Molecular and Cellular Mechanisms Influenced by Postbiotics
Abstract
1. Introduction
2. The Impact of Postbiotics on Immunomodulatory Pathways
2.1. NF-κB and MAPK Pathways
2.1.1. Molecular Mechanism of NF-κB Signalling
2.1.2. MAPK Signalling Pathway
2.1.3. Postbiotics’ Influence on NF-κB and MAPK Pathways
2.2. Other Mechanisms and Molecular Pathways
3. Molecular Targets of Postbiotics
3.1. Toll-like Receptors
3.1.1. TLR Ligands: LTA
3.1.2. TLR Ligands: Flagellins
3.1.3. TLR Ligands: Other Cell Wall-Associated Proteins and Fimbriae
3.1.4. TLR Ligands: Oligonucleotides
3.1.5. TLR Ligands: Bacteriocins
3.1.6. TLR—Not Only Intestine Receptors
3.2. NOD like Receptors (NLR)
3.2.1. NOD and the Microbiome Crosstalk
3.2.2. Other Regulatory Roles of NOD Receptors
3.2.3. NOD and Postbiotic Molecules
- The polysaccharide-PGN complex (connected by phosphodiester bond), described for the first time in L. casei Shirota (LcS) [278], suppresses IL-6 production via the inhibition of NF- κB phosphorylation and enhances NOD2 mRNA levels in RAW264 macrophages in vitro [279]. Furthermore, it provides resistance to lysis, which determines amplified IL-12 production in macrophages. This effect is observed only for intact cell walls consisting of PGN and cell wall polysaccharides. Purified PGN has a weak ability to induce IL-12 production, whereas minimal structural units of PGN (6-O-stearoyl-muramyl dipeptide) abolish the effect of intact cell walls [247].
3.2.4. Inflammasomes and Postbiotic Molecules
3.2.5. NLR and TLR Interplay
3.3. Other Receptors
C-Type Lectin Receptor (CLR)—DC-SIGN
4. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Tierney, B.T.; Yang, Z.; Luber, J.M.; Beaudin, M.; Wibowo, M.C.; Baek, C.; Mehlenbacher, E.; Patel, C.J.; Kostic, A.D. The Landscape of Genetic Content in the Gut and Oral Human Microbiome. Cell Host. Microbe 2019, 26, 283–295.e8. [Google Scholar] [CrossRef]
- Creely, S.J.; McTernan, P.G.; Kusminski, C.M.; Fisher, M.; Da Silva, N.F.; Khanolkar, M.; Evans, M.; Harte, A.L.; Kumar, S. Lipopolysaccharide activates an innate immune system response in human adipose tissue in obesity and type 2 diabetes. Am. J. Physiol. Endocrinol. Metab. 2007, 292, E740–E747. [Google Scholar] [CrossRef] [PubMed]
- Corrêa-Oliveira, R.; Fachi, J.L.; Vieira, A.; Sato, F.T.; Vinolo, M.A.R. Regulation of immune cell function by short-chain fatty acids. Clin. Transl. Immunol. 2016, 5, e73. [Google Scholar] [CrossRef]
- Cortés-Martín, A.; Selma, M.V.; Tomás-Barberán, F.A.; González-Sarrías, A.; Espín, J.C. Where to Look into the Puzzle of Polyphenols and Health? The Postbiotics and Gut Microbiota Associated with Human Metabotypes. Mol. Nutr. Food Res. 2020, 64, e1900952. [Google Scholar] [CrossRef] [PubMed]
- Ostaff, M.J.; Stange, E.F.; Wehkamp, J. Antimicrobial peptides and gut microbiota in homeostasis and pathology: Homeostasis in the gut. EMBO Mol. Med 2013, 5, 1465–1483. [Google Scholar] [CrossRef]
- Kubinak, J.L.; Stephens, W.Z.; Soto, R.; Petersen, C.; Chiaro, T.; Gogokhia, L.; Bell, R.; Ajami, N.J.; Petrosino, J.F.; Morrison, L.; et al. MHC variation sculpts individualized microbial communities that control susceptibility to enteric infection. Nat. Commun. 2015, 6, 8642. [Google Scholar] [CrossRef] [PubMed]
- Vieira, A.T.; Fukumori, C.; Ferreira, C.M. New insights into therapeutic strategies for gut microbiota modulation in inflammatory diseases. Clin. Transl. Immunol. 2016, 5, e87. [Google Scholar] [CrossRef]
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G.R.; Merenstein, D.J.; Pot, B.; Morelli, L.; Canani, R.B.; Flint, H.J.; Salminen, S.; et al. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506–514. [Google Scholar] [CrossRef] [PubMed]
- Azad, M.A.K.; Sarker, M.; Wan, D. Immunomodulatory Effects of Probiotics on Cytokine Profiles. Biomed. Res. Int. 2018, 2018, 8063647. [Google Scholar] [CrossRef] [PubMed]
- Ganji-Arjenaki, M.; Rafieian-Kopaei, M. Probiotics are a good choice in remission of inflammatory bowel diseases: A meta analysis and systematic review. J. Cell Physiol. 2018, 233, 2091–2103. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.A.; Kim, J. Effect of Probiotics on Blood Lipid Concentrations: A Meta-Analysis of Randomized Controlled Trials. Medicine 2015, 94, e1714. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Bae, J.-H. Probiotics for weight loss: A systematic review and meta-analysis. Nutr. Res. 2015, 35, 566–575. [Google Scholar] [CrossRef] [PubMed]
- Qi, D.; Nie, X.-L.; Zhang, J.-J. The effect of probiotics supplementation on blood pressure: A systemic review and meta-analysis. Lipids Health Dis. 2020, 19, 79. [Google Scholar] [CrossRef]
- Drago, L. Probiotics and Colon Cancer. Microorganisms 2019, 7, 66. [Google Scholar] [CrossRef]
- Akbari, E.; Asemi, Z.; Daneshvar Kakhaki, R.; Bahmani, F.; Kouchaki, E.; Tamtaji, O.R.; Hamidi, G.A.; Salami, M. Effect of Probiotic Supplementation on Cognitive Function and Metabolic Status in Alzheimer’s Disease: A Randomized, Double-Blind and Controlled Trial. Front. Aging. Neurosci. 2016, 8, 256. [Google Scholar] [CrossRef]
- Li, W.; Guo, J.; Shen, Y.; Huang, L.; Leng, B.; Fan, D.; Shui, L.; Chen, C. Probiotics, prebiotics, and synbiotics for the treatment of dementia: Protocol for a systematic review. Medicine 2020, 99, e18608. [Google Scholar] [CrossRef]
- Mishra, V.; Shah, C.; Mokashe, N.; Chavan, R.; Yadav, H.; Prajapati, J. Probiotics as potential antioxidants: A systematic review. J. Agric. Food Chem. 2015, 63, 3615–3626. [Google Scholar] [CrossRef]
- Doron, S.; Snydman, D.R. Risk and safety of probiotics. Clin. Infect. Dis. 2015, 60, S129–S134. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.; Zhang, R.; Tian, X.; Zhou, X.; Pan, X.; Wong, A. Assessing the Risk of Probiotic Dietary Supplements in the Context of Antibiotic Resistance. Front. Microbiol. 2017, 8, 908. [Google Scholar] [CrossRef]
- Imperial, I.C.V.J.; Ibana, J.A. Addressing the Antibiotic Resistance Problem with Probiotics: Reducing the Risk of Its Double-Edged Sword Effect. Front. Microbiol. 2016, 7, 1983. [Google Scholar] [CrossRef]
- Cicenia, A.; Scirocco, A.; Carabotti, M.; Pallotta, L.; Marignani, M.; Severi, C. Postbiotic activities of lactobacilli-derived factors. J. Clin. Gastroenterol. 2014, 48, S18–S22. [Google Scholar] [CrossRef]
- Suez, J.; Elinav, E. The path towards microbiome-based metabolite treatment. Nat. Microbiol. 2017, 2, 17075. [Google Scholar] [CrossRef] [PubMed]
- Salminen, S.; Collado, M.C.; Endo, A.; Hill, C.; Lebeer, S.; Quigley, E.M.M.; Sanders, M.E.; Shamir, R.; Swann, J.R.; Szajewska, H.; et al. The International Scientific Association of Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of postbiotics. Nat. Rev. Gastroenterol. Hepatol. 2021, 18, 1–19. [Google Scholar] [CrossRef]
- Taverniti, V.; Guglielmetti, S. The immunomodulatory properties of probiotic microorganisms beyond their viability (ghost probiotics: Proposal of paraprobiotic concept). Genes Nutr. 2011, 6, 261–274. [Google Scholar] [CrossRef]
- Aguilar-Toalá, J.E.; Garcia-Varela, R.; Garcia, H.S.; Mata-Haro, V.; González-Córdova, A.F.; Vallejo-Cordoba, B.; Hernán-dez-Mendoza, A. Postbiotics: An evolving term within the functional foods field. Trends Food Sci. Technol. 2018, 75, 105–114. [Google Scholar] [CrossRef]
- Wegh, C.A.M.; Geerlings, S.Y.; Knol, J.; Roeselers, G.; Belzer, C. Postbiotics and Their Potential Applications in Early Life Nutrition and Beyond. Int. J. Mol. Sci. 2019, 20, 4673. [Google Scholar] [CrossRef]
- Mayorgas, A.; Dotti, I.; Salas, A. Microbial Metabolites, Postbiotics, and Intestinal Epithelial Function. Mol. Nutr. Food Res. 2021, 65, e2000188. [Google Scholar] [CrossRef]
- Nataraj, B.H.; Ali, S.A.; Behare, P.V.; Yadav, H. Postbiotics-parabiotics: The new horizons in microbial biotherapy and functional foods. Microb. Cell Fact 2020, 19, 168. [Google Scholar] [CrossRef]
- Dicks, L.M.T.; Dreyer, L.; Smith, C.; van Staden, A.D. A Review: The Fate of Bacteriocins in the Human Gastro-Intestinal Tract: Do They Cross the Gut–Blood Barrier? Front. Microbiol. 2018, 9, 2297. [Google Scholar] [CrossRef]
- Hörmannsperger, G.; von Schillde, M.-A.; Haller, D. Lactocepin as a protective microbial structure in the context of IBD. Gut Microbes 2013, 4, 152–157. [Google Scholar] [CrossRef]
- von Schillde, M.-A.; Hörmannsperger, G.; Weiher, M.; Alpert, C.-A.; Hahne, H.; Bäuerl, C.; van Huynegem, K.; Steidler, L.; Hrncir, T.; Pérez-Martínez, G.; et al. Lactocepin secreted by Lactobacillus exerts anti-inflammatory effects by selectively degrading proinflammatory chemokines. Cell Host Microbe 2012, 11, 387–396. [Google Scholar] [CrossRef]
- Zhang, L.; Li, N.; Caicedo, R.; Neu, J. Alive and Dead Lactobacillus Rhamnosus GG Decrease Tumor Necrosis Factor-Alpha-Induced Interleukin-8 Production in Caco-2 Cells. J. Nutr. 2005, 135, 1752–1756. [Google Scholar] [CrossRef]
- Ivanov, D.; Emonet, C.; Foata, F.; Affolter, M.; Delley, M.; Fisseha, M.; Blum-Sperisen, S.; Kochhar, S.; Arigoni, F. A Serpin from the Gut Bacterium Bifidobacterium Longum Inhibits Eukaryotic Elastase-like Serine Proteases. J. Biol. Chem. 2006, 281, 17246–17252. [Google Scholar] [CrossRef]
- Hidalgo-Cantabrana, C.; Moro-García, M.A.; Blanco-Míguez, A.; Fdez-Riverola, F.; Oliván, M.; Royo, L.J.; Riestra, S.; Margolles, A.; Lourenço, A.; Alonso-Arias, R.; et al. The Extracellular Proteins of Lactobacillus Acidophilus DSM 20079T Display Anti-Inflammatory Effect in Both in Piglets, Healthy Human Donors and Crohn’s Disease Patients. J. Funct. Foods 2020, 64, 103660. [Google Scholar] [CrossRef]
- Choi, J.H.; Moon, C.M.; Shin, T.-S.; Kim, E.K.; McDowell, A.; Jo, M.-K.; Joo, Y.H.; Kim, S.-E.; Jung, H.-K.; Shim, K.-N.; et al. Lactobacillus Paracasei-Derived Extracellular Vesicles Attenuate the Intestinal Inflammatory Response by Augmenting the Endoplasmic Reticulum Stress Pathway. Exp. Mol. Med. 2020, 52, 423–437. [Google Scholar] [CrossRef]
- Patil, S.; Sawant, S.; Hauff, K.; Hampp, G. Validated Postbiotic Screening Confirms Presence of Physiologically-Active Metabolites, Such as Short-Chain Fatty Acids, Amino Acids and Vitamins in Hylak® Forte. Probiotics Antimicrob. Proteins 2019, 11, 1124–1131. [Google Scholar] [CrossRef]
- Ewaschuk, J.B.; Walker, J.W.; Diaz, H.; Madsen, K.L. Bioproduction of Conjugated Linoleic Acid by Probiotic Bacteria Occurs in Vitro and in Vivo in Mice. J. Nutr. 2006, 136, 1483–1487. [Google Scholar] [CrossRef]
- Kepert, I.; Fonseca, J.; Müller, C.; Milger, K.; Hochwind, K.; Kostric, M.; Fedoseeva, M.; Ohnmacht, C.; Dehmel, S.; Nathan, P.; et al. D-Tryptophan from Probiotic Bacteria Influences the Gut Microbiome and Allergic Airway Disease. J. Allergy Clin. Immunol. 2017, 139, 1525–1535. [Google Scholar] [CrossRef]
- Venkatesh, M.; Mukherjee, S.; Wang, H.; Li, H.; Sun, K.; Benechet, A.P.; Qiu, Z.; Maher, L.; Redinbo, M.R.; Phillips, R.S.; et al. Symbiotic Bacterial Metabolites Regulate Gastrointestinal Barrier Function via the Xenobiotic Sensor PXR and Toll-like Receptor 4. Immunity 2014, 41, 296–310. [Google Scholar] [CrossRef]
- Ehrlich, A.M.; Henrick, B.; Pacheco, A.; Taft, D.; Xu, G.; Huda, N.; Lozada-Contreras, M.; Goodson, M.; Slupsky, C.; Mills, D.; et al. Bifidobacterium Grown on Human Milk Oligosaccharides Produce Tryptophan Metabolite Indole-3-Lactic Acid That Significantly Decreases Inflammation in Intestinal Cells in Vitro. FASEB J. 2018, 32, lb359. [Google Scholar] [CrossRef]
