Non-Coding RNAs: Novel Players in Insulin Resistance and Related Diseases
Abstract
1. Introduction
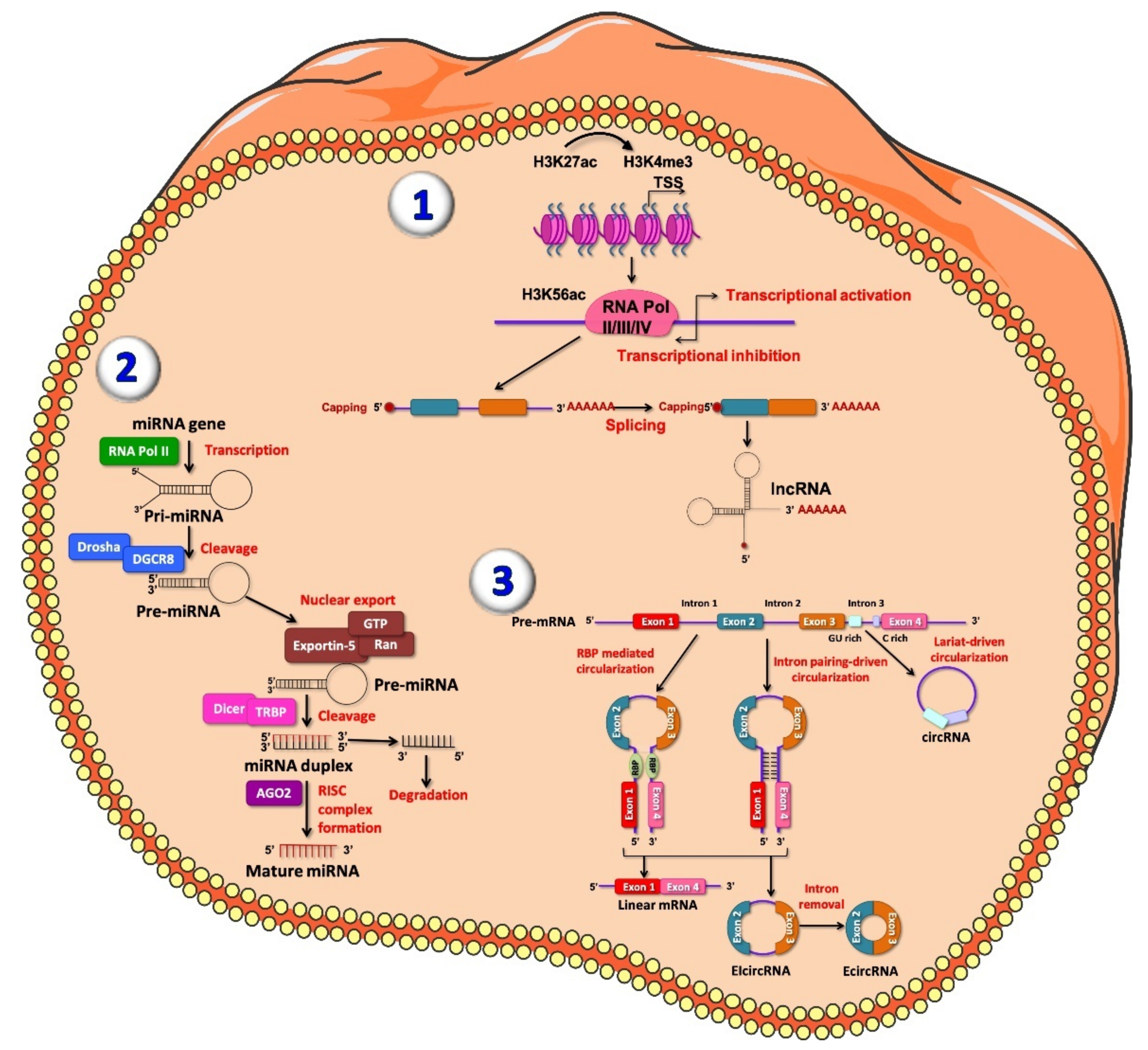
2. Non-Coding RNAs Biogenesis and Function
2.1. Long Non-Coding RNAs
2.2. MicroRNAs
2.3. Circular RNAs
3. Non-Coding RNAs in Metabolic Diseases: State of Art
3.1. Long Non-Coding RNAs
3.1.1. Long Non-Coding RNAs and Obesity
3.1.2. Long Non-Coding RNAs and Type 2 Diabetes Mellitus
3.1.3. Long Non-Coding RNAs and NAFLD/NASH
3.2. Micro RNAs
3.2.1. MicroRNAs and Obesity
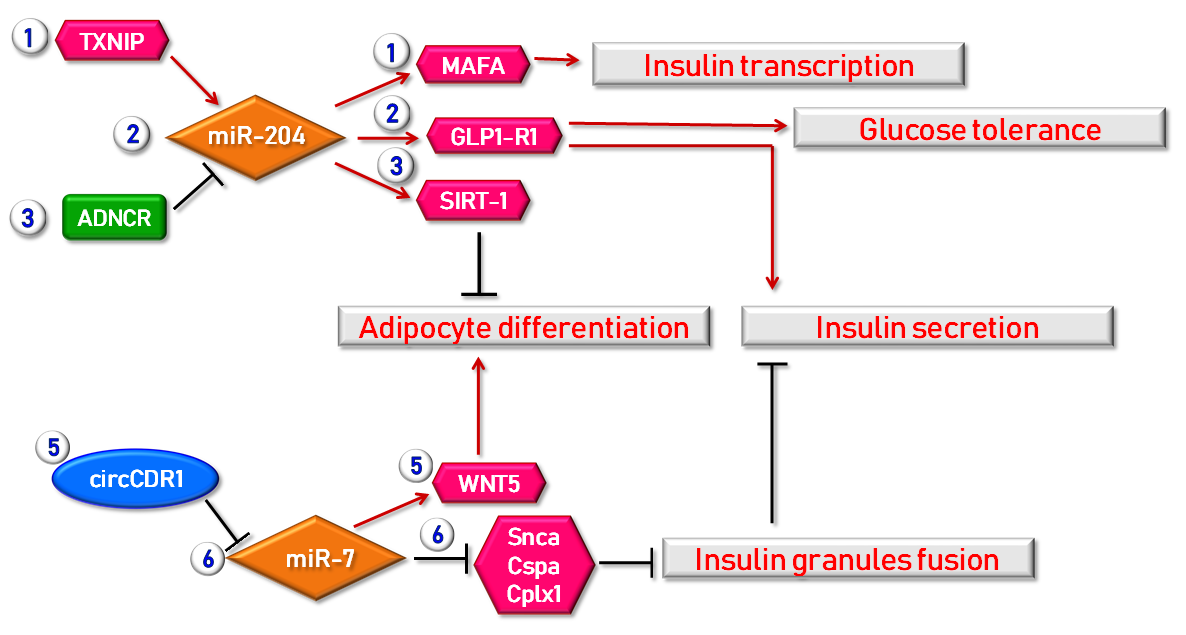
3.2.2. MicroRNAs and Type 2 Diabetes Mellitus
3.2.3. MicroRNAs and NAFLD/NASH
3.3. Circular RNAs
3.3.1. Circular RNAs and Obesity
3.3.2. Circular RNAs and Type 2 Diabetes Mellitus
3.3.3. Circular RNAs and NAFLD/NASH
4. Potential Clinical Application of Non-Coding RNAs
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Al-Tobasei, R.; Paneru, B.; Salem, M. Genome-Wide Discovery of Long Non-Coding RNAs in Rainbow Trout. PLoS ONE 2016, 11, e0148940. [Google Scholar] [CrossRef] [PubMed]
- Dragomir, M.P.; Knutsen, E.; Calin, G.A. SnapShot: Unconventional miRNA Functions. Cell 2018, 174, 1038–1038.e1. [Google Scholar] [CrossRef] [PubMed]
- Knoll, M.; Lodish, H.F.; Sun, L. Long non-coding RNAs as regulators of the endocrine system. Nat. Rev. Endocrinol. 2015, 11, 151–160. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, J.; Acuña, S.; Aoki, J.; Floeter-Winter, L.; Muxel, S. Long Non-Coding RNAs in the Regulation of Gene Expression: Physiology and Disease. Non-Coding RNA 2019, 5, 17. [Google Scholar] [CrossRef]
- Zaiou, M. The Emerging Role and Promise of Circular RNAs in Obesity and Related Metabolic Disorders. Cells 2020, 9, 1473. [Google Scholar] [CrossRef]
- Hangauer, M.J.; Vaughn, I.W.; McManus, M.T. Pervasive Transcription of the Human Genome Produces Thousands of Previously Unidentified Long Intergenic Noncoding RNAs. PLoS Genet. 2013, 9, e1003569. [Google Scholar] [CrossRef]
- Taheri, M.; Eghtedarian, R.; Ghafouri-Fard, S.; Omrani, M.D. Non-coding RNAs and type 2 diabetes mellitus. Arch. Physiol. Biochem. 2020, 1–10. [Google Scholar] [CrossRef]
- Ardekani, A.M.; Naeini, M.M. The Role of MicroRNAs in Human Diseases. In miRNomics: MicroRNA Biology and Computational Analysis; Humana Press: Totowa, NJ, USA, 2010; Volume 2, pp. 161–179. [Google Scholar]
- Sathishkumar, C.; Prabu, P.; Mohan, V.; Balasubramanyam, M. Linking a role of lncRNAs (long non-coding RNAs) with insulin resistance, accelerated senescence, and inflammation in patients with type 2 diabetes. Hum. Genom. 2018, 12, 1–9. [Google Scholar] [CrossRef]
- Ghasemi, H.; Sabati, Z.; Ghaedi, H.; Salehi, Z.; Alipoor, B. Circular RNAs in β-cell function and type 2 diabetes-related complications: A potential diagnostic and therapeutic approach. Mol. Biol. Rep. 2019, 46, 5631–5643. [Google Scholar] [CrossRef]
- Zimmet, P.; Magliano, D.; Matsuzawa, Y.; Alberti, G.; Shaw, J. The Metabolic Syndrome: A Global Public Health Problem and A New Definition. J. Atheroscler. Thromb. 2005, 12, 295–300. [Google Scholar] [CrossRef]
- Swinburn, B.A.; Sacks, G.; Hall, K.D.; McPherson, K.; Finegood, D.T.; Moodie, M.L.; Gortmaker, S.L. The global obesity pandemic: Shaped by global drivers and local environments. Lancet 2011, 378, 804–814. [Google Scholar] [CrossRef]
