Increased Frequency of Copy Number Variations Revealed by Array Comparative Genomic Hybridization in the Offspring of Male Mice Exposed to Low Dose-Rate Ionizing Radiation
Abstract
1. Introduction
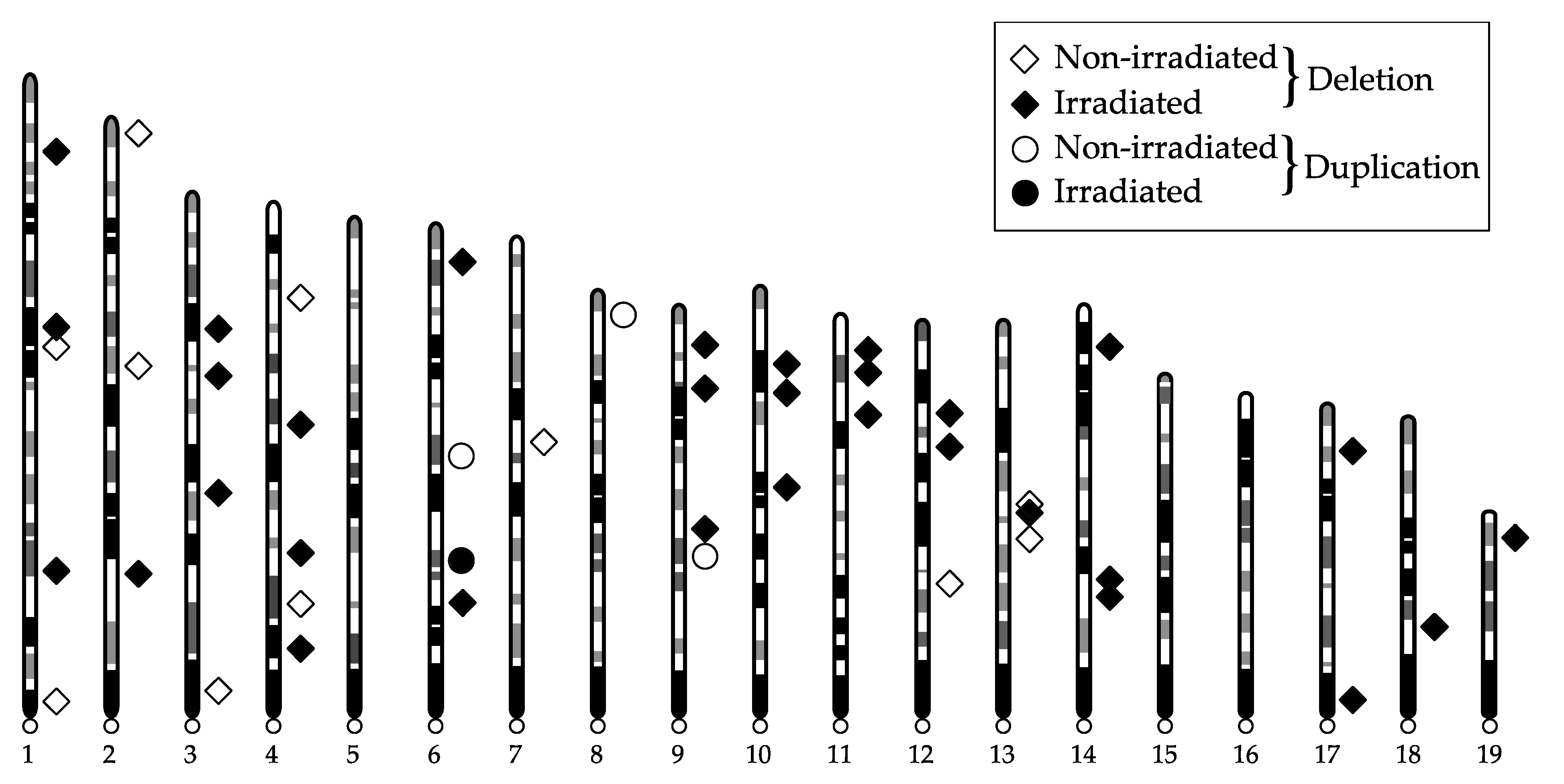
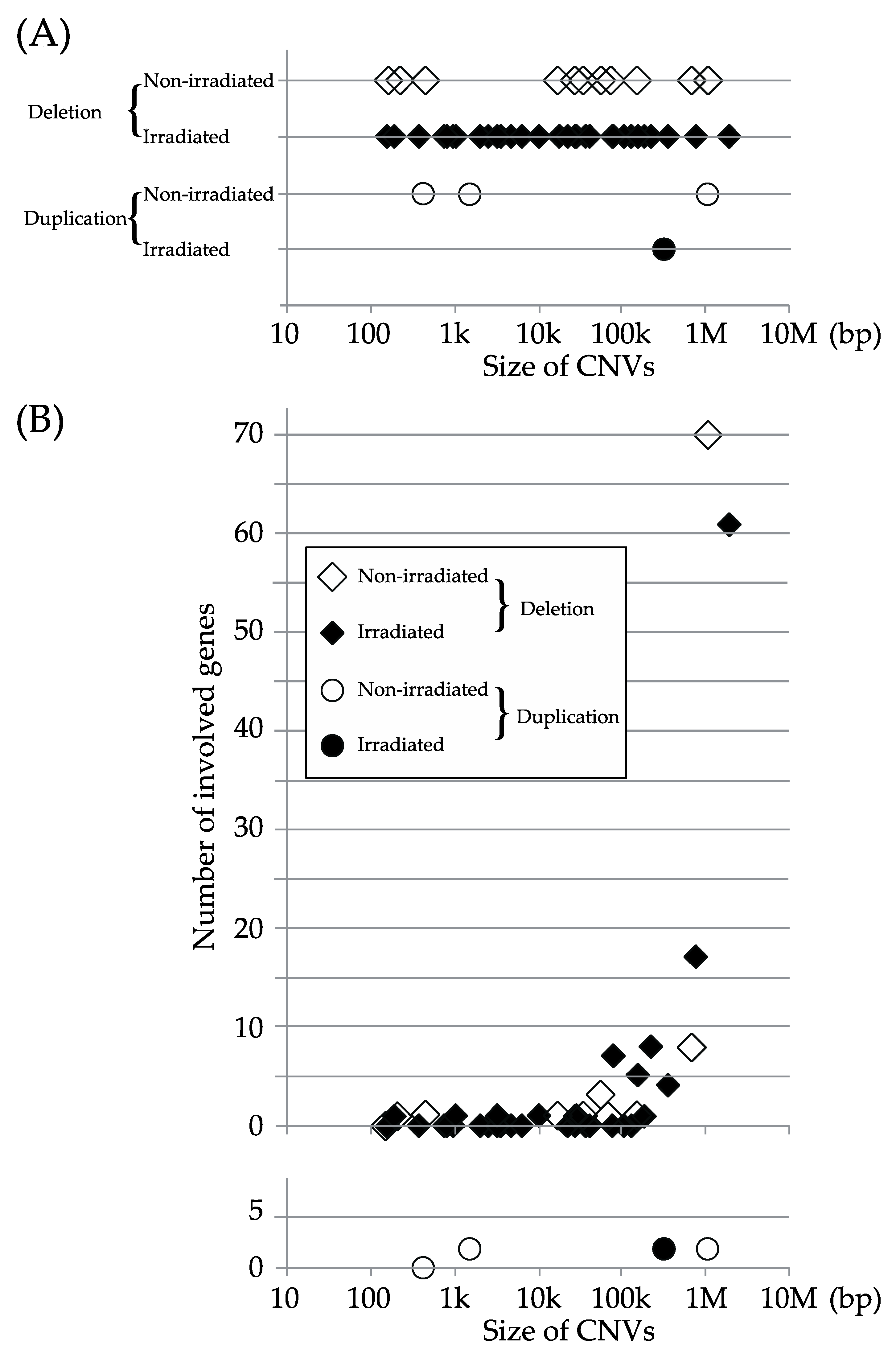
2. Results
3. Discussion
4. Materials and Methods
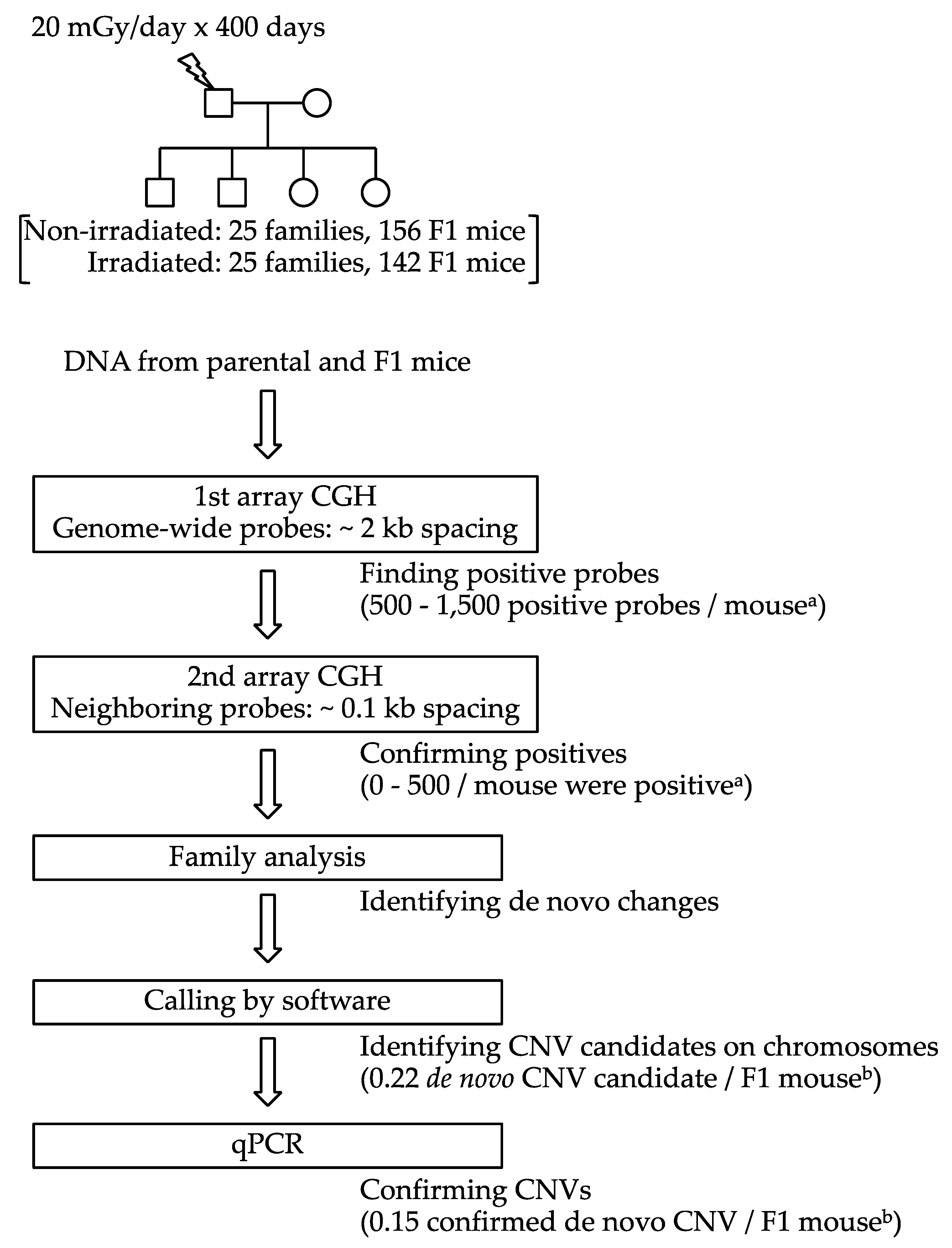
4.1. Animals, Irradiation and DNA Isolation
4.2. Array CGH
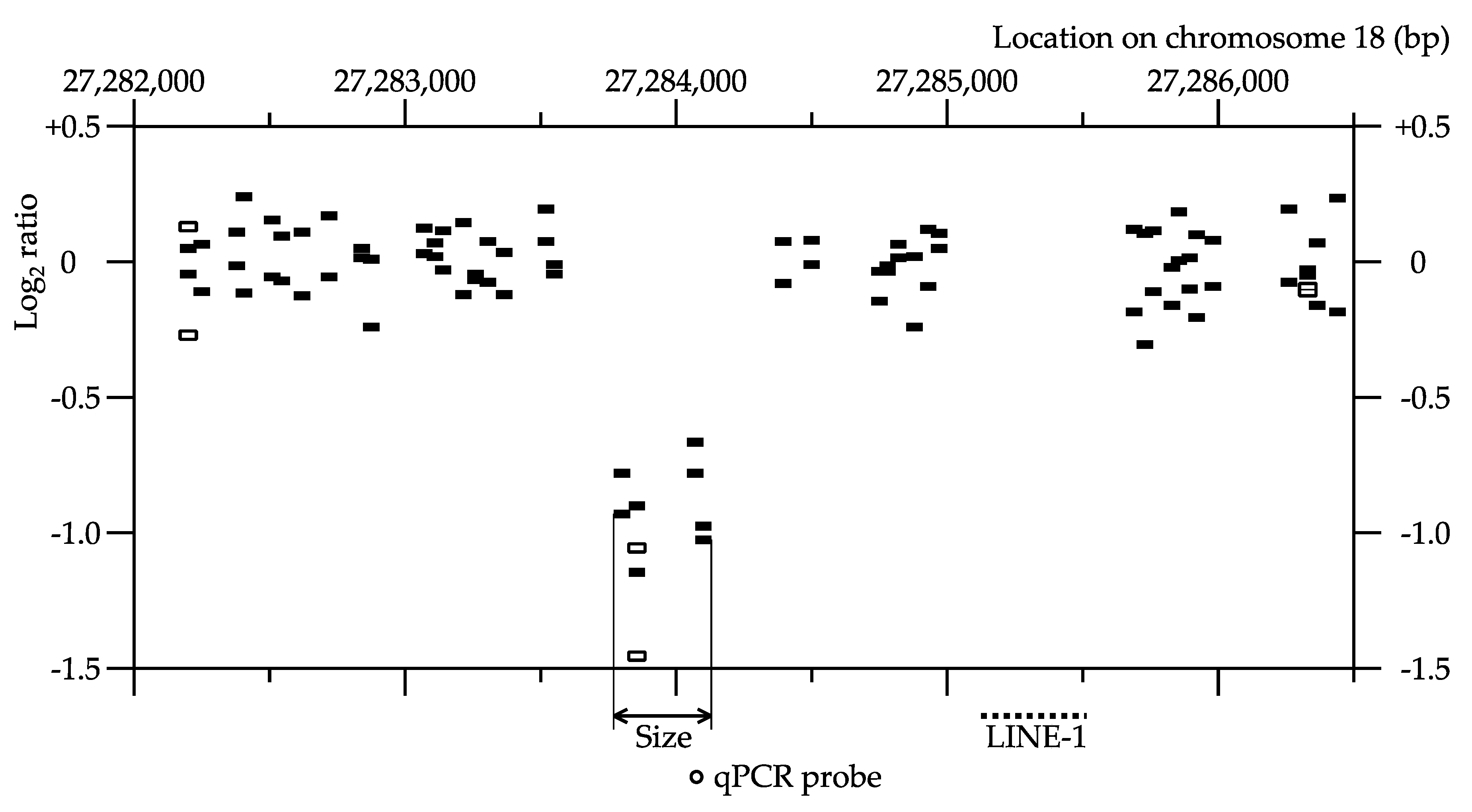
4.3. Quantitative PCR (qPCR)
4.4. Statistics
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Muller, H.J. Artificial transmutation of the gene. Science 1927, 66, 84–87. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, N.; Suyama, A.; Noda, A.; Kodama, Y. Radiation effects on human heredity. Annu. Rev. Genet. 2013, 47, 33–50. [Google Scholar] [CrossRef][Green Version]
- Nakamura, N. Why genetic effects of radiation are observed in mice but not in humans. Radiat. Res. 2018, 189, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, N. History of radiation genetics: Light and darkness. Int. J. Radiat. Biol. 2019, 95, 999–1014. [Google Scholar] [CrossRef] [PubMed]
- United Nations Scientific Committee on the Effects of Atomic Radiation. UNSCEAR 2001 Report to the General Assembly, with Scientific Annex, Hereditary Effects of Radiation; United Nations: New York, NY, USA, 2001. [Google Scholar]
- International Commission on Radiological Protection. ICRP Publication 103, The 2007 Recommendations of the International Commission on Radiological Protection. Ann. ICRP 2007, 37, 2–4. [Google Scholar]
- Russell, W.L.; Kelly, E.M. Mutation frequencies in male mice and the estimation of genetic hazards of radiation in men. Proc. Natl. Acad. Sci. USA 1982, 79, 542–544. [Google Scholar] [CrossRef]
- Lyon, M.F.; Morris, T. Gene and chromosome mutation after large fractionated or unfractionated radiation doses to mouse spermatogonia. Mutat. Res. 1969, 8, 191–198. [Google Scholar] [CrossRef]
- Awa, A.A.; Honda, T.; Neriishi, S.; Sufuni, T.; Shimba, H.; Ohtaki, K.; Nakano, M.; Kodama, Y.; Itoh, M.; Hamilton, H.B. Cytogenetic Study of the offspring of atomic bomb survivors, Hiroshima and Nagasaki. In The Children of Atomic Bomb Survivors: A Genetic Study; Neel, J.V., Schull, W.J., Eds.; National Academies Press: Washington, DC, USA, 1991; pp. 344–362. [Google Scholar]
- Neel, J.V.; Satoh, C.; Goriki, K.; Asakawa, J.; Fujita, M.; Takahashi, N.; Kageoka, T.; Hazama, R. Search for mutations altering protein charge and/or function in children of atomic bomb survivors: Final report. Am. J. Hum. Genet. 1988, 42, 663–676. [Google Scholar]