- Segawa, S.; Fujiya, M.; Konishi, H.; Ueno, N.; Kobayashi, N.; Shigyo, T.; Kohgo, Y. Probiotic-Derived Polyphosphate Enhances the Epithelial Barrier Function and Maintains Intestinal Homeostasis through Integrin–p38 MAPK Pathway. PLoS ONE 2011, 6, e23278. [Google Scholar] [CrossRef]
- Feng, G.; Feng, Y.; Guo, T.; Yang, Y.; Guo, W.; Huang, M.; Wu, H.; Zeng, M. Biogenic Polyphosphate Nanoparticles from Synechococcus Sp. PCC 7002 Exhibit Intestinal Protective Potential in Human Intestinal Epithelial Cells In Vitro and Murine Small Intestine Ex Vivo. J. Agric. Food Chem. 2018, 66, 8026–8035. [Google Scholar] [CrossRef]
- Kato-Mori, Y.; Orihashi, T.; Kanai, Y.; Sato, M.; Sera, K.; Hagiwara, K. Fermentation Metabolites from Lactobacillus Gasseri and Propionibacterium Freudenreichii Exert Bacteriocidal Effects in Mice. J. Med. Food 2010, 13, 1460–1467. [Google Scholar] [CrossRef]
- Johnson, B.; Selle, K.; O’Flaherty, S.; Goh, Y.J.; Klaenhammer, T. Identification of Extracellular Surface-Layer Associated Proteins in Lactobacillus Acidophilus NCFM. Microbiology 2013, 159, 2269–2282. [Google Scholar] [CrossRef]
- Taverniti, V.; Stuknyte, M.; Minuzzo, M.; Arioli, S.; De Noni, I.; Scabiosi, C.; Cordova, Z.M.; Junttila, I.; Hämäläinen, S.; Turpeinen, H.; et al. S-Layer Protein Mediates the Stimulatory Effect of Lactobacillus Helveticus MIMLh5 on Innate Immunity. Appl. Environ. Microbiol. 2013, 79, 1221–1231. [Google Scholar] [CrossRef]
- Prado Acosta, M.; Ruzal, S.M.; Cordo, S.M. S-Layer Proteins from Lactobacillus Sp. Inhibit Bacterial Infection by Blockage of DC-SIGN Cell Receptor. Int. J. Biol. Macromol. 2016, 92, 998–1005. [Google Scholar] [CrossRef]
- Prado Acosta, M.; Geoghegan, E.M.; Lepenies, B.; Ruzal, S.; Kielian, M.; Martinez, M.G. Surface (S) Layer Proteins of Lactobacillus Acidophilus Block Virus Infection via DC-SIGN Interaction. Front. Microbiol. 2019, 10, 810. [Google Scholar] [CrossRef]
- Kobatake, E.; Kabuki, T. S-Layer Protein of Lactobacillus Helveticus SBT2171 Promotes Human β-Defensin 2 Expression via TLR2–JNK Signaling. Front. Microbiol. 2019, 10, 2414. [Google Scholar] [CrossRef]
- Lebeer, S.; Claes, I.J.J.; Vanderleyden, J. Anti-Inflammatory Potential of Probiotics: Lipoteichoic Acid Makes a Difference. Trends Microbiol. 2012, 20, 5–10. [Google Scholar] [CrossRef]
- Macho Fernandez, E.; Valenti, V.; Rockel, C.; Hermann, C.; Pot, B.; Boneca, I.G.; Grangette, C. Anti-Inflammatory Capacity of Selected Lactobacilli in Experimental Colitis Is Driven by NOD2-Mediated Recognition of a Specific Peptidoglycan-Derived Muropeptide. Gut 2011, 60, 1050–1059. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Qiao, Y.; Peng, Q.; Shi, B.; Dia, V.P. Antioxidant and Immunomodulatory Properties of Partially Purified Exopolysaccharide from Lactobacillus Casei Isolated from Chinese Northeast Sauerkraut. Immunol. Investig. 2021, 1–18. [Google Scholar] [CrossRef]
- Górska, S.; Hermanova, P.; Ciekot, J.; Schwarzer, M.; Srutkova, D.; Brzozowska, E.; Kozakova, H.; Gamian, A. Chemical Characterization and Immunomodulatory Properties of Polysaccharides Isolated from Probiotic Lactobacillus Casei LOCK 0919. Glycobiology 2017, 27, 275–277. [Google Scholar]
- Fanning, S.; Hall, L.J.; van Sinderen, D. Bifidobacterium Breve UCC2003 Surface Exopolysaccharide Production Is a Beneficial Trait Mediating Commensal-Host Interaction through Immune Modulation and Pathogen Protection. Gut Microbes 2012, 3, 420–425. [Google Scholar] [CrossRef]
- Górska, S.; Schwarzer, M.; Jachymek, W.; Srutkova, D.; Brzozowska, E.; Kozakova, H.; Gamian, A. Distinct Immunomodulation of Bone Marrow-Derived Dendritic Cell Responses to Lactobacillus Plantarum WCFS1 by Two Different Polysaccharides Isolated from Lactobacillus Rhamnosus LOCK 0900. Appl. Environ. Microbiol. 2014, 80, 6506–6516. [Google Scholar] [CrossRef]
- Ai, L.; Zhang, H.; Guo, B.; Chen, W.; Wu, Z.; Wu, Y. Preparation, Partial Characterization and Bioactivity of Exopolysaccharides from Lactobacillus Casei LC2W. Carbohydr. Polym. 2008, 74, 353–357. [Google Scholar] [CrossRef]
- Gao, K.; Wang, C.; Liu, L.; Dou, X.; Liu, J.; Yuan, L.; Zhang, W.; Wang, H. Immunomodulation and Signaling Mechanism of Lactobacillus Rhamnosus GG and Its Components on Porcine Intestinal Epithelial Cells Stimulated by Lipopolysaccharide. J. Microbiol. Immunol. Infect. 2017, 50, 700–713. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhou, Z.; Li, Y.; Zhou, L.; Ding, Q.; Xu, L. Isolated Exopolysaccharides from Lactobacillus Rhamnosus GG Alleviated Adipogenesis Mediated by TLR2 in Mice. Sci. Rep. 2016, 6, 36083. [Google Scholar] [CrossRef]
- Agostoni, C.; Goulet, O.; Kolacek, S.; Koletzko, B.; Moreno, L.; Puntis, J.; Rigo, J.; Shamir, R.; Szajewska, H.; Turck, D. Fermented Infant Formulae Without Live Bacteria. J. Pediatr. Gastroenterol. Nutr. 2007, 44, 392–397. [Google Scholar] [CrossRef]
- Szajewska, H.; Skórka, A.; Pieścik-Lech, M. Fermented Infant Formulas without Live Bacteria: A Systematic Review. Eur. J. Pediatr. 2015, 174, 1413–1420. [Google Scholar] [CrossRef]
- Jeong, K.; Kim, M.; Jeon, S.A.; Kim, Y.-H.; Lee, S. A Randomized Trial of Lactobacillus Rhamnosus IDCC 3201 Tyndallizate (RHT3201) for Treating Atopic Dermatitis. Pediatr. Allergy Immunol. 2020. [Google Scholar] [CrossRef]
- Vinolo, M.A.R.; Rodrigues, H.G.; Nachbar, R.T.; Curi, R. Regulation of Inflammation by Short Chain Fatty Acids. Nutrients 2011, 3, 858–876. [Google Scholar] [CrossRef]
- Sun, M.; Wu, W.; Chen, L.; Yang, W.; Huang, X.; Ma, C.; Chen, F.; Xiao, Y.; Zhao, Y.; Ma, C.; et al. Microbiota-Derived Short-Chain Fatty Acids Promote Th1 Cell IL-10 Production to Maintain Intestinal Homeostasis. Nat. Commun. 2018, 9, 3555. [Google Scholar] [CrossRef]
- McDaniel, D.K.; Eden, K.; Ringel, V.M.; Allen, I.C. Emerging Roles for Noncanonical NF-κB Signaling in the Modulation of Inflammatory Bowel Disease Pathobiology. Inflamm. Bowel Dis. 2016, 22, 2265–2279. [Google Scholar] [CrossRef]
- Kim, E.K.; Choi, E.-J. Pathological Roles of MAPK Signaling Pathways in Human Diseases. Biochim. Biophys. Acta 2010, 1802, 396–405. [Google Scholar] [CrossRef] [PubMed]
- Atreya, I.; Atreya, R.; Neurath, M.F. NF-κB in Inflammatory Bowel Disease. J. Intern. Med. 2008, 263, 591–596. [Google Scholar] [CrossRef]
- Broom, O.J.; Widjaya, B.; Troelsen, J.; Olsen, J.; Nielsen, O.H. Mitogen Activated Protein Kinases: A Role in Inflammatory Bowel Disease? Clin. Exp. Immunol. 2009, 158, 272–280. [Google Scholar] [CrossRef]
- Schulze-Osthoff, K.; Ferrari, D.; Riehemann, K.; Wesselborg, S. Regulation of NF-Kappa B Activation by MAP Kinase Cascades. Immunobiology 1997, 198, 35–49. [Google Scholar] [CrossRef]
- Shi, J.-H.; Sun, S.-C. Tumor Necrosis Factor Receptor-Associated Factor Regulation of Nuclear Factor κB and Mitogen-Activated Protein Kinase Pathways. Front. Immunol. 2018, 9, 1849. [Google Scholar] [CrossRef] [PubMed]
- Kitazawa, H.; Villena, J.; Alvarez, S. Probiotics: Immunobiotics and Immunogenics; CRC Press: Boca Raton, FL, USA, 2013; ISBN 9781482206852. [Google Scholar]
- Virginia Rodriguez, A.; Griet, M. NF-κB in Anti-Inflammatory Activity of Probiotics: An Update. Curr. Immunol. Rev. 2017, 12, 74–82. [Google Scholar] [CrossRef]
- Wang, H.; Li, S.; Li, H.; Du, F.; Guan, J.; Wu, Y. Mechanism of Probiotic VSL#3 Inhibiting NF-κB and TNF-α on Colitis through TLR4-NF-κB Signal Pathway. Iran. J. Public Health 2019, 48, 1292–1300. [Google Scholar]
- Plaza-Díaz, J.; Ruiz-Ojeda, F.J.; Vilchez-Padial, L.M.; Gil, A. Evidence of the Anti-Inflammatory Effects of Probiotics and Synbiotics in Intestinal Chronic Diseases. Nutrients 2017, 9, 555. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, K.; Karin, M. NF-κB, Inflammation, Immunity and Cancer: Coming of Age. Nat. Rev. Immunol. 2018, 18, 309–324. [Google Scholar] [CrossRef] [PubMed]
- Oeckinghaus, A.; Ghosh, S. The NF-kappaB Family of Transcription Factors and Its Regulation. Cold Spring Harb. Perspect. Biol. 2009, 1, a000034. [Google Scholar] [CrossRef]
- Giuliani, C.; Bucci, I.; Napolitano, G. The Role of the Transcription Factor Nuclear Factor-Kappa B in Thyroid Autoimmunity and Cancer. Front. Endocrinol. 2018, 9, 471. [Google Scholar] [CrossRef]
- Dong, C.; Davis, R.J.; Flavell, R.A. MAP Kinases in the Immune Response. Annu. Rev. Immunol. 2002, 20, 55–72. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Dong, C. Regulatory Mechanisms of Mitogen-Activated Kinase Signaling. Cell. Mol. Life Sci. 2007, 64, 2771–2789. [Google Scholar] [CrossRef]
- Houde, M.; Laprise, P.; Jean, D.; Blais, M.; Asselin, C.; Rivard, N. Intestinal Epithelial Cell Differentiation Involves Activation of p38 Mitogen-Activated Protein Kinase That Regulates the Homeobox Transcription Factor CDX2. J. Biol. Chem. 2001, 276, 21885–21894. [Google Scholar] [CrossRef] [PubMed]
- Wakeman, D.; Schneider, J.E.; Liu, J.; Wandu, W.S.; Erwin, C.R.; Guo, J.; Stappenbeck, T.S.; Warner, B.W. Deletion of p38-Alpha Mitogen-Activated Protein Kinase within the Intestinal Epithelium Promotes Colon Tumorigenesis. Surgery 2012, 152, 286–293. [Google Scholar] [CrossRef]
- Morrison, D.K. MAP Kinase Pathways. Cold Spring Harb. Perspect. Biol. 2012, 4. [Google Scholar] [CrossRef]
- Cargnello, M.; Roux, P.P. Activation and Function of the MAPKs and Their Substrates, the MAPK-Activated Protein Kinases. Microbiol. Mol. Biol. Rev. 2011, 75, 50–83. [Google Scholar] [CrossRef]
- Lee, S.; Rauch, J.; Kolch, W. Targeting MAPK Signaling in Cancer: Mechanisms of Drug Resistance and Sensitivity. Int. J. Mol. Sci. 2020, 21, 1102. [Google Scholar] [CrossRef]
- Jones, S.E.; Versalovic, J. Probiotic Lactobacillus Reuteri Biofilms Produce Antimicrobial and Anti-Inflammatory Factors. BMC Microbiol. 2009, 9, 35. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, R.; Vasiljevic, T.; Smith, S.C.; Donkor, O.N. Effect of Cell-Surface Components and Metabolites of Lactic Acid Bacteria and Probiotic Organisms on Cytokine Production and Induction of CD25 Expression in Human Peripheral Mononuclear Cells. J. Dairy Sci. 2014, 97, 2542–2558. [Google Scholar] [CrossRef] [PubMed]
- Uchinaka, A.; Azuma, N.; Mizumoto, H.; Nakano, S.; Minamiya, M.; Yoneda, M.; Aoyama, K.; Komatsu, Y.; Yamada, Y.; Murohara, T.; et al. Anti-Inflammatory Effects of Heat-Killed Lactobacillus Plantarum L-137 on Cardiac and Adipose Tissue in Rats with Metabolic Syndrome. Sci. Rep. 2018, 8, 8156. [Google Scholar] [CrossRef]
- De Marco, S.; Sichetti, M.; Muradyan, D.; Piccioni, M.; Traina, G.; Pagiotti, R.; Pietrella, D. Probiotic Cell-Free Supernatants Exhibited Anti-Inflammatory and Antioxidant Activity on Human Gut Epithelial Cells and Macrophages Stimulated with LPS. Evid. Based. Complement. Alternat. Med. 2018, 2018, 1756308. [Google Scholar] [CrossRef]
- Lakhdari, O.; Tap, J.; Béguet-Crespel, F.; Le Roux, K.; de Wouters, T.; Cultrone, A.; Nepelska, M.; Lefèvre, F.; Doré, J.; Blottière, H.M. Identification of NF-κB Modulation Capabilities within Human Intestinal Commensal Bacteria. J. Biomed. Biotechnol. 2011, 2011, 282356. [Google Scholar] [CrossRef]
- Imaoka, A.; Shima, T.; Kato, K.; Mizuno, S.; Uehara, T.; Matsumoto, S.; Setoyama, H.; Hara, T.; Umesaki, Y. Anti-Inflammatory Activity of Probiotic Bifidobacterium: Enhancement of IL-10 Production in Peripheral Blood Mononuclear Cells from Ulcerative Colitis Patients and Inhibition of IL-8 Secretion in HT-29 Cells. World J. Gastroenterol. 2008, 14, 2511–2516. [Google Scholar] [CrossRef]
