Delivery of Metabolically Neuroactive Probiotics to the Human Gut
Abstract
:1. Introduction
2. Results
2.1. Capsule Design & Delivery of Metabolically Active Probiotic Cells to the Intestine
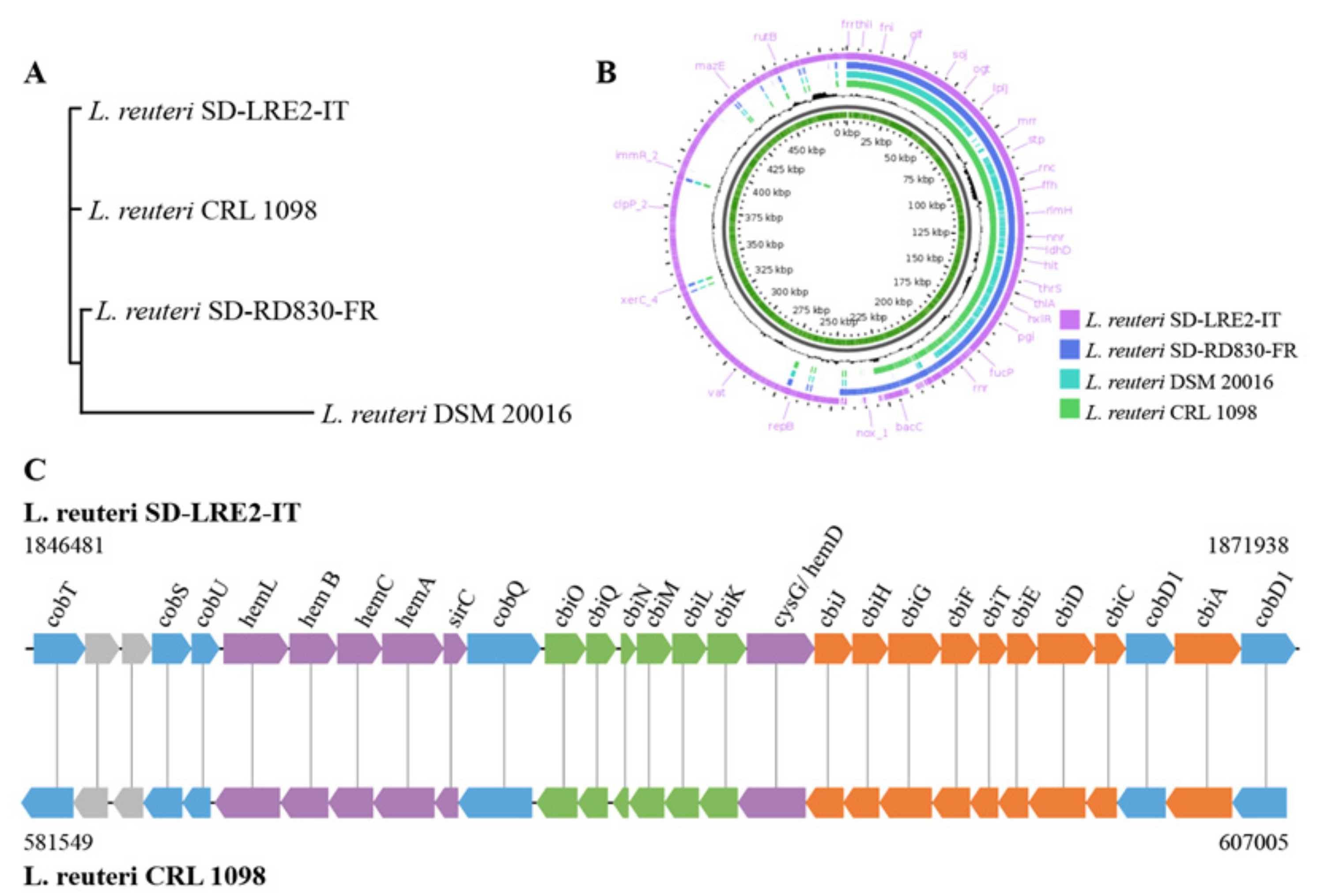
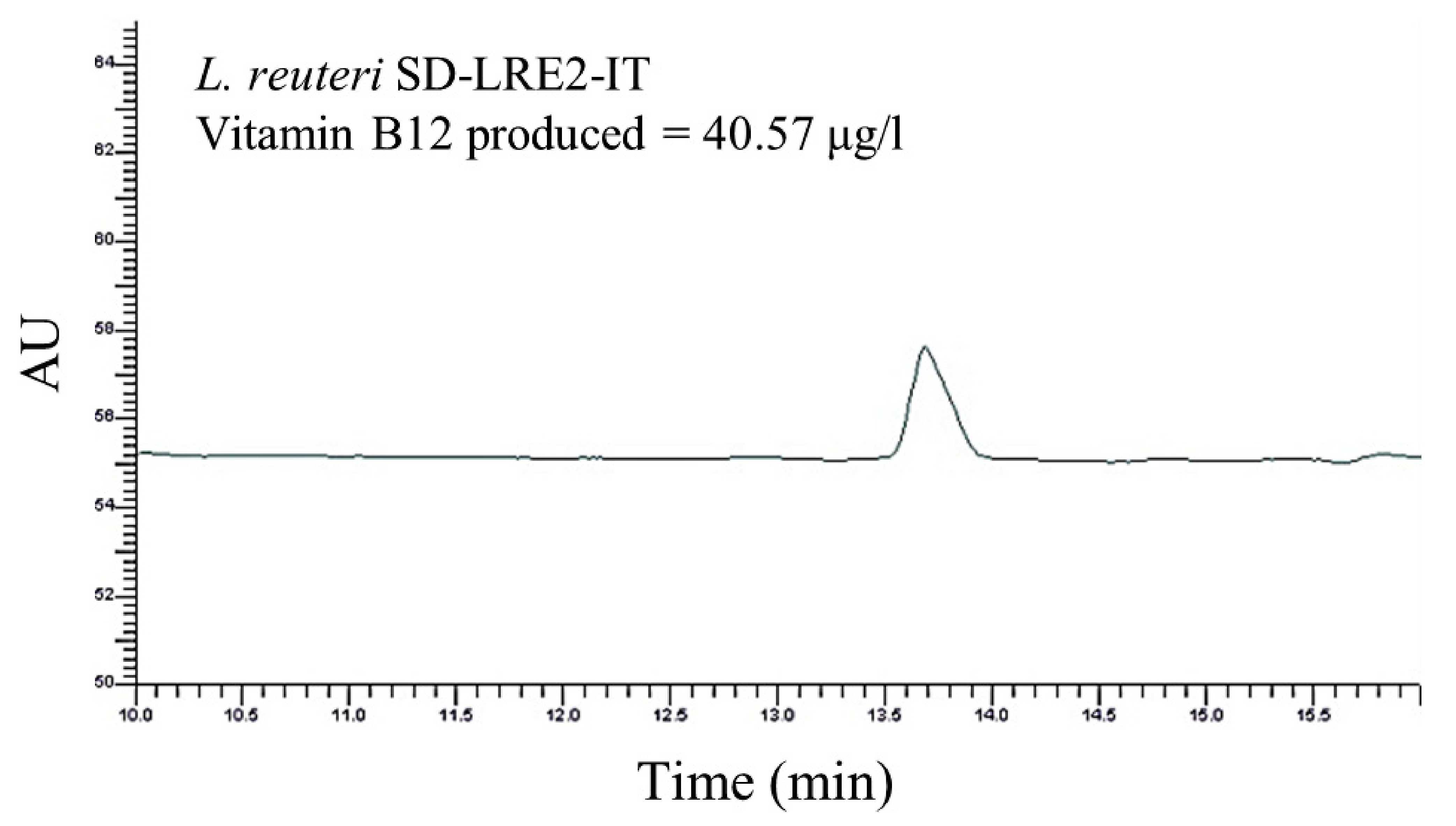
2.2. Genomic & Functional Analysis of Vitamin B12 Production by L. reuteri SD-LRE2-IT
3. Discussion
4. Materials and Methods
4.1. Probiotic Product
4.2. Upper GIT Model and Bacterial Release from the Capsule
4.3. Probiotic Survivability
4.4. Sequencing and Analysis of L. reuteri Genomes for Vitamin B12 Production Capacity
4.5. Functional Assessment of Vitamin B12 Production by L. reuteri
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yeung, A.W.K.; Goto, T.K.; Leung, W.K. The changing landscape of neuroscience research, 2006–2015: A bibliometric study. Front. Neurosci. 2017, 11, 120. [Google Scholar] [CrossRef]
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G.R.; Merenstein, D.J.; Pot, B.; Morelli, L.; Canani, R.B.; Flint, H.J.; Salminen, S.; et al. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506–514. [Google Scholar] [CrossRef] [Green Version]
- Bercik, P.; Park, A.J.; Sinclair, D.; Khoshdel, A.; Lu, J.; Huang, X.; Deng, Y.; Blennerhassett, P.A.; Fahnestock, M.; Moine, D.; et al. The anxiolytic effect of Bifidobacterium longum NCC3001 involves vagal pathways for gut-brain communication. Neurogastroenterol. Motil. 2011, 23, 1132–1139. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rice, M.W.; Pandya, J.D.; Shear, D.A. Gut microbiota as a therapeutic target to ameliorate the biochemical, neuroanatomical, and behavioral effects of traumatic brain injuries. Front. Neurol. 2019, 10, 875. [Google Scholar] [CrossRef] [PubMed]
