Structural Features and Phylogenetic Implications of Four New Mitogenomes of Caliscelidae (Hemiptera: Fulgoromorpha)
Abstract
1. Introduction
2. Materials and Methods
2.1. Species Collection and Taxonomic Identification
2.2. DNA Extraction and Sequencing
2.3. Genome Assembly, Annotation, and Analysis
2.4. Phylogenetic Analyses
3. Results
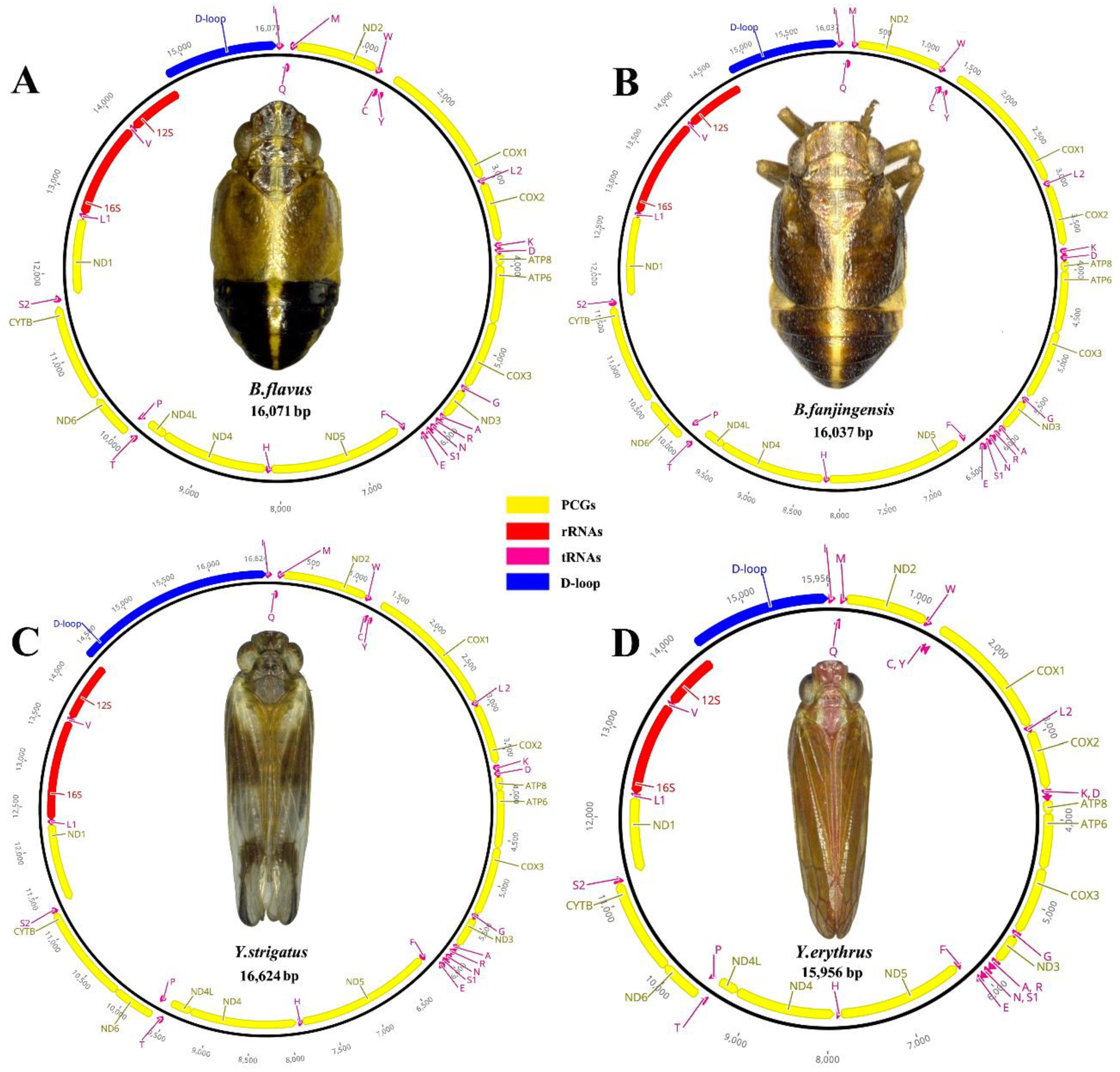
3.1. Genome Organization and Composition
3.2. PCGs and Codon Usage
3.3. Ribosomal RNA and Transfer RNA Genes
3.4. Overlapping and Intergenic Spacer Regions
3.5. AT-Rich Region
3.6. Phylogenetic Relationships
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gnezdilov, V.M. A Modern System of the Family Caliscelidae Amyot et Serville (Homoptera, Fulgoroidea). Зоологический Журнал/Entomol. Rev. 2013, 92, 1309–1311. [Google Scholar] [CrossRef]
- Bourgoin, T. FLOW (Fulgoromorpha Lists on The Web): A World Knowledge Base Dedicated to Fulgoromorpha. Version 8 Updated 1 October 2020. Available online: http://hemiptera-databases.org/flow/ (accessed on 17 October 2020).
- Chen, X.-S.; Zhang, Z.-G. Bambusicaliscelis, a New Bamboo-Feeding Planthopper Genus of Caliscelini (Hemiptera: Fulgoroidea: Caliscelidae: Caliscelinae), with Descriptions of Two New Species and Their Fifth-Instar Nymphs from Southwestern China. Ann. Entomol. Soc. Am. 2011, 104, 95–104. [Google Scholar] [CrossRef]
- Gong, N.; Yang, L.; Chen, X.-S. Youtuus, a new bamboo-feeding genus of the tribe Augilini with two new species from China (Hemiptera, Fulgoromorpha, Caliscelidae). ZooKeys 2018, 783, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.-Y.; Long, J.-K.; Chen, X.-S. Comparative analysis of the complete mitochondrial genomes of five Achilidae species (Hemiptera: Fulgoroidea) and other Fulgoroidea reveals conserved mitochondrial genome organization. PeerJ 2019, 7, e6659. [Google Scholar] [CrossRef] [PubMed]
- Urban, J.M.; Cryan, J.R. Evolution of the planthoppers (Insecta: Hemiptera: Fulgoroidea). Mol. Phylogenet. Evol. 2007, 42, 556–572. [Google Scholar] [CrossRef]
- Gwiazdowski, R.A.; Foottit, R.G.; Maw, H.E.L.; Hebert, P.D.N. The Hemiptera (Insecta) of Canada: Constructing a Reference Library of DNA Barcodes. PLoS ONE 2015, 10, e0125635. [Google Scholar] [CrossRef]
