Time-Dependent Image Restoration of Low-SNR Live-Cell Ca2 Fluorescence Microscopy Data
Abstract
:1. Introduction
2. Results
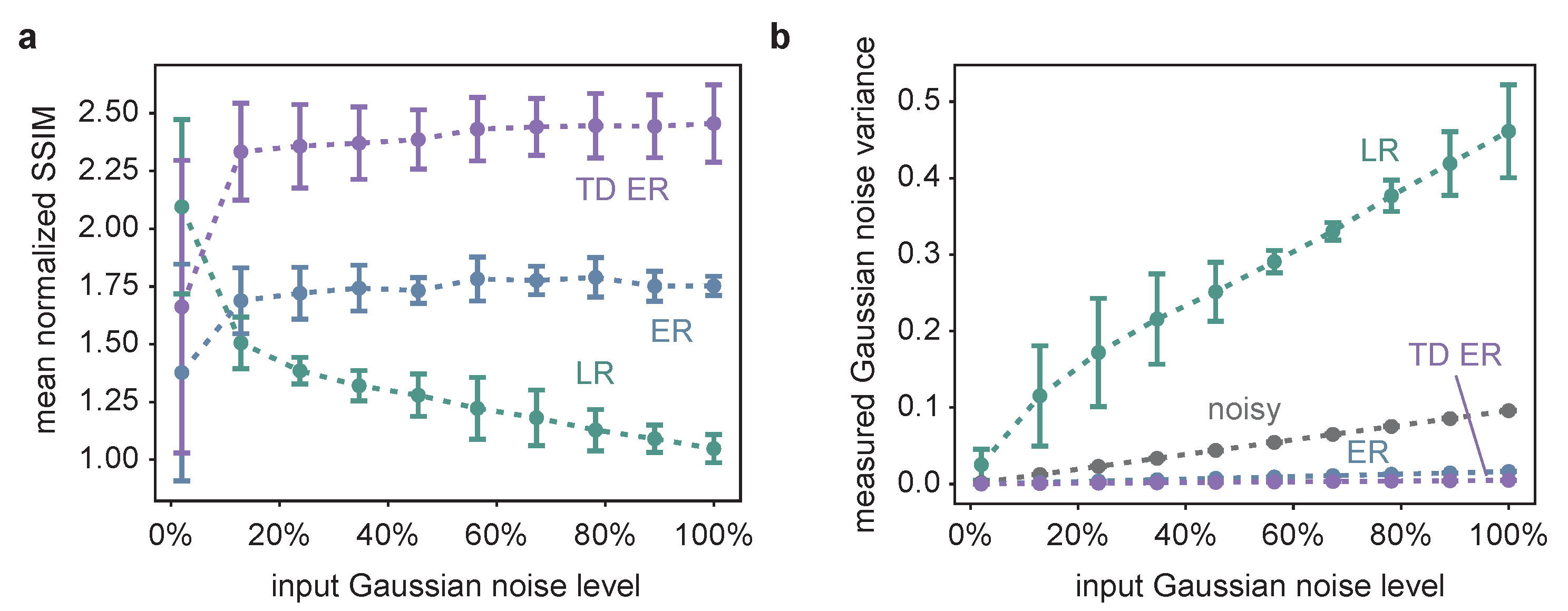
2.1. Synthetic Data
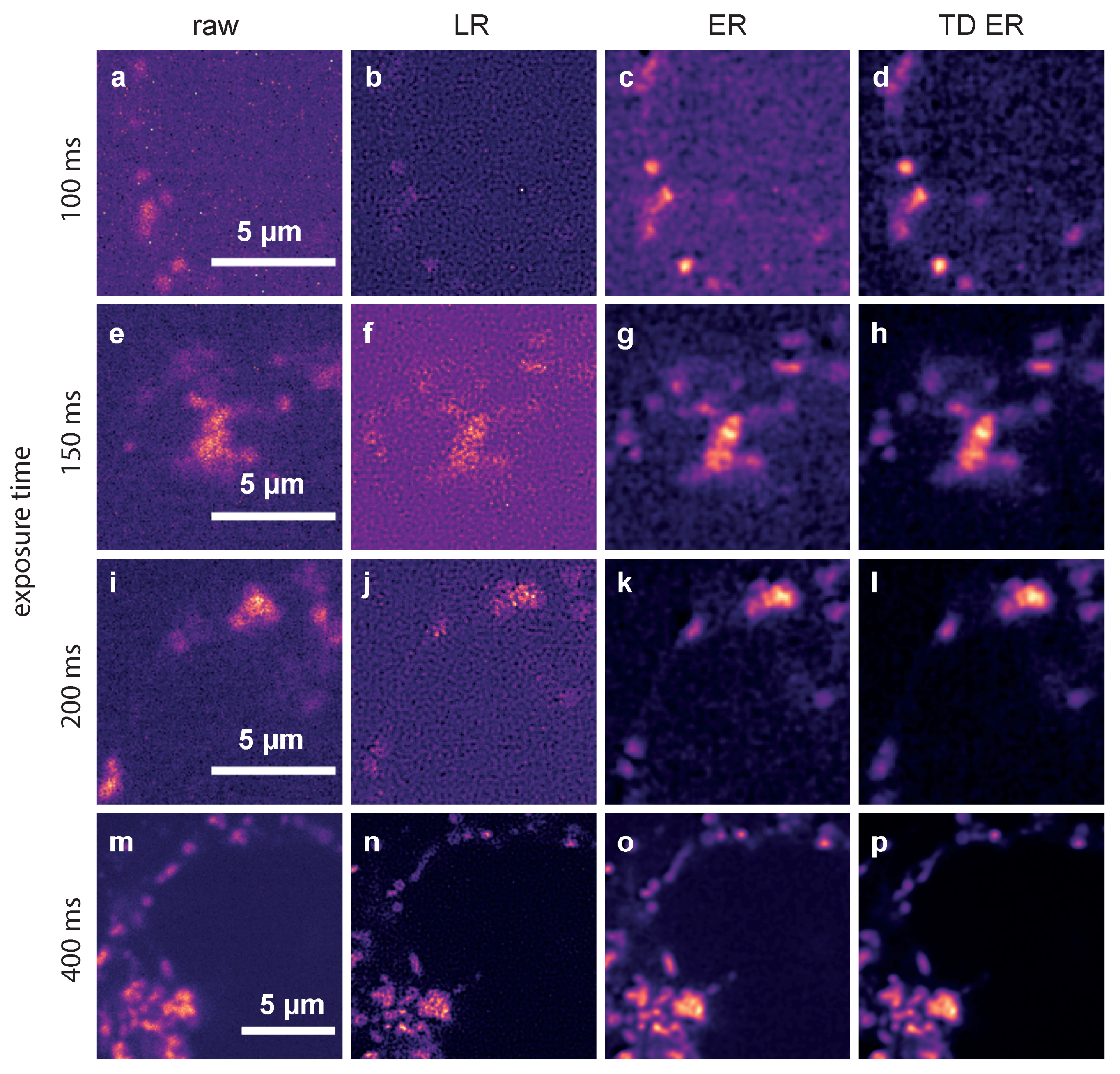
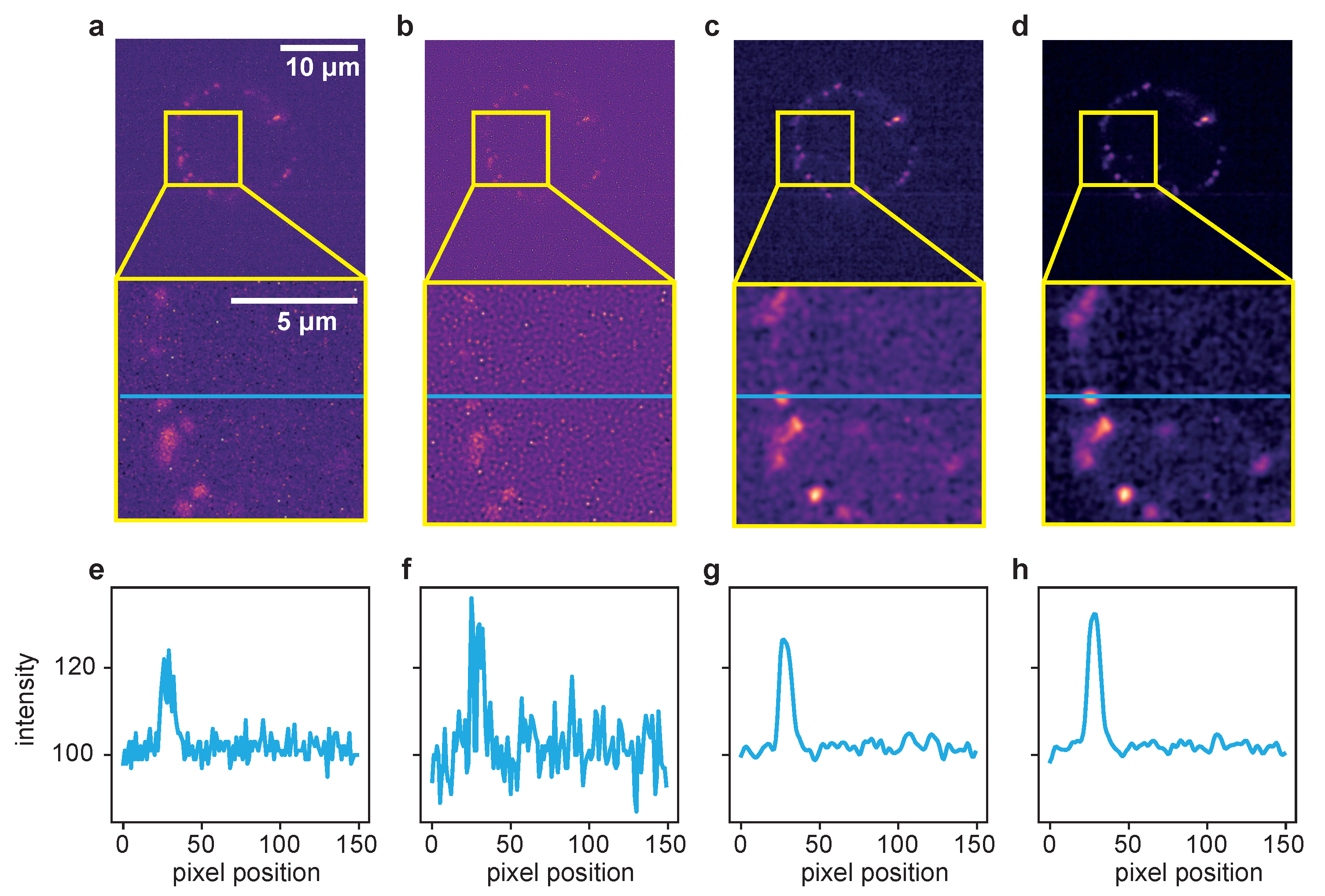
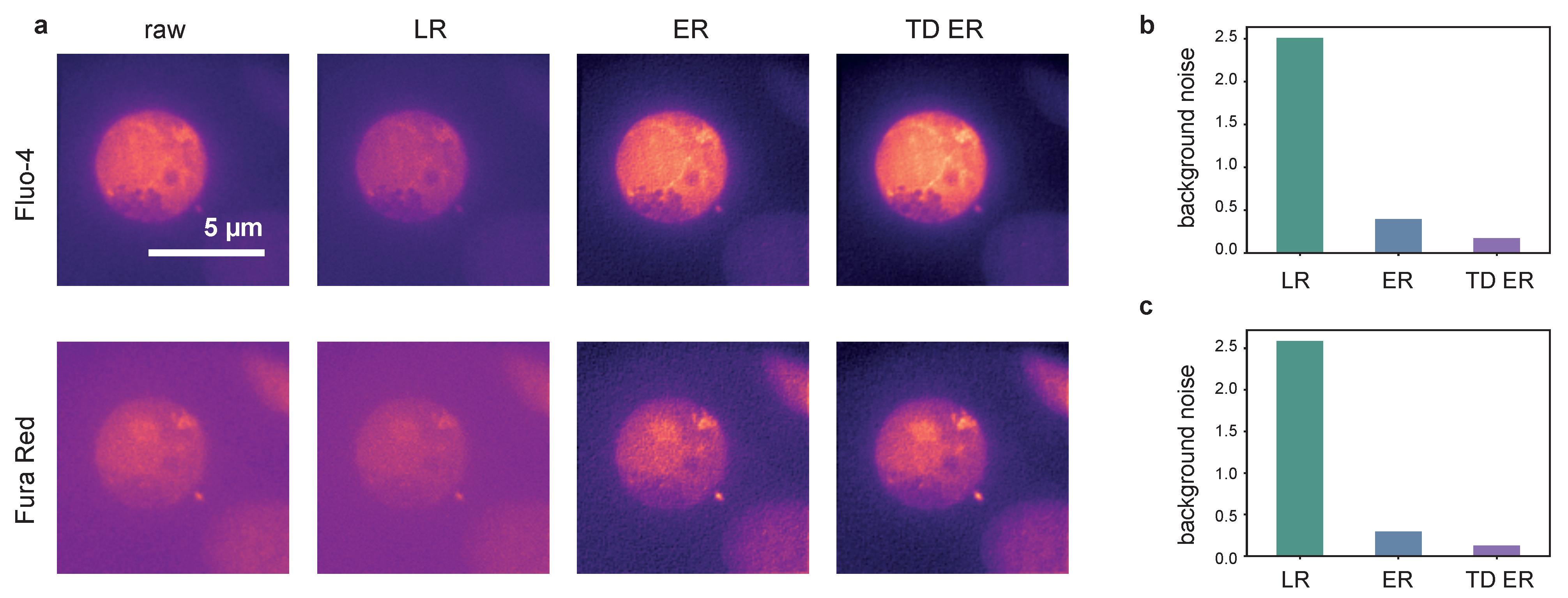
2.2. Live-Cell Fluorescence Microscopy
3. Discussion
4. Materials and Methods
4.1. Mathematical Formulation
4.2. Experiments: Imaging Data and Evaluation
4.2.1. Dataset 1: Synthetic Image Data
4.2.2. Dataset 2: Genetically Encoded Ca Indicator for Optimal Imaging (GECO) Tagged to Lysosomal TPC2 in Jurkat T-Cells
4.2.3. Dataset 3: Free Cytosolic Ca Concentration Imaging in Jurkat T-Cells
4.2.4. Dataset 4: Confocal Ca Imaging in Astrocytes In Situ
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| ER | static entropy deconvolution |
| GECO | genetically encoded Ca indicator for optimal imaging |
| LR | Lucy–Richardson |
| PSF | Point spread function |
| ROI | Region of interest |
| SNR | Signal-to-noise ratio |
| SSIM | Structural similarity index |
| TD ER | time-dependent entropy deconvolution |
| TPC | two pore channel |
Appendix A. Algorithm Details
Appendix A.1. Minimization of Cost Functional (uid13)
