Pharmacological Approaches for the Modulation of the Potassium Channel KV4.x and KChIPs
Abstract
1. Introduction
2. Potassium Channel Interacting Proteins (KChIPs)
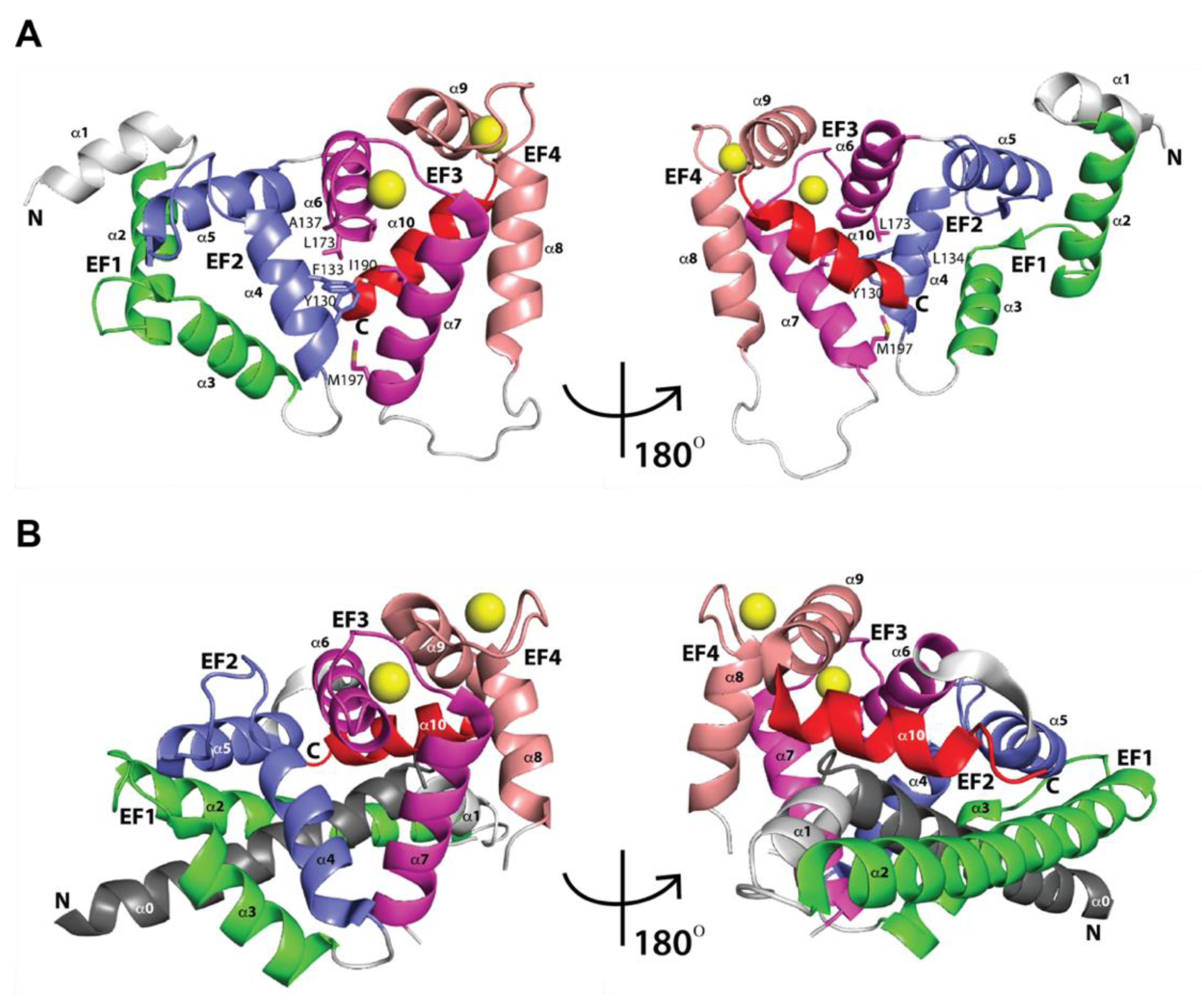
2.1. Three-Dimensional Structures of Free KChIPs
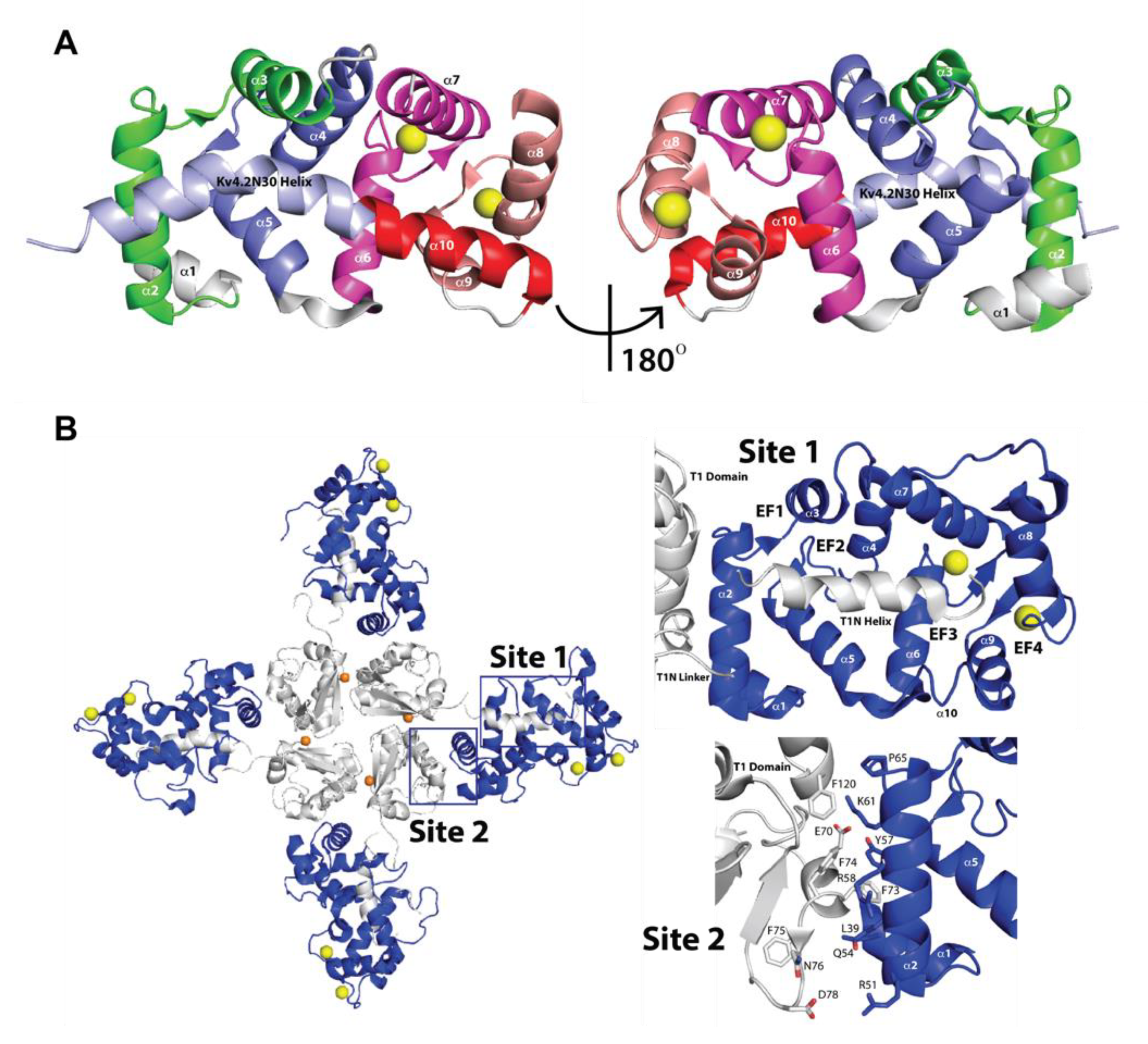
2.2. Three-Dimensional Structures of KChIPs-KV4.2/KV4.3 Channels
3. KChIP Ligands
4. Concluding Remarks
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| 1,8-ANS | 8-anilino-1-naphthalene sulfonate |
| DPPs | Dipeptidyl-peptidase-like proteins |
| ISA | Somato-dendritic subthreshold-activating A-type K+ current |
| ITO | Transient outward current |
| KV | Voltage-gated potassium channels |
| KChIPs | Potassium channel interacting proteins |
| MAPKs | Mitogen-activated protein kinase |
| NMR | Nuclear magnetic resonance |
| PI4K-beta | Phosphatidylinositol 4-kinase beta |
| PKA | Protein kinase A |
| PKC | Protein kinase C |
| SPR | Surface plasmon resonance |
References
- June-Bum, K. Channelopathies. Korean J. Pediatr. 2014, 57, 18. [Google Scholar] [CrossRef]
- Isbrandt, D.; Leicher, T.; Waldschütz, R.; Zhu, X.; Luhmann, U.; Michel, U.; Sauter, K.; Pongs, O. Gene structures and expression profiles of three human KCND (Kv4) potassium channels mediating A-type currents I(TO) and I(SA). Genomics 2000, 64, 144–154. [Google Scholar] [CrossRef] [PubMed]
- Birnbaum, S.G.; Varga, A.W.; Yuan, L.L.; Anderson, A.E.; Sweatt, J.D.; Schrader, L.A. Structure and function of Kv4-family transient potassium channels. Physiol. Rev. 2004, 84, 803–833. [Google Scholar] [CrossRef] [PubMed]
- Nanao, M.H.; Zhou, W.; Pfaffinger, P.J.; Choe, S. Determining the basis of channel-tetramerization specificity by X-ray crystallography and a sequence-comparison algorithm: Family values (FamVal). Proc. Natl. Acad. Sci. USA 2003, 100, 8670–8675. [Google Scholar] [CrossRef] [PubMed]
- Strang, C.; Cushman, S.J.; DeRubeis, D.; Peterson, D.; Pfaffinger, P.J. A Central Role for the T1 Domain in Voltage-gated Potassium Channel Formation and Function. J. Biol. Chem. 2001, 276, 28493–28502. [Google Scholar] [CrossRef]
- Ohya, S.; Tanaka, M.; Oku, T.; Asai, Y.; Watanabe, M.; Giles, W.R.; Imaizumi, Y. Molecular cloning and tissue distribution of an alternatively spliced variant of an A-type K+ channel α-subunit, Kv4.3 in the rat. FEBS Lett. 1997, 420, 47–53. [Google Scholar] [CrossRef]
- Kong, W.; Po, S.; Yamagishi, T.; Ashen, M.D.; Stetten, G.; Tomaselli, G.F. Isolation and characterization of the human gene encoding I(to): Further diversity by alternative mRNA splicing. Am. J. Physiol. Heart Circ. Physiol. 1998, 275, 1963–1970. [Google Scholar] [CrossRef]