- Zaiou, M. Circular RNAs as Potential Biomarkers and Therapeutic Targets for Metabolic Diseases. In Reviews on Biomarker Studies of Metabolic and Metabolism-Related Disorders; Advances in Experimental Medicine and Biology; Guest, P.C., Ed.; Springer International Publishing: Cham, Switzerland, 2019; Volume 1134, pp. 177–191. ISBN 978-3-030-12667-4. [Google Scholar]
- Reaven, G.M. The Insulin Resistance Syndrome: Definition and Dietary Approaches to Treatment. Annu. Rev. Nutr. 2005, 25, 391–406. [Google Scholar] [CrossRef]
- Martí, A.; Martinez-González, M.A.; Martínez, J.A. Interaction between genes and lifestyle factors on obesity: Nutrition Society Silver Medal Lecture. Proc. Nutr. Soc. 2008, 67, 1–8. [Google Scholar] [CrossRef]
- Lumeng, C.N.; Saltiel, A. Inflammatory links between obesity and metabolic disease. J. Clin. Investig. 2011, 121, 2111–2117. [Google Scholar] [CrossRef]
- Friedman, J.M. Obesity in the new millennium. Nat. Cell Biol. 2000, 404, 632–634. [Google Scholar] [CrossRef]
- American Diabetes Association. Classification and Diagnosis of Diabetes: Standards of Medical Care in Diabetes—2018. Diabetes Care 2021, 44, S15–S33. [Google Scholar] [CrossRef]
- European Association for the Study of the Liver (EASL); European Association for the Study of Diabetes (EASD). European Association for the Study of Obesity (EASO) EASL–EASD–EASO Clinical Practice Guidelines for the management of non-alcoholic fatty liver disease. J. Hepatol. 2016, 64, 1388–1402. [Google Scholar] [CrossRef]
- Cobbina, E.; Akhlaghi, F. Non-alcoholic fatty liver disease (NAFLD)-pathogenesis, classification, and effect on drug metabolizing enzymes and transporters. Drug Metab. Rev. 2017, 49, 197–211. [Google Scholar] [CrossRef]
- Khalifa, O.; Errafii, K.; Al-Akl, N.S.; Arredouani, A. Noncoding RNAs in Nonalcoholic Fatty Liver Disease: Potential Diagnosis and Prognosis Biomarkers. Dis. Markers 2020, 2020, 8822859. [Google Scholar] [CrossRef]
- Friedman, S.L.; Neuschwander-Tetri, B.A.; Rinella, M.; Sanyal, A.J. Mechanisms of NAFLD development and therapeutic strategies. Nat. Med. 2018, 24, 908–922. [Google Scholar] [CrossRef]
- López-Pastor, A.R.; Infante-Menéndez, J.; Escribano, Ó.; Gómez-Hernández, A. miRNA Dysregulation in the Development of Non-Alcoholic Fatty Liver Disease and the Related Disorders Type 2 Diabetes Mellitus and Cardiovascular Disease. Front. Med. 2020, 7, 527059. [Google Scholar] [CrossRef]
- Byrne, C.D.; Targher, G. NAFLD: A multisystem disease. J. Hepatol. 2015, 62, S47–S64. [Google Scholar] [CrossRef]
- Perumpail, B.J.; Khan, M.A.; Yoo, E.R.; Cholankeril, G.; Kim, D.; Ahmed, A. Clinical epidemiology and disease burden of nonalcoholic fatty liver disease. World J. Gastroenterol. 2017, 23, 8263–8276. [Google Scholar] [CrossRef]
- Diehl, A.M.; Day, C. Cause, Pathogenesis, and Treatment of Nonalcoholic Steatohepatitis. N. Engl. J. Med. 2017, 377, 2063–2072. [Google Scholar] [CrossRef]
- Marra, F.; Gastaldelli, A.; Baroni, G.S.; Tell, G.; Tiribelli, C. Molecular basis and mechanisms of progression of non-alcoholic steatohepatitis. Trends Mol. Med. 2008, 14, 72–81. [Google Scholar] [CrossRef]
- Petersen, M.C.; Shulman, G.I. Mechanisms of Insulin Action and Insulin Resistance. Physiol. Rev. 2018, 98, 2133–2223. [Google Scholar] [CrossRef] [PubMed]