- Mörkl, S.; Butler, M.I.; Holl, A.; Cryan, J.F.; Dinan, T.G. Probiotics and the microbiota-gut-brain axis: Focus on psychiatry. Curr. Nutr. Rep. 2020, 9, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Erny, D.; De Angelis, A.L.; Jaitin, D.; Wieghofer, P.; Staszewski, O.; David, E.; Keren-Shaul, H.; Mahlakoiv, T.; Jakobshagen, K.; Buch, T.; et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 2015, 18, 965–977. [Google Scholar] [CrossRef] [PubMed]
- Peterson, L.; Artis, D. Intestinal epithelial cells: Regulators of barrier function and immune homeostasis. Nat. Rev. Immunol. 2014, 14, 141–153. [Google Scholar] [CrossRef]
- El Aidy, S.; Bogert, B.V.D.; Kleerebezem, M. The small intestine microbiota, nutritional modulation and relevance for health. Curr. Opin. Biotechnol. 2015, 32, 14–20. [Google Scholar] [CrossRef]
- van Baarlen, P.; Troost, F.; van der Meer, C.; Hooiveld, G.; Boekschoten, M.; Brummer, R.J.M.; Kleerebezem, M. Human mucosal in vivo transcriptome responses to three lactobacilli indicate how probiotics may modulate human cellular pathways. Proc. Natl. Acad. Sci. USA 2011, 108 (Suppl. 1), 4562–4569. [Google Scholar] [CrossRef] [Green Version]
- van Baarlen, P.; Troost, F.J.; van Hemert, S.; van der Meer, C.; de Vos, W.M.; de Groot, P.J.; Hooiveld, G.J.E.J.; Brummer, R.-J.M.; Kleerebezem, M. Differential NF-kappaB pathways induction by Lactobacillus plantarum in the duodenum of healthy humans correlating with immune tolerance. Proc. Natl. Acad. Sci. USA 2009, 106, 2371–2376. [Google Scholar] [CrossRef] [Green Version]
