Phage Therapy as a Focused Management Strategy in Aquaculture
Abstract
1. Phage Biology and Spatial Distribution
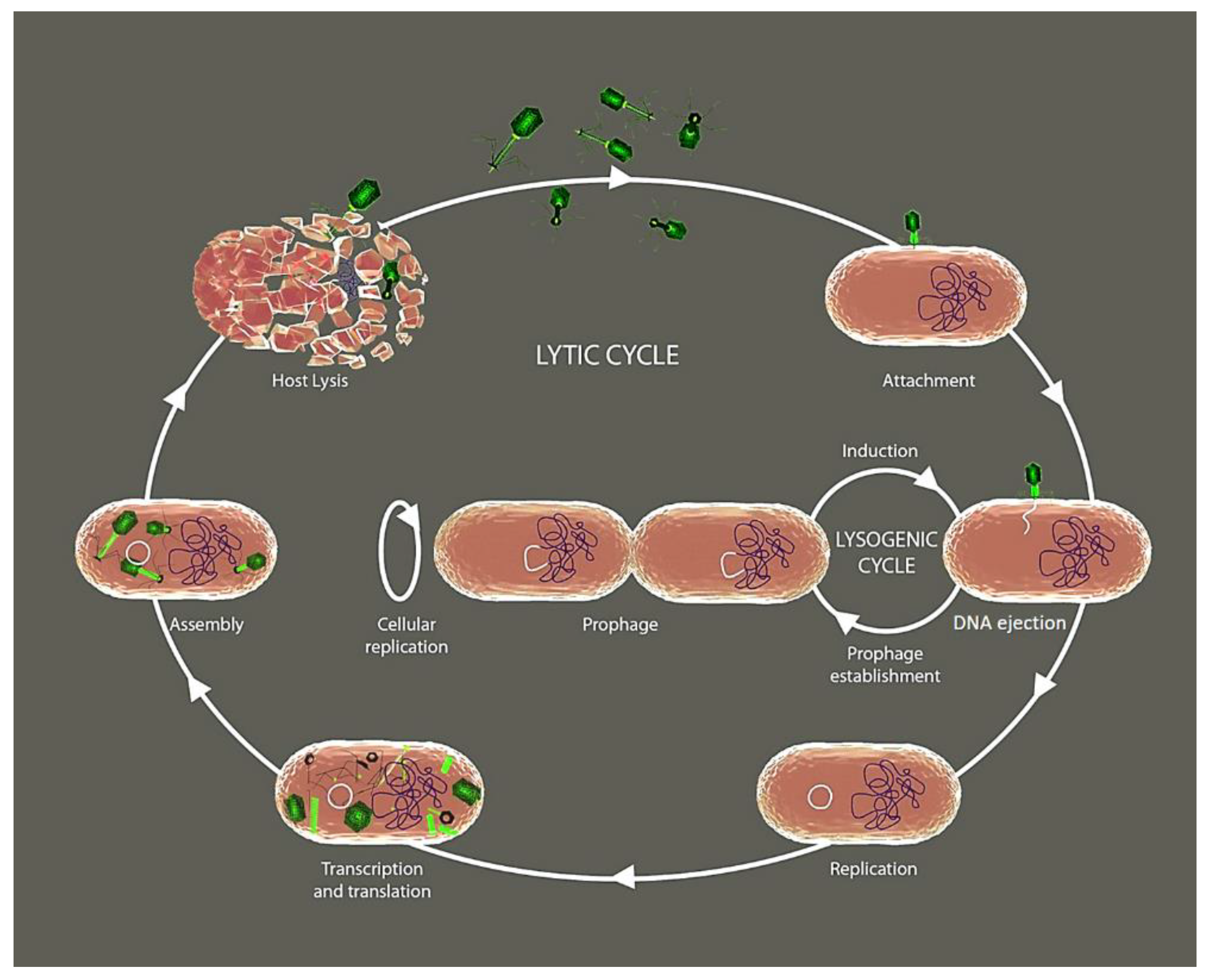
2. Phage’s Life Cycle
3. Phage Lytic Enzymes and Depolymerases
4. Interactions between Phage and the Fish Immune System
4.1. Phage-Mediated Activation of Inflammation
4.2. Phage-Specific Adaptive Responses
5. Phage Diversity in the Aquatic Environment
6. Bacteriophages against Biofilms in Aquaculture Facilities
| Gram-Negative Targets | Source | Enrichment ɸ | Characterization Method | Phage Strains Name | Family * | Genome Length | References |
|---|---|---|---|---|---|---|---|
| Aeromonas hydrophila | River water | No | TEM | ɸ2 and ɸ5 | Myoviridae | ~20 kb | [100] |
| Fishponds; Polluted rivers | Single | TEM | N21, W3, G65, Y71 and Y81 | Myoviridae; Podoviridae | n.d. | [101] | |
| Stream water | Single | TEM, dsDNA | pAh-1 | Myoviridae | ~64 kb | [102] | |
| Sea water | Single | TEM, DNA sequencing | Akh-2 | Siphoviridae | 114,901 bp | [103] | |
| Carp tissues | Single | TEM | AHP-1 | Myoviridae | n.d. | [104] | |
| Lake water | Single | TEM, dsDNA, DNA sequencing | AhyVDH1 | Myxoviridae | 39,175 bp | [105] | |
| River water | No | TEM, dsDNA, DNA sequencing | MJG | Podoviridae | 45,057 bp | [106] | |
| Sewage water | Single | TEM | AH1 | n.d. | n.d. | [107] | |
| Striped catfish pond water | Single | TEM, dsDNA, DNA sequencing | PVN02 | Myoviridae | 51,668 bp | [108,109] | |
| River water | TEM, dsDNA | pAh1-C pAh6-C | Myoviridae | 55 kb 58 kb | [110] | ||
| Wastewater | No | TEM, dsDNA, DNA sequencing | Ahp1 | Podoviridae | ~42 kb | [111] | |
| Aeromonas punctata | Stream water | Single | TEM, dsDNA | IHQ1 | Myoviridae | 25–28 kb | [112] |
| Aeromonas salmonicida | River waters, two passing through fish farms | Single | TEM, DNA sequencing | SW69-9 L9-6 Riv-10 | Myoviridae | 173,097 bp, 173,578 bp and 174,311 bp | [113] |
| River water | Single | TEM, DNA sequencing | phiAS5 | Myoviridae | 225,268 bp | [114] | |
| Sediment of a Rainbow trout culture farm | Single | TEM, dsDNA, DNA sequencing | PAS-1 | Myoviridae | ~48 kb | [115] | |
| Wastewater from a seafood market | No | TEM, DNA sequencing | AsXd-1 | Siphoviridae | 39,014 bp | [116] | |
| Sewage network water from a lift station | Single | TEM | AS-A AS-D AS-E | Myoviridae | n.d. | [117,118] | |
| River water | No | TEM | HER 110 | Myoviridae | n.d. | [119,120] | |
| Aeromonas spp. | Gastrointestinal content of variated fish species | No | TEM, DNA sequencing | phiA8-29 | Myoviridae | 144,974 bp | [66,121] |
| Citrobacter freundii | Sewage water | No | TEM, DNA sequencing | IME-JL8 | Siphoviridae | 49,838 bp | [122] |
| Edwardsiella ictaluri | Water from catfish ponds | Single | TEM, dsDNA, DNA sequencing | eiAU eiDWF eiMSLS | Siphoviridae | 42.80 kbp 42.12 kbp 42.69 kbp | [123,124] |
| River water | Multiple | DNA Sequencing | PEi21 | Myoviridae | 43,378 bp | [125,126] | |
| Striped catfish kidney and liver | Single | TEM, dsDNA | MK7 | Myoviridae | ~34 kb | [127] | |
| Edwardsiella tarda | Seawater | Single | TEM, dsDNA | ETP-1 | Podoviridae | ~40 kb | [54] |
| River water | No | TEM, DNA sequencing | pEt-SU | Myoviridae | 276,734 bp | [128] | |
| Wastewater | Single | DNA sequencing | PETp9 | Myoviridae | 89,762 bp | [129] | |
| Fish tissues and rearing seawater | No | TEM, DNA sequencing | GF-2 | Myoviridae | 43,129 bp | [130] | |
| Flavobacterium columnare | River water | Single | TEM, DNA sequencing | FCL-2 | Myoviridae | 47,142 bp | [38,131,132] |
| Fishpond’s water and bottom sediments | No | TEM, dsDNA | FCP1-FCP9 | Podoviridae | n.d. | [133] | |
| Flavobacterium psychrophilum | Rainbow trout farm water | Single/double | TEM, dsDNA | ø (FpV-1 to FpV-22) | Podoviridae Siphoviridae Myoviridae | (~8 to ~90 kb) | [134,135] |
| Ayu kidneys and pondwater collected from ayu farms | Multiple | TEM, dsDNA | PFpW-3, PFpC-Y PFpW-6, PFpW-7 PFpW-8 | Myoviridae; Podoviridae; Siphoviridae | n.d. | [136] | |
| Photobacterium damselae subsp. damselae | Raw oysters | Single | TEM, dsDNA | Phda1 | Myoviridae | 35.2–39.5 kb | [137] |
| Gastrointestinal tract of lollipop catshark | Single | TEM, DNA sequencing | vB_Pd_PDCC-1 | Myoviridae | 237,509 bp | [138] | |
| Pseudomonas plecoglossicida | Ayu pond water and diseased fish | No | TEM, DNA sequencing | PPpW-3 PPpW-4 | Myoviridae Podoviridae | 43,564 bp 41,386 bp | [139,140] |
| Pseudomonas aeruginosa | Wastewater | No | TEM, DNA sequencing | MBL | n.d. | 42,519 bp | [141] |
| Shewanella spp. | Wastewater from a marketplace | Single | TEM, DNA sequencing | SppYZU01 to SppYZU10 | Myoviridae; Siphoviridae. | SppYZU01 (43.567 bp) SppYZU5 (54.319 bp) | [142] |
| Tenacibaculum maritimum | Seawater | Multiple | TEM, DNA sequencing | PTm1 PTm5 | Myoviridae | 224,680 bp 226,876 bp | [143] |
| Vibrio alginolyticus | Aquaculture tank water | Single | TEM, DNA sequencing | VEN | Podoviridae | 44,603 bp | [144] |
| Marine sediment | No | TEM, DNA sequencing | ValKK3 | Myoviridae | 248,088 bp | [145] | |
| Marine water | Single | TEM, dsDNA | St2 Grn1 | Myoviridae | 250,485 bp 248,605 bp | [146] | |
| Vibrio anguillarum | Soft tissues from clams and mussels | No | TEM, dsDNA | 309 ALMED CHOED ALME CHOD CHOB | Several shapes | ~47–48 kb | [147] |
| Sewage water | Double | dsDNA | VP-2 VA-1 | n.d. | n.d. | [148] | |
| Water samples from fish farms | Multiple | TEM, DNA sequencing | ø H1, H7, S4-7, H4, H5 H8, H20 S4-18, 2E-1, H2 | Myoviridae Siphoviridae Podoviridae | ~194–195 kb ~50 kb ~45–51 kb | [149] | |
| Vibrio campbellii | Host strain (V. campbellii) isolated form a dead shrimp | No | TEM, DNA sequencing | HY01 | Siphoviridae | 41.772 bp | [150] |
| Hepatopancreas of Pacific white shrimp | Single | dsDNA, DNA sequencing | vB_Vc_SrVc9 | Autographiviridae | ~43.15 kb | [151] | |
| Vibrio harveyi | Shrimp farm, hatcheries and marine water | Multiple | TEM, dsDNA | A | Siphoviridae | n.d. | [152] |
| Vibrio harveyi | No | TEM, dsDNA | VHML | Myovirus-like | n.d. | [153] | |
| Shrimp pond water | Single | TEM, dsDNA | PW2 | Siphoviridae | ~46 kb | [154] | |
| Water and sediment samples | Single | TEM, dsDNA | VHM1, VHM2 VHS1 | Myoviridae, Siphoviridae | ~55 kb, ~66 kb ~69 kb | [155] | |
| Hatchery water and oyster tissues | Single | TEM, dsDNA | vB_VhaS-a vB_VhaS-tm | Siphoviridae | ~82 kb ~59 kb | [156] | |
| Commercial clam samples | Multiple | Genomic analysis, dsDNA | ø VhCCS-01 VhCCS-02 VhCCS-04 VhCCS-06 VhCCS-17 VhCCS-20 VhCCS-19 VhCCS-21 | Siphoviridae, Myoviridae | n.d. | [157] | |
| Oyster, clam, shrimp, and seawater samples | No | TEM, DNA sequencing | VHP6b | Siphoviridae | 78,081 bp | [158] | |
| shrimp hatchery and farm water, oysters from estuaries, coastal sea water | Multiple | TEM, dsDNA | Viha10 Viha8 Viha9 Viha11 Viha1 to Viha7 | Siphoviridae - Siphoviridae Myoviridae (Viha4) | n.d. ~44–94 kb ~85 kb (Viha4) | [159,160] | |
| Seawater sample | Single | TEM | VhKM4 | Myoviridae | n.d. | [161] | |
| Vibrio ordalii | Macerated specimens of mussels | No | TEM, DNA sequencing | B_VorS-PVo5 | Siphoviridae | 80,578 bp | [162] |
| Vibrio parahaemolyticus | Sewage sample | No | TEM, dsDNA | VPp1 | Tectiviridae | ~15 kb | [163] |
| Polluted seawater | No | TEM, dsDNA | KVP40 KVP41 | Myoviridae | n.d. | [164,165] | |
| Seawater or mussels | Single | dsDNA | SPA2 SPA3 | n.d. | ~21 kb | [166] | |
| Coastal water | Single | TEM, DNA sequencing | pVP-1 | Siphoviridae | 111,506 bp | [167,168] | |
| V. parahaemolyticus isolated from sewage samples collected from an aquatic product market | No | TEM, DNA sequencing | vB_VpS_BA3 vB_VpS_CA8 | Siphoviridae | 58,648 bp 58,480 bp | [169] | |
| Shrimp pond water | Single | TEM, DNA sequencing | VP-1 | Myoviridae | 150,764 bp | [170] | |
| Coastal sand sediment | double | TEM, DNA sequencing | VpKK5 | Siphoviridae | 56,637 bp | [171,172] | |
| Vibrio splendidus | Raw sewage obtained from local hatcheries | Single | TEM | PVS-1, PVS-2 PVS-3 | Myoviridae; Siphoviridae | n.d. | [173] |
| Seawater near a fish farm cage | Single | TEM, DNA sequencing | vB_VspP_pVa5 | Podoviridae | 78,145 bp | [174] | |