- Boore, J.L. Animal mitochondrial genomes. Nucleic Acids Res. 1999, 27, 1767–1780. [Google Scholar] [CrossRef]
- Wolstenholme, D.R. Animal Mitochondrial DNA: Structure and Evolution. Adv. Clin. Chem. 1992, 141, 173–216. [Google Scholar] [CrossRef]
- Cao, S.-Y.; Wu, X.-B.; Yan, P.; Hu, Y.-L.; Su, X.; Jiang, Z. Complete nucleotide sequences and gene organization of mitochondrial genome of Bufo gargarizans. Mitochondrion 2006, 6, 186–193. [Google Scholar] [CrossRef]
- E Cook, C.; Yue, Q.; Akam, M. Mitochondrial genomes suggest that hexapods and crustaceans are mutually paraphyletic. Proc. R. Soc. B Boil. Sci. 2005, 272, 1295–1304. [Google Scholar] [CrossRef]
- Ma, C.; Yang, P.; Jiang, F.; Chapuis, M.; Shali, Y.; Sword, G.A.; Kang, L. Mitochondrial genomes reveal the global phylogeography and dispersal routes of the migratory locust. Mol. Ecol. 2012, 21, 4344–4358. [Google Scholar] [CrossRef] [PubMed]
- Nardi, F.; Spinsanti, G.; Boore, J.L.; Carapelli, A.; Dallai, R.; Frati, F. Hexapod origins: Monophyletic or paraphyletic. Science 2003, 299, 1887–1889. [Google Scholar] [CrossRef] [PubMed]
- Saccone, C.; de Giorgi, C.; Gissi, C.; Pesole, G.; Reyes, A. Evolutionary genomics in Metazoa: The mitochondrial DNA as a model system. Gene 1999, 238, 195–209. [Google Scholar] [CrossRef]
- Timmermans, M.J.T.N.; Lees, D.C.; Simonsen, T.J. Towards a mitogenomic phylogeny of Lepidoptera. Mol. Phylogenet. Evol. 2014, 79, 169–178. [Google Scholar] [CrossRef]
- Zhang, P.; Zhou, H.; Liang, D.; Liu, Y.-F.; Chen, Y.-Q.; Qu, L.-H. The complete mitochondrial genome of a tree frog, Polypedates megacephalus (Amphibia: Anura: Rhacophoridae), and a novel gene organization in living amphibians. Gene 2005, 346, 133–143. [Google Scholar] [CrossRef]
- Gong, N.; Yang, L.; Chen, X.-S. Two new species of the bamboo-feeding genus Bambusicaliscelis Chen & Zhang, 2011 from China (Hemiptera, Fulgoromorpha, Caliscelidae). ZooKeys 2018, 776, 81–89. [Google Scholar] [CrossRef]
- Meng, G.; Li, Y.; Yang, C.; Liu, S. MitoZ: A toolkit for animal mitochondrial genome assembly, annotation and visualization. Nucleic Acids Res. 2019, 47, e63. [Google Scholar] [CrossRef]
