Monoclonal Antibodies against Nucleocapsid Protein of SARS-CoV-2 Variants for Detection of COVID-19
Abstract
:1. Introduction
2. Results
2.1. Generation and Identification of mAbs against SARS-CoV-2 NP
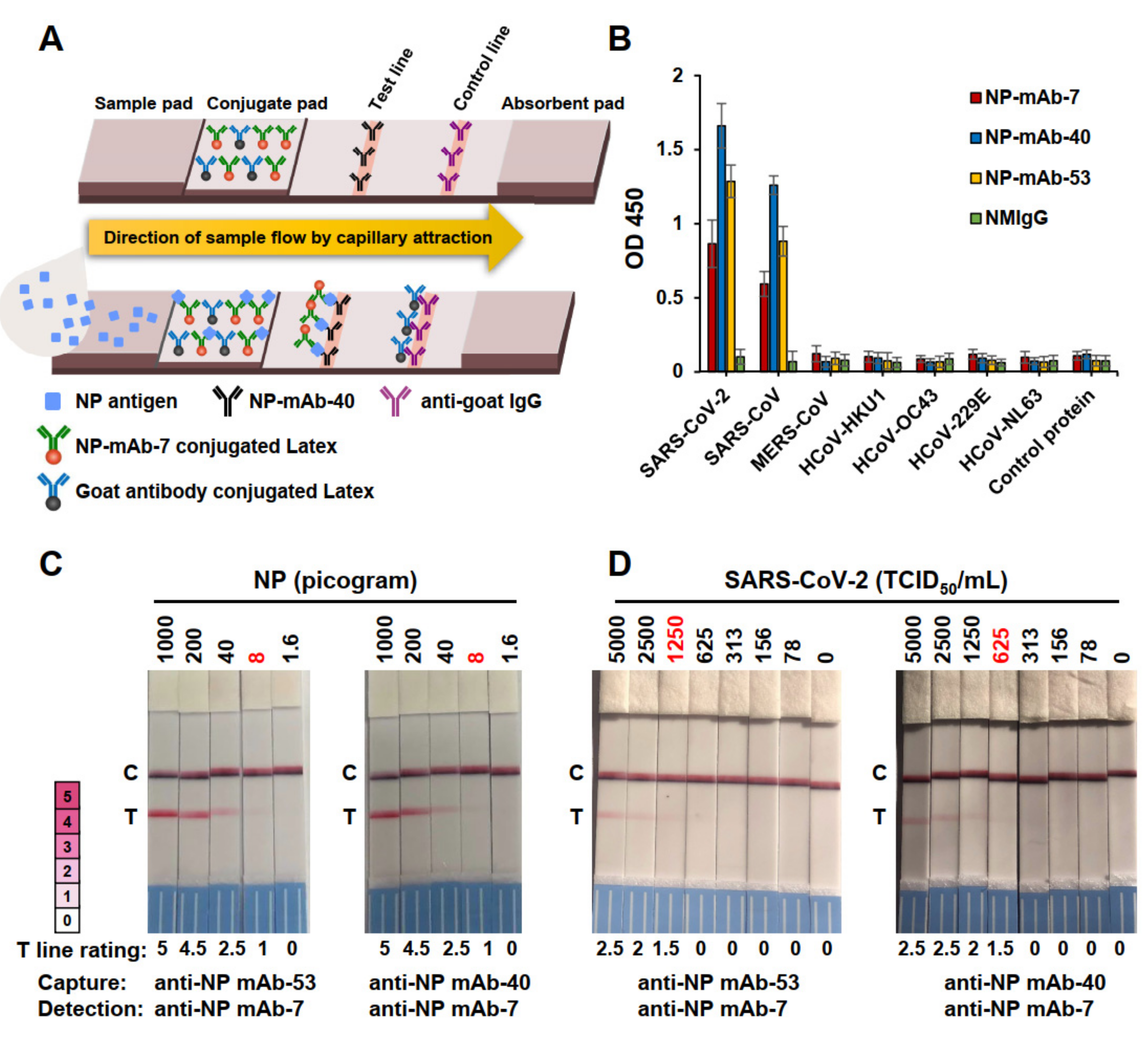
2.2. Screening of Antibody Pairs in Lateral Flow Immunoassays (LFIAs) to Detect NP Antigen
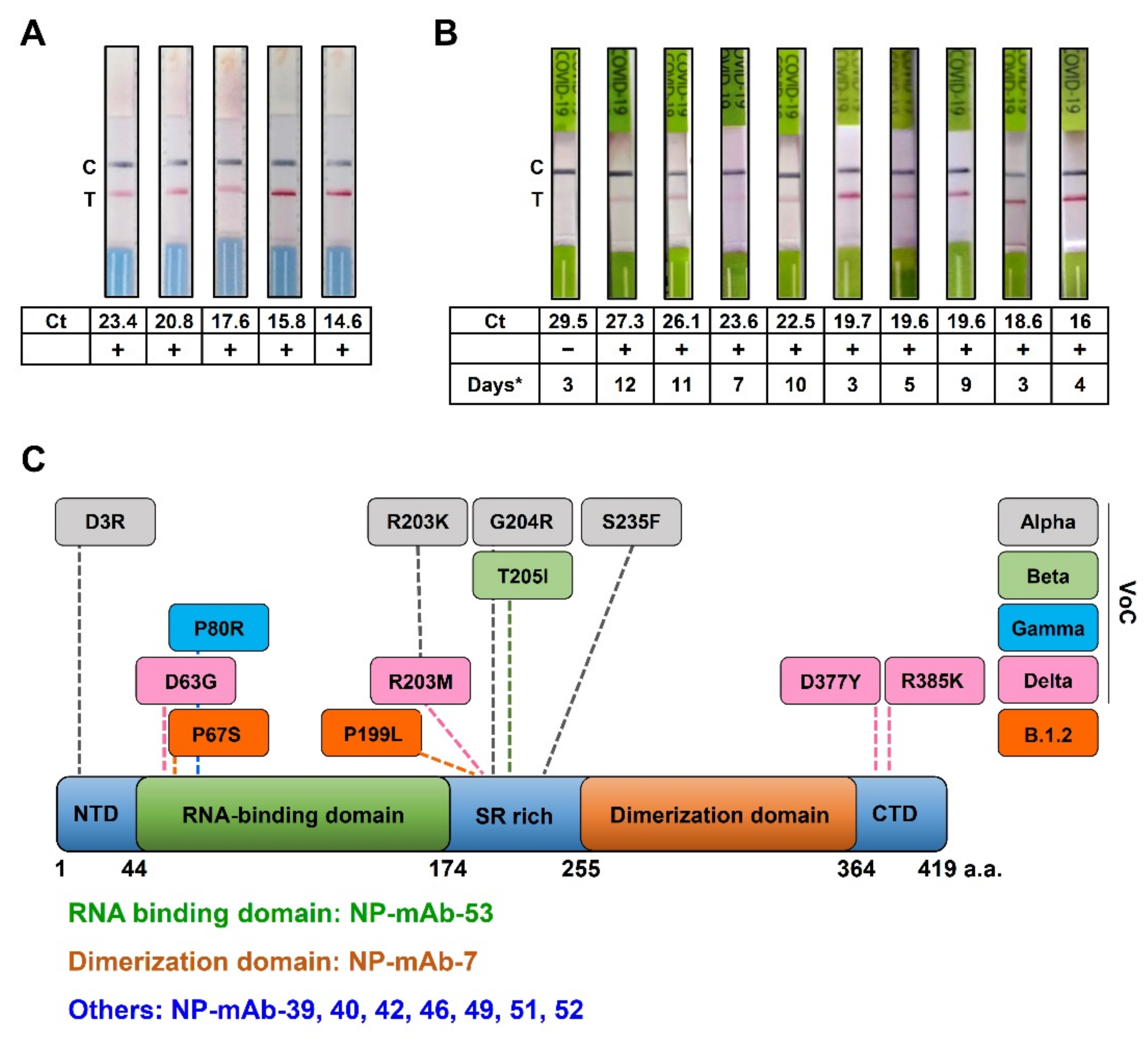
2.3. Performance of NP-mAb-40/7 LFIA Strips for NP of SARS-CoV-2 Variants
2.4. Specificity of NP-mAb-40/7 LFIA Strips
2.5. Clinical Studies of NP-mAb-40/7 LFIA Strips
3. Discussion
4. Materials and Methods
4.1. Recombinant Proteins and Reagents
4.2. Generation and Purification of mAbs
4.3. Screening of mAbs against NP by ELISA
4.4. Western Blot Analysis
4.5. Lateral Flow Immunoassay (LFIA) Strip Preparation
4.6. Immunofluorescence Assay (IFA)
4.7. Detection of NP from SARS-CoV-2-Infected Cells by NP-mAb LFIA
4.8. LFIA limit of Detection (LOD)
4.9. Cross-Reactivity Assays of Human Respiratory-Associated Pathogens
4.10. Cross-Reactivity and Interference Assays
4.11. Accelerated Stability Testing
4.12. Clinical Studies
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gupta, A.; Madhavan, M.V.; Sehgal, K.; Nair, N.; Mahajan, S.; Sehrawat, T.S.; Bikdeli, B.; Ahluwalia, N.; Ausiello, J.C.; Wan, E.Y.; et al. Extrapulmonary manifestations of COVID-19. Nat. Med. 2020, 26, 1017–1032. [Google Scholar] [CrossRef]
- Letko, M.; Marzi, A.; Munster, V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat. Microb. 2020, 5, 562–569. [Google Scholar] [CrossRef] [Green Version]