- Serodio, P.; Kentros, C.; Rudy, B. Identification of molecular components of A-type channels activating at subthreshold potentials. J. Neurophysiol. 1994, 72, 1516–1529. [Google Scholar] [CrossRef]
- Serôdio, P.; Vega-Saenz De Miera, E.; Rudy, B. Cloning of a novel component of A-type K+ channels operating at subthreshold potentials with unique expression in heart and brain. J. Neurophysiol. 1996, 75, 2174–2179. [Google Scholar] [CrossRef]
- Yang, K.C.; Nerbonne, J.M. Mechanisms contributing to myocardial potassium channel diversity, regulation and remodeling. Trends Cardiovasc. Med. 2016, 26, 209–218. [Google Scholar] [CrossRef]
- Dabrowska, J.; Rainnie, D.G. Expression and distribution of Kv4 potassium channel subunits and potassium channel interacting proteins in subpopulations of interneurons in the basolateral amygdala. Neuroscience 2010, 171, 721–733. [Google Scholar] [CrossRef] [PubMed]
- Alfaro-Ruíz, R.; Aguado, C.; Martín-Belmonte, A.; Moreno-Martínez, A.E.; Luján, R. Expression, cellular and subcellular localisation of Kv4.2 and Kv4.3 channels in the rodent hippocampus. Int. J. Mol. Sci. 2019, 20, 246. [Google Scholar] [CrossRef] [PubMed]
- MacDonald, J.F.; Belrose, J.C.; Xie, Y.-F.; Jackson, M.F. Nonselective cation channels and links to hippocampal ischemia, aging, and dementia. Adv. Exp. Med. Biol. 2013, 961, 433–447. [Google Scholar] [CrossRef] [PubMed]
- Bernard, C.; Anderson, A.; Becker, A.; Poolos, N.P.; Deck, H.; Johnston, D. Acquired dendritic channelopathy in temporal lobe epilepsy. Science 2004, 305, 532–535. [Google Scholar] [CrossRef] [PubMed]
- Breen, M.S.; Dobbyn, A.; Li, Q.; Roussos, P.; Hoffman, G.E.; Stahl, E.; Chess, A.; Sklar, P.; Li, J.B.; Devlin, B.; et al. Global landscape and genetic regulation of RNA editing in cortical samples from individuals with schizophrenia. Nat. Neurosci. 2019, 22, 1402–1412. [Google Scholar] [CrossRef]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2—A multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef] [PubMed]
- Nerbonne, J.M. Molecular basis of functional voltage-gated K+ channel diversity in the mammalian myocardium. J. Physiol. 2000, 525, 285–298. [Google Scholar] [CrossRef]
- Radicke, S.; Cotella, D.; Graf, E.M.; Banse, U.; Jost, N.; Varró, A.; Tseng, G.N.; Ravens, U.; Wettwer, E. Functional modulation of the transient outward current Ito by KCNE β-subunits and regional distribution in human non-failing and failing hearts. Cardiovasc. Res. 2006, 71, 695–703. [Google Scholar] [CrossRef]
- Radicke, S.; Cotella, D.; Graf, E.M.; Ravens, U.; Wettwer, E. Expression and function of dipeptidyl-aminopeptidase-like protein 6 as a putative β-subunit of human cardiac transient outward current encoded by Kv4.3. J. Physiol. 2005, 565, 751–756. [Google Scholar] [CrossRef]