- Bernt, M.; Donath, A.; Jühling, F.; Gärtner, F.; Florentz, C.; Fritzsch, G.; Pütz, J.; Middendorf, M.; Stadler, P.F. MITOS: Improved de novo metazoan mitochondrial genome annotation. Mol. Phylogenet. Evol. 2013, 69, 313–319. [Google Scholar] [CrossRef]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A Program for Improved Detection of Transfer RNA Genes in Genomic Sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Zhang, D.; Gao, F.; Li, W.-X.; Jakovlić, I.; Zou, H.; Zhang, J.; Wang, G.-T. PhyloSuite: An integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. bioRxiv 2018, 489088. [Google Scholar] [CrossRef]
- Bivand, R.; Anselin, L.; Berke, O.; Bernat, A.; Carvalho, M.; Chun, Y.; Dormann, C.F.; Dray, S.; Halbersma, R.; Lewin-Koh, N. spdep: Spatial dependence: Weighting schemes, statistics and models. GeoHealth 2019, 4, e2020GH000276. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Perna, N.T.; Kocher, T.D. Patterns of nucleotide composition at fourfold degenerate sites of animal mitochondrial genomes. J. Mol. Evol. 1995, 41, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 1999, 27, 573–580. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Vaidya, G.; Lohman, D.J.; Meier, R. SequenceMatrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 2011, 27, 171–180. [Google Scholar] [CrossRef]
- Lanfear, R.; Frandsen, P.B.; Wright, A.M.; Senfeld, T.; Calcott, B. PartitionFinder 2: New Methods for Selecting Partitioned Models of Evolution for Molecular and Morphological Phylogenetic Analyses. Mol. Biol. Evol. 2017, 34, 772–773. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2014, 32, 268–274. [Google Scholar] [CrossRef]
- Minh, B.Q.; Nguyen, M.A.T.; von Haeseler, A. Ultrafast Approximation for Phylogenetic Bootstrap. Mol. Biol. Evol. 2013, 30, 1188–1195. [Google Scholar] [CrossRef]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed]
- Mousavi, S.A.; Österman, J.; Wahlberg, N.; Nesme, X.; Lavire, C.; Vial, L.; Paulin, L.; de Lajudie, P.; Lindström, K. Phy-logeny of the Rhizobium-Allorhizobium-Agrobacterium clade supports the delineation of Neorhizobium gen. nov. Sys-Temat. Appl. Microbiol. 2014, 37, 208–215. [Google Scholar]
- Dai, L.-S.; Zhu, B.-J.; Qian, C.; Zhang, C.-F.; Li, J.; Wang, L.; Wei, G.-Q.; Liu, C. The complete mitochondrial genome of the diamondback moth, Plutella xylostella (Lepidoptera: Plutellidae). Mitochondrial DNA 2016, 27, 1512–1513. [Google Scholar] [CrossRef] [PubMed]