| Vibrio coralliilyticus | sewage in oyster hatchery | Single | TEM | pVco-14 | Siphoviridae | n.d. | [175] |
| Vibrio vulnificus | Seawater sample | Single | TEM, DNA sequencing | SSP002 | Siphoviridae | 76,350 bp | [176,177] |
| Abalone samples | No | TEM, sequencing | VVPoo1 | Siphoviridae | 76,423 bp | [178] | |
| Initial host strain (V. vulnificus) | No | TEM | VV1 VV2 VV3 VV4 | Tectiviridae | n.d. | [179] | |
| Vibrio sp. | Sewage draining exits | Single | TEM, DNA sequencing | VspDsh-1 VpaJT-1 ValLY-3 ValSw4-1 VspSw-1 | Siphoviridae | 46,692 bp 60,177 bp 76,310 bp 79,545 bp 113,778 bp | [180] |
| Yersinia ruckeri | Wastewater containing suspended trout feces from a settling pond at a trout farm | Single | TEM | NC10 | Podoviridae | n.d. | [181] |
| Sewage | No | TEM | YerA41 (several phages) | icosahedral head, contractile tail | n.d. | [182] | |
| Sewage | No | TEM, DNA sequencing, dsDNA | R1-37 | Myoviridae | ~270 kb | [183,184] |
| Gram-Positive Targets | Source | Enrichment ɸ | Characterization Method | Phage Strains Name | Family * | Genome Length | References |
|---|---|---|---|---|---|---|---|
| Lactococcus garvieae | L. garvieae isolated from diseased yellowtail | No | TEM, dsDNA | PLgY(16) | Siphoviridae | n.d. | [185] |
| Yellowtail (Y) Water (W) Sediments (S) | Single | TEM, dsDNA | PLgW1-6 PLgY16 PLgY30 PLgY886 PLgS1 | Siphoviridae | >20 kbp | [186,187,188] | |
| Domestic compost | Single | TEM, DNA sequencing | GE1 | Siphoviridae | 24,847 bp | [189] | |
| L. garvieae host | No | TEM, DNA sequencing | PLgT-1 | Siphoviridae | 29,284 bp | [190,191,192] | |
| Rainbow trout farm water | Single | TEM, DNA sequencing | WP-2 | Picovirinae | 18,899 bp | [193] | |
| Streptococcus agalactiae | Tilapia pond | No | TEM | HN48 | Caudoviridae | n.d. | [194] |
| S. iniae | S. iniae host | No | TEM, dsDNA | vB_SinS-44 vB_SinS-45 vB_SinS-46 vB_SinS-48 | Siphoviridae | ~51.7 kb ~28.4 kb ~66.3 kb ~27.5 kb | [195] |
| Weissella ceti | W. ceti host strain | No | TEM | PWc | Siphoviridae | 38,783 bp | [196] |
7. Potential of Phage Therapy in Aquaculture Settings
8. Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Conflicts of Interest
References
- Ghosh, C.; Sarkar, P.; Issa, R.; Haldar, J. Alternatives to Conventional Antibiotics in the Era of Antimicrobial Resistance. Trends Microbiol. 2019, 27, 323–338. [Google Scholar] [CrossRef]
- Golkar, Z.; Bagasra, O.; Pace, D.G. Bacteriophage therapy: A potential solution for the antibiotic resistance crisis. J. Infect. Dev. Ctries 2014, 8, 129–136. [Google Scholar] [CrossRef]
- Wigington, C.H.; Sonderegger, D.; Brussaard, C.P.; Buchan, A.; Finke, J.F.; Fuhrman, J.A.; Lennon, J.T.; Middelboe, M.; Suttle, C.A.; Stock, C.; et al. Re-examination of the relationship between marine virus and microbial cell abundances. Nat. Microbiol. 2016, 1, 15024. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Li, Y.; Yan, W.; Wang, Y.; Cai, L.; Luo, T.; Li, H.; Weinbauer, M.G.; Jiao, N. Viral control of biomass and diversity of bacterioplankton in the deep sea. Commun. Biol. 2020, 3, 256. [Google Scholar] [CrossRef] [PubMed]
- Brum, J.R.; Schenck, R.O.; Sullivan, M.B. Global morphological analysis of marine viruses shows minimal regional variation and dominance of non-tailed viruses. ISME J. 2013, 7, 1738–1751. [Google Scholar] [CrossRef]
- Dion, M.B.; Oechslin, F.; Moineau, S. Phage diversity, genomics and phylogeny. Nat. Rev. Microbiol. 2020, 18, 125–138. [Google Scholar] [CrossRef] [PubMed]
- Washizaki, A.; Yonesaki, T.; Otsuka, Y. Characterization of the interactions between Escherichia coli receptors, LPS and OmpC, and bacteriophage T4 long tail fibers. Microbiologyopen 2016, 5, 1003–1015. [Google Scholar] [CrossRef] [PubMed]
- Dunne, M.; Hupfeld, M.; Klumpp, J.; Loessner, M.J. Molecular Basis of Bacterial Host Interactions by Gram-Positive Targeting Bacteriophages. Viruses 2018, 10, 397. [Google Scholar] [CrossRef] [PubMed]
- Bertozzi Silva, J.; Storms, Z.; Sauvageau, D. Host receptors for bacteriophage adsorption. FEMS Microbiol. Lett. 2016, 363, fnw002. [Google Scholar] [CrossRef]
- Los, M.; Wegrzyn, G. Pseudolysogeny. Adv. Virus Res. 2012, 82, 339–349. [Google Scholar] [PubMed]
- Fischetti, V.A. Development of Phage Lysins as Novel Therapeutics: A Historical Perspective. Viruses 2018, 10, 310. [Google Scholar] [CrossRef]
- Trudil, D. Phage lytic enzymes: A history. Virol. Sin. 2015, 30, 26–32. [Google Scholar] [CrossRef]
- Coffey, B.; Mills, S.; Coffey, A.; McAuliffe, O.; Ross, R.P. Phage and Their Lysins as Biocontrol Agents for Food Safety Applications. Annu. Rev. Food Sci. Technol. 2010, 1, 449–468. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, C.; Costa, A.R.; Silva, F.; Oliveira, A. Bacteriophages and their derivatives for the treatment and control of food-producing animal infections. Crit. Rev. Microbiol. 2017, 43, 583–601. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Rubio, L.; Gutiérrez, D.; Donovan, D.M.; Martínez, B.; Rodríguez, A.; García, P. Phage lytic proteins: Biotechnological applications beyond clinical antimicrobials. Crit. Rev. Biotechnol. 2016, 36, 542–552. [Google Scholar] [CrossRef]
- Vázquez, R.; García, E.; García, P. Phage Lysins for Fighting Bacterial Respiratory Infections: A New Generation of Antimicrobials. Front. Immunol. 2018, 9, 2252. [Google Scholar] [CrossRef] [PubMed]
- Broendum, S.S.; Buckle, A.M.; McGowan, S. Catalytic diversity and cell wall binding repeats in the phage-encoded endolysins. Mol. Microbiol. 2018, 110, 879–896. [Google Scholar] [CrossRef]
- Cahill, J.; Young, R. Phage Lysis: Multiple Genes for Multiple Barriers. Adv. Virus Res. 2019, 103, 33–70. [Google Scholar] [PubMed]
- Rodriguez-Rubio, L.; Martínez, B.; Donovan, D.M.; Rodriguez, A.; García, P. Bacteriophage virion-associated peptidoglycan hydrolases: Potential new enzybiotics. Crit. Rev. Microbiol. 2013, 39, 427–434. [Google Scholar] [CrossRef] [PubMed]
- Ghose, C.; Euler, C.W. Gram-Negative Bacterial Lysins. Antibiotics 2020, 9, 74. [Google Scholar] [CrossRef]
- Oliveira, A.; Leite, M.; Kluskens, L.D.; Santos, S.B.; Melo, L.D.; Azeredo, J. The First Paenibacillus larvae Bacteriophage Endolysin (PlyPl23) with High Potential to Control American Foulbrood. PLoS ONE 2015, 10, e0132095. [Google Scholar]
- Fenton, M.; Ross, P.; Mcauliffe, O.; O’Mahony, J.; Coffey, A. Recombinant bacteriophage lysins as antibacterials. Bioeng. Bugs 2010, 1, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Fischetti, V.A. Bacteriophage endolysins: A novel anti-infective to control Gram-positive pathogens. Int. J. Med. Microbiol. 2010, 300, 357–362. [Google Scholar] [CrossRef] [PubMed]
- Sharma, U.; Vipra, A.; Channabasappa, S. Phage-derived lysins as potential agents for eradicating biofilms and persisters. Drug Discov. Today 2018, 23, 848–856. [Google Scholar] [CrossRef]
- Chan, B.K.; Abedon, S.T. Bacteriophages and Their Enzymes in Biofilm Control. Curr. Pharm. Des. 2015, 21, 85–99. [Google Scholar] [CrossRef]
- Meng, X.; Shi, Y.; Ji, W.; Zhang, J.; Wang, H.; Lu, C.; Sun, J.; Yan, Y. Application of a bacteriophage lysin to disrupt biofilms formed by the animal pathogen Streptococcus suis. Appl. Environ. Microbiol. 2011, 77, 8272–8279. [Google Scholar] [CrossRef] [PubMed]
- Gerstmans, H.; Grimon, D.; Gutierrez, D.; Lood, C.; Rodriguez, A.; van Noort, V.; Lammertyn, J.; Lavigne, R.; Briers, Y. A VersaTile-driven platform for rapid hit-to-lead development of engineered lysins. Sci. Adv. 2020, 6, eaaz1136. [Google Scholar] [CrossRef]
- Gerstmans, H.; Rodriguez-Rubio, L.; Lavigne, R.; Briers, Y. From endolysins to Artilysin(R)s: Novel enzyme-based approaches to kill drug-resistant bacteria. Biochem. Soc. Trans. 2016, 44, 123–128. [Google Scholar] [CrossRef]
- Wang, Z.; Kong, L.; Liu, Y.; Fu, Q.; Cui, Z.; Wang, J.; Ma, J.; Wang, H.; Yan, Y.; Sun, J. A Phage Lysin Fused to a Cell-Penetrating Peptide Kills Intracellular Methicillin-Resistant Staphylococcus aureus in Keratinocytes and Has Potential as a Treatment for Skin Infections in Mice. Appl. Environ. Microbiol. 2018, 84, e00380-18. [Google Scholar] [CrossRef]