- Auwaerter, P.M.D.; Coronavirus COVID-19 (SARS-CoV-2). Johns Hopkins ABX Guide. The Johns Hopkins University. Available online: https://www.hopkinsguides.com/hopkins/view/Johns_Hopkins_ABX_Guide/540747/all/Coronavirus_COVID_19__SARS_CoV_2_ (accessed on 11 November 2021).
- Andreadakis, Z.; Kumar, A.; Román, R.G.; Tollefsen, S.; Saville, M.; Mayhew, S. The COVID-19 vaccine development landscape. Nat. Rev. Drug Discov. 2020, 19, 305–306. [Google Scholar]
- Taylor, P.C.; Adams, A.C.; Hufford, M.M.; de la Torre, I.; Winthrop, K.; Gottlieb, R.L. Neutralizing monoclonal antibodies for treatment of COVID-19. Nat. Rev. Immunol. 2021, 21, 382–393. [Google Scholar] [CrossRef]
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.-M.; Wang, W.; Song, Z.-G.; Hu, Y.; Tao, Z.-W.; Tian, J.-H.; Pei, Y.-Y. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cubuk, J.; Alston, J.J.; Incicco, J.J.; Singh, S.; Stuchell-Brereton, M.D.; Ward, M.D.; Zimmerman, M.I.; Vithani, N.; Griffith, D.; Wagoner, J.A.; et al. The SARS-CoV-2 nucleocapsid protein is dynamic, disordered, and phase separates with RNA. Nat. Commun. 2021, 12, 1936. [Google Scholar] [CrossRef] [PubMed]
- Zeng, W.; Liu, G.; Ma, H.; Zhao, D.; Yang, Y.; Liu, M.; Mohammed, A.; Zhao, C.; Yang, Y.; Xie, J.; et al. Biochemical characterization of SARS-CoV-2 nucleocapsid protein. Biochem. Biophys. Res. Commun. 2020, 527, 618–623. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Zhu, F.; Zhao, M.; Shao, F.; Yu, D.; Ma, J.; Zhang, X.; Li, W.; Qian, Y.; Zhang, Y.; et al. SARS-CoV-2 nucleocapsid suppresses host pyroptosis by blocking Gasdermin D cleavage. EMBO J. 2021, 40, e108249. [Google Scholar] [CrossRef]