- Petrof, E.O.; Kojima, K.; Ropeleski, M.J.; Musch, M.W.; Tao, Y.; De Simone, C.; Chang, E.B. Probiotics Inhibit Nuclear Factor-kappaB and Induce Heat Shock Proteins in Colonic Epithelial Cells through Proteasome Inhibition. Gastroenterology 2004, 127, 1474–1487. [Google Scholar] [CrossRef]
- Heuvelin, E.; Lebreton, C.; Grangette, C.; Pot, B.; Cerf-Bensussan, N.; Heyman, M. Mechanisms Involved in Alleviation of Intestinal Inflammation by Bifidobacterium Breve Soluble Factors. PLoS ONE 2009, 4, e5184. [Google Scholar] [CrossRef] [PubMed]
- Kaci, G.; Lakhdari, O.; Doré, J.; Dusko Ehrlich, S.; Renault, P.; Blottière, H.M.; Delorme, C. Inhibition of the NF-κB Pathway in Human Intestinal Epithelial Cells by Commensal Streptococcus Salivarius. Appl. Environ. Microbiol. 2011, 77, 4681–4684. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, J.; Cheng, Y.; Liu, X.; Huang, Y. Secreted Factors from Bifidobacterium Animalis Subsp. Lactis Inhibit NF-κB-Mediated Interleukin-8 Gene Expression in Caco-2 Cells. Appl. Environ. Microbiol. 2011, 77, 8171–8174. [Google Scholar] [CrossRef] [PubMed]
- Petrof, E.O.; Claud, E.C.; Sun, J.; Abramova, T.; Guo, Y.; Waypa, T.S.; He, S.-M.; Nakagawa, Y.; Chang, E.B. Bacteria-Free Solution Derived from Lactobacillus Plantarum Inhibits Multiple NF-kappaB Pathways and Inhibits Proteasome Function. Inflamm. Bowel Dis. 2009, 15, 1537–1547. [Google Scholar] [CrossRef]
- Ismaeil, H.; Abdo, W.; Amer, S.; Tahoun, A.; Massoud, D.; Zanaty, E.; Bin-Jumah, M.; Mahmoud, A.M. Ameliorative Effect of Heat-Killed Lactobacillus Plantarum L.137 And/or Aloe Vera against Colitis in Mice. Processes 2020, 8, 225. [Google Scholar] [CrossRef]
- Bermudez-Brito, M.; Muñoz-Quezada, S.; Gomez-Llorente, C.; Romero, F.; Gil, A. Lactobacillus Rhamnosus and Its Cell-Free Culture Supernatant Differentially Modulate Inflammatory Biomarkers in Escherichia Coli-Challenged Human Dendritic Cells. Br. J. Nutr. 2014, 111, 1727–1737. [Google Scholar] [CrossRef] [PubMed]
- Chon, H.; Choi, B.; Jeong, G.; Lee, E.; Lee, S. Suppression of Proinflammatory Cytokine Production by Specific Metabolites of Lactobacillus Plantarum 10hk2 via Inhibiting NF-κB and p38 MAPK Expressions. Comp. Immunol. Microbiol. Infect. Dis. 2010, 33, e41–e49. [Google Scholar] [CrossRef]
- Ghadimi, D.; de Vrese, M.; Heller, K.J.; Schrezenmeir, J. Effect of Natural Commensal-Origin DNA on Toll-like Receptor 9 (TLR9) Signaling Cascade, Chemokine IL-8 Expression, and Barrier Integritiy of Polarized Intestinal Epithelial Cells. Inflamm. Bowel Dis. 2010, 16, 410–427. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.-S.; Lee, N.-K.; Choi, A.-J.; Choe, J.-S.; Bae, C.H.; Paik, H.-D. Anti-Inflammatory Potential of Probiotic Strain Weissella Cibaria JW15 Isolated from Kimchi through Regulation of NF-κB and MAPKs Pathways in LPS-Induced RAW 264.7 Cells. J. Microbiol. Biotechnol. 2019, 29, 1022–1032. [Google Scholar] [CrossRef]
- Jung, J.-I.; Baek, S.-M.; Nguyen, T.H.; Kim, J.W.; Kang, C.-H.; Kim, S.; Imm, J.-Y. Effects of Probiotic Culture Supernatant on Cariogenic Biofilm Formation and RANKL-Induced Osteoclastogenesis in RAW 264.7 Macrophages. Molecules 2021, 26, 733. [Google Scholar] [CrossRef]
- Rocha-Ramírez, L.M.; Pérez-Solano, R.A.; Castañón-Alonso, S.L.; Moreno Guerrero, S.S.; Ramírez Pacheco, A.; García Garibay, M.; Eslava, C. Probiotic Lactobacillus Strains Stimulate the Inflammatory Response and Activate Human Macrophages. J. Immunol. Res. 2017, 2017, 4607491. [Google Scholar] [CrossRef]
- Karlsson, M.; Scherbak, N.; Reid, G.; Jass, J. Lactobacillus Rhamnosus GR-1 Enhances NF-kappaB Activation in Escherichia Coli-Stimulated Urinary Bladder Cells through TLR4. BMC Microbiol. 2012, 12, 15. [Google Scholar] [CrossRef]
- Tao, Y.; Drabik, K.A.; Waypa, T.S.; Musch, M.W.; Alverdy, J.C.; Schneewind, O.; Chang, E.B.; Petrof, E.O. Soluble Factors from Lactobacillus GG Activate MAPKs and Induce Cytoprotective Heat Shock Proteins in Intestinal Epithelial Cells. Am. J. Physiol. Cell Physiol. 2006, 290, C1018–C1030. [Google Scholar] [CrossRef]
- Fujiya, M.; Musch, M.W.; Nakagawa, Y.; Hu, S.; Alverdy, J.; Kohgo, Y.; Schneewind, O.; Jabri, B.; Chang, E.B. The Bacillus Subtilis Quorum-Sensing Molecule CSF Contributes to Intestinal Homeostasis via OCTN2, a Host Cell Membrane Transporter. Cell Host Microbe 2007, 1, 299–308. [Google Scholar] [CrossRef]
- Schlee, M.; Harder, J.; Köten, B.; Stange, E.F.; Wehkamp, J.; Fellermann, K. Probiotic Lactobacilli and VSL# 3 Induce Enterocyte β-Defensin 2. Clin. Exp. Immunol. 2008, 151, 528–535. [Google Scholar]
- Bidmon-Fliegenschnee, B.; Lederhuber, H.C.; Csaicsich, D.; Pichler, J.; Herzog, R.; Memaran-Dadgar, N.; Huber, W.-D.; Aufricht, C.; Kratochwill, K. Overexpression of Hsp70 Confers Cytoprotection during Gliadin Exposure in Caco-2 Cells. Pediatr. Res. 2015, 78, 358–364. [Google Scholar] [CrossRef]
- Koeninger, L.; Armbruster, N.S.; Brinch, K.S.; Kjaerulf, S.; Andersen, B.; Langnau, C.; Autenrieth, S.E.; Schneidawind, D.; Stange, E.F.; Malek, N.P.; et al. Human β-Defensin 2 Mediated Immune Modulation as Treatment for Experimental Colitis. Front. Immunol. 2020, 11, 93. [Google Scholar] [CrossRef] [PubMed]
- Iyer, C.; Kosters, A.; Sethi, G.; Kunnumakkara, A.B.; Aggarwal, B.B.; Versalovic, J. Probiotic Lactobacillus Reuteri Promotes TNF-Induced Apoptosis in Human Myeloid Leukemia-Derived Cells by Modulation of NF-κB and MAPK Signalling. Cell. Microbiol. 2008, 10, 1442–1452. [Google Scholar] [CrossRef] [PubMed]
- Jeong, M.; Kim, J.H.; Yang, H.; Kang, S.D.; Song, S.; Lee, D.; Lee, J.S.; Yoon Park, J.H.; Byun, S.; Lee, K.W. Heat-Killed Lactobacillus Plantarum KCTC 13314BP Enhances Phagocytic Activity and Immunomodulatory Effects Via Activation of MAPK and STAT3 Pathways. J. Microbiol. Biotechnol. 2019, 29, 1248–1254. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Jiang, Y.-J.; Yang, X.-Y.; Liu, Y.; Wang, J.-Y.; Man, C.-X. Immunoregulatory Effects on Caco-2 Cells and Mice of Exopolysaccharides Isolated from Lactobacillus Acidophilus NCFM. Food Funct. 2014, 5, 3261–3268. [Google Scholar] [CrossRef]
- Wang, H.; Cheng, X.; Zhang, L.; Xu, S.; Zhang, Q.; Lu, R. A Surface-Layer Protein from Lactobacillus Acidophilus NCFM Induces Autophagic Death in HCT116 Cells Requiring ROS-Mediated Modulation of mTOR and JNK Signaling Pathways. Food Funct. 2019, 10, 4102–4112. [Google Scholar] [CrossRef]
- Wang, H.; Zhang, L.; Xu, S.; Pan, J.; Zhang, Q.; Lu, R. Surface-Layer Protein from Lactobacillus Acidophilus NCFM Inhibits Lipopolysaccharide-Induced Inflammation through MAPK and NF-κB Signaling Pathways in RAW264.7 Cells. J. Agric. Food Chem. 2018, 66, 7655–7662. [Google Scholar] [CrossRef]
- Liu, Q.; Yu, Z.; Tian, F.; Zhao, J.; Zhang, H.; Zhai, Q.; Chen, W. Surface Components and Metabolites of Probiotics for Regulation of Intestinal Epithelial Barrier. Microb. Cell Fact. 2020, 19, 23. [Google Scholar] [CrossRef] [PubMed]
- Wagner, C.E.; Wheeler, K.M.; Ribbeck, K. Mucins and Their Role in Shaping the Functions of Mucus Barriers. Annu. Rev. Cell Dev. Biol. 2018, 34, 189–215. [Google Scholar] [CrossRef]
- Osaki, M.; Oshimura, M.; Ito, H. PI3K-Akt Pathway: Its Functions and Alterations in Human Cancer. Apoptosis 2004, 9, 667–676. [Google Scholar] [CrossRef]
- Schultze, S.M.; Hemmings, B.A.; Niessen, M.; Tschopp, O. PI3K/AKT, MAPK and AMPK Signalling: Protein Kinases in Glucose Homeostasis. Expert Rev. Mol. Med. 2012, 14, e1. [Google Scholar] [CrossRef]
- Paveljšek, D.; Ivičak-Kocjan, K.; Treven, P.; Benčina, M.; Jerala, R.; Rogelj, I. Distinctive Probiotic Features Share Common TLR2-Dependent Signalling in Intestinal Epithelial Cells. Cell. Microbiol. 2021, 23, e13264. [Google Scholar] [CrossRef] [PubMed]
- Mohseni, A.H.; Casolaro, V.; Bermúdez-Humarán, L.G.; Keyvani, H.; Taghinezhad-S, S. Modulation of the PI3K/Akt/mTOR Signaling Pathway by Probiotics as a Fruitful Target for Orchestrating the Immune Response. Gut Microbes 2021, 13, 1–17. [Google Scholar] [CrossRef]
- Zhang, M.; Wang, C.; Wang, C.; Zhao, H.; Zhao, C.; Chen, Y.; Wang, Y.; McClain, C.; Feng, W. Enhanced AMPK Phosphorylation Contributes to the Beneficial Effects of Lactobacillus Rhamnosus GG Supernatant on Chronic-Alcohol-Induced Fatty Liver Disease. J. Nutr. Biochem. 2015, 26, 337–344. [Google Scholar] [CrossRef]
- Lew, L.C.; Hor, Y.Y.; Jaafar, M.H.; Lau, A.S.Y.; Ong, J.S.; Chuah, L.O.; Yap, K.P.; Azzam, G.; Azlan, A.; Liong, M.T. Lactobacilli Modulated AMPK Activity and Prevented Telomere Shortening in Ageing Rats. Benef. Microbes 2019, 10, 883–892. [Google Scholar] [CrossRef]
- Ventura-Sobrevilla, J.M.; Boone-Villa, V.D.; Jiménez-Villarreal, J.; Nevarez-Moorillón, G.V. Influence of Probiotics Over AMPK-Dependent Health Activity: A Look into Its Molecular Mechanisms. In Advances in Probiotics for Sustainable Food and Medicine; Goel, G., Kumar, A., Eds.; Springer Singapore: Singapore, 2021; pp. 213–223. ISBN 9789811567957. [Google Scholar]
- Yan, F.; Polk, D.B. Probiotic Bacterium Prevents Cytokine-Induced Apoptosis in Intestinal Epithelial Cells. J. Biol. Chem. 2002, 277, 50959–50965. [Google Scholar] [CrossRef]
- Yan, F.; Cao, H.; Cover, T.L.; Whitehead, R.; Washington, M.K.; Polk, D.B. Soluble Proteins Produced by Probiotic Bacteria Regulate Intestinal Epithelial Cell Survival and Growth. Gastroenterology 2007, 132, 562–575. [Google Scholar] [CrossRef]
- Ammoscato, F.; Scirocco, A.; Altomare, A.; Matarrese, P.; Petitta, C.; Ascione, B.; Caronna, R.; Guarino, M.; Marignani, M.; Cicala, M.; et al. Lactobacillus Rhamnosus Protects Human Colonic Muscle from Pathogen Lipopolysaccharide-Induced Damage. Neurogastroenterol. Motil. 2013, 25, 984-e777. [Google Scholar] [CrossRef]
- Caballero-Franco, C.; Keller, K.; De Simone, C.; Chadee, K. The VSL#3 Probiotic Formula Induces Mucin Gene Expression and Secretion in Colonic Epithelial Cells. Am. J. Physiol.-Gastrointest. Liver Physiol. 2007, 292, G315–G322. [Google Scholar]
- Gao, J.; Li, Y.; Wan, Y.; Hu, T.; Liu, L.; Yang, S.; Gong, Z.; Zeng, Q.; Wei, Y.; Yang, W.; et al. A Novel Postbiotic From Lactobacillus Rhamnosus GG With a Beneficial Effect on Intestinal Barrier Function. Front. Microbiol. 2019, 10, 477. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Ajuwon, K.M. Butyrate Modifies Intestinal Barrier Function in IPEC-J2 Cells through a Selective Upregulation of Tight Junction Proteins and Activation of the Akt Signaling Pathway. PLoS ONE 2017, 12, e0179586. [Google Scholar] [CrossRef] [PubMed]
- Izuddin, W.I.; Loh, T.C.; Foo, H.L.; Samsudin, A.A.; Humam, A.M. Postbiotic L. Plantarum RG14 Improves Ruminal Epithelium Growth, Immune Status and Upregulates the Intestinal Barrier Function in Post-Weaning Lambs. Sci. Rep. 2019, 9, 9938. [Google Scholar]
- Han, N.; Jia, L.; Su, Y.; Du, J.; Guo, L.; Luo, Z.; Liu, Y. Lactobacillus Reuteri Extracts Promoted Wound Healing via PI3K/AKT/β-catenin/TGFβ1 Pathway. Stem Cell Res. Ther. 2019, 10, 243. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Sharma, A.; Gupta, M.; Padwad, Y.; Sharma, R. Cell-Free Culture Supernatant of Probiotic Lactobacillus Fermentum Protects Against H2O2-Induced Premature Senescence by Suppressing ROS-Akt-mTOR Axis in Murine Preadipocytes. Probiotics Antimicrob. Proteins 2020, 12, 563–576. [Google Scholar] [CrossRef]
- Takeuchi, O.; Akira, S. Pattern Recognition Receptors and Inflammation. Cell 2010, 140, 805–820. [Google Scholar] [CrossRef]
