Identification of Rhythmically Expressed LncRNAs in the Zebrafish Pineal Gland and Testis
Abstract
:1. Introduction
2. Results
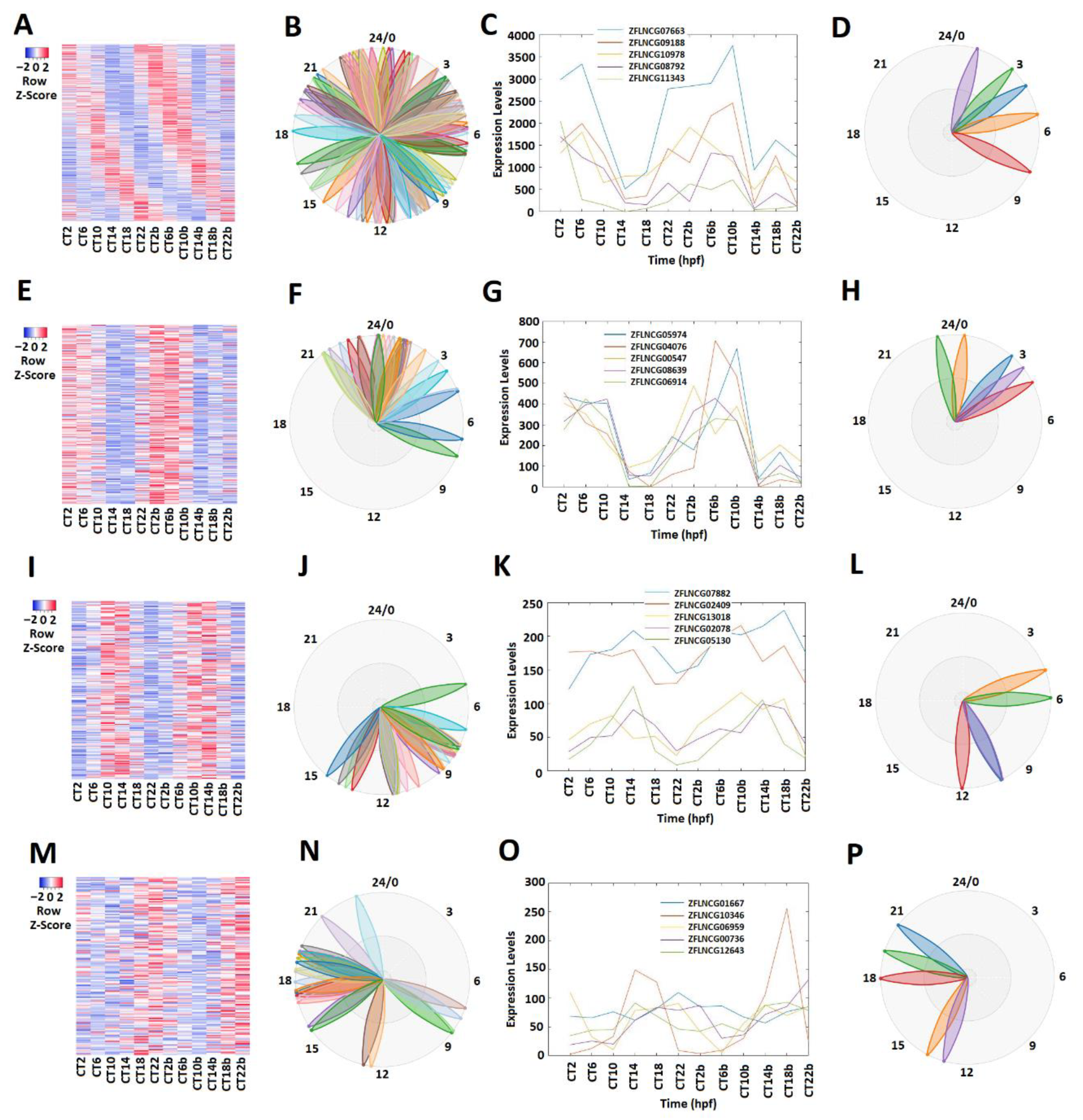
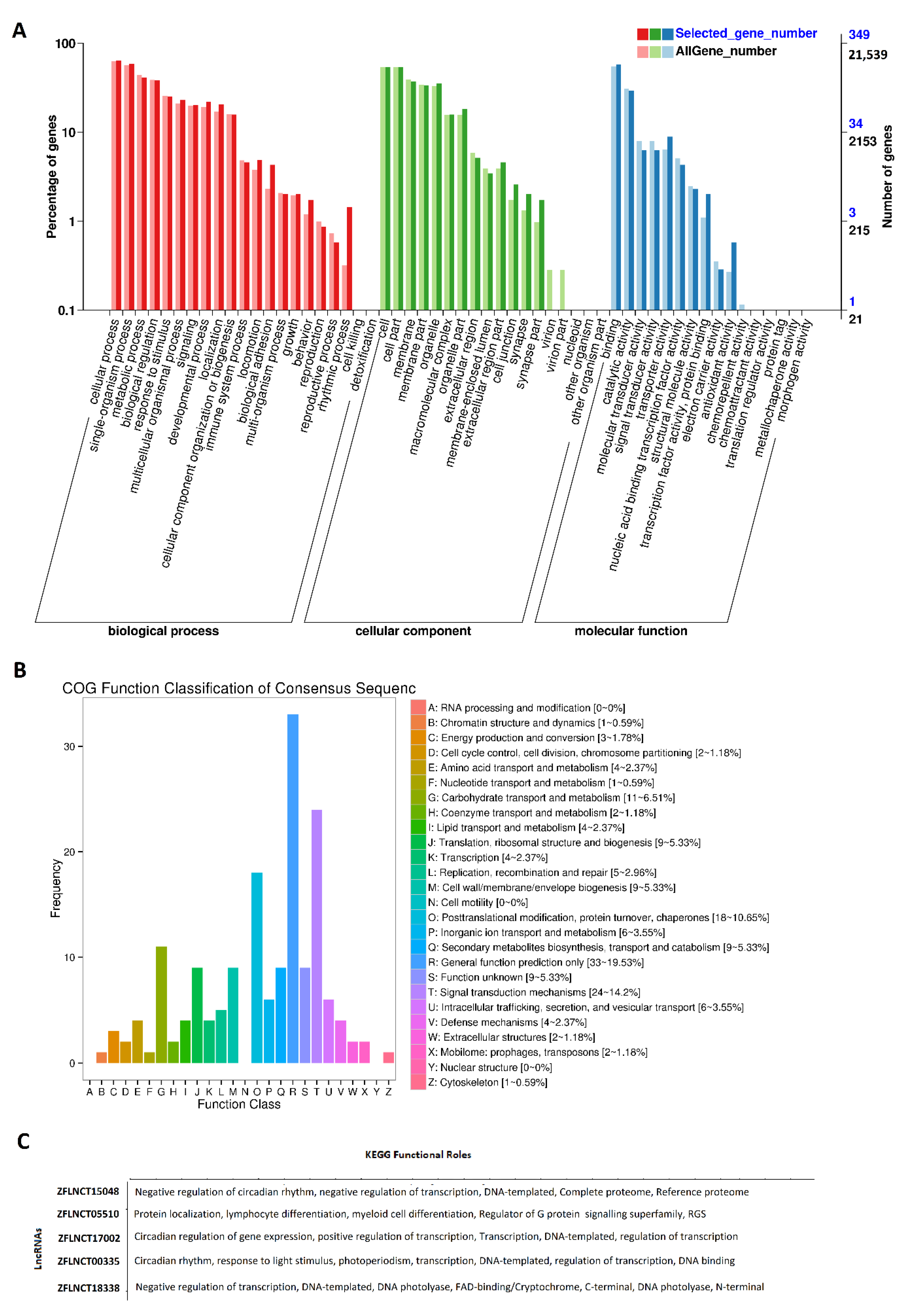
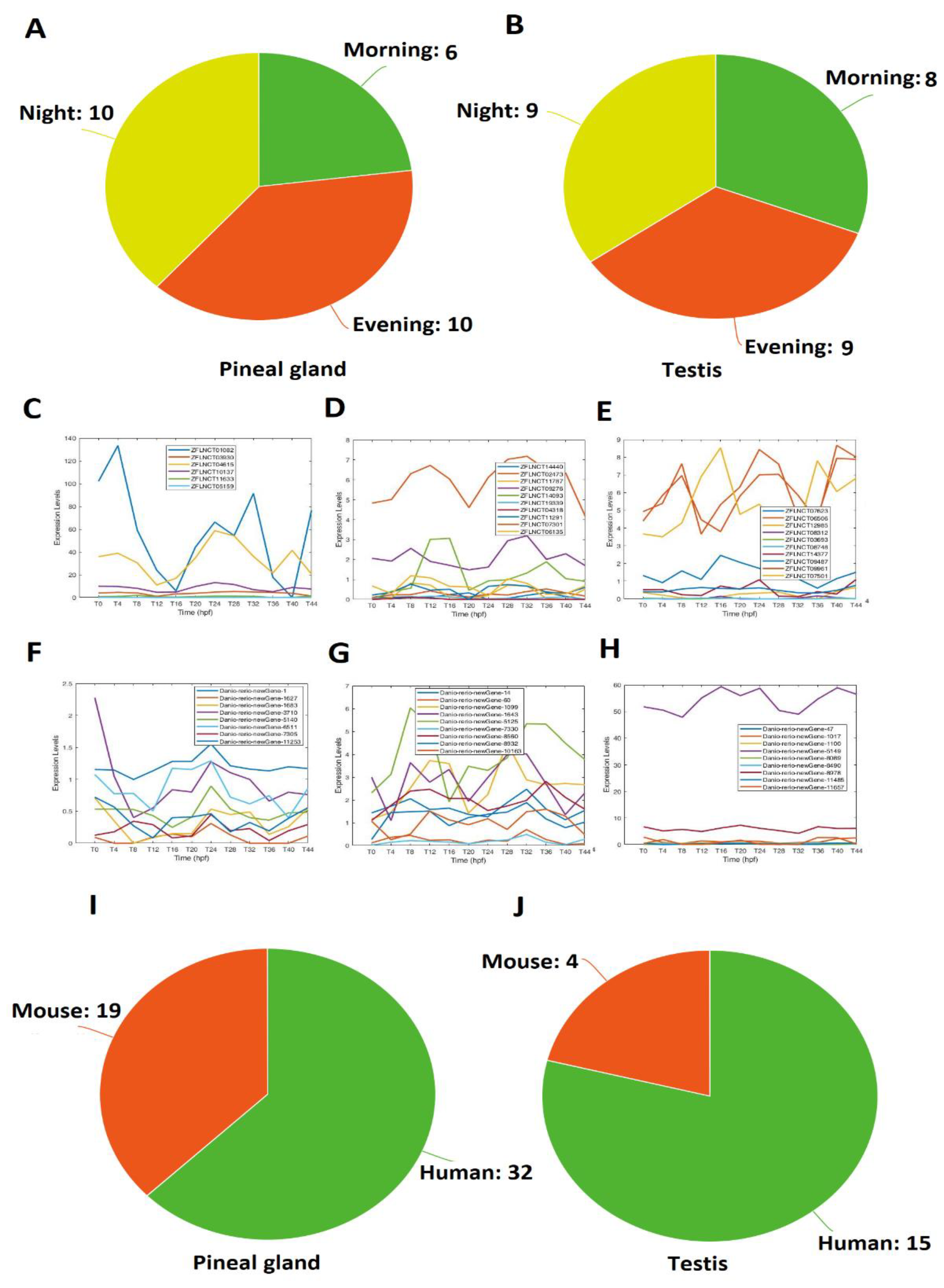
2.1. Rhythmically Expressed LncRNAs in the Zebrafish Pineal Gland, and Their GO, COG, and KEGG Analyses
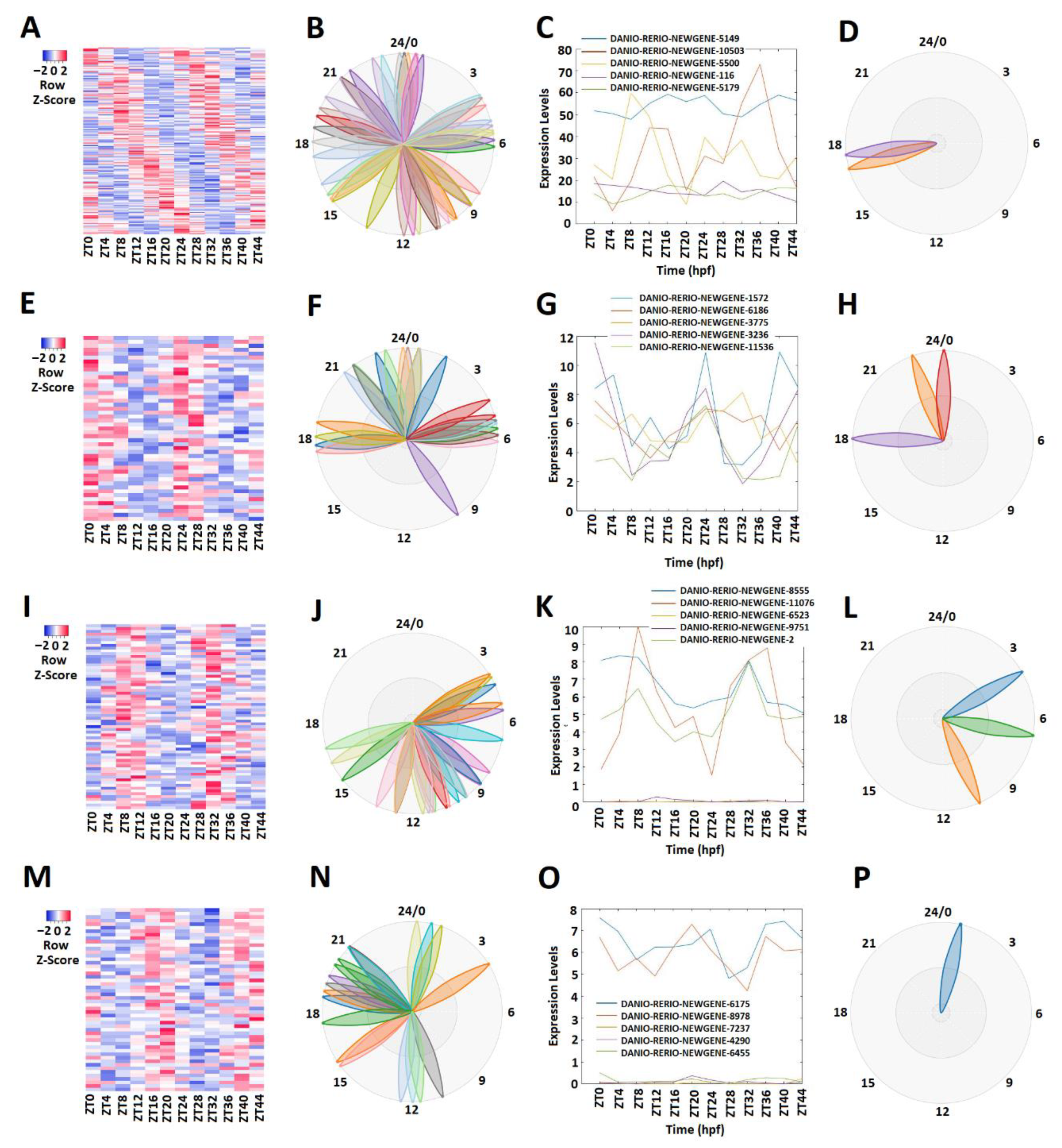
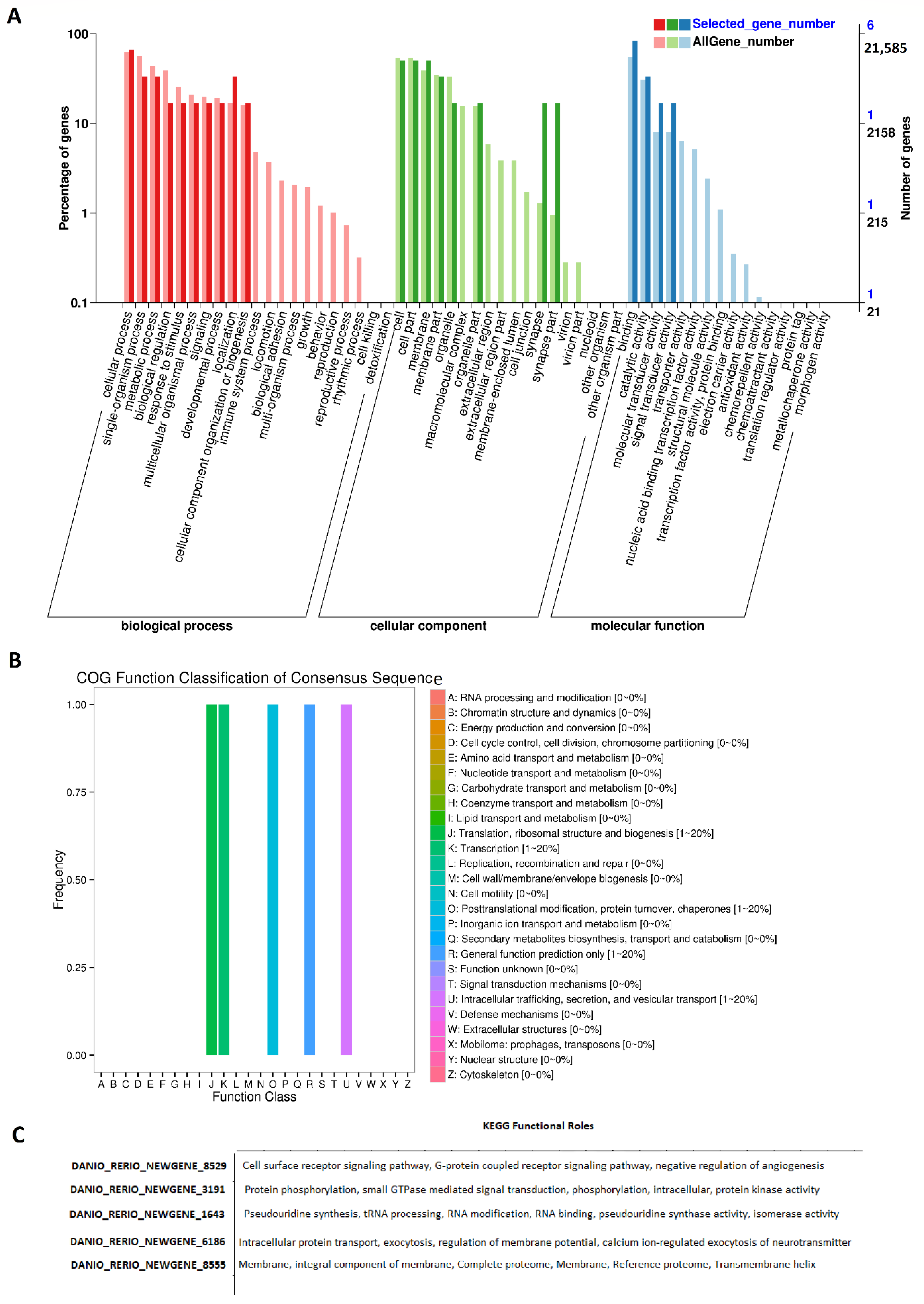
2.2. Rhythmically Expressed lncRNAs in the Zebrafish Testis, and Their GO, COG, and KEGG Analyses
2.3. Rhythmically Expressed LncRNAs Shared between the Pineal Gland and the Testis
2.4. Conservation of Zebrafish Rhythmically Expressed LncRNAs with Mice and Humans
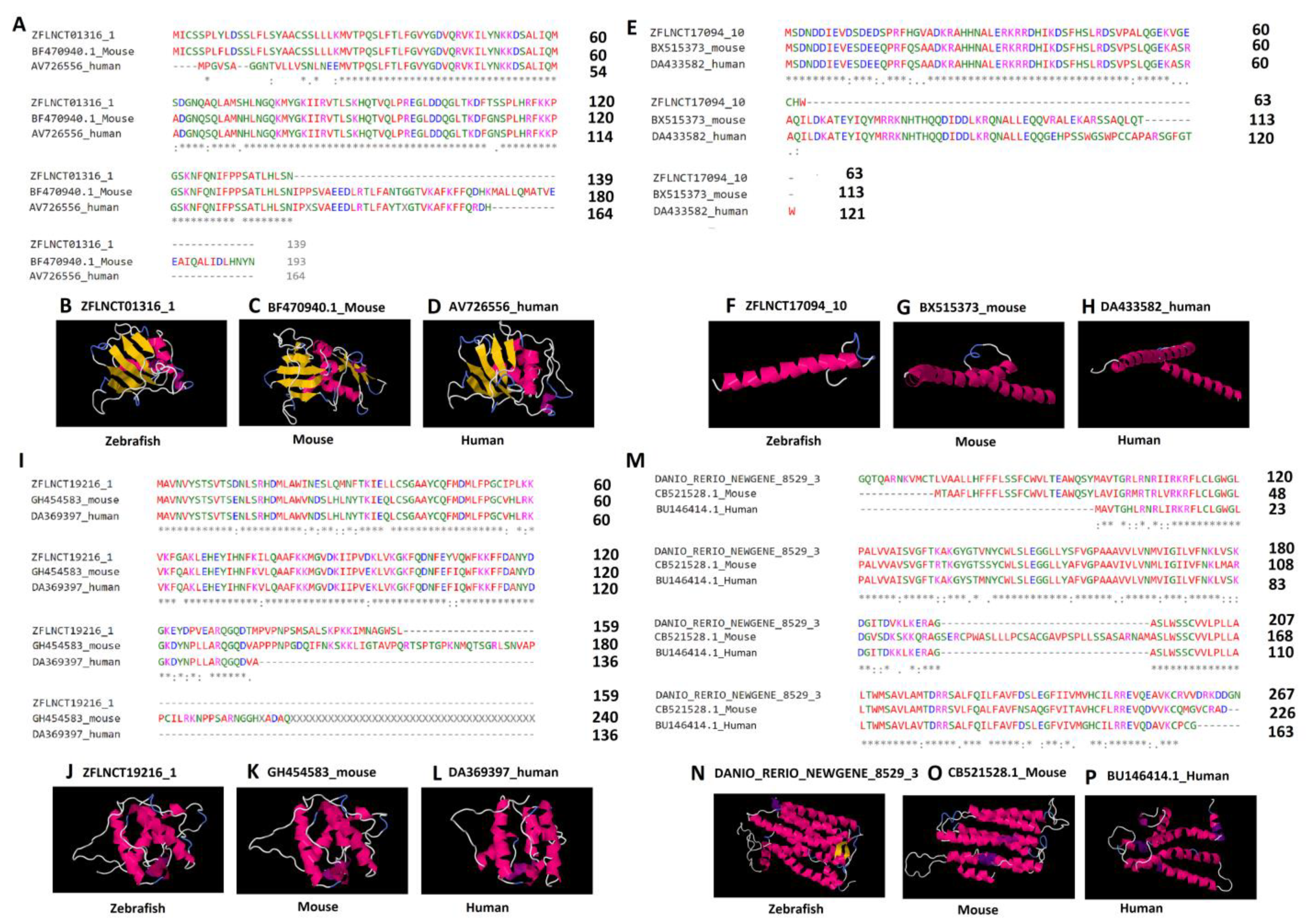
2.5. Conserved LncRNA-Encoded Peptides and Their Predicted 3D Structures
3. Discussion
4. Materials and Methods
4.1. Zebrafish Rhythmically Expressed RNA-Seq Datasets for the Pineal Gland and Testis
4.2. Collection of Zebrafish Pineal Gland LncRNAs
4.3. Identification of Zebrafish Testis LncRNAs
4.4. Rhythmicity Analysis
4.5. Searching for E-Box, D-Box, and RORE Regulatory Motifs in the Promoter Regions of the Rhythmically Expressed LncRNAs
4.6. GO Analysis, COG Analysis, and KEGG Pathway Analysis of Rhythmically Expressed LncRNAs
4.7. Principal Component Analysis (PCA)
4.8. Conservation Analysis with NCBI BLAST
4.9. Prediction of lncRNA-Encoded Peptides’ 3D Structure and Functions
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- The FANTOM Consortium; Carninci, P.; Kasukawa, T.; Katayama, S.; Gough, J.; Frith, M.C.; Maeda, N.; Oyama, R.; Ravasi, T.; Lenhard, B.; et al. The Transcriptional Landscape of the Mammalian Genome. Science 2005, 309, 1559–1563. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ørom, U.A.; Derrien, T.; Beringer, M.; Gumireddy, K.; Gardini, A.; Bussotti, G.; Lai, F.; Zytnicki, M.; Notredame, C.; Huang, Q.; et al. Long Noncoding RNAs with Enhancer-like Function in Human Cells. Cell 2010, 143, 46–58. [Google Scholar] [CrossRef] [Green Version]