- Yan, G.; Liu, J.; Ma, Q.; Zhu, R.; Guo, Z.; Gao, C.; Wang, S.; Yu, L.; Gu, J.; Hu, D.; et al. The N-terminal and central domain of colicin A enables phage lysin to lyse Escherichia coli extracellularly. Antonie Van Leeuwenhoek 2017, 110, 1627–1635. [Google Scholar] [CrossRef]
- Cheleuitte-Nieves, C.; Heselpoth, R.D.; Westblade, L.F.; Lipman, N.S.; Fischetti, V.A. Searching for a Bacteriophage Lysin to Treat Corynebacterium bovis in Immunocompromised Mice. Comp. Med. 2020, 70, 328–335. [Google Scholar] [CrossRef]
- Heselpoth, R.D.; Euler, C.W.; Schuch, R.; Fischetti, V.A. Lysocins: Bioengineered Antimicrobials That Deliver Lysins across the Outer Membrane of Gram-Negative Bacteria. Antimicrob. Agents Chemother. 2019, 63, e00342-19. [Google Scholar] [CrossRef]
- Pires, D.P.; Oliveira, H.; Melo, L.D.; Sillankorva, S.; Azeredo, J. Bacteriophage-encoded depolymerases: Their diversity and biotechnological applications. Appl. Microbiol. Biotechnol. 2016, 100, 2141–2151. [Google Scholar] [CrossRef]
- Volozhantsev, N.V.; Shpirt, A.M.; Borzilov, A.I.; Komisarova, E.V.; Krasilnikova, V.M.; Shashkov, A.S.; Verevkin, V.V.; Knirel, Y.A. Characterization and Therapeutic Potential of Bacteriophage-Encoded Polysaccharide Depolymerases with β Galactosidase Activity against Klebsiella pneumoniae K57 Capsular Type. Antibiotics 2020, 9, 732. [Google Scholar] [CrossRef]
- Lin, H.; Paff, M.L.; Molineux, I.J.; Bull, J.J. Antibiotic Therapy Using Phage Depolymerases: Robustness across a Range of Conditions. Viruses 2018, 10, 622. [Google Scholar] [CrossRef] [PubMed]
- Dabrowska, K.; Switała-Jelen, K.; Opolski, A.; Weber-Dabrowska, B.; Gorski, A. Bacteriophage Penetration in Vertebrates. J. Appl. Microbiol. 2005, 98, 7–13. [Google Scholar] [CrossRef]
- Barr, J.J.; Auro, R.; Furlan, M.; Whiteson, K.L.; Erb, M.L.; Pogliano, J.; Stotland, A.; Wolkowicz, R.; Cutting, A.S.; Doran, K.S. Bacteriophage Adhering to Mucus Provide a Non–Host-Derived Immunity. Proc. Natl. Acad. Sci. USA 2013, 110, 10771–10776. [Google Scholar] [CrossRef] [PubMed]
- Almeida, G.M.; Laanto, E.; Ashrafi, R.; Sundberg, L.-R. Bacteriophage Adherence to Mucus Mediates Preventive Protection against Pathogenic Bacteria. MBio 2019, 10, e01984-19. [Google Scholar] [CrossRef]
- Fattal, B.; Dotan, A.; Tchorsh, Y.; Parpari, L.; Shuval, H. Penetration of E. coli and F2 Bacteriophage into Fish Tissues. Schr. Ver. Wasser- Boden-und Lufthyg. 1988, 78, 27–38. [Google Scholar]
- Nguyen, S.; Baker, K.; Padman, B.S.; Patwa, R.; Dunstan, R.A.; Weston, T.A.; Schlosser, K.; Bailey, B.; Lithgow, T.; Lazarou, M. Bacteriophage Transcytosis Provides a Mechanism to Cross Epithelial Cell Layers. MBio 2017, 8, e01874-17. [Google Scholar] [CrossRef]
- Carroll-Portillo, A.; Lin, H.C. Bacteriophage and the Innate Immune System: Access and Signaling. Microorganisms 2019, 7, 625. [Google Scholar] [CrossRef]
- Galindo-Villegas, J.; García-Moreno, D.; De Oliveira, S.; Meseguer, J.; Mulero, V. Regulation of Immunity and Disease Resistance by Commensal Microbes and Chromatin Modifications during Zebrafish Development. Proc. Natl. Acad. Sci. USA 2012, 109, E2605–E2614. [Google Scholar] [CrossRef] [PubMed]
- Gorski, A.; Dabrowska, K.; Switala-Jeleń, K.; Nowaczyk, M.; Weber-Dabrowska, B.; Boratynski, J.; Wietrzyk, J.; Opolski, A. New Insights into the Possible Role of Bacteriophages in Host Defense and Disease. Med. Immunol. 2003, 2, 1–5. [Google Scholar] [CrossRef] [PubMed]
- An, T.-W.; Kim, S.-J.; Lee, Y.-D.; Park, J.-H.; Chang, H.-I. The Immune-Enhancing Effect of the Cronobacter sakazakii ES2 Phage Results in the Activation of Nuclear Factor-kB and Dendritic Cell Maturation via the Activation of IL-12p40 in the Mouse Bone Marrow. Immunol. Lett. 2014, 157, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Kurzępa, A.; Dąbrowska, K.; Skaradziński, G.; Górski, A. Bacteriophage Interactions with Phagocytes and Their Potential Significance in Experimental Therapy. Clin. Exp. Med. 2009, 9, 93–100. [Google Scholar] [CrossRef]
- Górski, A.; Dąbrowska, K.; Międzybrodzki, R.; Weber-Dąbrowska, B.; Łusiak-Szelachowska, M.; Jończyk-Matysiak, E.; Borysowski, J. Phages and Immunomodulation. Future Microbiol. 2017, 12, 905–914. [Google Scholar] [CrossRef]
- Van Belleghem, J.D.; Clement, F.; Merabishvili, M.; Lavigne, R.; Vaneechoutte, M. Pro-and Anti-Inflammatory Responses of Peripheral Blood Mononuclear Cells Induced by Staphylococcus aureus and Pseudomonas aeruginosa Phages. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef]
- Dufour, N.; Delattre, R.; Chevallereau, A.; Ricard, J.-D.; Debarbieux, L. Phage Therapy of Pneumonia Is Not Associated with an Overstimulation of the Inflammatory Response Compared to Antibiotic Treatment in Mice. Antimicrob. Agents Chemother. 2019, 63, e00379-19. [Google Scholar] [CrossRef]
- Khan Mirzaei, M.; Haileselassie, Y.; Navis, M.; Cooper, C.; Sverremark-Ekström, E.; Nilsson, A.S. Morphologically Distinct Escherichia coli Bacteriophages Differ in Their Efficacy and Ability to Stimulate Cytokine Release in vitro. Front. Microbiol. 2016, 7, 437. [Google Scholar]
- Secor, P.R.; Michaels, L.A.; Smigiel, K.S.; Rohani, M.G.; Jennings, L.K.; Hisert, K.B.; Arrigoni, A.; Braun, K.R.; Birkland, T.P.; Lai, Y. Filamentous Bacteriophage Produced by Pseudomonas aeruginosa Alters the Inflammatory Response and Promotes Noninvasive Infection in vivo. Infect. Immun. 2017, 85, e00648-16. [Google Scholar] [CrossRef]
- Trend, S.; Chang, B.J.; O’Dea, M.; Stick, S.M.; Kicic, A.; WAERP; AusREC; AREST CF. Use of a Primary Epithelial Cell Screening Tool to Investigate Phage Therapy in Cystic Fibrosis. Front. Pharmacol. 2018, 9, 1330. [Google Scholar] [CrossRef]
- Cafora, M.; Deflorian, G.; Forti, F.; Ferrari, L.; Binelli, G.; Briani, F.; Ghisotti, D.; Pistocchi, A. Phage Therapy against Pseudomonas aeruginosa Infections in a Cystic Fibrosis Zebrafish Model. Sci. Rep. 2019, 9, 1–10. [Google Scholar] [CrossRef]
- Cafora, M.; Forti, F.; Briani, F.; Ghisotti, D.; Pistocchi, A. Phage Therapy Application to Counteract Pseudomonas aeruginosa Infection in Cystic Fibrosis Zebrafish Embryos. JoVE (J. Vis. Exp.) 2020, e61275. [Google Scholar] [CrossRef]
- Nikapitiya, C.; Chandrarathna, H.; Dananjaya, S.; De Zoysa, M.; Lee, J. Isolation and Characterization of Phage (ETP-1) Specific to Multidrug Resistant Pathogenic Edwardsiella tarda and Its in vivo Biocontrol Efficacy in Zebrafish (Danio rerio). Biologicals 2020, 63, 14–23. [Google Scholar] [CrossRef]
- Yun, S.; Jun, J.W.; Giri, S.S.; Kim, H.J.; Chi, C.; Kim, S.G.; Kim, S.W.; Kang, J.W.; Han, S.J.; Kwon, J. Immunostimulation of Cyprinus carpio Using Phage Lysate of Aeromonas hydrophila. Fish Shellfish. Immunol. 2019, 86, 680–687. [Google Scholar] [CrossRef]
- Schulz, P.; Robak, S.; Dastych, J.; Siwicki, A.K. Influence of Bacteriophages Cocktail on European Eel (Anguilla anguilla) Immunity and Survival after Experimental Challenge. Fish Shellfish. Immunol. 2019, 84, 28–37. [Google Scholar] [CrossRef]
- Lin, H.; Caywood, B.E.; Rowlands, D., Jr. Primary and Secondary Immune Responses of the Marine Toad (Bufo marinus) to Bacterophage F2. Immunology 1971, 20, 373. [Google Scholar]
- Bradley, S.; Kim, Y.; Watson, D. Immune Response by the Mouse to Orally Administered Actinophage. Proc. Soc. Exp. Biol. Med. 1963, 113, 686–688. [Google Scholar] [CrossRef]
- Young, R.; Ruddle, F.H. Inactivation of T-2 Bacteriophage by Sensitized Leucocytes in vitro. Nature 1965, 208, 1105–1106. [Google Scholar] [CrossRef]
- Van Belleghem, J.D.; Dąbrowska, K.; Vaneechoutte, M.; Barr, J.J.; Bollyky, P.L. Interactions between Bacteriophage, Bacteria, and the Mammalian Immune System. Viruses 2019, 11, 10. [Google Scholar] [CrossRef]
- Bekeredjian-Ding, I.B.; Wagner, M.; Hornung, V.; Giese, T.; Schnurr, M.; Endres, S.; Hartmann, G. Plasmacytoid Dendritic Cells Control TLR7 Sensitivity of Naive B Cells via Type I IFN. J. Immunol. 2005, 174, 4043–4050. [Google Scholar] [CrossRef]
- Hashiguchi, S.; Yamaguchi, Y.; Takeuchi, O.; Akira, S.; Sugimura, K. Immunological Basis of M13 Phage Vaccine: Regulation under MyD88 and TLR9 Signaling. Biochem. Biophys. Res. Commun. 2010, 402, 19–22. [Google Scholar] [CrossRef]
