Identification of Potential Long Non-Coding RNA Candidates that Contribute to Triple-Negative Breast Cancer in Humans through Computational Approach
Abstract
:1. Introduction
2. Results
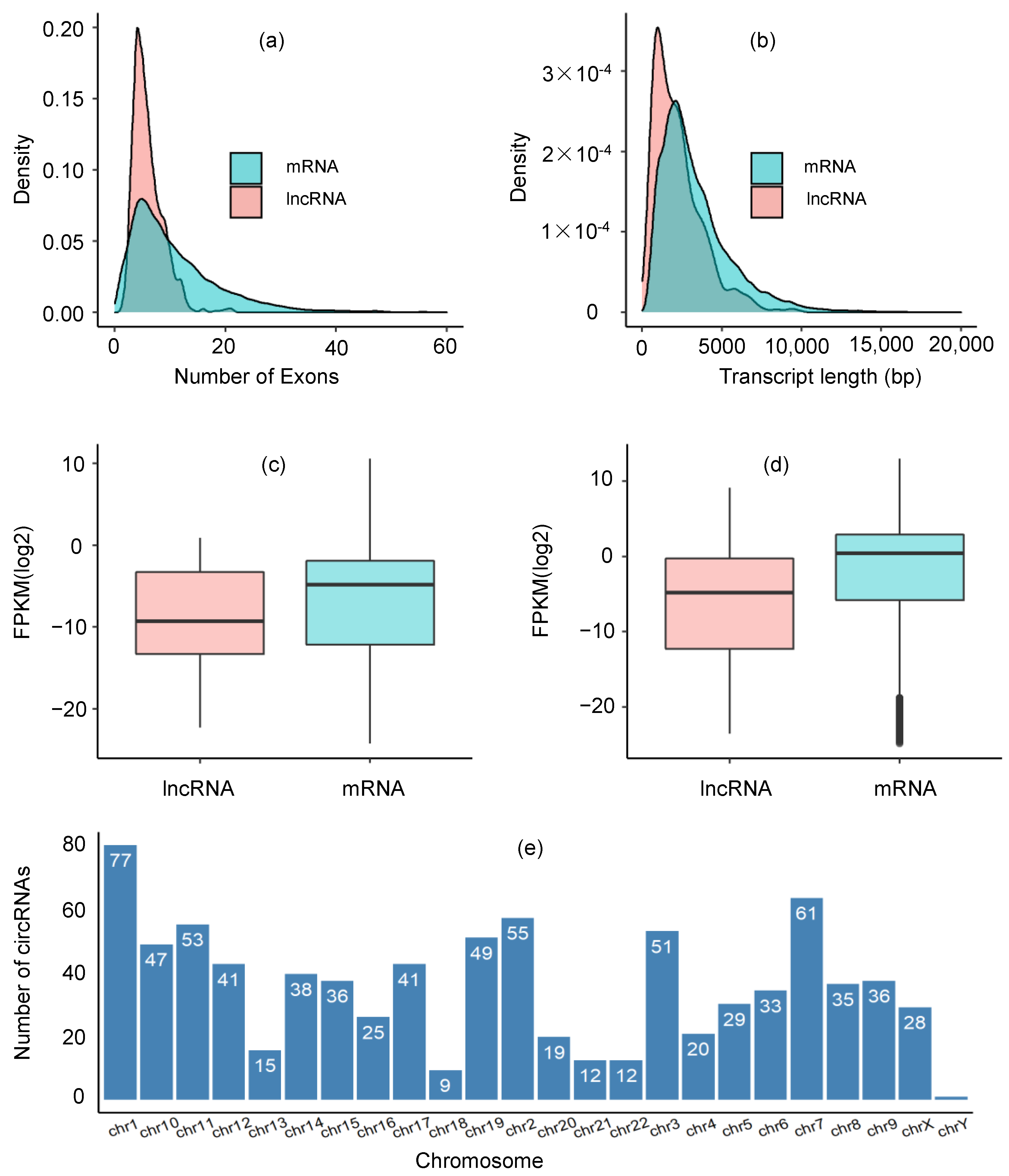
2.1. Identification of lncRNA Transcripts
2.2. Expression Profiles of lncRNAs and mRNAs
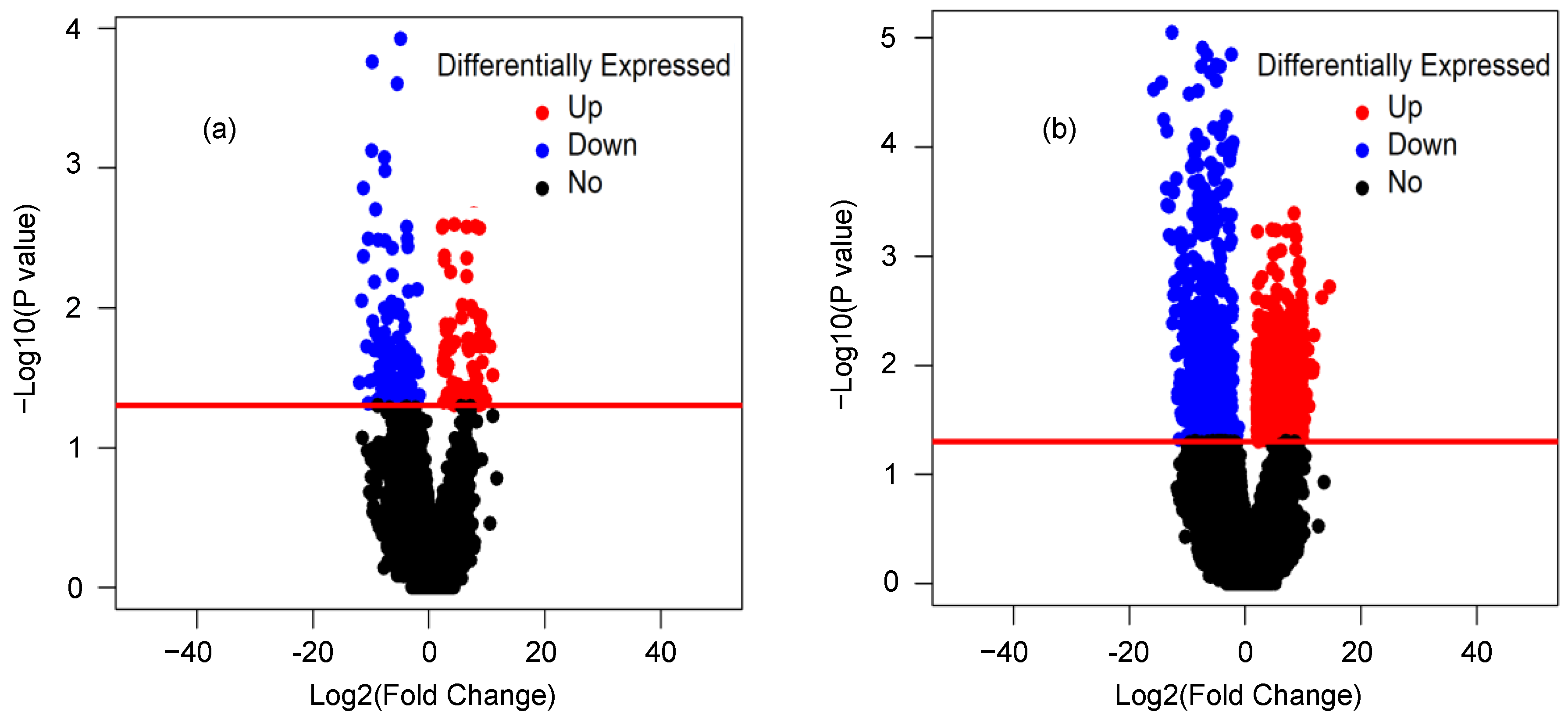
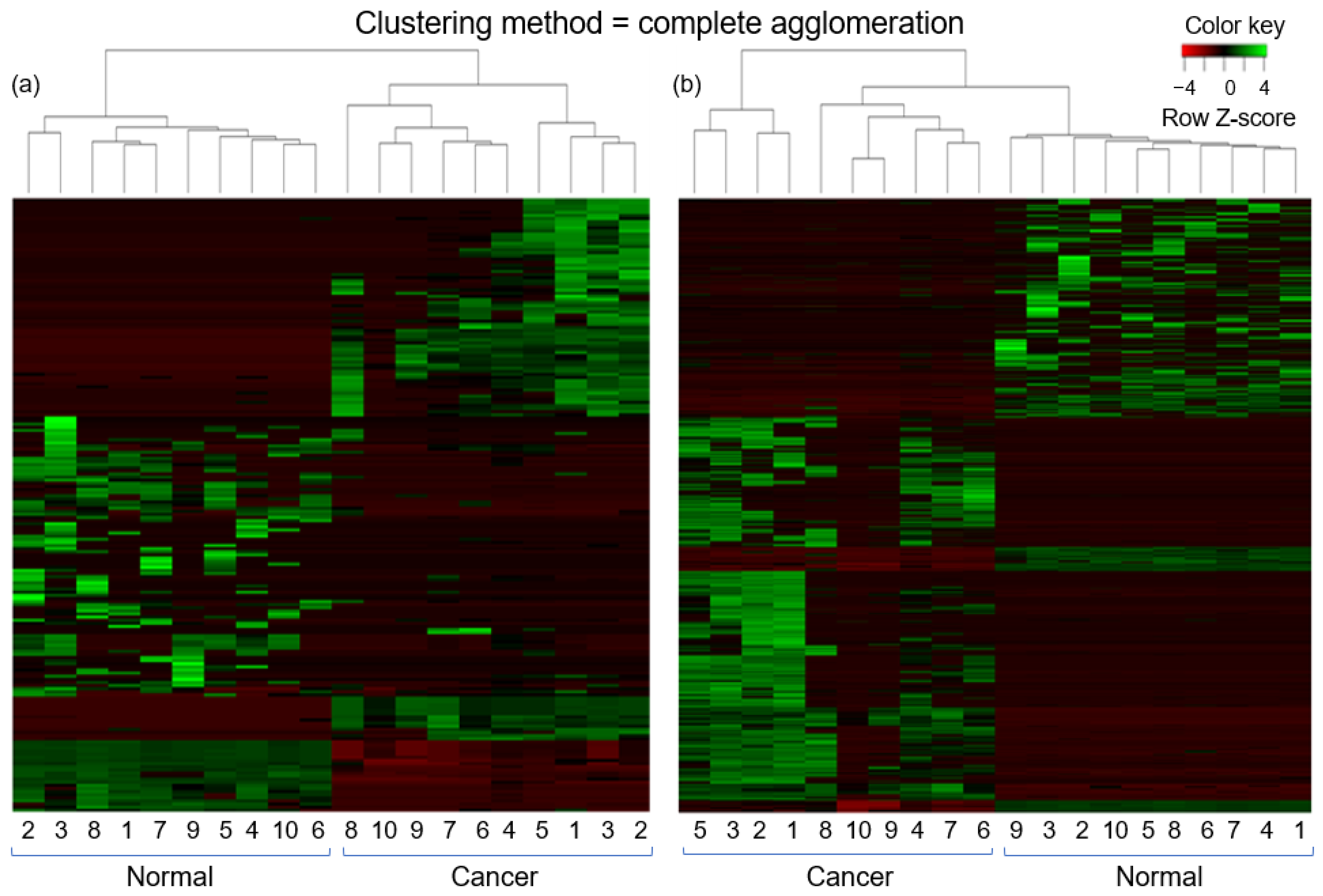
2.3. Differential Expression Analysis of lncRNAs and mRNAs
2.4. Analysis of lncRNA-miRNA-Gene Interaction
2.5. GO Term and KEGG Pathway Enrichment Analysis
3. Discussion
4. Materials and Methods
4.1. Data Collection
4.2. Quality Control, Mapping, and Transcript Assembly
4.3. Transcripts Filtration
4.4. Assessment of the Coding Potential
4.5. Differential Expression Analysis
4.6. GO and KEGG Enrichment Analysis
4.7. LncRNA–miRNA-Gene Interaction Network Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chen, W.; Zheng, R.; Baade, P. Cancer statistics in China. CA Cancer J. Clin. 2016, 66, 115–132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bray, F.; McCarron, P.; Parkin, D.M. The changing global patterns of female breast cancer incidence and mortality. Breast Cancer Res. 2004, 6, 229–239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, L.; Zhang, Y.; Lu, J. The roles of long noncoding RNAs in breast cancer metastasis. Cell Death Dis. 2020, 11, 749. [Google Scholar] [CrossRef] [PubMed]
- Gupta, G.P.; Massagué, J. Cancer metastasis: Building a framework. Cell 2006, 127, 679–695. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [Green Version]
- Koboldt, D.C.; Fulton, R.S.; McLellan, M.D.; Schmidt, H.; Kalicki-Veizer, J.; McMichael, J.F.; Fulton, L.L.; Dooling, D.J.; Ding, L.; Mardis, E.R.; et al. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61–70. [Google Scholar] [CrossRef] [Green Version]
- Perou, C.M.; Sørile, T.; Eisen, M.B.; Van De Rijn, M.; Jeffrey, S.S.; Ress, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef]
- Rakha, E.A.; El-Sayed, M.E.; Green, A.R.; Lee, A.H.S.; Robertson, J.F.; Ellis, I.O. Prognostic markers in triple-negative breast cancer. Cancer 2007, 109, 25–32. [Google Scholar] [CrossRef]
- Pusztai, L.; Karn, T.; Safonov, A.; Abu-Khalaf, M.M.; Bianchini, G. New strategies in breast cancer: Immunotherapy. Clin. Cancer Res. 2016, 22, 2105–2110. [Google Scholar] [CrossRef] [Green Version]
- Klinge, C.M. Non-coding RNAs in breast cancer: Intracellular and intercellular communication. Non-Coding RNA 2018, 4, 40. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, S.; He, Y.; Yang, S.; Hu, J.; Zhang, Q.; Chen, W.; Xu, H.; Zhang, H.; Zhong, S.; Zhao, J.; et al. The regulatory roles of lncRNAs in the process of breast cancer invasion and metastasis. Biosci. Rep. 2018, 38, BSR20180772. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tomar, D.; Yadav, A.S.; Kumar, D.; Bhadauriya, G.; Kundu, G.C. Non-coding RNAs as potential therapeutic targets in breast cancer. Biochim. Biophys. Acta Gene Regul. Mech. 2020, 1863, 194378. [Google Scholar] [CrossRef]
