Regulation of p53 and Cancer Signaling by Heat Shock Protein 40/J-Domain Protein Family Members
Abstract
1. Introduction
2. Effects of HSP40/JDPs on p53 Activity and Cancer Signaling
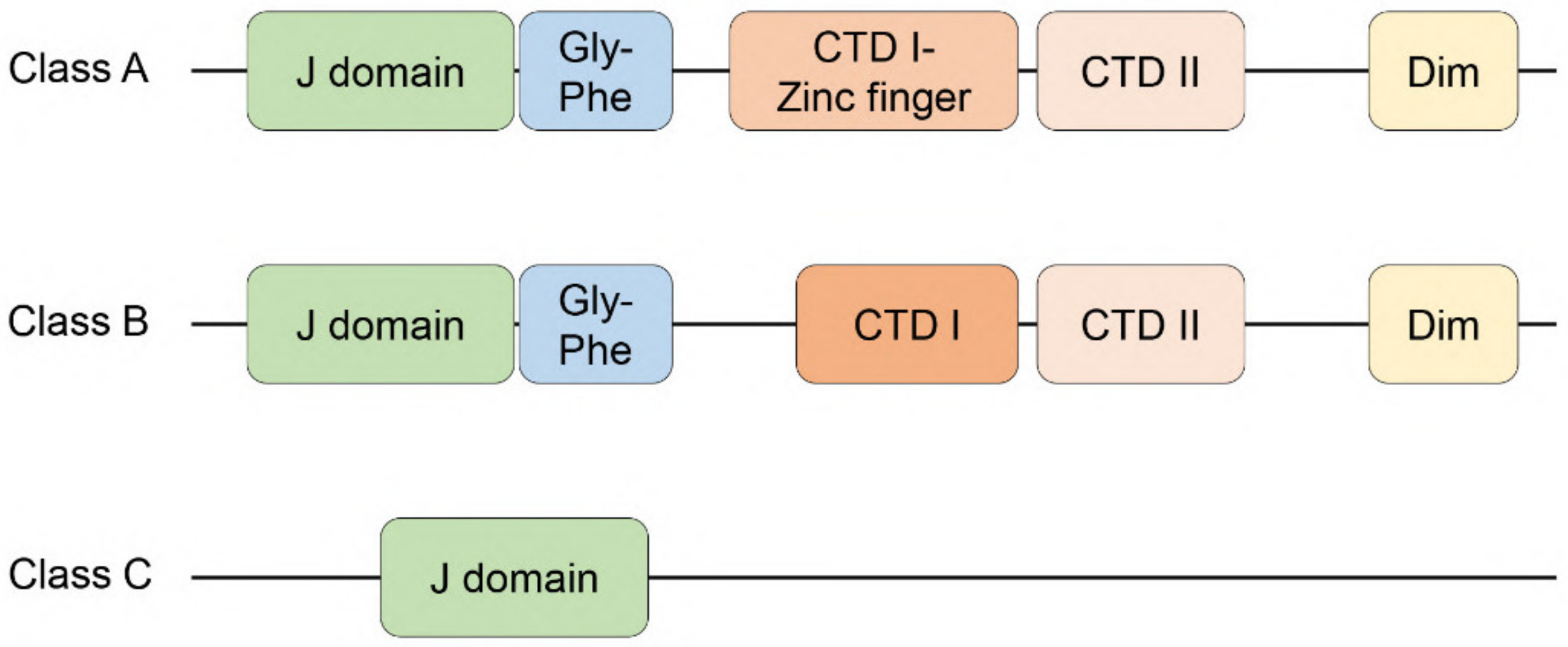
2.1. Class A HSP40/JDPs (DNAJA Proteins)
2.1.1. DNAJA1/HDJ2
- DNAJA1/HDJ2 and cancer
- DNAJA1/HDJ2 and p53
- DNAJA1/HDJ2 as a cancer therapeutic target
2.1.2. DNAJA3 (Tid1: Tumorous Imaginal Disc 1)
- DNAJA3/Tid1 and cancer
- DNAJA3/Tid1 and p53
2.2. Class B HSP40/JDPs (DNAJB Proteins)
2.2.1. DNAJB1/HDJ1
- DNAJB1/HDJ1 and cancer
- DNAJB1/HDJ1 and p53
2.2.2. DNAJB9/MDG1/ERdj4
- DNAJB9/MDG1/ERdj41 and cancer
- DNAJB9/MDG1/ERdj41 and p53
2.2.3. Other DNAJB Members and Cancer
- DNAJB4/HLJ1
- DNAJB6/MRJ
- DNAJB8
- DNAJB11/ERdj3/HEDJ
- DNAJB12
2.3. Class C HSP40/JDPs (DNAJC Proteins)
2.3.1. DNAJC2/ZRF1
- DNAJC2/ZRF1 and cancer
- DNAJC2/ZRF1 and p53
2.3.2. DNAJC7/TPR2
- DNAJC7/TPR2 and cancer
- DNAJC7/TPR2 and p53
2.3.3. DNAJC9
- DNAJC9 and cancer
- DNAJC9 and p53
2.3.4. Other DNAJC Members and Cancer
- DNAJC12
- DNAJC15/MCJ
- DNAJC25
3. Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vogelstein, B.; Lane, D.; Levine, A.J. Surfing the p53 network. Nature 2000, 408, 307–310. [Google Scholar] [CrossRef] [PubMed]
- Lane, D.; Levine, A. p53 Research: The past thirty years and the next thirty years. Cold Spring Harb. Perspect. Biol. 2010, 2, a000893. [Google Scholar] [CrossRef]
- Bieging, K.T.; Mello, S.S.; Attardi, L.D. Unravelling mechanisms of p53-mediated tumour suppression. Nat. Rev. Cancer 2014, 14, 359–370. [Google Scholar] [CrossRef] [PubMed]
- Deng, C.; Zhang, P.; Harper, J.W.; Elledge, S.J.; Leder, P. Mice lacking p21CIP1/WAF1 undergo normal development, but are defective in G1 checkpoint control. Cell 1995, 82, 675–684. [Google Scholar] [CrossRef]
- Hollander, M.C.; Sheikh, M.S.; Bulavin, D.V.; Lundgren, K.; Augeri-Henmueller, L.; Shehee, R.; Molinaro, T.A.; Kim, K.E.; Tolosa, E.; Ashwell, J.D.; et al. Genomic instability in Gadd45a-deficient mice. Nat. Genet. 1999, 23, 176–184. [Google Scholar] [CrossRef]
- Fridman, J.S.; Lowe, S.W. Control of apoptosis by p53. Oncogene 2003, 22, 9030–9040. [Google Scholar] [CrossRef] [PubMed]
- Rufini, A.; Tucci, P.; Celardo, I.; Melino, G. Senescence and aging: The critical roles of p53. Oncogene 2013, 32, 5129–5143. [Google Scholar] [CrossRef]
- Green, D.R.; Kroemer, G. Cytoplasmic functions of the tumour suppressor p53. Nature 2009, 458, 1127–1130. [Google Scholar] [CrossRef]
- Liang, S.H.; Clarke, M.F. Regulation of p53 localization. Eur. J. Biochem. 2001, 268, 2779–2783. [Google Scholar] [CrossRef]
- Giorgi, C.; Bonora, M.; Sorrentino, G.; Missiroli, S.; Poletti, F.; Suski, J.M.; Galindo Ramirez, F.; Rizzuto, R.; Di Virgilio, F.; Zito, E.; et al. p53 at the endoplasmic reticulum regulates apoptosis in a Ca2+-dependent manner. Proc. Natl. Acad. Sci. USA 2015, 112, 1779–1784. [Google Scholar] [CrossRef]
- Marchenko, N.D.; Zaika, A.; Moll, U.M. Death signal-induced localization of p53 protein to mitochondria. A potential role in apoptotic signaling. J. Biol. Chem. 2000, 275, 16202–16212. [Google Scholar] [CrossRef] [PubMed]
- Mihara, M.; Erster, S.; Zaika, A.; Petrenko, O.; Chittenden, T.; Pancoska, P.; Moll, U.M. p53 has a direct apoptogenic role at the mitochondria. Mol. Cell 2003, 11, 577–590. [Google Scholar] [CrossRef]
- Baugh, E.H.; Ke, H.; Levine, A.J.; Bonneau, R.A.; Chan, C.S. Why are there hotspot mutations in the TP53 gene in human cancers? Cell Death Differ. 2018, 25, 154–160. [Google Scholar] [CrossRef]
- Parrales, A.; Iwakuma, T. Targeting Oncogenic Mutant p53 for Cancer Therapy. Front. Oncol. 2015, 5, 288. [Google Scholar] [CrossRef] [PubMed]
- Walerych, D.; Lisek, K.; Del Sal, G. Mutant p53: One, No One, and One Hundred Thousand. Front. Oncol. 2015, 5, 289. [Google Scholar] [CrossRef]
- Kim, M.P.; Zhang, Y.; Lozano, G. Mutant p53: Multiple Mechanisms Define Biologic Activity in Cancer. Front. Oncol. 2015, 5, 249. [Google Scholar] [CrossRef]
- Lang, G.A.; Iwakuma, T.; Suh, Y.A.; Liu, G.; Rao, V.A.; Parant, J.M.; Valentin-Vega, Y.A.; Terzian, T.; Caldwell, L.C.; Strong, L.C.; et al. Gain of function of a p53 hot spot mutation in a mouse model of Li-Fraumeni syndrome. Cell 2004, 119, 861–872. [Google Scholar] [CrossRef]
- Adorno, M.; Cordenonsi, M.; Montagner, M.; Dupont, S.; Wong, C.; Hann, B.; Solari, A.; Bobisse, S.; Rondina, M.B.; Guzzardo, V.; et al. A Mutant-p53/Smad complex opposes p63 to empower TGFbeta-induced metastasis. Cell 2009, 137, 87–98. [Google Scholar] [CrossRef]
- Muller, P.A.; Caswell, P.T.; Doyle, B.; Iwanicki, M.P.; Tan, E.H.; Karim, S.; Lukashchuk, N.; Gillespie, D.A.; Ludwig, R.L.; Gosselin, P.; et al. Mutant p53 drives invasion by promoting integrin recycling. Cell 2009, 139, 1327–1341. [Google Scholar] [CrossRef]
- Parrales, A.; Thoenen, E.; Iwakuma, T. The interplay between mutant p53 and the mevalonate pathway. Cell Death Differ. 2018, 25, 460–470. [Google Scholar] [CrossRef]
- Yamamoto, S.; Iwakuma, T. Regulators of Oncogenic Mutant TP53 Gain of Function. Cancers 2018, 11, 4. [Google Scholar] [CrossRef] [PubMed]
- Terzian, T.; Suh, Y.A.; Iwakuma, T.; Post, S.M.; Neumann, M.; Lang, G.A.; Van Pelt, C.S.; Lozano, G. The inherent instability of mutant p53 is alleviated by Mdm2 or p16INK4a loss. Genes Dev. 2008, 22, 1337–1344. [Google Scholar] [CrossRef]
- Parrales, A.; Ranjan, A.; Iyer, S.V.; Padhye, S.; Weir, S.J.; Roy, A.; Iwakuma, T. DNAJA1 controls the fate of misfolded mutant p53 through the mevalonate pathway. Nat. Cell Biol. 2016, 18, 1233–1243. [Google Scholar] [CrossRef]
- Tracz-Gaszewska, Z.; Klimczak, M.; Biecek, P.; Herok, M.; Kosinski, M.; Olszewski, M.B.; Czerwinska, P.; Wiech, M.; Wiznerowicz, M.; Zylicz, A.; et al. Molecular chaperones in the acquisition of cancer cell chemoresistance with mutated TP53 and MDM2 up-regulation. Oncotarget 2017, 8, 82123–82143. [Google Scholar] [CrossRef]
- Wawrzynow, B.; Zylicz, A.; Zylicz, M. Chaperoning the guardian of the genome. The two-faced role of molecular chaperones in p53 tumor suppressor action. Biochim. Biophys. Acta (BBA) Rev. Cancer 2018, 1869, 161–174. [Google Scholar] [CrossRef] [PubMed]
- Lukashchuk, N.; Vousden, K.H. Ubiquitination and degradation of mutant p53. Mol. Cell Biol. 2007, 27, 8284–8295. [Google Scholar] [CrossRef] [PubMed]
- Jolly, C.; Morimoto, R.I. Role of the heat shock response and molecular chaperones in oncogenesis and cell death. J. Natl. Cancer Inst. 2000, 92, 1564–1572. [Google Scholar] [CrossRef]
- Saibil, H. Chaperone machines for protein folding, unfolding and disaggregation. Nat. Rev. Mol. Cell Biol. 2013, 14, 630–642. [Google Scholar] [CrossRef] [PubMed]
- Doyle, S.M.; Genest, O.; Wickner, S. Protein rescue from aggregates by powerful molecular chaperone machines. Nat. Rev. Mol. Cell Biol. 2013, 14, 617–629. [Google Scholar] [CrossRef]
- Hartl, F.U.; Bracher, A.; Hayer-Hartl, M. Molecular chaperones in protein folding and proteostasis. Nature 2011, 475, 324–332. [Google Scholar] [CrossRef]
- Rosenzweig, R.; Nillegoda, N.B.; Mayer, M.P.; Bukau, B. The Hsp70 chaperone network. Nat. Rev. Mol. Cell Biol. 2019, 20, 665–680. [Google Scholar] [CrossRef]
- Milarski, K.L.; Morimoto, R.I. Expression of human HSP70 during the synthetic phase of the cell cycle. Proc. Natl. Acad. Sci. USA 1986, 83, 9517–9521. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, E.; Parcellier, A.; Gurbuxani, S.; Cande, C.; Hammann, A.; Morales, M.C.; Hunt, C.R.; Dix, D.J.; Kroemer, R.T.; Giordanetto, F.; et al. Chemosensitization by a non-apoptogenic heat shock protein 70-binding apoptosis-inducing factor mutant. Cancer Res. 2003, 63, 8233–8240. [Google Scholar] [PubMed]