- Kawai, T.; Akira, S. The Role of Pattern-Recognition Receptors in Innate Immunity: Update on Toll-like Receptors. Nat. Immunol. 2010, 11, 373–384. [Google Scholar] [CrossRef]
- Franchi, L.; Warner, N.; Viani, K. Function of Nod-like Receptors in Microbial Recognition and Host Defense. Immunol. Rev. 2009, 227, 106–128. [Google Scholar] [CrossRef] [PubMed]
- Geijtenbeek, T.B.H.; Gringhuis, S.I. Signalling through C-Type Lectin Receptors: Shaping Immune Responses. Nat. Rev. Immunol. 2009, 9, 465–479. [Google Scholar] [CrossRef]
- Lebeer, S.; Vanderleyden, J.; De Keersmaecker, S.C.J. Host Interactions of Probiotic Bacterial Surface Molecules: Comparison with Commensals and Pathogens. Nat. Rev. Microbiol. 2010, 8, 171–184. [Google Scholar] [CrossRef]
- Rakoff-Nahoum, S.; Paglino, J.; Eslami-Varzaneh, F.; Edberg, S.; Medzhitov, R. Recognition of Commensal Microflora by Toll-like Receptors Is Required for Intestinal Homeostasis. Cell 2004, 118, 229–241. [Google Scholar] [CrossRef]
- Lee, J.; Mo, J.-H.; Katakura, K.; Alkalay, I.; Rucker, A.N.; Liu, Y.-T.; Lee, H.-K.; Shen, C.; Cojocaru, G.; Shenouda, S.; et al. Maintenance of Colonic Homeostasis by Distinctive Apical TLR9 Signalling in Intestinal Epithelial Cells. Nat. Cell Biol. 2006, 8, 1327–1336. [Google Scholar] [CrossRef] [PubMed]
- Bermudez-Brito, M.; Plaza-Díaz, J.; Muñoz-Quezada, S.; Gómez-Llorente, C.; Gil, A. Probiotic Mechanisms of Action. Ann. Nutr. Metab. 2012, 61, 160–174. [Google Scholar] [CrossRef] [PubMed]
- Abreu, M.T.; Fukata, M.; Arditi, M. TLR Signaling in the Gut in Health and Disease. J. Immunol. 2005, 174, 4453–4460. [Google Scholar] [CrossRef]
- Bell, J.K.; Mullen, G.E.D.; Leifer, C.A.; Mazzoni, A.; Davies, D.R.; Segal, D.M. Leucine-Rich Repeats and Pathogen Recognition in Toll-like Receptors. Trends Immunol. 2003, 24, 528–533. [Google Scholar] [CrossRef]
- Gao, W.; Xiong, Y.; Li, Q.; Yang, H. Inhibition of Toll-Like Receptor Signaling as a Promising Therapy for Inflammatory Diseases: A Journey from Molecular to Nano Therapeutics. Front. Physiol. 2017, 8, 508. [Google Scholar] [CrossRef]
- Rosenfeld, Y.; Shai, Y. Lipopolysaccharide (Endotoxin)-Host Defense Antibacterial Peptides Interactions: Role in Bacterial Resistance and Prevention of Sepsis. Biochim. Biophys. Acta 2006, 1758, 1513–1522. [Google Scholar] [CrossRef]
- Kawasaki, T.; Kawai, T. Toll-like Receptor Signaling Pathways. Front. Immunol. 2014, 5, 461. [Google Scholar] [CrossRef]
- Xu, Y.; Tao, X.; Shen, B.; Horng, T.; Medzhitov, R.; Manley, J.L.; Tong, L. Structural Basis for Signal Transduction by the Toll/interleukin-1 Receptor Domains. Nature 2000, 408, 111–115. [Google Scholar] [CrossRef]
- Dunne, A.; Ejdebäck, M.; Ludidi, P.L.; O’Neill, L.A.J.; Gay, N.J. Structural Complementarity of Toll/Interleukin-1 Receptor Domains in Toll-like Receptors and the Adaptors Mal and MyD88. J. Biol. Chem. 2003, 278, 41443–41451. [Google Scholar] [CrossRef]
- Miguel, R.N.; Wong, J.; Westoll, J.F.; Brooks, H.J.; O’Neill, L.A.J.; Gay, N.J.; Bryant, C.E.; Monie, T.P. A Dimer of the Toll-Like Receptor 4 Cytoplasmic Domain Provides a Specific Scaffold for the Recruitment of Signalling Adaptor Proteins. PLoS ONE 2007, 2, e788. [Google Scholar]
- Deguine, J.; Barton, G.M. MyD88: A Central Player in Innate Immune Signaling. F1000Prime Rep. 2014, 6, 97. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Chen, T.; Zhang, J.; Yang, M.; Li, N.; Xu, X.; Cao, X. The E3 Ubiquitin Ligase Nrdp1’preferentially’promotes TLR-Mediated Production of Type I Interferon. Nat. Immunol. 2009, 10, 744. [Google Scholar] [CrossRef]
- Tseng, P.-H.; Matsuzawa, A.; Zhang, W.; Mino, T.; Vignali, D.A.A.; Karin, M. Different Modes of Ubiquitination of the Adaptor TRAF3 Selectively Activate the Expression of Type I Interferons and Proinflammatory Cytokines. Nat. Immunol. 2010, 11, 70–75. [Google Scholar] [CrossRef]
- Gabhann, J.N.; Jefferies, C.A. TLR-Induced Activation of Btk--Role for Endosomal MHC Class II Molecules Revealed. Cell Res. 2011, 21, 998–1001. [Google Scholar] [CrossRef][Green Version]
- Frei, R.; Steinle, J.; Birchler, T.; Loeliger, S.; Roduit, C.; Steinhoff, D.; Seibl, R.; Büchner, K.; Seger, R.; Reith, W.; et al. MHC Class II Molecules Enhance Toll-like Receptor Mediated Innate Immune Responses. PLoS ONE 2010, 5, e8808. [Google Scholar] [CrossRef]
- Hassan, G.S.; Mourad, W. An Unexpected Role for MHC Class II. Nat. Immunol. 2011, 12, 375–376. [Google Scholar] [CrossRef]
- Negi, S.; Singh, H.; Mukhopadhyay, A. Gut Bacterial Peptides with Autoimmunity Potential as Environmental Trigger for Late Onset Complex Diseases: In-Silico Study. PLoS ONE 2017, 12, e0180518. [Google Scholar] [CrossRef] [PubMed]
- Matsuguchi, T.; Takagi, A.; Matsuzaki, T.; Nagaoka, M.; Ishikawa, K.; Yokokura, T.; Yoshikai, Y. Lipoteichoic Acids from Lactobacillus Strains Elicit Strong Tumor Necrosis Factor Alpha-Inducing Activities in Macrophages through Toll-like Receptor 2. Clin. Diagn. Lab. Immunol. 2003, 10, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.G.; Lee, S.Y.; Kim, N.-R.; Ko, M.Y.; Lee, J.M.; Yi, T.-H.; Chung, S.K.; Chung, D.K. Inhibitory Effects of Lactobacillus Plantarum Lipoteichoic Acid (LTA) on Staphylococcus Aureus LTA-Induced Tumor Necrosis Factor-Alpha Production. J. Microbiol. Biotechnol. 2008, 18, 1191–1196. [Google Scholar] [PubMed]
- Kim, H.G.; Kim, N.-R.; Gim, M.G.; Lee, J.M.; Lee, S.Y.; Ko, M.Y.; Kim, J.Y.; Han, S.H.; Chung, D.K. Lipoteichoic Acid Isolated from Lactobacillus Plantarum Inhibits Lipopolysaccharide-Induced TNF-α Production in THP-1 Cells and Endotoxin Shock in Mice. J. Immunol. 2008, 180, 2553–2561. [Google Scholar] [CrossRef]
- Kim, H.G.; Lee, S.Y.; Kim, N.R.; Lee, H.Y.; Ko, M.Y.; Jung, B.J.; Kim, C.M.; Lee, J.M.; Park, J.H.; Han, S.H.; et al. Lactobacillus Plantarum Lipoteichoic Acid down-Regulated Shigella Flexneri Peptidoglycan-Induced Inflammation. Mol. Immunol. 2011, 48, 382–391. [Google Scholar] [CrossRef]
- Claes, I.J.J.; Lebeer, S.; Shen, C.; Verhoeven, T.L.A.; Dilissen, E.; De Hertogh, G.; Bullens, D.M.A.; Ceuppens, J.L.; Van Assche, G.; Vermeire, S.; et al. Impact of Lipoteichoic Acid Modification on the Performance of the Probiotic Lactobacillus Rhamnosus GG in Experimental Colitis. Clin. Exp. Immunol. 2010, 162, 306–314. [Google Scholar] [CrossRef] [PubMed]
- Grangette, C.; Nutten, S.; Palumbo, E.; Morath, S.; Hermann, C.; Dewulf, J.; Pot, B.; Hartung, T.; Hols, P.; Mercenier, A. Enhanced Antiinflammatory Capacity of a Lactobacillus Plantarum Mutant Synthesizing Modified Teichoic Acids. Proc. Natl. Acad. Sci. USA 2005, 102, 10321–10326. [Google Scholar] [CrossRef]
- Jang, K.-S.; Baik, J.E.; Han, S.H.; Chung, D.K.; Kim, B.-G. Multi-Spectrometric Analyses of Lipoteichoic Acids Isolated from Lactobacillus Plantarum. Biochem. Biophys. Res. Commun. 2011, 407, 823–830. [Google Scholar] [CrossRef]
- Hayashi, F.; Smith, K.D.; Ozinsky, A.; Hawn, T.R.; Yi, E.C.; Goodlett, D.R.; Eng, J.K.; Akira, S.; Underhill, D.M.; Aderem, A. The Innate Immune Response to Bacterial Flagellin Is Mediated by Toll-like Receptor 5. Nature 2001, 410, 1099–1103. [Google Scholar] [CrossRef]
- Gewirtz, A.T. Flag in the Crossroads: Flagellin Modulates Innate and Adaptive Immunity. Curr. Opin. Gastroenterol. 2006, 22, 8–12. [Google Scholar] [CrossRef]
- Ren, T.; Zamboni, D.S.; Roy, C.R.; Dietrich, W.F.; Vance, R.E. Flagellin-Deficient Legionella Mutants Evade Caspase-1- and Naip5-Mediated Macrophage Immunity. PLoS Pathogens 2006, 2, e18. [Google Scholar] [CrossRef]
- Akira, S.; Uematsu, S.; Takeuchi, O. Pathogen Recognition and Innate Immunity. Cell 2006, 124, 783–801. [Google Scholar] [CrossRef] [PubMed]
- Cullender, T.C.; Chassaing, B.; Janzon, A.; Kumar, K.; Muller, C.E.; Werner, J.J.; Angenent, L.T.; Bell, M.E.; Hay, A.G.; Peterson, D.A.; et al. Innate and Adaptive Immunity Interact to Quench Microbiome Flagellar Motility in the Gut. Cell Host Microbe 2013, 14, 571–581. [Google Scholar] [CrossRef] [PubMed]
- Fulde, M.; Sommer, F.; Chassaing, B.; van Vorst, K.; Dupont, A.; Hensel, M.; Basic, M.; Klopfleisch, R.; Rosenstiel, P.; Bleich, A.; et al. Publisher Correction: Neonatal Selection by Toll-like Receptor 5 Influences Long-Term Gut Microbiota Composition. Nature 2018, 563, E25. [Google Scholar] [CrossRef]
- Vijay-Kumar, M.; Aitken, J.D.; Sanders, C.J.; Frias, A.; Sloane, V.M.; Xu, J.; Neish, A.S.; Rojas, M.; Gewirtz, A.T. Flagellin Treatment Protects against Chemicals, Bacteria, Viruses, and Radiation. J. Immunol. 2008, 180, 8280–8285. [Google Scholar] [CrossRef] [PubMed]
- Chassaing, B.; Ley, R.E.; Gewirtz, A.T. Intestinal Epithelial Cell Toll-like Receptor 5 Regulates the Intestinal Microbiota to Prevent Low-Grade Inflammation and Metabolic Syndrome in Mice. Gastroenterology 2014, 147, 1363–1377.e17. [Google Scholar] [CrossRef]
- Lopetuso, L.R.; Jia, R.; Wang, X.-M.; Jia, L.-G.; Petito, V.; Goodman, W.A.; Meddings, J.B.; Cominelli, F.; Reuter, B.K.; Pizarro, T.T. Epithelial-Specific Toll-like Receptor (TLR)5 Activation Mediates Barrier Dysfunction in Experimental Ileitis. Inflamm. Bowel Dis. 2017, 23, 392–403. [Google Scholar] [CrossRef]
- Vijay-Kumar, M.; Aitken, J.D.; Carvalho, F.A.; Cullender, T.C.; Mwangi, S.; Srinivasan, S.; Sitaraman, S.V.; Knight, R.; Ley, R.E.; Gewirtz, A.T. Metabolic Syndrome and Altered Gut Microbiota in Mice Lacking Toll-like Receptor 5. Science 2010, 328, 228–231. [Google Scholar] [CrossRef] [PubMed]
- Pekkala, S.; Munukka, E.; Kong, L.; Pöllänen, E.; Autio, R.; Roos, C.; Wiklund, P.; Fischer-Posovszky, P.; Wabitsch, M.; Alen, M.; et al. Toll-like Receptor 5 in Obesity: The Role of Gut Microbiota and Adipose Tissue Inflammation: Gut Microbiota, TLR5, Inflammation, and Obesity. Obesity 2015, 23, 581–590. [Google Scholar] [CrossRef]
- Zhang, W.; Hartmann, R.; Tun, H.M.; Elson, C.O.; Khafipour, E.; Garvey, W.T. Deletion of the Toll-Like Receptor 5 Gene Per Se Does Not Determine the Gut Microbiome Profile That Induces Metabolic Syndrome: Environment Trumps Genotype. PLoS ONE 2016, 11, e0150943. [Google Scholar] [CrossRef]
- Neville, B.A.; Forde, B.M.; Claesson, M.J.; Darby, T.; Coghlan, A.; Nally, K.; Ross, R.P.; O’Toole, P.W. Characterization of pro-Inflammatory Flagellin Proteins Produced by Lactobacillus Ruminis and Related Motile Lactobacilli. PLoS ONE 2012, 7, e40592. [Google Scholar] [CrossRef]
- Kajikawa, A.; Midorikawa, E.; Masuda, K.; Kondo, K.; Irisawa, T.; Igimi, S.; Okada, S. Characterization of Flagellins Isolated from a Highly Motile Strain of Lactobacillus Agilis. BMC Microbiol. 2016, 16, 49. [Google Scholar] [CrossRef]
- Voogdt, C.G.P.; Bouwman, L.I.; Kik, M.J.L.; Wagenaar, J.A.; van Putten, J.P.M. Reptile Toll-like Receptor 5 Unveils Adaptive Evolution of Bacterial Flagellin Recognition. Sci. Rep. 2016, 6, 19046. [Google Scholar] [CrossRef] [PubMed]
- Schlee, M.; Wehkamp, J.; Altenhoefer, A.; Oelschlaeger, T.A.; Stange, E.F.; Fellermann, K. Induction of Human β-Defensin 2 by the Probiotic Escherichia Coli Nissle 1917 Is Mediated through Flagellin. Infect. Immun. 2007, 75, 2399–2407. [Google Scholar] [CrossRef]
- Telford, J.L.; Barocchi, M.A.; Margarit, I.; Rappuoli, R.; Grandi, G. Pili in Gram-Positive Pathogens. Nat. Rev. Microbiol. 2006, 4, 509–519. [Google Scholar] [CrossRef]
- Proft, T.; Baker, E.N. Pili in Gram-Negative and Gram-Positive Bacteria—structure, Assembly and Their Role in Disease. Cell. Mol. Life Sci. 2009, 66, 613. [Google Scholar] [CrossRef]