- Bin, X.; Hongjian, Y.; Xiping, Z.; Bo, C.; Shifeng, Y.; Binbin, T. Research progresses in roles of LncRNA and its relationships with breast cancer. Cancer Cell Int. 2018, 18, 179. [Google Scholar] [CrossRef]
- Bhan, A.; Soleimani, M.; Mandal, S.S. Long noncoding RNA and cancer: A new paradigm. Cancer Res. 2017, 77, 3965–3981. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iyer, M.K.; Niknafs, Y.S.; Malik, R.; Singhal, U.; Sahu, A.; Hosono, Y.; Barrette, T.R.; Prensner, J.R.; Evans, J.R.; Zhao, S.; et al. The landscape of long noncoding RNAs in the human transcriptome. Nat. Genet. 2015, 47, 199–208. [Google Scholar] [CrossRef]
- Zhang, T.; Hu, H.; Yan, G.; Wu, T.; Liu, S.; Chen, W.; Ning, Y.; Lu, Z. Long non-coding RNA and breast cancer. Technol. Cancer Res. Treat. 2019, 18, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Peng, W.X.; Koirala, P.; Mo, Y.Y. LncRNA-mediated regulation of cell signaling in cancer. Oncogene 2017, 36, 5661–5667. [Google Scholar] [CrossRef]
- Li, Z.; Hou, P.; Fan, D.; Dong, M.; Ma, M.; Li, H.; Yao, R.; Li, Y.; Wang, G.; Geng, P.; et al. The degradation of EZH2 mediated by lncRNA ANCR attenuated the invasion and metastasis of breast cancer. Cell Death Differ. 2017, 24, 59–71. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lou, K.X.; Li, Z.H.; Wang, P.; Liu, Z.; Chen, Y.; Wang, X.L.; Cui, H.X. Long non-coding RNA BANCR indicates poor prognosis for breast cancer and promotes cell proliferation and invasion. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 1358–1365. [Google Scholar] [CrossRef]
- Li, W.; Jia, G.; Qu, Y.; Du, Q.; Liu, B.; Liu, B. Long non-coding RNA (LncRNA) HOXA11-AS promotes breast cancer invasion and metastasis by regulating epithelial-mesenchymal transition. Med. Sci. Monit. 2017, 23, 3393–3403. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hansji, H.; Leung, E.Y.; Baguley, B.C.; Finlay, G.J.; Askarian-Amiri, M.E. Keeping abreast with long non-coding RNAs in mammary gland development and breast cancer. Front. Genet. 2014, 5, 379. [Google Scholar] [CrossRef] [Green Version]
- Tripathi, M.K.; Doxtater, K.; Keramatnia, F.; Zacheaus, C.; Yallapu, M.M.; Jaggi, M.; Chauhan, S.C. Role of lncRNAs in ovarian cancer: Defining new biomarkers for therapeutic purposes. Drug Discov. Today 2018, 23, 1635–1643. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.M.; Tsang, F.H.; Ng, I.O.L. Non-coding RNAs in hepatocellular carcinoma: Molecular functions and pathological implications. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 137–151. [Google Scholar] [CrossRef]