- Kodaira, M.; Shizue Izumi, S.; Takahashi, N.; Nakamura, N. No evidence of radiation effect on mutation rates at hypervariable minisatellite loci in the germ cells of atomic bomb survivors. Radiat. Res. 2004, 162, 350–356. [Google Scholar] [CrossRef]
- Horai, M.; Mishima, H.; Hayashida, C.; Kinoshita, A.; Nakane, Y.; Matsuo, T.; Tsuruda, K.; Yanagihara, K.; Sato, S.; Imanishi, D.; et al. Detection of de novo single nucleotide variants in offspring of atomic-bomb survivors close to the hypocenter by whole-genome sequencing. J. Hum. Genet. 2018, 63, 357–363. [Google Scholar] [CrossRef]
- Winther, J.F.; Olsen, J.H.; Wu, H.; Shyr, Y.; Mulvihill, J.J.; Stovall, M.; Nielsen, A.; Schmiegelow, M.; Boice, J.B.J. Genetic disease in the children of Danish survivors of childhood and adolescent cancer. J. Clin. Oncol. 2012, 30, 27–33. [Google Scholar] [CrossRef]
- Holtgrewe, M.; Knaus, A.; Hildebrand, G.; Pantel, J.T.; de los Santos, M.R.; Neveling, K.; Goldmann, J.; Schubach, M.; Jäger, M.; Coutelier, M.; et al. Multisite de novo mutations in human offspring after paternal exposure to ionizing radiation. Sci. Rep. 2018, 8, 14611. [Google Scholar] [CrossRef]
- Dubrova, Y.E.; Nesterov, V.N.; Krouchinsky, N.G.; Valdislav, A.; Ostapenko, V.A.; Neumann, R.; Neil, D.L.; Jeffreys, A.J. Human minisatellite mutation rate after the Chernobyl accident. Nature 1996, 380, 683–686. [Google Scholar] [CrossRef] [PubMed]
- Yeager, M.; Machiela, M.J.; Kothiyal, P.; Dean, M.; Bodelon, C.; Suman, S.; Wang, M.; Mirabello, L.; Nelson, C.W.; Zhou, W.; et al. Lack of transgenerational effects of ionizing radiation exposure from the Chernobyl accident. Science 2021, 372, 725–729. [Google Scholar] [CrossRef] [PubMed]
- Costa, E.O.A.; Pinto, I.P.; Gonçalves, M.W.; da Silva, J.F.; Oliveira, L.G.; da Cruz, A.S.; Silva, D.M.E.; da Silva, C.C.; Pereira, R.W.; da Cruz, A.D. Small de novo CNVs as biomarkers of parental exposure to low doses of ionizing radiation of caesium-137. Sci. Rep. 2018, 8, 5914. [Google Scholar] [CrossRef] [PubMed]
- Leite Filho, H.P.; Pinto, I.P.; Oliveira, L.G.; Costa, E.O.A.; da Cruz, A.S.; Silva, D.M.; da Silva, C.C.; Caetano, A.R.; da Cruz, A.D. Deviation from Mendelian transmission of autosomal SNPs can be used to estimate germline mutations in humans exposed to ionizing radiation. PLoS ONE 2020, 15, e0233941. [Google Scholar] [CrossRef]
- Adewoye, A.B.; Lindsay, S.J.; Dubrova, Y.E.; Matthew, E.; Hurles, M.E. The genome-wide effects of ionizing radiation on mutation induction in the mammalian germline. Nat. Commun. 2015, 6, 6684. [Google Scholar] [CrossRef]
