Affibody Functionalized Beads for the Highly Sensitive Detection of Cancer Cell-Derived Exosomes
Abstract
:1. Introduction
2. Results
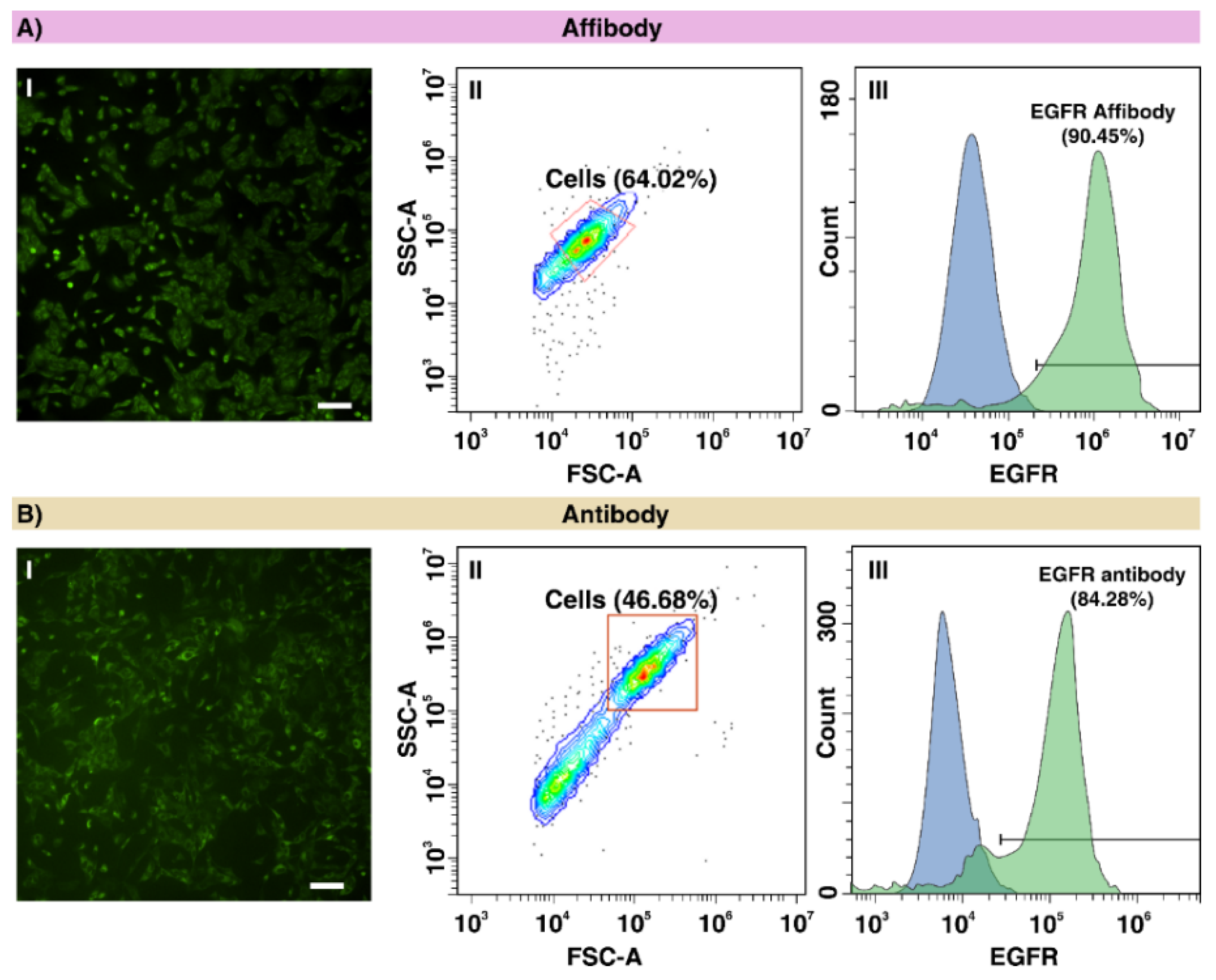
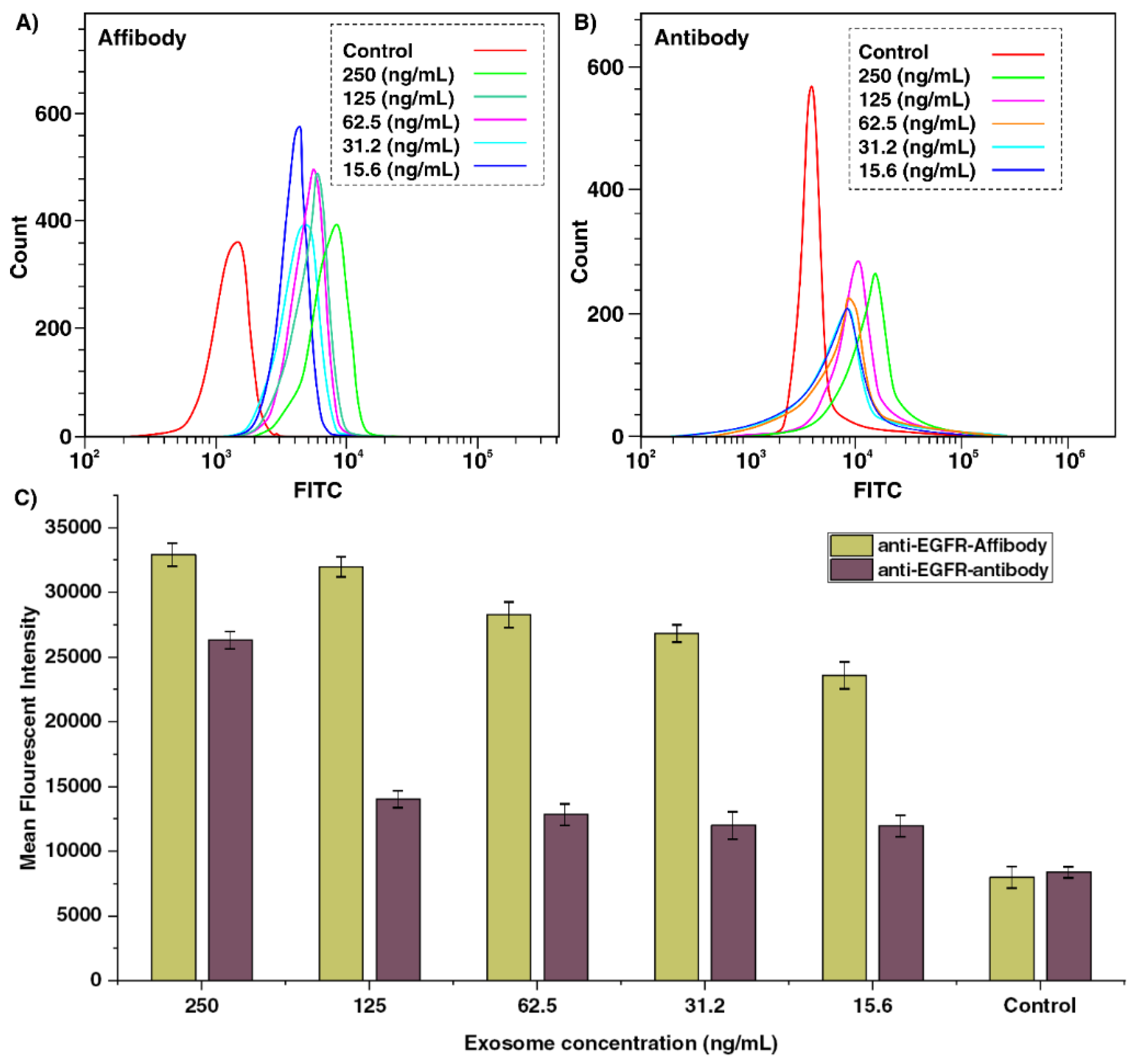
2.1. Evaluating the EGFR Expression in A549 Cells Using Anti-EGFR Affibody and Antibody
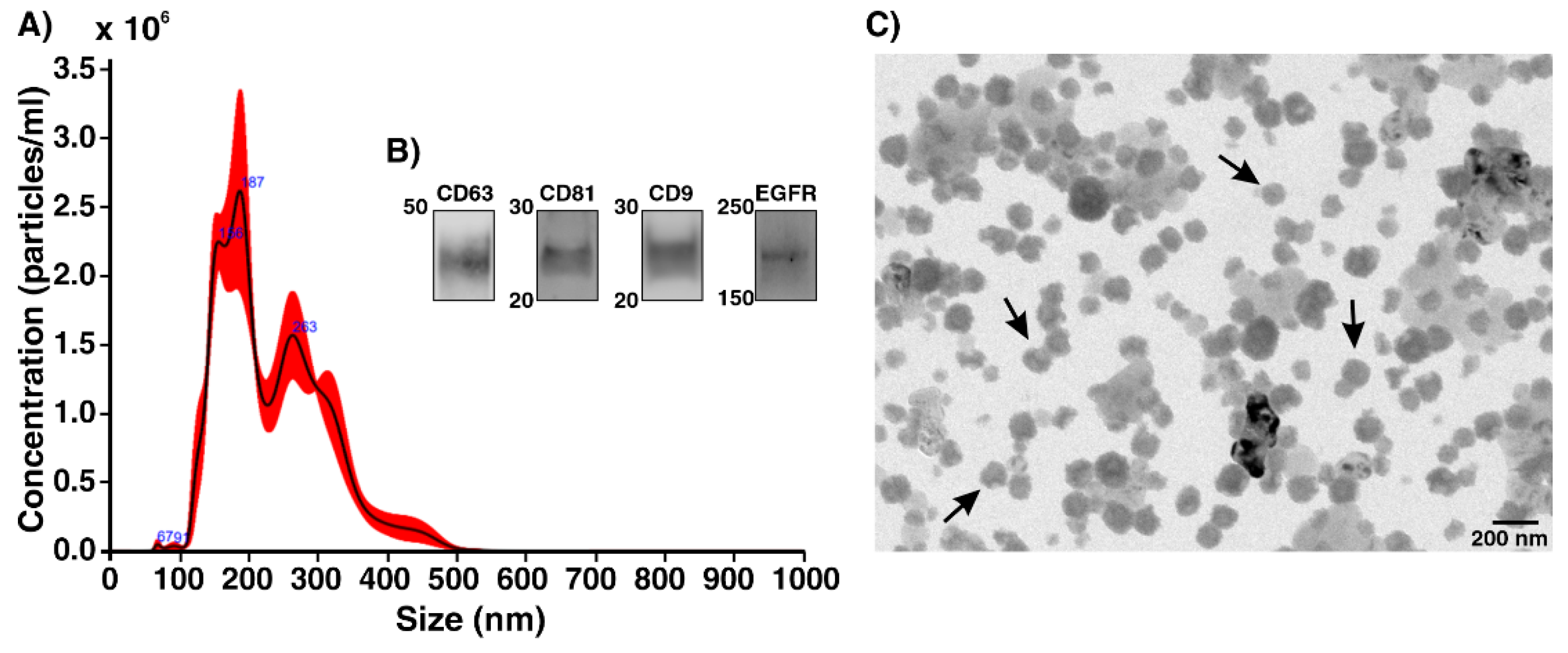
2.2. Characterization of Human Lung Cancer Cell-Derived Exosomes
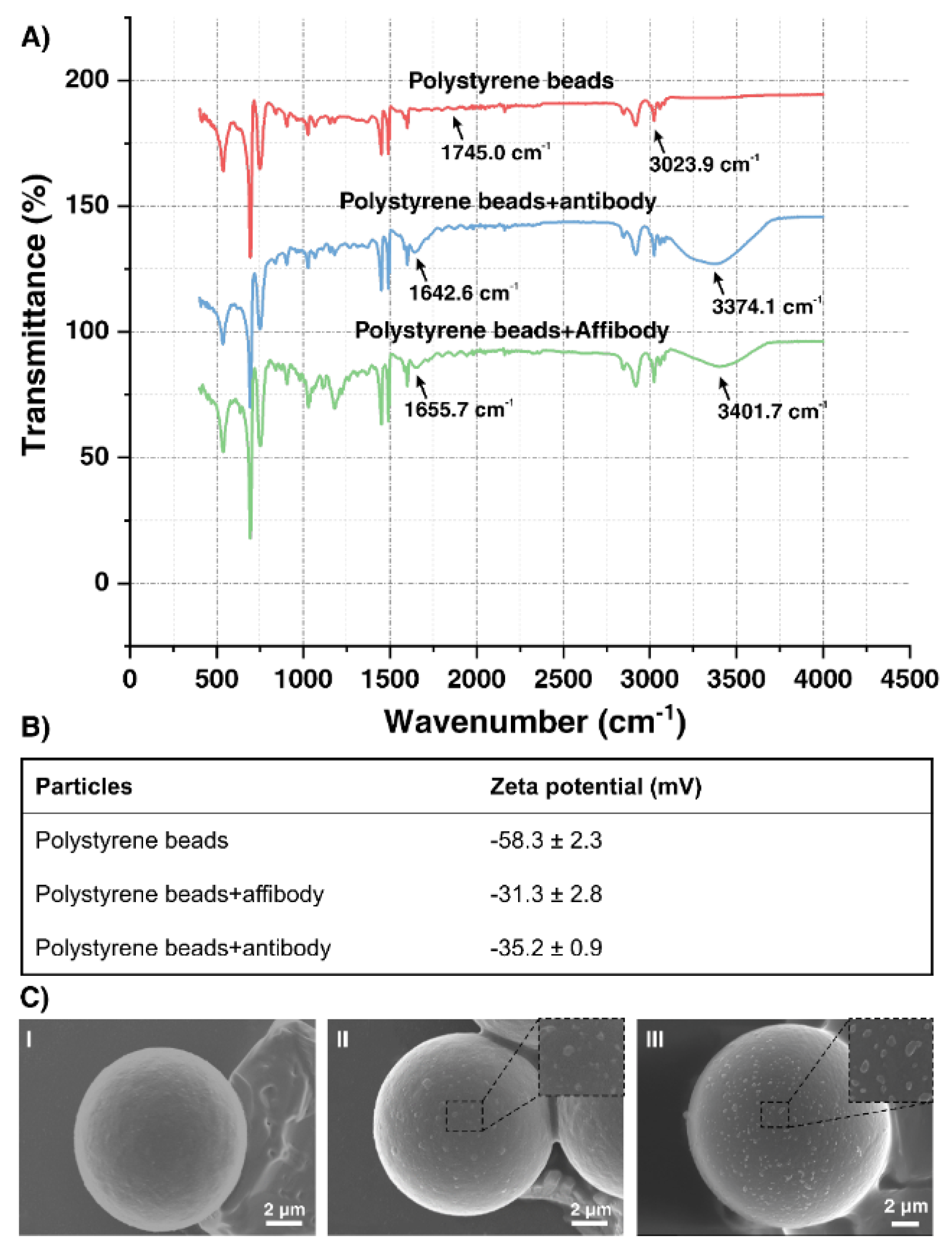
2.3. Surface Characterization of Functionalized Polystyrene Microbeads
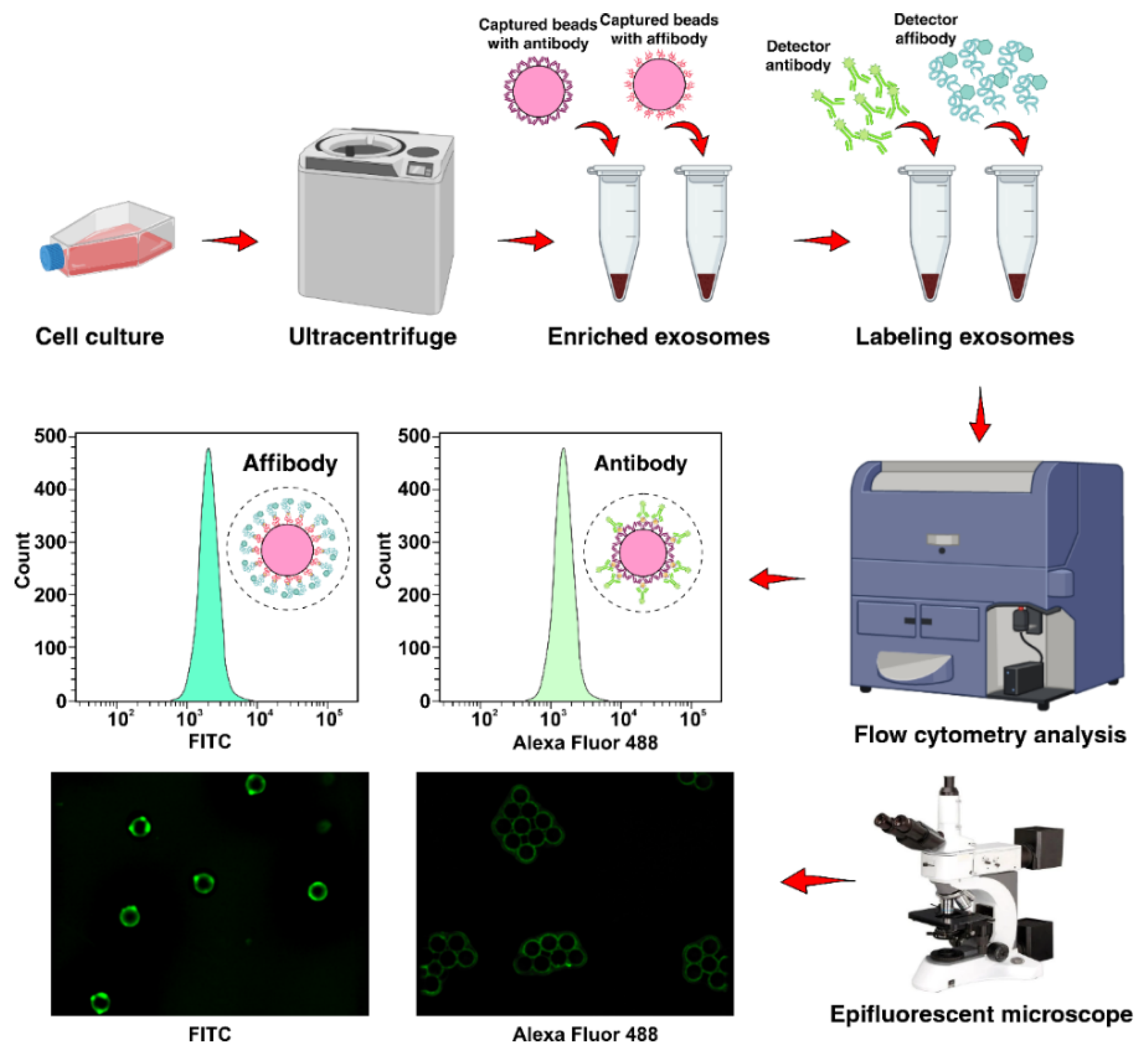
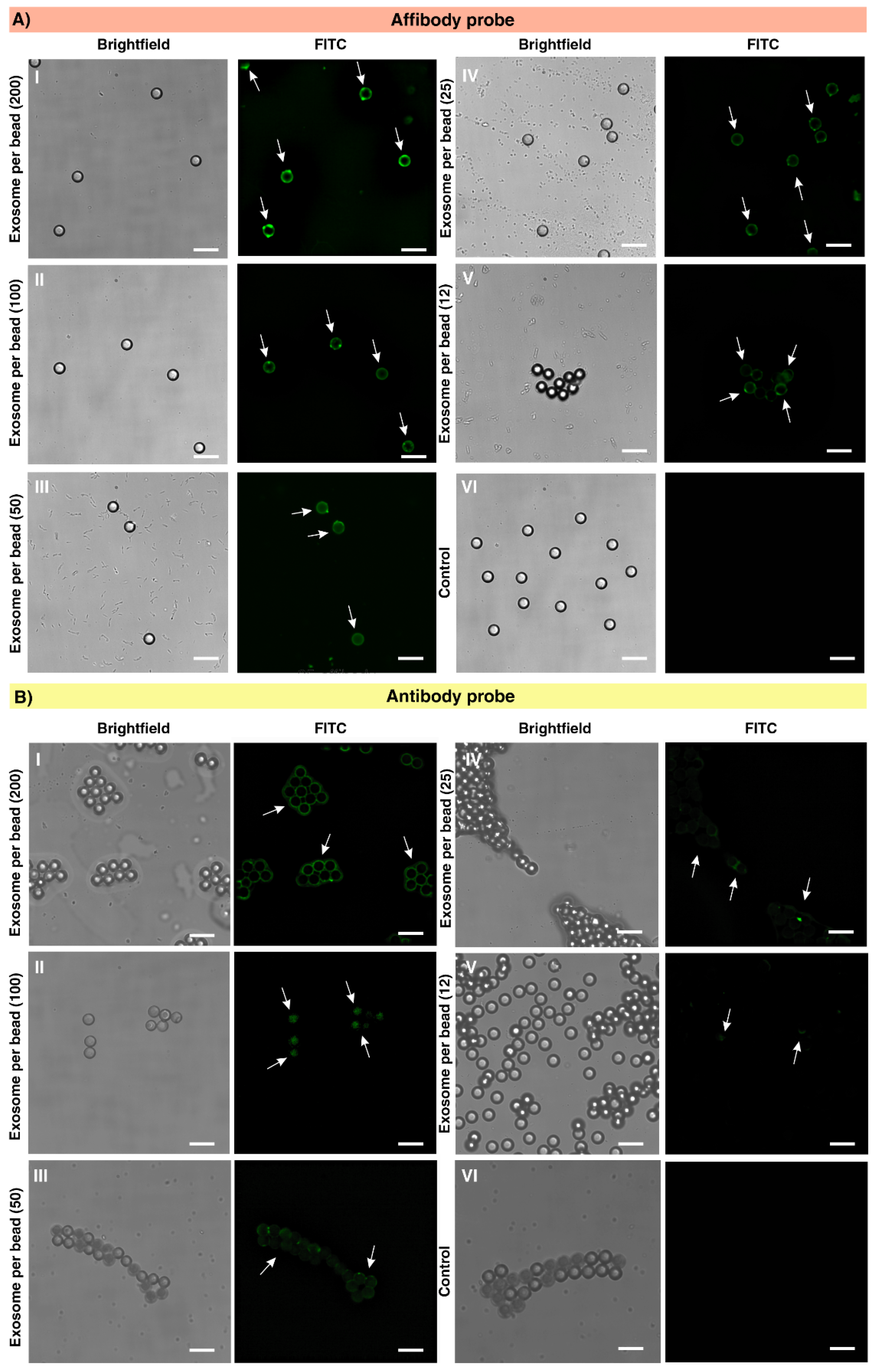
2.4. Exosome Capture Efficiency and Calculation of Limit of Detection (LOD)
3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Cell Culture
4.3. Immunofluorescence Staining of A549 Cells Using Anti-EGFR Antibody and Affibody
4.4. The Preparation of Conditioned Medium
4.5. Isolation of Extracellular Vesicles by Ultracentrifugation
4.6. Nanoparticle Tracking Analysis
4.7. Western Blot
4.8. Transmission Electron Microscopy (TEM)
4.9. Scanning Electron Microscope (SEM) Images of AffiBeads with Captured Exosomes
4.10. Surface Characterization
4.11. Capturing EGFR Positive Exosome Using AffiBeads
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kalluri, R.; LeBleu, V.S. The biology, function, and biomedical applications of exosomes. Science 2020, 367, eaau6977. [Google Scholar] [CrossRef]
- Wang, W.; Luo, J.; Wang, S. Recent progress in isolation and detection of extracellular vesicles for cancer diagnostics. Adv. Healthc. Mater. 2018, 7, 1800484. [Google Scholar] [CrossRef]
- Théry, C.; Ostrowski, M.; Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 2009, 9, 581–593. [Google Scholar] [CrossRef]