- Aligue, R.; Akhavan-Niak, H.; Russell, P. A role for Hsp90 in cell cycle control: Wee1 tyrosine kinase activity requires interaction with Hsp90. EMBO J. 1994, 13, 6099–6106. [Google Scholar] [CrossRef] [PubMed]
- Burrows, F.; Zhang, H.; Kamal, A. Hsp90 activation and cell cycle regulation. Cell Cycle 2004, 3, 1530–1536. [Google Scholar] [CrossRef]
- Minami, Y.; Hohfeld, J.; Ohtsuka, K.; Hartl, F.U. Regulation of the heat-shock protein 70 reaction cycle by the mammalian DnaJ homolog, Hsp40. J. Biol. Chem. 1996, 271, 19617–19624. [Google Scholar] [CrossRef]
- Kampinga, H.H.; Craig, E.A. The HSP70 chaperone machinery: J proteins as drivers of functional specificity. Nat. Rev. Mol. Cell Biol. 2010, 11, 579–592. [Google Scholar] [CrossRef]
- Kityk, R.; Kopp, J.; Mayer, M.P. Molecular Mechanism of J-Domain-Triggered ATP Hydrolysis by Hsp70 Chaperones. Mol. Cell 2018, 69, 227–237.e224. [Google Scholar] [CrossRef]
- Cyr, D.M. Cooperation of the molecular chaperone Ydj1 with specific Hsp70 homologs to suppress protein aggregation. FEBS Lett. 1995, 359, 129–132. [Google Scholar] [CrossRef]
- Lu, Z.; Cyr, D.M. Protein folding activity of Hsp70 is modified differentially by the hsp40 co-chaperones Sis1 and Ydj1. J. Biol. Chem. 1998, 273, 27824–27830. [Google Scholar] [CrossRef] [PubMed]
- Qi, M.; Zhang, J.; Zeng, W.; Chen, X. DNAJB1 stabilizes MDM2 and contributes to cancer cell proliferation in a p53-dependent manner. Biochim. Biophys. Acta 2014, 1839, 62–69. [Google Scholar] [CrossRef]
- Hageman, J.; Rujano, M.A.; van Waarde, M.A.; Kakkar, V.; Dirks, R.P.; Govorukhina, N.; Oosterveld-Hut, H.M.; Lubsen, N.H.; Kampinga, H.H. A DNAJB chaperone subfamily with HDAC-dependent activities suppresses toxic protein aggregation. Mol. Cell 2010, 37, 355–369. [Google Scholar] [CrossRef]
- Yang, C.; Owen, H.A.; Yang, P. Dimeric heat shock protein 40 binds radial spokes for generating coupled power strokes and recovery strokes of 9 + 2 flagella. J. Cell Biol. 2008, 180, 403–415. [Google Scholar] [CrossRef] [PubMed]
- Ushioda, R.; Hoseki, J.; Araki, K.; Jansen, G.; Thomas, D.Y.; Nagata, K. ERdj5 is required as a disulfide reductase for degradation of misfolded proteins in the ER. Science 2008, 321, 569–572. [Google Scholar] [CrossRef] [PubMed]
- Sahi, C.; Lee, T.; Inada, M.; Pleiss, J.A.; Craig, E.A. Cwc23, an essential J protein critical for pre-mRNA splicing with a dispensable J domain. Mol. Cell Biol. 2010, 30, 33–42. [Google Scholar] [CrossRef]
- Kim Chiaw, P.; Hantouche, C.; Wong, M.J.H.; Matthes, E.; Robert, R.; Hanrahan, J.W.; Shrier, A.; Young, J.C. Hsp70 and DNAJA2 limit CFTR levels through degradation. PLoS ONE 2019, 14, e0220984. [Google Scholar] [CrossRef] [PubMed]
- Mok, S.A.; Condello, C.; Freilich, R.; Gillies, A.; Arhar, T.; Oroz, J.; Kadavath, H.; Julien, O.; Assimon, V.A.; Rauch, J.N.; et al. Mapping interactions with the chaperone network reveals factors that protect against tau aggregation. Nat. Struct. Mol. Biol. 2018, 25, 384–393. [Google Scholar] [CrossRef]
- Baaklini, I.; Gonçalves, C.C.; Lukacs, G.L.; Young, J.C. Selective Binding of HSC70 and its Co-Chaperones to Structural Hotspots on CFTR. Sci. Rep. 2020, 10, 4176. [Google Scholar] [CrossRef]
- Pohl, S.; Pervaiz, S.; Dharmarajan, A.; Agostino, M. Gene expression analysis of heat-shock proteins and redox regulators reveals combinatorial prognostic markers in carcinomas of the gastrointestinal tract. Redox Biol. 2019, 25, 101060. [Google Scholar] [CrossRef]
- Mahoney, S.E.; Yao, Z.; Keyes, C.C.; Tapscott, S.J.; Diede, S.J. Genome-wide DNA methylation studies suggest distinct DNA methylation patterns in pediatric embryonal and alveolar rhabdomyosarcomas. Epigenetics 2012, 7, 400–408. [Google Scholar] [CrossRef]
- Tukaj, S.; Kotlarz, A.; Jozwik, A.; Smolenska, Z.; Bryl, E.; Witkowski, J.M.; Lipinska, B. Hsp40 proteins modulate humoral and cellular immune response in rheumatoid arthritis patients. Cell Stress Chaperones 2010, 15, 555–566. [Google Scholar] [CrossRef]
- Kotlarz, A.; Tukaj, S.; Krzewski, K.; Brycka, E.; Lipinska, B. Human Hsp40 proteins, DNAJA1 and DNAJA2, as potential targets of the immune response triggered by bacterial DnaJ in rheumatoid arthritis. Cell Stress Chaperones 2013, 18, 653–659. [Google Scholar] [CrossRef]
- Wyttenbach, A.; Carmichael, J.; Swartz, J.; Furlong, R.A.; Narain, Y.; Rankin, J.; Rubinsztein, D.C. Effects of heat shock, heat shock protein 40 (HDJ-2), and proteasome inhibition on protein aggregation in cellular models of Huntington’s disease. Proc. Natl. Acad. Sci. USA 2000, 97, 2898–2903. [Google Scholar] [CrossRef]
- Rodríguez-González, C.; Lin, S.; Arkan, S.; Hansen, C. Co-chaperones DNAJA1 and DNAJB6 are critical for regulation of polyglutamine aggregation. Sci. Rep. 2020, 10, 8130. [Google Scholar] [CrossRef]
- Xu, D.; Tong, X.; Sun, L.; Li, H.; Jones, R.D.; Liao, J.; Yang, G.Y. Inhibition of mutant Kras and p53-driven pancreatic carcinogenesis by atorvastatin: Mainly via targeting of the farnesylated DNAJA1 in chaperoning mutant p53. Mol. Carcinog. 2019, 58, 2052–2064. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Ren, X.; Liang, Y.; Yan, Y.; Zhou, Y.; Hu, J.; Wang, Z.; Song, F.; Wang, F.; Liao, W.; et al. KNK437 restricts the growth and metastasis of colorectal cancer via targeting DNAJA1/CDC45 axis. Oncogene 2020, 39, 249–261. [Google Scholar] [CrossRef]
- Stark, J.L.; Mehla, K.; Chaika, N.; Acton, T.B.; Xiao, R.; Singh, P.K.; Montelione, G.T.; Powers, R. Structure and function of human DnaJ homologue subfamily a member 1 (DNAJA1) and its relationship to pancreatic cancer. Biochemistry 2014, 53, 1360–1372. [Google Scholar] [CrossRef]
- Chung, Y.T.; Matkowskyj, K.A.; Li, H.; Bai, H.; Zhang, W.; Tsao, M.S.; Liao, J.; Yang, G.Y. Overexpression and oncogenic function of aldo-keto reductase family 1B10 (AKR1B10) in pancreatic carcinoma. Mod. Pathol. 2012, 25, 758–766. [Google Scholar] [CrossRef] [PubMed]
- Meshalkina, D.A.; Shevtsov, M.A.; Dobrodumov, A.V.; Komarova, E.Y.; Voronkina, I.V.; Lazarev, V.F.; Margulis, B.A.; Guzhova, I.V. Knock-down of Hdj2/DNAJA1 co-chaperone results in an unexpected burst of tumorigenicity of C6 glioblastoma cells. Oncotarget 2016, 7, 22050–22063. [Google Scholar] [CrossRef] [PubMed]
- Ileri, F.C.; Acun, T. High expression of DNAJA1 (HDJ2) predicts unfavorable survival outcomes in breast cancer. Biomark. Med. 2021, 15, 941–950. [Google Scholar] [CrossRef]
- Roth, H.E.; Bhinderwala, F.; Franco, R.; Zhou, Y.; Powers, R. DNAJA1 Dysregulates Metabolism Promoting an Antiapoptotic Phenotype in Pancreatic Ductal Adenocarcinoma. J. Proteome Res. 2021, 20, 3925–3939. [Google Scholar] [CrossRef] [PubMed]
- Ergulen, E.; Becsi, B.; Csomos, I.; Fesus, L.; Kanchan, K. Identification of DNAJA1 as a novel interacting partner and a substrate of human transglutaminase 2. Biochem. J. 2016, 473, 3889–3901. [Google Scholar] [CrossRef] [PubMed]
- Gotoh, T.; Terada, K.; Oyadomari, S.; Mori, M. hsp70-DnaJ chaperone pair prevents nitric oxide- and CHOP-induced apoptosis by inhibiting translocation of Bax to mitochondria. Cell Death Differ. 2004, 11, 390–402. [Google Scholar] [CrossRef]
- Tong, X.; Xu, D.; Mishra, R.K.; Jones, R.D.; Sun, L.; Schiltz, G.E.; Liao, J.; Yang, G.Y. Identification of a druggable protein-protein interaction site between mutant p53 and its stabilizing chaperone DNAJA1. J. Biol. Chem. 2020, 296, 100098. [Google Scholar] [CrossRef] [PubMed]
