Plasmidome of Listeria spp.—The repA-Family Business
Abstract
1. Introduction
2. Results and Discussion
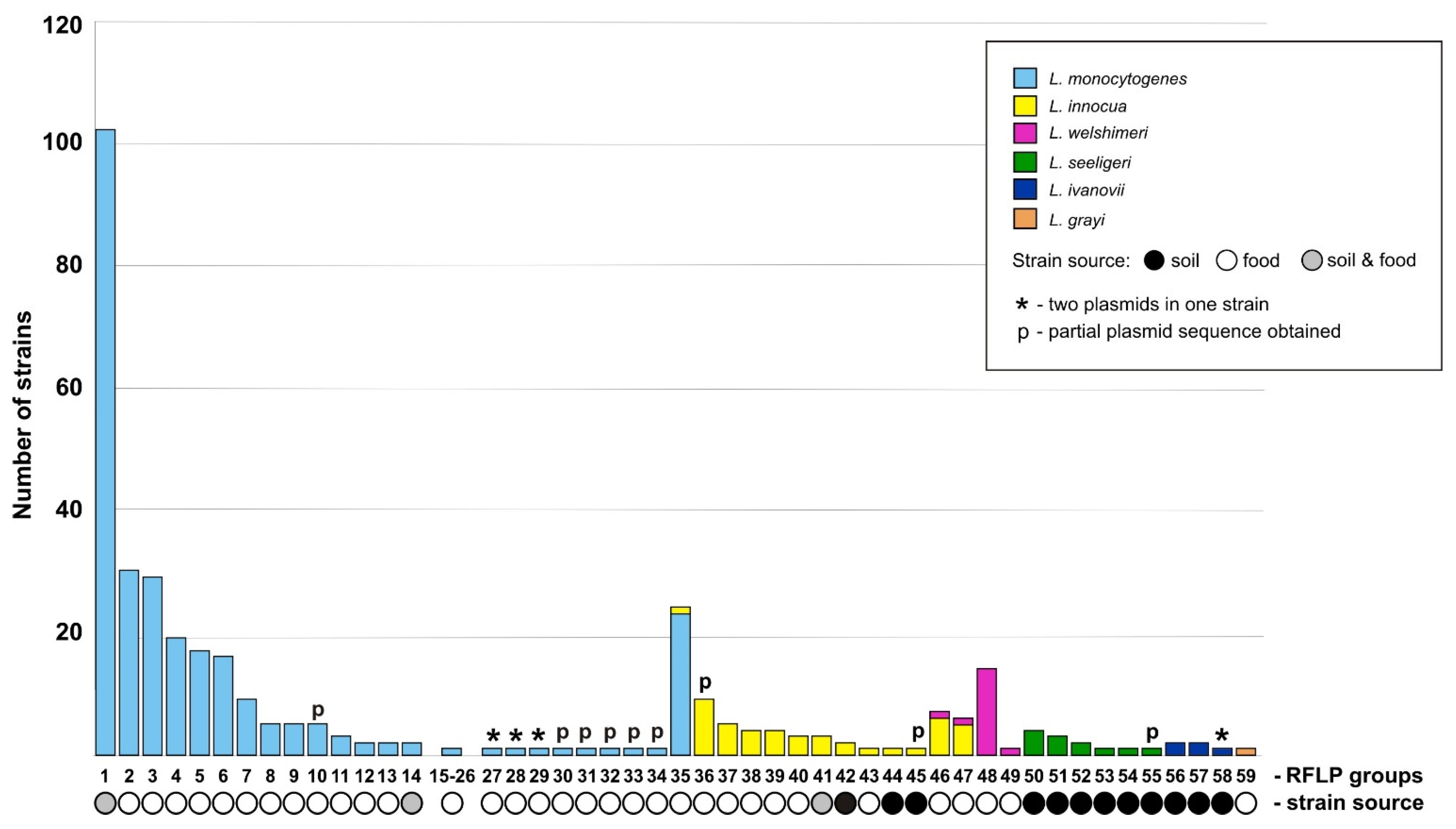
2.1. Identification and Classification of Plasmids Occurring in 782 Listeria spp. Strains
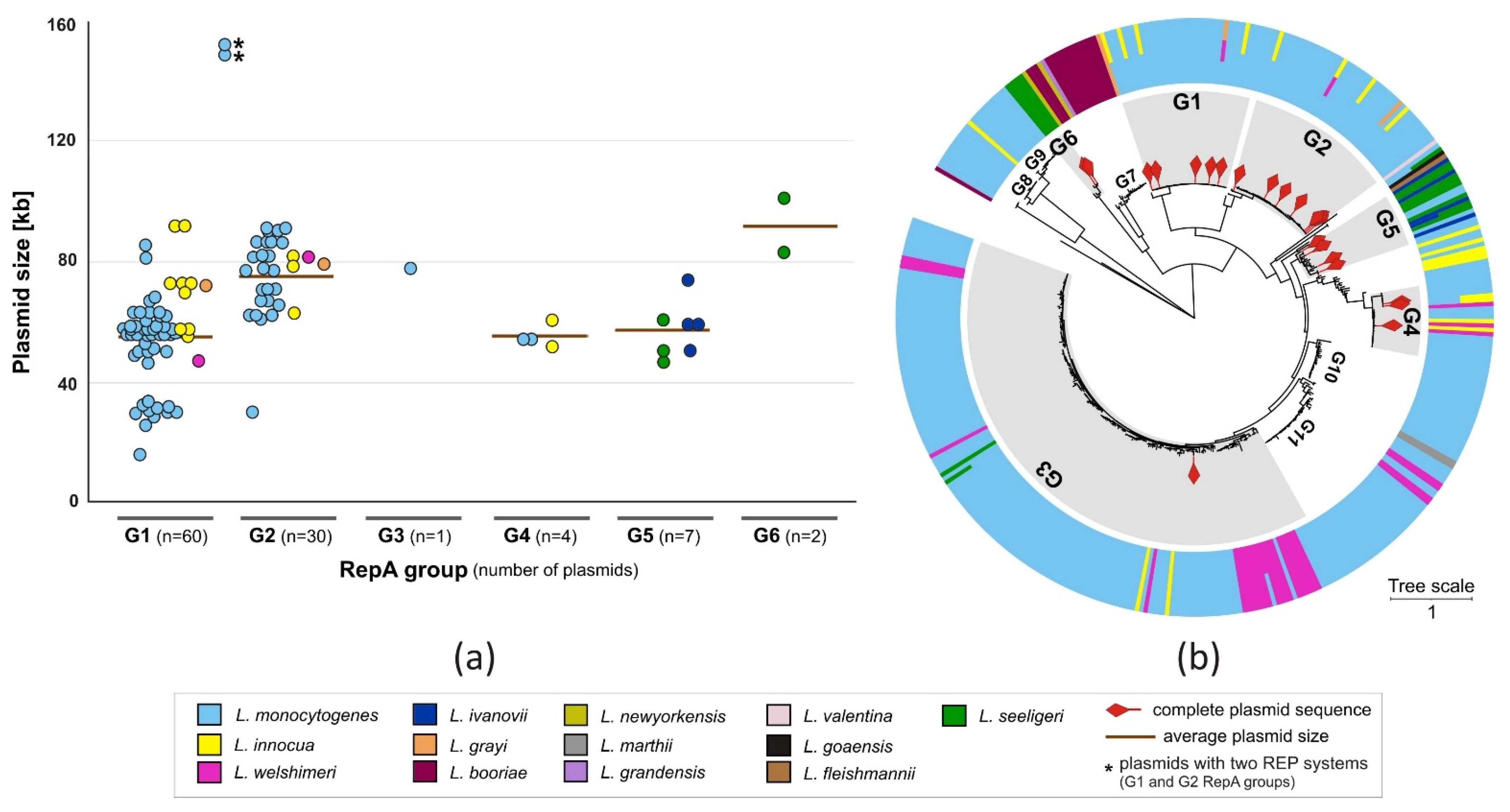
2.2. A Dataset of Fully Sequenced Listeria spp. Plasmids
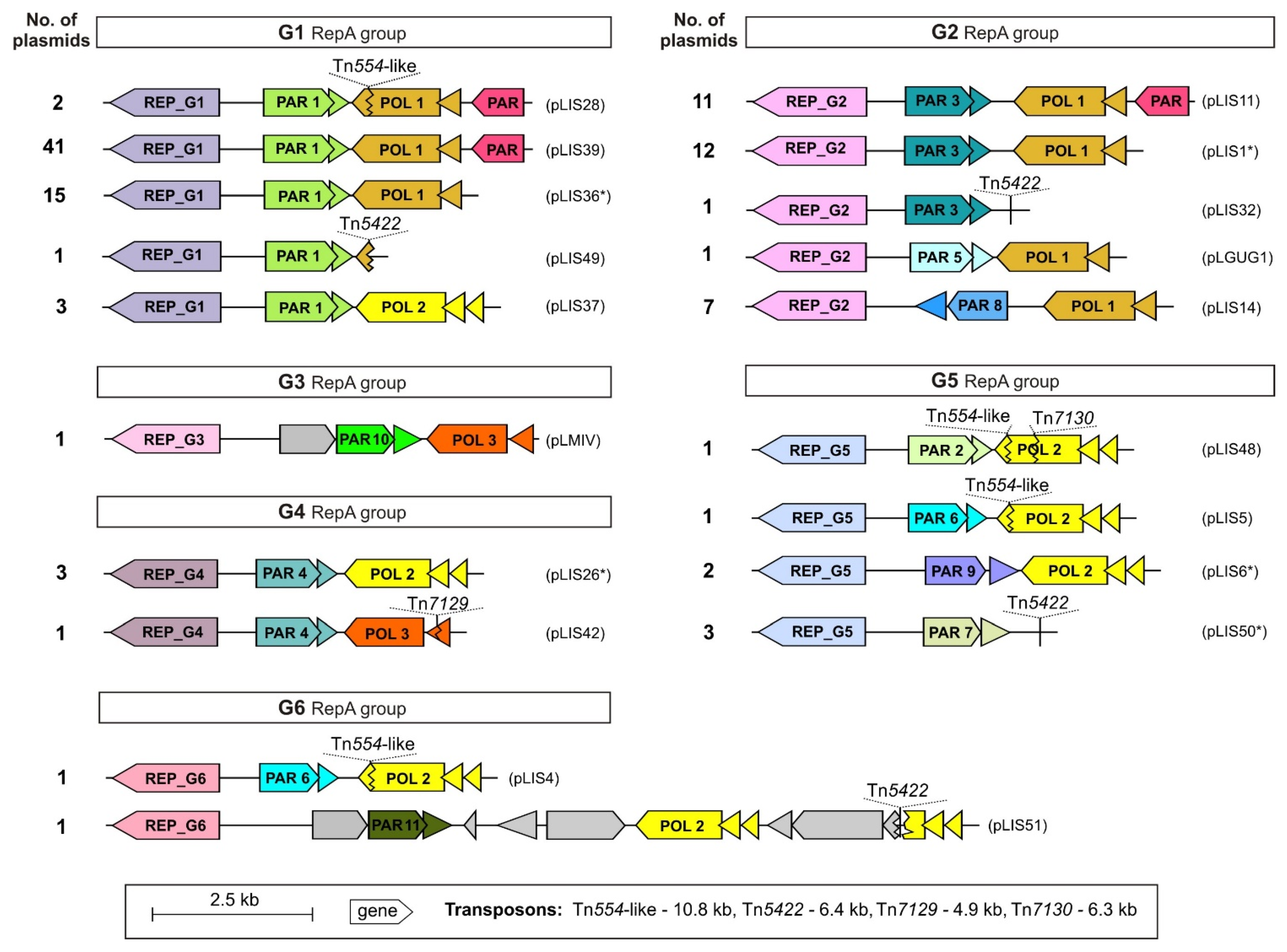
2.3. Conserved Backbone and Structure of the repA-Family Plasmids
2.3.1. Replication Systems
2.3.2. Stable Maintenance Systems
2.3.3. Y-Family DNA Polymerase Modules
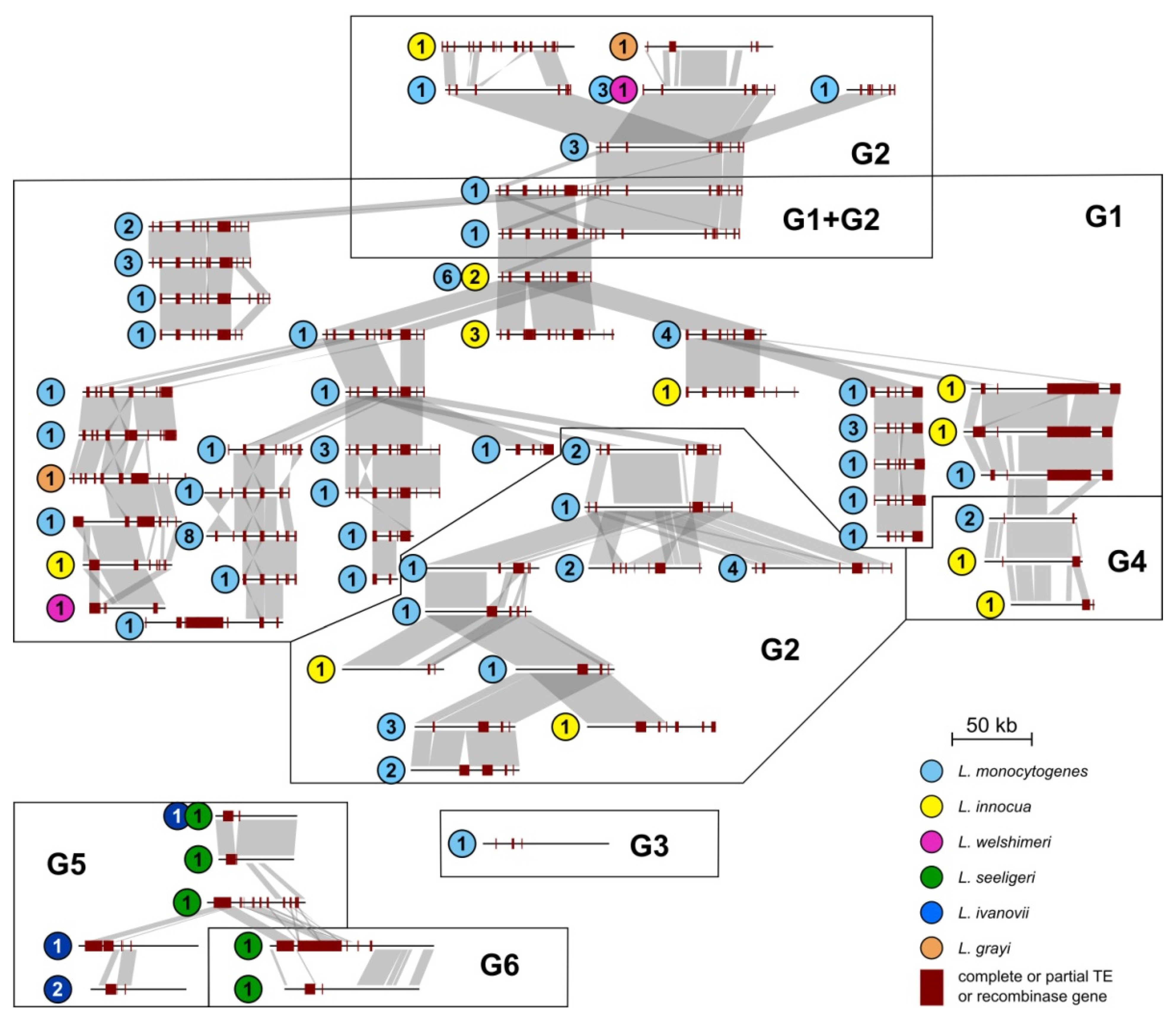
2.3.4. Conserved Structure of the repA-Family Plasmids
2.4. Conjugative Transfer Modules of Listeria spp. Plasmids
2.5. Adaptive Genes of Listeria spp. Plasmids
2.5.1. Heavy Metal and Metalloid Resistance
2.5.2. Sanitizer Resistance
2.5.3. Antibiotic Resistance
2.5.4. Heat Stress
2.5.5. Oxidative, Acid and High Salinity Stresses
2.5.6. Other Genes of Potential Adaptive Value
2.5.7. Putative Virulence Factors
2.6. Listeria Plasmids Pangenome Estimation
3. Materials and Methods
3.1. Bacterial Strains and Growth Conditions
3.2. Plasmid DNA Isolation and Restriction Fragment Length Polymorphism (RFLP) Profile Analysis
3.3. DNA Sequencing
3.4. Heavy Metal, Metalloid, Benzalkonium Chloride and Crystal Violet Susceptibility
3.5. Antimicrobial Susceptibility
3.6. Shuttle Vector Construction
3.7. Assessing Host Range of Shuttle Vectors
3.8. Bioinformatic Analyses
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Carlin, C.R.; Liao, J.; Weller, D.; Guo, X.; Orsi, R.; Wiedmann, M. Listeria cossartiae sp. nov., Listeria immobilis sp. nov., Listeria portnoyi sp. nov. and Listeria rustica sp. nov., isolated from agricultural water and natural environments. Int. J. Syst. Evol. Microbiol. 2021, 71, 004795. [Google Scholar] [CrossRef]
- Bergholz, T.M.; den Bakker, H.C.; Fortes, E.D.; Boor, K.J.; Wiedmann, M. Salt stress phenotypes in Listeria monocytogenes vary by genetic lineage and temperature. Foodborne Pathog. Dis. 2010, 7, 1537–1549. [Google Scholar] [CrossRef] [PubMed]
- Durack, J.; Ross, T.; Bowman, J.P. Characterisation of the transcriptomes of genetically diverse Listeria monocytogenes exposed to hyperosmotic and low temperature conditions reveal global stress-adaptation mechanisms. PLoS ONE 2013, 8, e73603. [Google Scholar] [CrossRef] [PubMed]
- Korsak, D.; Szuplewska, M. Characterization of nonpathogenic Listeria species isolated from food and food processing environment. Int. J. Food Microbiol. 2016, 238, 274–280. [Google Scholar] [CrossRef] [PubMed]
- Hingston, P.; Chen, J.; Dhillon, B.K.; Laing, C.; Bertelli, C.; Gannon, V.; Tasara, T.; Allen, K.; Brinkman, F.S.L.; Truelstrup Hansen, L.; et al. Genotypes Associated with Listeria monocytogenes Isolates Displaying Impaired or Enhanced Tolerances to Cold, Salt, Acid, or Desiccation Stress. Front. Microbiol. 2017, 8, 369. [Google Scholar] [CrossRef]
- Charpentier, E.; Gerbaud, G.; Courvalin, P. Conjugative mobilization of the rolling-circle plasmid pIP823 from Listeria monocytogenes BM4293 among gram-positive and gram-negative bacteria. J. Bacteriol. 1999, 181, 3368–3374. [Google Scholar] [CrossRef]
- Yan, H.; Yu, R.; Li, D.; Shi, L.; Schwarz, S.; Yao, H.; Li, X.-S.; Du, X.-D. A novel multiresistance gene cluster located on a plasmid-borne transposon in Listeria monocytogenes. J. Antimicrob. Chemother. 2020, 75, 868–872. [Google Scholar] [CrossRef]
- Kremer, P.H.C.; Lees, J.A.; Koopmans, M.M.; Ferwerda, B.; Arends, A.W.M.; Feller, M.M.; Schipper, K.; Valls Seron, M.; van der Ende, A.; Brouwer, M.C.; et al. Benzalkonium tolerance genes and outcome in Listeria monocytogenes meningitis. Clin. Microbiol. Infect. 2017, 23, 265.e1–265.e7. [Google Scholar] [CrossRef] [PubMed]