- Théry, C.; Zitvogel, L.; Amigorena, S. Exosomes: Composition, biogenesis and function. Nat. Rev. Immunol. 2002, 2, 569–579. [Google Scholar] [CrossRef]
- Vlassov, A.V.; Magdaleno, S.; Setterquist, R.; Conrad, R. Exosomes: Current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim. Biophys. Acta (BBA) Gen. Subj. 2012, 1820, 940–948. [Google Scholar] [CrossRef]
- Ko, J.; Carpenter, E.; Issadore, D. Detection and isolation of circulating exosomes and microvesicles for cancer monitoring and diagnostics using micro-/nano-based devices. Analyst 2016, 141, 450–460. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, C.; Jiang, F.; Ma, Y.; Wang, J.; Li, H.; Zhang, J. Isolation and Detection Technologies of Extracellular Vesicles and Application on Cancer Diagnostic. Dose-Response 2019, 17, 1559325819891004. [Google Scholar] [CrossRef] [Green Version]
- Gardiner, C.; Di Vizio, D.; Sahoo, S.; Théry, C.; Witwer, K.W.; Wauben, M.; Hill, A.F. Techniques used for the isolation and characterization of extracellular vesicles: Results of a worldwide survey. J. Extracell. Vesicles 2016, 5, 32945. [Google Scholar] [CrossRef] [PubMed]
- Rekker, K.; Saare, M.; Roost, A.M.; Kubo, A.-L.; Zarovni, N.; Chiesi, A.; Salumets, A.; Peters, M. Comparison of serum exosome isolation methods for microRNA profiling. Clin. Biochem. 2014, 47, 135–138. [Google Scholar] [CrossRef] [PubMed]
- de Menezes-Neto, A.; Sáez, M.J.F.; Lozano-Ramos, I.; Segui-Barber, J.; Martin-Jaular, L.; Ullate, J.M.E.; Fernandez-Becerra, C.; Borrás, F.E.; Del Portillo, H.A. Size-exclusion chromatography as a stand-alone methodology identifies novel markers in mass spectrometry analyses of plasma-derived vesicles from healthy individuals. J. Extracell. Vesicles 2015, 4, 27378. [Google Scholar] [CrossRef] [Green Version]