- Hodyra-Stefaniak, K.; Miernikiewicz, P.; Drapała, J.; Drab, M.; Jończyk-Matysiak, E.; Lecion, D.; Kaźmierczak, Z.; Beta, W.; Majewska, J.; Harhala, M. Mammalian Host-Versus-Phage Immune Response Determines Phage Fate in vivo. Sci. Rep. 2015, 5, 1–13. [Google Scholar] [CrossRef]
- Krut, O.; Bekeredjian-Ding, I. Contribution of the Immune Response to Phage Therapy. J. Immunol. 2018, 200, 3037–3044. [Google Scholar] [CrossRef]
- Bettarel, Y.; Combe, M.; Adingra, A.; Ndiaye, A.; Bouvier, T.; Panfili, J.; Durand, J.-D. Hordes of Phages in the Gut of the Tilapia Sarotherodon melanotheron. Sci. Rep. 2018, 8, 1–6. [Google Scholar]
- He, Y.; Yang, H. The Gastrointestinal Phage Communities of the Cultivated Freshwater Fishes. FEMS Microbiol. Lett. 2015, 362, fnu027. [Google Scholar] [CrossRef]
- Luo, E.; Aylward, F.O.; Mende, D.R.; DeLong, E.F. Bacteriophage Distributions and Temporal Variability in the Ocean’s Interior. MBio 2017, 8, e01903-17. [Google Scholar] [CrossRef]
- Pietilä, M.K.; Demina, T.A.; Atanasova, N.S.; Oksanen, H.M.; Bamford, D.H. Archaeal Viruses and Bacteriophages: Comparisons and Contrasts. Trends Microbiol. 2014, 22, 334–344. [Google Scholar] [CrossRef]
- Ackermann, H.-W. Classification of Bacteriophages. Bacteriophages 2006, 2, 8–16. [Google Scholar]
- Ackermann, H.-W. 5500 Phages Examined in the Electron Microscope. Arch. Virol. 2007, 152, 227–243. [Google Scholar] [CrossRef] [PubMed]
- Ackermann, H.-W. Phage classification and characterization. In Bacteriophages; Springer: Cham, Switzerland, 2009; pp. 127–140. [Google Scholar]
- Adriaenssens, E.M.; Edwards, R.; Nash, J.H.; Mahadevan, P.; Seto, D.; Ackermann, H.-W.; Lavigne, R.; Kropinski, A.M. Integration of Genomic and Proteomic Analyses in the Classification of the Siphoviridae Family. Virology 2015, 477, 144–154. [Google Scholar] [CrossRef]
- Lavigne, R.; Darius, P.; Summer, E.J.; Seto, D.; Mahadevan, P.; Nilsson, A.S.; Ackermann, H.W.; Kropinski, A.M. Classification of Myoviridae Bacteriophages Using Protein Sequence Similarity. BMC Microbiol. 2009, 9, 1–16. [Google Scholar] [CrossRef]
- Lavigne, R.; Seto, D.; Mahadevan, P.; Ackermann, H.-W.; Kropinski, A.M. Unifying Classical and Molecular Taxonomic Classification: Analysis of the Podoviridae Using BLASTP-Based Tools. Res. Microbiol. 2008, 159, 406–414. [Google Scholar] [CrossRef]
- Paterson, W.; Douglas, R.; Grinyer, I.; McDermott, L. Isolation and Preliminary Characterization of Some Aeromonas salmonicida Bacteriophages. J. Fish. Board Can. 1969, 26, 629–632. [Google Scholar] [CrossRef]
- Carlson, K. Working with Bacteriophages: Common Techniques and Methodological Approaches; CRC Press: Boca Raton, FL, USA, 2005; Volume 1. [Google Scholar]
- Dubrovin, E.V.; Voloshin, A.G.; Kraevsky, S.V.; Ignatyuk, T.E.; Abramchuk, S.S.; Yaminsky, I.V.; Ignatov, S.G. Atomic Force Microscopy Investigation of Phage Infection of Bacteria. Langmuir 2008, 24, 13068–13074. [Google Scholar] [CrossRef]
- Zago, M.; Scaltriti, E.; Fornasari, M.E.; Rivetti, C.; Grolli, S.; Giraffa, G.; Ramoni, R.; Carminati, D. Epifluorescence and Atomic Force Microscopy: Two Innovative Applications for Studying Phage–Host Interactions in Lactobacillus helveticus. J. Microbiol. Methods 2012, 88, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Dubrovin, E.V.; Popova, A.V.; Kraevskiy, S.V.; Ignatov, S.G.; Ignatyuk, T.E.; Yaminsky, I.V.; Volozhantsev, N.V. Atomic Force Microscopy Analysis of the Acinetobacter baumannii Bacteriophage AP22 Lytic Cycle. PLoS ONE 2012, 7, e47348. [Google Scholar] [CrossRef] [PubMed]
- Remuzgo-Martínez, S.; Lázaro-Díez, M.; Padilla, D.; Vega, B.; El Aamri, F.; Icardo, J.M.; Acosta, F.; Ramos-Vivas, J. New Aspects in the Biology of Photobacterium damselae subsp. piscicida: Pili, Motility and Adherence to Solid Surfaces. Vet. Microbiol. 2014, 174, 247–254. [Google Scholar] [CrossRef]
- Vivas, J.; Padilla, D.; Real, F.; Bravo, J.; Grasso, V.; Acosta, F. Influence of Environmental Conditions on Biofilm Formation by Hafnia alvei Strains. Vet. Microbiol. 2008, 129, 150–155. [Google Scholar] [CrossRef]
- Cai, W.; Arias, C.R. Biofilm Formation on Aquaculture Substrates by Selected Bacterial Fish Pathogens. J. Aquat. Anim. Health 2017, 29, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Hall-Stoodley, L.; Costerton, J.W.; Stoodley, P. Bacterial Biofilms: From the Natural Environment to Infectious Diseases. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef]
- Acosta, F.; Montero, D.; Izquierdo, M.; Galindo-Villegas, J. High-Level Biocidal Products Effectively Eradicate Pathogenic γ-Proteobacteria Biofilms from Aquaculture Facilities. Aquaculture 2021, 532, 736004. [Google Scholar] [CrossRef]
- Koo, H.; Allan, R.N.; Howlin, R.P.; Stoodley, P.; Hall-Stoodley, L. Targeting Microbial Biofilms: Current and Prospective Therapeutic Strategies. Nat. Rev. Microbiol. 2017, 15, 740–755. [Google Scholar] [CrossRef]
- Clutterbuck, A.; Woods, E.; Knottenbelt, D.; Clegg, P.; Cochrane, C.; Percival, S. Biofilms and Their Relevance to Veterinary Medicine. Vet. Microbiol. 2007, 121, 1–17. [Google Scholar] [CrossRef]
- MacKenzie, K.D.; Palmer, M.B.; Köster, W.L.; White, A.P. Examining the Link between Biofilm Formation and the Ability of Pathogenic Salmonella Strains to Colonize Multiple Host Species. Front. Vet. Sci. 2017, 4, 138. [Google Scholar] [CrossRef]
- Curtin, J.J.; Donlan, R.M. Using Bacteriophages to Reduce Formation of Catheter-Associated Biofilms by Staphylococcus epidermidis. Antimicrob. Agents Chemother. 2006, 50, 1268–1275. [Google Scholar] [CrossRef] [PubMed]
- Melo, L.D.; Ferreira, R.; Costa, A.R.; Oliveira, H.; Azeredo, J. Efficacy and Safety Assessment of Two Enterococci Phages in an in vitro Biofilm Wound Model. Sci. Rep. 2019, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Shlezinger, M.; Friedman, M.; Houri-Haddad, Y.; Hazan, R.; Beyth, N. Phages in a Thermoreversible Sustained-Release Formulation Targeting E. faecalis in vitro and in vivo. PLoS ONE 2019, 14, e0219599. [Google Scholar] [CrossRef]
- Gutierrez, D.; Ruas-Madiedo, P.; Martínez, B.; Rodríguez, A.; García, P. Effective Removal of Staphylococcal Biofilms by the Endolysin LysH5. PLoS ONE 2014, 9, e107307. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, D.; Fernández, L.; Martínez, B.; Ruas-Madiedo, P.; García, P.; Rodríguez, A. Real-Time Assessment of Staphylococcus aureus Biofilm Disruption by Phage-Derived Proteins. Front. Microbiol. 2017, 8, 1632. [Google Scholar] [CrossRef] [PubMed]
- Fu, W.; Forster, T.; Mayer, O.; Curtin, J.J.; Lehman, S.M.; Donlan, R.M. Bacteriophage Cocktail for the Prevention of Biofilm Formation by Pseudomonas aeruginosa on Catheters in an in vitro Model System. Antimicrob. Agents Chemother. 2010, 54, 397–404. [Google Scholar] [CrossRef] [PubMed]
- Magin, V.; Garrec, N.; Andrés, Y. Selection of Bacteriophages to Control in vitro 24 h Old Biofilm of Pseudomonas aeruginosa Isolated from Drinking and Thermal Water. Viruses 2019, 11, 749. [Google Scholar] [CrossRef]
- Cano, E.J.; Caflisch, K.M.; Bollyky, P.L.; Van Belleghem, J.D.; Patel, R.; Fackler, J.; Brownstein, M.J.; Horne, B.; Biswas, B.; Henry, M. Phage Therapy for Limb-Threatening Prosthetic Knee Klebsiella pneumoniae Infection: Case Report and in vitro Characterization of Anti-Biofilm Activity. Clin. Infect. Dis. 2021, 73, e144–e151. [Google Scholar] [CrossRef] [PubMed]
- Fong, S.A.; Drilling, A.; Morales, S.; Cornet, M.E.; Woodworth, B.A.; Fokkens, W.J.; Psaltis, A.J.; Vreugde, S.; Wormald, P.-J. Activity of Bacteriophages in Removing Biofilms of Pseudomonas aeruginosa Isolates from Chronic Rhinosinusitis Patients. Front. Cell. Infect. Microbiol. 2017, 7, 418. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, A.; Sousa, J.C.; Silva, A.C.; Melo, L.D.; Sillankorva, S. Chestnut Honey and Bacteriophage Application to Control Pseudomonas Aeruginosa and Escherichia coli Biofilms: Evaluation in an ex vivo Wound Model. Front. Microbiol. 2018, 9, 1725. [Google Scholar] [CrossRef]