- Satoh, Y.; Asakawa, J.; Nishimura, M.; Kuo, T.; Shinkai, N.; Cullings, H.M.; Minakuchi, Y.; Sese, J.; Toyoda, A.; Shimada, Y.; et al. Characteristics of induced mutations in offspring derived from irradiated mouse spermatogonia and mature oocytes. Sci. Rep. 2020, 10, 37. [Google Scholar] [CrossRef]
- Asakawa, J.; Kodaira, M.; Miura, A.; Tsuji, T.; Nakamoto, Y.; Imanaka, M.; Kitamura, J.; Cullings, H.; Nishimura, M.; Shimada, Y.; et al. Genome-wide deletion screening with the array CGH method in mouse offspring derived from irradiated spermatogonia indicates that mutagenic responses are highly variable among genes. Radiat. Res. 2016, 186, 568–576. [Google Scholar] [CrossRef]
- Kodaira, M.; Asakawa, J.; Nakamura, N. Radiation-induced deletions in mouse spermatogonia are usually large (over 200 kb) and contain little sequence similarity at the junctions. Radiat. Res. 2017, 187, 722–731. [Google Scholar] [CrossRef]
- Strobel, M.C.; Seperack, P.K.; Copeland, N.G.; Jenkins, N.A. Molecular analysis of two mouse dilute locus deletion mutations: Spontaneous dilute lethal20J and radiation-induced dilute prenatal lethal Aa2 alleles. Mol. Cell. Biol. 1990, 10, 501–509. [Google Scholar]
- Russell, L.B.; Hunsicker, P.R. The effect of dose rate on the frequency of specific-locus mutations induced in mouse spermatogonia is restricted to larger lesions; a retrospective analysis of historical data. Radiat. Res. 2012, 177, 555–564. [Google Scholar] [CrossRef] [PubMed]
- Sankaranarayanan, K.; Wassom, J.S. Ionizing radiation and genetic risks XIV. Potential research directions in the post-genome era based on knowledge of repair of radiation-induced DNA double-strand breaks in mammalian somatic cells and the origin of deletions associated with human genomic disorders. Mutat. Res. 2005, 578, 333–370. [Google Scholar]
- Collis, S.J.; Schwaninger, J.M.; Ntambi, A.J.; Keller, T.W.; Nelson, W.G.; Dillehay, L.E.; DeWeese, T.L. Evasion of early cellular response mechanisms following low level radiation-induced DNA damage. J. Biol. Chem. 2004, 279, 49624–49632. [Google Scholar] [CrossRef]
- Grudzenski, S.; Raths, A.; Conrad, S.; Rübe, C.E.; Löbrich, M. Inducible response required for repair of low-dose radiation damage in human fibroblasts. Proc. Natl. Acad. Sci. USA 2010, 107, 14205–14210. [Google Scholar] [CrossRef]
- Lieber, M.R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 2010, 79, 181–211. [Google Scholar] [CrossRef]