- Capera, J.; Serrano-Novillo, C.; Navarro-Pérez, M.; Cassinelli, S.; Felipe, A. The potassium channel odyssey: Mechanisms of traffic and membrane arrangement. Int. J. Mol. Sci. 2019, 20, 734. [Google Scholar] [CrossRef]
- Mao, L.M.; Wang, J.Q. Synaptically Localized Mitogen-Activated Protein Kinases: Local Substrates and Regulation. Mol. Neurobiol. 2016, 53, 6309–6315. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, T.Y.; Nakao, S.; Wakabayashi, S. Emerging roles of neuronal Ca2+ sensor-1 in cardiac and neuronal tissues: A mini review. Front. Mol. Neurosci. 2019, 12, 56. [Google Scholar] [CrossRef] [PubMed]
- Van der Heyden, M.A.G.; Wijnhoven, T.J.M.; Opthof, T. Molecular aspects of adrenergic modulation of the transient outward current. Cardiovasc. Res. 2006, 71, 430–442. [Google Scholar] [CrossRef] [PubMed]
- Portero, V.; Wilders, R.; Casini, S.; Charpentier, F.; Verkerk, A.O.; Remme, C.A. KV4.3 Expression Modulates NaV1.5 Sodium Current. Front. Physiol. 2018, 9, 178. [Google Scholar] [CrossRef] [PubMed]
- Clatot, J.; Neyroud, N.; Cox, R.; Souil, C.; Huang, J.; Guicheney, P.; Antzelevitch, C. Inter-regulation of kv 4.3 and voltage-gated sodium channels underlies predisposition to cardiac and neuronal channelopathies. Int. J. Mol. Sci. 2020, 21, 5057. [Google Scholar] [CrossRef] [PubMed]
- Turner, R.W.; Zamponi, G.W. T-type channels buddy up. Pflugers Arch. Eur. J. Physiol. 2014, 466, 661–675. [Google Scholar] [CrossRef][Green Version]
- Anderson, D.; Mehaffey, W.H.; Iftinca, M.; Rehak, R.; Engbers, J.D.T.; Hameed, S.; Zamponi, G.W.; Turner, R.W. Regulation of neuronal activity by Cav3-Kv4 channel signaling complexes. Nat. Neurosci. 2010, 13, 333–337. [Google Scholar] [CrossRef]
- Alfaro-Ruíz, R.; Aguado, C.; Martín-Belmonte, A.; Moreno-Martínez, A.E.; Luján, R. Cellular and subcellular localisation of kv4-associated kchip proteins in the rat cerebellum. Int. J. Mol. Sci. 2020, 21, 6403. [Google Scholar] [CrossRef]
- Abbott, G.W. β subunits functionally differentiate human Kv4.3 potassium channel splice variants. Front. Physiol. 2017, 8, 1–15. [Google Scholar] [CrossRef]
- Melnyk, P.; Ehrlich, J.R.; Pourrier, M.; Villeneuve, L.; Cha, T.J.; Nattel, S. Comparison of ion channel distribution and expression in cardiomyocytes of canine pulmonary veins versus left atrium. Cardiovasc. Res. 2005, 65, 104–116. [Google Scholar] [CrossRef]
- McCrossan, Z.A.; Abbott, G.W. The MinK-related peptides. Neuropharmacology 2004, 47, 787–821. [Google Scholar] [CrossRef] [PubMed]
- Isom, L.L.; De Jongh, K.S.; Catterall, W.A. Auxiliary subunits of voltage-gated ion channels. Neuron 1994, 12, 1183–1194. [Google Scholar] [CrossRef]