- Elhanafi, D.; Dutta, V.; Kathariou, S. Genetic characterization of plasmid-associated benzalkonium chloride resistance determinants in a Listeria monocytogenes strain from the 1998–1999 outbreak. Appl. Environ. Microbiol. 2010, 76, 8231–8238. [Google Scholar] [CrossRef]
- Lebrun, M.; Audurier, A.; Cossart, P. Plasmid-borne cadmium resistance genes in Listeria monocytogenes are present on Tn5422, a novel transposon closely related to Tn917. J. Bacteriol. 1994, 176, 3049–3061. [Google Scholar] [CrossRef] [PubMed]
- Dutta, V.; Elhanafi, D.; Kathariou, S. Conservation and distribution of the benzalkonium chloride resistance cassette bcrABC in Listeria monocytogenes. Appl. Environ. Microbiol. 2013, 79, 6067–6074. [Google Scholar] [CrossRef] [PubMed]
- Kuenne, C.; Voget, S.; Pischimarov, J.; Oehm, S.; Goesmann, A.; Daniel, R.; Hain, T.; Chakraborty, T. Comparative analysis of plasmids in the genus Listeria. PLoS ONE 2010, 5, e12511. [Google Scholar] [CrossRef] [PubMed]
- Pöntinen, A.; Aalto-Araneda, M.; Lindström, M.; Korkeala, H. Heat Resistance Mediated by pLM58 Plasmid-Borne ClpL in Listeria monocytogenes. mSphere 2017, 2, e00364-17. [Google Scholar] [CrossRef]
- Fox, E.M.; Allnutt, T.; Bradbury, M.I.; Fanning, S.; Chandry, P.S. Comparative genomics of the Listeria monocytogenes ST204 subgroup. Front. Microbiol. 2016, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Schmitz-Esser, S.; Anast, J.M.; Cortes, B.W. A large-scale sequencing-based survey of plasmids in Listeria monocytogenes reveals global dissemination of plasmids. Front. Microbiol. 2021, 12, 653155. [Google Scholar] [CrossRef]
- Hingston, P.; Brenner, T.; Truelstrup Hansen, L.; Wang, S. Comparative analysis of Listeria monocytogenes plasmids and expression levels of plasmid-encoded genes during growth under salt and acid stress conditions. Toxins 2019, 11, 426. [Google Scholar] [CrossRef]
- Rychli, K.; Wagner, E.M.; Ciolacu, L.; Zaiser, A.; Tasara, T.; Wagner, M.; Schmitz-Esser, S. Comparative genomics of human and non-human Listeria monocytogenes sequence type 121 strains. PLoS ONE 2017, 12, e0176857. [Google Scholar] [CrossRef]
- Palma, F.; Brauge, T.; Radomski, N.; Mallet, L.; Felten, A.; Mistou, M.-Y.; Brisabois, A.; Guillier, L.; Midelet-Bourdin, G. Dynamics of mobile genetic elements of Listeria monocytogenes persisting in ready-to-eat seafood processing plants in France. BMC Genom. 2020, 21, 130. [Google Scholar] [CrossRef]
- Schmitz-Esser, S.; Müller, A.; Stessl, B.; Wagner, M. Genomes of sequence type 121 Listeria monocytogenes strains harbor highly conserved plasmids and prophages. Front. Microbiol. 2015, 6, 380. [Google Scholar] [CrossRef]
- Korsak, D.; Chmielowska, C.; Szuplewska, M.; Bartosik, D. Prevalence of plasmid-borne benzalkonium chloride resistance cassette bcrABC and cadmium resistance cadA genes in nonpathogenic Listeria spp. isolated from food and food-processing environments. Int. J. Food Microbiol. 2019, 290, 247–253. [Google Scholar] [CrossRef]
- Katharios-Lanwermeyer, S.; Rakic-Martinez, M.; Elhanafi, D.; Ratani, S.; Tiedje, J.M.; Kathariou, S. Coselection of cadmium and benzalkonium chloride resistance in conjugative transfers from nonpathogenic Listeria spp. to other Listeriae. Appl. Environ. Microbiol. 2012, 78, 7549–7556. [Google Scholar] [CrossRef]
- Bertsch, D.; Anderegg, J.; Lacroix, C.; Meile, L.; Stevens, M.J.A. PDB2011, a 7.6kb multidrug resistance plasmid from Listeria innocua replicating in Gram-positive and Gram-negative hosts. Plasmid 2013, 70, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Chmielowska, C.; Korsak, D.; Szmulkowska, B.; Krop, A.; Lipka, K.; Krupińska, M.; Bartosik, D. Genetic carriers and genomic distribution of cadA6—A novel variant of a cadmium resistance determinant identified in Listeria spp. Int. J. Mol. Sci. 2020, 21, 8713. [Google Scholar] [CrossRef]