- Han, H.-S.; Kang, G.; Kim, J.S.; Choi, B.H.; Koo, S.-H. Regulation of glucose metabolism from a liver-centric perspective. Exp. Mol. Med. 2016, 48, e218. [Google Scholar] [CrossRef]
- Beale, E.G. Insulin Signaling and Insulin Resistance. J. Investig. Med. 2013, 61, 11–14. [Google Scholar] [CrossRef]
- Fuchsberger, C.; Flannick, J.; Teslovich, T.M.; Mahajan, A.; Agarwala, V.; Gaulton, K.J.; Ma, C.; Fontanillas, P.; Moutsianas, L.; McCarthy, D.; et al. The genetic architecture of type 2 diabetes. Nat. Cell Biol. 2016, 536, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo, A.G.; Crujeiras, A.B. Obesity-Related Epigenetic Changes after Bariatric Surgery. Front. Endocrinol. 2019, 10, 232. [Google Scholar] [CrossRef]
- Lee, J.T. Lessons from X-chromosome inactivation: Long ncRNA as guides and tethers to the epigenome. Genes Dev. 2009, 23, 1831–1842. [Google Scholar] [CrossRef] [PubMed]
- Flynn, R.; Chang, H.Y. Long Noncoding RNAs in Cell-Fate Programming and Reprogramming. Cell Stem Cell 2014, 14, 752–761. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.-H.; Hwang, H.-J.; Cho, J.-Y. Long Non-Coding RNA Associated with Cholesterol Homeostasis and Its Involvement in Metabolic Diseases. Int. J. Mol. Sci. 2020, 21, 8337. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.; Du, M.; Jiang, Z.; Hausman, G.J.; Zhang, L.; Dodson, M.V. Long noncoding RNAs in regulating adipogenesis: New RNAs shed lights on obesity. Cell. Mol. Life Sci. 2016, 73, 2079–2087. [Google Scholar] [CrossRef]
- Pielok, A.; Marycz, K. Non-Coding RNAs as Potential Novel Biomarkers for Early Diagnosis of Hepatic Insulin Resistance. Int. J. Mol. Sci. 2020, 21, 4182. [Google Scholar] [CrossRef]
- Assmann, T.S.; Cuevas-Sierra, A.; Salas-Pérez, F.; Riezu-Boj, J.I.; Milagro, F.I.; Martínez, J.A. Crosstalk between circulating microRNAs and chronotypical features in subjects with metabolic syndrome. Chronobiol. Int. 2020, 37, 1048–1058. [Google Scholar] [CrossRef]
- Guo, X.; Gao, L.; Wang, Y.; Chiu, D.K.Y.; Wang, T.; Deng, Y. Advances in long noncoding RNAs: Identification, structure prediction and function annotation. Brief. Funct. Genom. 2016, 15, 38–46. [Google Scholar] [CrossRef]
- Statello, L.; Guo, C.-J.; Chen, L.-L.; Huarte, M. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 2021, 22, 96–118. [Google Scholar] [CrossRef]
- Dhanoa, J.K.; Sethi, R.S.; Verma, R.; Arora, J.S.; Mukhopadhyay, C.S. Long non-coding RNA: Its evolutionary relics and biological implications in mammals: A review. J. Anim. Sci. Technol. 2018, 60, 25. [Google Scholar] [CrossRef]
- Huang, Y.; Liu, N.; Wang, J.P.; Wang, Y.Q.; Yu, X.L.; Bin Wang, Z.; Cheng, X.C.; Zou, Q. Regulatory long non-coding RNA and its functions. J. Physiol. Biochem. 2012, 68, 611–618. [Google Scholar] [CrossRef]
- Mercer, T.; Dinger, M.; Mattick, J. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef]
- Hung, T.; Wang, Y.; Lin, M.F.; Koegel, A.K.; Kotake, Y.; Grant, G.D.; Horlings, H.M.; Shah, N.; Umbricht, C.; Wang, P.; et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat. Genet. 2011, 43, 621–629. [Google Scholar] [CrossRef]
- Yap, K.L.; Li, S.; Muñoz-Cabello, A.M.; Raguz, S.; Zeng, L.; Mujtaba, S.; Gil, J.; Walsh, M.J.; Zhou, M.-M. Molecular Interplay of the Noncoding RNA ANRIL and Methylated Histone H3 Lysine 27 by Polycomb CBX7 in Transcriptional Silencing of INK4a. Mol. Cell 2010, 38, 662–674. [Google Scholar] [CrossRef]