- Decher, N.; Barth, A.S.; Gonzalez, T.; Steinmeyer, K.; Sanguinetti, M.C. Novel KChlP2 isoforms increase functional diversity of transient outward potassium currents. J. Physiol. 2004, 557, 761–772. [Google Scholar] [CrossRef] [PubMed]
- Pongs, O.; Schwarz, J.R. Ancillary subunits associated with voltage-dependent K+ channels. Physiol. Rev. 2010, 90, 755–796. [Google Scholar] [CrossRef] [PubMed]
- Bett, G.C.L.; Rasmusson, R.L. Modification of K+ channel-drug interactions by ancillary subunits. J. Physiol. 2008, 586, 929–950. [Google Scholar] [CrossRef] [PubMed]
- An, W.F.; Bowlby, M.R.; Betty, M.; Cao, J.; Ling, H.P.; Mendoza, G.; Hinson, J.W.; Mattsson, K.I.; Strassle, B.W.; Trimmer, J.S.; et al. Modulation of A-type potassium channels by a family of calcium sensors. Nature 2000, 403, 553–556. [Google Scholar] [CrossRef]
- Buxbaum, J.D.; Choi, E.K.; Luo, Y.; Lilliehook, C.; Crowley, A.C.; Merriam, D.E.; Wasco, W. Calsenilin: A calcium-binding protein that interacts with the presenilins and regulates the levels of a presenilin fragment. Nat. Med. 1998, 4, 1177–1181. [Google Scholar] [CrossRef]
- Carrión, A.M.; Link, W.A.; Ledo, F.; Mellström, B.; Naranjo, J.R. DREAM is a Ca2+-regulated transcriptional repressor. Nature 1999, 398, 80–84. [Google Scholar] [CrossRef]
- Morohashi, Y.; Hatano, N.; Ohya, S.; Takikawa, R.; Watabiki, T.; Takasugi, N.; Imaizumi, Y.; Tomita, T.; Iwatsubo, T. Molecular cloning and characterization of CALP/KChIP4, a novel EF-hand protein interacting with presenilin 2 and voltage-gated potassium channel subunit Kv4. J. Biol. Chem. 2002, 277, 14965–14975. [Google Scholar] [CrossRef]
- Kuo, H.C.; Cheng, C.F.; Clark, R.B.; Lin, J.J.C.; Lin, J.L.C.; Hoshijima, M.; Nguye-Trân, V.T.B.; Gu, Y.; Ikeda, Y.; Chu, P.H.; et al. A defect in the Kv channel-interacting protein 2 (KChIP2) gene leads to a complete loss of Ito and confers susceptibility to ventricular tachycardia. Cell 2001, 107, 801–813. [Google Scholar] [CrossRef]
- Ames, J.B.; Lim, S. Molecular structure and target recognition of neuronal calcium sensor proteins. Biochim. Biophys. Acta Gen. Subj. 2012, 1820, 1205–1213. [Google Scholar] [CrossRef] [PubMed]
- Burgoyne, R.D. Neuronal calcium sensor proteins: Generating diversity in neuronal Ca2+ signalling. Nat. Rev. Neurosci. 2007, 8, 182–193. [Google Scholar] [CrossRef] [PubMed]
- Bähring, R. Kv channel-interacting proteins as neuronal and non-neuronal calcium sensors. Channels 2018, 12, 187–200. [Google Scholar] [CrossRef] [PubMed]
- Naranjo, J.R.; Zhang, H.; Villar, D.; González, P.; Dopazo, X.M.; Morón-Oset, J.; Higueras, E.; Oliveros, J.C.; Arrabal, M.D.; Prieto, A.; et al. Activating transcription factor 6 derepression mediates neuroprotection in Huntington disease. J. Clin. Investig. 2016, 126, 627–638. [Google Scholar] [CrossRef]