- Kaida, A.; Yamamoto, S.; Parrales, A.; Young, E.D.; Ranjan, A.; Alalem, M.A.; Morita, K.I.; Oikawa, Y.; Harada, H.; Ikeda, T.; et al. DNAJA1 promotes cancer metastasis through interaction with mutant p53. Oncogene 2021, 40, 5013–5025. [Google Scholar] [CrossRef]
- Terada, K.; Mori, M. Human DnaJ homologs dj2 and dj3, and bag-1 are positive cochaperones of hsc70. J. Biol. Chem. 2000, 275, 24728–24734. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.C.; Liao, Y.P.; Mischel, P.S.; Iwamoto, K.S.; Cacalano, N.A.; McBride, W.H. HDJ-2 as a target for radiosensitization of glioblastoma multiforme cells by the farnesyltransferase inhibitor R115777 and the role of the p53/p21 pathway. Cancer Res. 2006, 66, 6756–6762. [Google Scholar] [CrossRef] [PubMed]
- Ahn, B.Y.; Trinh, D.L.; Zajchowski, L.D.; Lee, B.; Elwi, A.N.; Kim, S.W. Tid1 is a new regulator of p53 mitochondrial translocation and apoptosis in cancer. Oncogene 2010, 29, 1155–1166. [Google Scholar] [CrossRef]
- Trinh, D.L.; Elwi, A.N.; Kim, S.W. Direct interaction between p53 and Tid1 proteins affects p53 mitochondrial localization and apoptosis. Oncotarget 2010, 1, 396–404. [Google Scholar] [CrossRef]
- Cui, X.; Choi, H.K.; Choi, Y.S.; Park, S.Y.; Sung, G.J.; Lee, Y.H.; Lee, J.; Jun, W.J.; Kim, K.; Choi, K.C.; et al. DNAJB1 destabilizes PDCD5 to suppress p53-mediated apoptosis. Cancer Lett. 2015, 357, 307–315. [Google Scholar] [CrossRef]
- Hiraki, M.; Hwang, S.Y.; Cao, S.; Ramadhar, T.R.; Byun, S.; Yoon, K.W.; Lee, J.H.; Chu, K.; Gurkar, A.U.; Kolev, V.; et al. Small-Molecule Reactivation of Mutant p53 to Wild-Type-like p53 through the p53-Hsp40 Regulatory Axis. Chem. Biol. 2015, 22, 1206–1216. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.J.; Kim, J.M.; Kim, K.H.; Heo, J.I.; Kwak, S.J.; Han, J.A. Genotoxic stress/p53-induced DNAJB9 inhibits the pro-apoptotic function of p53. Cell Death Differ. 2015, 22, 86–95. [Google Scholar] [CrossRef]
- Lee, H.J.; Jung, Y.J.; Lee, S.; Kim, J.I.; Han, J.A. DNAJB9 Inhibits p53-Dependent Oncogene-Induced Senescence and Induces Cell Transformation. Mol. Cells 2020, 43, 397–407. [Google Scholar] [CrossRef] [PubMed]
- Lazare, C.; Zhi, W.; Dai, J.; Cao, C.; Sookha, R.R.; Wang, L.; Meng, Y.; Gao, P.; Wu, P.; Wei, J.; et al. A pilot study comparing the genetic molecular biology of gestational and non-gestational choriocarcinoma. Am. J. Transl. Res. 2019, 11, 7049–7062. [Google Scholar]
- Imamura, T.; Komatsu, S.; Ichikawa, D.; Miyamae, M.; Okajima, W.; Ohashi, T.; Kiuchi, J.; Nishibeppu, K.; Kosuga, T.; Konishi, H.; et al. Overexpression of ZRF1 is related to tumor malignant potential and a poor outcome of gastric carcinoma. Carcinogenesis 2018, 39, 263–271. [Google Scholar] [CrossRef]
- Kubo, N.; Wu, D.; Yoshihara, Y.; Sang, M.; Nakagawara, A.; Ozaki, T. Co-chaperon DnaJC7/TPR2 enhances p53 stability and activity through blocking the complex formation between p53 and MDM2. Biochem. Biophys. Res. Commun. 2013, 430, 1034–1039. [Google Scholar] [CrossRef]
- Tebaldi, T.; Zaccara, S.; Alessandrini, F.; Bisio, A.; Ciribilli, Y.; Inga, A. Whole-genome cartography of p53 response elements ranked on transactivation potential. BMC Genom. 2015, 16, 464. [Google Scholar] [CrossRef] [PubMed]
- Isermann, T.; Sener, O.C.; Stender, A.; Klemke, L.; Winkler, N.; Neesse, A.; Li, J.; Wegwitz, F.; Moll, U.M.; Schulz-Heddergott, R. Suppression of HSF1 activity by wildtype p53 creates a driving force for p53 loss-of-heterozygosity. Nat. Commun. 2021, 12, 4019. [Google Scholar] [CrossRef] [PubMed]
- Nitika; Blackman, J.S.; Knighton, L.E.; Takakuwa, J.E.; Calderwood, S.K.; Truman, A.W. Chemogenomic screening identifies the Hsp70 co-chaperone DNAJA1 as a hub for anticancer drug resistance. Sci. Rep. 2020, 10, 13831. [Google Scholar] [CrossRef]
- Masunaga, S.; Ono, K.; Takahashi, A.; Ohnishi, T.; Kinashi, Y.; Takagaki, M. Radiobiological characteristics of solid tumours depending on the p53 status of the tumour cells, with emphasis on the response of intratumour quiescent cells. Eur. J. Cancer 2002, 38, 718–727. [Google Scholar] [CrossRef]
- Masunaga, S.I.; Kobayashi, J.; Tano, K.; Sanada, Y.; Suzuki, M.; Ono, K. The Effect of p53 Status on Radio-Sensitivity of Quiescent Tumor Cell Population Irradiated With γ-Rays at Various Dose Rates. J. Clin. Med. Res. 2018, 10, 815–821. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Okaichi, K.; Ide-Kanematsu, M.; Izumi, N.; Morita, N.; Okumura, Y.; Ihara, M. Variations in sensitivity to ionizing radiation in relation to p53 mutation point. Anticancer Res. 2008, 28, 2687–2690. [Google Scholar]
- Skinner, H.D.; Sandulache, V.C.; Ow, T.J.; Meyn, R.E.; Yordy, J.S.; Beadle, B.M.; Fitzgerald, A.L.; Giri, U.; Ang, K.K.; Myers, J.N. TP53 disruptive mutations lead to head and neck cancer treatment failure through inhibition of radiation-induced senescence. Clin. Cancer Res. 2012, 18, 290–300. [Google Scholar] [CrossRef] [PubMed]
- Wisén, S.; Bertelsen, E.B.; Thompson, A.D.; Patury, S.; Ung, P.; Chang, L.; Evans, C.G.; Walter, G.M.; Wipf, P.; Carlson, H.A.; et al. Binding of a small molecule at a protein-protein interface regulates the chaperone activity of hsp70-hsp40. ACS Chem. Biol. 2010, 5, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Moses, M.A.; Kim, Y.S.; Rivera-Marquez, G.M.; Oshima, N.; Watson, M.J.; Beebe, K.E.; Wells, C.; Lee, S.; Zuehlke, A.D.; Shao, H.; et al. Targeting the Hsp40/Hsp70 Chaperone Axis as a Novel Strategy to Treat Castration-Resistant Prostate Cancer. Cancer Res. 2018, 78, 4022–4035. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.F.; Huang, K.H.; Tseng, W.C.; Lo, J.F.; Li, A.F.; Fang, W.L.; Chen, C.F.; Yeh, T.S.; Chang, Y.L.; Chou, Y.C.; et al. DNAJA3/Tid1 Is Required for Mitochondrial DNA Maintenance and Regulates Migration and Invasion of Human Gastric Cancer Cells. Cancers 2020, 12, 3463. [Google Scholar] [CrossRef] [PubMed]
- Syken, J.; De-Medina, T.; Munger, K. TID1, a human homolog of the Drosophila tumor suppressor l(2)tid, encodes two mitochondrial modulators of apoptosis with opposing functions. Proc. Natl. Acad. Sci. USA 1999, 96, 8499–8504. [Google Scholar] [CrossRef]
- Chen, C.Y.; Chiou, S.H.; Huang, C.Y.; Jan, C.I.; Lin, S.C.; Hu, W.Y.; Chou, S.H.; Liu, C.J.; Lo, J.F. Tid1 functions as a tumour suppressor in head and neck squamous cell carcinoma. J. Pathol. 2009, 219, 347–355. [Google Scholar] [CrossRef] [PubMed]
- Tarunina, M.; Alger, L.; Chu, G.; Munger, K.; Gudkov, A.; Jat, P.S. Functional genetic screen for genes involved in senescence: Role of Tid1, a homologue of the Drosophila tumor suppressor l(2)tid, in senescence and cell survival. Mol. Cell Biol. 2004, 24, 10792–10801. [Google Scholar] [CrossRef]
- Cheng, L.H.; Hung, K.F.; Lee, T.C.; Huang, C.Y.; Chiu, W.T.; Lo, J.F.; Huang, T.F. Mitochondrial co-chaperone protein Tid1 is required for energy homeostasis during skeletal myogenesis. Stem. Cell Res. Ther. 2016, 7, 185. [Google Scholar] [CrossRef]
- Lu, B.; Garrido, N.; Spelbrink, J.N.; Suzuki, C.K. Tid1 isoforms are mitochondrial DnaJ-like chaperones with unique carboxyl termini that determine cytosolic fate. J. Biol. Chem. 2006, 281, 13150–13158. [Google Scholar] [CrossRef]
- Maletzko, A.; Key, J.; Wittig, I.; Gispert, S.; Koepf, G.; Canet-Pons, J.; Torres-Odio, S.; West, A.P.; Auburger, G. Increased presence of nuclear DNAJA3 and upregulation of cytosolic STAT1 and of nucleic acid sensors trigger innate immunity in the ClpP-null mouse. Neurogenetics 2021, 22, 297–312. [Google Scholar] [CrossRef]