- Duncan, J.R.; Lieber, M.R.; Adachi, N.; Wahl, R.L. Radiation dose does matter: Mechanistic insights into DNA damage and repair support the linear no-threshold model of low-dose radiation health risks. J. Nucl. Med. 2018, 59, 1014–1016. [Google Scholar] [CrossRef]
- Dubrova, Y.E. Radiation-induced transgenerational instability. Oncogene 2003, 22, 7087–7093. [Google Scholar] [CrossRef]
- Niwa, O. Radiation Induced Dynamic Mutations and Transgenerational Effects. J. Radiat. Res. 2006, 47 (Suppl. B), 25–30. [Google Scholar] [CrossRef]
- Vilenchik, M.M.; Knudson, A.G.J. Inverse radiation dose-rate effects on somatic and germ-line mutations and DNA damage rates. Proc. Natl. Acad. Sci. USA 2000, 97, 5381–5386. [Google Scholar] [CrossRef]
- Ogura, K.; Magae, J.; Kawakami, Y.; Koana, T. Reduction in mutation frequency by very low-dose gamma irradiation of Drosophila melanogaster germ cells. Radiat. Res. 2009, 171, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, I.B., III; Tanaka, S.; Kohda, A.; Takai, D.; Nakamura, S.; Ono, T.; Tanaka, K.; Komura, J. Experimental studies on the biological effects of chronic low dose-rate radiation exposure in mice: Overview of the studies at the Institute for Environmental Sciences. Int. J. Radiat. Biol. 2018, 94, 423–433. [Google Scholar] [CrossRef] [PubMed]
- The R Project for Statistical Computing. Available online: https://www.R-project.org/ (accessed on 18 June 2021).
| F1 Mouse | No. of Positive Probes | Size (bp) | Chromosomal Location (GRCm38) | qPCR Probe Location (GRCm38) | Involved Genes | ||||
|---|---|---|---|---|---|---|---|---|---|
| ID | Sex | 1st Array CGH | 2nd Array CGH | ||||||
| Non-Irradiated | Deletion | 0mGyX3 | M | 11 | >50 | 1,103,254 | Chr13:66,699,838–67,803,092 | 66,700,397 | Gm40988, Gm48404, Gm40989, Gm7896, Gm48413, Gm40989, Vmn2r-ps108, Gm48412, Gm48414, Gm10323, Vmn2r-ps109, Cbx3-ps4, Gm5451, Gm7911, Uqcrb, Gm10767, Mterf3, Ptdss1, 4933433G19Rik, Zfp712, Gm48705, Gm46440, Zfp708, Gm28557, Gm17938, Rslcan18, Zfp759, Gm48732, Gm48733, Rsl1, Gm49646, Gm9626, Zfp455, Gm49064, Zfp458, F630042J09Rik, Zfp457, Gm48824, Zfp595, Gm28044, Zfp593, Zfp456, Gm17039, Zfp953, Gm28041, Zfp429, Gm48900, Zfp459, Zfp874a, Gm38307, Gm7928, Zfp874b, Zfp58, Gm26965, Zfp87, Zfp748, 9430065F17Rik, Zfp729b, Gm49345, Zfp729a, Zfp738, Gm48095, Gm26875, Zfp65, Gm48894, Zfp85, Gm9894, Zfp493, 4930525G20Rik |
| 0mGyH5 a | M | 7 | >50 | 666,200 | Chr2:177,329,278–177,995,478 | 178,041,392 | Gm14414, Zfp970, Gm14403, Gm14324, Gm14322, Gm14326, Gm14327, Rps8-ps5 | ||
| 0mGyH5 a | M | 22 | >50 | 146,136 | Chr12:40,469,827–40,615,963 | 40,596,701 | Dock4 | ||