- Scannevin, R.H.; Wang, K.W.; Jow, F.; Megules, J.; Kopsco, D.C.; Edris, W.; Carroll, K.C.; Lü, Q.; Xu, W.; Xu, Z.; et al. Two N-Terminal Domains of Kv4 K+ Channels Regulate Binding to and Modulation by KChIP1. Neuron 2004, 41, 587–598. [Google Scholar] [CrossRef]
- Bourne, Y.; Dannenberg, J.; Pollmann, V.; Marchot, P.; Pongs, O. Immunocytochemical Localization and Crystal Structure of Human Frequenin (Neuronal Calcium Sensor 1). J. Biol. Chem. 2001, 276, 11949–11955. [Google Scholar] [CrossRef]
- Flaherty, K.M.; Zozulya, S.; Stryer, L.; McKay, D.B. Three-dimensional structure of recoverin, a calcium sensor in vision. Cell 1993, 75, 709–716. [Google Scholar] [CrossRef]
- Vijay-Kumar, S.; Kumar, V.D. Crystal structure of recombinant bovine neurocalcin. Nat. Struct. Biol. 1999, 6, 80–88. [Google Scholar] [CrossRef]
- Babu, Y.S.; Bugg, C.E.; Cook, W.J. Structure of calmodulin refined at 2.2 Å resolution. J. Mol. Biol. 1988, 204, 191–204. [Google Scholar] [CrossRef]
- Yu, L.; Sun, C.; Mendoza, R.; Wang, J.; Matayoshi, E.D.; Hebert, E.; Pereda-Lopez, A.; Hajduk, P.J.; Olejniczak, E.T. Solution structure and calcium-binding properties of EF-hands 3 and 4 of calsenilin. Protein Sci. 2007, 16, 2502–2509. [Google Scholar] [CrossRef]
- Lusin, J.D.; Vanarotti, M.; Dace, A.; Li, C.; Valiveti, A.; Ames, J.B. NMR structure of DREAM: Implications for Ca2+-dependent DNA binding and protein dimerization (Biochemistry (2008) 47, 8, (2252–2265)). Biochemistry 2008, 47, 6088. [Google Scholar] [CrossRef][Green Version]
- Liang, P.; Wang, H.; Chen, H.; Cui, Y.; Gu, L.; Chai, J.; Wang, K.W. Structural insights into KChIP4a modulation of Kv4.3 inactivation. J. Biol. Chem. 2009, 284, 4960–4967. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Qian, Y.; Kunjilwar, K.; Pfaffinger, P.J.; Choe, S. Structural Insights into the Functional Interaction of KChIP1 with Shal-Type K+ Channels. Neuron 2004, 41, 573–586. [Google Scholar] [CrossRef]
- Kim, L.A.; Furst, J.; Gutierrez, D.; Butler, M.H.; Xu, S.; Goldstein, S.A.N.; Grigorieff, N. Three-Dimensional Structure of Ito: Kv4.2-KChIP2 Ion Channels by Electron Microscopy at 21 Å Resolution. Neuron 2004, 41, 513–519. [Google Scholar] [CrossRef]
- Pioletti, M.; Findeisen, F.; Hura, G.L.; Minor, D.L. Three-dimensional structure of the KChIP1-Kv4.3 T1 complex reveals a cross-shaped octamer. Nat. Struct. Mol. Biol. 2006, 13, 987–995. [Google Scholar] [CrossRef] [PubMed]
- Bähring, R.; Dannenberg, J.; Peters, H.C.; Leicher, T.; Pongs, O.; Isbrandt, D. Conserved Kv4 N-terminal Domain Critical for Effects of Kv Channel-interacting Protein 2.2 on Channel Expression and Gating. J. Biol. Chem. 2001, 276, 23888–23894. [Google Scholar] [CrossRef]