- Dai, L.-S.; Zhu, B.-J.; Zhao, Y.; Zhang, C.-F.; Liu, C.-L. Comparative Mitochondrial Genome Analysis of Eligma narcissus and other Lepidopteran Insects Reveals Conserved Mitochondrial Genome Organization and Phylogenetic Relationships. Sci. Rep. 2016, 6, 26387. [Google Scholar] [CrossRef]
- Kim, I.; Lee, E.M.; Seol, K.Y.; Yun, E.Y.; Lee, Y.B.; Hwang, J.S.; Jin, B.R. The mitochondrial genome of the Korean hairstreak, Coreana raphaelis (Lepidoptera: Lycaenidae). Insect Mol. Biol. 2006, 15, 217–225. [Google Scholar] [CrossRef]
- Boore, J.L. Complete Mitochondrial Genome Sequence of the Polychaete Annelid Platynereis dumerilii. Mol. Biol. Evol. 2001, 18, 1413–1416. [Google Scholar] [CrossRef]
- Ojala, D.; Montoya, J.; Attardi, G. tRNA punctuation model of RNA processing in human mitochondria. Nat. Cell Biol. 1981, 290, 470–474. [Google Scholar] [CrossRef]
- Muir, F. XLVIII—On the classification of the Fulgoroidea. J. Nat. Hist. 1930, 6, 461–478. [Google Scholar] [CrossRef]
- Asche, M. Preliminary thoughts on the phylogeny of Fulgoromorpha (Homoptera: Auchenorrhyncha). In Proceedings of the 6th Auchenorrhyncha, Turin, Italy, 7–11 September 1987; pp. 47–53. [Google Scholar]
- Bourgoin, T.; Steffen-Campbell, J.; Campbell, B. Molecular Phylogeny of Fulgoromorpha (Insecta, Hemiptera, Archaeorrhyncha). The Enigmatic Tettigometridae: Evolutionary Affiliations and Historical Biogeography. Cladistics 1997, 13, 207–224. [Google Scholar] [CrossRef]
- Yeh, W.B.; Yang, C.T.; Hui, C.F. A molecular phylogeny of planthoppers (Hemiptera: Fulgoroidea) inferred from mito-chondrial 16S rDNA sequences. Zool. Stud. 2005, 44, 519–535. [Google Scholar]
- Gnezdilov, V.M. Notes on the phylogenetic relationships of planthoppers of the family issidae (Hemiptera, Fulgoroidea) of the Western Palaearctic fauna, with descriptions of two new genera. Entomol. Rev. 2016, 96, 332–347. [Google Scholar] [CrossRef]
- Sheffield, N.C.; Song, H.; Cameron, S.L.; Whiting, M.F. A Comparative Analysis of Mitochondrial Genomes in Coleoptera (Arthropoda: Insecta) and Genome Descriptions of Six New Beetles. Mol. Biol. Evol. 2008, 25, 2499–2509. [Google Scholar] [CrossRef]