- Yoo, C.E.; Kim, G.; Kim, M.; Park, D.; Kang, H.J.; Lee, M.; Huh, N. A direct extraction method for microRNAs from exosomes captured by immunoaffinity beads. Anal. Biochem. 2012, 431, 96–98. [Google Scholar] [CrossRef]
- Gardiner, C.; Ferreira, Y.J.; Dragovic, R.A.; Redman, C.W.; Sargent, I.L. Extracellular vesicle sizing and enumeration by nanoparticle tracking analysis. J. Extracell. Vesicles 2013, 2, 19671. [Google Scholar] [CrossRef] [PubMed]
- Ueda, K.; Ishikawa, N.; Tatsuguchi, A.; Saichi, N.; Fujii, R.; Nakagawa, H. Antibody-coupled monolithic silica microtips for highthroughput molecular profiling of circulating exosomes. Sci. Rep. 2014, 4, 6232. [Google Scholar] [CrossRef] [PubMed]
- Arraud, N.; Gounou, C.; Turpin, D.; Brisson, A.R. Fluorescence triggering: A general strategy for enumerating and phenotyping extracellular vesicles by flow cytometry. Cytom. Part A 2016, 89, 184–195. [Google Scholar] [CrossRef]
- Zhang, Z.; Cheng, H.J. Recent development in sample preparation and analytical techniques for determination of quinolone residues in food products. Crit. Rev. Anal. Chem. 2017, 47, 223–250. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.; Ning, J.; Peng, D.; Chen, T.; Ahmad, I.; Ali, A.; Lei, Z.; Shabbir, M.A.B.; Cheng, G.; Yuan, Z. Current advances in immunoassays for the detection of antibiotics residues: A review. Food Agric. Immunol. 2020, 31, 268–290. [Google Scholar] [CrossRef]
- Liu, J.; Cui, D.; Jiang, Y.; Li, Y.; Liu, Z.; Tao, L.; Zhao, Q.; Diao, A. Selection and characterization of a novel affibody peptide and its application in a two-site ELISA for the detection of cancer biomarker alpha-fetoprotein. Int. J. Biol. Macromol. 2021, 166, 884–892. [Google Scholar] [CrossRef] [PubMed]