- Callsen, B.; Isbrandt, D.; Sauter, K.; Hartmann, L.S.; Pongs, O.; Bähring, R. Contribution of N- and C-terminal channel domains to Kv channel interacting proteins in a mammalian cell line. J. Physiol. 2005, 568, 397–412. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Yan, Y.; Liu, Q.; Huang, Y.; Shen, Y.; Chen, L.; Chen, Y.; Yang, Q.; Hao, Q.; Wang, K.W.; et al. Structural basis for modulation of Kv4 K+ channels by auxiliary KChIP subunits. Nat. Neurosci. 2007, 10, 32–39. [Google Scholar] [CrossRef]
- Bowlby, M.R.; Chanda, P.; Edris, W.; Hinson, J.; Jow, F.; Katz, A.H.; Kennedy, J.; Krishnamurthy, G.; Pitts, K.; Ryan, K.; et al. Identification and characterization of small molecule modulators of KChIP/Kv4 function. Bioorg. Med. Chem. 2005, 13, 6112–6119. [Google Scholar] [CrossRef]
- Holmqvist, M.H.; Cao, J.; Knoppers, M.H.; Jurman, M.E.; Distefano, P.S.; Rhodes, K.J.; Xie, Y.; An, W.F. Kinetic modulation of Kv4-mediated A-current by arachidonic acid is dependent on potassium channel interacting proteins. J. Neurosci. 2001, 21, 4154–4161. [Google Scholar] [CrossRef]
- Boland, L.M.; Drzewiecki, M.M.; Timoney, G.; Casey, E. Inhibitory effects of polyunsaturated fatty acids on Kv4/KChIP potassium channels. Am. J. Physiol. Cell Physiol. 2009, 296. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, W.G.; Miksovska, J. Application of ANS fluorescent probes to identify hydrophobic sites on the surface of DREAM. Biochim. Biophys. Acta Proteins Proteom. 2014, 1844, 1472–1480. [Google Scholar] [CrossRef] [PubMed]
- Lundby, A.; Jespersen, T.; Schmitt, N.; Grunnet, M.; Olesen, S.-P.; Cordeiro, J.M.; Calloe, K. Effect of the Ito activator NS5806 on cloned Kv4 channels depends on the accessory protein KChIP2. Br. J. Pharmacol. 2010, 160, 2028–2044. [Google Scholar] [CrossRef] [PubMed]
- Calloe, K.; Soltysinska, E.; Jespersen, T.; Lundby, A.; Antzelevitch, C.; Olesen, S.P.; Cordeiro, J.M. Differential effects of the transient outward K+ current activator NS5806 in the canine left ventricle. J. Mol. Cell. Cardiol. 2010, 48, 191–200. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Witzel, K.; Fischer, P.; Bähring, R. Hippocampal A-type current and Kv4.2 channel modulation by the sulfonylurea compound NS5806. Neuropharmacology 2012, 63, 1389–1403. [Google Scholar] [CrossRef]
- Gonzalez, W.G.; Pham, K.; Miksovska, J. Modulation of the voltage-gated potassium channel (Kv4.3) and the auxiliary protein (KChIP3) interactions by the current activator NS5806. J. Biol. Chem. 2014. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, H.; Wang, C.; Wang, Y.; Zou, R.; Shi, C.; Guan, B.; Gamper, N.; Xu, Y. Auxiliary subunits control biophysical properties and response to compound NS5806 of the Kv4 potassium channel complex. FASEB J. 2020, 34, 807–821. [Google Scholar] [CrossRef]