- Oliveira, A.; Ribeiro, H.G.; Silva, A.C.; Silva, M.D.; Sousa, J.C.; Rodrigues, C.F.; Melo, L.D.; Henriques, A.F.; Sillankorva, S. Synergistic Antimicrobial Interaction between Honey and Phage against Escherichia coli Biofilms. Front. Microbiol. 2017, 8, 2407. [Google Scholar] [CrossRef]
- Papadopoulou, A.; Dalsgaard, I.; Wiklund, T. Inhibition Activity of Compounds and Bacteriophages against Flavobacterium psychrophilum Biofilms in vitro. J. Aquat. Anim. Health 2019, 31, 225–238. [Google Scholar] [CrossRef] [PubMed]
- Le, T.S.; Nguyen, T.H.; Vo, H.P.; Doan, V.C.; Nguyen, H.L.; Tran, M.T.; Tran, T.T.; Southgate, P.C.; Kurtböke, D.İ. Protective Effects of Bacteriophages against Aeromonas hydrophila Causing Motile Aeromonas septicemia (MAS) in Striped Catfish. Antibiotics 2018, 7, 16. [Google Scholar] [CrossRef]
- Liu, J.; Gao, S.; Dong, Y.; Lu, C.; Liu, Y. Isolation and Characterization of Bacteriophages against Virulent Aeromonas hydrophila. BMC Microbiol. 2020, 20, 1–13. [Google Scholar] [CrossRef]
- Easwaran, M.; Dananjaya, S.; Park, S.C.; Lee, J.; Shin, H.; De Zoysa, M. Characterization of Bacteriophage PAh-1 and Its Protective Effects on Experimental Infection of Aeromonas hydrophila in Zebrafish (Danio Rerio). J. Fish Dis. 2017, 40, 841–846. [Google Scholar] [CrossRef]
- Akmal, M.; Rahimi-Midani, A.; Hafeez-ur-Rehman, M.; Hussain, A.; Choi, T.-J. Isolation, Characterization, and Application of a Bacteriophage Infecting the Fish Pathogen Aeromonas hydrophila. Pathogens 2020, 9, 215. [Google Scholar] [CrossRef]
- Chandrarathna, H.; Nikapitiya, C.; Dananjaya, S.; De Silva, B.; Heo, G.-J.; De Zoysa, M.; Lee, J. Isolation and Characterization of Phage AHP-1 and Its Combined Effect with Chloramphenicol to Control Aeromonas hydrophila. Braz. J. Microbiol. 2020, 51, 409–416. [Google Scholar] [CrossRef]
- Cheng, Y.; Gao, D.; Xia, Y.; Wang, Z.; Bai, M.; Luo, K.; Cui, X.; Wang, Y.; Zhang, S.; Xiao, W. Characterization of Novel Bacteriophage AhyVDH1 and Its Lytic Activity Against Aeromonas hydrophila. Curr. Microbiol. 2021, 78, 329–337. [Google Scholar] [CrossRef]
- Cao, Y.; Li, S.; Wang, D.; Zhao, J.; Xu, L.; Liu, H.; Lu, T.; Mou, Z. Genomic Characterization of a Novel Virulent Phage Infecting the Aeromonas hydrophila Isolated from Rainbow Trout (Oncorhynchus mykiss). Virus Res. 2019, 273, 197764. [Google Scholar] [CrossRef]
- Wu, J.-L.; Lin, H.-M.; Jan, L.; Hsu, Y.-L.; CHANG, L.-H. Biological Control of Fish Bacterial Pathogen, Aeromonas hydrophila, by Bacteriophage AH 1. Fish Pathol. 1981, 15, 271–276. [Google Scholar] [CrossRef]
- Tu, V.Q.; Nguyen, T.-T.; Tran, X.T.; Millard, A.D.; Phan, H.T.; Le, N.P.; Dang, O.T.; Hoang, H.A. Complete Genome Sequence of a Novel Lytic Phage Infecting Aeromonas hydrophila, an Infectious Agent in Striped Catfish (Pangasianodon hypophthalmus). Arch. Virol. 2020, 165, 2973–2977. [Google Scholar] [CrossRef] [PubMed]
- Hoang Hoang, A.; Xuan Tran, T.T.; Nga, L.E.P.; Oanh Dang, T.H. Selection of Phages to Control Aeromonas hydrophila–an Infectious Agent in Striped Catfish. Biocontrol Sci. 2019, 24, 23–28. [Google Scholar]
- Jun, J.W.; Kim, J.H.; Shin, S.P.; Han, J.E.; Chai, J.Y.; Park, S.C. Protective Effects of the Aeromonas Phages PAh1-C and PAh6-C against Mass Mortality of the Cyprinid Loach (Misgurnus anguillicaudatus) Caused by Aeromonas hydrophila. Aquaculture 2013, 416, 289–295. [Google Scholar] [CrossRef]
- Wang, J.-B.; Lin, N.-T.; Tseng, Y.-H.; Weng, S.-F. Genomic Characterization of the Novel Aeromonas hydrophila Phage Ahp1 Suggests the Derivation of a New Subgroup from PhiKMV-like Family. PLoS ONE 2016, 11, e0162060. [Google Scholar] [CrossRef]
- Haq, I.U.; Chaudhry, W.N.; Andleeb, S.; Qadri, I. Isolation and Partial Characterization of a Virulent Bacteriophage IHQ1 Specific for Aeromonas punctata from Stream Water. Microb. Ecol. 2012, 63, 954–963. [Google Scholar] [CrossRef] [PubMed]
- Vincent, A.T.; Paquet, V.E.; Bernatchez, A.; Tremblay, D.M.; Moineau, S.; Charette, S.J. Characterization and Diversity of Phages Infecting Aeromonas salmonicida subsp. salmonicida. Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Kim, J.H.; Son, J.S.; Choi, Y.J.; Choresca, C.H., Jr.; Shin, S.P.; Han, J.E.; Jun, J.W.; Park, S.C. Complete Genome Sequence and Characterization of a Broad-Host Range T4-like Bacteriophage PhiAS5 Infecting Aeromonas salmonicida subsp. salmonicida. Vet. Microbiol. 2012, 157, 164–171. [Google Scholar] [CrossRef]
- Kim, J.; Son, J.; Choi, Y.; Choresca, C.; Shin, S.; Han, J.; Jun, J.; Kang, D.; Oh, C.; Heo, S. Isolation and Characterization of a Lytic Myoviridae Bacteriophage PAS-1 with Broad Infectivity in Aeromonas salmonicida. Curr. Microbiol. 2012, 64, 418–426. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Yuan, S.; Chen, L.; Liu, Q.; Zhang, H.; Ma, Y.; Wei, T.; Huang, S. Complete Genome Analysis of Bacteriophage AsXd-1, a New Member of the Genus Hk97virus, Family Siphoviridae. Arch. Virol. 2018, 163, 3195–3197. [Google Scholar] [CrossRef] [PubMed]
- Silva, Y.J.; Moreirinha, C.; Pereira, C.; Costa, L.; Rocha, R.J.; Cunha, Â.; Gomes, N.C.; Calado, R.; Almeida, A. Biological Control of Aeromonas salmonicida Infection in Juvenile Senegalese Sole (Solea senegalensis) with Phage AS-A. Aquaculture 2016, 450, 225–233. [Google Scholar] [CrossRef]
- Duarte, J.; Pereira, C.; Costa, P.; Almeida, A. Bacteriophages with Potential to Inactivate Aeromonas hydrophila in Cockles: In vitro and in vivo Preliminary Studies. Antibiotics 2021, 10, 710. [Google Scholar] [CrossRef] [PubMed]
- Imbeault, S.; Parent, S.; Lagacé, M.; Uhland, C.F.; Blais, J.-F. Using Bacteriophages to Prevent Furunculosis Caused by Aeromonas salmonicida in Farmed Brook Trout. J. Aquat. Anim. Health 2006, 18, 203–214. [Google Scholar] [CrossRef]
- Petrov, V.; Karam, J. Diversity of Structure and Function of DNA Polymerase (Gp43) of T4-Related Bacteriophages. Biochemistry (Moscow) 2004, 69, 1213–1218. [Google Scholar] [CrossRef]
- He, Y.; Huang, Z.; Zhang, X.; Zhang, Z.; Gong, M.; Pan, X.; Wei, D.; Yang, H. Characterization of a Novel Lytic Myophage, PhiA8-29, Infecting Aeromonas Strains. Arch. Virol. 2019, 164, 893–896. [Google Scholar] [CrossRef]
- Jia, K.; Yang, N.; Zhang, X.; Cai, R.; Zhang, Y.; Tian, J.; Raza, S.H.A.; Kang, Y.; Qian, A.; Li, Y. Genomic, Morphological and Functional Characterization of Virulent Bacteriophage IME-JL8 Targeting Citrobacter freundii. Front. Microbiol. 2020, 11, 2967. [Google Scholar] [CrossRef]
- Walakira, J.; Carrias, A.; Hossain, M.; Jones, E.; Terhune, J.; Liles, M. Identification and Characterization of Bacteriophages Specific to the Catfish Pathogen, Edwardsiella ictaluri. J. Appl. Microbiol. 2008, 105, 2133–2142. [Google Scholar] [CrossRef]
- Carrias, A.; Welch, T.J.; Waldbieser, G.C.; Mead, D.A.; Terhune, J.S.; Liles, M.R. Comparative Genomic Analysis of Bacteriophages Specific to the Channel Catfish Pathogen Edwardsiella ictaluri. Virol. J. 2011, 8, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Yasuike, M.; Kai, W.; Nakamura, Y.; Fujiwara, A.; Kawato, Y.; Hassan, E.S.; Mahmoud, M.M.; Nagai, S.; Kobayashi, T.; Ototake, M. Complete Genome Sequence of the Edwardsiella ictaluri-Specific Bacteriophage PEi21, Isolated from River Water in Japan. Genome Announc. 2014, 2, e00228-14. [Google Scholar] [CrossRef]
- Hassan, E.S.; Mahmoud, M.M.; Kawato, Y.; Nagai, T.; Kawaguchi, O.; Iida, Y.; Yuasa, K.; Nakai, T. Subclinical Edwardsiella ictaluri Infection of Wild Ayu Plecoglossus Altivelis. Fish Pathol. 2012, 47, 64–73. [Google Scholar] [CrossRef]
- Hoang, H.A.; Yen, M.H.; Ngoan, V.T.; Nga, L.P.; Oanh, D.T. Virulent Bacteriophage of Edwardsiella ictaluri Isolated from Kidney and Liver of Striped Catfish Pangasianodon Hypophthalmus in Vietnam. Dis. Aquat. Org. 2018, 132, 49–56. [Google Scholar] [CrossRef]
- Kim, S.G.; Giri, S.S.; Yun, S.; Kim, H.J.; Kim, S.W.; Kang, J.W.; Han, S.J.; Kwon, J.; Jun, J.W.; Oh, W.T. Genomic Characterization of Bacteriophage PEt-SU, a Novel PhiKZ-Related Virus Infecting Edwardsiella tarda. Arch. Virol. 2020, 165, 219–222. [Google Scholar] [CrossRef]