- Austerberry, J.; Dajani, R.; Panova, S.; Roberts, D.; Golovanov, A.P.; Pluen, A.; van der Walle, C.F.; Uddin, S.; Warwicker, J.; Derrick, J.P.; et al. The effect of charge mutations on the stability and aggregation of a human single chain Fv fragment. Eur. J. Pharm. Biopharm. 2017, 115, 18–30. [Google Scholar] [CrossRef] [Green Version]
- Frejd, F.Y.; Kim, K.-T. Affibody molecules as engineered protein drugs. Exp. Mol. Med. 2017, 49, e306. [Google Scholar] [CrossRef] [Green Version]
- Nord, K.; Gunneriusson, E.; Ringdahl, J.; Ståhl, S.; Uhlén, M.; Nygren, P.-Å. Binding proteins selected from combinatorial libraries of an α-helical bacterial receptor domain. Nat. Biotechnol. 1997, 15, 772–777. [Google Scholar] [CrossRef] [PubMed]
- Ståhl, S.; Gräslund, T.; Karlström, A.E.; Frejd, F.Y.; Nygren, P.-Å.; Löfblom, J. Affibody molecules in biotechnological and medical applications. Trends Biotechnol. 2017, 35, 691–712. [Google Scholar] [CrossRef] [PubMed]
- Löfblom, J.; Feldwisch, J.; Tolmachev, V.; Carlsson, J.; Ståhl, S.; Frejd, F.Y. Affibody molecules: Engineered proteins for therapeutic, diagnostic and biotechnological applications. FEBS Lett. 2010, 584, 2670–2680. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Case, B.A.; Kruziki, M.A.; Stern, L.A.; Hackel, B.J. Evaluation of affibody charge modification identified by synthetic consensus design in molecular PET imaging of epidermal growth factor receptor. Mol. Syst. Des. Eng. 2018, 3, 171–182. [Google Scholar] [CrossRef]
- Gebauer, M.; Skerra, A. Engineered protein scaffolds as Next-Generation therapeutics. Annu. Rev. Pharmacol. Toxicol. 2020, 60, 391–415. [Google Scholar] [CrossRef]
- Yarden, Y.; Sliwkowski, M.X. Untangling the ErbB signalling network. Nat. Rev. Mol. Cell Biol. 2001, 2, 127–137. [Google Scholar] [CrossRef]