- Kang, S.; Yang, M.; He, S.; Wang, Y.; Chen, X.; Chen, Y.-Q.; Hong, Z.; Liu, J.; Jiang, G.; Chen, Q.; et al. A SARS-CoV-2 antibody curbs viral nucleocapsid protein-induced complement hyperactivation. Nat. Commun. 2021, 12, 2697. [Google Scholar] [CrossRef]
- Shan, D.; Johnson, J.M.; Fernandes, S.C.; Suib, H.; Hwang, S.; Wuelfing, D.; Mendes, M.; Holdridge, M.; Burke, E.M.; Beauregard, K.; et al. N-protein presents early in blood, dried blood and saliva during asymptomatic and symptomatic SARS-CoV-2 infection. Nat. Commun. 2021, 12, 1931. [Google Scholar] [CrossRef]
- Gutmann, C.; Takov, K.; Burnap, S.A.; Singh, B.; Ali, H.; Theofilatos, K.; Reed, E.; Hasman, M.; Nabeebaccus, A.; Fish, M.; et al. SARS-CoV-2 RNAemia and proteomic trajectories inform prognostication in COVID-19 patients admitted to intensive care. Nat. Commun. 2021, 12, 3406. [Google Scholar] [CrossRef] [PubMed]
- Djukic, T.; Mladenovic, M.; Stanic-Vucinic, D.; Radosavljevic, J.; Smiljanic, K.; Sabljic, L.; Devic, M.; Cujic, D.; Vasovic, T.; Simovic, A.; et al. Expression, purification and immunological characterization of recombinant nucleocapsid protein fragment from SARS-CoV-2. Virology 2021, 557, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Grzelak, L.; Temmam, S.; Planchais, C.; Demeret, C.; Tondeur, L.; Huon, C.; Guivel-Benhassine, F.; Staropoli, I.; Chazal, M.; Dufloo, J.; et al. A comparison of four serological assays for detecting anti–SARS-CoV-2 antibodies in human serum samples from different populations. Sci. Translat. Med. 2020, 12, eabc3103. [Google Scholar] [CrossRef] [PubMed]