- Cui, H.; Zhang, J.; Cong, C.; Wang, L.; Li, X.; Murtaza, B.; Xu, Y. Complete Genome Analysis of the Novel Edwardsiella tarda Phage VB_EtaM_ET-ABTNL-9. Arch. Virol. 2020, 165, 1241–1244. [Google Scholar] [CrossRef]
- Yasuike, M.; Nishiki, I.; Iwasaki, Y.; Nakamura, Y.; Fujiwara, A.; Sugaya, E.; Kawato, Y.; Nagai, S.; Kobayashi, T.; Ototake, M. Full-Genome Sequence of a Novel Myovirus, GF-2, Infecting Edwardsiella tarda: Comparison with Other Edwardsiella myoviral Genomes. Arch. Virol. 2015, 160, 2129–2133. [Google Scholar] [CrossRef]
- Laanto, E.; Bamford, J.K.; Ravantti, J.J.; Sundberg, L.-R. The Use of Phage FCL-2 as an Alternative to Chemotherapy against Columnaris Disease in Aquaculture. Front. Microbiol. 2015, 6, 829. [Google Scholar] [CrossRef] [PubMed]
- Laanto, E.; Sundberg, L.-R.; Bamford, J.K. Phage Specificity of the Freshwater Fish Pathogen Flavobacterium columnare. Appl. Environ. Microbiol. 2011, 77, 7868–7872. [Google Scholar] [CrossRef] [PubMed]
- Prasad, Y.; Kumar, D.; Sharma, A. Lytic Bacteriophages Specific to Flavobacterium columnare Rescue Catfish, Clarias batrachus (Linn.) from Columnaris Disease. J. Environ. Biol. 2011, 32, 161–168. [Google Scholar] [PubMed]
- Christiansen, R.H.; Dalsgaard, I.; Middelboe, M.; Lauritsen, A.H.; Madsen, L. Detection and Quantification of Flavobacterium psychrophilum-Specific Bacteriophages in vivo in Rainbow Trout upon Oral Administration: Implications for Disease Control in Aquaculture. Appl. Environ. Microbiol. 2014, 80, 7683–7693. [Google Scholar] [CrossRef]
- Stenholm, A.R.; Dalsgaard, I.; Middelboe, M. Isolation and Characterization of Bacteriophages Infecting the Fish Pathogen Flavobacterium psychrophilum. Appl. Environ. Microbiol. 2008, 74, 4070–4078. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Gomez, D.K.; Nakai, T.; Park, S.C. Isolation and Identification of Bacteriophages Infecting Ayu Plecoglossus altivelis Altivelis Specific Flavobacterium psychrophilum. Vet. Microbiol. 2010, 140, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Yamaki, S.; Kawai, Y.; Yamazaki, K. Characterization of a Novel Bacteriophage, Phda1, Infecting the Histamine-producing Photobacterium damselae subsp. damselae. J. Appl. Microbiol. 2015, 118, 1541–1550. [Google Scholar] [CrossRef]
- Veyrand-Quirós, B.; Gómez-Gil, B.; Lomeli-Ortega, C.O.; Escobedo-Fregoso, C.; Millard, A.D.; Tovar-Ramírez, D.; Balcázar, J.L.; Quiroz-Guzmán, E. Use of Bacteriophage VB_Pd_PDCC-1 as Biological Control Agent of Photobacterium Damselae Subsp. Damselae during Hatching of Longfin Yellowtail (Seriola rivoliana) Eggs. J. Appl. Microbiol. 2020, 129, 1497–1510. [Google Scholar] [CrossRef]
- Kawato, Y.; Yasuike, M.; Nakamura, Y.; Shigenobu, Y.; Fujiwara, A.; Sano, M.; Nakai, T. Complete Genome Sequence Analysis of Two Pseudomonas plecoglossicida Phages, Potential Therapeutic Agents. Appl. Environ. Microbiol. 2015, 81, 874–881. [Google Scholar] [CrossRef]
- Park, S.C.; Shimamura, I.; Fukunaga, M.; Mori, K.-I.; Nakai, T. Isolation of Bacteriophages Specific to a Fish Pathogen, Pseudomonas plecoglossicida, as a Candidate for Disease Control. Appl. Environ. Microbiol. 2000, 66, 1416–1422. [Google Scholar] [CrossRef]
- Khairnar, K.; Raut, M.P.; Chandekar, R.H.; Sanmukh, S.G.; Paunikar, W.N. Novel Bacteriophage Therapy for Controlling Metallo-Beta-Lactamase Producing Pseudomonas aeruginosa Infection in Catfish. BMC Vet. Res. 2013, 9, 1–9. [Google Scholar] [CrossRef]
- Yang, Z.; Tao, X.; Zhang, H.; Rao, S.; Gao, L.; Pan, Z.; Jiao, X. Isolation and Characterization of Virulent Phages Infecting Shewanella baltica and Shewanella putrefaciens, and Their Application for Biopreservation of Chilled Channel Catfish (Ictalurus punctatus). Int. J. Food Microbiol. 2019, 292, 107–117. [Google Scholar] [CrossRef]
- Kawato, Y.; Istiqomah, I.; Gaafar, A.Y.; Hanaoka, M.; Ishimaru, K.; Yasuike, M.; Nishiki, I.; Nakamura, Y.; Fujiwara, A.; Nakai, T. A Novel Jumbo Tenacibaculum maritimum Lytic Phage with Head-Fiber-like Appendages. Arch. Virol. 2020, 165, 303–311. [Google Scholar] [CrossRef] [PubMed]
- Kokkari, C.; Sarropoulou, E.; Bastias, R.; Mandalakis, M.; Katharios, P. Isolation and Characterization of a Novel Bacteriophage Infecting Vibrio alginolyticus. Arch. Microbiol. 2018, 200, 707–718. [Google Scholar] [CrossRef] [PubMed]
- Lal, T.M.; Sano, M.; Hatai, K.; Ransangan, J. Complete Genome Sequence of a Giant Vibrio Phage ValKK3 Infecting Vibrio alginolyticus. Genom. Data 2016, 8, 37–38. [Google Scholar] [CrossRef]
- Kalatzis, P.G.; Bastías, R.; Kokkari, C.; Katharios, P. Isolation and Characterization of Two Lytic Bacteriophages, ΦSt2 and ΦGrn1; Phage Therapy Application for Biological Control of Vibrio alginolyticus in Aquaculture Live Feeds. PLoS ONE 2016, 11, e0151101. [Google Scholar] [CrossRef] [PubMed]
- Higuera, G.; Bastías, R.; Tsertsvadze, G.; Romero, J.; Espejo, R.T. Recently Discovered Vibrio anguillarum Phages Can Protect against Experimentally Induced Vibriosis in Atlantic Salmon, Salmo salar. Aquaculture 2013, 392, 128–133. [Google Scholar] [CrossRef]
- Silva, Y.J.; Costa, L.; Pereira, C.; Mateus, C.; Cunha, A.; Calado, R.; Gomes, N.C.; Pardo, M.A.; Hernandez, I.; Almeida, A. Phage Therapy as an Approach to Prevent Vibrio anguillarum Infections in Fish Larvae Production. PLoS ONE 2014, 9, e114197. [Google Scholar] [CrossRef]
- Tan, D.; Gram, L.; Middelboe, M. Vibriophages and Their Interactions with the Fish Pathogen Vibrio anguillarum. Appl. Environ. Microbiol. 2014, 80, 3128–3140. [Google Scholar] [CrossRef]
- Nuidate, T.; Kuaphiriyakul, A.; Surachat, K.; Mittraparp-Arthorn, P. Induction and Genome Analysis of HY01, a Newly Reported Prophage from an Emerging Shrimp Pathogen Vibrio campbellii. Microorganisms 2021, 9, 400. [Google Scholar] [CrossRef]
- Lomelí-Ortega, C.O.; Martínez-Sández, A.; Barajas-Sandoval, D.R.; Reyes, A.G.; Magallón-Barajas, F.; Veyrand-Quíros, B.; Gannon, L.; Harrison, C.; Michniewski, S.; Millard, A. Isolation and Characterization of Vibriophage VB_Vc_SrVc9: An Effective Agent in Preventing Vibrio campbellii Infections in Brine Shrimp Nauplii (Artemia franciscana). J. Appl. Microbiol. 2021, 131, 36–49. [Google Scholar] [CrossRef]
- Vinod, M.; Shivu, M.; Umesha, K.; Rajeeva, B.; Krohne, G.; Karunasagar, I.; Karunasagar, I. Isolation of Vibrio harveyi Bacteriophage with a Potential for Biocontrol of Luminous Vibriosis in Hatchery Environments. Aquaculture 2006, 255, 117–124. [Google Scholar] [CrossRef]
- Oakey, H.; Owens, L. A New Bacteriophage, VHML, Isolated from a Toxin-producing Strain of Vibrio harveyi in Tropical Australia. J. Appl. Microbiol. 2000, 89, 702–709. [Google Scholar] [CrossRef]
- Phumkhachorn, P.; Rattanachaikunsopon, P. Isolation and Partial Characterization of a Bacteriophage Infecting the Shrimp Pathogen Vibrio harveyi. Afr. J. Microbiol. Res 2010, 4, 1794–1800. [Google Scholar]
- Stalin, N.; Srinivasan, P. Efficacy of Potential Phage Cocktails against Vibrio harveyi and Closely Related Vibrio Species Isolated from Shrimp Aquaculture Environment in the South East Coast of India. Vet. Microbiol. 2017, 207, 83–96. [Google Scholar] [CrossRef]
- Wang, Y.; Barton, M.; Elliott, L.; Li, X.; Abraham, S.; O’Dea, M.; Munro, J. Bacteriophage Therapy for the Control of Vibrio harveyi in Greenlip Abalone (Haliotis laevigata). Aquaculture 2017, 473, 251–258. [Google Scholar] [CrossRef]
- Crothers-Stomps, C.; Høj, L.; Bourne, D.; Hall, M.; Owens, L. Isolation of Lytic Bacteriophage against Vibrio harveyi. J. Appl. Microbiol. 2010, 108, 1744–1750. [Google Scholar] [CrossRef]
- Patil, J.R.; Desai, S.N.; Roy, P.; Durgaiah, M.; Saravanan, R.S.; Vipra, A. Simulated Hatchery System to Assess Bacteriophage Efficacy against Vibrio harveyi. Dis. Aquat. Org. 2014, 112, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Karunasagar, I.; Shivu, M.; Girisha, S.; Krohne, G.; Karunasagar, I. Biocontrol of Pathogens in Shrimp Hatcheries Using Bacteriophages. Aquaculture 2007, 268, 288–292. [Google Scholar] [CrossRef]