- Schilling, B.; De-Medina, T.; Syken, J.; Vidal, M.; Münger, K. A novel human DnaJ protein, hTid-1, a homolog of the Drosophila tumor suppressor protein Tid56, can interact with the human papillomavirus type 16 E7 oncoprotein. Virology 1998, 247, 74–85. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Cenciarelli, C.; Shao, Z.; Vidal, M.; Parks, W.P.; Pagano, M.; Cheng-Mayer, C. Human T cell leukemia virus type 1 Tax associates with a molecular chaperone complex containing hTid-1 and Hsp70. Curr. Biol. 2001, 11, 1771–1775. [Google Scholar] [CrossRef]
- Cheng, H.; Cenciarelli, C.; Tao, M.; Parks, W.P.; Cheng-Mayer, C. HTLV-1 Tax-associated hTid-1, a human DnaJ protein, is a repressor of Ikappa B kinase beta subunit. J. Biol. Chem. 2002, 277, 20605–20610. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Yang, F.; Zhu, Z.; Yang, Y.; Wang, Z.; Cao, W.; Dang, W.; Li, L.; Mao, R.; Liu, Y.; et al. Cellular DNAJA3, a Novel VP1-Interacting Protein, Inhibits Foot-and-Mouth Disease Virus Replication by Inducing Lysosomal Degradation of VP1 and Attenuating Its Antagonistic Role in the Beta Interferon Signaling Pathway. J. Virol. 2019, 93, e00588-19. [Google Scholar] [CrossRef]
- Hayashi, M.; Imanaka-Yoshida, K.; Yoshida, T.; Wood, M.; Fearns, C.; Tatake, R.J.; Lee, J.D. A crucial role of mitochondrial Hsp40 in preventing dilated cardiomyopathy. Nat. Med. 2006, 12, 128–132. [Google Scholar] [CrossRef]
- Chen, Y.S.; Chang, C.W.; Tsay, Y.G.; Huang, L.Y.; Wu, Y.C.; Cheng, L.H.; Yang, C.C.; Wu, C.H.; Teo, W.H.; Hung, K.F.; et al. HSP40 co-chaperone protein Tid1 suppresses metastasis of head and neck cancer by inhibiting Galectin-7-TCF3-MMP9 axis signaling. Theranostics 2018, 8, 3841–3855. [Google Scholar] [CrossRef]
- Jan, C.I.; Yu, C.C.; Hung, M.C.; Harn, H.J.; Nieh, S.; Lee, H.S.; Lou, M.A.; Wu, Y.C.; Chen, C.Y.; Huang, C.Y.; et al. Tid1, CHIP and ErbB2 interactions and their prognostic implications for breast cancer patients. J. Pathol. 2011, 225, 424–437. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.Y.; Jan, C.I.; Lo, J.F.; Yang, S.C.; Chang, Y.L.; Pan, S.H.; Wang, W.L.; Hong, T.M.; Yang, P.C. Tid1-L inhibits EGFR signaling in lung adenocarcinoma by enhancing EGFR Ubiquitinylation and degradation. Cancer Res. 2013, 73, 4009–4019. [Google Scholar] [CrossRef]
- Chen, K.Y.; Huang, Y.H.; Teo, W.H.; Chang, C.W.; Chen, Y.S.; Yeh, Y.C.; Lee, C.J.; Lo, J.F. Loss of Tid1/DNAJA3 Co-Chaperone Promotes Progression and Recurrence of Hepatocellular Carcinoma after Surgical Resection: A Novel Model to Stratify Risk of Recurrence. Cancers 2021, 13, 138. [Google Scholar] [CrossRef]
- Bae, M.K.; Jeong, J.W.; Kim, S.H.; Kim, S.Y.; Kang, H.J.; Kim, D.M.; Bae, S.K.; Yun, I.; Trentin, G.A.; Rozakis-Adcock, M.; et al. Tid-1 interacts with the von Hippel-Lindau protein and modulates angiogenesis by destabilization of HIF-1alpha. Cancer Res. 2005, 65, 2520–2525. [Google Scholar] [CrossRef]
- Kurzik-Dumke, U.; Horner, M.; Czaja, J.; Nicotra, M.R.; Simiantonaki, N.; Koslowski, M.; Natali, P.G. Progression of colorectal cancers correlates with overexpression and loss of polarization of expression of the htid-1 tumor suppressor. Int. J. Mol. Med. 2008, 21, 19–31. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.H.; Lin, Y.H.; Yang, S.C.; Chang, P.C.; Wang, T.C.; Chen, C.Y. Tid1-S regulates the mitochondrial localization of EGFR in non-small cell lung carcinoma. Oncogenesis 2017, 6, e361. [Google Scholar] [CrossRef]
- Faust, O.; Abayev-Avraham, M.; Wentink, A.S.; Maurer, M.; Nillegoda, N.B.; London, N.; Bukau, B.; Rosenzweig, R. HSP40 proteins use class-specific regulation to drive HSP70 functional diversity. Nature 2020, 587, 489–494. [Google Scholar] [CrossRef]
- Gess, B.; Auer-Grumbach, M.; Schirmacher, A.; Strom, T.; Zitzelsberger, M.; Rudnik-Schöneborn, S.; Röhr, D.; Halfter, H.; Young, P.; Senderek, J. HSJ1-related hereditary neuropathies: Novel mutations and extended clinical spectrum. Neurology 2014, 83, 1726–1732. [Google Scholar] [CrossRef] [PubMed]
- Lupo, V.; García-García, F.; Sancho, P.; Tello, C.; García-Romero, M.; Villarreal, L.; Alberti, A.; Sivera, R.; Dopazo, J.; Pascual-Pascual, S.I.; et al. Assessment of Targeted Next-Generation Sequencing as a Tool for the Diagnosis of Charcot-Marie-Tooth Disease and Hereditary Motor Neuropathy. J. Mol. Diagn 2016, 18, 225–234. [Google Scholar] [CrossRef]
- Sanchez, E.; Darvish, H.; Mesias, R.; Taghavi, S.; Firouzabadi, S.G.; Walker, R.H.; Tafakhori, A.; Paisán-Ruiz, C. Identification of a Large DNAJB2 Deletion in a Family with Spinal Muscular Atrophy and Parkinsonism. Hum. Mutat. 2016, 37, 1180–1189. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Hayashi, Y.K.; Oya, Y.; Kondo, T.; Sugie, K.; Kaneda, D.; Houzen, H.; Yabe, I.; Sasaki, H.; Noguchi, S.; et al. DNAJB6 myopathy in an Asian cohort and cytoplasmic/nuclear inclusions. Neuromuscul. Disord. 2013, 23, 269–276. [Google Scholar] [CrossRef]
- Harms, M.B.; Sommerville, R.B.; Allred, P.; Bell, S.; Ma, D.; Cooper, P.; Lopate, G.; Pestronk, A.; Weihl, C.C.; Baloh, R.H. Exome sequencing reveals DNAJB6 mutations in dominantly-inherited myopathy. Ann. Neurol. 2012, 71, 407–416. [Google Scholar] [CrossRef]
- Sarparanta, J.; Jonson, P.H.; Golzio, C.; Sandell, S.; Luque, H.; Screen, M.; McDonald, K.; Stajich, J.M.; Mahjneh, I.; Vihola, A.; et al. Mutations affecting the cytoplasmic functions of the co-chaperone DNAJB6 cause limb-girdle muscular dystrophy. Nat. Genet. 2012, 44, s451–452. [Google Scholar] [CrossRef] [PubMed]
- Palmio, J.; Jonson, P.H.; Evilä, A.; Auranen, M.; Straub, V.; Bushby, K.; Sarkozy, A.; Kiuru-Enari, S.; Sandell, S.; Pihko, H.; et al. Novel mutations in DNAJB6 gene cause a very severe early-onset limb-girdle muscular dystrophy 1D disease. Neuromuscul. Disord. 2015, 25, 835–842. [Google Scholar] [CrossRef]
- Tsai, P.C.; Tsai, Y.S.; Soong, B.W.; Huang, Y.H.; Wu, H.T.; Chen, Y.H.; Lin, K.P.; Liao, Y.C.; Lee, Y.C. A novel DNAJB6 mutation causes dominantly inherited distal-onset myopathy and compromises DNAJB6 function. Neuromuscul. Disord. 2017, 92, 150–157. [Google Scholar] [CrossRef]
- El Khouri, E.; Thomas, L.; Jeanson, L.; Bequignon, E.; Vallette, B.; Duquesnoy, P.; Montantin, G.; Copin, B.; Dastot-Le Moal, F.; Blanchon, S.; et al. Mutations in DNAJB13, Encoding an HSP40 Family Member, Cause Primary Ciliary Dyskinesia and Male Infertility. Am. J. Hum. Genet. 2016, 99, 489–500. [Google Scholar] [CrossRef] [PubMed]
- Honeyman, J.N.; Simon, E.P.; Robine, N.; Chiaroni-Clarke, R.; Darcy, D.G.; Lim, I.; Gleason, C.E.; Murphy, J.M.; Rosenberg, B.R.; Teegan, L.; et al. Detection of a recurrent DNAJB1-PRKACA chimeric transcript in fibrolamellar hepatocellular carcinoma. Science 2014, 343, 1010–1014. [Google Scholar] [CrossRef]
- Chen, C.H.; Chang, W.H.; Su, K.Y.; Ku, W.H.; Chang, G.C.; Hong, Q.S.; Hsiao, Y.J.; Chen, H.C.; Chen, H.Y.; Wu, R.; et al. HLJ1 is an endogenous Src inhibitor suppressing cancer progression through dual mechanisms. Oncogene 2016, 35, 5674–5685. [Google Scholar] [CrossRef] [PubMed]
- Miao, W.; Li, L.; Wang, Y. A Targeted Proteomic Approach for Heat Shock Proteins Reveals DNAJB4 as a Suppressor for Melanoma Metastasis. Anal. Chem. 2018, 90, 6835–6842. [Google Scholar] [CrossRef] [PubMed]
- Uretmen Kagiali, Z.C.; Sanal, E.; Karayel, O.; Polat, A.N.; Saatci, O.; Ersan, P.G.; Trappe, K.; Renard, B.Y.; Onder, T.T.; Tuncbag, N.; et al. Systems-level Analysis Reveals Multiple Modulators of Epithelial-mesenchymal Transition and Identifies DNAJB4 and CD81 as Novel Metastasis Inducers in Breast Cancer. Mol. Cell Proteom. 2019, 18, 1756–1771. [Google Scholar] [CrossRef] [PubMed]