- Strozzi, G.P.; Mogna, L. Quantification of folic acid in human feces after administration of Bifidobacterium probiotic strains. J. Clin. Gastroenterol. 2008, 42 (Suppl. 3), S179–S184. [Google Scholar] [CrossRef]
- Black, M.M. Effects of vitamin B12 and folate deficiency on brain development in children. Food Nutr. Bull. 2008, 29, S126–S131. [Google Scholar] [CrossRef] [Green Version]
- Ma, F.; Zhou, X.; Li, Q.; Zhao, J.; Song, A.; An, P.; Du, Y.; Xu, W.; Huang, G. Effects of folic acid and vitamin B12, alone and in combination on cognitive function and inflammatory factors in the elderly with mild cognitive impairment: A single-blind experimental design. Curr. Alzheimer Res. 2019, 16, 622–632. [Google Scholar] [CrossRef]
- Ma, F.; Wu, T.; Zhao, J.; Ji, L.; Song, A.; Zhang, M.; Huang, G. Plasma homocysteine and serum folate and vitamin B12 levels in mild cognitive impairment and Alzheimer’s disease: A case-control study. Nutrients 2017, 9, 725. [Google Scholar] [CrossRef] [Green Version]
- Hervouet, E.; Debien, E.; Campion, L.; Charbord, J.; Menanteau, J.; Vallette, F.; Cartron, P.-F. Folate supplementation limits the aggressiveness of glioma via the remethylation of DNA repeats element and genes governing apoptosis and proliferation. Clin. Cancer Res. 2009, 15, 3519–3529. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mu, Q.; Tavella, V.J.; Luo, X.M. Role of Lactobacillus reuteri in human health and diseases. Front. Microbiol. 2018, 9, 757. [Google Scholar] [CrossRef]
- Santos, F.; Wegkamp, A.; de Vos, W.M.; Smid, E.J.; Hugenholtz, J. High-level folate production in fermented foods by the B 12 producer Lactobacillus reuteri JCM1112. Appl. Environ. Microbiol. 2008, 74, 3291–3294. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Santos, F.; Vera, J.L.; van der Heijden, R.; Valdez, G.; de Vos, W.M.; Sesma, F.; Hugenholtz, J. The complete coenzyme B12 biosynthesis gene cluster of Lactobacillus reuteri CRL1098. Microbiology 2008, 154, 81–93. [Google Scholar] [CrossRef] [Green Version]