- Chmielowska, C.; Korsak, D.; Szuplewska, M.; Grzelecka, M.; Maćkiw, E.; Stasiak, M.; Macion, A.; Skowron, K.; Bartosik, D. Benzalkonium chloride and heavy metal resistance profiles of Listeria monocytogenes strains isolated from fish, fish products and food-producing factories in Poland. Food Microbiol. 2021, 98, 103756. [Google Scholar] [CrossRef] [PubMed]
- Fagerlund, A.; Langsrud, S.; Moen, B.; Heir, E.; Møretrø, T. Complete genome sequences of six Listeria monocytogenes sequence type 9 isolates from meat processing plants in Norway. Genome Announc. 2018, 6, e00016-18. [Google Scholar] [CrossRef]
- Orsini, M.; Ordinelli, A.; Cornacchia, A.; Acciari, V.; Centorame, P.; Torresi, M.; Pompei, A.; Marcacci, M.; Ancora, M.; Di Domenico, M.; et al. Whole-genome sequence of Listeria monocytogenes serovar 1/2a strain IZSAM_Lm_15_17439_A144, representative of a human outbreak in 2008. Microbiol. Resour. Announc. 2019, 8, e00074-19. [Google Scholar] [CrossRef] [PubMed]
- Lanza, V.F.; Tedim, A.P.; Martínez, J.L.; Baquero, F.; Coque, T.M. The plasmidome of firmicutes: Impact on the emergence and the spread of resistance to antimicrobials. Microbiol. Spectr. 2015, 3, 1–37. [Google Scholar] [CrossRef] [PubMed]
- Halbedel, S.; Wilking, H.; Holzer, A.; Kleta, S.; Fischer, M.A.; Lüth, S.; Pietzka, A.; Huhulescu, S.; Lachmann, R.; Krings, A.; et al. Large nationwide outbreak of invasive listeriosis associated with blood sausage, Germany, 2018–2019. Emerg. Infect. Dis. 2020, 26, 1456–1464. [Google Scholar] [CrossRef]
- Castro, H.; Douillard, F.P.; Korkeala, H.; Lindström, M. Mobile elements harboring heavy metal and bacitracin resistance genes are common among Listeria monocytogenes strains persisting on dairy farms. mSphere 2021, 6, e0038321. [Google Scholar] [CrossRef]
- Kamruzzaman, M.; Wu, A.Y.; Iredell, J.R. Biological functions of type II toxin-antitoxin systems in bacteria. Microorganisms 2021, 9, 1276. [Google Scholar] [CrossRef]
- Agüero, J.A.; Akarsu, H.; Aguilar-Bultet, L.; Oevermann, A.; Falquet, L. Large-scale comparison of toxin and antitoxins in Listeria monocytogenes. Toxins 2020, 12, 29. [Google Scholar] [CrossRef] [PubMed]
- Curtis, T.; Takeuchi, I.; Gram, L.; Knudsen, G. The influence of the toxin/antitoxin mazEF on growth and survival of Listeria monocytogenes under stress. Toxins 2017, 9, 31. [Google Scholar] [CrossRef] [PubMed]
- Kalani, B.S.; Irajian, G.; Lotfollahi, L.; Abdollahzadeh, E.; Razavi, S. Putative type II toxin-antitoxin systems in Listeria monocytogenes isolated from clinical, food, and animal samples in Iran. Microb. Pathog. 2018, 122, 19–24. [Google Scholar] [CrossRef] [PubMed]
- Permina, E.A.; Mironov, A.A.; Gelfand, M.S. Damage-repair error-prone polymerases of eubacteria: Association with mobile genome elements. Gene 2002, 293, 133–140. [Google Scholar] [CrossRef]
- Timinskas, K.; Venclovas, Č. New insights into the structures and interactions of bacterial Y-family DNA polymerases. Nucleic Acids Res. 2019, 47, 4393–4405. [Google Scholar] [CrossRef]
- Yang, W. An overview of Y-family DNA polymerases and a case study of human DNA Polymerase η. Biochemistry 2014, 53, 2793–2803. [Google Scholar] [CrossRef]
- Koskiniemi, S.; Andersson, D.I. Translesion DNA polymerases are required for spontaneous deletion formation in Salmonella typhimurium. Proc. Natl. Acad. Sci. USA 2009, 106, 10248–10253. [Google Scholar] [CrossRef]
- Van der Veen, S.; van Schalkwijk, S.; Molenaar, D.; de Vos, W.M.; Abee, T.; Wells-Bennik, M.H.J. The SOS response of Listeria monocytogenes is involved in stress resistance and mutagenesis. Microbiology 2010, 156, 374–384. [Google Scholar] [CrossRef] [PubMed]