- Guttman, M.; Amit, I.; Garber, M.; French, C.; Lin, M.F.; Feldser, D.; Huarte, M.; Zuk, O.; Carey, B.W.; Cassady, J.P.; et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nat. Cell Biol. 2009, 458, 223–227. [Google Scholar] [CrossRef] [PubMed]
- Yunusov, D.; Anderson, L.; DaSilva, L.F.; Wysocka, J.; Ezashi, T.; Roberts, R.M.; Verjovski-Almeida, S. HIPSTR and thousands of lncRNAs are heterogeneously expressed in human embryos, primordial germ cells and stable cell lines. Sci. Rep. 2016, 6, 32753. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, Y.; Liu, N.; Wang, J.P.; Wang, Y.Q.; Yu, X.L.; Bin Wang, Z.; Cheng, X.C.; Zou, Q. Regulatory long non-coding RNA and its functions. J. Physiol. Biochem. 2012, 68, 611–618. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Xu, J.; Liu, L.; Bai, J.; Wang, L.; Xiao, Y.; Li, X.; Zhang, L. Computational identification of epigenetically regulated lncRNAs and their associated genes based on integrating genomic data. FEBS Lett. 2015, 589, 521–531. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.; Wang, Y.; Wang, C.; Hu, J.-F.; Li, W. LncRNA Functions as a New Emerging Epigenetic Factor in Determining the Fate of Stem Cells. Front. Genet. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Anderson, D.M.; Anderson, K.M.; Chang, C.-L.; Makarewich, C.A.; Nelson, B.R.; McAnally, J.R.; Kasaragod, P.; Shelton, J.M.; Liou, J.; Bassel-Duby, R.; et al. A Micropeptide Encoded by a Putative Long Noncoding RNA Regulates Muscle Performance. Cell 2015, 160, 595–606. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, T.; Xu, J.; Liu, L.; Bai, J.; Xu, C.; Xiao, Y.; Li, X.; Zhang, L. Identification of cancer-related lncRNAs through integrating genome, regulome and transcriptome features. Mol. BioSyst. 2014, 11, 126–136. [Google Scholar] [CrossRef] [PubMed]