- Veen, H.V.B.-V.D.; Van Swam, I.; Wels, M.; Bron, P.A.; Kleerebezem, M. Congruent strain specific intestinal persistence of Lactobacillus plantarum in an intestine-mimicking in vitro system and in human volunteers. PLoS ONE 2012, 7, e44588. [Google Scholar]
- Piano, M.D.; Carmagnola, S.; Ballarè, M.; Balzarini, M.; Montino, F.; Pagliarulo, M.; Anderloni, A.; Orsello, M.; Tari, R.; Sforza, F.; et al. Comparison of the kinetics of intestinal colonization by associating 5 probiotic bacteria assumed either in a microencapsulated or in a traditional, uncoated form. J. Clin. Gastroenterol. 2012, 46, S85–S92. [Google Scholar] [CrossRef] [PubMed]
- Del Piano, M.; Carmagnola, S.; Andorno, S.; Pagliarulo, M.; Tari, R.; Mogna, L.; Strozzi, G.P.; Sforza, F.; Capurso, L. Evaluation of the intestinal colonization by microencapsulated probiotic bacteria in comparison with the same uncoated strains. J. Clin. Gastroenterol. 2010, 44 (Suppl. 1), S42–S46. [Google Scholar] [CrossRef]
- Ajlouni, S.; Ranadheera, C.S.; Chua, E.L. Encapsulation increases the in vitro bioaccessibility of probiotics in yoghurt. Int. J. Dairy Technol 2021, 74, 118–127. [Google Scholar] [CrossRef]
- Molly, K.; Woestyne, M.V.; Verstraete, W. Development of a 5-step multi-chamber reactor as a simulation of the human intestinal microbial ecosystem. Appl. Microbiol. Biotechnol. 1993, 39, 254–258. [Google Scholar] [CrossRef] [PubMed]