- Kankainen, M.; Paulin, L.; Tynkkynen, S.; von Ossowski, I.; Reunanen, J.; Partanen, P.; Satokari, R.; Vesterlund, S.; Hendrickx, A.P.A.; Lebeer, S.; et al. Comparative Genomic Analysis of Lactobacillus Rhamnosus GG Reveals Pili Containing a Human- Mucus Binding Protein. Proc. Natl. Acad. Sci. USA 2009, 106, 17193–17198. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Jaatinen, A.; Rintahaka, J.; Hynönen, U.; Lyytinen, O.; Kant, R.; Åvall-Jääskeläinen, S.; von Ossowski, I.; Palva, A. Human Gut-Commensalic Lactobacillus Ruminis ATCC 25644 Displays Sortase-Assembled Surface Piliation: Phenotypic Characterization of Its Fimbrial Operon through In Silico Predictive Analysis and Recombinant Expression in Lactococcus Lactis. PLoS ONE 2015, 10, e0145718. [Google Scholar] [CrossRef]
- Lebeer, S.; Claes, I.; Tytgat, H.L.P.; Verhoeven, T.L.A.; Marien, E.; von Ossowski, I.; Reunanen, J.; Palva, A.; de Vos, W.M.; Keersmaecker, S.C.J.D.; et al. Functional Analysis of Lactobacillus Rhamnosus GG Pili in Relation to Adhesion and Immunomodulatory Interactions with Intestinal Epithelial Cells. Appl. Environ. Microbiol. 2012, 78, 185–193. [Google Scholar] [CrossRef]
- Chaurasia, P.; Pratap, S.; von Ossowski, I.; Palva, A.; Krishnan, V. New Insights about Pilus Formation in Gut-Adapted Lactobacillus Rhamnosus GG from the Crystal Structure of the SpaA Backbone-Pilin Subunit. Sci. Rep. 2016, 6, 28664. [Google Scholar] [CrossRef]
- Granato, D.; Bergonzelli, G.E.; Pridmore, R.D.; Marvin, L.; Rouvet, M.; Corthésy-Theulaz, I.E. Cell Surface-Associated Elongation Factor Tu Mediates the Attachment of Lactobacillus Johnsonii NCC533 (La1) to Human Intestinal Cells and Mucins. Infect. Immun. 2004, 72, 2160–2169. [Google Scholar] [CrossRef] [PubMed]
- Bergonzelli, G.E.; Granato, D.; Pridmore, R.D.; Marvin-Guy, L.F.; Donnicola, D.; Corthésy-Theulaz, I.E. GroEL of Lactobacillus Johnsonii La1 (NCC 533) Is Cell Surface Associated: Potential Role in Interactions with the Host and the Gastric Pathogen Helicobacter Pylori. Infect. Immun. 2006, 74, 425–434. [Google Scholar] [CrossRef] [PubMed]
- Finberg, R.W.; Kurt-Jones, E.A. CD14: Chaperone or Matchmaker? Immunity 2006, 24, 127–129. [Google Scholar] [CrossRef] [PubMed]
- Zanoni, I.; Ostuni, R.; Marek, L.R.; Barresi, S.; Barbalat, R.; Barton, G.M.; Granucci, F.; Kagan, J.C. CD14 Controls the LPS-Induced Endocytosis of Toll-like Receptor 4. Cell 2011, 147, 868–880. [Google Scholar] [CrossRef] [PubMed]
- Hsu, K.; Chung, Y.M.; Endoh, Y.; Geczy, C.L. TLR9 Ligands Induce S100A8 in Macrophages via a STAT3-Dependent Pathway Which Requires IL-10 and PGE2. PLoS ONE 2014, 9, e103629. [Google Scholar] [CrossRef] [PubMed]
- Hornung, V.; Rothenfusser, S.; Britsch, S.; Krug, A.; Jahrsdörfer, B.; Giese, T.; Endres, S.; Hartmann, G. Quantitative Expression of Toll-like Receptor 1-10 mRNA in Cellular Subsets of Human Peripheral Blood Mononuclear Cells and Sensitivity to CpG Oligodeoxynucleotides. J. Immunol. 2002, 168, 4531–4537. [Google Scholar] [CrossRef]
- Kadowaki, N.; Ho, S.; Antonenko, S.; Malefyt, R.W.; Kastelein, R.A.; Bazan, F.; Liu, Y.J. Subsets of Human Dendritic Cell Precursors Express Different Toll-like Receptors and Respond to Different Microbial Antigens. J. Exp. Med. 2001, 194, 863–869. [Google Scholar] [CrossRef]
- Schneberger, D.; Caldwell, S.; Kanthan, R.; Singh, B. Expression of Toll-like Receptor 9 in Mouse and Human Lungs. J. Anat. 2013, 222, 495–503. [Google Scholar] [CrossRef]
- Dalpke, A.; Frank, J.; Peter, M.; Heeg, K. Activation of Toll-like Receptor 9 by DNA from Different Bacterial Species. Infect. Immun. 2006, 74, 940–946. [Google Scholar] [CrossRef]
- Klinman, D.M.; Yamshchikov, G.; Ishigatsubo, Y. Contribution of CpG Motifs to the Immunogenicity of DNA Vaccines. J. Immunol. 1997, 158, 3635–3639. [Google Scholar]
- Bode, C.; Zhao, G.; Steinhagen, F.; Kinjo, T. CpG DNA as a Vaccine Adjuvant. Rev. Vaccines 2011, 10, 499–511. [Google Scholar] [CrossRef]
- Zhong, Y.; Huang, J.; Tang, W.; Chen, B.; Cai, W. Effects of Probiotics, Probiotic DNA and the CpG Oligodeoxynucleotides on Ovalbumin-Sensitized Brown-Norway Rats via TLR9/NF-κB Pathway. FEMS Immunol. Med. Microbiol. 2012, 66, 71–82. [Google Scholar] [CrossRef]
- Iliev, I.D.; Kitazawa, H.; Shimosato, T.; Katoh, S.; Morita, H.; He, F.; Hosoda, M.; Saito, T. Strong Immunostimulation in Murine Immune Cells by Lactobacillus Rhamnosus GG DNA Containing Novel Oligodeoxynucleotide Pattern. Cell. Microbiol. 2005, 7, 403–414. [Google Scholar] [CrossRef]
- Iliev, I.D.; Tohno, M.; Kurosaki, D.; Shimosato, T.; He, F.; Hosoda, M.; Saito, T.; Kitazawa, H. Immunostimulatory Oligodeoxynucleotide Containing TTTCGTTT Motif from Lactobacillus Rhamnosus GG DNA Potentially Suppresses OVA-Specific IgE Production in Mice. Scand. J. Immunol. 2008, 67, 370–376. [Google Scholar] [CrossRef] [PubMed]
- Kant, R.; de Vos, W.M.; Palva, A.; Satokari, R. Immunostimulatory CpG Motifs in the Genomes of Gut Bacteria and Their Role in Human Health and Disease. J. Med. Microbiol. 2014, 63, 293–308. [Google Scholar] [CrossRef]
- Watson, J.L.; McKay, D.M. The Immunophysiological Impact of Bacterial CpG DNA on the Gut. Clin. Chim. Acta 2006, 364, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Ghadimi, D.; Fölster-Holst, R.; de Vrese, M.; Winkler, P.; Heller, K.J.; Schrezenmeir, J. Effects of Probiotic Bacteria and Their Genomic DNA on TH1/TH2-Cytokine Production by Peripheral Blood Mononuclear Cells (PBMCs) of Healthy and Allergic Subjects. Immunobiology 2008, 213, 677–692. [Google Scholar] [CrossRef] [PubMed]
- Grüber, C.; Wendt, M.; Sulser, C.; Lau, S.; Kulig, M.; Wahn, U.; Werfel, T.; Niggemann, B. Randomized, Placebo-Controlled Trial of Lactobacillus Rhamnosus GG as Treatment of Atopic Dermatitis in Infancy. Allergy 2007, 62, 1270–1276. [Google Scholar] [CrossRef]
- Odamaki, T.; Xiao, J.-Z.; Iwabuchi, N.; Sakamoto, M.; Takahashi, N.; Kondo, S.; Miyaji, K.; Iwatsuki, K.; Togashi, H.; Enomoto, T.; et al. Influence of Bifidobacterium Longum BB536 Intake on Faecal Microbiota in Individuals with Japanese Cedar Pollinosis during the Pollen Season. J. Med. Microbiol. 2007, 56, 1301–1308. [Google Scholar] [CrossRef]
- Qi, S.R.; Cui, Y.J.; Liu, J.X.; Luo, X.; Wang, H.F. Lactobacillus Rhamnosus GG Components, SLP, gDNA and CpG, Exert Protective Effects on Mouse Macrophages upon Lipopolysaccharide Challenge. Lett. Appl. Microbiol. 2020, 70, 118–127. [Google Scholar] [CrossRef]
- Hee Kim, C.; Geun Kim, H.; Yun Kim, J.; Ra Kim, N.; Jun Jung, B.; Hye Jeong, J.; Kyun Chung, D. Probiotic Genomic DNA Reduces the Production of pro-Inflammatory Cytokine Tumor Necrosis Factor-Alpha. FEMS Microbiol. Lett. 2012, 328, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Hassan, M.; Kjos, M.; Nes, I.F.; Diep, D.B.; Lotfipour, F. Natural Antimicrobial Peptides from Bacteria: Characteristics and Potential Applications to Fight against Antibiotic Resistance. J. Appl. Microbiol. 2012, 113, 723–736. [Google Scholar] [CrossRef]
- Dobson, A.; Cotter, P.D.; Ross, R.P.; Hill, C. Bacteriocin Production: A Probiotic Trait? Appl. Environ. Microbiol. 2012, 78, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-H.; Karamychev, V.N.; Kozyavkin, S.A.; Mills, D.; Pavlov, A.R.; Pavlova, N.V.; Polouchine, N.N.; Richardson, P.M.; Shakhova, V.V.; Slesarev, A.I.; et al. Comparative Genomic Analysis of the Gut Bacterium Bifidobacterium Longum Reveals Loci Susceptible to Deletion during Pure Culture Growth. BMC Genom. 2008, 9, 247. [Google Scholar] [CrossRef] [PubMed]
- Walsh, M.C.; Gardiner, G.E.; Hart, O.M.; Lawlor, P.G.; Daly, M.; Lynch, B.; Richert, B.T.; Radcliffe, S.; Giblin, L.; Hill, C.; et al. Predominance of a Bacteriocin-Producing Lactobacillus Salivarius Component of a Five-Strain Probiotic in the Porcine Ileum and Effects on Host Immune Phenotype. FEMS Microbiol. Ecol. 2008, 64, 317–327. [Google Scholar] [CrossRef]
- O’Shea, E.F.; Gardiner, G.E.; O’Connor, P.M.; Mills, S.; Paul Ross, R.; Hill, C. Characterization of Enterocin- and Salivaricin-Producing Lactic Acid Bacteria from the Mammalian Gastrointestinal Tract. FEMS Microbiol. Lett. 2009, 291, 24–34. [Google Scholar] [CrossRef] [PubMed]
- O’Shea, E.F.; O’Connor, P.M.; Raftis, E.J.; O’Toole, P.W.; Stanton, C.; Cotter, P.D.; Ross, R.P.; Hill, C. Production of Multiple Bacteriocins from a Single Locus by Gastrointestinal Strains of Lactobacillus Salivarius. J. Bacteriol. 2011, 193, 6973–6982. [Google Scholar] [CrossRef]
- Gordon, D.M.; O’Brien, C.L. Bacteriocin Diversity and the Frequency of Multiple Bacteriocin Production in Escherichia Coli. Microbiology 2006, 152, 3239–3244. [Google Scholar] [CrossRef]
- Drissi, F.; Buffet, S.; Raoult, D.; Merhej, V. Common Occurrence of Antibacterial Agents in Human Intestinal Microbiota. Front. Microbiol. 2015, 6, 1–8. [Google Scholar] [CrossRef]
- Lakshminarayanan, B.; Guinane, C.M.; O’Connor, P.M.; Coakley, M.; Hill, C.; Stanton, C.; O’Toole, P.W.; Ross, R.P. Isolation and Characterization of Bacteriocin-Producing Bacteria from the Intestinal Microbiota of Elderly Irish Subjects. J. Appl. Microbiol. 2013, 114, 886–898. [Google Scholar] [CrossRef]
- Håversen, L.; Ohlsson, B.G.; Hahn-Zoric, M.; Hanson, L.Å.; Mattsby-Baltzer, I. Lactoferrin down-Regulates the LPS-Induced Cytokine Production in Monocytic Cells via NF-κB. Cell. Immunol. 2002, 220, 83–95. [Google Scholar] [CrossRef]
- Davidson, D.J.; Currie, A.J.; Reid, G.S.D.; Bowdish, D.M.E.; MacDonald, K.L.; Ma, R.C.; Hancock, R.E.W.; Speert, D.P. The Cationic Antimicrobial Peptide LL-37 Modulates Dendritic Cell Differentiation and Dendritic Cell-Induced T Cell Polarization. J. Immunol. 2004, 172, 1146–1156. [Google Scholar] [CrossRef] [PubMed]
- Mookherjee, N.; Brown, K.L.; Bowdish, D.M.E.; Doria, S.; Falsafi, R.; Hokamp, K.; Roche, F.M.; Mu, R.; Doho, G.H.; Pistolic, J.; et al. Modulation of the TLR-Mediated Inflammatory Response by the Endogenous Human Host Defense Peptide LL-37. J. Immunol. 2006, 176, 2455–2464. [Google Scholar] [CrossRef]
- Lai, Y.; Gallo, R.L. AMPed up Immunity: How Antimicrobial Peptides Have Multiple Roles in Immune Defense. Trends Immunol. 2009, 30, 131–141. [Google Scholar] [CrossRef] [PubMed]
- van der Does, A.M.; Bogaards, S.J.P.; Ravensbergen, B.; Beekhuizen, H.; van Dissel, J.T.; Nibbering, P.H. Antimicrobial Peptide hLF1-11 Directs Granulocyte-Macrophage Colony-Stimulating Factor-Driven Monocyte Differentiation toward Macrophages with Enhanced Recognition and Clearance of Pathogens. Antimicrob. Agents Chemother. 2010, 54, 811–816. [Google Scholar] [CrossRef]
- Yeung, A.T.Y.; Gellatly, S.L.; Hancock, R.E.W. Multifunctional Cationic Host Defence Peptides and Their Clinical Applications. Cell. Mol. Life Sci. 2011, 68, 2161–2176. [Google Scholar] [CrossRef]
- Choi, K.-Y.; Chow, L.N.Y.; Mookherjee, N. Cationic Host Defence Peptides: Multifaceted Role in Immune Modulation and Inflammation. J. Innate Immun. 2012, 4, 361–370. [Google Scholar] [CrossRef]
- Scheenstra, M.R.; van Harten, R.M.; Veldhuizen, E.J.A.; Haagsman, H.P.; Coorens, M. Cathelicidins Modulate TLR-Activation and Inflammation. Front. Immunol. 2020, 11, 1137. [Google Scholar] [CrossRef]
- Nagaoka, I.; Hirota, S.; Niyonsaba, F.; Hirata, M.; Adachi, Y.; Tamura, H.; Tanaka, S.; Heumann, D. Augmentation of the Lipopolysaccharide-Neutralizing Activities of Human Cathelicidin CAP18/LL-37-Derived Antimicrobial Peptides by Replacement with Hydrophobic and Cationic Amino Acid Residues. Clin. Vaccine Immunol. 2002, 9, 972–982. [Google Scholar] [CrossRef]