- Weissleder, R.; Lee, H.; Ko, J.; Pittet, M.J. COVID-19 diagnostics in context. Sci. Translat. Med. 2020, 12, eabc1931. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, R.; He, Q.; Pascual, M. Quantifying asymptomatic infection and transmission of COVID-19 in New York City using observed cases, serology, and testing capacity. Proc. Natl. Acad. Sci. USA 2021, 118, 2019716118. [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.A.; Quandelacy, T.M.; Kada, S.; Prasad, P.V.; Steele, M.; Brooks, J.T.; Slayton, R.B.; Biggerstaff, M.; Butler, J.C. SARS-CoV-2 Transmission from People Without COVID-19 Symptoms. JAMA Netw. Open 2021, 4, e2035057. [Google Scholar] [CrossRef]
- Afzal, A. Molecular diagnostic technologies for COVID-19: Limitations and challenges. J. Adv. Res. 2020, 26, 149–159. [Google Scholar] [CrossRef]
- Carter, L.J.; Garner, L.V.; Smoot, J.W.; Li, Y.; Zhou, Q.; Saveson, C.J.; Sasso, J.M.; Gregg, A.C.; Soares, D.J.; Beskid, T.R.; et al. Assay Techniques and Test Development for COVID-19 Diagnosis. ACS Cent. Sci. 2020, 6, 591–605. [Google Scholar] [CrossRef]
- U.S. Food & Drug Administration. SARS-CoV-2 Viral Mutations: Impact on COVID-19 Tests. Available online: https://www.fda.gov/medical-devices/coronavirus-covid-19-and-medical-devices/sars-cov-2-viral-mutations-impact-covid-19-tests (accessed on 23 September 2021).