- Marzorati, M.; Possemiers, S.; Verhelst, A.; Cadé, D.; Madit, N.; Van de Wiele, T. A novel hypromellose capsule, with acid resistance properties, permits the targeted delivery of acid-sensitive products to the intestine. LWT Food Sci. Technol. 2015, 60, 544–551. [Google Scholar] [CrossRef]
- Miyamoto-Shinohara, Y.; Sukenobe, J.; Imaizumi, T.; Nakahara, T. Survival of freeze-dried bacteria. J. Gen. Appl. Microbiol. 2008, 54, 9–24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, M.; Kissoon-Singh, V.; Coria, A.L.; Moreau, F.; Chadee, K. Probiotic mixture VSL#3 reduces colonic inflammation and improves intestinal barrier function in Muc2 mucin-deficient mice. Am. J. Physiol. Gastrointest. Liver Physiol. 2017, 312, G34–G45. [Google Scholar]
- Del Piano, M.; Carmagnola, S.; Anderloni, A.; Andorno, S.; Ballarè, M.; Balzarini, M.; Montino, F.; Orsello, M.; Pagliarulo, M.; Sartori, M.; et al. The use of probiotics in healthy volunteers with evacuation disorders and hard stools: A double-blind, randomized, placebo-controlled study. J. Clin. Gastroenterol. 2010, 44 (Suppl. 1), S30–S34. [Google Scholar] [CrossRef]
- Ogata, T.; Nakamura, T.; Anjitsu, K.; Yaeshima, T.; Takahashi, S.; Fukuwatari, Y.; Ishibashi, N.; Hayasawa, H.; Fujisawa, T.; Iino, H. Effect of Bifidobacterium longum BB536 administration on the intestinal environment, defecation frequency and fecal characteristics of human volunteers. Biosci. Microflora 1997, 16, 53–58. [Google Scholar] [CrossRef] [Green Version]
- Ma, Z.F.; Yusof, N.; Hamid, N.; Lawenko, R.; Mohammad, W.W.; Liong, M.; Sugahara, H.; Odamaki, T.; Xiao, J.; Lee, Y.Y. Bifidobacterium infantis M-63 improves mental health in victims with irritable bowel syndrome developed after a major flood disaster. Benef. Microbes 2019, 10, 111–120. [Google Scholar] [CrossRef]
