 91–100. [Google Scholar] [CrossRef]
- Renner, P.; Eggenhofer, E.; Rosenauer, A.; Popp, F.; Steinmann, J.; Slowik, P.; Geissler, E.; Piso, P.; Schlitt, H.; Dahlke, M. Mesenchymal Stem Cells Require a Sufficient, Ongoing Immune Response to Exert Their Immunosuppressive Function. Transplant. Proc. 2009, 41, 2607–2611. [Google Scholar] [CrossRef]
- Fang, X.; Abbott, J.; Cheng, L.; Colby, J.K.; Lee, J.W.; Levy, B.D.; Matthay, M.A. Human Mesenchymal Stem (Stromal) Cells Promote the Resolution of Acute Lung Injury in Part through Lipoxin A4. J. Immunol. 2015, 195, 875–881. [Google Scholar] [CrossRef]
- Levy, B.D.; Clish, C.B.; Schmidt, B.A.; Gronert, K.; Serhan, C.N. Lipid mediator class switching during acute inflammation: Signals in resolution. Nat. Immunol. 2001, 2, 612–619. [Google Scholar] [CrossRef]
- Lands, W. Fish, Omega-3 and Human Health, 2nd ed.; AOCS Publishing: New York, NY, USA, 2005; ISBN 1893997812. [Google Scholar]
- Tigistu-Sahle, F.; Lampinen, M.; Kilpinen, L.; Holopainen, M.; Lehenkari, P.; Laitinen, S.; Käkelä, R. Metabolism and phospholipid assembly of polyunsaturated fatty acids in human bone marrow mesenchymal stromal cells. J. Lipid Res. 2017, 58, 92–110. [Google Scholar] [CrossRef]
- Bijlsma, J.W.J.; Berenbaum, F.; Lafeber, F.P.J.G. Osteoarthritis: An update with relevance for clinical practice. Lancet 2011, 377, 2115–2126. [Google Scholar] [CrossRef]
- Loef, M.; Schoones, J.W.; Kloppenburg, M.; Ioan-Facsinay, A. Fatty acids and osteoarthritis: Different types, different effects. Jt. Bone Spine 2019, 86, 451–458. [Google Scholar] [CrossRef]
- Brouwers, H.; Von Hegedus, J.; Toes, R.E.M.; Kloppenburg, M.; Ioan-Facsinay, A. Lipid mediators of inflammation in rheumatoid arthritis and osteoarthritis. Best Pract. Res. Clin. Rheumatol. 2015, 29, 741–755. [Google Scholar] [CrossRef]
- Barnes, I.; Krasnokutsky, S.; Statnikov, A.; Samuels, J.; Li, Z.; Friese, O.; Le Graverand-Gastineau, M.-P.H.; Rybak, L.; Kraus, V.B.; Jordan, J.M.; et al. Low-Grade Inflammation in Symptomatic Knee Osteoarthritis: Prognostic Value of Inflammatory Plasma Lipids and Peripheral Blood Leukocyte Biomarkers. Arthritis Rheumatol. 2015, 67, 2905–2915. [Google Scholar] [CrossRef]
- Scanzello, C.R.; Plaas, A.; Crow, M.K. Innate immune system activation in osteoarthritis: Is osteoarthritis a chronic wound? Curr. Opin. Rheumatol. 2008, 20, 565–572. [Google Scholar] [CrossRef]
- Baker, K.R.; Matthan, N.R.; Lichtenstein, A.; Niu, J.; Guermazi, A.; Roemer, F.; Grainger, A.; Nevitt, M.; Clancy, M.; Lewis, C.E.; et al. Association of plasma n-6 and n-3 polyunsaturated fatty acids with synovitis in the knee: The MOST study. Osteoarthr. Cartil. 2012, 20, 382–387. [Google Scholar] [CrossRef]
- Zainal, Z.; Longman, A.J.; Hurst, S.; Duggan, K.; Caterson, B.; Hughes, C.E.; Harwood, J.L. Relative efficacies of omega-3 polyunsaturated fatty acids in reducing expression of key proteins in a model system for studying osteoarthritis. Osteoarthr. Cartil. 2009, 17, 896–905. [Google Scholar] [CrossRef]
- Wang, Z.; Guo, A.; Ma, L.; Yu, H.; Zhang, L.; Meng, H.; Cui, Y.; Yu, F.; Yang, B. Docosahexenoic acid treatment ameliorates cartilage degeneration via a p38 MAPK-dependent mechanism. Int. J. Mol. Med. 2016, 37, 1542–1550. [Google Scholar] [CrossRef]
- Hardy, M.M.; Seibert, K.; Manning, P.T.; Currie, M.G.; Woerner, B.M.; Edwards, D.; Koki, A.; Tripp, C.S. Cyclooxygenase 2-dependent prostaglandin E2 modulates cartilage proteoglycan degradation in human osteoarthritis explants. Arthritis Rheum. 2002, 46, 1789–1803. [Google Scholar] [CrossRef]
- Shimpo, H.; Sakai, T.; Kondo, S.; Mishima, S.; Yoda, M.; Hiraiwa, H.; Ishiguro, N. Regulation of prostaglandin E2 synthesis in cells derived from chondrocytes of patients with osteoarthritis. J. Orthop. Sci. 2009, 14, 611–617. [Google Scholar] [CrossRef]
- Benabdoun, H.A.; Rondon, E.-P.; Shi, Q.; Fernandes, J.; Ranger, P.; Fahmi, H.; Benderdour, M. The role of resolvin D1 in the regulation of inflammatory and catabolic mediators in osteoarthritis. Inflamm. Res. 2016, 65, 635–645. [Google Scholar] [CrossRef]
- Demoor, M.; Ollitrault, D.; Gomez-Leduc, T.; Bouyoucef, M.; Hervieu, M.; Fabre, H.; Lafont, J.; Denoix, J.-M.; Audigié, F.; Mallein-Gerin, F.; et al. Cartilage tissue engineering: Molecular control of chondrocyte differentiation for proper cartilage matrix reconstruction. Biochim. Biophys. Acta (BBA) Gen. Subj. 2014, 1840, 2414–2440. [Google Scholar] [CrossRef]