- Menezes, M.E.; Mitra, A.; Shevde, L.A.; Samant, R.S. DNAJB6 governs a novel regulatory loop determining Wnt/beta-catenin signalling activity. Biochem. J. 2012, 444, 573–580. [Google Scholar] [CrossRef]
- Mitra, A.; Fillmore, R.A.; Metge, B.J.; Rajesh, M.; Xi, Y.; King, J.; Ju, J.; Pannell, L.; Shevde, L.A.; Samant, R.S. Large isoform of MRJ (DNAJB6) reduces malignant activity of breast cancer. Breast Cancer Res. 2008, 10, R22. [Google Scholar] [CrossRef]
- Tien, J.F.; Mazloomian, A.; Cheng, S.G.; Hughes, C.S.; Chow, C.C.T.; Canapi, L.T.; Oloumi, A.; Trigo-Gonzalez, G.; Bashashati, A.; Xu, J.; et al. CDK12 regulates alternative last exon mRNA splicing and promotes breast cancer cell invasion. Nucleic Acids Res. 2017, 45, 6698–6716. [Google Scholar] [CrossRef] [PubMed]
- Yu, V.Z.; Wong, V.C.; Dai, W.; Ko, J.M.; Lam, A.K.; Chan, K.W.; Samant, R.S.; Lung, H.L.; Shuen, W.H.; Law, S.; et al. Nuclear Localization of DNAJB6 Is Associated With Survival of Patients With Esophageal Cancer and Reduces AKT Signaling and Proliferation of Cancer Cells. Gastroenterology 2015, 149, 1825–1836.e1825. [Google Scholar] [CrossRef]
- Zhang, T.T.; Jiang, Y.Y.; Shang, L.; Shi, Z.Z.; Liang, J.W.; Wang, Z.; Zhang, Y.; Hao, J.J.; Jia, X.M.; Xu, X.; et al. Overexpression of DNAJB6 promotes colorectal cancer cell invasion through an IQGAP1/ERK-dependent signaling pathway. Mol. Carcinog. 2015, 54, 1205–1213. [Google Scholar] [CrossRef]
- Nishizawa, S.; Hirohashi, Y.; Torigoe, T.; Takahashi, A.; Tamura, Y.; Mori, T.; Kanaseki, T.; Kamiguchi, K.; Asanuma, H.; Morita, R.; et al. HSP DNAJB8 controls tumor-initiating ability in renal cancer stem-like cells. Cancer Res. 2012, 72, 2844–2854. [Google Scholar] [CrossRef] [PubMed]
- Kusumoto, H.; Hirohashi, Y.; Nishizawa, S.; Yamashita, M.; Yasuda, K.; Murai, A.; Takaya, A.; Mori, T.; Kubo, T.; Nakatsugawa, M.; et al. Cellular stress induces cancer stem-like cells through expression of DNAJB8 by activation of heat shock factor 1. Cancer Sci. 2018, 109, 741–750. [Google Scholar] [CrossRef]
- Pan, J.; Cao, D.; Gong, J. The endoplasmic reticulum co-chaperone ERdj3/DNAJB11 promotes hepatocellular carcinoma progression through suppressing AATZ degradation. Future Oncol. 2018, 14, 3001–3013. [Google Scholar] [CrossRef] [PubMed]
- Sopha, P.; Ren, H.Y.; Grove, D.E.; Cyr, D.M. Endoplasmic reticulum stress-induced degradation of DNAJB12 stimulates BOK accumulation and primes cancer cells for apoptosis. J. Biol. Chem. 2017, 292, 11792–11803. [Google Scholar] [CrossRef]
- Gao, X.; Carroni, M.; Nussbaum-Krammer, C.; Mogk, A.; Nillegoda, N.B.; Szlachcic, A.; Guilbride, D.L.; Saibil, H.R.; Mayer, M.P.; Bukau, B. Human Hsp70 Disaggregase Reverses Parkinson’s-Linked alpha-Synuclein Amyloid Fibrils. Mol. Cell 2015, 59, 781–793. [Google Scholar] [CrossRef]
- Irwin, R.; Faust, O.; Petrovic, I.; Wolf, S.G.; Hofmann, H.; Rosenzweig, R. Hsp40s play complementary roles in the prevention of tau amyloid formation. Elife 2021, 10, e69601. [Google Scholar] [CrossRef]
- Takashima, K.; Oshiumi, H.; Matsumoto, M.; Seya, T. DNAJB1/HSP40 Suppresses Melanoma Differentiation-Associated Gene 5-Mitochondrial Antiviral Signaling Protein Function in Conjunction with HSP70. J. Innate Immun. 2018, 10, 44–55. [Google Scholar] [CrossRef]
- Batra, J.; Tripathi, S.; Kumar, A.; Katz, J.M.; Cox, N.J.; Lal, R.B.; Sambhara, S.; Lal, S.K. Human Heat shock protein 40 (Hsp40/DnaJB1) promotes influenza A virus replication by assisting nuclear import of viral ribonucleoproteins. Sci. Rep. 2016, 6, 19063. [Google Scholar] [CrossRef]
- Park, S.Y.; Choi, H.K.; Seo, J.S.; Yoo, J.Y.; Jeong, J.W.; Choi, Y.; Choi, K.C.; Yoon, H.G. DNAJB1 negatively regulates MIG6 to promote epidermal growth factor receptor signaling. Biochim. Biophys. Acta 2015, 1853, 2722–2730. [Google Scholar] [CrossRef]
- Kastenhuber, E.R.; Lalazar, G.; Houlihan, S.L.; Tschaharganeh, D.F.; Baslan, T.; Chen, C.C.; Requena, D.; Tian, S.; Bosbach, B.; Wilkinson, J.E.; et al. DNAJB1-PRKACA fusion kinase interacts with beta-catenin and the liver regenerative response to drive fibrolamellar hepatocellular carcinoma. Proc. Natl. Acad. Sci. USA 2017, 114, 13076–13084. [Google Scholar] [CrossRef]
- Hackel, P.O.; Gishizky, M.; Ullrich, A. Mig-6 is a negative regulator of the epidermal growth factor receptor signal. Biol. Chem. 2001, 382, 1649–1662. [Google Scholar] [CrossRef] [PubMed]
- Ren, H.; Luo, M.; Chen, J.; Zhou, Y.; Li, X.; Zhan, Y.; Shen, D.; Chen, B. Identification of TPD52 and DNAJB1 as two novel bile biomarkers for cholangiocarcinoma by iTRAQbased quantitative proteomics analysis. Oncol. Rep. 2019, 42, 2622–2634. [Google Scholar] [CrossRef] [PubMed]
- King, F.W.; Wawrzynow, A.; Hohfeld, J.; Zylicz, M. Co-chaperones Bag-1, Hop and Hsp40 regulate Hsc70 and Hsp90 interactions with wild-type or mutant p53. EMBO J. 2001, 20, 6297–6305. [Google Scholar] [CrossRef] [PubMed]
- Sugito, K.; Yamane, M.; Hattori, H.; Hayashi, Y.; Tohnai, I.; Ueda, M.; Tsuchida, N.; Ohtsuka, K. Interaction between hsp70 and hsp40, eukaryotic homologues of DnaK and DnaJ, in human cells expressing mutant-type p53. FEBS Lett. 1995, 358, 161–164. [Google Scholar] [CrossRef]
- Silva, G.; Marins, M.; Chaichanasak, N.; Yoon, Y.; Fachin, A.L.; Pinhanelli, V.C.; Regasini, L.O.; Dos Santos, M.B.; Ayusso, G.M.; Marques, B.C.; et al. Trans-chalcone increases p53 activity via DNAJB1/HSP40 induction and CRM1 inhibition. PLoS ONE 2018, 13, e0202263. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Meunier, L.; Hendershot, L.M. Identification and characterization of a novel endoplasmic reticulum (ER) DnaJ homologue, which stimulates ATPase activity of BiP in vitro and is induced by ER stress. J. Biol. Chem. 2002, 277, 15947–15956. [Google Scholar] [CrossRef]
- Kurisu, J.; Honma, A.; Miyajima, H.; Kondo, S.; Okumura, M.; Imaizumi, K. MDG1/ERdj4, an ER-resident DnaJ family member, suppresses cell death induced by ER stress. Genes Cells 2003, 8, 189–202. [Google Scholar] [CrossRef] [PubMed]
- Dong, M.; Bridges, J.P.; Apsley, K.; Xu, Y.; Weaver, T.E. ERdj4 and ERdj5 are required for endoplasmic reticulum-associated protein degradation of misfolded surfactant protein C. Mol. Biol. Cell 2008, 19, 2620–2630. [Google Scholar] [CrossRef]
- Huang, Y.; Arora, K.; Mun, K.S.; Yang, F.; Moon, C.; Yarlagadda, S.; Jegga, A.; Weaver, T.; Naren, A.P. Targeting DNAJB9, a novel ER luminal co-chaperone, to rescue ΔF508-CFTR. Sci. Rep. 2019, 9, 9808. [Google Scholar] [CrossRef] [PubMed]
- Sun, F.; Liao, Y.; Qu, X.; Xiao, X.; Hou, S.; Chen, Z.; Huang, H.; Li, P.; Fu, S. Hepatic DNAJB9 Drives Anabolic Biasing to Reduce Steatosis and Obesity. Cell Rep. 2020, 30, 1835–1847.e1839. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.Y.; Kim, Y.M.; Hong, S. DNAJB9 suppresses the metastasis of triple-negative breast cancer by promoting FBXO45-mediated degradation of ZEB1. Cell Death Dis. 2021, 12, 461. [Google Scholar] [CrossRef]
- Lei, J.X.; Cassone, C.G.; Luebbert, C.; Liu, Q.Y. A novel neuron-enriched protein SDIM1 is down regulated in Alzheimer’s brains and attenuates cell death induced by DNAJB4 over-expression in neuro-progenitor cells. Mol. Neurodegener. 2011, 6, 9. [Google Scholar] [CrossRef]
- Snider, J.C.; Riley, L.A.; Mallory, N.T.; Bersi, M.R.; Umbarkar, P.; Gautam, R.; Zhang, Q.; Mahadevan-Jansen, A.; Hatzopoulos, A.K.; Maroteaux, L.; et al. Targeting 5-HT(2B) Receptor Signaling Prevents Border Zone Expansion and Improves Microstructural Remodeling After Myocardial Infarction. Circulation 2021, 143, 1317–1330. [Google Scholar] [CrossRef]