- Alvarez-Dominguez, J.R.; Lodish, H.F. Emerging mechanisms of long noncoding RNA function during normal and malignant hematopoiesis. Blood 2017, 130, 1965–1975. [Google Scholar] [CrossRef] [PubMed]
- Collina, F.; Aquino, G.; Brogna, M.; Cipolletta, S.; Buonfanti, G.; De Laurentiis, M.; Di Bonito, M.; Cantile, M.; Botti, G. LncRNA HOTAIR up-regulation is strongly related with lymph nodes metastasis and LAR subtype of triple negative breast cancer. J. Cancer 2019, 10, 2018–2024. [Google Scholar] [CrossRef] [Green Version]
- Haffty, B.G.; Yang, Q.; Reiss, M.; Kearney, T.; Higgins, S.A.; Weidhaas, J.; Harris, L.; Hait, W.; Toppmeyer, D. Locoregional relapse and distant metastasis in conservatively managed triple negative early-stage breast cancer. J. Clin. Oncol. 2006, 24, 5664. [Google Scholar] [CrossRef]
- Chen, V.E.; Gillespie, E.F.; Zakeri, K.; Murphy, J.D.; Yashar, C.M.; Lu, S.; Einck, J.P. Pathologic response after neoadjuvant chemotherapy predicts locoregional control in patients with triple negative breast cancer. Adv. Radiat. Oncol. 2017, 2, 105–109. [Google Scholar] [CrossRef] [Green Version]
- Fan, H.; Yuan, J.; Li, X.; Ma, Y.; Wang, X.; Xu, B.; Li, X. LncRNA LINC00173 enhances triple-negative breast cancer progression by suppressing miR-490-3p expression. Biomed. Pharmacother. 2020, 125, 109987. [Google Scholar] [CrossRef] [PubMed]
- Han, C.; Fu, Y.; Zeng, N.; Yin, J.; Li, Q. LncRNA FAM83H-AS1 promotes triple-negative breast cancer progression by regulating the miR-136-5p/metadherin axis. Aging 2020, 12, 3594. [Google Scholar] [CrossRef]
- Hua, K.; Deng, X.; Hu, J.; Ji, C.; Yu, Y.; Li, J.; Wang, X.; Fang, L. Long noncoding RNA HOST2, working as a competitive endogenous RNA, promotes STAT3-mediated cell proliferation and migration via decoying of let-7b in triple-negative breast cancer. J. Exp. Clin. Cancer Res. 2020, 39, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Dong, S.Y.; Lv, L.; Liu, Y.H.; Yao, Z.H.; Zhang, X.H.; Wang, O.C. Long non-coding RNA AFAP1-AS1 was up-regulated in triple-negative breast cancer and regulated proliferation and invasion. Int. J. Clin. Exp. Pathol. 2016, 9, 6378–6384. [Google Scholar]
- Zuo, Y.; Li, Y.; Zhou, Z.; Ma, M.; Fu, K. Long non-coding RNA MALAT1 promotes proliferation and invasion via targeting miR-129-5p in triple-negative breast cancer. Biomed. Pharmacother. 2017, 95, 922–928. [Google Scholar] [CrossRef] [PubMed]