- Mao, P.; Wang, Y.; Gan, L.; Sun, H.; Wang, Y.; Li, L.; Ji, S.; Song, Z.; Jiang, H.; Ye, C. Function and distribution of the conjugative plasmid pLM1686 in foodborne Listeria monocytogenes in China. Int. J. Food Microbiol. 2021, 352, 109261. [Google Scholar] [CrossRef]
- Luo, L.; Chen, X.; Payne, M.; Cao, X.; Wang, Y.; Zhang, J.; Deng, J.; Wang, H.; Zhang, Z.; Li, Q.; et al. Case report: Whole genome sequencing based investigation of maternal-neonatal listeriosis in Sichuan, China. BMC Infect. Dis. 2019, 19, 893. [Google Scholar] [CrossRef]
- Den Bakker, H.C.; Bowen, B.M.; Rodriguez-Rivera, L.D.; Wiedmann, M. FSL J1-208, a virulent uncommon phylogenetic lineage IV Listeria monocytogenes strain with a small chromosome size and a putative virulence plasmid carrying internalin-like genes. Appl. Environ. Microbiol. 2012, 78, 1876–1889. [Google Scholar] [CrossRef] [PubMed]
- Ramachandran, G.; Miguel-Arribas, A.; Abia, D.; Singh, P.K.; Crespo, I.; Gago-Córdoba, C.; Hao, J.A.; Luque-Ortega, J.R.; Alfonso, C.; Wu, L.J.; et al. Discovery of a new family of relaxases in Firmicutes bacteria. PLoS Genet. 2017, 13, e1006586. [Google Scholar] [CrossRef] [PubMed]
- Lebrun, M.; Audurier, A.; Cossart, P. Plasmid-borne cadmium resistance genes in Listeria monocytogenes are similar to cadA and cadC of Staphylococcus aureus and are induced by cadmium. J. Bacteriol. 1994, 176, 3040–3048. [Google Scholar] [CrossRef]
- Kuenne, C.; Billion, A.; Mraheil, M.A.; Strittmatter, A.; Daniel, R.; Goesmann, A.; Barbuddhe, S.; Hain, T.; Chakraborty, T. Reassessment of the Listeria monocytogenes pan-genome reveals dynamic integration hotspots and mobile genetic elements as major components of the accessory genome. BMC Genom. 2013, 14, 47. [Google Scholar] [CrossRef] [PubMed]
- Parsons, C.; Lee, S.; Jayeola, V.; Kathariou, S. Novel cadmium resistance determinant in Listeria monocytogenes. Appl. Environ. Microbiol. 2017, 83, e02580-16. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Ward, T.J.; Jima, D.D.; Parsons, C.; Kathariou, S. The arsenic resistance-associated Listeria genomic island LGI2 exhibits sequence and integration site diversity and a propensity for three Listeria monocytogenes clones with enhanced virulence. Appl. Environ. Microbiol. 2017, 83, e01189-17. [Google Scholar] [CrossRef]
- Lee, S.; Parsons, C.; Chen, Y.; Hanafy, Z.; Brown, E.; Kathariou, S. Identification and characterization of a novel genomic island harboring cadmium and arsenic resistance genes in Listeria welshimeri. Biomolecules 2021, 11, 560. [Google Scholar] [CrossRef]
- Crupper, S.S.; Worrell, V.; Stewart, G.C.; Iandolo, J.J. Cloning and expression of cadD, a new cadmium resistance gene of Staphylococcus aureus. J. Bacteriol. 1999, 181, 4071–4075. [Google Scholar] [CrossRef]
- Ji, G.; Silver, S. Regulation and expression of the arsenic resistance operon from Staphylococcus aureus plasmid pI258. J. Bacteriol. 1992, 174, 3684–3694. [Google Scholar] [CrossRef]
- Francis, M.S.; Thomas, C.J. Mutants in the CtpA copper transporting P-type ATPase reduce virulence of Listeria monocytogenes. Microb. Pathog. 1997, 22, 67–78. [Google Scholar] [CrossRef]
- Francis, M.S.; Thomas, C.J. The Listeria monocytogenes gene ctpA encodes a putative P-type ATPase involved in copper transport. Mol. Gen. Genet. MGG 1997, 253, 484–491. [Google Scholar] [CrossRef]
- Bell, F.Y. Copper Tolerance of Listeria monocytogenes Strain DRDC8. Ph.D. Thesis, University of Adelaide, Adelaide, Australia, 2010. [Google Scholar]
- Zapotoczna, M.; Riboldi, G.P.; Moustafa, A.M.; Dickson, E.; Narechania, A.; Morrissey, J.A.; Planet, P.J.; Holden, M.T.G.; Waldron, K.J.; Geoghegan, J.A. Mobile-genetic-element-encoded hypertolerance to copper protects Staphylococcus aureus from killing by host phagocytes. MBio 2018, 9, e00550-18. [Google Scholar] [CrossRef]