- Kluiver, J.; van den Berg, A.; de Jong, D.; Blokzijl, T.; Harms, G.; Bouwman, E.; Zhou, H.; Wang, H.; Wang, Z.; Bai, J.; et al. A micropeptide encoded by lncRNA MIR155HG suppresses autoimmune inflammation via modulating antigen presentation. Sci. Adv. 2020, 26, 3769–3776. [Google Scholar]
- Lin, Y.-F.; Xiao, M.-H.; Chen, H.-X.; Meng, Y.; Zhao, N.; Yang, L.; Tang, H.; Wang, J.-L.; Liu, X.; Zhu, Y.; et al. A novel mitochondrial micropeptide MPM enhances mitochondrial respiratory activity and promotes myogenic differentiation. Cell Death Dis. 2019, 10, 528. [Google Scholar] [CrossRef]
- Sousa, M.E.; Farkas, M.H. Micropeptide. PLoS Genet. 2018, 14, e1007764. [Google Scholar] [CrossRef] [PubMed]
- Boon, R.A.; Jaé, N.; Holdt, L.; Dimmeler, S. Long Noncoding RNAs from Clinical Genetics to Therapeutic Targets? J. Am. Coll. Cardiol. 2016, 67, 1214–1226. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wu, S.; Zhu, X.; Zhang, L.; Deng, J.; Li, F.; Guo, B.; Zhang, S.; Wu, R.; Zhang, Z.; et al. LncRNA-encoded polypeptide ASRPS inhibits triple-negative breast cancer angiogenesis. J. Exp. Med. 2020, 217. [Google Scholar] [CrossRef] [PubMed]
- Anfossi, S.; Calin, G.A. When non-coding is not enough. J. Exp. Med. 2020, 217, 217. [Google Scholar] [CrossRef] [PubMed]
- Ye, M.; Zhang, J.; Wei, M.; Liu, B.; Dong, K. Emerging role of long noncoding RNA-encoded micropeptides in cancer. Cancer Cell Int. 2020, 20, 506. [Google Scholar] [CrossRef]
- Cui, M.; Zheng, M.; Sun, B.; Wang, Y.; Ye, L.; Zhang, X. A Long Noncoding RNA Perturbs the Circadian Rhythm of Hepatoma Cells to Facilitate Hepatocarcinogenesis. Neoplasia 2015, 17, 79–88. [Google Scholar] [CrossRef] [Green Version]
- Fan, Z.; Zhao, M.; Joshi, P.D.; Li, P.; Zhang, Y.; Guo, W.; Xu, Y.; Wang, H.; Zhao, Z.; Yan, J. A class of circadian long non-coding RNAs mark enhancers modulating long-range circadian gene regulation. Nucleic Acids Res. 2017, 45, 5720–5738. [Google Scholar] [CrossRef] [Green Version]