- Keniry, A.; Oxley, D.; Monnier, P.; Kyba, M.; Dandolo, L.; Smits, G.; Reik, W. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat. Cell Biol. 2012, 14, 659–665. [Google Scholar] [CrossRef]
- Patil, V.S.; Zhou, R.; Rana, T.M. Gene regulation by non-coding RNAs. Crit. Rev. Biochem. Mol. Biol. 2014, 49, 16–32. [Google Scholar] [CrossRef]
- Andersen, A.; Panning, B. Epigenetic gene regulation by noncoding RNAs. Curr. Opin. Cell Biol. 2003, 15, 281–289. [Google Scholar] [CrossRef]
- Wang, K.C.; Chang, H.Y. Molecular Mechanisms of Long Noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef]
- Rinn, J.L.; Chang, H.Y. Genome Regulation by Long Noncoding RNAs. Annu. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef]
- Tsai, M.-C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long Noncoding RNA as Modular Scaffold of Histone Modification Complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef]
- Pasman, Z.; Been, M.D.; Garcia-Blanco, M.A. Exon circularization in mammalian nuclear extracts. RNA 1996, 2, 603–610. [Google Scholar]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef]
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.-H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004, 23, 4051–4060. [Google Scholar] [CrossRef]
- Denli, A.M.; Tops, B.; Plasterk, R.H.A.; Ketting, R.F.; Hannon, G.J. Processing of primary microRNAs by the Microprocessor complex. Nat. Cell Biol. 2004, 432, 231–235. [Google Scholar] [CrossRef]
- Yi, R.; Qin, Y.; Macara, I.G.; Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 2003, 17, 3011–3016. [Google Scholar] [CrossRef]
- Okada, C.; Yamashita, E.; Lee, S.J.; Shibata, S.; Katahira, J.; Nakagawa, A.; Yoneda, Y.; Tsukihara, T. A High-Resolution Structure of the Pre-microRNA Nuclear Export Machinery. Science 2009, 326, 1275–1279. [Google Scholar] [CrossRef]
- Yoda, M.; Kawamata, T.; Paroo, Z.; Ye, X.; Iwasaki, S.; Liu, Q.; Tomari, Y. ATP-dependent human RISC assembly pathways. Nat. Struct. Mol. Biol. 2010, 17, 17–23. [Google Scholar] [CrossRef]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef]
- Yang, J.-S.; Maurin, T.; Robine, N.; Rasmussen, K.D.; Jeffrey, K.; Chandwani, R.; Papapetrou, E.P.; Sadelain, M.; O’Carroll, D.; Lai, E.C. Conserved vertebrate mir-451 provides a platform for Dicer-independent, Ago2-mediated microRNA biogenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 15163–15168. [Google Scholar] [CrossRef] [PubMed]
- Xie, M.; Li, M.; Vilborg, A.; Lee, N.; Shu, M.-D.; Yartseva, V.; Šestan, N.; Steitz, J.A. Mammalian 5′-Capped MicroRNA Precursors that Generate a Single MicroRNA. Cell 2013, 155, 1568–1580. [Google Scholar] [CrossRef] [PubMed]
- Kawamata, T.; Tomari, Y. Making RISC. Trends Biochem. Sci. 2010, 35, 368–376. [Google Scholar] [CrossRef] [PubMed]
- Felekkis, K.; Touvana, E.; Stefanou, C.; Deltas, C. microRNAs: A newly described class of encoded molecules that play a role in health and disease. Hippokratia 2010, 14, 236–240. [Google Scholar]
- Meijer, H.A.; Smith, E.; Bushell, M. Regulation of miRNA strand selection: Follow the leader? Biochem. Soc. Trans. 2014, 42, 1135–1140. [Google Scholar] [CrossRef]
- Catalanotto, C.; Cogoni, C.; Zardo, G. MicroRNA in Control of Gene Expression: An Overview of Nuclear Functions. Int. J. Mol. Sci. 2016, 17, 1712. [Google Scholar] [CrossRef]