| 0mGyG2 | M | 19 | >50 | 74,519 | Chr1:112,845,241–112,919,760 | 112,877,741 | Gm8204 | ||
| 0mGyS6 | F | 14 | >50 | 56,272 | Chr7:102,192,468–102,248,740 | 102,193,027 | Nup98, Pgap2, Rhog | ||
| 0mGyK4 | F | 9 | >50 | 34,160 | Chr3:5,533,449–5,567,609 | 5,560,033 | Pex2 | ||
| 0mGyK5 | F | 10 | >50 | 25,412 | Chr13:54,919,455–54,944,867 | 54,942,291 | 4930526F13Rik | ||
| 0mGyJ3 | F | 6 | >50 | 17,346 | Chr4:128,415,985–128,433,331 | 128,416,544 | Csmd2 | ||
| 0mGyV2 | M | 1 | 2 | 461 | Chr2:126,675,275–126,675,736 | 126,675,329 | Gabpb1 | ||
| 0mGyA3 | F | 1 | 2 | 212 | Chr4:34,844,952–34,845,164 | 34,845,005 | Zfp292 | ||
| 0mGyO7 | F | 1 | 2 | 153 | Chr1:7,323,388–7,323,541 | 7,323,321 | |||
| Duplication | 0mGyZ1 | M | 158 | >50 | 1,100,262 | Chr6:79,988,064–81,088,326 | 79,988,623 | Lrrtm4, Gm43900 | |
| 0mGyE4 | F | 1 | 13 | 1535 | Chr8:123,427,701–123,429,236 | 123,429,010 | Galnt2, Def8 | ||
| 0mGyG5 | F | 1 | 3 | 412 | Chr9:49,721,288–49,721,700 | 49,721,288 | Ncam1 | ||
| Irradiated | Deletion | 20mGyL5 | M | 575 | >50 | 1,908,155 | Chr13:63,399,812–65,307,967 | 65,307,023 | Fancc, patch1, A930032L01 Rik, Gm30655, Gm30709, 1700024I08 Rik, Gm47387, Gm47390, Gm47417, Gm47418, Ercc6l2, Gm7695, Hsd17b3, Slc35d2, Zfp367, Gm47513, Habp4, Cdc14b, Gm46424, 1810034E14 Rik, Gm47004,Gm47003, Gm25654, Prxl2c, Zfp182, Gm47123, Gm49230, Ctsl, 1700015C15 Rik, Cdk20, Gm31218, Fam240b, Gm7712, Gm47190, Gm4810, Gm47191, Gm47193, Gm5791, Gm4935, Gm47194, Gm7065, Cntnap3, Spata31, Eif1-ps2, Prss47, Mfsd14b, Spata31d1c, Gm6888, Gm3785, Olfr465-ps1, Olfr466, Gm24130, Nlrp4f, Gm47249, Gm36445, Gm47250, Gm47251, Gm47254, Gm10775, Zfp369, Gm47258 |
| 20mGyI5 | F | 200 | >50 | 742,233 | Chr10:99,214,798–99,957,031 | 99,959,613 | Gm34777, Gm34574, Gm48884, Dusp6, Gm48089, B530045E10 Rik, Gm34921, Gad1, Gm34983, Gm35035, Gm48085, Gm20110, Gm18409, Gm35101, Gm47578, Csl, Gm47579 | ||
| 20mGyL8 | F | 145 | >50 | 353,159 | Chr3:105,510,798–105,863,957 | 105,511,357 | AK018929, Kcnd3 | ||
| 20mGyE1 | M | 83 | >50 | 235,104 | Chr9:57,625,823–57,860,927 | 57,854,366 | Csk, Cyp1a2, Cyp1a1, Edc3, Clk3, Gm17231, Arid3b | ||
| 20mGyL2 | M | 36 | >50 | 145,401 | Chr14:37,914,599–38,060,000 | 37,936,448 | Gm47974 | ||
| 20mGyA1 | M | 12 | >50 | 133,572 | Chr9:114,085,760–114,219,332 | 114,213,599 | |||
| 20mGyAA4 a | F | 34 | >50 | 102,921 | Chr1:117,376,753–117,479,674 | 117,377,055 | |||
| 20mGyU6 | F | 35 | >50 | 76,287 | Chr1:172,286,528–172,362,815 | 172,287,087 | Igsf8, Atp1a2, Kcnj9, Gm36937, Kcnj10 | ||