- Baraniak, P.R.; McDevitt, T.C. Stem cell paracrine actions and tissue regeneration. Regen. Med. 2010, 5, 121–143. [Google Scholar] [CrossRef]
- Niada, S.; Giannasi, C.; Gomarasca, M.; Stanco, D.; Casati, S.; Brini, A.T. Adipose-derived stromal cell secretome reduces TNFα-induced hypertrophy and catabolic markers in primary human articular chondrocytes. Stem Cell Res. 2019, 38, 101463. [Google Scholar] [CrossRef]
- Giannasi, C.; Niada, S.; Magagnotti, C.; Ragni, E.; Andolfo, A.; Brini, A.T. Comparison of two ASC-derived therapeutics in an in vitro OA model: Secretome versus extracellular vesicles. Stem Cell Res. Ther. 2020, 11, 1–15. [Google Scholar] [CrossRef]
- Amodeo, G.; Niada, S.; Moschetti, G.; Franchi, S.; Savadori, P.; Brini, A.; Sacerdote, P. Human adipose mesenchymal stem cell secretome relieves pain and neuroinflammation independently of the route of administration in the MIA murine model of osteoarthritis. Brain Behav. Immun. Under review.
- Bruno, S.; Grange, C.; Collino, F.; Deregibus, M.C.; Cantaluppi, V.; Biancone, L.; Tetta, C.; Camussi, G. Microvesicles Derived from Mesenchymal Stem Cells Enhance Survival in a Lethal Model of Acute Kidney Injury. PLoS ONE 2012, 7, e33115. [Google Scholar] [CrossRef]
- Phinney, D.G.; Pittenger, M.F. Concise Review: MSC-Derived Exosomes for Cell-Free Therapy. Stem Cells 2017, 35, 851–858. [Google Scholar] [CrossRef]
- Ratajczak, M.Z. The emerging role of microvesicles in cellular therapies for organ/tissue regeneration. Nephrol. Dial. Transplant. 2011, 26, 1453–1456. [Google Scholar] [CrossRef]
- Haraszti, R.A.; Didiot, M.-C.; Sapp, E.; Leszyk, J.; Shaffer, S.A.; Rockwell, H.E.; Gao, F.; Narain, N.R.; DiFiglia, M.; Kiebish, M.A.; et al. High-resolution proteomic and lipidomic analysis of exosomes and microvesicles from different cell sources. J. Extracell. Vesicles 2016, 5, 32570. [Google Scholar] [CrossRef]
- Llorente, A.; Skotland, T.; Sylvänne, T.; Kauhanen, D.; Róg, T.; Orłowski, A.; Vattulainen, I.; Ekroos, K.; Sandvig, K. Molecular lipidomics of exosomes released by PC-3 prostate cancer cells. Biochim. Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2013, 1831, 1302–1309. [Google Scholar] [CrossRef]
- Xiang, C.; Yang, K.; Liang, Z.; Wan, Y.; Cheng, Y.; Ma, N.; Zhang, H.; Hou, W.; Fu, P. Sphingosine-1-phosphate mediates the therapeutic effects of bone marrow mesenchymal stem cell-derived microvesicles on articular cartilage defect. Transl. Res. 2018, 193, 42–53. [Google Scholar] [CrossRef]
| Categories | Composition | Function | Classes or Examples |
|---|---|---|---|
| Polyunsaturated fatty acids | Carboxylic acid + hydrocarbon chain; synthesized by chain elongation of an acetyl-CoA with malonyl-CoA | Cell signaling; building blocks to complex lipids | AA, EPA, DHA |
| Endocannabinoids and related compounds | Ethanolamide or other head groups + FAs | Cell signaling | AEA, 2AG |
| Lysophospholipids | Polar head group + glycerol or sphingosine backbone | Membrane and lipoprotein composition, cell signaling | LGPLs, LSLs |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Casati, S.; Giannasi, C.; Niada, S.; Bergamaschi, R.F.; Orioli, M.; Brini, A.T. Bioactive Lipids in MSCs Biology: State of the Art and Role in Inflammation. Int. J. Mol. Sci. 2021, 22, 1481. https://doi.org/10.3390/ijms22031481
Casati S, Giannasi C, Niada S, Bergamaschi RF, Orioli M, Brini AT. Bioactive Lipids in MSCs Biology: State of the Art and Role in Inflammation. International Journal of Molecular Sciences. 2021; 22(3):1481. https://doi.org/10.3390/ijms22031481
Chicago/Turabian StyleCasati, Sara, Chiara Giannasi, Stefania Niada, Roberta F. Bergamaschi, Marica Orioli, and Anna T. Brini. 2021. "Bioactive Lipids in MSCs Biology: State of the Art and Role in Inflammation" International Journal of Molecular Sciences 22, no. 3: 1481. https://doi.org/10.3390/ijms22031481
APA StyleCasati, S., Giannasi, C., Niada, S., Bergamaschi, R. F., Orioli, M., & Brini, A. T. (2021). Bioactive Lipids in MSCs Biology: State of the Art and Role in Inflammation. International Journal of Molecular Sciences, 22(3), 1481. https://doi.org/10.3390/ijms22031481