- Costabile, A.; Buttarazzi, I.; Kolida, S.; Quercia, S.; Baldini, J.; Swann, J.; Brigidi, P.; Gibson, G.R. An in vivo assessment of the cholesterol-lowering efficacy of Lactobacillus plantarum ECGC 13110402 in normal to mildly hypercholesterolaemic adults. PLoS ONE 2017, 12, e0187964. [Google Scholar] [CrossRef]
- Salminen, S.; Collado, M.C.; Endo, A.; Hill, C.; Lebeer, S.; Quigley, E.M.M.; Sanders, M.E.; Shamir, R.; Swann, J.R.; Szajewska, H.; et al. The International Scientific Association of Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of postbiotics. Nat. Rev. Gastroenterol. Hepatol. 2021, 4, 1–19. [Google Scholar] [CrossRef]
- Ford, T.C.; Downey, L.A.; Simpson, T.; McPhee, G.; Oliver, C.; Stough, C. The effect of a high-dose vitamin B multivitamin supplement on the relationship between brain metabolism and blood biomarkers of oxidative stress: A randomized control trial. Nutrients 2018, 10, 1860. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Valles-Colomer, M.; Falony, G.; Darzi, Y.; Tigchelaar, E.F.; Wang, J.; Tito, R.Y.; Schiweck, C.; Kurilshikov, A.; Joossens, M.; Wijmenga, C.; et al. The neuroactive potential of the human gut microbiota in quality of life and depression. Nat. Microbiol. 2019, 4, 623–632. [Google Scholar] [CrossRef] [PubMed]
- Yano, J.M.; Yu, K.; Donaldson, G.P.; Shastri, G.G.; Ann, P.; Ma, L.; Nagler, C.R.; Ismagilov, R.F.; Mazmanian, S.K.; Hsiao, E.Y. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell 2015, 161, 264–276. [Google Scholar] [CrossRef] [Green Version]