- Cao, S.-S.; Du, Y.-Z. Characterization of the complete mitochondrial genome of Chilo auricilius and comparison with three other rice stem borers. Gene 2014, 548, 270–276. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.-J.; Zhu, W.-C.; Rong, X.; Liu, J.; Ding, X.-L.; Hong, X.-Y. The complete mitochondrial genome sequence of Sogatella furcifera (Horváth) and a comparative mitogenomic analysis of three predominant rice planthoppers. Gene 2014, 533, 100–109. [Google Scholar] [CrossRef] [PubMed]
- Song, N.; Liang, A.-P. A Preliminary Molecular Phylogeny of Planthoppers (Hemiptera: Fulgoroidea) Based on Nuclear and Mitochondrial DNA Sequences. PLoS ONE 2013, 8, e58400. [Google Scholar] [CrossRef]



| Gene | B. flavus | B. fanjingensis | Y. strigatus | Y. erythrus | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Size | Anticodons | Int. | Size | Anticodons | Int. | Size | Anticodons | Int. | Size | Anticodons | Int. | |
| Start/Stop | Start/Stop | Start/Stop | Start/Stop | |||||||||
| trnI | 67 | ATC | 67 | ATC | 65 | ATC | 65 | ATC | ||||
| trnQ | 69 | CAA | 7 | 69 | CAA | 9 | 69 | CAA | 7 | 69 | CAA | 7 |
| trnM | 67 | ATG | 13 | 66 | ATG | 20 | 64 | ATG | −1 | 64 | ATG | −1 |
| nad2 | 966 | ATT/TAA | 0 | 966 | ATT/TAA | 0 | 966 | ATT/TAA | 0 | 966 | ATT/TAA | 0 |
| trnW | 65 | TGA | −1 | 63 | TGA | 0 | 68 | TGA | −1 | 68 | TGA | −1 |
| trnC | 74 | TGC | −8 | 73 | TGC | −8 | 57 | TGC | −8 | 58 | TGC | −8 |
| trnY | 67 | TAC | 12 | 66 | TAC | 15 | 63 | TAC | 4 | 62 | TAC | 6 |
| cox1 | 1539 | ATG/TAA | 23 | 1536 | ATG/TAA | 23 | 1534 | ATG/T | 9 | 1534 | ATG/T | 10 |
| trnL2 | 70 | TTA | 1 | 70 | TTA | 8 | 66 | TTA | 0 | 68 | TTA | 0 |
| cox2 | 675 | ATT/TAA | 0 | 675 | ATT/TAA | 0 | 675 | ATA/TAA | 0 | 675 | ATA/TAA | 0 |
| trnK | 70 | AAG | 20 | 71 | AAG | 40 | 71 | AAG | 20 | 71 | AAG | 8 |
| trnD | 69 | GAC | 2 | 70 | GAC | 0 | 66 | GAC | 0 | 65 | GAC | 0 |
| atp8 | 134 | ATT/TAA | 0 | 119 | ATT/TAA | 2 | 156 | ATT/TAA | 0 | 156 | ATT/TAA | 0 |
| atp6 | 652 | ATA/T | −7 | 655 | ATA/T | −7 | 652 | ATG/T | −7 | 652 | ATG/T | −7 |
| cox3 | 783 | ATG/TAA | 0 | 783 | ATG/TAA | 0 | 781 | ATG/T | 0 | 781 | ATG/T | 0 |
| trnG | 65 | GGA | 7 | 66 | GGA | 26 | 66 | GGA | 0 | 63 | GGA | −1 |
| nad3 | 348 | ATA/TAA | 0 | 348 | ATA/TAA | 0 | 348 | ATT/TAG | 0 | 348 | ATT/TAA | −1 |
| trnA | 69 | GCA | −1 | 68 | GCA | 6 | 64 | GCA | −2 | 64 | GCG | 0 |