- Lanne, A.B.M.; Goode, A.; Prattley, C.; Kumari, D.; Drasbek, M.R.; Williams, P.; Conde-Álvarez, R.; Moriyón, I.; Bonev, B.B. Molecular Recognition of Lipopolysaccharide by the Lantibiotic Nisin. Biochim. Biophys. Acta Biomembr. 2019, 1861, 83–92. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.J.; Choi, H.J.; Kang, T.W.; Kim, H.O.; Chung, M.J.; Park, Y.M. CBT-SL5, a Bacteriocin from Enterococcus Faecalis, Suppresses the Expression of Interleukin-8 Induced by Propionibacterium Acnes in Cultured Human Keratinocytes. J. Microbiol. Biotechnol. 2008, 18, 1308–1316. [Google Scholar] [PubMed]
- Sand, S.L.; Oppegård, C.; Ohara, S.; Iijima, T.; Naderi, S.; Blomhoff, H.K.; Nissen-Meyer, J.; Sand, O. Plantaricin A, a Peptide Pheromone Produced by Lactobacillus Plantarum, Permeabilizes the Cell Membrane of Both Normal and Cancerous Lymphocytes and Neuronal Cells. Peptides 2010, 31, 1237–1244. [Google Scholar] [CrossRef] [PubMed]
- Pinto, D.; Marzani, B.; Minervini, F.; Calasso, M.; Giuliani, G.; Gobbetti, M.; De Angelis, M. Plantaricin A Synthesized by Lactobacillus Plantarum Induces in Vitro Proliferation and Migration of Human Keratinocytes and Increases the Expression of TGF-β1, FGF7, VEGF-A and IL-8 Genes. Peptides 2011, 32, 1815–1824. [Google Scholar] [CrossRef]
- Begde, D.; Bundale, S.; Mashitha, P.; Rudra, J.; Nashikkar, N.; Upadhyay, A. Immunomodulatory Efficacy of Nisin—a Bacterial Lantibiotic Peptide. J. Pept. Sci. 2011, 17, 438–444. [Google Scholar] [CrossRef] [PubMed]
- Kindrachuk, J.; Jenssen, H.; Elliott, M.; Nijnik, A.; Magrangeas-Janot, L.; Pasupuleti, M.; Thorson, L.; Ma, S.; Easton, D.M.; Bains, M.; et al. Manipulation of Innate Immunity by a Bacterial Secreted Peptide: Lantibiotic Nisin Z Is Selectively Immunomodulatory. Innate Immun. 2013, 19, 315–327. [Google Scholar] [CrossRef]
- Malvisi, M.; Stuknytė, M.; Magro, G.; Minozzi, G.; Giardini, A.; De Noni, I.; Piccinini, R. Antibacterial Activity and Immunomodulatory Effects on a Bovine Mammary Epithelial Cell Line Exerted by Nisin A-Producing Lactococcus Lactis Strains. J. Dairy Sci. 2016, 99, 2288–2296. [Google Scholar] [CrossRef] [PubMed]
- Molhoek, E.M.; Margo Molhoek, E.; den Hertog, A.L.; de Vries, A.-M.B.C.; Nazmi, K.; Veerman, E.C.I.; Hartgers, F.C.; Yazdanbakhsh, M.; Bikker, F.J.; van der Kleij, D. Structure-Function Relationship of the Human Antimicrobial Peptide LL-37 and LL-37 Fragments in the Modulation of TLR Responses. Biol. Chem. 2009, 390, 295–303. [Google Scholar] [CrossRef] [PubMed]
- Nagaoka, I.; Hirota, S.; Niyonsaba, F.; Hirata, M.; Adachi, Y.; Tamura, H.; Heumann, D. Cathelicidin Family of Antibacterial Peptides CAP18 and CAP11 Inhibit the Expression of TNF-α by Blocking the Binding of LPS to CD14 Cells. J. Immunol. 2001, 167, 3329–3338. [Google Scholar] [CrossRef] [PubMed]
- Rosenfeld, Y.; Papo, N.; Shai, Y. Endotoxin (lipopolysaccharide) Neutralization by Innate Immunity Host-Defense Peptides. Peptide Properties and Plausible Modes of Action. J. Biol. Chem. 2006, 281, 1636–1643. [Google Scholar] [CrossRef] [PubMed]
- Jia, Z.; He, M.; Wang, C.; Chen, A.; Zhang, X.; Xu, J.; Fu, H.; Liu, B. Nisin Reduces Uterine Inflammation in Rats by Modulating Concentrations of Pro- and Anti-inflammatory Cytokines. Am. J. Reprod. Immunol. 2019, 81, e13096. [Google Scholar] [CrossRef]
- Anitha, M.; Vijay–Kumar, M.; Sitaraman, S.V.; Gewirtz, A.T.; Srinivasan, S. Gut Microbial Products Regulate Murine Gastrointestinal Motility via Toll-Like Receptor 4 Signaling. Gastroenterology 2012, 143, 1006–1016.e4. [Google Scholar] [CrossRef]
- Brun, P.; Giron, M.C.; Qesari, M.; Porzionato, A.; Caputi, V.; Zoppellaro, C.; Banzato, S.; Grillo, A.R.; Spagnol, L.; De Caro, R.; et al. Toll-Like Receptor 2 Regulates Intestinal Inflammation by Controlling Integrity of the Enteric Nervous System. Gastroenterology 2013, 145, 1323–1333. [Google Scholar] [CrossRef] [PubMed]
- Sepehri, Z.; Kiani, Z.; Kohan, F.; Alavian, S.M.; Ghavami, S. Toll like Receptor 4 and Hepatocellular Carcinoma; A Systematic Review. Life Sci. 2017, 179, 80–87. [Google Scholar] [CrossRef]
- Etienne-Mesmin, L.; Vijay-Kumar, M.; Gewirtz, A.T.; Chassaing, B. Hepatocyte Toll-Like Receptor 5 Promotes Bacterial Clearance and Protects Mice Against High-Fat Diet–Induced Liver Disease. Cell. Mol. Gastroenterol. Hepatol. 2016, 2, 584–604. [Google Scholar] [CrossRef]
- Zou, H.; Wang, W.-K.; Liu, Y.-L.; Braddock, M.; Zheng, M.-H.; Huang, D.-S. Toll-like Receptors in Hepatocellular Carcinoma: Potential Novel Targets for Pharmacological Intervention. Expert Opin. Ther. Targets 2016, 20, 1127–1135. [Google Scholar] [CrossRef]
- Lee, Y.-S.; Kim, Y.-H.; Jung, Y.S.; Kim, K.-S.; Kim, D.-K.; Na, S.-Y.; Lee, J.-M.; Lee, C.-H.; Choi, H.-S. Hepatocyte Toll-like Receptor 4 Mediates Lipopolysaccharide-Induced Hepcidin Expression. Exp. Mol. Med. 2017, 49, e408. [Google Scholar] [CrossRef]
- Zhang, C.; Yang, M.; Ericsson, A.C. Antimicrobial Peptides: Potential Application in Liver Cancer. Front. Microbiol. 2019, 10, 1257. [Google Scholar] [CrossRef] [PubMed]
- Spadoni, I.; Zagato, E.; Bertocchi, A.; Paolinelli, R.; Hot, E.; Di Sabatino, A.; Caprioli, F.; Bottiglieri, L.; Oldani, A.; Viale, G.; et al. A Gut-Vascular Barrier Controls the Systemic Dissemination of Bacteria. Science 2015, 350, 830–834. [Google Scholar] [CrossRef]
- Dreyer, L. The Ability of Antimicrobial Peptides to Migrate across the Gastrointestinal Epithelial and Vascular Endothelial Barriers; Stellenbosch University: Stellenbosch, South Africa, 2018. [Google Scholar]
- Girardin, S.E.; Boneca, I.G.; Carneiro, L.A.M.; Antignac, A.; Jéhanno, M.; Viala, J.; Tedin, K.; Taha, M.-K.; Labigne, A.; Zähringer, U.; et al. Nod1 Detects a Unique Muropeptide from Gram-Negative Bacterial Peptidoglycan. Science 2003, 300, 1584–1587. [Google Scholar] [CrossRef]
- Zhao, Y.; Yang, J.; Shi, J.; Gong, Y.-N.; Lu, Q.; Xu, H.; Liu, L.; Shao, F. The NLRC4 Inflammasome Receptors for Bacterial Flagellin and Type III Secretion Apparatus. Nature 2011, 477, 596–600. [Google Scholar] [CrossRef] [PubMed]
- Rayamajhi, M.; Zak, D.E.; Chavarria-Smith, J.; Vance, R.E.; Miao, E.A. Cutting Edge: Mouse NAIP1 Detects the Type III Secretion System Needle Protein. J. Immunol. 2013, 191, 3986–3989. [Google Scholar] [CrossRef]
- Liwinski, T.; Zheng, D.; Elinav, E. The Microbiome and Cytosolic Innate Immune Receptors. Immunol. Rev. 2020, 297, 207–224. [Google Scholar] [CrossRef]
- Christgen, S.; Kanneganti, T.-D. Inflammasomes and the Fine Line between Defense and Disease. Curr. Opin. Immunol. 2020, 62, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Swanson, K.V.; Deng, M.; Ting, J.P.-Y. The NLRP3 Inflammasome: Molecular Activation and Regulation to Therapeutics. Nat. Rev. Immunol. 2019, 19, 477–489. [Google Scholar] [CrossRef] [PubMed]
- Wells, J.M.; Rossi, O.; Meijerink, M. Epithelial Crosstalk at the Microbiota–mucosal Interface. Proc. Natl. Acad. Sci. USA 2011, 108, 4607–4614. [Google Scholar] [CrossRef]
- Mukherjee, T.; Hovingh, E.S.; Foerster, E.G.; Abdel-Nour, M.; Philpott, D.J.; Girardin, S.E. NOD1 and NOD2 in Inflammation, Immunity and Disease. Arch. Biochem. Biophys. 2019, 670, 69–81. [Google Scholar] [CrossRef] [PubMed]
- Cario, E. Bacterial Interactions with Cells of the Intestinal Mucosa: Toll-like Receptors and NOD2. Gut 2005, 54, 1182–1193. [Google Scholar] [CrossRef]
- McDaniel, M.M.; Kottyan, L.C.; Singh, H.; Pasare, C. Suppression of Inflammasome Activation by IRF8 and IRF4 in cDCs Is Critical for T Cell Priming. Cell Rep. 2020, 31, 107604. [Google Scholar]
- Nagyőszi, P.; Nyúl-Tóth, Á; Fazakas, C.; Wilhelm, I.; Kozma, M.; Molnár, J.; Haskó, J.; Krizbai, I.A. Regulation of NOD-like Receptors and Inflammasome Activation in Cerebral Endothelial Cells. J. Neurochem. 2015, 135, 551–564. [Google Scholar] [CrossRef] [PubMed]
- Clarke, T.B.; Davis, K.M.; Lysenko, E.S.; Zhou, A.Y.; Yu, Y.; Weiser, J.N. Recognition of Peptidoglycan from the Microbiota by Nod1 Enhances Systemic Innate Immunity. Nat. Med. 2010, 16, 228–231. [Google Scholar] [CrossRef]
- Chen, G.Y.; Shaw, M.H.; Redondo, G.; Núñez, G. The Innate Immune Receptor Nod1 Protects the Intestine from Inflammation-Induced Tumorigenesis. Cancer Res. 2008, 68, 10060–10067. [Google Scholar] [CrossRef]
- Jung, C.; Hugot, J.-P.; Barreau, F. Peyer’s Patches: The Immune Sensors of the Intestine. Int. J. Inflam. 2010, 2010, 823710. [Google Scholar] [CrossRef]
- Barreau, F.; Madre, C.; Meinzer, U.; Berrebi, D.; Dussaillant, M.; Merlin, F.; Eckmann, L.; Karin, M.; Sterkers, G.; Bonacorsi, S.; et al. Nod2 Regulates the Host Response towards Microflora by Modulating T Cell Function and Epithelial Permeability in Mouse Peyer’s Patches. Gut 2010, 59, 207–217. [Google Scholar] [CrossRef]
- Kobayashi, K.S.; Chamaillard, M.; Ogura, Y.; Henegariu, O.; Inohara, N.; Nuñez, G.; Flavell, R.A. Nod2-Dependent Regulation of Innate and Adaptive Immunity in the Intestinal Tract. Science 2005, 307, 731–734. [Google Scholar] [CrossRef] [PubMed]
- Al Nabhani, Z.; Lepage, P.; Mauny, P.; Montcuquet, N.; Roy, M.; Le Roux, K.; Dussaillant, M.; Berrebi, D.; Hugot, J.-P.; Barreau, F. Nod2 Deficiency Leads to a Specific and Transmissible Mucosa-Associated Microbial Dysbiosis Which Is Independent of the Mucosal Barrier Defect. J. Crohns. Colitis 2016, 10, 1428–1436. [Google Scholar] [CrossRef]
- Ramanan, D.; Tang, M.S.; Bowcutt, R.; Loke, P. ’ng; Cadwell, K. Bacterial Sensor Nod2 Prevents Inflammation of the Small Intestine by Restricting the Expansion of the Commensal Bacteroides Vulgatus. Immunity 2014, 41, 311–324. [Google Scholar] [CrossRef] [PubMed]
- Hugot, J.P.; Chamaillard, M.; Zouali, H.; Lesage, S.; Cézard, J.P.; Belaiche, J.; Almer, S.; Tysk, C.; O’Morain, C.A.; Gassull, M.; et al. Association of NOD2 Leucine-Rich Repeat Variants with Susceptibility to Crohn’s Disease. Nature 2001, 411, 599–603. [Google Scholar] [CrossRef] [PubMed]
- Imhann, F.; Vich Vila, A.; Bonder, M.J.; Fu, J.; Gevers, D.; Visschedijk, M.C.; Spekhorst, L.M.; Alberts, R.; Franke, L.; van Dullemen, H.M.; et al. Interplay of Host Genetics and Gut Microbiota Underlying the Onset and Clinical Presentation of Inflammatory Bowel Disease. Gut 2018, 67, 108–119. [Google Scholar] [CrossRef]
- Schertzer, J.D.; Tamrakar, A.K.; Magalhães, J.G.; Pereira, S.; Bilan, P.J.; Fullerton, M.D.; Liu, Z.; Steinberg, G.R.; Giacca, A.; Philpott, D.J.; et al. NOD1 Activators Link Innate Immunity to Insulin Resistance. Diabetes 2011, 60, 2206–2215. [Google Scholar] [CrossRef] [PubMed]
- Chan, K.L.; Tam, T.H.; Boroumand, P.; Prescott, D.; Costford, S.R.; Escalante, N.K.; Fine, N.; Tu, Y.; Robertson, S.J.; Prabaharan, D.; et al. Circulating NOD1 Activators and Hematopoietic NOD1 Contribute to Metabolic Inflammation and Insulin Resistance. Cell Rep. 2017, 18, 2415–2426. [Google Scholar] [CrossRef]
- Denou, E.; Lolmède, K.; Garidou, L.; Pomie, C.; Chabo, C.; Lau, T.C.; Fullerton, M.D.; Nigro, G.; Zakaroff-Girard, A.; Luche, E.; et al. Defective NOD2 Peptidoglycan Sensing Promotes Diet-Induced Inflammation, Dysbiosis, and Insulin Resistance. EMBO Mol. Med. 2015, 7, 259–274. [Google Scholar] [CrossRef]