- Chen, X.; Ba, Y.; Ma, L.; Cai, X.; Yin, Y.; Wang, K.; Guo, J.; Zhang, Y.; Chen, J.; Guo, X.; et al. Characterization of microRNAs in serum: A novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008, 18, 997–1006. [Google Scholar] [CrossRef]
- Arroyo, J.; Chevillet, J.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.; Bennett, C.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef]
- Gallo, A.; Tandon, M.; Alevizos, I.; Illei, G.G. The Majority of MicroRNAs Detectable in Serum and Saliva Is Concentrated in Exosomes. PLoS ONE 2012, 7, e30679. [Google Scholar] [CrossRef]
- Zhou, Q.; Li, M.; Wang, X.; Li, Q.; Wang, T.; Zhou, X.; Wang, X.; Gao, X.; Li, X. Immune-related MicroRNAs are Abundant in Breast Milk Exosomes. Int. J. Biol. Sci. 2012, 8, 118–123. [Google Scholar] [CrossRef]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; Huang, K.-H.; Lee, M.-J.; Galas, D.J.; Wang, K. The MicroRNA Spectrum in 12 Body Fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef]
- Théry, C.; Zitvogel, L.; Amigorena, S. Exosomes: Composition, biogenesis and function. Nat. Rev. Immunol. 2002, 2, 569–579. [Google Scholar] [CrossRef]
- Théry, C.; Amigorena, S.; Raposo, G.; Clayton, A. Isolation and Characterization of Exosomes from Cell Culture Supernatants and Biological Fluids. Curr. Protoc. Cell Biol. 2006, 30, 3.22.1–3.22.29. [Google Scholar] [CrossRef]
- Crescitelli, R.; Lässer, C.; Szabó, T.G.; Kittel, A.; Eldh, M.; Dianzani, I.; Buzás, E.I.; Lotvall, J. Distinct RNA profiles in subpopulations of extracellular vesicles: Apoptotic bodies, microvesicles and exosomes. J. Extracell. Vesicles 2013, 2. [Google Scholar] [CrossRef]
- Geekiyanage, H.; Rayatpisheh, S.; Wohlschlegel, J.A.; Brown, R.; Ambros, V. Extracellular microRNAs in human circulation are associated with miRISC complexes that are accessible to anti-AGO2 antibody and can bind target mimic oligonucleotides. Proc. Natl. Acad. Sci. USA 2020, 117, 24213–24223. [Google Scholar] [CrossRef]
- Grieco, G.E.; Sebastiani, G.; Eandi, C.M.; Neri, G.; Nigi, L.; Brusco, N.; D’Aurizio, R.; Posarelli, M.; Bacci, T.; De Benedetto, E.; et al. MicroRNA Expression in the Aqueous Humor of Patients with Diabetic Macular Edema. Int. J. Mol. Sci. 2020, 21, 7328. [Google Scholar] [CrossRef]
- Guarino, E.; Poggi, C.D.; Grieco, G.E.; Cenci, V.; Ceccarelli, E.; Crisci, I.; Sebastiani, G.; Dotta, F. Circulating MicroRNAs as Biomarkers of Gestational Diabetes Mellitus: Updates and Perspectives. Int. J. Endocrinol. 2018, 2018, 6380463. [Google Scholar] [CrossRef]
- Sebastiani, G.; Nigi, L.; Grieco, G.E.; Mancarella, F.; Ventriglia, G.; Dotta, F. Circulating microRNAs and diabetes mellitus: A novel tool for disease prediction, diagnosis, and staging? J. Endocrinol. Investig. 2017, 40, 591–610. [Google Scholar] [CrossRef] [PubMed]
- Finnegan, E.F.; Pasquinelli, A.E. MicroRNA biogenesis: Regulating the regulators. Crit. Rev. Biochem. Mol. Biol. 2013, 48, 51–68. [Google Scholar] [CrossRef]
- Wilczynska, A.; Bushell, M. The complexity of miRNA-mediated repression. Cell Death Differ. 2015, 22, 22–33. [Google Scholar] [CrossRef]
- Winter, J.; Jung, S.; Keller, S.; Gregory, R.I.; Diederichs, S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat. Cell Biol. 2009, 11, 228–234. [Google Scholar] [CrossRef]