- Al-Nedawi, K.; Meehan, B.; Kerbel, R.S.; Allison, A.C.; Rak, J. Endothelial expression of autocrine VEGF upon the uptake of tumor-derived microvesicles containing oncogenic EGFR. Proc. Natl. Acad. Sci. USA 2009, 106, 3794–3799. [Google Scholar] [CrossRef] [Green Version]
- Yamashita, T.; Kamada, H.; Kanasaki, S.; Maeda, Y.; Nagano, K.; Abe, Y.; Yoshioka, Y.; Tsutsumi, Y.; Katayama, S.; Inoue, M.; et al. Epidermal growth factor receptor localized to exosome membranes as a possible biomarker for lung cancer diagnosis. Die Pharm. Int. J. Pharm. Sci. 2013, 68, 969–973. [Google Scholar]
- Castellanos-Rizaldos, E.; Grimm, D.; Tadigotla, V.; Hurley, J.; Healy, J.; Neal, P.L.; Sher, M.; Venkatesan, R.; Karlovich, C.; Raponi, M.; et al. Exosome-Based Detection of EGFR T790M in Plasma from Non–Small Cell Lung Cancer Patients. Clin. Cancer Res. 2018, 24, 2944–2950. [Google Scholar] [CrossRef] [Green Version]
- Gong, H.; Kovar, J.; Little, G.; Chen, H.; Olive, D.M. In Vivo Imaging of Xenograft Tumors Using an Epidermal Growth Factor Receptor-Specific Affibody Molecule Labeled with a Near-infrared Fluorophore. Neoplasia 2010, 12, 139–149. [Google Scholar] [CrossRef] [Green Version]
- Higginbotham, J.N.; Zhang, Q.; Jeppesen, D.; Scott, A.; Manning, H.C.; Ochieng, J.; Franklin, J.L.; Coffey, R.J. Identification and characterization of EGF receptor in individual exosomes by fluorescence-activated vesicle sorting. J. Extracell. Vesicles 2016, 5, 29254. [Google Scholar] [CrossRef]
- Cavallaro, S.; Horak, J.; Hååg, P.; Gupta, D.; Stiller, C.; Sahu, S.S.; Görgens, A.; Gatty, H.K.; Viktorsson, K.; El-Andaloussi, S.; et al. Label-Free Surface Protein Profiling of Extracellular Vesicles by an Electrokinetic Sensor. ACS Sens. 2019, 4, 1399–1408. [Google Scholar] [CrossRef]