- Blacher, E.; Bashiardes, S.; Shapiro, H.; Rothschild, D.; Mor, U.; Dori-Bachash, M.; Kleimeyer, C.; Moresi, C.; Harnik, Y.; Zur, M.; et al. Potential roles of gut microbiome and metabolites in modulating ALS in mice. Nature 2019, 572, 474–480. [Google Scholar] [CrossRef] [PubMed]
- Segers, M.E.; Lebeer, S. Towards a better understanding of Lactobacillus rhamnosus GG—host interactions. Microb. Cell Factories 2014, 13, S7. [Google Scholar] [CrossRef] [Green Version]
- Bron, P.A.; Kleerebezem, M.; Brummer, R.-J.; Cani, P.D.; Mercenier, A.; Macdonald, T.T.; Garcia-Ródenas, C.L.; Wells, J.M. Can probiotics modulate human disease by impacting intestinal barrier function? Br. J. Nutr. 2017, 117, 93–107. [Google Scholar] [CrossRef] [PubMed]
- Hu, P.-L.; Yuan, Y.-H.; Yue, T.-L.; Guo, C.-F. Bile acid patterns in commercially available oxgall powders used for the evaluation of the bile tolerance ability of potential probiotics. PLoS ONE 2018, 13, e0192964. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef]
- Hyatt, D.; Chen, G.-L.; Locascio, P.F.; Land, M.L.; Larimer, F.W.; Hauser, L.J. Prodigal: Prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010, 11, 119. [Google Scholar] [CrossRef] [Green Version]
- Page, A.J.; Cummins, C.A.; Hunt, M.; Wong, V.K.; Reuter, S.; Holden, M.T.G.; Fookes, M.; Falush, D.; Keane, J.A.; Parkhill, J. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 2015, 31, 3691–3693. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
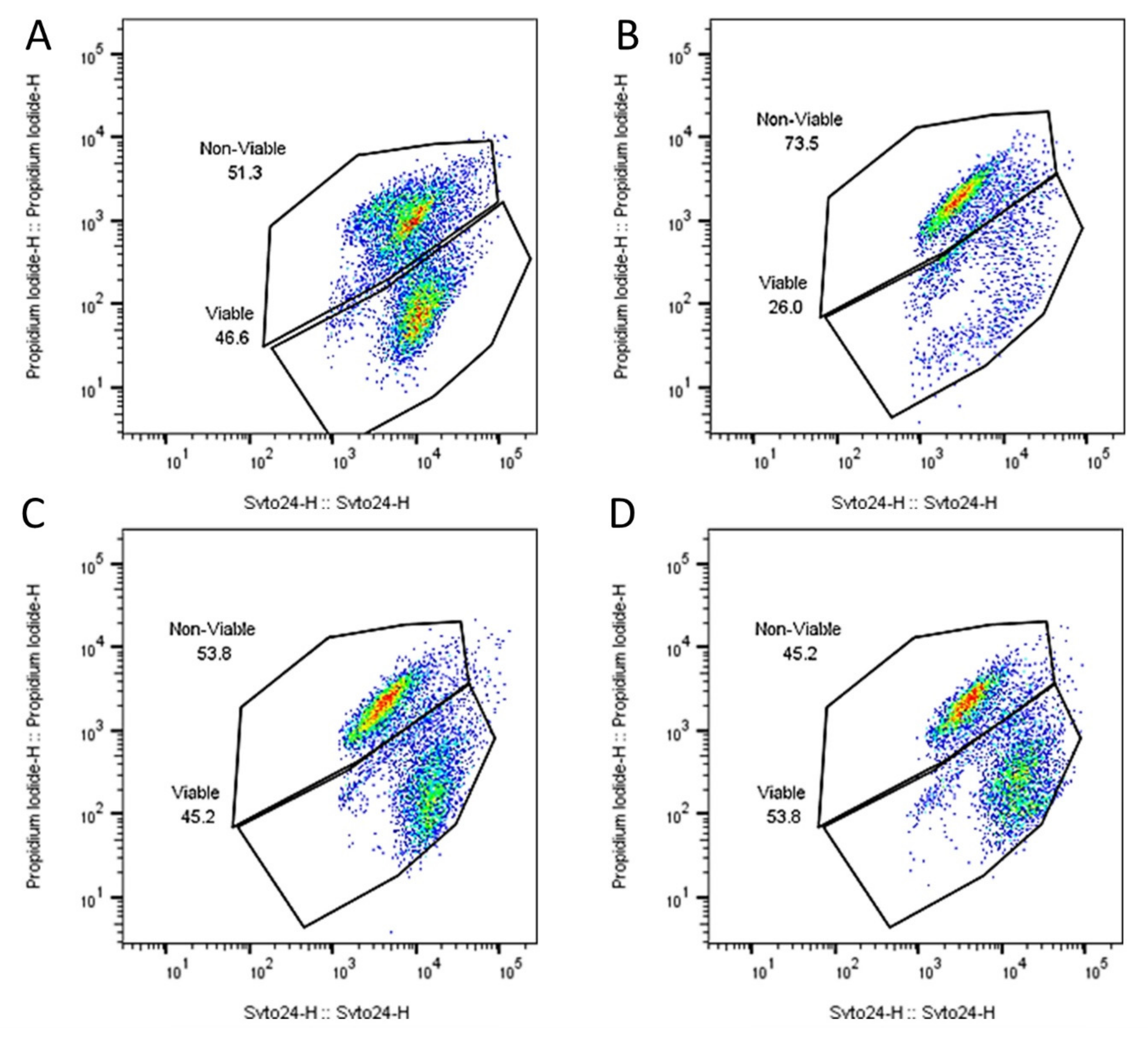
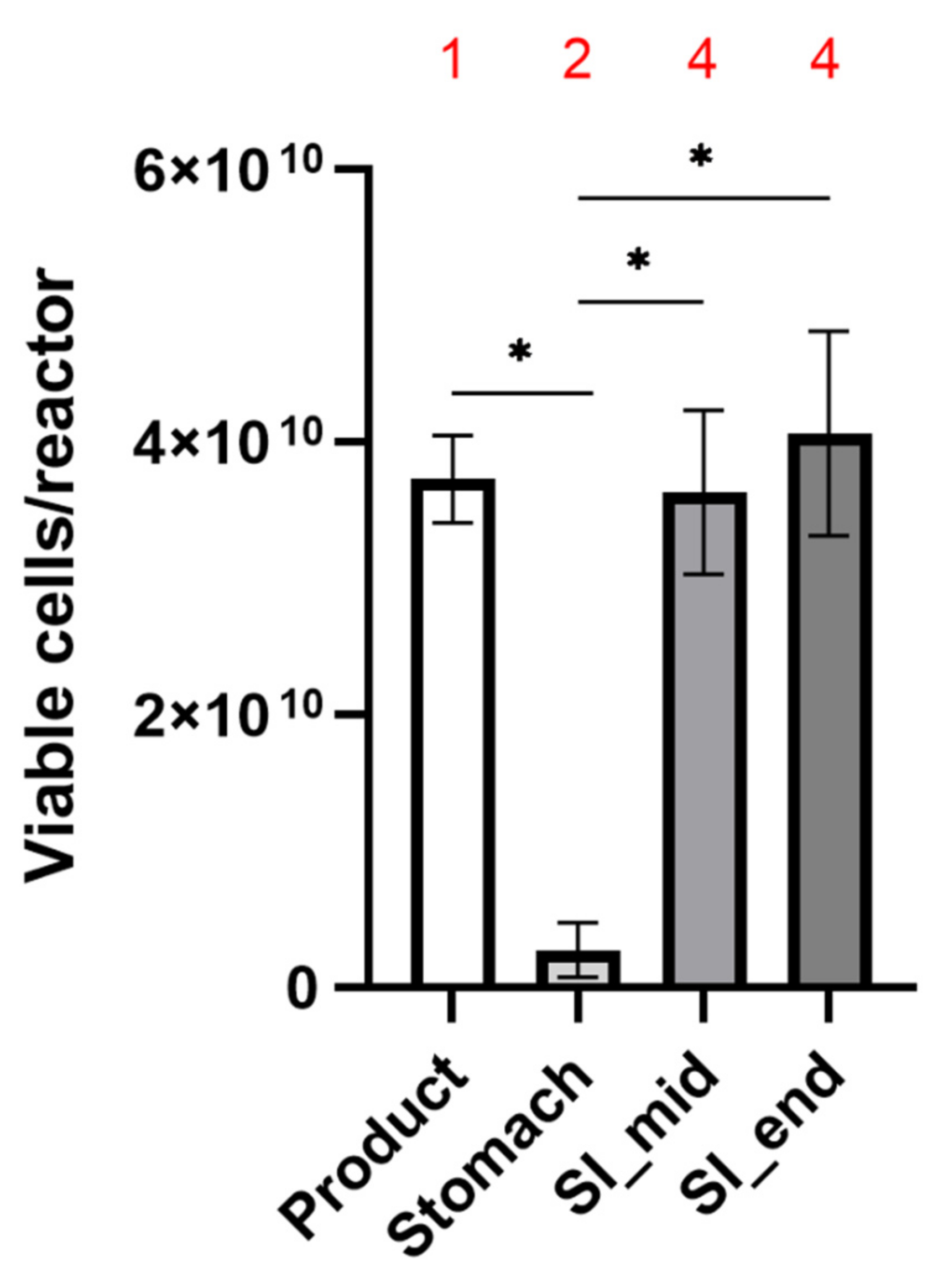
| Counts | Percentage of Cells Viable | |||||
|---|---|---|---|---|---|---|
| Viable | ||||||
| Product | 3.73 × 1010 | ± | 3.21 × 109 | 46.03 | ± | 0.54 |
| ST end | 2.77 × 109 | ± | 1.98 × 109 | 25.99 | ± | 4.03 |