- Lainez, S.; Doray, A.; Hancox, J.C.; Cannell, M.B. Regulation of Kv4.3 and hERG potassium channels by KChIP2 isoforms and DPP6 and response to the dual K+ channel activator NS3623. Biochem. Pharmacol. 2018, 150, 120–130. [Google Scholar] [CrossRef]
- Lopez-Hurtado, A.; Peraza, D.A.; Cercos, P.; Lagartera, L.; Gonzalez, P.; Dopazo, X.M.; Herranz, R.; Gonzalez, T.; Martin-Martinez, M.; Mellström, B.; et al. Targeting the neuronal calcium sensor DREAM with small-molecules for Huntington’s disease treatment. Sci. Rep. 2019, 9, 7260. [Google Scholar] [CrossRef]
- Peraza, D.A.; Cercós, P.; Miaja, P.; Merinero, Y.G.; Lagartera, L.; Socuéllamos, P.G.; Izquierdo García, C.; Sánchez, S.A.; López-Hurtado, A.; Martín-Martínez, M.; et al. Identification of IQM-266, a Novel DREAM Ligand That Modulates KV4 Currents. Front. Mol. Neurosci. 2019, 12, 11. [Google Scholar] [CrossRef]
- Calloe, K.; Cordeiro, J.M.; Di Diego, J.M.; Hansen, R.S.; Grunnet, M.; Olesen, S.P.; Antzelevitch, C. A transient outward potassium current activator recapitulates the electrocardiographic manifestations of Brugada syndrome. Cardiovasc. Res. 2009, 81, 686–694. [Google Scholar] [CrossRef] [PubMed]
- Bennekou, P.; De Franceschi, L.; Pedersen, O.; Lian, L.; Asakura, T.; Evans, G.; Brugnara, C.; Christophersen, P. Treatment with NS3623, a novel Cl-conductance blocker, ameliorates erythrocyte dehydration in transgenic SAD mice: A possible new therapeutic approach for sickle cell disease. Blood 2001, 97, 1451–1457. [Google Scholar] [CrossRef] [PubMed]
- Hansen, R.S.; Diness, T.G.; Christ, T.; Wettwer, E.; Ravens, U.; Olesen, S.P.; Grunnet, M. Biophysical characterization of the new human ether-a-go-go-related gene channel opener NS3623 [N-(4-Bromo-2-(1H-tetrazol-5-yl)-phenyl)-N′-(3′-trifluoromethylphenyl)urea]. Mol. Pharmacol. 2006, 70, 1319–1329. [Google Scholar] [CrossRef] [PubMed]
- Calloe, K.; Di Diego, J.M.; Hansen, R.S.; Nagle, S.A.; Treat, J.A.; Cordeiro, J.M. A dual potassium channel activator improves repolarization reserve and normalizes ventricular action potentials. Biochem. Pharmacol. 2016, 108, 36–46. [Google Scholar] [CrossRef]
- Caballero, R.; de la Fuente, M.G.; Gómez, R.; Barana, A.; Amorós, I.; Dolz-Gaitón, P.; Osuna, L.; Almendral, J.; Atienza, F.; Fernández-Avilés, F.; et al. In Humans, Chronic Atrial Fibrillation Decreases the Transient Outward Current and Ultrarapid Component of the Delayed Rectifier Current Differentially on Each Atria and Increases the Slow Component of the Delayed Rectifier Current in Both. J. Am. Coll. Cardiol. 2010, 55, 2346–2354. [Google Scholar] [CrossRef]