- Cortes, B.W.; Naditz, A.L.; Anast, J.M.; Schmitz-Esser, S. Transcriptome sequencing of Listeria monocytogenes reveals major gene expression changes in response to lactic acid stress exposure but a less pronounced response to oxidative stress. Front. Microbiol. 2020, 10, 3110. [Google Scholar] [CrossRef]
- Mullapudi, S.; Siletzky, R.M.; Kathariou, S. Heavy-metal and benzalkonium chloride resistance of Listeria monocytogenes isolates from the environment of Turkey-processing plants. Appl. Environ. Microbiol. 2008, 74, 1464–1468. [Google Scholar] [CrossRef]
- Ratani, S.S.; Siletzky, R.M.; Dutta, V.; Yildirim, S.; Osborne, J.A.; Lin, W.; Hitchins, A.D.; Ward, T.J.; Kathariou, S. Heavy metal and disinfectant resistance of Listeria monocytogenes from foods and food processing plants. Appl. Environ. Microbiol. 2012, 78, 6938–6945. [Google Scholar] [CrossRef]
- Pirone-Davies, C.; Chen, Y.; Pightling, A.; Ryan, G.; Wang, Y.; Yao, K.; Hoffmann, M.; Allard, M.W. Genes significantly associated with lineage II food isolates of Listeria monocytogenes. BMC Genom. 2018, 19, 708. [Google Scholar] [CrossRef] [PubMed]
- Müller, A.; Rychli, K.; Muhterem-Uyar, M.; Zaiser, A.; Stessl, B.; Guinane, C.M.; Cotter, P.D.; Wagner, M.; Schmitz-Esser, S. Tn6188—A novel transposon in Listeria monocytogenes responsible for tolerance to benzalkonium chloride. PLoS ONE 2013, 8, e76835. [Google Scholar] [CrossRef] [PubMed]
- Poyart-Salmeron, C.; Carlier, C.; Trieu-Cuot, P.; Courtieu, A.; Courvalin, P. Transferable plasmid-mediated antibiotic resistance in Listeria monocytogenes. Lancet 1990, 335, 1422–1426. [Google Scholar] [CrossRef]
- Charpentier, E.; Courvalin, P. Emergence of the trimethoprim resistance gene dfrD in Listeria monocytogenes BM4293. Antimicrob. Agents Chemother. 1997, 41, 1134–1136. [Google Scholar] [CrossRef]
- Hadorn, K.; Hächler, H.; Schaffner, A.; Kayser, F.H. Genetic characterization of plasmid-encoded multiple antibiotic resistance in a strain of Listeria monocytogenes causing endocarditis. Eur. J. Clin. Microbiol. Infect. Dis. 1993, 12, 928–937. [Google Scholar] [CrossRef]
- Bertsch, D.; Uruty, A.; Anderegg, J.; Lacroix, C.; Perreten, V.; Meile, L. Tn6198, a novel transposon containing the trimethoprim resistance gene dfrG embedded into a Tn916 element in Listeria monocytogenes. J. Antimicrob. Chemother. 2013, 68, 986–991. [Google Scholar] [CrossRef]
- Naditz, A.L.; Dzieciol, M.; Wagner, M.; Schmitz-Esser, S. Plasmids contribute to food processing environment–associated stress survival in three Listeria monocytogenes ST121, ST8, and ST5 strains. Int. J. Food Microbiol. 2019, 299, 39–46. [Google Scholar] [CrossRef]
- Patchett, R.A.; Kelly, A.F.; Kroll, R.G. Effect of sodium chloride on the intracellular solute pools of Listeria monocytogenes. Appl. Environ. Microbiol. 1992, 58, 3959–3963. [Google Scholar] [CrossRef] [PubMed]
- Beumer, R.R.; Te Giffel, M.C.; Cox, L.J.; Rombouts, F.M.; Abee, T. Effect of exogenous proline, betaine, and carnitine on growth of Listeria monocytogenes in a minimal medium. Appl. Environ. Microbiol. 1994, 60, 1359–1363. [Google Scholar] [CrossRef]
- Ko, R.; Smith, L.T.; Smith, G.M. Glycine betaine confers enhanced osmotolerance and cryotolerance on Listeria monocytogenes. J. Bacteriol. 1994, 176, 426–431. [Google Scholar] [CrossRef] [PubMed]
- Ko, R.; Smith, L.T. Identification of an ATP-driven, osmoregulated glycine betaine transport system in Listeria monocytogenes. Appl. Environ. Microbiol. 1999, 65, 4040–4048. [Google Scholar] [CrossRef] [PubMed]
- Brøndsted, L.; Kallipolitis, B.H.; Ingmer, H.; Knöchel, S. kdpE and a putative RsbQ homologue contribute to growth of Listeria monocytogenes at high osmolarity and low temperature. FEMS Microbiol. Lett. 2003, 219, 233–239. [Google Scholar] [CrossRef]