- Harvey, W.T.; Carabelli, A.M.; Jackson, B.; Gupta, R.K.; Thomson, E.C.; Harrison, E.M.; Ludden, C.; Reeve, R.; Rambaut, A.; Peacock, S.J.; et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat. Rev. Microbiol. 2021, 19, 409–424. [Google Scholar] [CrossRef]
- Troyano-Hernáez, P.; Reinosa, R.; Holguín, Á. Evolution of SARS-CoV-2 Envelope, Membrane, Nucleocapsid, and Spike Structural Proteins from the Beginning of the Pandemic to September 2020: A Global and Regional Approach by Epidemiological Week. Viruses 2021, 13, 243. [Google Scholar] [CrossRef]
- Chen, A.T.; Altschuler, K.; Zhan, S.H.; Chan, Y.A.; Deverman, B.E. COVID-19 CG enables SARS-CoV-2 mutation and lineage tracking by locations and dates of interest. eLife 2021, 10, e63409. [Google Scholar] [CrossRef]
- Bekliz, M.; Adea, K.; Essaidi-Laziosi, M.; Sacks, J.A.; Escadafal, C.; Kaiser, L.; Eckerle, I. SARS-CoV-2 rapid diagnostic tests for emerging variants. Lancet Microbe 2021, 2, e351. [Google Scholar] [CrossRef]
- Jungnick, S.; Hobmaier, B.; Mautner, L.; Hoyos, M.; Haase, M.; Baiker, A.; Lahne, H.; Eberle, U.; Wimmer, C.; Hepner, S.; et al. Detection of the new SARS-CoV-2 variants of concern B.1.1.7 and B.1.351 in five SARS-CoV-2 rapid antigen tests (RATs), Germany, March 2021. Euro Surveill. 2021, 26, 2100413. [Google Scholar] [CrossRef]
- Mullen, J.L.; Tsueng, G.; Latif, A.A.; Alkuzweny, M.; Cano, M.; Haag, E.; Zhou, J.; Zeller, M.; Hufbauer, E.; Matteson, N.; et al. A Standardized, Open-Source Database of COVID-19 Resources and Epidemiology Data. Available online: https://outbreak.info/ (accessed on 23 September 2020).
- Tao, K.; Tzou, P.L.; Nouhin, J.; Gupta, R.K.; de Oliveira, T.; Pond, S.L.K.; Fera, D.; Shafer, R.W. The biological and clinical significance of emerging SARS-CoV-2 variants. Nat. Rev. Genet. 2021, 22, 757–773. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, V.; de Jesus, R.; Adelino, T.; Reis, A.B.; de Souza, B.B.; Ribeiro, A.A.; Guimarães, N.R.; Livorati, M.T.F.P.; Neto, D.F.D.L.; Kato, R.B.; et al. Genomic evidence of SARS-CoV-2 reinfection case with the emerging B.1.2 variant in Brazil. J. Infect. 2021, 83, 237–279. [Google Scholar] [CrossRef]
- Bajaj, S.; Singla, D.; Sakhuja, N. Stability testing of pharmaceutical products. J. App. Pharmaceut. Sci. 2012, 2, 129–138. [Google Scholar]
- Zou, L.; Ruan, F.; Huang, M.; Liang, L.; Huang, H.; Hong, Z.; Yu, J.; Kang, M.; Song, Y.; Xia, J.; et al. SARS-CoV-2 Viral Load in Upper Respiratory Specimens of Infected Patients. N. Engl. J. Med. 2020, 382, 1177–1179. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.W.; Schmitz, J.E.; Persing, D.H.; Stratton, C.W. Laboratory Diagnosis of COVID-19: Current Issues and Challenges. J. Clin. Microbiol. 2020, 58, e00512-20. [Google Scholar] [CrossRef] [Green Version]
- Ulinici, M.; Covantev, S.; Wingfield-Digby, J.; Beloukas, A.; Mathioudakis, A.G.; Corlateanu, A. Screening, Diagnostic and Prognostic Tests for COVID-19: A Comprehensive Review. Life 2021, 11, 561. [Google Scholar] [CrossRef]