- Tsai, M.F.; Wang, C.C.; Chang, G.C.; Chen, C.Y.; Chen, H.Y.; Cheng, C.L.; Yang, Y.P.; Wu, C.Y.; Shih, F.Y.; Liu, C.C.; et al. A new tumor suppressor DnaJ-like heat shock protein, HLJ1, and survival of patients with non-small-cell lung carcinoma. J. Natl. Cancer Inst. 2006, 98, 825–838. [Google Scholar] [CrossRef]
- Liu, Y.; Zhou, J.; Zhang, C.; Fu, W.; Xiao, X.; Ruan, S.; Zhang, Y.; Luo, X.; Tang, M. HLJ1 is a novel biomarker for colorectal carcinoma progression and overall patient survival. Int. J. Clin. Exp. Pathol. 2014, 7, 969–977. [Google Scholar] [PubMed]
- Acun, T.; Doberstein, N.; Habermann, J.K.; Gemoll, T.; Thorns, C.; Oztas, E.; Ried, T. HLJ1 (DNAJB4) Gene Is a Novel Biomarker Candidate in Breast Cancer. OMICS 2017, 21, 257–265. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Peng, N.; Zhuang, H.; Zhang, D.; Wang, Y.; Hua, Z.C. Heat shock proteins HSP70 and MRJ cooperatively regulate cell adhesion and migration through urokinase receptor. BMC Cancer 2014, 14, 639. [Google Scholar] [CrossRef] [PubMed]
- Mitra, A.; Menezes, M.E.; Shevde, L.A.; Samant, R.S. DNAJB6 induces degradation of beta-catenin and causes partial reversal of mesenchymal phenotype. J. Biol. Chem. 2010, 285, 24686–24694. [Google Scholar] [CrossRef] [PubMed]
- Mitra, A.; Menezes, M.E.; Pannell, L.K.; Mulekar, M.S.; Honkanen, R.E.; Shevde, L.A.; Samant, R.S. DNAJB6 chaperones PP2A mediated dephosphorylation of GSK3beta to downregulate beta-catenin transcription target, osteopontin. Oncogene 2012, 31, 4472–4483. [Google Scholar] [CrossRef] [PubMed]
- Gillis, J.; Schipper-Krom, S.; Juenemann, K.; Gruber, A.; Coolen, S.; van den Nieuwendijk, R.; van Veen, H.; Overkleeft, H.; Goedhart, J.; Kampinga, H.H.; et al. The DNAJB6 and DNAJB8 protein chaperones prevent intracellular aggregation of polyglutamine peptides. J. Biol. Chem. 2013, 288, 17225–17237. [Google Scholar] [CrossRef]
- Månsson, C.; Kakkar, V.; Monsellier, E.; Sourigues, Y.; Härmark, J.; Kampinga, H.H.; Melki, R.; Emanuelsson, C. DNAJB6 is a peptide-binding chaperone which can suppress amyloid fibrillation of polyglutamine peptides at substoichiometric molar ratios. Cell Stress Chaperones 2014, 19, 227–239. [Google Scholar] [CrossRef] [PubMed]
- Morita, R.; Nishizawa, S.; Torigoe, T.; Takahashi, A.; Tamura, Y.; Tsukahara, T.; Kanaseki, T.; Sokolovskaya, A.; Kochin, V.; Kondo, T.; et al. Heat shock protein DNAJB8 is a novel target for immunotherapy of colon cancer-initiating cells. Cancer Sci. 2014, 105, 389–395. [Google Scholar] [CrossRef]
- Yamashita, M.; Hirohashi, Y.; Torigoe, T.; Kusumoto, H.; Murai, A.; Imagawa, T.; Sato, N. Dnajb8, a Member of the Heat Shock Protein 40 Family Has a Role in the Tumor Initiation and Resistance to Docetaxel but Is Dispensable for Stress Response. PLoS ONE 2016, 11, e0146501. [Google Scholar] [CrossRef]
- Yu, M.; Haslam, R.H.; Haslam, D.B. HEDJ, an Hsp40 co-chaperone localized to the endoplasmic reticulum of human cells. J. Biol. Chem. 2000, 275, 24984–24992. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.L.; Genereux, J.C.; Pankow, S.; Aerts, J.M.; Yates, J.R., 3rd; Kelly, J.W. ERdj3 is an endoplasmic reticulum degradation factor for mutant glucocerebrosidase variants linked to Gaucher’s disease. Chem. Biol. 2014, 21, 967–976. [Google Scholar] [CrossRef] [PubMed]
- Cornec-Le Gall, E.; Olson, R.J.; Besse, W.; Heyer, C.M.; Gainullin, V.G.; Smith, J.M.; Audrezet, M.P.; Hopp, K.; Porath, B.; Shi, B.; et al. Monoallelic Mutations to DNAJB11 Cause Atypical Autosomal-Dominant Polycystic Kidney Disease. Am. J. Hum. Genet. 2018, 102, 832–844. [Google Scholar] [CrossRef] [PubMed]
- Tousson-Abouelazm, N.; Papillon, J.; Guillemette, J.; Cybulsky, A.V. Urinary ERdj3 and mesencephalic astrocyte-derived neutrophic factor identify endoplasmic reticulum stress in glomerular disease. Lab. Investig. 2020, 100, 945–958. [Google Scholar] [CrossRef]
- Hsu, C.W.; Yu, J.S.; Peng, P.H.; Liu, S.C.; Chang, Y.S.; Chang, K.P.; Wu, C.C. Secretome profiling of primary cells reveals that THBS2 is a salivary biomarker of oral cavity squamous cell carcinoma. J. Proteome Res. 2014, 13, 4796–4807. [Google Scholar] [CrossRef] [PubMed]
- Wen, K.W.; Damania, B. Hsp90 and Hsp40/Erdj3 are required for the expression and anti-apoptotic function of KSHV K1. Oncogene 2010, 29, 3532–3544. [Google Scholar] [CrossRef]
- Grove, D.E.; Fan, C.Y.; Ren, H.Y.; Cyr, D.M. The endoplasmic reticulum-associated Hsp40 DNAJB12 and Hsc70 cooperate to facilitate RMA1 E3-dependent degradation of nascent CFTRDeltaF508. Mol. Biol. Cell 2011, 22, 301–314. [Google Scholar] [CrossRef]
- Sterrenberg, J.N.; Blatch, G.L.; Edkins, A.L. Human DNAJ in cancer and stem cells. Cancer Lett. 2011, 312, 129–142. [Google Scholar] [CrossRef] [PubMed]
- Farhan, S.M.K.; Howrigan, D.P.; Abbott, L.E.; Klim, J.R.; Topp, S.D.; Byrnes, A.E.; Churchhouse, C.; Phatnani, H.; Smith, B.N.; Rampersaud, E.; et al. Exome sequencing in amyotrophic lateral sclerosis implicates a novel gene, DNAJC7, encoding a heat-shock protein. Nat. Neurosci. 2019, 22, 1966–1974. [Google Scholar] [CrossRef] [PubMed]
- Roosen, D.A.; Blauwendraat, C.; Cookson, M.R.; Lewis, P.A. DNAJC proteins and pathways to parkinsonism. FEBS J. 2019, 286, 3080–3094. [Google Scholar] [CrossRef]
- Hundley, H.A.; Walter, W.; Bairstow, S.; Craig, E.A. Human Mpp11 J protein: Ribosome-tethered molecular chaperones are ubiquitous. Science 2005, 308, 1032–1034. [Google Scholar] [CrossRef]
- Aloia, L.; Demajo, S.; Di Croce, L. ZRF1: A novel epigenetic regulator of stem cell identity and cancer. Cell Cycle 2015, 14, 510–515. [Google Scholar] [CrossRef] [PubMed]
- Richly, H.; Rocha-Viegas, L.; Ribeiro, J.D.; Demajo, S.; Gundem, G.; Lopez-Bigas, N.; Nakagawa, T.; Rospert, S.; Ito, T.; Di Croce, L. Transcriptional activation of polycomb-repressed genes by ZRF1. Nature 2010, 468, 1124–1128. [Google Scholar] [CrossRef] [PubMed]
- Richly, H.; Di Croce, L. The flip side of the coin: Role of ZRF1 and histone H2A ubiquitination in transcriptional activation. Cell Cycle 2011, 10, 745–750. [Google Scholar] [CrossRef]
- Ribeiro, J.D.; Morey, L.; Mas, A.; Gutierrez, A.; Luis, N.M.; Mejetta, S.; Richly, H.; Benitah, S.A.; Keyes, W.M.; Di Croce, L. ZRF1 controls oncogene-induced senescence through the INK4-ARF locus. Oncogene 2013, 32, 2161–2168. [Google Scholar] [CrossRef]
- Demajo, S.; Uribesalgo, I.; Gutierrez, A.; Ballare, C.; Capdevila, S.; Roth, M.; Zuber, J.; Martin-Caballero, J.; Di Croce, L. ZRF1 controls the retinoic acid pathway and regulates leukemogenic potential in acute myeloid leukemia. Oncogene 2014, 33, 5501–5510. [Google Scholar] [CrossRef] [PubMed]
- Brychzy, A.; Rein, T.; Winklhofer, K.F.; Hartl, F.U.; Young, J.C.; Obermann, W.M. Cofactor Tpr2 combines two TPR domains and a J domain to regulate the Hsp70/Hsp90 chaperone system. EMBO J 2003, 22, 3613–3623. [Google Scholar] [CrossRef]
- Jih, K.Y.; Tsai, P.C.; Tsai, Y.S.; Liao, Y.C.; Lee, Y.C. Rapid progressive ALS in a patient with a DNAJC7 loss-of-function mutation. Neurol. Genet. 2020, 6, e503. [Google Scholar] [CrossRef]
- Hou, Z.; Wydorski, P.M.; Perez, V.A.; Mendoza-Oliva, A.; Ryder, B.D.; Mirbaha, H.; Kashmer, O.; Joachimiak, L.A. DnaJC7 binds natively folded structural elements in tau to inhibit amyloid formation. bioRxiv 2020, 12, 5338. [Google Scholar] [CrossRef]
- Moffatt, N.S.; Bruinsma, E.; Uhl, C.; Obermann, W.M.; Toft, D. Role of the cochaperone Tpr2 in Hsp90 chaperoning. Biochemistry 2008, 47, 8203–8213. [Google Scholar] [CrossRef] [PubMed]