| trnR | 66 | CGA | 3 | 65 | CGA | 12 | 63 | CGA | 3 | 59 | CGA | 4 |
| trnN | 71 | AAC | 23 | 71 | AAC | 3 | 67 | AAC | 4 | 67 | AAC | 8 |
| trnS1 | 60 | AGC | −1 | 61 | AGC | −1 | 59 | AGC | −1 | 58 | AGC | −1 |
| trnE | 67 | GAA | 18 | 71 | GAA | 10 | 62 | GAA | −1 | 62 | GAA | −1 |
| trnF | 65 | TTC | 37 | 62 | TTC | 34 | 63 | TTC | 0 | 64 | TTC | 0 |
| nad5 | 1713 | GTG/TAA | 48 | 1671 | GTG/TAA | 58 | 1678 | GTG/T | 4 | 1654 | ATG/T | 3 |
| trnH | 72 | CAC | 0 | 70 | CAC | 0 | 60 | CAC | 0 | 62 | CAC | 0 |
| nad4 | 1366 | ATG/T | 1 | 1321 | ATG/T | 1 | 1317 | ATG/TAA | 21 | 1308 | ATA/TAG | 27 |
| nad4l | 270 | ATG/TAA | −7 | 270 | ATG/TAA | −7 | 273 | ATG/TAA | −7 | 273 | ATG/TAA | 1 |
| trnT | 69 | ACA | 2 | 69 | ACA | 38 | 67 | ACA | 2 | 67 | ACA | 2 |
| trnP | 63 | CCA | 4 | 66 | CCA | 26 | 63 | CCA | 9 | 66 | CCA | 4 |
| nad6 | 504 | ATA/TAA | 1 | 501 | ATA/TAA | 1 | 492 | ATA/TAA | 1 | 492 | ATA/TAA | 1 |
| cob | 1140 | ATG/TAA | 32 | 1134 | ATG/TAA | 33 | 1119 | ATG/TAA | −8 | 1117 | ATG/T | −8 |
| trnS2 | 68 | TCA | 48 | 70 | TCA | 11 | 62 | TCA | 2 | 64 | TCA | 0 |
| nad1 | 949 | ATG/T | −4 | 963 | ATG/TAA | 16 | 942 | ATG/TAA | 6 | 945 | ATG/TAA | −1 |
| trnL1 | 67 | CTA | 3 | 63 | CTA | 1 | 65 | CTA | 1 | 64 | CTA | 1 |
| rrnL | 1227 | - | 0 | 1221 | - | 0 | 1222 | - | 0 | 1222 | - | 0 |
| trnV | 56 | GTA | 0 | 61 | GTA | 0 | 58 | GTA | 0 | 56 | GTA | 0 |
| rrnS | 717 | - | 0 | 738 | - | 0 | 732 | - | 0 | 732 | - | 0 |
| CR | 1318 | 0 | 1230 | 0 | 2288 | 0 | 1609 | 0 | ||||
| Total | 16053 | 15979 | 16640 | 15922 | ||||||||
| Regions | Size (bp) | T(U) | C | A | G | A+T (%) | G+C (%) | AT Skewness | GC Skewness |
|---|---|---|---|---|---|---|---|---|---|
| Full Genome | |||||||||
| B. flavus | 16053 | 32 | 13.2 | 47.5 | 7.2 | 79.5 | 20.4 | 0.194 | −0.293 |
| B. fanjingensis | 15979 | 32.2 | 12.6 | 48.1 | 7 | 80.3 | 19.6 | 0.197 | −0.288 |
| Y. strigatus | 16640 | 33 | 12.5 | 46.8 | 7.7 | 79.8 | 20.2 | 0.173 | −0.238 |
| Y. erythrus | 15922 | 33.3 | 12.7 | 46.4 | 7.6 | 79.7 | 20.3 | 0.164 | −0.251 |
| 13 PCGs | |||||||||
| B. flavus | 11039 | 44.5 | 11.2 | 33.9 | 10.3 | 78.4 | 21.5 | −0.135 | −0.042 |
| B. fanjingensis | 10942 | 44.5 | 11.2 | 34.3 | 9.9 | 78.8 | 21.1 | −0.129 | −0.062 |