- Coon, S.L.; Munson, P.J.; Cherukuri, P.F.; Sugden, D.; Rath, M.; Møller, M.; Clokie, S.; Fu, C.; Olanich, M.E.; Rangel, Z.; et al. Circadian changes in long noncoding RNAs in the pineal gland. Proc. Natl. Acad. Sci. USA 2012, 109, 13319–13324. [Google Scholar] [CrossRef] [Green Version]
- Klein, D.C.; Weller, J. Input and output signals in a model neural system: The regulation of melatonin production in the pineal gland. In Vitro 1970, 6, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Bittman, E.L. Timing in the Testis. J. Biol. Rhythm. 2015, 31, 12–36. [Google Scholar] [CrossRef] [Green Version]
- Morse, D.; Cermakian, N.; Brancorsini, S.; Parvinen, M.; Sassone-Corsi, P. No Circadian Rhythms in Testis: Period1 Expression Is Clock Independent and Developmentally Regulated in the Mouse. Mol. Endocrinol. 2003, 17, 141–151. [Google Scholar] [CrossRef] [Green Version]
- Alvarez, J.D.; Sehgal, A. The Thymus Is Similar to the Testis in Its Pattern of Circadian Clock Gene Expression. J. Biol. Rhythm. 2005, 20, 111–121. [Google Scholar] [CrossRef]
- Vatine, G.; Vallone, D.; Gothilf, Y.; Foulkes, N.S. It’s time to swim! Zebrafish and the circadian clock. FEBS Lett. 2011, 585, 1485–1494. [Google Scholar] [CrossRef] [PubMed]
- Tataroglu, O.; Emery, P. Studying circadian rhythms in Drosophila melanogaster. Methods 2014, 68, 140–150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ben-Moshe, Z.; Foulkes, N.S.; Gothilf, Y. Functional Development of the Circadian Clock in the Zebrafish Pineal Gland. BioMed Res. Int. 2014, 2014, 506. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choi, D. Potency of Melatonin in Living Beings. Dev. Reprod. 2013, 17, 149–177. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tamai, T.K.; Young, L.C.; Whitmore, D. Light signaling to the zebrafish circadian clock by Cryptochrome 1a. Proc. Natl. Acad. Sci. USA 2007, 104, 14712–14717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duan, F.A.; Pauley, M.; Spindel, E.R.; Zhang, L.; Norgren, R.B. Large scale analysis of positional effects of single-base mismatches on microarray gene expression data. BioData Min. 2010, 3, 2. [Google Scholar] [CrossRef] [Green Version]
- Cabili, M.N.; Trapnell, C.; Goff, L.; Koziol, M.; Tazon-Vega, B.; Regev, A.; Rinn, J.L. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 2011, 25, 1915–1927. [Google Scholar] [CrossRef] [Green Version]
- Tovin, A.; Alon, S.; Ben-Moshe, Z.; Mracek, P.; Vatine, G.; Foulkes, N.S.; Jacob-Hirsch, J.; Rechavi, G.; Toyama, R.; Coon, S.L.; et al. Systematic Identification of Rhythmic Genes Reveals camk1gb as a New Element in the Circadian Clockwork. PLoS Genet. 2012, 8, e1003116. [Google Scholar] [CrossRef]
- Barik, S. Molecular Interactions between Pathogens and the Circadian Clock. Int. J. Mol. Sci. 2019, 20, 5824. [Google Scholar] [CrossRef] [Green Version]
- Thiriet, M. Control of Cell Fate in the Circulatory and Ventilatory Systems. In Control of Cell Fate in the Circulatory and Ventilatory Systems; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2012; Volume 2. [Google Scholar] [CrossRef] [Green Version]
- Doherty, C.J.; Kay, S.A. Circadian Control of Global Gene Expression Patterns. Annu. Rev. Genet. 2010, 44, 419–444. [Google Scholar] [CrossRef] [Green Version]
- Grant, C.E.; Bailey, T.L.; Noble, W.S. FIMO: Scanning for occurrences of a given motif. Bioinformatics 2011, 27, 1017–1018. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Galperin, M.; Kristensen, D.M.; Makarova, K.S.; Wolf, Y.; Koonin, E.V. Microbial genome analysis: The COG approach. Brief. Bioinform. 2019, 20, 1063–1070. [Google Scholar] [CrossRef] [PubMed]