- Mo, Y.-Y. MicroRNA regulatory networks and human disease. Cell. Mol. Life Sci. 2012, 69, 3529–3531. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Ding, Y.; Liang, B.; Lin, J.; Kim, T.-K.; Yu, H.; Hang, H.; Wang, K. A Systematic Study of Dysregulated MicroRNA in Type 2 Diabetes Mellitus. Int. J. Mol. Sci. 2017, 18, 456. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-O.; Wang, H.-B.; Zhang, Y.; Lu, X.; Chen, L.-L.; Yang, L. Complementary Sequence-Mediated Exon Circularization. Cell 2014, 159, 134–147. [Google Scholar] [CrossRef] [PubMed]
- Arnberg, A.; Van Ommen, G.-J.; Grivell, L.; Van Bruggen, E.; Borst, P. Some yeast mitochondrial RNAs are circular. Cell 1980, 19, 313–319. [Google Scholar] [CrossRef]
- Kos, A.; Dijkema, R.; Arnberg, A.C.; Van Der Meide, P.H.; Schellekens, H. The hepatitis delta (δ) virus possesses a circular RNA. Nat. Cell Biol. 1986, 323, 558–560. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.-L. The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat. Rev. Mol. Cell Biol. 2020, 21, 475–490. [Google Scholar] [CrossRef]
- Kristensen, L.S.; Andersen, M.S.; Stagsted, L.V.W.; Ebbesen, K.K.; Hansen, T.B.; Kjems, J. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 2019, 20, 675–691. [Google Scholar] [CrossRef]
- Chen, L.-L.; Yang, L. Regulation of circRNA biogenesis. RNA Biol. 2015, 12, 381–388. [Google Scholar] [CrossRef]
- Zaphiropoulos, P. Circular RNAs from transcripts of the rat cytochrome P450 2C24 gene: Correlation with exon skipping. Proc. Natl. Acad. Sci. USA 1996, 93, 6536–6541. [Google Scholar] [CrossRef]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, E.N. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013, 19, 141–157. [Google Scholar] [CrossRef]
- Li, X.; Liu, S.; Zhang, L.; Issaian, A.; Hill, R.C.; Espinosa, S.; Shi, S.; Cui, Y.; Kappel, K.; Das, R.; et al. A unified mechanism for intron and exon definition and back-splicing. Nat. Cell Biol. 2019, 573, 375–380. [Google Scholar] [CrossRef]
- Zhao, X.; Cai, Y.; Xu, J. Circular RNAs: Biogenesis, Mechanism, and Function in Human Cancers. Int. J. Mol. Sci. 2019, 20, 3926. [Google Scholar] [CrossRef]
- Yu, C.-Y.; Kuo, H.-C. The emerging roles and functions of circular RNAs and their generation. J. Biomed. Sci. 2019, 26, 29. [Google Scholar] [CrossRef]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, X.-O.; Chen, T.; Xiang, J.-F.; Yin, Q.-F.; Xing, Y.-H.; Zhu, S.; Yang, L.; Chen, L.-L. Circular Intronic Long Noncoding RNAs. Mol. Cell 2013, 51, 792–806. [Google Scholar] [CrossRef]
- Payne, J.M.; Laybourn, P.J.; Dahmus, M.E. The transition of RNA polymerase II from initiation to elongation is associated with phosphorylation of the carboxyl-terminal domain of subunit IIa. J. Biol. Chem. 1989, 264, 19621–19629. [Google Scholar] [CrossRef]
- Latreille, M.; Hausser, J.; Stützer, I.; Zhang, Q.; Hastoy, B.; Gargani, S.; Kerr-Conte, J.; Pattou, F.; Zavolan, M.; Esguerra, J.; et al. MicroRNA-7a regulates pancreatic β cell function. J. Clin. Investig. 2014, 124, 2722–2735. [Google Scholar] [CrossRef]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Xu, H.; Guo, S.; Li, W.; Yu, P. The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci. Rep. 2015, 5, 12453. [Google Scholar] [CrossRef]