- Kaur, H.; Kumar, S.; Kukkar, D.; Kaur, I.; Singh, K.; Bharadwaj, L.M. Transportation of Drug-(Polystyrene Bead) Conjugate by Actomyosin Motor System. J. Biomed. Nanotechnol. 2010, 6, 279–286. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Zhang, D.; Mao, H.; Song, W.; Gao, G.; Liu, F. Preparation and characterization of Ag/AgO nanoshells on carboxylated polystyrene latex particles. J. Mater. Res. 2006, 21, 349–354. [Google Scholar] [CrossRef]
- Zhand, S.; Xiao, K.; Bazaz, S.R.; Zhu, Y.; Bordhan, P.; Jin, D.; Warkiani, M.E. Improving capture efficiency of human cancer cell derived exosomes with nanostructured metal organic framework functionalized beads. Appl. Mater. Today 2021, 23, 100994. [Google Scholar] [CrossRef]
- Campos-Silva, C.; Suárez, H.; Acevedo, R.J.; Linares-Espinós, E.; Martinez-Piñeiro, L.; Yáñez-Mó, M.; Valés-Gómez, M. High sensitivity detection of extracellular vesicles immune-captured from urine by conventional flow cytometry. Sci. Rep. 2019, 9, 2042. [Google Scholar] [CrossRef] [Green Version]
- Qi, R.; Zhao, Y.; Guo, Q.; Mi, X.; Cheng, M.; Hou, W.; Zheng, H.; Hua, B. Exosomes in the lung cancer microenvironment: Biological functions and potential use as clinical biomarkers. Cancer Cell Int. 2021, 21, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Lindborg, M.; Dubnovitsky, A.; Olesen, K.; Björkman, T.; Abrahmsén, L.; Feldwisch, J.; Härd, T. High-affinity binding to staphylococcal protein A by an engineered dimeric Affibody molecule. Protein Eng. Des. Sel. 2013, 26, 635–644. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoyer, W.; Grönwall, C.; Jonsson, A.; Ståhl, S.; Härd, T. Stabilization of a β-hairpin in monomeric Alzheimer’s amyloid-β peptide inhibits amyloid formation. Proc. Natl. Acad. Sci. USA 2008, 105, 5099–5104. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sörensen, J.; Velikyan, I.; Sandberg, D.; Wennborg, A.; Feldwisch, J.; Tolmachev, V.; Orlova, A.; Sandström, M.; Lubberink, M.; Olofsson, H.; et al. Measuring HER2-Receptor Expression In Metastatic Breast Cancer Using [68Ga]ABY-025 Affibody PET/CT. Theranostics 2016, 6, 262–271. [Google Scholar] [CrossRef]