| SI mid | 3.63 × 1010 | ± | 9.07 × 109 | 45.38 | ± | 8.56 |
| SI end | 4.07 × 1010 | ± | 7.51 × 109 | 46.15 | ± | 7.87 |
| Dead | ||||||
| Product | 3.57 × 109 | ± | 1.78 × 109 | 53.97 | ± | 0.54 |
| ST end | 8.80 × 109 | ± | 7.30 × 109 | 73.36 | ± | 4.54 |
| SI mid | 4.30 × 1010 | ± | 5.03 × 109 | 53.57 | ± | 8.36 |
| SI end | 4.67 × 1010 | ± | 6.11 × 109 | 53.13 | ± | 7.79 |
| Total | ||||||
| Product | 8.12 × 1010 | ± | 7.99 × 109 | 100 | ± | 0.5 |
| ST end | 1.16 × 1010 | ± | 9.29 × 109 | 99.35 | ± | 0.75 |
| SI mid | 8.02 × 1010 | ± | 4.36 × 109 | 98.95 | ± | 1.15 |
| SI end | 8.80 × 1010 | ± | 2.28 × 109 | 99.28 | ± | 0.35 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bron, P.A.; Catalayud, M.; Marzorati, M.; Pane, M.; Kartal, E.; Dhir, R.; Reid, G. Delivery of Metabolically Neuroactive Probiotics to the Human Gut. Int. J. Mol. Sci. 2021, 22, 9122. https://doi.org/10.3390/ijms22179122
Bron PA, Catalayud M, Marzorati M, Pane M, Kartal E, Dhir R, Reid G. Delivery of Metabolically Neuroactive Probiotics to the Human Gut. International Journal of Molecular Sciences. 2021; 22(17):9122. https://doi.org/10.3390/ijms22179122
Chicago/Turabian StyleBron, Peter A., Marta Catalayud, Massimo Marzorati, Marco Pane, Ece Kartal, Raja Dhir, and Gregor Reid. 2021. "Delivery of Metabolically Neuroactive Probiotics to the Human Gut" International Journal of Molecular Sciences 22, no. 17: 9122. https://doi.org/10.3390/ijms22179122
APA StyleBron, P. A., Catalayud, M., Marzorati, M., Pane, M., Kartal, E., Dhir, R., & Reid, G. (2021). Delivery of Metabolically Neuroactive Probiotics to the Human Gut. International Journal of Molecular Sciences, 22(17), 9122. https://doi.org/10.3390/ijms22179122