- Lorente-Cebrián, S.; González-Muniesa, P.; Milagro, F.; Martínez, J.A. MicroRNAs and other non-coding RNAs in adipose tissue and obesity: Emerging roles as biomarkers and therapeutic targets. Clin. Sci. 2019, 133, 23–40. [Google Scholar] [CrossRef]
- Squillaro, T.; Peluso, G.; Galderisi, U.; Di Bernardo, G. Long non-coding RNAs in regulation of adipogenesis and adipose tissue function. eLife 2020, 9, e59053. [Google Scholar] [CrossRef]
- Giroud, M.; Scheideler, M. Long Non-Coding RNAs in Metabolic Organs and Energy Homeostasis. Int. J. Mol. Sci. 2017, 1, 8. [Google Scholar] [CrossRef]
- Gao, H.; Kerr, A.; Jiao, H.; Hon, C.-C.; Rydén, M.; Dahlman, I.; Arner, P. Long Non-Coding RNAs Associated with Metabolic Traits in Human White Adipose Tissue. EBioMedicine 2018, 30, 248–260. [Google Scholar] [CrossRef]
- Li, M.; Sun, X.; Cai, H.; Sun, Y.; Plath, M.; Li, C.; Lan, X.; Lei, C.; Lin, F.; Bai, Y.; et al. Long non-coding RNA ADNCR suppresses adipogenic differentiation by targeting miR-204. Biochim. Biophys. Acta (BBA) Bioenerg. 2016, 1859, 871–882. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Zheng, Y.; Jin, C.; Li, X.; Jia, L.; Li, W. Long Non-coding RNA H19 Inhibits Adipocyte Differentiation of Bone Marrow Mesenchymal Stem Cells through Epigenetic Modulation of Histone Deacetylases. Sci. Rep. 2016, 6, 28897. [Google Scholar] [CrossRef]
- Divoux, A.; Karastergiou, K.; Xie, H.; Guo, W.; Perera, R.J.; Fried, S.K.; Smith, S.R. Identification of a novel lncRNA in gluteal adipose tissue and evidence for its positive effect on preadipocyte differentiation: LncRNA HOTAIR in Gluteal Adipose Tissue. Obesity 2014, 22, 1781–1785. [Google Scholar] [CrossRef]
- Zhao, X.-Y.; Li, S.; Wang, G.-X.; Yu, Q.; Lin, J.D. A Long Noncoding RNA Transcriptional Regulatory Circuit Drives Thermogenic Adipocyte Differentiation. Mol. Cell 2014, 55, 372–382. [Google Scholar] [CrossRef]
- Schmidt, E.; Dhaouadi, I.; Gaziano, I.; Oliverio, M.; Klemm, P.; Awazawa, M.; Mitterer, G.; Fernandez-Rebollo, E.; Pradas-Juni, M.; Wagner, W.; et al. LincRNA H19 protects from dietary obesity by constraining expression of monoallelic genes in brown fat. Nat. Commun. 2018, 9, 1–16. [Google Scholar] [CrossRef]
- Sun, J.; Ruan, Y.; Wang, M.; Chen, R.; Yu, N.; Sun, L.; Liu, T.; Chen, H. Differentially expressed circulating LncRNAs and mRNA identified by microarray analysis in obese patients. Sci. Rep. 2016, 6, 35421. [Google Scholar] [CrossRef]
- Liang, Y.; Yu, B.; Wang, Y.; Qiao, Z.; Cao, T.; Zhang, P. Duodenal long noncoding RNAs are associated with glycemic control after bariatric surgery in high-fat diet-induced diabetic mice. Surg. Obes. Relat. Dis. 2017, 13, 1212–1226. [Google Scholar] [CrossRef]
- Liu, C.; Yang, Z.; Wu, J.; Zhang, L.; Lee, S.; Shin, D.-J.; Tran, M.; Wang, L. Long noncoding RNA H19 interacts with polypyrimidine tract-binding protein 1 to reprogram hepatic lipid homeostasis. Hepatology 2018, 67, 1768–1783. [Google Scholar] [CrossRef]
- Tello-Flores, V.A.; Valladares-Salgado, A.; Ramírez-Vargas, M.A.; Cruz, M.; Del-Moral-Hernández, O.; Cahua-Pablo, J.Á.; Ramírez-Ruano, M.; Hernández-Sotelo, D.; Armenta-Solis, A.; Flores-Alfaro, E. Altered levels of MALAT1 and H19 derived from serum or serum exosomes associated with type-2 diabetes. Non-Coding RNA Res. 2020, 5, 71–76. [Google Scholar] [CrossRef]