- Igloi, Z.; Velzing, J.; van Beek, J.; van de Vijver, D.; Aron, G.; Ensing, R.; Benschop, K.; Han, W.; Boelsums, T.; Koopmans, M.; et al. Clinical Evaluation of Roche SD Biosensor Rapid Antigen Test for SARS-CoV-2 in Municipal Health Service Testing Site, the Netherlands. Emerg. Infect. Dis. 2021, 27, 1323–1329. [Google Scholar] [CrossRef]
- Teo, A.K.J.; Choudhury, Y.; Tan, I.B.; Cher, C.Y.; Chew, S.H.; Wan, Z.Y.; Cheng, L.T.E.; Oon, L.L.E.; Tan, M.H.; Chan, K.S.; et al. Saliva is more sensitive than nasopharyngeal or nasal swabs for diagnosis of asymptomatic and mild COVID-19 infection. Sci. Rep. 2021, 11, 3134. [Google Scholar] [CrossRef] [PubMed]
- Hayer, J.; Kasapic, D.; Zemmrich, C. Real-world clinical performance of commercial SARS-CoV-2 rapid antigen tests in suspected COVID-19: A systematic meta-analysis of available data as of November 20, 2020. Int. J. Infect. Dis. 2021, 108, 592–602. [Google Scholar] [CrossRef] [PubMed]
- Salvagno, G.L.; Gianfilippi, G.; Bragantini, D.; Henry, B.M.; Lippi, G. Clinical assessment of the Roche SARS-CoV-2 rapid antigen test. Diagnosis 2021, 8, 322–326. [Google Scholar] [CrossRef] [PubMed]
- Okoye, N.C.; Barker, A.P.; Curtis, K.; Orlandi, R.R.; Snavely, E.A.; Wright, C.; Hanson, K.E.; Pearson, L.N. Performance Characteristics of BinaxNOW COVID-19 Antigen Card for Screening Asymptomatic Individuals in a University Setting. J. Clin. Microbiol. 2021, 59, e03282-20. [Google Scholar] [CrossRef]
- Lauring, A.S.; Hodcroft, E.B. Genetic Variants of SARS-CoV-2—What Do They Mean? JAMA 2021, 325, 529–531. [Google Scholar] [CrossRef]
- Abdool Karim, S.S.; de Oliveira, T. New SARS-CoV-2 Variants—Clinical, Public Health, and Vaccine Implications. N. Engl. J. Med. 2021, 384, 1866–1868. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Yang, M.; Hong, Z.; Zhang, L.; Huang, Z.; Chen, X.; He, S.; Zhou, Z.; Zhou, Z.; Chen, Q.; et al. Crystal structure of SARS-CoV-2 nucleocapsid protein RNA binding domain reveals potential unique drug targeting sites. Acta Pharm. Sin. B 2020, 10, 1228–1238. [Google Scholar] [CrossRef]
- Yu, I.M.; Gustafson, C.L.; Diao, J.; Burgner, J.W., 2nd; Li, Z.; Zhang, J.; Chen, J. Recombinant severe acute respiratory syndrome (SARS) coronavirus nucleocapsid protein forms a dimer through its C-terminal domain. J. Biol. Chem. 2005, 280, 23280–23286. [Google Scholar] [CrossRef] [Green Version]
- MacRaild, C.A.; Richards, J.S.; Anders, R.F.; Norton, R.S. Antibody Recognition of Disordered Antigens. Structure 2016, 24, 148–157. [Google Scholar] [CrossRef] [Green Version]
- Su, S.C.; Yang, T.J.; Yu, P.Y.; Liang, K.H.; Chen, W.Y.; Yang, C.W.; Lin, H.T.; Wang, M.J.; Lu, R.M.; Tso, H.C.; et al. Structure-guided antibody cocktail for prevention and treatment of COVID-19. PLoS Pathog. 2021, 17, e1009704. [Google Scholar] [CrossRef]