- Shivu, M.M.; Rajeeva, B.C.; Girisha, S.K.; Karunasagar, I.; Krohne, G.; Karunasagar, I. Molecular Characterization of Vibrio harveyi Bacteriophages Isolated from Aquaculture Environments along the Coast of India. Environ. Microbiol. 2007, 9, 322–331. [Google Scholar] [CrossRef] [PubMed]
- Lal, T.M.; Sano, M.; Ransangan, J. Isolation and Characterization of Large Marine Bacteriophage (Myoviridae), VhKM4 Infecting Vibrio harveyi. J. Aquat. Anim. Health 2017, 29, 26–30. [Google Scholar] [CrossRef] [PubMed]
- Echeverría-Vega, A.; Morales-Vicencio, P.; Saez-Saavedra, C.; Ceh, J.; Araya, R. The Complete Genome Sequence and Analysis of VB_VorS-PVo5, a Vibrio Phage Infectious to the Pathogenic Bacterium Vibrio Ordalii ATCC-33509. Stand. Genom. Sci. 2016, 11, 1–8. [Google Scholar] [CrossRef]
- Peng, Y.; Ding, Y.; Lin, H.; Wang, J. Isolation, Identification and Lysis Properties Analysis of a Vibrio parahaemolyticus Phage VPp1. Mar. Sci. 2013, 37, 96–101. [Google Scholar]
- Matsuzaki, S.; Inoue, T.; Tanaka, S.; Koga, T.; Kuroda, M.; Kimura, S.; Imai, S. Characterization of a Novel Vibrio parahaemolyticus Phage, KVP241, and Its Relatives Frequently Isolated from Seawater. Microbiol. Immunol. 2000, 44, 953–956. [Google Scholar] [CrossRef]
- Matsuzaki, S.; Tanaka, S.; Koga, T.; Kawata, T. A Broad-host-range Vibriophage, KVP40, Isolated from Sea Water. Microbiol. Immunol. 1992, 36, 93–97. [Google Scholar] [CrossRef]
- Onarinde, B.A.; Dixon, R.A. Prospects for Biocontrol of Vibrio parahaemolyticus Contamination in Blue Mussels (Mytilus edulus)—A Year-Long Study. Front. Microbiol. 2018, 9, 1043. [Google Scholar] [CrossRef]
- Kim, J.H.; Jun, J.W.; Choresca, C.H.; Shin, S.P.; Han, J.E.; Park, S.C. Complete Genome Sequence of a Novel Marine Siphovirus, PVp-1, Infecting Vibrio parahaemolyticus. J. Virol. 2012, 86, 7013–7014. [Google Scholar] [CrossRef] [PubMed]
- Jun, J.W.; Kim, H.J.; Yun, S.K.; Chai, J.Y.; Park, S.C. Eating Oysters without Risk of Vibriosis: Application of a Bacteriophage against Vibrio parahaemolyticus in Oysters. Int. J. Food Microbiol. 2014, 188, 31–35. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Liang, Y.; Huang, S.; Zhang, J.; Wang, J.; Chen, H.; Ye, Y.; Gao, X.; Wu, Q.; Tan, Z. Isolation and Characterization of the Novel Phages VB_VpS_BA3 and VB_VpS_CA8 for Lysing Vibrio parahaemolyticus. Front. Microbiol. 2020, 11, 259. [Google Scholar] [CrossRef] [PubMed]
- Matamp, N.; Bhat, S.G. Genome Characterization of Novel Lytic Myoviridae Bacteriophage ΦVP-1 Enhances Its Applicability against MDR-Biofilm-Forming Vibrio parahaemolyticus. Arch. Virol. 2020, 165, 387–396. [Google Scholar] [CrossRef] [PubMed]
- Lal, T.M.; Sano, M.; Ransangan, J. Genome Characterization of a Novel Vibriophage VpKK5 (Siphoviridae) Specific to Fish Pathogenic Strain of Vibrio parahaemolyticus. J. Basic Microbiol. 2016, 56, 872–888. [Google Scholar] [CrossRef]
- Lal, T.M.; Ransangan, J. Complete Genome Sequence of VpKK5, a Novel Vibrio parahaemolyticus Lytic Siphophage. Genome Announc. 2015, 3, e01381-14. [Google Scholar] [CrossRef]
- Li, Z.; Li, X.; Zhang, J.; Wang, X.; Wang, L.; Cao, Z.; Xu, Y. Use of Phages to Control Vibrio splendidus Infection in the Juvenile Sea Cucumber Apostichopus japonicus. Fish Shellfish. Immunol. 2016, 54, 302–311. [Google Scholar] [CrossRef]
- Katharios, P.; Kalatzis, P.G.; Kokkari, C.; Sarropoulou, E.; Middelboe, M. Isolation and Characterization of a N4-like Lytic Bacteriophage Infecting Vibrio splendidus, a Pathogen of Fish and Bivalves. PLoS ONE 2017, 12, e0190083. [Google Scholar] [CrossRef]
- Kim, H.J.; Jun, J.W.; Giri, S.S.; Chi, C.; Yun, S.; Kim, S.G.; Kim, S.W.; Kang, J.W.; Han, S.J.; Kwon, J. Application of the Bacteriophage PVco-14 to Prevent Vibrio coralliilyticus Infection in Pacific Oyster (Crassostrea gigas) Larvae. J. Invertebr. Pathol. 2019, 167, 107244. [Google Scholar] [CrossRef]
- Lee, H.S.; Choi, S.; Choi, S.H. Complete Genome Sequence of Vibrio vulnificus Bacteriophage SSP002. J. Virol. 2012, 86, 7711. [Google Scholar] [CrossRef]
- Lee, H.S.; Choi, S.; Shin, H.; Lee, J.-H.; Choi, S.H. Vibrio vulnificus Bacteriophage SSP002 as a Possible Biocontrol Agent. Appl. Environ. Microbiol. 2014, 80, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-J.; Kim, Y.-T.; Kim, H.B.; Choi, S.H.; Lee, J.-H. Characterization of Bacteriophage VVP001 and Its Application for the Inhibition of Vibrio Vulnificus Causing Seafood-Borne Diseases. Food Microbiol. 2021, 94, 103630. [Google Scholar] [CrossRef] [PubMed]
- Srinivasan, P.; Ramasamy, P. Morphological Characterization and Biocontrol Effects of Vibrio vulnificus Phages against Vibriosis in the Shrimp Aquaculture Environment. Microb. Pathog. 2017, 111, 472–480. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Fan, J.; Yan, T.; Liu, Q.; Yuan, S.; Zhang, H.; Yang, J.; Deng, D.; Huang, S.; Ma, Y. Isolation and Characterization of Specific Phages to Prepare a Cocktail Preventing Vibrio Sp. Va-F3 Infections in Shrimp (Litopenaeus vannamei). Front. Microbiol. 2019, 10, 2337. [Google Scholar] [CrossRef] [PubMed]
- Welch, T.J. Characterization of a Novel Yersinia ruckeri Serotype O1-specific Bacteriophage with Virulence-neutralizing Activity. J. Fish Dis. 2020, 43, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Stevenson, R.; Airdrie, D. Isolation of Yersinia ruckeri Bacteriophages. Appl. Environ. Microbiol. 1984, 47, 1201–1205. [Google Scholar] [CrossRef]
- Kiljunen, S.; Hakala, K.; Pinta, E.; Huttunen, S.; Pluta, P.; Gador, A.; Lönnberg, H.; Skurnik, M. Yersiniophage ΦR1-37 Is a Tailed Bacteriophage Having a 270 Kb DNA Genome with Thymidine Replaced by Deoxyuridine. Microbiology 2005, 151, 4093–4102. [Google Scholar] [CrossRef]
- Leskinen, K.; Pajunen, M.I.; Vilanova, M.V.G.-R.; Kiljunen, S.; Nelson, A.; Smith, D.; Skurnik, M. YerA41, a Yersinia ruckeri Bacteriophage: Determination of a Non-Sequencable DNA Bacteriophage Genome via RNA-Sequencing. Viruses 2020, 12, 620. [Google Scholar] [CrossRef]
- Park, K.; Matsuoka, S.; Nakai, T.; Muroga, K. A Virulent Bacteriophage of Lactococcus garvieae (Formerly Enterococcus Seriolicida) Isolated from Yellowtail Seriola quinqueradiata. Dis. Aquat. Org. 1997, 29, 145–149. [Google Scholar] [CrossRef]
- Park, K.H.; Kato, H.; Nakai, T.; Muroga, K. Phage Typing of Lactococcus garvieae (Formerly Enterococcus Seriolicida) a Pathogen of Cultured Yellowtail. Fish. Sci. 1998, 64, 62–64. [Google Scholar] [CrossRef]
- Nakai, T.; Sugimoto, R.; Park, K.-H.; Matsuoka, S.; Mori, K.; Nishioka, T.; Maruyama, K. Protective Effects of Bacteriophage on Experimental Lactococcus garvieae Infection in Yellowtail. Dis. Aquat. Org. 1999, 37, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Ooyama, T.; Hirokawa, Y.; Minami, T.; Yasuda, H.; Nakai, T.; Endo, M.; Ruangpan, L.; Yoshida, T. Cell-Surface Properties of Lactococcus Garvieae Strains and Their Immunogenicity in the Yellowtail Seriola quinqueradiata. Dis. Aquat. Org. 2002, 51, 169–177. [Google Scholar] [CrossRef] [PubMed]
- Eraclio, G.; Tremblay, D.M.; Lacelle-Côté, A.; Labrie, S.J.; Fortina, M.G.; Moineau, S. A Virulent Phage Infecting Lactococcus garvieae, with Homology to Lactococcus lactis Phages. Appl. Environ. Microbiol. 2015, 81, 8358–8365. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hoai, T.D.; Nishiki, I.; Yoshida, T. Properties and Genomic Analysis of Lactococcus garvieae Lysogenic Bacteriophage PLgT-1, a New Member of Siphoviridae, with Homology to Lactococcus lactis Phages. Virus Res. 2016, 222, 13–23. [Google Scholar] [CrossRef]
- Hoai, T.; Yoshida, T. Induction and Characterization of a Lysogenic Bacteriophage of Lactococcus garvieae Isolated from Marine Fish Species. J. Fish Dis. 2016, 39, 799–808. [Google Scholar] [CrossRef]
- Hoai, T.D.; Nishiki, I.; Fujiwara, A.; Yoshida, T.; Nakai, T. Comparative Genomic Analysis of Three Lytic Lactococcus garvieae Phages, Novel Phages with Genome Architecture Linking the 936 Phage Species of Lactococcus lactis. Mar. Genom. 2019, 48, 100696. [Google Scholar] [CrossRef]