- Xiang, S.L.; Kumano, T.; Iwasaki, S.I.; Sun, X.; Yoshioka, K.; Yamamoto, K.C. The J domain of Tpr2 regulates its interaction with the proapoptotic and cell-cycle checkpoint protein, Rad9. Biochem. Biophys. Res. Commun. 2001, 287, 932–940. [Google Scholar] [CrossRef]
- Hatakeyama, S.; Matsumoto, M.; Yada, M.; Nakayama, K.I. Interaction of U-box-type ubiquitin-protein ligases (E3s) with molecular chaperones. Genes Cells 2004, 9, 533–548. [Google Scholar] [CrossRef]
- Timsit, Y.E.; Negishi, M. Coordinated regulation of nuclear receptor CAR by CCRP/DNAJC7, HSP70 and the ubiquitin-proteasome system. PLoS ONE 2014, 9, e96092. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wang, J.; Hao, J.; Dong, B.; Li, Y.; Zhu, X.; Ding, J.; Ren, S.; Zhao, H.; Wu, S.; et al. Reduced cytosolic carboxypeptidase 6 (CCP6) level leads to accumulation of serum polyglutamylated DNAJC7 protein: A potential biomarker for renal cell carcinoma early detection. Oncotarget 2016, 7, 22385–22396. [Google Scholar] [CrossRef]
- Han, C.; Chen, T.; Li, N.; Yang, M.; Wan, T.; Cao, X. HDJC9, a novel human type C DnaJ/HSP40 member interacts with and cochaperones HSP70 through the J domain. Biochem. Biophys. Res. Commun. 2007, 353, 280–285. [Google Scholar] [CrossRef]
- Hammond, C.M.; Bao, H.; Hendriks, I.A.; Carraro, M.; García-Nieto, A.; Liu, Y.; Reverón-Gómez, N.; Spanos, C.; Chen, L.; Rappsilber, J.; et al. DNAJC9 integrates heat shock molecular chaperones into the histone chaperone network. Mol. Cell 2021, 81, 2533–2548.e2539. [Google Scholar] [CrossRef] [PubMed]
- Oliver, V.F.; van Bysterveldt, K.A.; Cadzow, M.; Steger, B.; Romano, V.; Markie, D.; Hewitt, A.W.; Mackey, D.A.; Willoughby, C.E.; Sherwin, T.; et al. A COL17A1 Splice-Altering Mutation Is Prevalent in Inherited Recurrent Corneal Erosions. Ophthalmology 2016, 123, 709–722. [Google Scholar] [CrossRef]
- Liu, C.M.; Fann, C.S.; Chen, C.Y.; Liu, Y.L.; Oyang, Y.J.; Yang, W.C.; Chang, C.C.; Wen, C.C.; Chen, W.J.; Hwang, T.J.; et al. ANXA7, PPP3CB, DNAJC9, and ZMYND17 genes at chromosome 10q22 associated with the subgroup of schizophrenia with deficits in attention and executive function. Biol. Psychiatry 2011, 70, 51–58. [Google Scholar] [CrossRef]
- Pan, Z.; Chen, S.; Pan, X.; Wang, Z.; Han, H.; Zheng, W.; Wang, X.; Li, F.; Qu, S.; Shao, R. Differential gene expression identified in Uigur women cervical squamous cell carcinoma by suppression subtractive hybridization. Neoplasma 2010, 57, 123–128. [Google Scholar] [CrossRef]
- Liu, X.; Li, K.; Wang, L.; Zhang, M.; Wang, X. Variations of human heat shock proteins in multiple cancers. Clin. Transl. Med. 2021, 11, e320. [Google Scholar] [CrossRef] [PubMed]
- Mandriani, B.; Castellana, S.; Rinaldi, C.; Manzoni, M.; Venuto, S.; Rodriguez-Aznar, E.; Galceran, J.; Nieto, M.A.; Borsani, G.; Monti, E.; et al. Identification of p53-target genes in Danio rerio. Sci. Rep. 2016, 6, 32474. [Google Scholar] [CrossRef]
- Anikster, Y.; Haack, T.B.; Vilboux, T.; Pode-Shakked, B.; Thony, B.; Shen, N.; Guarani, V.; Meissner, T.; Mayatepek, E.; Trefz, F.K.; et al. Biallelic Mutations in DNAJC12 Cause Hyperphenylalaninemia, Dystonia, and Intellectual Disability. Am. J. Hum. Genet. 2017, 100, 257–266. [Google Scholar] [CrossRef]
- Li, M.; Yang, Q.; Yi, S.; Qin, Z.; Luo, J.; Fan, X. Two novel mutations in DNAJC12 identified by whole-exome sequencing in a patient with mild hyperphenylalaninemia. Mol. Genet. Genom. Med. 2020, 8, e1303. [Google Scholar] [CrossRef]
- Gallego, D.; Leal, F.; Gámez, A.; Castro, M.; Navarrete, R.; Sanchez-Lijarcio, O.; Vitoria, I.; Bueno-Delgado, M.; Belanger-Quintana, A.; Morais, A.; et al. Pathogenic variants of DNAJC12 and evaluation of the encoded cochaperone as a genetic modifier of hyperphenylalaninemia. Hum. Mutat. 2020, 41, 1329–1338. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Yang, Z.H.; Li, F.; Hu, X.C.; Yue, Y.W.; Yang, J.; Liu, Y.T.; Liu, H.; Wang, Y.L.; Shi, C.H.; et al. DNAJC12 mutation is rare in Chinese Han population with Parkinson’s disease. Neurobiol. Aging. 2018, 68, e151–e159. [Google Scholar] [CrossRef]
- He, H.L.; Lee, Y.E.; Chen, H.P.; Hsing, C.H.; Chang, I.W.; Shiue, Y.L.; Lee, S.W.; Hsu, C.T.; Lin, L.C.; Wu, T.F.; et al. Overexpression of DNAJC12 predicts poor response to neoadjuvant concurrent chemoradiotherapy in patients with rectal cancer. Exp. Mol. Pathol. 2015, 98, 338–345. [Google Scholar] [CrossRef]
- Uno, Y.; Kanda, M.; Miwa, T.; Umeda, S.; Tanaka, H.; Tanaka, C.; Kobayashi, D.; Suenaga, M.; Hattori, N.; Hayashi, M.; et al. Increased Expression of DNAJC12 is Associated with Aggressive Phenotype of Gastric Cancer. Ann. Surg. Oncol. 2019, 26, 836–844. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, M.; Jin, F.; Liu, J.; Chen, M.; Yin, J. DNAJC12 promotes lung cancer growth by regulating the activation of β-catenin. Int. J. Mol. Med. 2021, 47, 105. [Google Scholar] [CrossRef]
- Hatle, K.M.; Gummadidala, P.; Navasa, N.; Bernardo, E.; Dodge, J.; Silverstrim, B.; Fortner, K.; Burg, E.; Suratt, B.T.; Hammer, J.; et al. MCJ/DnaJC15, an endogenous mitochondrial repressor of the respiratory chain that controls metabolic alterations. Mol. Cell Biol. 2013, 33, 2302–2314. [Google Scholar] [CrossRef]
- Sinha, D.; D’Silva, P. Chaperoning mitochondrial permeability transition: Regulation of transition pore complex by a J-protein, DnaJC15. Cell Death Dis. 2014, 5, e1101. [Google Scholar] [CrossRef]
- Lindsey, J.C.; Lusher, M.E.; Strathdee, G.; Brown, R.; Gilbertson, R.J.; Bailey, S.; Ellison, D.W.; Clifford, S.C. Epigenetic inactivation of MCJ (DNAJD1) in malignant paediatric brain tumours. Int. J. Cancer 2006, 118, 346–352. [Google Scholar] [CrossRef]
- Lau, D.T.; Hesson, L.B.; Norris, M.D.; Marshall, G.M.; Haber, M.; Ashton, L.J. Prognostic significance of promoter DNA methylation in patients with childhood neuroblastoma. Clin. Cancer Res. 2012, 18, 5690–5700. [Google Scholar] [CrossRef]
- Schusdziarra, C.; Blamowska, M.; Azem, A.; Hell, K. Methylation-controlled J-protein MCJ acts in the import of proteins into human mitochondria. Hum. Mol. Genet. 2013, 22, 1348–1357. [Google Scholar] [CrossRef] [PubMed]
- Alessandrini, F.; Pezze, L.; Menendez, D.; Resnick, M.A.; Ciribilli, Y. ETV7-Mediated DNAJC15 Repression Leads to Doxorubicin Resistance in Breast Cancer Cells. Neoplasia 2018, 20, 857–870. [Google Scholar] [CrossRef] [PubMed]
- Witham, J.; Vidot, S.; Agarwal, R.; Kaye, S.B.; Richardson, A. Transient ectopic expression as a method to detect genes conferring drug resistance. Int. J. Cancer 2008, 122, 2641–2645. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Cabezudo, M.J.; Faour, I.; Jones, K.; Champagne, D.P.; Jaloudi, M.A.; Mohamed, Y.A.; Bashir, G.; Almarzooqi, S.; Albawardi, A.; Hashim, M.J.; et al. Deficiency of mitochondrial modulator MCJ promotes chemoresistance in breast cancer. JCI Insight 2016, 1, e86873. [Google Scholar] [CrossRef] [PubMed]
- Hatle, K.M.; Neveu, W.; Dienz, O.; Rymarchyk, S.; Barrantes, R.; Hale, S.; Farley, N.; Lounsbury, K.M.; Bond, J.P.; Taatjes, D.; et al. Methylation-controlled J protein promotes c-Jun degradation to prevent ABCB1 transporter expression. Mol. Cell Biol. 2007, 27, 2952–2966. [Google Scholar] [CrossRef]
- Liu, T.; Jiang, W.; Han, D.; Yu, L. DNAJC25 is downregulated in hepatocellular carcinoma and is a novel tumor suppressor gene. Oncol. Lett. 2012, 4, 1274–1280. [Google Scholar] [CrossRef]
- Acun, T. DNAJC25 as a tumor suppressor candidate gene in breast cancer. Hacet. J. Biol. Chem. 2021, 49, 305. [Google Scholar] [CrossRef]