- Lu, R.M.; Hwang, Y.C.; Liu, I.J.; Lee, C.C.; Tsai, H.Z.; Li, H.J.; Wu, H.C. Development of therapeutic antibodies for the treatment of diseases. J. Biomed. Sci. 2020, 27, 1. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.M.; Chiu, C.Y.; Liu, I.J.; Chang, Y.L.; Liu, Y.J.; Wu, H.C. Novel human Ab against vascular endothelial growth factor receptor 2 shows therapeutic potential for leukemia and prostate cancer. Cancer Sci. 2019, 110, 3773–3787. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pierson-Perry, J.; Altaie, S.; Danielson, S.; Jorgensen, B.; Poetsch, B.; Savol, R.; Vaks, J.; Budd, J.; De Vore, K.; Magari, R. EP25-A Evaluation of Stability of In Vitro Diagnostic Reagents; Approved Guideline; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2009; Volume 29, Chapter 7; pp. 15–17. [Google Scholar]
- Garrett, P.E.; Lasky, F.D.; Meier, K.L. EP12-A2 User Protocol for Evaluation of Qualitative Test Performance; Approved Guideline; Clinical and Laboratory Standard Institute: Wayne, PA, USA, 2008; Volume 28, Chapter 10; pp. 17–25. [Google Scholar]


| mAbs | Isotype | ELISA* | WB | IFA | Cross-Reactivity with NPs of Human CoV Strains | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| SARS-CoV | MERS-CoV | HCoV-HKU1 | HCoV-OC43 | HCoV-229E | HCoV-NL63 | |||||
| NP-mAb-1 | n.d. | + | + | n.d. | + | − | − | − | − | − |
| NP-mAb-2 | n.d. | + | − | n.d. | + | − | − | − | − | − |
| NP-mAb-3 | n.d. | ++ | − | n.d. | + | − | − | − | − | − |
| NP-mAb-4 | n.d. | + | + | n.d. | + | − | − | − | − | − |
| NP-mAb-5 | n.d. | ++ | + | n.d. | − | − | − | − | − | − |
| NP-mAb-6 | n.d. | + | − | n.d. | + | − | − | − | − | − |
| NP-mAb-7 | IgG1, κ | ++ | + | + | + | − | − | − | − | − |
| NP-mAb-8 | n.d. | + | + | n.d. | + | − | − | − | − | − |
| NP-mAb-9 | n.d. | + | + | n.d. | − | − | − | − | − | − |
| NP-mAb-11 | n.d. | + | − | n.d. | + | − | − | − | − | − |
| NP-mAb-12 | n.d. | + | + | n.d. | − | − | − | − | − | − |
| NP-mAb-13 | n.d. | + | − | n.d. | + | − | − | − | − | − |
| NP-mAb-14 | n.d. | + | − | n.d. | − | − | − | − | − | − |
| NP-mAb-15 | n.d. | + | − | n.d. | − | − | − | − | − | − |
| NP-mAb-18 | n.d. | ++ | − | n.d. | + | − | − | − | − | − |
| NP-mAb-19 | n.d. | + | − | n.d. | n.d. | n.d. | n.d. | n.d. | n.d. | n.d. |
| NP-mAb-21 | n.d. | + | + | n.d. | − | − | − | − | − | − |
| NP-mAb-22 | n.d. | ++ | − | n.d. | + | − | − | − | − | − |
| NP-mAb-23 | n.d. | + | + | n.d. | + | − | − | − | − | − |
| NP-mAb-24 | n.d. | ++ | − | n.d. | + | − | − | − | − | − |
| NP-mAb-25 | n.d. | ++ | − | n.d. | − | − | − | − | − | − |
| NP-mAb-26 | n.d. | + | + | n.d. | + | − | − | − | − | − |
| NP-mAb-27 | n.d. | + | + | n.d. | + | − | − | − | − | − |
| NP-mAb-30 | n.d. | ++ | − | n.d. | + | − | − | − | − | − |
| NP-mAb-33 | n.d. | ++ | − | n.d. | + | − | − | − | − | − |
| NP-mAb-35 | n.d. | + | − | n.d. | − | − | − | − | − | − |
| NP-mAb-39 | IgG1, κ | +++ | + | − | − | − | − | − | − | − |