| Ligand | KV4.x Channel | KChIPx | Effect | Ref. |
|---|---|---|---|---|
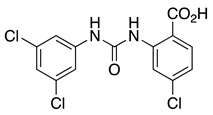 CL-888 (1) | KV4.2 KV4.3 | KChIP1 KChIP3 | Inhibition | [44,59] |
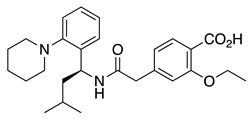 Repaglinide (2) | KV4.3 | KChIP3 | Inhibition | [44] |
| Polyunsaturated fatty acids (PUFAs) | Kv4.3 Kv4.2 | KChIP1 | Inhibition | [60,61,62] |
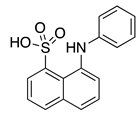 1,8-ANS (3) | --- | KChIP3 | Inhibition | [62] |
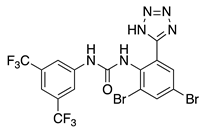 NS5806 (4) | KV4.3 | KChIP2KChIP3 | Inhibition/Activation | [63,64,65,66,67] |
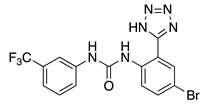 NS3623 (5) | KV4.3 | KChIP2 | Inhibition/Activation | [68] |
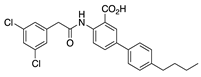 IQM-PC330 (6) | KV4.3 | KChIP3 | Inhibition | [69] |
 IQM-PC332 (7) | KV4.3 | KChIP3 | Inhibition | [69] |
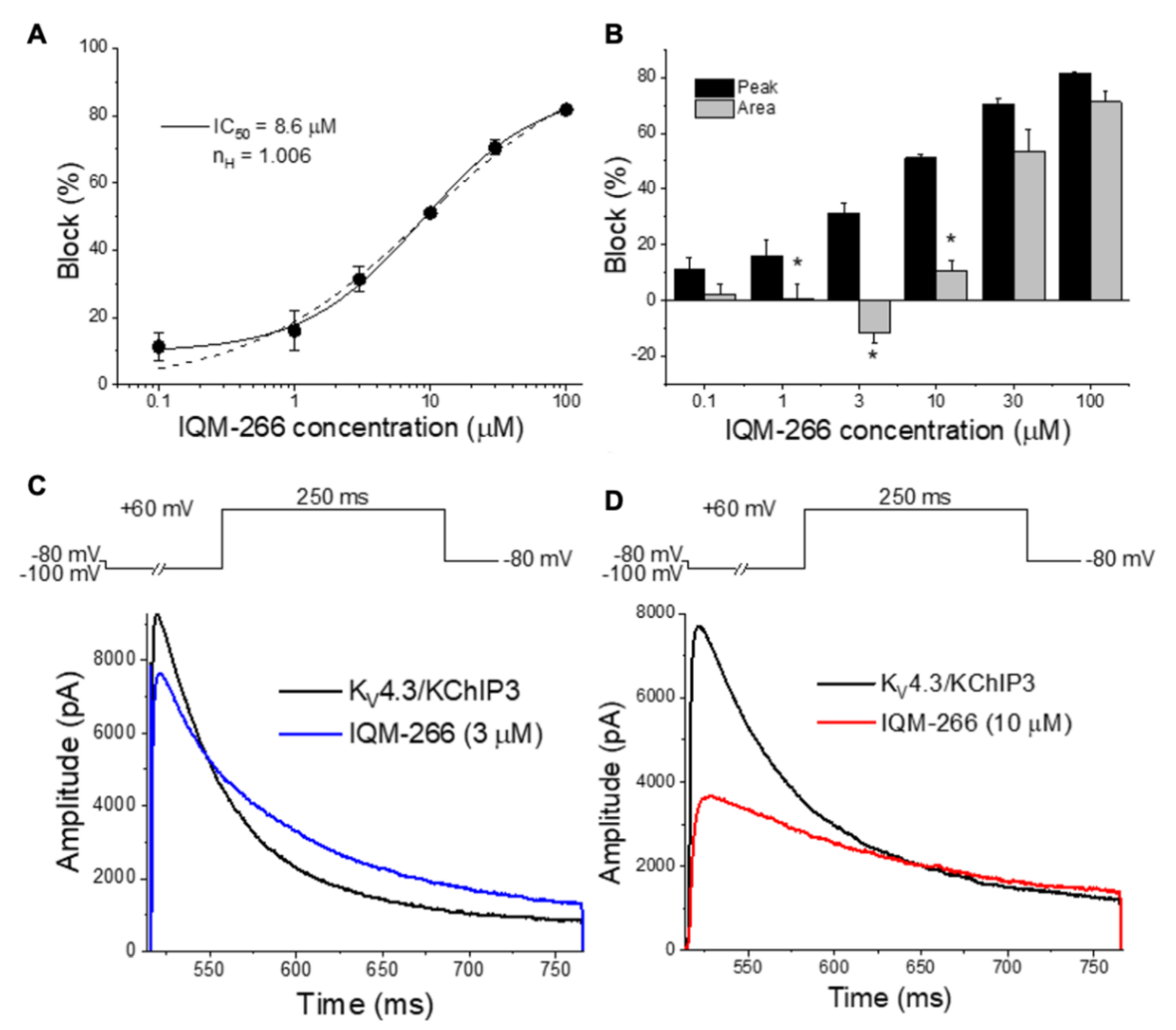
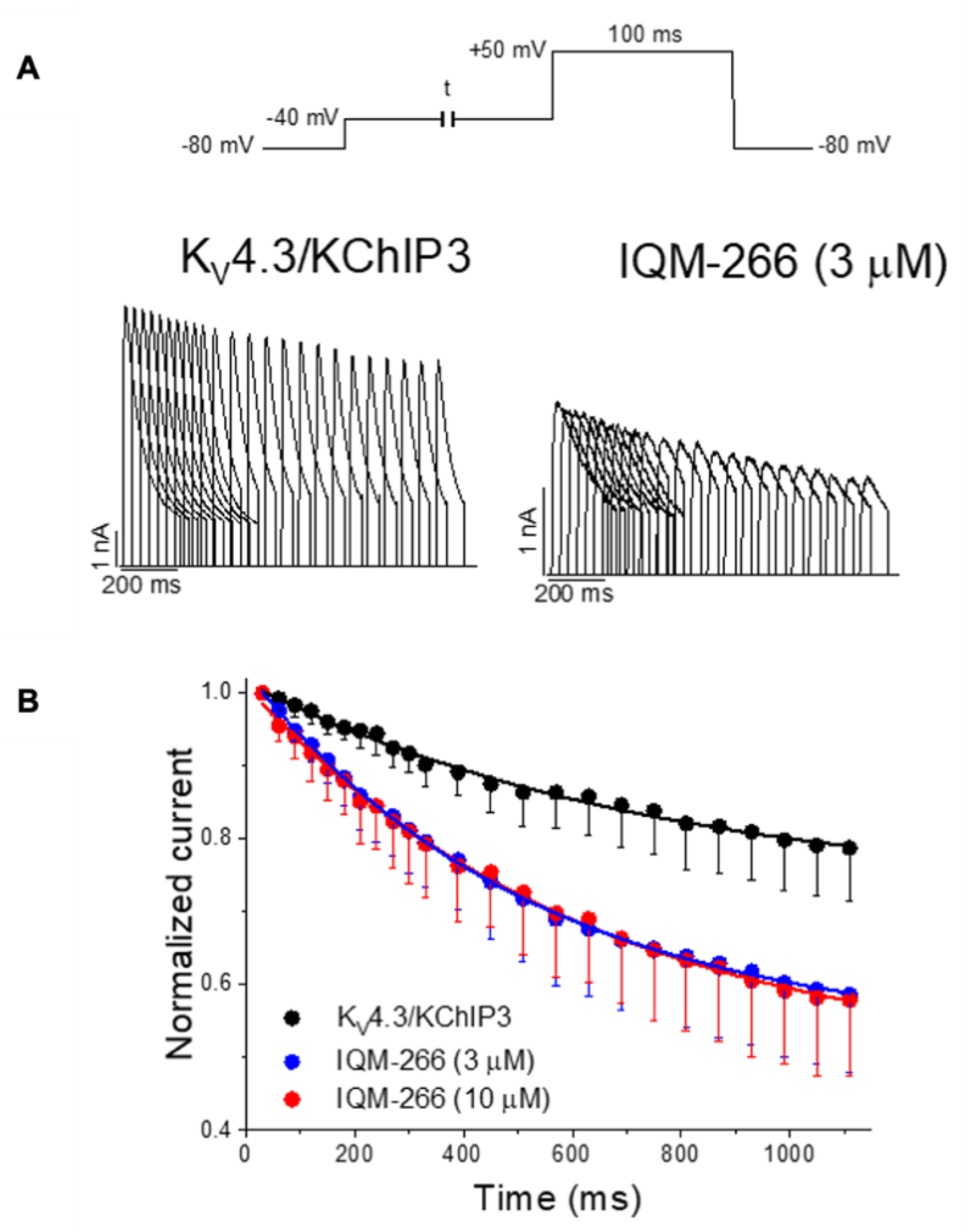
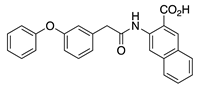 IQM-266 (8) | KV4.3 | KChIP3 | Inhibition/Activation | [70] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cercós, P.; Peraza, D.A.; Benito-Bueno, A.d.; Socuéllamos, P.G.; Aziz-Nignan, A.; Arrechaga-Estévez, D.; Beato, E.; Peña-Acevedo, E.; Albert, A.; González-Vera, J.A.; et al. Pharmacological Approaches for the Modulation of the Potassium Channel KV4.x and KChIPs. Int. J. Mol. Sci. 2021, 22, 1419. https://doi.org/10.3390/ijms22031419
Cercós P, Peraza DA, Benito-Bueno Ad, Socuéllamos PG, Aziz-Nignan A, Arrechaga-Estévez D, Beato E, Peña-Acevedo E, Albert A, González-Vera JA, et al. Pharmacological Approaches for the Modulation of the Potassium Channel KV4.x and KChIPs. International Journal of Molecular Sciences. 2021; 22(3):1419. https://doi.org/10.3390/ijms22031419
Chicago/Turabian StyleCercós, Pilar, Diego A. Peraza, Angela de Benito-Bueno, Paula G. Socuéllamos, Abdoul Aziz-Nignan, Dariel Arrechaga-Estévez, Escarle Beato, Emilio Peña-Acevedo, Armando Albert, Juan A. González-Vera, and et al. 2021. "Pharmacological Approaches for the Modulation of the Potassium Channel KV4.x and KChIPs" International Journal of Molecular Sciences 22, no. 3: 1419. https://doi.org/10.3390/ijms22031419
APA StyleCercós, P., Peraza, D. A., Benito-Bueno, A. d., Socuéllamos, P. G., Aziz-Nignan, A., Arrechaga-Estévez, D., Beato, E., Peña-Acevedo, E., Albert, A., González-Vera, J. A., Rodríguez, Y., Martín-Martínez, M., Valenzuela, C., & Gutiérrez-Rodríguez, M. (2021). Pharmacological Approaches for the Modulation of the Potassium Channel KV4.x and KChIPs. International Journal of Molecular Sciences, 22(3), 1419. https://doi.org/10.3390/ijms22031419