| Y. strigatus | 10933 | 43.9 | 11.1 | 35.2 | 9.8 | 79.1 | 20.9 | −0.11 | −0.062 |
| Y. erythrus | 10901 | 44.4 | 11.0 | 34.6 | 10.0 | 79 | 21 | −0.124 | −0.048 |
| 2 rRNAs | |||||||||
| B. flavus | 1944 | 47.7 | 6.6 | 32.8 | 12.9 | 80.5 | 19.5 | −0.185 | 0.323 |
| B. fanjingensis | 1959 | 46.7 | 6.5 | 34 | 12.8 | 80.7 | 19.3 | −0.157 | 0.328 |
| Y. strigatus | 1954 | 48.7 | 6.7 | 32.2 | 12.4 | 80.9 | 19.1 | −0.203 | 0.298 |
| Y. erythrus | 1954 | 47.5 | 6.9 | 33.5 | 12.1 | 81 | 19 | −0.172 | 0.278 |
| 22 tRNAs | |||||||||
| B. flavus | 1476 | 37.8 | 9.1 | 42.1 | 11.0 | 79.9 | 20.1 | 0.054 | 0.095 |
| B. fanjingensis | 1478 | 37.1 | 8.9 | 43.4 | 10.7 | 80.5 | 19.6 | 0.078 | 0.092 |
| Y. strigatus | 1408 | 37.4 | 8.8 | 42.0 | 11.8 | 79.4 | 20.6 | 0.058 | 0.146 |
| Y. erythrus | 1406 | 38.0 | 8.6 | 41.1 | 12.3 | 79.1 | 20.9 | 0.039 | 0.177 |
| AT-rich regions | |||||||||
| B. flavus | 1318 | 39.3 | 11.1 | 44.9 | 4.7 | 84.2 | 15.8 | 0.067 | −0.405 |
| B. fanjingensis | 1230 | 41.5 | 7.4 | 47.6 | 3.5 | 89.1 | 10.9 | 0.068 | −0.358 |
| Y. strigatus | 2288 | 38.8 | 10.3 | 43.5 | 7.3 | 82.3 | 17.7 | 0.057 | −0.169 |
| Y. erythrus | 1609 | 40.9 | 11.5 | 41.8 | 5.8 | 82.7 | 17.3 | 0.011 | −0.329 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gong, N.; Yang, L.; Chen, X.-S. Structural Features and Phylogenetic Implications of Four New Mitogenomes of Caliscelidae (Hemiptera: Fulgoromorpha). Int. J. Mol. Sci. 2021, 22, 1348. https://doi.org/10.3390/ijms22031348
Gong N, Yang L, Chen X-S. Structural Features and Phylogenetic Implications of Four New Mitogenomes of Caliscelidae (Hemiptera: Fulgoromorpha). International Journal of Molecular Sciences. 2021; 22(3):1348. https://doi.org/10.3390/ijms22031348
Chicago/Turabian StyleGong, Nian, Lin Yang, and Xiang-Sheng Chen. 2021. "Structural Features and Phylogenetic Implications of Four New Mitogenomes of Caliscelidae (Hemiptera: Fulgoromorpha)" International Journal of Molecular Sciences 22, no. 3: 1348. https://doi.org/10.3390/ijms22031348
APA StyleGong, N., Yang, L., & Chen, X.-S. (2021). Structural Features and Phylogenetic Implications of Four New Mitogenomes of Caliscelidae (Hemiptera: Fulgoromorpha). International Journal of Molecular Sciences, 22(3), 1348. https://doi.org/10.3390/ijms22031348