- Niu, L.; Fan, Q.; Yan, M.; Wang, L. LncRNA NRON down-regulates lncRNA snaR and inhibits cancer cell proliferation in TNBC. Biosci. Rep. 2019, 39, BSR20190468. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Liu, D.; Wu, X.; Zeng, Y.; Li, L.; Hou, Y.; Li, W.; Liu, Z. Long non-coding RNA (LncRNA) RMST in triple-negative breast cancer (TNBC): Expression analysis and biological roles research. J. Cell. Physiol. 2018, 233, 6603–6612. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Ke, H.; Zhang, H.; Ma, Y.; Ao, L.; Zou, L.; Yang, Q.; Zhu, H.; Nie, J.; Wu, C.; et al. LncRNA MIR100HG promotes cell proliferation in triple-negative breast cancer through triplex formation with p27 loci. Cell Death Dis. 2018, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Tang, H.; Ling, L.; Li, N.; Jia, X.; Zhang, Z.; Wang, X.; Shi, L.; Yin, J.; Qiu, N.; et al. LINC01638 lncRNA activates MTDH-Twist1 signaling by preventing SPOP-mediated c-Myc degradation in triple-negative breast cancer. Oncogene 2018, 37, 1178. [Google Scholar] [CrossRef]
- Liang, Y.; Hu, J.; Li, J.; Liu, Y.; Yu, J.; Zhuang, X.; Mu, L.; Kong, X.; Hong, D.; Yang, Q.; et al. Epigenetic activation of TWIST1 by MTDH promotes cancer stem-like cell traits in breast cancer. Cancer Res. 2015, 75, 3672–3680. [Google Scholar] [CrossRef] [Green Version]
- Wang, O.C.; Yang, F.; Liu, Y.H.; Lv, L.; Ma, R.; Chen, C.; Wang, J.; Tan, Q.; Cheng, Y.; Xia, E.; et al. C-MYC-induced upregulation of lncRNA SNHG12 regulates cell proliferation, apoptosis and migration in triple-negative breast cancer. Am. J. Transl. Res. 2017, 9, 533. [Google Scholar]
- Cech, T.R.; Steitz, J.A. The noncoding RNA revolution—Trashing old rules to forge new ones. Cell 2014, 157, 77–94. [Google Scholar] [CrossRef] [Green Version]
- Pang, K.C.; Frith, M.C.; Mattick, J.S. Rapid evolution of noncoding RNAs: Lack of conservation does not mean lack of function. Trends Genet. 2006, 22, 1–5. [Google Scholar] [CrossRef]
- Weikard, R.; Demasius, W.; Kuehn, C. Mining long noncoding RNA in livestock. Anim. Genet. 2017, 48, 3–18. [Google Scholar] [CrossRef] [PubMed]
- Dent, R.; Trudeau, M.; Pritchard, K.I.; Hanna, W.M.; Kahn, H.K.; Sawka, C.A.; Lickley, L.A.; Rawlinson, E.; Sun, P.; Narod, S.A. Triple-negative breast cancer: Clinical features and patterns of recurrence. Clin. Cancer Res. 2007, 13, 4429–4434. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.X.; Wang, L.J.; Hou, J.; Liu, H.Y.; Wang, R.; Wang, C.; Xie, W.H. Identification of long noncoding RNAs as predictors of survival in triple-negative breast cancer based on network analysis. Biomed. Res. Int. 2020, 2020, 8970340. [Google Scholar] [CrossRef] [PubMed]
- Weikard, R.; Hadlich, F.; Kuehn, C. Identification of novel transcripts and noncoding RNAs in bovine skin by deep next generation sequencing. BMC Genom. 2013, 14, 789. [Google Scholar] [CrossRef] [Green Version]