| 20mGyV6 | F | 29 | >50 | 75,893 | Chr2:45,606,923–45,682,816 | 45,607,482 | |||
| 20mGyW1 | M | 14 | >50 | 40,175 | Chr19:54,228,002–54,268,177 | 54,228,561 | |||
| 20mGyL4 | M | 8 | >50 | 32,881 | Chr6:138,709,973–138,742,854 | 138,741,398 | |||
| 20mGyU8 | F | 7 | >50 | 29,202 | Chr9:100,793,927–100,823,129 | 100,794,486 | Stag1 | ||
| 20mGyX1 | M | 12 | >50 | 28,507 | Chr12:92,023,170–92,051,677 | 92,047,867 | |||
| 20mGyV1 | M | 8 | >50 | 22,299 | Chr11:111,762,690–111,784,989 | 111,763,249 | |||
| 20mGyN4 a | M | 6 | 32 | 10,224 | Chr6:34,932,670–34,942,894 | 34,941,090 | Stra8 | ||
| 20mGyV4 | F | 4 | >50 | 9424 | Chr10:107,833,661–107843085 | 107,834,220 | Otogl | ||
| 20mGyX2 | F | 3 | 40 | 5965 | Chr14:112,914,840–112,920,805 | 112,919,755 | |||
| 20mGyAA4a | F | 2 | 33 | 4495 | Chr17:81,065,337–81,069,832 | 81,069,694 | |||
| 20mGyU5 | M | 2 | 23 | 3469 | Chr11:92,816,645–92,820,114 | 92,817,204 | |||
| 20mGyY1 a | M | 2 | 16 | 3370 | Chr11:105,950,869–105,954,239 | 105,951,424 | |||
| 20mGyG2 | M | 2 | 16 | 3286 | Chr14:40,993,918–40,997,204 | 40,996,166 | Prxl2a | ||
| 20mGyN4 a | M | 1 | 19 | 2367 | Chr4:50,541,636–50,544,003 | 50,543,175 | |||
| 20mGyG7 | F | 1 | 17 | 1878 | Chr4:90,232,441–90,234,319 | 90,234,260 | |||
| 20mGyB5 | F | 1 | 3 | 1040 | Chr10:69,420,081–69,421,121 | 69,420,140 | Ank3-246 | ||
| 20mGyG5 | F | 1 | 9 | 948 | Chr1:44,804,456–44,805,404 | 44,804,512 | |||
| 20mGyX7 | F | 1 | 5 | 732 | Chr4:21,163,257–21,163,989 | 21,163,816 | |||
| 20mGyH3 | M | 1 | 8 | 716 | Chr3:119,702,208–119,702,924 | 119,702,263 | |||
| 20mGyY1 a | M | 1 | 4 | 356 | Chr18:27,125,326–27,125,682 | 27,125,885 | |||
| 20mGyX5 | F | 1 | 2 | 191 | Chr12:81,490,882–81,491,073 | 81,491,441 | Synj2bp | ||
| 20mGyB2 | F | 1 | 2 | 156 | Chr17:4,507,025–4,507,181 | 4,507,139 | |||
| Duplication | 20mGyE5 | F | 86 | >50 | 305,969 | Chr6:48,055,971–48,361,940 | 48,360,175 | Zfp746, Gm16630 | |
| No. of Positive Probes in 1st Array CGH | ≥2 (Type L) | 1 (Type S) | (Total) | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No. of Positive Probes in 2nd Array CGH | ≥5 | 4 | 3 | 2 | (Total) | ||||||||||
| Deletion (del) or Duplication (dup) | Del | Dup | Del | Dup | Del | Dup | Del | Dup | Del | Dup | Del | Dup | Del | Dup | |
| Non-irradiated (n = 156) | Detected by array CGH | 8 | 1 | 0 | 1 | 1 | 0 | 0 | 2 | 13 | 4 | 14 | 7 | 22 | 8 |
| Confirmed by qPCR | 8 | 1 | 0 | 1 | 0 | 0 | 0 | 1 | 3 | 0 | 3 | 2 | 11 | 3 | |
| Irradiated (n = 142) | Detected by array CGH | 22 | 1 | 5 | 0 | 2 | 0 | 2 | 0 | 3 | 0 | 12 | 0 | 34 | 1 |