- Ballal, A.; Basu, B.; Apte, S.K. The Kdp-ATPase system and its regulation. J. Biosci. 2007, 32, 559–568. [Google Scholar] [CrossRef]
- Dutta, V.; Elhanafi, D.; Osborne, J.; Martinez, M.R.; Kathariou, S. Genetic characterization of plasmid-associated triphenylmethane reductase in Listeria monocytogenes. Appl. Environ. Microbiol. 2014, 80, 5379–5385. [Google Scholar] [CrossRef]
- Verdon, E.; Bessiral, M.; Chotard, M.-P.; Couëdor, P.; Fourmond, M.-P.; Fuselier, R.; Gaugain, M.; Gautier, S.; Hurtaud-Pessel, D.; Laurentie, M.; et al. The monitoring of triphenylmethane dyes in aquaculture products through the european union network of official control laboratories. J. AOAC Int. 2015, 98, 649–657. [Google Scholar] [CrossRef]
- Kalmokoff, M.L.; Banerjee, S.K.; Cyr, T.; Hefford, M.A.; Gleeson, T. Identification of a new plasmid-encodedsec-dependent bacteriocin produced by Listeria innocua 743. Appl. Environ. Microbiol. 2001, 67, 4041–4047. [Google Scholar] [CrossRef]
- Kropac, A.C.; Eshwar, A.K.; Stephan, R.; Tasara, T. New insights on the role of the pLMST6 plasmid in Listeria monocytogenes biocide tolerance and virulence. Front. Microbiol. 2019, 10, 1538. [Google Scholar] [CrossRef]
- Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 1983, 166, 557–580. [Google Scholar] [CrossRef]
- Platt, R.; Drescher, C.; Park, S.-K.; Phillips, G.J. Genetic system for reversible integration of DNA constructs and lacZ gene fusions into the Escherichia coli chromosome. Plasmid 2000, 43, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Demarre, G.; Guérout, A.-M.; Matsumoto-Mashimo, C.; Rowe-Magnus, D.A.; Marlière, P.; Mazel, D. A new family of mobilizable suicide plasmids based on broad host range R388 plasmid (IncW) and RP4 plasmid (IncPα) conjugative machineries and their cognate Escherichia coli host strains. Res. Microbiol. 2005, 156, 245–255. [Google Scholar] [CrossRef]
- Bishop, D.K.; Hinrichs, D.J. Adoptive transfer of immunity to Listeria monocytogenes. The influence of in vitro stimulation on lymphocyte subset requirements. J. Immunol. 1987, 139, 2005–2009. [Google Scholar]
- Burkholder, P.R.; Giles, N.H. Induced biochemical mutations in Bacillus subtilis. Am. J. Bot. 1947, 34, 345–348. [Google Scholar] [CrossRef] [PubMed]
- Bartosik, D.; Baj, J.; Bartosik, A.A.; Wlodarczyk, M. Characterization of the replicator region of megaplasmid pTAV3 of Paracoccus versutus and search for plasmid-encoded traits. Microbiology 2002, 148, 871–881. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Anderson, D.G.; McKay, L.L. Simple and rapid method for isolating large plasmid DNA from lactic streptococci. Appl. Environ. Microbiol. 1983, 46, 549–552. [Google Scholar] [CrossRef]
- Wick, R.R.; Judd, L.M.; Gorrie, C.L.; Holt, K.E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 2017, 13, e1005595. [Google Scholar] [CrossRef]
- Wattam, A.R.; Davis, J.J.; Assaf, R.; Boisvert, S.; Brettin, T.; Bun, C.; Conrad, N.; Dietrich, E.M.; Disz, T.; Gabbard, J.L.; et al. Improvements to PATRIC, the all-bacterial bioinformatics database and analysis resource center. Nucleic Acids Res. 2017, 45, D535–D542. [Google Scholar] [CrossRef] [PubMed]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed]
- Rutherford, K.; Parkhill, J.; Crook, J.; Horsnell, T.; Rice, P.; Rajandream, M.-A.; Barrell, B. Artemis: Sequence visualization and annotation. Bioinformatics 2000, 16, 944–945. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, L.; Stephens, A.; Nam, S.-Z.; Rau, D.; Kübler, J.; Lozajic, M.; Gabler, F.; Söding, J.; Lupas, A.N.; Alva, V. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 2018, 430, 2237–2243. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef]