- Billerey, C.; Boussaha, M.; Esquerré, D.; Rebours, E.; Djari, A.; Meersseman, C.; Klopp, C.; Gautheret, D.; Rocha, D. Identification of large intergenic non-coding RNAs in bovine muscle using next-generation transcriptomic sequencing. BMC Genom. 2014, 15, 499. [Google Scholar] [CrossRef] [Green Version]
- Gupta, P.; Peter, S.; Jung, M.; Lewin, A.; Hemmrich-Stanisak, G.; Franke, A.; von Kleist, M.; Schütte, C.; Einspanier, R.; Sharbati, S.; et al. Analysis of long non-coding RNA and mRNA expression in bovine macrophages brings up novel aspects of Mycobacterium avium subspecies paratuberculosis infections. Sci. Rep. 2019, 9, 1571. [Google Scholar] [CrossRef] [Green Version]
- Delás, M.J.; Hannon, G.J. lncRNAs in development and disease: From functions to mechanisms. Open Biol. 2017, 7, 170121. [Google Scholar] [CrossRef] [Green Version]
- Li, Z.; Zhao, W.; Wang, M.; Zhou, X. The Role of Long Noncoding RNAs in Gene Expression Regulation; IntechOpen: London, UK, 2019. [Google Scholar]
- Bonofiglio, D.; Cione, E.; Vizza, D.; Perri, M.; Pingitore, A.; Qi, H.; Catalano, S.; Rovito, D.; Genchi, G.; Andò, S. Bid as a potential target of apoptotic effects exerted by low doses of PPARγ and RXR ligands in breast cancer cells. Cell Cycle 2011, 10, 2344–2354. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Memon, A.; Lee, W.K. KLF10 as a tumor suppressor gene and its TGF-β signaling. Cancers 2018, 10, 161. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Subramaniam, M.; Hawse, J.R.; Rajamannan, N.M.; Ingle, J.N.; Spelsberg, T.C. Functional role of KLF10 in multiple disease processes. BioFactors 2010, 36, 8–18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karaayvaz, M.; Cristea, S.; Gillespie, S.M.; Patel, A.P.; Mylvaganam, R.; Luo, C.C.; Specht, M.C.; Bernstein, B.E.; Michor, F.; Ellisen, L.W. Unravelling subclonal heterogeneity and aggressive disease states in TNBC through single-cell RNA-seq. Nat. Commun. 2018, 9, 3588. [Google Scholar] [CrossRef] [Green Version]
- Gao, R.; Kim, C.; Sei, E.; Foukakis, T.; Crosetto, N.; Chan, L.K.; Srinivasan, M.; Zhang, H.; Meric-Bernstam, F.; Navin, N. Nanogrid single-nucleus RNA sequencing reveals phenotypic diversity in breast cancer. Nat. Commun. 2017, 8, 228. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [Green Version]
- Koufariotis, L.T.; Chen, Y.P.P.; Chamberlain, A.; Jagt, C.V.; Hayes, B.J. A catalogue of novel bovine long noncoding RNA across 18 tissues. PLoS ONE 2015, 10, e0141225. [Google Scholar] [CrossRef]