- Cavallari, J.F.; Fullerton, M.D.; Duggan, B.M.; Foley, K.P.; Denou, E.; Smith, B.K.; Desjardins, E.M.; Henriksbo, B.D.; Kim, K.J.; Tuinema, B.R.; et al. Muramyl Dipeptide-Based Postbiotics Mitigate Obesity-Induced Insulin Resistance via IRF4. Cell Metab. 2017, 25, 1063–1074.e3. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, T.; Asano, N.; Meng, G.; Yamashita, K.; Arai, Y.; Sakurai, T.; Kudo, M.; Fuss, I.J.; Kitani, A.; Shimosegawa, T.; et al. NOD2 Downregulates Colonic Inflammation by IRF4-Mediated Inhibition of K63-Linked Polyubiquitination of RICK and TRAF6. Mucosal Immunol. 2014, 7, 1312–1325. [Google Scholar] [CrossRef]
- Cavallari, J.F.; Barra, N.G.; Foley, K.P.; Lee, A.; Duggan, B.M.; Henriksbo, B.D.; Anhê, F.F.; Ashkar, A.A.; Schertzer, J.D. Postbiotics for NOD2 Require Nonhematopoietic RIPK2 to Improve Blood Glucose and Metabolic Inflammation in Mice. Am. J. Physiol. Endocrinol. Metab. 2020, 318, E579–E585. [Google Scholar] [CrossRef]
- Zeuthen, L.H.; Fink, L.N.; Frøkiaer, H. Toll-like Receptor 2 and Nucleotide-Binding Oligomerization Domain-2 Play Divergent Roles in the Recognition of Gut-Derived Lactobacilli and Bifidobacteria in Dendritic Cells. Immunology 2008, 124, 489–502. [Google Scholar] [CrossRef]
- Foligne, B.; Zoumpopoulou, G.; Dewulf, J.; Ben Younes, A.; Chareyre, F.; Sirard, J.-C.; Pot, B.; Grangette, C. A Key Role of Dendritic Cells in Probiotic Functionality. PLoS ONE 2007, 2, e313. [Google Scholar] [CrossRef] [PubMed]
- Kwon, H.K.; Lee, C.G.; So, J.S.; Chae, C.S. Generation of Regulatory Dendritic Cells and CD4+ Foxp3+ T Cells by Probiotics Administration Suppresses Immune Disorders. Proc. Natl. Acad. Sci. USA 2010, 107, 2159–2164. [Google Scholar] [CrossRef]
- Kolling, Y.; Salva, S.; Villena, J.; Marranzino, G.; Alvarez, S. Non-Viable Immunobiotic Lactobacillus Rhamnosus CRL1505 and Its Peptidoglycan Improve Systemic and Respiratory Innate Immune Response during Recovery of Immunocompromised-Malnourished Mice. Int. Immunopharmacol. 2015, 25, 474–484. [Google Scholar] [CrossRef]
- Kolling, Y.; Salva, S.; Villena, J.; Alvarez, S. Are the Immunomodulatory Properties of Lactobacillus Rhamnosus CRL1505 Peptidoglycan Common for All Lactobacilli during Respiratory Infection in Malnourished Mice? PLoS ONE 2018, 13, e0194034. [Google Scholar] [CrossRef]
- Melnyk, J.E.; Mohanan, V.; Schaefer, A.K.; Hou, C.-W.; Grimes, C.L. Peptidoglycan Modifications Tune the Stability and Function of the Innate Immune Receptor Nod2. J. Am. Chem. Soc. 2015, 137, 6987–6990. [Google Scholar] [CrossRef] [PubMed]
- Vollmer, W. Structural Variation in the Glycan Strands of Bacterial Peptidoglycan. FEMS Microbiol. Rev. 2008, 32, 287–306. [Google Scholar] [CrossRef]
- Wolf, A.J.; Reyes, C.N.; Liang, W.; Becker, C.; Shimada, K.; Wheeler, M.L.; Cho, H.C.; Popescu, N.I.; Coggeshall, K.M.; Arditi, M.; et al. Hexokinase Is an Innate Immune Receptor for the Detection of Bacterial Peptidoglycan. Cell 2016, 166, 624–636. [Google Scholar] [CrossRef]
- Bernard, E.; Rolain, T.; Courtin, P.; Hols, P.; Chapot-Chartier, M.-P. Identification of the Amidotransferase AsnB1 as Being Responsible Formeso-Diaminopimelic Acid Amidation in Lactobacillus Plantarum Peptidoglycan. J. Bacteriol. 2011, 193, 6323–6330. [Google Scholar] [CrossRef] [PubMed]
- Vijayrajratnam, S.; Pushkaran, A.C.; Balakrishnan, A.; Vasudevan, A.K.; Biswas, R.; Mohan, C.G. Bacterial Peptidoglycan with Amidated Meso-Diaminopimelic Acid Evades NOD1 Recognition: An Insight into NOD1 Structure–recognition. Biochem. J 2016, 473, 4573–4592. [Google Scholar] [CrossRef]
- Bernard, E.; Rolain, T.; David, B.; André, G.; Dupres, V.; Dufrêne, Y.F.; Hallet, B.; Chapot-Chartier, M.-P.; Hols, P. Dual Role for the O-Acetyltransferase OatA in Peptidoglycan Modification and Control of Cell Septation in Lactobacillus Plantarum. PLoS ONE 2012, 7, e47893. [Google Scholar] [CrossRef] [PubMed]
- Nagaoka, M.; Muto, M.; Nomoto, K.; Matuzaki, T.; Watanabe, T.; Yokokura, T. Structure of Polysaccharide-Peptidoglycan Complex from the Cell Wall of Lactobacillus Casei YIT9018. J. Biochem. 1990, 108, 568–571. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, S.; Hara, T.; Nagaoka, M.; Mike, A.; Mitsuyama, K.; Sako, T.; Yamamoto, M.; Kado, S.; Takada, T. A Component of Polysaccharide Peptidoglycan Complex on Lactobacillus Induced an Improvement of Murine Model of Inflammatory Bowel Disease and Colitis-Associated Cancer. Immunology 2009, 128, e170–e180. [Google Scholar] [CrossRef] [PubMed]
- Shida, K.; Kiyoshima-Shibata, J.; Kaji, R.; Nagaoka, M.; Nanno, M. Peptidoglycan from Lactobacilli Inhibits Interleukin-12 Production by Macrophages Induced by Lactobacillus Casei through Toll-like Receptor 2-Dependent and Independent Mechanisms. Immunology 2009, 128, e858–e869. [Google Scholar] [CrossRef]
- Wolf, A.J.; Underhill, D.M. Peptidoglycan Recognition by the Innate Immune System. Nat. Rev. Immunol. 2018, 18, 243–254. [Google Scholar] [CrossRef]
- Clua, P.; Kanmani, P.; Zelaya, H.; Tada, A.; Kober, A.K.M.H.; Salva, S.; Alvarez, S.; Kitazawa, H.; Villena, J. Peptidoglycan from Immunobiotic Lactobacillus Rhamnosus Improves Resistance of Infant Mice to Respiratory Syncytial Viral Infection and Secondary Pneumococcal Pneumonia. Front. Immunol. 2017, 8, 948. [Google Scholar] [CrossRef]
- Kozakova, H.; Schwarzer, M.; Tuckova, L.; Srutkova, D.; Czarnowska, E.; Rosiak, I.; Hudcovic, T.; Schabussova, I.; Hermanova, P.; Zakostelska, Z.; et al. Colonization of Germ-Free Mice with a Mixture of Three Lactobacillus Strains Enhances the Integrity of Gut Mucosa and Ameliorates Allergic Sensitization. Cell. Mol. Immunol. 2016, 13, 251–262. [Google Scholar] [CrossRef]
- Guo, H.; Gibson, S.A.; Ting, J.P.Y. Gut Microbiota, NLR Proteins, and Intestinal Homeostasis. J. Exp. Med. 2020, 217. [Google Scholar] [CrossRef]
- Zhen, Y.; Zhang, H. NLRP3 Inflammasome and Inflammatory Bowel Disease. Front. Immunol. 2019, 10, 276. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Elinav, E.; Huber, S.; Strowig, T.; Hao, L.; Hafemann, A.; Jin, C.; Wunderlich, C.; Wunderlich, T.; Eisenbarth, S.C.; et al. Microbiota-Induced Activation of Epithelial IL-6 Signaling Links Inflammasome-Driven Inflammation with Transmissible Cancer. Proc. Natl. Acad. Sci. USA 2013, 110, 9862–9867. [Google Scholar] [CrossRef] [PubMed]
- Kelley, N.; Jeltema, D.; Duan, Y.; He, Y. The NLRP3 Inflammasome: An Overview of Mechanisms of Activation and Regulation. Int. J. Mol. Sci. 2019, 20, 3328. [Google Scholar] [CrossRef]
- Andrade-Oliveira, V.; Foresto-Neto, O.; Watanabe, I.K.M.; Zatz, R.; Câmara, N.O.S. Inflammation in Renal Diseases: New and Old Players. Front. Pharmacol. 2019, 10, 1192. [Google Scholar] [CrossRef] [PubMed]
- Valiño-Rivas, L.; Cuarental, L.; Nuñez, G.; Sanz, A.B.; Ortiz, A.; Sanchez-Niño, M.D. Loss of NLRP6 Expression Increases the Severity of Acute Kidney Injury. Nephrol. Dial. Transpl. 2020, 35, 587–598. [Google Scholar] [CrossRef] [PubMed]
- Inserra, A.; Rogers, G.B.; Licinio, J.; Wong, M.-L. The Microbiota-Inflammasome Hypothesis of Major Depression. BioEssays 2018, 40, 1800027. [Google Scholar] [CrossRef]
- Truax, A.D.; Chen, L.; Tam, J.W.; Cheng, N.; Guo, H.; Koblansky, A.A.; Chou, W.-C.; Wilson, J.E.; Brickey, W.J.; Petrucelli, A.; et al. The Inhibitory Innate Immune Sensor NLRP12 Maintains a Threshold against Obesity by Regulating Gut Microbiota Homeostasis. Cell Host Microbe 2018, 24, 364–378.e6. [Google Scholar] [CrossRef]
- Singh, V.; Yeoh, B.S.; Walker, R.E.; Xiao, X.; Saha, P.; Golonka, R.M.; Cai, J.; Bretin, A.C.A.; Cheng, X.; Liu, Q.; et al. Microbiota Fermentation-NLRP3 Axis Shapes the Impact of Dietary Fibres on Intestinal Inflammation. Gut 2019, 68, 1801–1812. [Google Scholar] [CrossRef]
- Hara, H.; Seregin, S.S.; Yang, D.; Fukase, K.; Chamaillard, M.; Alnemri, E.S.; Inohara, N.; Chen, G.Y.; Núñez, G. The NLRP6 Inflammasome Recognizes Lipoteichoic Acid and Regulates Gram-Positive Pathogen Infection. Cell 2018, 175, 1651–1664.e14. [Google Scholar] [CrossRef] [PubMed]
- Venuprasad, K.; Theiss, A.L. NLRP6 in Host Defense and Intestinal Inflammation. Cell Rep. 2021, 35, 109043. [Google Scholar] [CrossRef]
- Levy, M.; Thaiss, C.A.; Zeevi, D.; Dohnalová, L.; Zilberman-Schapira, G.; Mahdi, J.A.; David, E.; Savidor, A.; Korem, T.; Herzig, Y.; et al. Microbiota-Modulated Metabolites Shape the Intestinal Microenvironment by Regulating NLRP6 Inflammasome Signaling. Cell 2015, 163, 1428–1443. [Google Scholar] [CrossRef]
- Frei, R.; Ferstl, R.; Konieczna, P.; Ziegler, M.; Simon, T.; Rugeles, T.M.; Mailand, S.; Watanabe, T.; Lauener, R.; Akdis, C.A.; et al. Histamine Receptor 2 Modifies Dendritic Cell Responses to Microbial Ligands. J. Allergy Clin. Immunol. 2013, 132, 194–204. [Google Scholar] [CrossRef] [PubMed]
- Ferstl, R.; Frei, R.; Schiavi, E.; Konieczna, P.; Barcik, W.; Ziegler, M.; Lauener, R.P.; Chassard, C.; Lacroix, C.; Akdis, C.A.; et al. Histamine Receptor 2 Is a Key Influence in Immune Responses to Intestinal Histamine-Secreting Microbes. J. Allergy Clin. Immunol. 2014, 134, 744–746. [Google Scholar] [CrossRef]
- Angosto-Bazarra, D.; Molina-López, C.; Peñín-Franch, A.; Hurtado-Navarro, L.; Pelegrín, P. Techniques to Study Inflammasome Activation and Inhibition by Small Molecules. Molecules 2021, 26, 1704. [Google Scholar] [CrossRef]
- Fritz, J.H.; Girardin, S.E.; Fitting, C.; Werts, C.; Mengin-Lecreulx, D.; Caroff, M.; Cavaillon, J.-M.; Philpott, D.J.; Adib-Conquy, M. Synergistic Stimulation of Human Monocytes and Dendritic Cells by Toll-like Receptor 4 and NOD1- and NOD2-Activating Agonists. Eur. J. Immunol. 2005, 35, 2459–2470. [Google Scholar] [CrossRef]
- Tada, H.; Aiba, S.; Shibata, K.-I.; Ohteki, T.; Takada, H. Synergistic Effect of Nod1 and Nod2 Agonists with Toll-like Receptor Agonists on Human Dendritic Cells to Generate Interleukin-12 and T Helper Type 1 Cells. Infect. Immun. 2005, 73, 7967–7976. [Google Scholar] [CrossRef] [PubMed]
- van Heel, D.A.; Ghosh, S.; Butler, M.; Hunt, K.; Foxwell, B.M.J.; Mengin-Lecreulx, D.; Playford, R.J. Synergistic Enhancement of Toll-like Receptor Responses by NOD1 Activation. Eur. J. Immunol. 2005, 35, 2471–2476. [Google Scholar] [CrossRef]
- Schirbel, A.; Kessler, S.; Rieder, F.; West, G.; Rebert, N.; Asosingh, K.; McDonald, C.; Fiocchi, C. Pro-Angiogenic Activity of TLRs and NLRs: A Novel Link between Gut Microbiota and Intestinal Angiogenesis. Gastroenterology 2013, 144, 613–623.e9. [Google Scholar] [CrossRef] [PubMed]
- Bäuerl, C.; Coll-Marqués, J.M.; Tarazona-González, C.; Pérez-Martínez, G. Lactobacillus Casei Extracellular Vesicles Stimulate EGFR Pathway Likely due to the Presence of Proteins P40 and P75 Bound to Their Surface. Sci. Rep. 2020, 10, 19237. [Google Scholar] [CrossRef]
- Bäuerl, C.; Abitayeva, G.; Sosa-Carrillo, S.; Mencher-Beltrán, A.; Navarro-Lleó, N.; Coll-Marqués, J.M.; Zúñiga-Cabrera, M.; Shaikhin, S.; Pérez-Martinez, G. P40 and P75 Are Singular Functional Muramidases Present in the Lactobacillus Casei /paracasei/rhamnosus Taxon. Front. Microbiol. 2019, 10, 1420. [Google Scholar] [CrossRef]
- Giani, A.D.; De Giani, A.; Bovio, F.; Forcella, M.; Fusi, P.; Sello, G.; Di Gennaro, P. Identification of a Bacteriocin-like Compound from Lactobacillus Plantarum with Antimicrobial Activity and Effects on Normal and Cancerogenic Human Intestinal Cells. AMB Express 2019, 9. [Google Scholar] [CrossRef]
- Wentworth, C.C.; Jones, R.M.; Kwon, Y.M.; Nusrat, A.; Neish, A.S. Commensal-Epithelial Signaling Mediated via Formyl Peptide Receptors. Am. J. Pathol. 2010, 177, 2782–2790. [Google Scholar] [CrossRef]