- Gilchrist, C.L.M.; Chooi, Y.-H. clinker & clustermap.js: Automatic generation of gene cluster comparison figures. Bioinformatics 2021, 37, 2473–2475. [Google Scholar] [CrossRef]
- Siguier, P.; Perochon, J.; Lestrade, L.; Mahillon, J.; Chandler, M. ISfinder: The reference centre for bacterial insertion sequences. Nucleic Acids Res. 2006, 34, D32–D36. [Google Scholar] [CrossRef] [PubMed]
- Tansirichaiya, S.; Rahman, M.A.; Roberts, A.P. The Transposon Registry. Mob. DNA 2019, 10, 40. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef] [PubMed]
- Tonkin-Hill, G.; MacAlasdair, N.; Ruis, C.; Weimann, A.; Horesh, G.; Lees, J.A.; Gladstone, R.A.; Lo, S.; Beaudoin, C.; Floto, R.A.; et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 2020, 21, 180. [Google Scholar] [CrossRef]
- Galperin, M.Y.; Makarova, K.S.; Wolf, Y.I.; Koonin, E.V. Expanded microbial genome coverage and improved protein family annotation in the COG database. Nucleic Acids Res. 2015, 43, D261–D269. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Jacomy, M.; Venturini, T.; Heymann, S.; Bastian, M. ForceAtlas2, a continuous graph layout algorithm for handy network visualization designed for the Gephi software. PLoS ONE 2014, 9, e98679. [Google Scholar] [CrossRef]
- Bastian, M.; Heymann, S.; Jacomy, M. Gephi: An open source software for exploring and manipulating networks. In Proceedings of the International AAAI Conference on Web and Social Media, San Jose, CA, USA, 17–20 May 2009; Volume 3, pp. 361–362. [Google Scholar]
- Bruand, C.; Le Chatelier, E.; Ehrlich, S.D.; Janniere, L. A fourth class of theta-replicating plasmids: The pAM beta 1 family from gram-positive bacteria. Proc. Natl. Acad. Sci. USA 1993, 90, 11668–11672. [Google Scholar] [CrossRef] [PubMed]
- Flamm, R.K.; Hinrichs, D.J.; Thomashow, M.F. Introduction of pAM beta 1 into Listeria monocytogenes by conjugation and homology between native L. monocytogenes plasmids. Infect. Immun. 1984, 44, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Vicente, M.F.; Baquero, F.; Pérez-Diaz, J.C. Conjugative acquisition and expression of antibiotic resistance determinants in Listeria spp. J. Antimicrob. Chemother. 1988, 21, 309–318. [Google Scholar] [CrossRef]
- den Bakker, H.C.; Cummings, C.A.; Ferreira, V.; Vatta, P.; Orsi, R.H.; Degoricija, L.; Barker, M.; Petrauskene, O.; Furtado, M.R.; Wiedmann, M. Comparative genomics of the bacterial genus Listeria: Genome evolution is characterized by limited gene acquisition and limited gene loss. BMC Genom. 2010, 11, 688. [Google Scholar] [CrossRef]
- Pombinho, R.; Camejo, A.; Vieira, A.; Reis, O.; Carvalho, F.; Almeida, M.T.; Pinheiro, J.C.; Sousa, S.; Cabanes, D. Listeria monocytogenes CadC regulates cadmium efflux and fine-tunes lipoprotein localization to escape the host immune response and promote infection. J. Infect. Dis. 2017, 215, 1468–1479. [Google Scholar] [CrossRef]
- Schmitz-Esser, S.; Gram, L.; Wagner, M. Complete genome sequence of the persistent Listeria monocytogenes strain R479a. Genome Announc. 2015, 3, e00150-15. [Google Scholar] [CrossRef]
- Gilmour, M.W.; Graham, M.; Van Domselaar, G.; Tyler, S.; Kent, H.; Trout-Yakel, K.M.; Larios, O.; Allen, V.; Lee, B.; Nadon, C. High-throughput genome sequencing of two Listeria monocytogenes clinical isolates during a large foodborne outbreak. BMC Genom. 2010, 11, 120. [Google Scholar] [CrossRef]
- Canchaya, C.; Giubellini, V.; Ventura, M.; de los Reyes-Gavilán, C.G.; Margolles, A. Mosaic-like sequences containing transposon, phage, and plasmid elements among Listeria monocytogenes plasmids. Appl. Environ. Microbiol. 2010, 76, 4851–4857. [Google Scholar] [CrossRef]