- Jorquera, R.; Ortiz, R.; Ossandon, F.; Cárdenas, J.P.; Sepúlveda, R.; González, C.; Holmes, D.S. SinEx DB: A database for single exon coding sequences in mammalian genomes. Database 2016, 2016, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Sun, M.; Gadad, S.S.; Kim, D.S.; Kraus, W.L. Discovery, annotation, and functional analysis of long noncoding RNAs controlling cell-cycle gene expression and proliferation in breast cancer cells. Mol. Cell 2015, 59, 698–711. [Google Scholar] [CrossRef] [Green Version]
- Bush, S.J.; Muriuki, C.; McCulloch, M.E.B.; Farquhar, I.L.; Clark, E.L.; Hume, D.A. Cross-species inference of long non-coding RNAs greatly expands the ruminant transcriptome. Genet. Sel. Evol. 2018, 50, 20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rédei, G.P. Encyclopedia of genetics, genomics, proteomics and informatics. In Encyclopedia of Genetics, Genomics, Proteomics and Informatics; Springer: Berlin/Heidelberg, Germany, 2008; p. 285. [Google Scholar]
- Kong, L.; Zhang, Y.; Ye, Z.Q.; Liu, X.Q.; Zhao, S.Q.; Wei, L.; Gao, G. CPC: Assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007, 35, 345–349. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]


| mRNA ID | Location | No. of Exons | Length | Base Mean | LFC | p-Value | Gene |
|---|---|---|---|---|---|---|---|
| TCONS_00027847 | chr16:72054592-72061056 | 7 | 1448 | 3419.94 | Inf | 0.00041 | HP |
| TCONS_00062848 | chr5:150401637-150412936 | 9 | 1681 | 2201.54 | Inf | 0.00058 | CD74 |
| TCONS_00027845 | chr16:72054592-72061056 | 5 | 1271 | 2210.50 | Inf | 0.00058 | HP |
| TCONS_00062849 | chr5:150401637-150412936 | 8 | 1489 | 2166.03 | Inf | 0.00058 | CD74 |
| TCONS_00037794 | chr19:6677835-6720682 | 41 | 5132 | 2131.98 | Inf | 0.00059 | C3 |
| TCONS_00027846 | chr16:72054592-72061056 | 5 | 1271 | 2125.11 | Inf | 0.00060 | HP |
| TCONS_00030302 | chr17:34255277-34257201 | 3 | 747 | 1907.13 | Inf | 0.00067 | CCL2 |
| TCONS_00008098 | chr10:58513144-58831437 | 21 | 5471 | 1317.61 | Inf | 0.00087 | BICC1 |
| TCONS_00062847 | chr5:150401637-150412936 | 6 | 1305 | 1274.15 | Inf | 0.00089 | CD74 |
| TCONS_00005374 | chr1:89052304-89065360 | 11 | 3050 | 1164.10 | Inf | 0.00096 | GBP1 |
| TCONS_00065563 | chr6:27866792-27867581 | 1 | 790 | 301.37 | −12.6 | 8.98 × 10−06 | HIST1H1B |
| TCONS_00028779 | chr16:30104810-30113557 | 10 | 1098 | 105.09 | −Inf | 1.25 × 10−05 | GDPD3 |
| TCONS_00010969 | chr10:128096661-128126204 | 14 | 11,415 | 114.64 | −Inf | 1.44 × 10−05 | MKI67 |
| TCONS_00010970 | chr10:128096661-128126204 | 15 | 12,495 | 119.33 | −Inf | 1.45 × 10−05 | MKI67 |
| TCONS_00012090 | chr11:62270155-62273157 | 3 | 506 | 117.08 | −Inf | 1.80 × 10−05 | SCGB2A2 |
| TCONS_00031424 | chr17:79778178-79787650 | 5 | 4265 | 42.19 | −Inf | 1.84 × 10−05 | CBX2 |
| TCONS_00082965 | chrX:154651972-154653579 | 3 | 764 | 24.78 | −Inf | 1.85 × 10−05 | CTAG2 |
| TCONS_00054384 | chr3:170037947-170085395 | 4 | 2013 | 18.08 | −Inf | 2.11 × 10−05 | GPR160 |