- Fang, G.; Bhardwaj, N.; Robilotto, R.; Gerstein, M.B. Getting Started in Gene Orthology and Functional Analysis. PLoS Comput. Biol. 2010, 6, e1000703. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, H.; Hao, Y.; Dong, X.; Gong, Q.; Chen, J.; Zhang, J.; Tian, W. Molecular Mechanisms and Function Prediction of Long Noncoding RNA. Sci. World J. 2012, 2012, 541786. [Google Scholar] [CrossRef]
- Yan, C.; Zhang, Z.; Bao, S.; Hou, P.; Zhou, M.; Xu, C.; Sun, J. Computational Methods and Applications for Identifying Disease-Associated lncRNAs as Potential Biomarkers and Therapeutic Targets. Mol. Ther. Nucleic Acids 2020, 21, 156–171. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Fan, J.; Han, L.; Qi, H.; Wang, Y.; Wang, H.; Chen, S.; Du, L.; Li, S.; Zhang, Y.; et al. The micropeptide LEMP plays an evolutionarily conserved role in myogenesis. Cell Death Dis. 2020, 11, 357. [Google Scholar] [CrossRef]
- Ozsolak, F.; Milos, P.M. RNA sequencing: Advances, challenges and opportunities. Nat. Rev. Genet. 2010, 12, 87–98. [Google Scholar] [CrossRef]
- Yu, F.; Zhang, Y.; Cheng, C.; Wang, W.; Zhou, Z.; Rang, W.; Yu, H.; Wei, Y.; Wu, Q.; Zhang, Y. Poly(A)-seq: A method for direct sequencing and analysis of the transcriptomic poly(A)-tails. PLoS ONE 2020, 15, e0234696. [Google Scholar] [CrossRef]
- Huang, G.; Zhang, F.; Ye, Q.; Wang, H. The circadian clock regulates autophagy directly through the nuclear hormone receptor Nr1d1/Reverbα and indirectly via Cebpb/(C/ebpβ) in zebrafish. Autophagy 2016, 2, 1292–1309. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Vitaterna, M.H.; Takahashi, J.; Turek, F.W. Overview of Circadian Rhythms. Alcohol Res. Health 2001, 25, 85–93. [Google Scholar] [PubMed]
- Hu, X.; Chen, W.; Li, J.; Huang, S.; Xu, X.; Zhang, X.; Xiang, S.; Liu, C. ZFLNC: A comprehensive and well-annotated database for zebrafish lncRNA. Database 2018, 2018. [Google Scholar] [CrossRef] [PubMed]