| Confirmed by qPCR | 22 | 1 | 5 | 0 | 1 | 0 | 1 | 0 | 2 | 0 | 9 | 0 | 31 | 1 | |
| No. of Positive Probes in 1st Array CGH | ≥2 (Type L) | 1 (Type S) | (Total) | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No. of Positive Probes in 2nd Array CGH | ≥5 | 4 | 3 | 2 | (Total) | |||||||||||
| Deletion (Del) or Duplication (Dup) | Del | Dup | Del | Dup | Del | Dup | Del | Dup | Del | Dup | Del | Dup | Del | Dup | ||
| Non-irradiated | 0mGyJ4 (F) | Detected by array CGH | 0 | 0 | 0 | 0 | 1 | 0 | 5 | 0 | 13 | 0 | 19 | 0 | 19 | 0 |
| Analyzed by qPCR | 0 | 0 | 0 | 0 | 1 | 0 | 5 | 0 | 1 | 0 | 7 | 0 | 7 | 0 | ||
| Confirmed by qPCR | 0 | 0 | 0 | 0 | 0 | 0 | 3 | 0 | 1 | 0 | 4 | 0 | 4 | 0 | ||
| 0mGyX5 (F) | Detected by array CGH | 0 | 0 | 0 | 0 | 3 | 0 | 9 | 0 | 43 | 0 | 55 | 0 | 55 | 0 | |
| Analyzed by qPCR | 0 | 0 | 0 | 0 | 3 | 0 | 2 | 0 | 5 | 0 | 10 | 0 | 10 | 0 | ||
| Confirmed by qPCR | 0 | 0 | 0 | 0 | 2 | 0 | 2 | 0 | 1 | 0 | 5 | 0 | 5 | 0 | ||
| Irradiated | 20mGyA2 (M) | Detected by array CGH | 0 | 0 | 14 | 0 | 18 | 0 | 27 | 0 | 49 | 0 | 108 | 0 | 108 | 0 |
| Analyzed by qPCR | 0 | 0 | 3 | 0 | 3 | 0 | 5 | 0 | 11 | 0 | 22 | 0 | 22 | 0 | ||
| Confirmed by qPCR | 0 | 0 | 3 | 0 | 1 | 0 | 1 | 0 | 6 | 0 | 11 | 0 | 11 | 0 | ||
| 20mGyF5 (F) | Detected by array CGH | 0 | 0 | 1 | 0 | 2 | 0 | 3 | 0 | 10 | 0 | 16 | 0 | 16 | 0 | |
| Analyzed by qPCR | 0 | 0 | 1 | 0 | 2 | 0 | 0 | 0 | 10 | 0 | 13 | 0 | 13 | 0 | ||
| Confirmed by qPCR | 0 | 0 | 1 | 0 | 2 | 0 | 0 | 0 | 2 | 0 | 5 | 0 | 5 | 0 | ||
| 20mGyL1 (M) | Detected by array CGH | 0 | 0 | 1 | 0 | 0 | 0 | 2 | 0 | 6 | 0 | 9 | 0 | 9 | 0 | |
| Analyzed by qPCR | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 0 | 6 | 0 | 8 | 0 | 8 | 0 | ||
| Confirmed by qPCR | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 3 | 0 | 4 | 0 | 4 | 0 | ||
| No. of Positive Probes in 1st Array CGH | ≥2 (Type L) | 1 (Type S) | (Total) | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Deletion (Del) or Duplication (Dup) | Del | Dup | Del | Dup | Del | Dup | ||||||||||||||
| Sex | n | n | % | pa | n | % | pa | n | % | pa | n | % | pa | n | % | pa | n | % | pa | |
| M | Non-irradiated | 75 | 4 | 5.3 | 0.018 | 1 | 1.3 | >0.1 | 1 | 1.3 | >0.1 | 0 | 0.0 | ND b | 5 | 6.6 | 0.024 | 1 | 1.3 | >0.1 |
| Irradiated | 75 | 13 | 17.3 | 0 | 0.0 | 3 | 4.0 | 0 | 0.0 | 14 c | 18.7 | 0 | 0.0 | |||||||
| F | Non-irradiated | 81 | 4 | 4.9 | >0.1 | 0 | 0.0 | >0.1 | 2 | 2.5 | 0.085 | 2 | 2.4 | >0.1 | 6 | 7.4 | 0.009 | 2 | 2.4 | >0.1 |
| Irradiated | 67 | 9 | 13.4 | 1 | 1.4 | 6 | 9.0 | 0 | 0.0 | 15 | 22.4 | 1 | 1.4 | |||||||