| Algorithm A1: Deconvolution. |
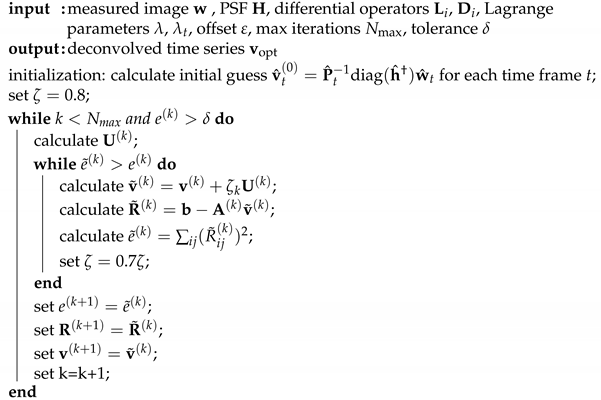 |
Appendix A.2. Practical Notes
References
- Wolf, I.M.A.; Diercks, B.P.; Gattkowski, E.; Czarniak, F.; Kempski, J.; Werner, R.; Schetelig, D.; Mittrücker, H.W.; Schumacher, V.; von Osten, M.; et al. Frontrunners of T cell activation: Initial, localized Ca2+ signals mediated by NAADP and the type 1 ryanodine receptor. Sci. Signal. 2015, 8, ra102. [Google Scholar] [CrossRef] [PubMed]
- Diercks, B.P.; Werner, R.; Weidemüller, P.; Czarniak, F.; Hernandez, L.; Lehmann, C.; Rosche, A.; Krüger, A.; Kaufmann, U.; Vaeth, M.; et al. ORAI1, STIM1/2, and RYR1 shape subsecond Ca2+ microdomains upon T cell activation. Sci. Signal. 2018, 11, eaat0358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ettinger, A.; Wittmann, T. Chapter 5—Fluorescence live cell imaging. In Methods in Cell Biology; Waters, J.C., Wittman, T., Eds.; Academic Press: Cambridge, MA, USA, 2014; Volume 123, pp. 77–94. [Google Scholar] [CrossRef] [Green Version]
- Fujiwara, K.; Tanaka, H.; Mani, H.; Nakagami, T.; Takamatsu, T. Burst Emergence of Intracellular Ca2+ Waves Evokes Arrhythmogenic Oscillatory Depolarization via the Na+–Ca2+ Exchanger. Circ. Res. 2008, 103, 509–518. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weigert, M.; Schmidt, U.; Boothe, T.; Müller, A.; Dibrov, A.; Jain, A.; Wilhelm, B.; Schmidt, D.; Broaddus, C.; Culley, S.; et al. Content-aware image restoration: Pushing the limits of fluorescence microscopy. Nat. Methods 2018, 15, 1090–1097. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.; Sasaki, H.; Lai, H.; Su, Y.; Liu, J.; Wu, Y.; Zhovmer, A.; Combs, C.A.; Rey-Suarez, I.; Chang, H.Y.; et al. Three-dimensional residual channel attention networks denoise and sharpen fluorescence microscopy image volumes. Nat. Methods 2021, 18, 678–687. [Google Scholar] [CrossRef] [PubMed]
- Belthangady, C.; Royer, L.A. Applications, promises, and pitfalls of deep learning for fluorescence image reconstruction. Nat. Methods 2019, 16, 1215–1225. [Google Scholar] [CrossRef] [PubMed]
- Wiener, N. Extrapolation, Interpolation, and Smoothing of Stationary Time Series: With Engineering Applications; MIT Press: Cambridge, MA, USA, 1964; Volume 8. [Google Scholar]
- Boyat, A.K.; Joshi, B.K. Image denoising using wavelet transform and wiener filter based on log energy distribution over Poisson-Gaussian noise model. In Proceedings of the 2014 IEEE International Conference on Computational Intelligence and Computing Research, Bhopal, India, 14–16 November 2014; pp. 1–6. [Google Scholar] [CrossRef]
- Richardson, W.H. Bayesian-Based Iterative Method of Image Restoration. J. Opt. Soc. Am. B 1972, 62, 55. [Google Scholar] [CrossRef]
- Lucy, L.B. An iterative technique for the rectification of observed distributions. Astron. J. 1974, 79, 745. [Google Scholar] [CrossRef] [Green Version]
- Sarder, P.; Nehorai, A. Deconvolution methods for 3-D fluorescence microscopy images. IEEE Signal Process. Mag. 2006, 23, 32–45. [Google Scholar] [CrossRef]
- Sage, D.; Donati, L.; Soulez, F.; Fortun, D.; Schmit, G.; Seitz, A.; Guiet, R.; Vonesch, C.; Unser, M. DeconvolutionLab2: An open-source software for deconvolution microscopy. Methods 2017, 115, 28–41. [Google Scholar] [CrossRef] [PubMed]
- Arigovindan, M.; Fung, J.C.; Elnatan, D.; Mennella, V.; Chan, Y.H.M.; Pollard, M.; Branlund, E.; Sedat, J.W.; Agard, D.A. High-resolution restoration of 3D structures from widefield images with extreme low signal-to-noise-ratio. Proc. Natl. Acad. Sci. USA 2013, 110, 17344–17349. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hugelier, S.; Eilers, P.H.C.; Devos, O.; Ruckebusch, C. Improved superresolution microscopy imaging by sparse deconvolution with an interframe penalty. J. Chemom. 2017, 31, e2847. [Google Scholar] [CrossRef]
- Ahmad, M.; Hugelier, S.; Vitale, R.; Eilers, P.; Ruckebusch, C. A criterion for automatic image deconvolution with L0-norm regularization. J. Chemom. 2020, 34, e3227. [Google Scholar] [CrossRef]
- Paul, P.; Duessmann, H.; Bernas, T.; Huber, H.; Kalamatianos, D. Automatic noise quantification for confocal fluorescence microscopy images. Comput. Med. Imaging Graph. 2010, 34, 426–434. [Google Scholar] [CrossRef] [PubMed]
- Dynes, J.L.; Amcheslavsky, A.; Cahalan, M.D. Genetically targeted single-channel optical recording reveals multiple Orai1 gating states and oscillations in calcium influx. Proc. Natl. Acad. Sci. USA 2016, 113, 440–445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lohr, C.; Beiersdorfer, A.; Fischer, T.; Hirnet, D.; Rotermund, N.; Sauer, J.; Schulz, K.; Gee, C.E. Using Genetically Encoded Calcium Indicators to Study Astrocyte Physiology: A Field Guide. Front. Cell. Neurosci. 2021, 15, 690147. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Woelk, L.