| TCONS_00082964 | chrX:154651972-154653579 | 2 | 993 | 20.37 | −Inf | 2.50 × 10−05 | CTAG2 |
| TCONS_00018204 | chr12:52806543-52814116 | 9 | 2147 | 45.76 | −14.4 | 2.61 × 10−05 | KRT4 |
| lncRNA ID | Location | No. of Exons | Length | Base Mean | LFC | p-Value | Gene |
|---|---|---|---|---|---|---|---|
| TCONS_00057817 | chr4:78645994-78684501 | 4 | 4214 | 362.9557 | Inf | 0.00254 | LINC01094 |
| TCONS_00057819 | chr4:78645994-78684501 | 3 | 4120 | 352.7188 | Inf | 0.00260 | LINC01094 |
| TCONS_00057820 | chr4:78645994-78684501 | 3 | 4105 | 345.6054 | Inf | 0.00263 | LINC01094 |
| TCONS_00057818 | chr4:78645994-78684501 | 4 | 4192 | 339.0413 | Inf | 0.00266 | LINC01094 |
| TCONS_00057821 | chr4:78645994-78684501 | 3 | 4066 | 338.1632 | Inf | 0.00268 | LINC01094 |
| TCONS_00057816 | chr4:78645994-78684501 | 5 | 4286 | 331.5815 | Inf | 0.00270 | LINC01094 |
| TCONS_00006590 | chr1:169690665-169708856 | 7 | 2206 | 176.9738 | Inf | 0.00426 | SELL |
| TCONS_00062278 | chr5:91368632-91383373 | 8 | 4332 | 208.4584 | Inf | 0.00443 | ARRDC3 |
| TCONS_00062275 | chr5:91368632-91380297 | 8 | 4621 | 200.0444 | Inf | 0.00463 | ARRDC3 |
| TCONS_00009315 | chr10:17214239-17229985 | 3 | 1875 | 121.2627 | Inf | 0.00557 | VIM-AS1 |
| TCONS_00082872 | chrX:152708261-152714549 | 5 | 825 | 2.665319 | −Inf | 0.00012 | CSAG3 |
| TCONS_00081533 | chrX:152753921-152760222 | 5 | 825 | 2.704539 | −Inf | 0.00018 | CSAG3 |
| TCONS_00035347 | chr19:751113-764319 | 4 | 1048 | 12.04587 | −Inf | 0.00025 | MISP |
| TCONS_00017579 | chr12:6848808-6851930 | 5 | 1615 | 5.361052 | −Inf | 0.00076 | CDCA3 |
| TCONS_00017578 | chr12:6848808-6851930 | 6 | 1792 | 5.745075 | −Inf | 0.00085 | CDCA3 |
| TCONS_00012837 | chr11:111912736-111926871 | 5 | 1527 | 30.57432 | −Inf | 0.00105 | HSPB2-C11orf52 |
| TCONS_00024012 | chr15:43593834-43599406 | 9 | 2842 | 10.85619 | −11.3 | 0.00140 | CKMT1B |
| TCONS_00081534 | chrX:152753921-152760222 | 5 | 789 | 0.786254 | −Inf | 0.00198 | CSAG3 |
| TCONS_00082873 | chrX:152708261-152714549 | 5 | 789 | 0.968629 | −Inf | 0.00266 | CSAG3 |
| TCONS_00082981 | chrX:155061625-155071272 | 6 | 2755 | 2.456605 | −Inf | 0.00322 | MTCP1 |
| lncRNA ID | Location | No. of Exons | Length | Base Mean | LFC | p-Value | Gene |
|---|---|---|---|---|---|---|---|
| TCONS_00076394 | chr8:102648777-102655902 | 4 | 2309 | 36.34 | Inf | 0.0146 | KLF10 |
| TCONS_00002947 | chr1:182789449-182830384 | 11 | 2728 | 32.38 | Inf | 0.0391 | NPL |
| TCONS_00051377 | chr22:17734140-17774665 | 5 | 1901 | 9.47 | Inf | 0.0434 | BID |
| TCONS_00082872 | chrX:152708261-152714549 | 5 | 825 | 2.67 | −Inf | 0.0001 | CSAG3 |
| TCONS_00081533 | chrX:152753921-152760222 | 5 | 825 | 2.70 | −Inf | 0.0002 | CSAG3 |