- Mishra, S.K.; Wang, H. Computational Analysis Predicts Hundreds of Coding lncRNAs in Zebrafis. Biology 2021, 10, 371. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Wang, Z.; Wang, D.; Qiu, C.; Liu, M.; Chen, X.; Zhang, Q.; Yan, G.; Cui, Q. CPAT: Coding-Potential As-sessment Tool using an alignment-free logistic regression model. Nucleic Acids Res. 2013, 41, 983–986. [Google Scholar] [CrossRef] [Green Version]
- Wittenbrink, N.; Bes, F.; Bartok, O.; Kunz, D.; de Zeeuw, J.; Ananthasubramaniam, B.; Nowozin, C.; Ashwal-Fluss, R.; Kramer, A. High-accuracy determination of internal circadian time from a single blood sample. J. Clin. Investig. 2018, 128, 3826–3839. [Google Scholar] [CrossRef]
- Wu, G.; Anafi, R.C.; Hughes, M.E.; Kornacker, K.; Hogenesch, J.B. MetaCycle: An Integrated R Package to Evalu-ate Periodicity in Large Scale Data”. Bioinformatics 2016, 32, 3351–3353. [Google Scholar] [CrossRef] [Green Version]
- Zielinski, T.; Moore, A.M.; Troup, E.; Halliday, K.J.; Millar, A.J. Strengths and Limitations of Period Estimation Methods for Circadian Data. PLoS ONE 2014, 9, e96462. [Google Scholar] [CrossRef] [Green Version]
- Mudunuri, U.; Che, A.; Yi, M.; Stephens, R.M. bioDBnet: The biological database network. Bioinformatics 2009, 25, 555–556. [Google Scholar] [CrossRef] [Green Version]
- Fornes, O.; Castro-Mondragon, J.A.; Khan, A.; Van Der Lee, R.; Zhang, X.; Richmond, P.A.; Modi, B.; Correard, S.; Gheorghe, M.; Baranašić, D.; et al. JASPAR 2020: Update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 2019, 48, D87–D92. [Google Scholar] [CrossRef]
- Lomax, J. Get ready to GO! A biologist’s guide to the Gene Ontology. Brief. Bioinform. 2005, 6, 298–304. [Google Scholar] [CrossRef] [Green Version]