- Tasara, T.; Klumpp, J.; Bille, J.; Stephan, R. Genome sequences of Listeria monocytogenes strains responsible for cheese- and cooked ham product-associated Swiss listeriosis outbreaks in 2005 and 2011. Genome Announc. 2016, 4, e00106-16. [Google Scholar] [CrossRef] [PubMed]
- Casey, A.; McAuliffe, O.; Fox, E.M.; Leong, D.; Gahan, C.G.M.; Jordan, K. Draft genome sequences of Listeria monocytogenes serotype 4b Strains 944 and 2993 and serotype 1/2c strains 198 and 2932. Genome Announc. 2016, 4, e00482-16. [Google Scholar] [CrossRef] [PubMed]
- Portmann, A.-C.; Fournier, C.; Gimonet, J.; Ngom-Bru, C.; Barretto, C.; Baert, L. A validation approach of an end-to-end whole genome sequencing workflow for source tracking of Listeria monocytogenes and Salmonella enterica. Front. Microbiol. 2018, 9, 446. [Google Scholar] [CrossRef] [PubMed]
- Orsini, M.; Cornacchia, A.; Patavino, C.; Torresi, M.; Centorame, P.; Acciari, V.A.; Ruolo, A.; Marcacci, M.; Ancora, M.; Di Domenico, M.; et al. Whole-genome sequences of two Listeria monocytogenes serovar 1/2a strains responsible for a severe listeriosis outbreak in central Italy. Genome Announc. 2018, 6, e00236-18. [Google Scholar] [CrossRef] [PubMed]
- Torresi, M.; Orsini, M.; Acciari, V.; Centorotola, G.; Di Lollo, V.; Di Domenico, M.; Bianchi, D.M.; Ziba, M.W.; Tramuta, C.; Cammà, C.; et al. Genetic characterization of a Listeria monocytogenes serotype IVb variant 1 strain isolated from vegetal matrix in Italy. Microbiol. Resour. Announc. 2020, 9, e00782-20. [Google Scholar] [CrossRef]
- Kwon, H.J.; Chen, Z.; Evans, P.; Meng, J.; Chen, Y. Characterization of mobile genetic elements using long-read sequencing for tracking Listeria monocytogenes from food processing environments. Pathogens 2020, 9, 822. [Google Scholar] [CrossRef]
- Bailey, T.W.; do Nascimento, N.C.; Bhunia, A.K. Genome sequence of Listeria monocytogenes strain F4244, a 4b serotype. Genome Announc. 2017, 5, e01324-17. [Google Scholar] [CrossRef]
- Peters, T.L.; Hudson, L.K.; Bryan, D.W.; Song, Y.; den Bakker, H.C.; Kucerova, Z.; Denes, T.G. Complete genome sequence of a serotype 7 Listeria monocytogenes strain, FSL R9-0915. Microbiol. Resour. Announc. 2021, 10, e01158-20. [Google Scholar] [CrossRef]
- Gonzalez-Escalona, N.; Kastanis, G.J.; Timme, R.; Roberson, D.; Balkey, M.; Tallent, S.M. Closed genome sequences of 28 foodborne pathogens from the CFSAN verification set, determined by a combination of long and short reads. Microbiol. Resour. Announc. 2020, 9, e00152-20. [Google Scholar] [CrossRef]
- Pightling, A.W.; Pagotto, F. Genome sequence of Listeria monocytogenes strain HPB5415, collected during a 2008 listeriosis outbreak in Canada. Genome Announc. 2015, 3, e00637-15. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chmielowska, C.; Korsak, D.; Chapkauskaitse, E.; Decewicz, P.; Lasek, R.; Szuplewska, M.; Bartosik, D. Plasmidome of Listeria spp.—The repA-Family Business. Int. J. Mol. Sci. 2021, 22, 10320. https://doi.org/10.3390/ijms221910320
Chmielowska C, Korsak D, Chapkauskaitse E, Decewicz P, Lasek R, Szuplewska M, Bartosik D. Plasmidome of Listeria spp.—The repA-Family Business. International Journal of Molecular Sciences. 2021; 22(19):10320. https://doi.org/10.3390/ijms221910320
Chicago/Turabian StyleChmielowska, Cora, Dorota Korsak, Elvira Chapkauskaitse, Przemysław Decewicz, Robert Lasek, Magdalena Szuplewska, and Dariusz Bartosik. 2021. "Plasmidome of Listeria spp.—The repA-Family Business" International Journal of Molecular Sciences 22, no. 19: 10320. https://doi.org/10.3390/ijms221910320
APA StyleChmielowska, C., Korsak, D., Chapkauskaitse, E., Decewicz, P., Lasek, R., Szuplewska, M., & Bartosik, D. (2021). Plasmidome of Listeria spp.—The repA-Family Business. International Journal of Molecular Sciences, 22(19), 10320. https://doi.org/10.3390/ijms221910320