| NP-mAb-40 | IgG1, κ | +++ | + | − | + | − | − | − | − | − |
| NP-mAb-42 | IgG1, κ | +++ | + | − | − | − | − | − | − | − |
| NP-mAb-43 | n.d. | + | + | n.d. | + | − | − | − | − | − |
| NP-mAb-46 | IgG1, κ | +++ | + | − | − | − | − | − | − | − |
| NP-mAb-49 | IgG1, κ | +++ | − | − | − | − | − | − | − | − |
| NP-mAb-50 | n.d. | ++ | − | n.d. | + | − | − | − | − | − |
| NP-mAb-51 | IgG1, κ | ++ | + | + | − | − | − | − | − | − |
| NP-mAb-52 | IgG1, κ | +++ | + | − | − | − | − | − | − | − |
| NP-mAb-53 | IgG1, κ | +++ | + | + | + | − | − | − | − | − |
| NP-mAb-54 | n.d. | + | + | n.d. | n.d. | − | − | − | − | − |
| NP-mAb-55 | n.d. | +++ | + | n.d. | n.d. | − | − | − | − | − |
| NP-mAb-56 | n.d. | ++ | + | n.d. | + | − | − | − | − | − |
| NP-mAb-61 | n.d. | ++ | + | n.d. | n.d. | − | − | − | − | − |
| NP-mAb-62 | n.d. | ++ | + | n.d. | n.d. | − | − | − | − | − |
| Inactivated SARS-CoV-2 Conc. (TCID50/mL) | Positive/Total | Positive Percentage | |
| 1 | 2.19 × 103 | 9/9 | 100% |
| 2 | 1.09 × 103 | 9/9 | 100% |
| 3 | 5.47 × 102 | 9/9 | 100% |
| 4 | 2.73 × 102 | 8/9 | 89% |
| Inactivated SARS-CoV-2 Conc. (TCID50/mL) | Positive/Total | Positive Percentage | |
| 5.47 × 102 | 20/20 | 100% | |
| Gender | |
| Male | 50% |
| Female | 50% |
| Age (Years) | |
| Range | 16–80 |
| Average | 45.3 |
| Median | 43 |
| IQR | 23.5 |
| RT-PCR-Positive Samples (n) | 10 |
| Ct values | |
| Range | 16–29.5 |
| Average | 22.5 |
| Median | 21.1 |
| RT-PCR | NP-mAb-40/7 LFIA Strips | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Positive | Negative | Sensitivity | Specificity | Accuracy | |||||
| % | 95% CI | % | 95% CI | % | 95% CI | ||||
| Positive | 10 | 9 | 1 | 90 | 55.5–99.8 | 100 | 92.9–100 | 98.3 | 91–100 |
| Negative | 50 | 0 | 50 | ||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, R.-M.; Ko, S.-H.; Chen, W.-Y.; Chang, Y.-L.; Lin, H.-T.; Wu, H.-C. Monoclonal Antibodies against Nucleocapsid Protein of SARS-CoV-2 Variants for Detection of COVID-19. Int. J. Mol. Sci. 2021, 22, 12412. https://doi.org/10.3390/ijms222212412
Lu R-M, Ko S-H, Chen W-Y, Chang Y-L, Lin H-T, Wu H-C. Monoclonal Antibodies against Nucleocapsid Protein of SARS-CoV-2 Variants for Detection of COVID-19. International Journal of Molecular Sciences. 2021; 22(22):12412. https://doi.org/10.3390/ijms222212412
Chicago/Turabian StyleLu, Ruei-Min, Shih-Han Ko, Wan-Yu Chen, Yu-Ling Chang, Hsiu-Ting Lin, and Han-Chung Wu. 2021. "Monoclonal Antibodies against Nucleocapsid Protein of SARS-CoV-2 Variants for Detection of COVID-19" International Journal of Molecular Sciences 22, no. 22: 12412. https://doi.org/10.3390/ijms222212412
APA StyleLu, R.-M., Ko, S.-H., Chen, W.-Y., Chang, Y.-L., Lin, H.-T., & Wu, H.-C. (2021). Monoclonal Antibodies against Nucleocapsid Protein of SARS-CoV-2 Variants for Detection of COVID-19. International Journal of Molecular Sciences, 22(22), 12412. https://doi.org/10.3390/ijms222212412