- Zhou, Q.; Rahimian, A.; Son, K.; Shin, D.-S.; Patel, T.; Revzin, A. Development of an aptasensor for electrochemical detection of exosomes. Methods 2016, 97, 88–93. [Google Scholar] [CrossRef] [Green Version]
- Zhao, X.; Zhang, W.; Qiu, X.; Mei, Q.; Luo, Y.; Fu, W. Rapid and sensitive exosome detection with CRISPR/Cas12a. Anal. Bioanal. Chem. 2020, 412, 601–609. [Google Scholar] [CrossRef]
- Zeng, X.; Yang, Y.; Zhang, N.; Ji, D.; Gu, X.; Jornet, J.M.; Wu, Y.; Gan, Q. Plasmonic Interferometer Array Biochip as a New Mobile Medical Device for Cancer Detection. IEEE J. Sel. Top. Quantum Electron. 2019, 25, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Thakur, A.; Qiu, G.; Siu-Pang, N.; Guan, J.; Yue, J.; Lee, Y.; Wu, C.M.L. Direct detection of two different tumor-derived extracellular vesicles by SAM-AuNIs LSPR biosensor. Biosens. Bioelectron. 2017, 94, 400–407. [Google Scholar] [CrossRef]
- Tian, Y.-F.; Ning, C.-F.; He, F.; Yin, B.-C.; Ye, B.-C.J.A. Highly sensitive detection of exosomes by SERS using gold nanostar@ Raman reporter@ nanoshell structures modified with a bivalent cholesterol-labeled DNA anchor. Analyst 2018, 143, 4915–4922. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Zhang, L.; Ye, Y.; Li, Y.; Luan, X.; Liu, J.; Cheng, J.; Zhao, Y.; Li, M.; Huang, C. Microsphere mediated exosome isolation and ultra-sensitive detection on a dielectrophoresis integrated microfluidic device. Analyst 2021, 146, 5962–5972. [Google Scholar] [CrossRef]