- Botstein, D.; Cherry, J.M.; Ashburner, M.; Ball, C.A.; Blake, J.A.; Butler, H.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene Ontology: Tool for the unification of biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Götz, S.; García-Gómez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talón, M.; Dopazo, J.; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID Bioinformatics Resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Lever, J.; Krzywinski, M.; Altman, N. Principal component analysis. Nat. Methods 2017, 14, 641–642. [Google Scholar] [CrossRef] [Green Version]
- Gasteiger, E.; Gattiker, A.; Hoogland, C.; Ivanyi, I.; Appel, R.D.; Bairoch, A. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 2003, 31, 3784–3788. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.-C.; Hwang, J.-K.; Yang, J.-M. (PS)2: Protein structure prediction server. Nucleic Acids Res. 2006, 34, W152–W157. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.-C.; Hwang, J.-K.; Yang, J.-M. (PS)2-v2: Template-based protein structure prediction server. BMC Bioinform. 2009, 10, 366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mishra, S.K.; Liu, T.; Wang, H. Identification of Rhythmically Expressed LncRNAs in the Zebrafish Pineal Gland and Testis. Int. J. Mol. Sci. 2021, 22, 7810. https://doi.org/10.3390/ijms22157810
Mishra SK, Liu T, Wang H. Identification of Rhythmically Expressed LncRNAs in the Zebrafish Pineal Gland and Testis. International Journal of Molecular Sciences. 2021; 22(15):7810. https://doi.org/10.3390/ijms22157810
Chicago/Turabian StyleMishra, Shital Kumar, Taole Liu, and Han Wang. 2021. "Identification of Rhythmically Expressed LncRNAs in the Zebrafish Pineal Gland and Testis" International Journal of Molecular Sciences 22, no. 15: 7810. https://doi.org/10.3390/ijms22157810
APA StyleMishra, S. K., Liu, T., & Wang, H. (2021). Identification of Rhythmically Expressed LncRNAs in the Zebrafish Pineal Gland and Testis. International Journal of Molecular Sciences, 22(15), 7810. https://doi.org/10.3390/ijms22157810