- Koczulla, R.; von Degenfeld, G.; Kupatt, C.; Krötz, F.; Zahler, S.; Gloe, T.; Issbrücker, K.; Unterberger, P.; Zaiou, M.; Lebherz, C.; et al. An Angiogenic Role for the Human Peptide Antibiotic LL-37/hCAP-18. J. Clin. Investig. 2003, 111, 1665–1672. [Google Scholar] [CrossRef]
- Yang, D.; Chertov, O.; Bykovskaia, S.N.; Chen, Q.; Buffo, M.J.; Shogan, J.; Anderson, M.; Schröder, J.M.; Wang, J.M.; Howard, O.M.; et al. Beta-Defensins: Linking Innate and Adaptive Immunity through Dendritic and T Cell CCR6. Science 1999, 286, 525–528. [Google Scholar] [CrossRef]
- Chambers, E.S.; Preston, T.; Frost, G.; Morrison, D.J. Role of Gut Microbiota-Generated Short-Chain Fatty Acids in Metabolic and Cardiovascular Health. Curr. Nutr. Rep. 2018, 7, 198–206. [Google Scholar] [CrossRef] [PubMed]
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly-Y, M.; Glickman, J.N.; Garrett, W.S. The Microbial Metabolites, Short-Chain Fatty Acids, Regulate Colonic Treg Cell Homeostasis. Science 2013, 341, 569–573. [Google Scholar] [CrossRef]
- Silva, Y.P.; Bernardi, A.; Frozza, R.L. The Role of Short-Chain Fatty Acids From Gut Microbiota in Gut-Brain Communication. Front. Endocrinol. 2020, 11, 25. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; van Esch, B.C.A.M.; Wagenaar, G.T.M.; Garssen, J.; Folkerts, G.; Henricks, P.A.J. Pro- and Anti-Inflammatory Effects of Short Chain Fatty Acids on Immune and Endothelial Cells. Eur. J. Pharmacol. 2018, 831, 52–59. [Google Scholar] [CrossRef]
- Hoving, J.C.; Wilson, G.J.; Brown, G.D. Signalling C-Type Lectin Receptors, Microbial Recognition and Immunity. Cell. Microbiol. 2014, 16, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.J.; Sancho, D.; Slack, E.C.; LeibundGut-Landmann, S.; Reis e Sousa, C. Myeloid C-Type Lectins in Innate Immunity. Nat. Immunol. 2006, 7, 1258–1265. [Google Scholar] [CrossRef]
- Dambuza, I.M.; Brown, G.D. C-Type Lectins in Immunity: Recent Developments. Curr. Opin. Immunol. 2015, 32, 21–27. [Google Scholar] [CrossRef]
- Mnich, M.E.; van Dalen, R.; van Sorge, N.M. C-Type Lectin Receptors in Host Defense Against Bacterial Pathogens. Front. Cell. Infect. Microbiol. 2020, 10, 309. [Google Scholar] [CrossRef] [PubMed]
- Kerrigan, A.M.; Brown, G.D. C-Type Lectins and Phagocytosis. Immunobiology 2009, 214, 562–575. [Google Scholar] [CrossRef]
- Chiffoleau, E. C-Type Lectin-Like Receptors As Emerging Orchestrators of Sterile Inflammation Represent Potential Therapeutic Targets. Front. Immunol. 2018, 9, 227. [Google Scholar] [CrossRef] [PubMed]
- Geijtenbeek, T.B.H.; Torensma, R.; van Vliet, S.J.; van Duijnhoven, G.C.F.; Adema, G.J.; van Kooyk, Y.; Figdor, C.G. Identification of DC-SIGN, a Novel Dendritic Cell–Specific ICAM-3 Receptor That Supports Primary Immune Responses. Cell 2000, 100, 575–585. [Google Scholar] [CrossRef]
- Lai, W.K.; Sun, P.J.; Zhang, J.; Jennings, A.; Lalor, P.F.; Hubscher, S.; McKeating, J.A.; Adams, D.H. Expression of DC-SIGN and DC-SIGNR on Human Sinusoidal Endothelium: A Role for Capturing Hepatitis C Virus Particles. Am. J. Pathol. 2006, 169, 200–208. [Google Scholar] [CrossRef] [PubMed]
- Tailleux, L.; Pham-Thi, N.; Bergeron-Lafaurie, A.; Herrmann, J.-L.; Charles, P.; Schwartz, O.; Scheinmann, P.; Lagrange, P.H.; de Blic, J.; Tazi, A.; et al. DC-SIGN Induction in Alveolar Macrophages Defines Privileged Target Host Cells for Mycobacteria in Patients with Tuberculosis. PLoS Med. 2005, 2, e381. [Google Scholar] [CrossRef] [PubMed]
- Svajger, U.; Anderluh, M.; Jeras, M.; Obermajer, N. C-Type Lectin DC-SIGN: An Adhesion, Signalling and Antigen-Uptake Molecule That Guides Dendritic Cells in Immunity. Cell. Signal. 2010, 22, 1397–1405. [Google Scholar] [CrossRef]
- Castro-Bravo, N.; Wells, J.M.; Margolles, A.; Ruas-Madiedo, P. Interactions of Surface Exopolysaccharides From Bifidobacterium and Lactobacillus Within the Intestinal Environment. Front. Microbiol. 2018, 9, 2426. [Google Scholar] [CrossRef]
- Castro-Bravo, N.; Margolles, A.; Wells, J.M.; Ruas-Madiedo, P. Exopolysaccharides Synthesized by Bifidobacterium Animalis Subsp. Lactis Interact with TLR4 in Intestinal Epithelial Cells. Anaerobe 2019, 56, 98–101. [Google Scholar] [CrossRef] [PubMed]
- López, P.; Monteserín, D.C.; Gueimonde, M.; de los Reyes-Gavilán, C.G.; Margolles, A.; Suárez, A.; Ruas-Madiedo, P. Exopolysaccharide-Producing Bifidobacterium Strains Elicit Different in Vitro Responses upon Interaction with Human Cells. Food Res. Int. 2012, 46, 99–107. [Google Scholar] [CrossRef]
- Górska, S.; Sandstrőm, C.; Wojas-Turek, J.; Rossowska, J.; Pajtasz-Piasecka, E.; Brzozowska, E.; Gamian, A. Structural and Immunomodulatory Differences among Lactobacilli Exopolysaccharides Isolated from Intestines of Mice with Experimentally Induced Inflammatory Bowel Disease. Sci. Rep. 2016, 6, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Bleau, C.; Monges, A.; Rashidan, K.; Laverdure, J.-P.; Lacroix, M.; Van Calsteren, M.-R.; Millette, M.; Savard, R.; Lamontagne, L. Intermediate Chains of Exopolysaccharides from Lactobacillus Rhamnosus RW-9595M Increase IL-10 Production by Macrophages. J. Appl. Microbiol. 2010, 108, 666–675. [Google Scholar] [CrossRef] [PubMed]
- Mazmanian, S.K.; Round, J.L.; Kasper, D.L. A Microbial Symbiosis Factor Prevents Intestinal Inflammatory Disease. Nature 2008, 453, 620–625. [Google Scholar] [CrossRef] [PubMed]
- Laiño, J.; Villena, J.; Kanmani, P.; Kitazawa, H. Immunoregulatory Effects Triggered by Lactic Acid Bacteria Exopolysaccharides: New Insights into Molecular Interactions with Host Cells. Microorganisms 2016, 4, 27. [Google Scholar] [CrossRef] [PubMed]
- Konieczna, P.; Schiavi, E.; Ziegler, M.; Groeger, D.; Healy, S.; Grant, R.; O’Mahony, L. Human Dendritic Cell DC-SIGN and TLR-2 Mediate Complementary Immune Regulatory Activities in Response to Lactobacillus Rhamnosus JB-1. PLoS ONE 2015, 10, e0120261. [Google Scholar] [CrossRef]
- Tytgat, H.L.P.; van Teijlingen, N.H.; Sullan, R.M.A.; Douillard, F.P.; Rasinkangas, P.; Messing, M.; Reunanen, J.; Satokari, R.; Vanderleyden, J.; Dufrêne, Y.F.; et al. Probiotic Gut Microbiota Isolate Interacts with Dendritic Cells via Glycosylated Heterotrimeric Pili. PLoS ONE 2016, 11, e0151824. [Google Scholar] [CrossRef]
- Fagan, R.P.; Fairweather, N.F. Biogenesis and Functions of Bacterial S-Layers. Nat. Rev. Microbiol. 2014, 12, 211–222. [Google Scholar] [CrossRef]
- Sleytr, U.B.; Schuster, B.; Egelseer, E.-M.; Pum, D. S-Layers: Principles and Applications. FEMS Microbiol. Rev. 2014, 38, 823–864. [Google Scholar] [CrossRef]
- Konstantinov, S.R.; Smidt, H.; de Vos, W.M.; Bruijns, S.C.M.; Singh, S.K.; Valence, F.; Molle, D.; Lortal, S.; Altermann, E.; Klaenhammer, T.R.; et al. S Layer Protein A of Lactobacillus Acidophilus NCFM Regulates Immature Dendritic Cell and T Cell Functions. Proc. Natl. Acad. Sci. USA 2008, 105, 19474–19479. [Google Scholar] [CrossRef] [PubMed]
- Chapot-Chartier, M.-P.; Kulakauskas, S. Cell Wall Structure and Function in Lactic Acid Bacteria. Microb. Cell Factories 2014, 13, S9. [Google Scholar] [CrossRef] [PubMed]
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial Biofilms: A Common Cause of Persistent Infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef] [PubMed]
- Hold, G.L.; Garrett, W.S. Gut Microbiota. Microbiota Organization—A Key to Understanding CRC Development. Nat. Rev. Gastroenterol. Hepatol. 2015, 12, 128–129. [Google Scholar] [CrossRef] [PubMed]
- Puccetti, M.; Xiroudaki, S.; Ricci, M.; Giovagnoli, S. Postbiotic-Enabled Targeting of the Host-Microbiota-Pathogen Interface: Hints of Antibiotic Decline? Pharmaceutics 2020, 12, 624. [Google Scholar] [CrossRef]
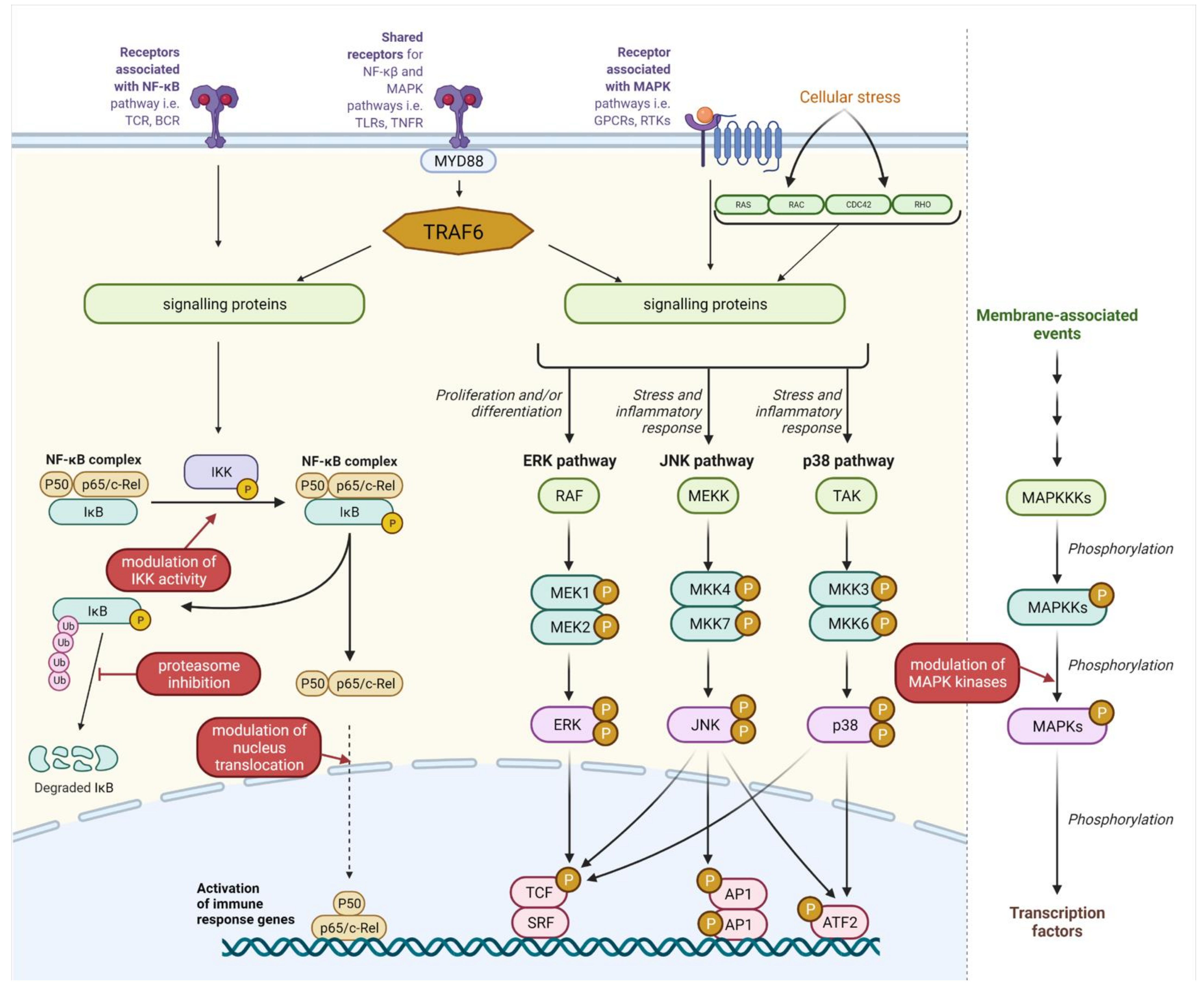
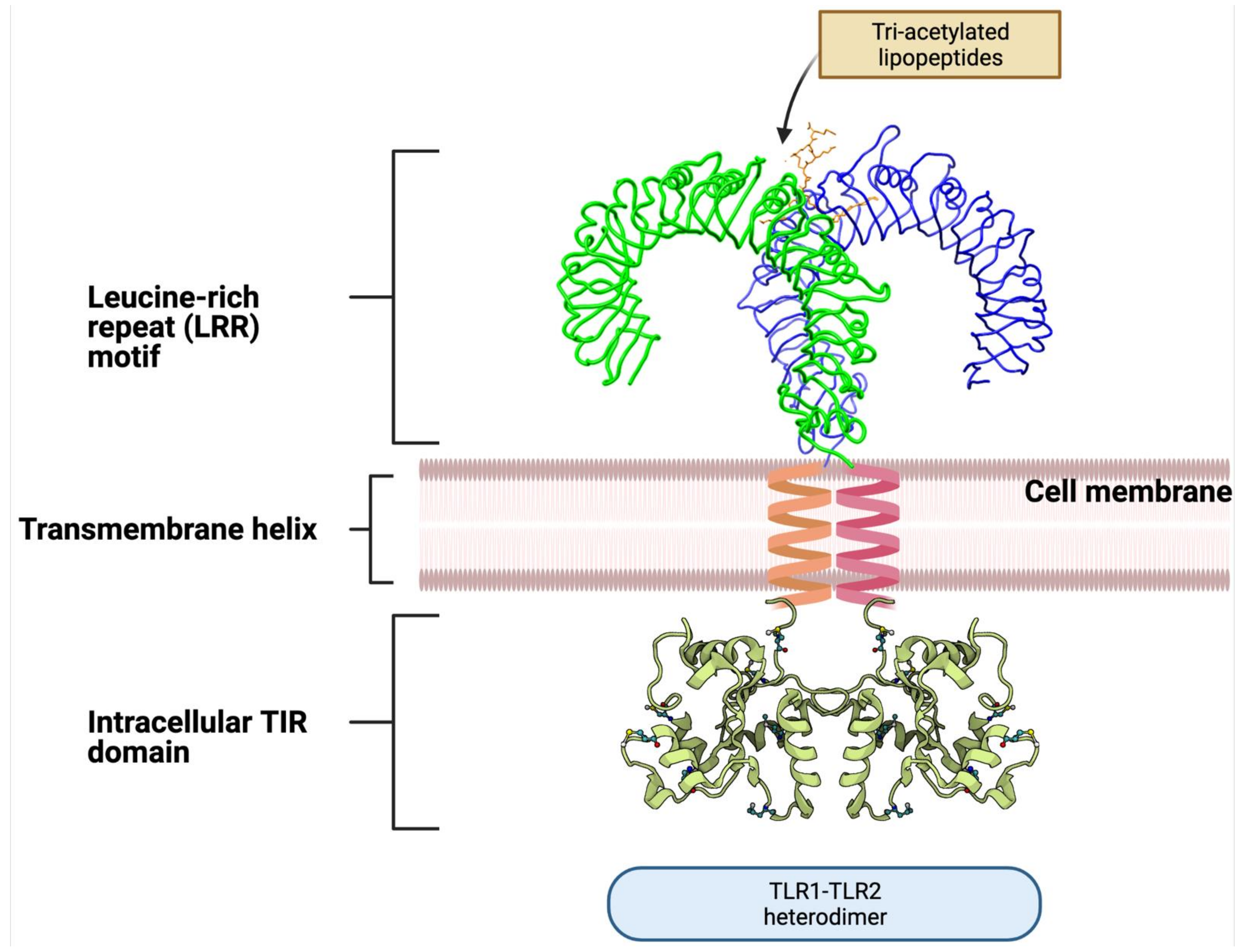
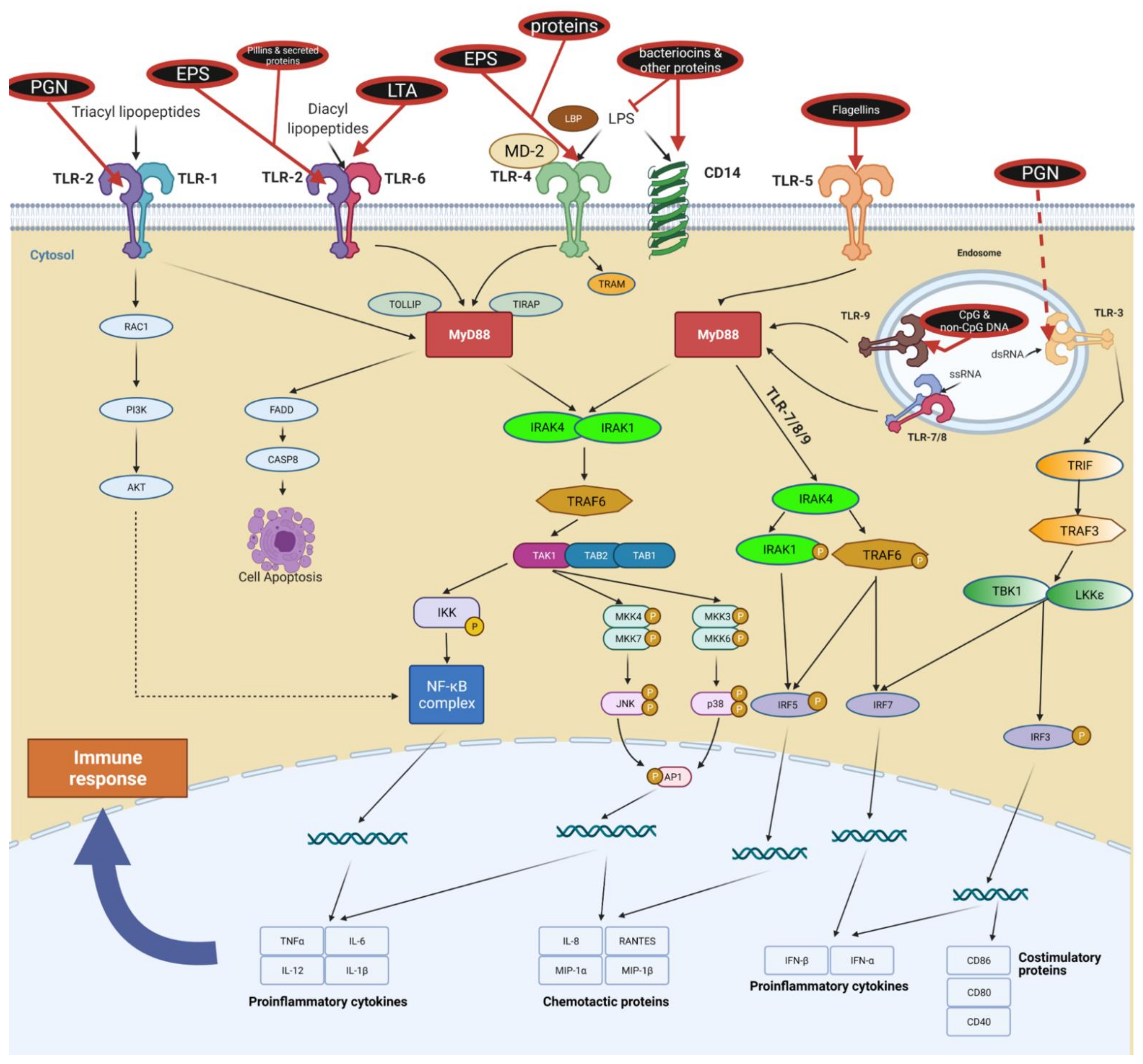
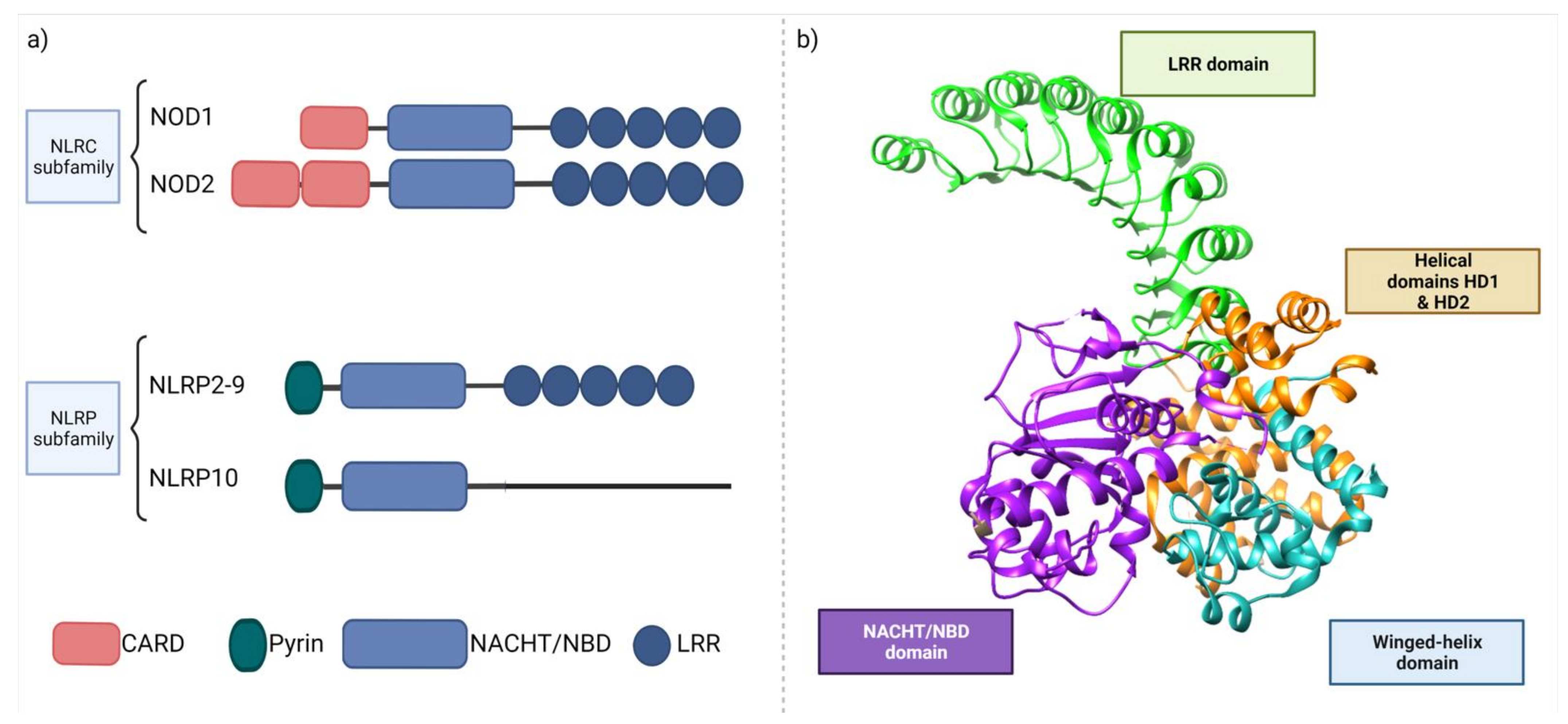

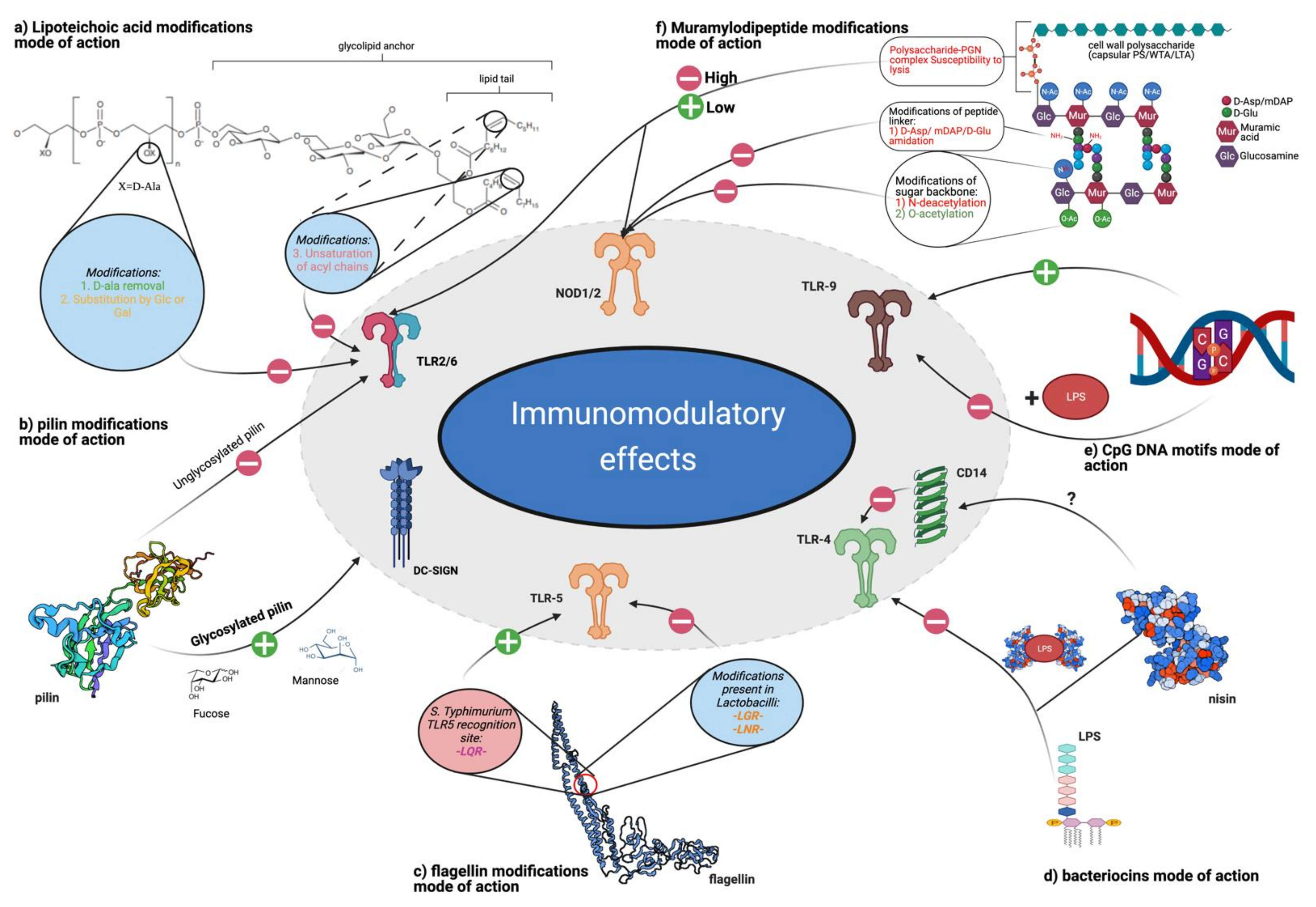
| Postbiotics and Biologically Active Bacterial Byproducts | |
|---|---|
| Supernatant-Derived (CFS) | Cell-Derived (HKB) |
| |
| |
| Receptor | Receptor Localization | Ligand | Origin of Ligand |
|---|---|---|---|
| TLR2 | Cell surface | Lipoteichoic acid | G (+) bacteria |
| Lipoprotein/lipopeptides | Various pathogens | ||
| Hemagglutinin protein | Viruses (Measles Virus) | ||
| Glycosyl-phosphatidylinositols | Parasites (Toxoplasma gondii) | ||
| Exopolysaccharides | G (+) bacteria | ||
| TLR2/1 | Cell surface | Triacyl lipopeptides | Bacteria and mycobacteria |
| TLR2/6 | Cell surface | Diacyl lipopeptides | Mycobacteria |
| Zymosan | Fungi | ||
| TLR3 | Cellular compartment (endosomes) | dsRNA | Viruses |
| TLR4 | Cell surface | Lipopolysaccharide | G (−) bacteria |
| Envelope proteins | Viruses (Respiratory Syncytial Virus) | ||
| Glycosyl-phosphatidylinositols | Parasites (Toxoplasma gondii) | ||
| TLR5 | Cell surface | Flagellin | Bacteria |
| TLR7/8 | Cellular compartment (endosomes) | ssRNA | Viruses |
| TLR9 | Cellular compartment (endosomes)/cell surface | CpG-containing DNA | Bacteria and viruses |
| TLR11 | Cell surface | Uropathogenic bacteria component | Bacteria (uropathogenic Escherichia coli) |
| Profilin | Parasites | ||
| NOD1 | Cell cytoplasm | Meso-diaminopimelic acid | PGN of G (−) and some G (+) bacteria |
| NOD2 | Cell cytoplasm | Muramyl dipeptide (MDP) | PGN of G (+) and G (−) bacteria |
| Receptor | Ligand | Outcome | Reference |
|---|---|---|---|
| EGFR | p40 and p75 plantaricin P1053 | phosphorylation of Akt and ERK kinases; increased viability of non-cancerogenic cells | [303,304,305] |
| formyl peptide receptors (FPRs) | N-formyl-Met-Leu-Phe; HK Lactobacillus | ERK phosphorylation without stimulating pro-inflammatory phospho-IκB or pro-apoptotic phospho-c-Jun NH2-terminal kinase | [306] |
| formyl peptide receptor-like 1 (FPRL1) | human bacteriocin LL-37 | upregulated angiogenesis via increased intracellular calcium levels | [307] |
| CCR6 | human bacteriocins beta-defensins | increased chemotaxis of dendritic and T cells | [308] |
| pregnane X receptor (PXR) | Indole derivatives (i.e., Indole-3-propionic acid) | improved tight junction functions by increased expression of claudins and occludins | [39] |
| aryl hydrogen receptors (AhRs) | indole-3-lactic | increased Il-22 expression | [40] |
| Short chain fatty acid receptors (GPR43, GPR41, GPR109a and Olfr78) | Short chain fatty acids (i.e., butyrate, propionate) | modulates AMP-kinase and NF-кkB activity regulates glucose homeostasis, immune processes, permeability of cell membrane, oxidative status, cell cycle or brain functions | [309,310,311,312] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jastrząb, R.; Graczyk, D.; Siedlecki, P. Molecular and Cellular Mechanisms Influenced by Postbiotics. Int. J. Mol. Sci. 2021, 22, 13475. https://doi.org/10.3390/ijms222413475
Jastrząb R, Graczyk D, Siedlecki P. Molecular and Cellular Mechanisms Influenced by Postbiotics. International Journal of Molecular Sciences. 2021; 22(24):13475. https://doi.org/10.3390/ijms222413475
Chicago/Turabian StyleJastrząb, Rafał, Damian Graczyk, and Pawel Siedlecki. 2021. "Molecular and Cellular Mechanisms Influenced by Postbiotics" International Journal of Molecular Sciences 22, no. 24: 13475. https://doi.org/10.3390/ijms222413475
APA StyleJastrząb, R., Graczyk, D., & Siedlecki, P. (2021). Molecular and Cellular Mechanisms Influenced by Postbiotics. International Journal of Molecular Sciences, 22(24), 13475. https://doi.org/10.3390/ijms222413475