-M.; Kannabiran , S.A.; Brock , V.J.; Gee , C.E.; Lohr , C.; Guse , A.H.; Diercks , B.-P.; Werner, R. Time-Dependent Image Restoration of Low-SNR Live-Cell Ca2 Fluorescence Microscopy Data. Int. J. Mol. Sci. 2021, 22, 11792. https://doi.org/10.3390/ijms222111792
Woelk L-M, Kannabiran SA, Brock VJ, Gee CE, Lohr C, Guse AH, Diercks B-P, Werner R. Time-Dependent Image Restoration of Low-SNR Live-Cell Ca2 Fluorescence Microscopy Data. International Journal of Molecular Sciences. 2021; 22(21):11792. https://doi.org/10.3390/ijms222111792
Chicago/Turabian StyleWoelk, Lena-Marie, Sukanya A. Kannabiran , Valerie J. Brock , Christine E. Gee , Christian Lohr , Andreas H. Guse , Björn-Philipp Diercks , and René Werner. 2021. "Time-Dependent Image Restoration of Low-SNR Live-Cell Ca2 Fluorescence Microscopy Data" International Journal of Molecular Sciences 22, no. 21: 11792. https://doi.org/10.3390/ijms222111792
APA StyleWoelk, L.-M., Kannabiran , S. A., Brock , V. J., Gee , C. E., Lohr , C., Guse , A. H., Diercks , B.-P., & Werner, R. (2021). Time-Dependent Image Restoration of Low-SNR Live-Cell Ca2 Fluorescence Microscopy Data. International Journal of Molecular Sciences, 22(21), 11792. https://doi.org/10.3390/ijms222111792