| Total | Non-irradiated | 156 | 8 | 5.1 | 0.004 | 1 | 0.6 | >0.1 | 3 | 1.9 | 0.075 | 2 | 1.2 | >0.1 | 11 | 7.1 | 0.001 | 3 | 1.9 | >0.1 |
| Irradiated | 142 | 22 | 15.5 | 1 | 0.7 | 9 | 6.3 | 0 | 0.0 | 29 | 20.4 | 1 | 0.7 | |||||||
| No. of Deletions | Deletion Size (bp) | |||
|---|---|---|---|---|
| GM a | GSD b | |||
| Non-irradiated | 11 | 18,762 | 12.181 | p = 0.6758 c |
| Irradiated | 31 | 12,674 | 21.484 | |
| Group of F1 Mice | No. of F1 Mice | Life Span (Days) | ||||
|---|---|---|---|---|---|---|
| (Mean ± SD) | ||||||
| M | Non-irradiated | CNV− b | 70 | 914 | ± | 170 |
| CNV+ c | 4 | 859 | ± | 161 | ||
| Irradiated | CNV- | 56 | 824 | ± | 163 | |
| CNV+ | 16 | 881 | ± | 155 | ||
| F | Non-irradiated | CNV- | 72 | 813 | ± | 132 |
| CNV+ | 8 | 772 | ± | 124 | ||
| Irradiated | CNV- | 54 | 833 | ± | 140 | |
| CNV+ | 10 | 738 | ± | 125 | ||
| Factor | Hazard Ratio | Z Value | p Value |
|---|---|---|---|
| Sex a | 0.507 | −5.40 | <0.0001 |
| Irradiation a | 0.851 | −1.32 | 0.187 |
| CNVs a | 0.697 | −2.00 | 0.045 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ogura, K.; Ayabe, Y.; Harada, C.; Tanaka, I.B., III; Tanaka, S.; Komura, J.-i. Increased Frequency of Copy Number Variations Revealed by Array Comparative Genomic Hybridization in the Offspring of Male Mice Exposed to Low Dose-Rate Ionizing Radiation. Int. J. Mol. Sci. 2021, 22, 12437. https://doi.org/10.3390/ijms222212437
Ogura K, Ayabe Y, Harada C, Tanaka IB III, Tanaka S, Komura J-i. Increased Frequency of Copy Number Variations Revealed by Array Comparative Genomic Hybridization in the Offspring of Male Mice Exposed to Low Dose-Rate Ionizing Radiation. International Journal of Molecular Sciences. 2021; 22(22):12437. https://doi.org/10.3390/ijms222212437
Chicago/Turabian StyleOgura, Keiji, Yoshiko Ayabe, Chihiro Harada, Ignacia Braga Tanaka, III, Satoshi Tanaka, and Jun-ichiro Komura. 2021. "Increased Frequency of Copy Number Variations Revealed by Array Comparative Genomic Hybridization in the Offspring of Male Mice Exposed to Low Dose-Rate Ionizing Radiation" International Journal of Molecular Sciences 22, no. 22: 12437. https://doi.org/10.3390/ijms222212437
APA StyleOgura, K., Ayabe, Y., Harada, C., Tanaka, I. B., III, Tanaka, S., & Komura, J.-i. (2021). Increased Frequency of Copy Number Variations Revealed by Array Comparative Genomic Hybridization in the Offspring of Male Mice Exposed to Low Dose-Rate Ionizing Radiation. International Journal of Molecular Sciences, 22(22), 12437. https://doi.org/10.3390/ijms222212437