| TCONS_00082981 | chrX:155061625-155071272 | 6 | 2755 | 2.46 | −Inf | 0.0032 | MTCP1 |
| TCONS_00026822 | chr16:4788397-4796491 | 6 | 883 | 22.91 | −7.6 | 0.0033 | SMIM22 |
| TCONS_00049160 | chr20:47298126-47356889 | 5 | 1346 | 2.07 | −3.5 | 0.0076 | ZMYND8 |
| TCONS_00054549 | chr3:184361714-184368595 | 5 | 1723 | 4.02 | −6.8 | 0.0220 | POLR2H |
| TCONS_00054548 | chr3:184361714-184368595 | 6 | 1871 | 4.60 | −5.9 | 0.0233 | POLR2H |
| GO/Pathway ID | GO/Pathway Name | No. of Genes | p-Value |
|---|---|---|---|
| Biological Process | |||
| GO:0051781 | Positive regulation of cell division | 3 | 0.0036 |
| GO:0030593 | Neutrophil chemotaxis | 3 | 0.0070 |
| GO:0006600 | Creatine metabolic process | 2 | 0.0208 |
| GO:0001934 | Positive regulation of protein phosphorylation | 3 | 0.0243 |
| GO:0042832 | Defense response to protozoan | 2 | 0.0356 |
| GO:0008285 | Negative regulation of cell proliferation | 4 | 0.0390 |
| Cellular Component | |||
| GO:0005743 | Mitochondrial inner membrane | 4 | 0.0449 |
| Molecular Function | |||
| GO:0016772 | Transferase activity, transferring phosphorus-containing groups | 2 | 0.0094 |
| GO:0016301 | Kinase activity | 4 | 0.0105 |
| GO:0004111 | Creatine kinase activity | 2 | 0.0113 |
| GO:0008083 | Growth factor activity | 3 | 0.0376 |
| KEGG pathway | |||
| hsa05215 | Prostate cancer | 3 | 0.0265 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rahman, M.M.; Hossain, M.T.; Reza, M.S.; Peng, Y.; Feng, S.; Wei, Y. Identification of Potential Long Non-Coding RNA Candidates that Contribute to Triple-Negative Breast Cancer in Humans through Computational Approach. Int. J. Mol. Sci. 2021, 22, 12359. https://doi.org/10.3390/ijms222212359
Rahman MM, Hossain MT, Reza MS, Peng Y, Feng S, Wei Y. Identification of Potential Long Non-Coding RNA Candidates that Contribute to Triple-Negative Breast Cancer in Humans through Computational Approach. International Journal of Molecular Sciences. 2021; 22(22):12359. https://doi.org/10.3390/ijms222212359
Chicago/Turabian StyleRahman, Md. Motiar, Md. Tofazzal Hossain, Md. Selim Reza, Yin Peng, Shengzhong Feng, and Yanjie Wei. 2021. "Identification of Potential Long Non-Coding RNA Candidates that Contribute to Triple-Negative Breast Cancer in Humans through Computational Approach" International Journal of Molecular Sciences 22, no. 22: 12359. https://doi.org/10.3390/ijms222212359
APA StyleRahman, M. M., Hossain, M. T., Reza, M. S., Peng, Y., Feng, S., & Wei, Y. (2021). Identification of Potential Long Non-Coding RNA Candidates that Contribute to Triple-Negative Breast Cancer in Humans through Computational Approach. International Journal of Molecular Sciences, 22(22), 12359. https://doi.org/10.3390/ijms222212359