- Bazaz, S.R.; Mashhadian, A.; Ehsani, A.; Saha, S.C.; Krüger, T.; Warkiani, M.E. Computational inertial microfluidics: A review. Lab Chip 2020, 20, 1023–1048. [Google Scholar] [CrossRef]
- Kanwar, S.S.; Dunlay, C.J.; Simeone, D.M.; Nagrath, S. Microfluidic device (ExoChip) for on-chip isolation, quantification and characterization of circulating exosomes. Lab Chip 2014, 14, 1891–1900. [Google Scholar] [CrossRef] [PubMed]
- Reátegui, E.; Van Der Vos, K.E.; Lai, C.P.; Zeinali, M.; Atai, N.A.; Aldikacti, B.; Floyd, F.P.; Khankhel, A.H.; Thapar, V.; Hochberg, F.H.; et al. Engineered nanointerfaces for microfluidic isolation and molecular profiling of tumor-specific extracellular vesicles. Nat. Commun. 2018, 9, 175. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sayyadi, N.; Zhand, S.; Razavi Bazaz, S.; Warkiani, M.E. Affibody Functionalized Beads for the Highly Sensitive Detection of Cancer Cell-Derived Exosomes. Int. J. Mol. Sci. 2021, 22, 12014. https://doi.org/10.3390/ijms222112014
Sayyadi N, Zhand S, Razavi Bazaz S, Warkiani ME. Affibody Functionalized Beads for the Highly Sensitive Detection of Cancer Cell-Derived Exosomes. International Journal of Molecular Sciences. 2021; 22(21):12014. https://doi.org/10.3390/ijms222112014
Chicago/Turabian StyleSayyadi, Nima, Sareh Zhand, Sajad Razavi Bazaz, and Majid Ebrahimi Warkiani. 2021. "Affibody Functionalized Beads for the Highly Sensitive Detection of Cancer Cell-Derived Exosomes" International Journal of Molecular Sciences 22, no. 21: 12014. https://doi.org/10.3390/ijms222112014
APA StyleSayyadi, N., Zhand, S., Razavi Bazaz, S., & Warkiani, M. E. (2021). Affibody Functionalized Beads for the Highly Sensitive Detection of Cancer Cell-Derived Exosomes. International Journal of Molecular Sciences, 22(21), 12014. https://doi.org/10.3390/ijms222112014