- Ghasemi, S.M.; Bouzari, M.; Yoon, B.H.; Chang, H.-I. Comparative Genomic Analysis of Lactococcus garvieae Phage WP-2, a New Member of Picovirinae Subfamily of Podoviridae. Gene 2014, 551, 222–229. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Liao, G.; Liu, C.; Jiang, X.; Lin, M.; Zhao, C.; Tao, J.; Huang, Z. Characterization of Bacteriophage HN 48 and Its Protective Effects in Nile Tilapia Oreochromis niloticus against Streptococcus agalactiae Infections. J. Fish Dis. 2018, 41, 1477–1484. [Google Scholar] [CrossRef] [PubMed]
- Wright, E.; Elliman, J.; Owens, L. Induction and Characterization of Lysogenic Bacteriophages from Streptococcus iniae. J. Appl. Microbiol. 2013, 114, 1616–1624. [Google Scholar] [CrossRef]
- Hoai, T.D.; Mitomi, K.; Nishiki, I.; Yoshida, T. A Lytic Bacteriophage of the Newly Emerging Rainbow Trout Pathogen Weissella Ceti. Virus Res. 2018, 247, 34–39. [Google Scholar] [CrossRef] [PubMed]
- Schulz, P.; Pajdak-Czaus, J.; Robak, S.; Dastych, J.; Siwicki, A.K. Bacteriophage-based Cocktail Modulates Selected Immunological Parameters and Post-challenge Survival of Rainbow Trout (Oncorhynchus mykiss). J. Fish Dis. 2019, 42, 1151–1160. [Google Scholar] [CrossRef]
- Moye, Z.D.; Woolston, J.; Abbeele, P.v.d.; Duysburgh, C.; Verstrepen, L.; Das, C.R.; Marzorati, M.; Sulakvelidze, A. A Bacteriophage Cocktail Eliminates Salmonella Typhimurium from the Human Colonic Microbiome While Preserving Cytokine Signaling and Preventing Attachment to and Invasion of Human Cells by Salmonella in vitro. J. Food Prot. 2019, 82, 1336–1349. [Google Scholar] [CrossRef] [PubMed]
- Moye, Z.D.; Woolston, J.; Sulakvelidze, A. Bacteriophage Applications for Food Production and Processing. Viruses 2018, 10, 205. [Google Scholar] [CrossRef]
- Fischer, S.; Kittler, S.; Klein, G.; Glünder, G. Impact of a Single Phage and a Phage Cocktail Application in Broilers on Reduction of Campylobacter Jejuni and Development of Resistance. PLoS ONE 2013, 8, e78543. [Google Scholar] [CrossRef]
- Sulakvelidze, A. Using Lytic Bacteriophages to Eliminate or Significantly Reduce Contamination of Food by Foodborne Bacterial Pathogens. J. Sci. Food Agric. 2013, 93, 3137–3146. [Google Scholar] [CrossRef]
- Hsu, C.; Lo, C.; Liu, J.; Lin, C. Control of the Eel (Anguilla Japonica) Pathogens, Aeromonas Hydrophila and Edwardsiella tarda, by Bacteriophages. J. Fish. Soc. Taiwan 2000, 27, 21–31. [Google Scholar]
- Blanco-Picazo, P.; Roscales, G.; Toribio-Avedillo, D.; Gómez-Gómez, C.; Avila, C.; Ballesté, E.; Muniesa, M.; Rodríguez-Rubio, L. Antibiotic Resistance Genes in Phage Particles from Antarctic and Mediterranean Seawater Ecosystems. Microorganisms 2020, 8, 1293. [Google Scholar] [CrossRef]
- Castillo, D.; Kauffman, K.; Hussain, F.; Kalatzis, P.; Rørbo, N.; Polz, M.F.; Middelboe, M. Widespread Distribution of Prophage-Encoded Virulence Factors in Marine Vibrio Communities. Sci. Rep. 2018, 8, 1–9. [Google Scholar] [CrossRef]
- Christiansen, R.H.; Madsen, L.; Dalsgaard, I.; Castillo, D.; Kalatzis, P.G.; Middelboe, M. Effect of Bacteriophages on the Growth of Flavobacterium psychrophilum and Development of Phage-Resistant Strains. Microb. Ecol. 2016, 71, 845–859. [Google Scholar] [CrossRef]
- Castillo, D.; Christiansen, R.H.; Dalsgaard, I.; Madsen, L.; Middelboe, M. Bacteriophage Resistance Mechanisms in the Fish Pathogen Flavobacterium psychrophilum: Linking Genomic Mutations to Changes in Bacterial Virulence Factors. Appl. Environ. Microbiol. 2015, 81, 1157–1167. [Google Scholar] [CrossRef]
- Middelboe, M.; Holmfeldt, K.; Riemann, L.; Nybroe, O.; Haaber, J. Bacteriophages Drive Strain Diversification in a Marine Flavobacterium: Implications for Phage Resistance and Physiological Properties. Environ. Microbiol. 2009, 11, 1971–1982. [Google Scholar] [CrossRef]
- Verner–Jeffreys, D.W.; Algoet, M.; Pond, M.J.; Virdee, H.K.; Bagwell, N.J.; Roberts, E.G. Furunculosis in Atlantic Salmon (Salmo salar L.) Is Not Readily Controllable by Bacteriophage Therapy. Aquaculture 2007, 270, 475–484. [Google Scholar] [CrossRef]
- Carrizo, J.C.; Griboff, J.; Bonansea, R.I.; Nimptsch, J.; Valdés, M.E.; Wunderlin, D.A.; Amé, M.V. Different Antibiotic Profiles in Wild and Farmed Chilean Salmonids. Which Is the Main Source for Antibiotic in Fish? Sci. Total. Environ. 2021, 800, 149516. [Google Scholar] [CrossRef] [PubMed]
- Firmino, J.P.; Galindo-Villegas, J.; Reyes-López, F.E.; Gisbert, E. Phytogenic bioactive compounds shape fish mucosal immunity. Front. Immunol. 2021, 12, 695973. [Google Scholar] [CrossRef] [PubMed]
- Siriyappagouder, P.; Galindo-Villegas, J.; Dhanasiri, A.K.; Zhang, Q.; Mulero, V.; Kiron, V.; Fernandes, J.M. Pseudozyma Priming Influences Expression of Genes Involved in Metabolic Pathways and Immunity in Zebrafish Larvae. Front. Immunol. 2020, 11, 978. [Google Scholar] [CrossRef]
- Monzón-Atienza, L.; Bravo, J.; Torrecillas, S.; Montero, D.; González-de Canales, A.F.; de la Banda, I.G.; Galindo-Villegas, J.; Ramos-Vivas, J.; Acosta, F. Isolation and Characterization of a Bacillus velezensis D-18 Strain, as a Potential Probiotic in European Seabass Aquaculture. Probiotics Antimicrob. Proteins 2021, 1–9. [Google Scholar]
- Montalban-Arques, A.; De Schryver, P.; Bossier, P.; Gorkiewicz, G.; Mulero, V.; Gatlin III, D.M.; Galindo-Villegas, J. Selective Manipulation of the Gut Microbiota Improves Immune Status in Vertebrates. Front. Immunol. 2015, 6, 512. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, M.; Sepahi, A.; Casadei, E.; Salinas, I. Symbiont-derived sphingolipids regulate inflammatory responses in rainbow trout (Oncorhynchus mykiss). Aquaculture 2018, 495, 932–939. [Google Scholar] [CrossRef] [PubMed]
- Galindo-Villegas, J.; Masumoto, T.; Hosokawa, H. Effect of Continuous and Interval Administration of Peptidoglycan on Innate Immune Response and Disease Resistance in Japanese Flounder Paralichthys olivaceus. Aquac. Sci. 2006, 54, 163–170. [Google Scholar]
- Rodrigues, M.V.; Zanuzzo, F.S.; Koch, J.F.A.; de Oliveira, C.A.F.; Sima, P.; Vetvicka, V. Development of Fish Immunity and the Role of β-Glucan in Immune Responses. Molecules 2020, 25, 5378. [Google Scholar] [CrossRef] [PubMed]
- Żaczek, M.; Weber-Dąbrowska, B.; Górski, A. Phages as a Cohesive Prophylactic and Therapeutic Approach in Aquaculture Systems. Antibiotics 2020, 9, 564. [Google Scholar] [CrossRef] [PubMed]
- Borin, J.M.; Avrani, S.; Barrick, J.E.; Petrie, K.L.; Meyer, J.R. Coevolutionary phage training leads to greater bacterial suppression and delays the evolution of phage resistance. Proc. Natl. Acad. Sci. USA 2021, 118, e2104592118. [Google Scholar] [CrossRef]
- Richards, G.P. Bacteriophage Remediation of Bacterial Pathogens in Aquaculture: A Review of the Technology. Bacteriophage 2014, 4, e975540. [Google Scholar] [CrossRef] [PubMed]
- Sieiro, C.; Areal-Hermida, L.; Pichardo-Gallardo, Á.; Almuiña-González, R.; de Miguel, T.; Sánchez, S.; Sánchez-Pérez, Á.; Villa, T.G. A Hundred Years of Bacteriophages: Can Phages Replace Antibiotics in Agriculture and Aquaculture? Antibiotics 2020, 9, 493. [Google Scholar] [CrossRef]
- Park, S.Y.; Han, J.E.; Kwon, H.; Park, S.C.; Kim, J.H. Recent Insights into Aeromonas salmonicida and Its Bacteriophages in Aquaculture: A Comprehensive Review. J. Microbiol. Biotechnol. 2020, 30, 1443–1457. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramos-Vivas, J.; Superio, J.; Galindo-Villegas, J.; Acosta, F. Phage Therapy as a Focused Management Strategy in Aquaculture. Int. J. Mol. Sci. 2021, 22, 10436. https://doi.org/10.3390/ijms221910436
Ramos-Vivas J, Superio J, Galindo-Villegas J, Acosta F. Phage Therapy as a Focused Management Strategy in Aquaculture. International Journal of Molecular Sciences. 2021; 22(19):10436. https://doi.org/10.3390/ijms221910436
Chicago/Turabian StyleRamos-Vivas, José, Joshua Superio, Jorge Galindo-Villegas, and Félix Acosta. 2021. "Phage Therapy as a Focused Management Strategy in Aquaculture" International Journal of Molecular Sciences 22, no. 19: 10436. https://doi.org/10.3390/ijms221910436
APA StyleRamos-Vivas, J., Superio, J., Galindo-Villegas, J., & Acosta, F. (2021). Phage Therapy as a Focused Management Strategy in Aquaculture. International Journal of Molecular Sciences, 22(19), 10436. https://doi.org/10.3390/ijms221910436