- Pinhasi-Kimhi, O.; Michalovitz, D.; Ben-Zeev, A.; Oren, M. Specific interaction between the p53 cellular tumour antigen and major heat shock proteins. Nature 1986, 320, 182–184. [Google Scholar] [CrossRef] [PubMed]
- Hinds, P.W.; Finlay, C.A.; Frey, A.B.; Levine, A.J. Immunological evidence for the association of p53 with a heat shock protein, hsc70, in p53-plus-ras-transformed cell lines. Mol. Cell Biol. 1987, 7, 2863–2869. [Google Scholar] [CrossRef]
- Hansen, S.; Hupp, T.R.; Lane, D.P. Allosteric regulation of the thermostability and DNA binding activity of human p53 by specific interacting proteins. CRC Cell Transformation Group. J. Biol. Chem. 1996, 271, 3917–3924. [Google Scholar] [CrossRef]
- Walerych, D.; Olszewski, M.B.; Gutkowska, M.; Helwak, A.; Zylicz, M.; Zylicz, A. Hsp70 molecular chaperones are required to support p53 tumor suppressor activity under stress conditions. Oncogene 2009, 28, 4284–4294. [Google Scholar] [CrossRef]
- Walerych, D.; Kudla, G.; Gutkowska, M.; Wawrzynow, B.; Muller, L.; King, F.W.; Helwak, A.; Boros, J.; Zylicz, A.; Zylicz, M. Hsp90 chaperones wild-type p53 tumor suppressor protein. J. Biol. Chem. 2004, 279, 48836–48845. [Google Scholar] [CrossRef] [PubMed]
- Boysen, M.; Kityk, R.; Mayer, M.P. Hsp70- and Hsp90-Mediated Regulation of the Conformation of p53 DNA Binding Domain and p53 Cancer Variants. Mol. Cell 2019, 74, 831–843.e834. [Google Scholar] [CrossRef]
- Trinidad, A.G.; Muller, P.A.; Cuellar, J.; Klejnot, M.; Nobis, M.; Valpuesta, J.M.; Vousden, K.H. Interaction of p53 with the CCT complex promotes protein folding and wild-type p53 activity. Mol. Cell 2013, 50, 805–817. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Deisseroth, A.B. Conformational change of p53 protein in growth factor-stimulated human myelogenous leukemia cells. Leuk. Lymphoma 1994, 14, 251–255. [Google Scholar] [CrossRef]
- Zheng, A.; Castren, K.; Saily, M.; Savolainen, E.R.; Koistinen, P.; Vahakangas, K. p53 status of newly established acute myeloid leukaemia cell lines. Br. J. Cancer 1999, 79, 407–415. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Arandkar, S.; Furth, N.; Elisha, Y.; Nataraj, N.B.; van der Kuip, H.; Yarden, Y.; Aulitzky, W.; Ulitsky, I.; Geiger, B.; Oren, M. Altered p53 functionality in cancer-associated fibroblasts contributes to their cancer-supporting features. Proc. Natl. Acad. Sci. USA 2018, 115, 6410–6415. [Google Scholar] [CrossRef]
- Benor, G.; Fuks, G.; Chin, S.F.; Rueda, O.M.; Mukherjee, S.; Arandkar, S.; Aylon, Y.; Caldas, C.; Domany, E.; Oren, M. Transcriptional profiling reveals a subset of human breast tumors that retain wt TP53 but display mutant p53-associated features. Mol. Oncol. 2020, 14, 1640–1652. [Google Scholar] [CrossRef]
- Tuval, A.B.Y.; Azogy, H.; Moshkovitz, Y.; Biezuner, T.; Leshkowitz, D.; Salame, T.M.; Minden, M.D.; Tal, P.; Rotter, V.; Oren, M.; et al. Pseudo-mutant p53 as a targetable phenotype of DNMT3A-mutated pre-leukemia. bioRxiv 2021. [Google Scholar] [CrossRef]
- Akakura, S.; Yoshida, M.; Yoneda, Y.; Horinouchi, S. A role for Hsc70 in regulating nucleocytoplasmic transport of a temperature-sensitive p53 (p53Val-135). J. Biol. Chem. 2001, 276, 14649–14657. [Google Scholar] [CrossRef] [PubMed]
- Nikolova, P.V.; Wong, K.B.; DeDecker, B.; Henckel, J.; Fersht, A.R. Mechanism of rescue of common p53 cancer mutations by second-site suppressor mutations. EMBO J. 2000, 19, 370–378. [Google Scholar] [CrossRef]
- Gomes, S.; Bosco, B.; Loureiro, J.B.; Ramos, H.; Raimundo, L.; Soares, J.; Nazareth, N.; Barcherini, V.; Domingues, L.; Oliveira, C.; et al. SLMP53-2 Restores Wild-Type-Like Function to Mutant p53 through Hsp70: Promising Activity in Hepatocellular Carcinoma. Cancers 2019, 11, 1151. [Google Scholar] [CrossRef]
- Selivanova, G.; Kawasaki, T.; Ryabchenko, L.; Wiman, K.G. Reactivation of mutant p53: A new strategy for cancer therapy. Semin. Cancer Biol. 1998, 8, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Rokaeus, N.; Klein, G.; Wiman, K.G.; Szekely, L.; Mattsson, K. PRIMA-1(MET) induces nucleolar accumulation of mutant p53 and PML nuclear body-associated proteins. Oncogene 2007, 26, 982–992. [Google Scholar] [CrossRef] [PubMed]
| HSP40/JDPs | Type of p53 Proteins Interacting with HSP40/JDPs | Effects on p53 | Dependency of HSP70 (UD: Undetermined) | References |
|---|---|---|---|---|
| DNAJA1/HDJ2 | p53R156P, p53V157F, p53R175H, p53C176F, p53Y220C | DNAJA1/HDJ2 promotes oncogenic functions (cell migration, colony-forming potential) by interacting with and stabilizing misfolded mutp53. It should be noted that p53R175H levels are not affected by HSC70 knockdown in CAL33 cells. | Stabilization of p53R175H could be HSC70-independent, but the dependency on HSP70 needs to be clarified. | [23,55,64,65] |
| p53R158L | DNAJA1/HDJ2 confers radio-resistance in SF763 cells. | UD | [67] | |
| DNAJA3/Tid1 | wtp53 | Hypoxic and genotoxic stresses result in formation of a complex of DNAJA3/Tid1 and wtp53 through the J domain to promote p53 mitochondrial localization and transcription-independent apoptosis. | UD | [68,69] |
| DNAJB1/HDJ1 | wtp53 (via MDM2) | The C-terminal region of DNAJB1/HDJ1 binds to, stabilizes, and inhibits MDM2, thus indirectly upregulating wtp53 activity. | Independent | [41] |
| wtp53 (via PDCD5) | DNAJB1/HDJ1 inhibits wtp53’s apoptotic function through interaction with PDCD5 and contributes to cancer progression. | UD | [70] | |
| p53R175H | DNAJB1/HDJ1 interacts with conformational mutp53 (R175H), TAp73, and HSP70. This complex enhances chemo-resistance. | UD | [24] | |
| p53R175H | The interaction of DNAJB1/HDJ1 with p53R175H is enhanced by chetomin, which restores the wtp53-like activity specifically in TOV-112D and CAL33 cells. | UD | [71] | |
| DNAJB9/MDG1/ERdj4 | wtp53 | DNAJB9/MDG1/ERdj4 inhibits apoptosis under genotoxic stress by interacting with wtp53 through the J domain. | UD | [72] |
| wtp53 | The interaction of DNAJB9/MDG1/ERdj4 with p53 inhibits the H-RASV12-induced senescence and promotes transformation in MEFs. | UD | [73] | |
| wtp53 | Site-specific mutations in the DNAJB9/MDG1/ERdj4 gene in JEG-3 cells reduce DNAJB9/MDG1/ERdj4 protein levels with increased wtp53 levels. | UD | [74] | |
| DNAJC2/ZRF1 | wtp53 | DNAJC2/ZRF1 inhibits wtp53’s activity in human gastric cancer cell lines carrying wtp53, but not p53 mutations or p53 null. | UD | [75] |
| DNAJC7/TPR2 | wtp53 | DNAJC7/TPR2 inhibits colony-forming potential by stabilizing wtp53 in H1299 cells exogenously expressing wtp53. | UD | [76] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaida, A.; Iwakuma, T. Regulation of p53 and Cancer Signaling by Heat Shock Protein 40/J-Domain Protein Family Members. Int. J. Mol. Sci. 2021, 22, 13527. https://doi.org/10.3390/ijms222413527
Kaida A, Iwakuma T. Regulation of p53 and Cancer Signaling by Heat Shock Protein 40/J-Domain Protein Family Members. International Journal of Molecular Sciences. 2021; 22(24):13527. https://doi.org/10.3390/ijms222413527
Chicago/Turabian StyleKaida, Atsushi, and Tomoo Iwakuma. 2021. "Regulation of p53 and Cancer Signaling by Heat Shock Protein 40/J-Domain Protein Family Members" International Journal of Molecular Sciences 22, no. 24: 13527. https://doi.org/10.3390/ijms222413527
APA StyleKaida, A., & Iwakuma, T. (2021). Regulation of p53 and Cancer Signaling by Heat Shock Protein 40/J-Domain Protein Family Members. International Journal of Molecular Sciences, 22(24), 13527. https://doi.org/10.3390/ijms222413527