- Gong, Y.; Zhu, Y.; Zhu, B.; Si, X.; Heng, D.; Tang, Y.; Sun, X.; Lin, L. LncRNA MALAT1 is up-regulated in diabetic gastroparesis and involved in high-glucose-induced cellular processes in human gastric smooth muscle cells. Biochem. Biophys. Res. Commun. 2018, 496, 401–406. [Google Scholar] [CrossRef] [PubMed]
- Leti, F.; Legendre, C.; Still, C.D.; Chu, X.; Petrick, A.; Gerhard, G.S.; DiStefano, J.K. Altered expression of MALAT1 lncRNA in nonalcoholic steatohepatitis fibrosis regulates CXCL5 in hepatic stellate cells. Transl. Res. 2017, 190, 25–39.e21. [Google Scholar] [CrossRef] [PubMed]
- Uchida, S.; Kauppinen, S. Long Non-Coding RNAs in Liver Cancer and Nonalcoholic Steatohepatitis. Non-Coding RNA 2020, 6, 34. [Google Scholar] [CrossRef]
- Tang, N.; Jiang, S.; Yang, Y.; Liu, S.; Ponnusamy, M.; Xin, H.; Yu, T. Noncoding RNAs as therapeutic targets in atherosclerosis with diabetes mellitus. Cardiovasc. Ther. 2018, 36, e12436. [Google Scholar] [CrossRef]
- Zhang, L.; Wang, Y.-M. Expression and function of lncRNA ANRIL in a mouse model of acute myocardial infarction combined with type 2 diabetes mellitus. J. Chin. Med. Assoc. 2019, 82, 685–692. [Google Scholar] [CrossRef]
- Wang, L.; Su, N.; Zhang, Y.; Wang, G. Clinical Significance of Serum lncRNA Cancer Susceptibility Candidate 2 (CASC2) for Chronic Renal Failure in Patients with Type 2 Diabetes. Med. Sci. Monit. 2018, 24, 6079–6084. [Google Scholar] [CrossRef]
- Yu, W.; Zhao, G.-Q.; Cao, R.-J.; Zhu, Z.; Li, K. LncRNA NONRATT021972 Was Associated with Neuropathic Pain Scoring in Patients with Type 2 Diabetes. Behav. Neurol. 2017, 2017, 2941297. [Google Scholar] [CrossRef]
- Zhang, M.; Chi, X.; Qu, N.; Wang, C. Long noncoding RNA lncARSR promotes hepatic lipogenesis via Akt/SREBP-1c pathway and contributes to the pathogenesis of nonalcoholic steatohepatitis. Biochem. Biophys. Res. Commun. 2018, 499, 66–70. [Google Scholar] [CrossRef]
- Atanasovska, B.; Rensen, S.S.; Van Der Sijde, M.R.; Marsman, G.; Kumar, V.; Jonkers, I.; Withoff, S.; Shiri-Sverdlov, R.; Greve, J.W.M.; Faber, K.N.; et al. A liver-specific long noncoding RNA with a role in cell viability is elevated in human nonalcoholic steatohepatitis. Hepatology 2017, 66, 794–808. [Google Scholar] [CrossRef]
- Hanson, A.; Wilhelmsen, D.; Distefano, J.K. The Role of Long Non-Coding RNAs (lncRNAs) in the Development and Progression of Fibrosis Associated with Nonalcoholic Fatty Liver Disease (NAFLD). Non-Coding RNA 2018, 4, 18. [Google Scholar] [CrossRef]
- De Rosa, S.; Arcidiacono, B.; Chiefari, E.; Brunetti, A.; Indolfi, C.; Foti, D.P. Type 2 Diabetes Mellitus and Cardiovascular Disease: Genetic and Epigenetic Links. Front. Endocrinol. 2018, 9, 2. [Google Scholar] [CrossRef]
- Guo, J.; Liu, Z.; Gong, R. Long noncoding RNA: An emerging player in diabetes and diabetic kidney disease. Clin. Sci. 2019, 133, 1321–1339. [Google Scholar] [CrossRef] [PubMed]
- Goyal, N.; Kesharwani, D.; Datta, M. Lnc-ing non-coding RNAs with metabolism and diabetes: Roles of lncRNAs. Cell. Mol. Life Sci. 2018, 75, 1827–1837. [Google Scholar] [CrossRef]
- Wang, X.; Chang, X.; Zhang, P.; Fan, L.; Zhou, T.; Sun, K. Aberrant Expression of Long Non-Coding RNAs in Newly Diagnosed Type 2 Diabetes Indicates Potential Roles in Chronic Inflammation and Insulin Resistance. Cell. Physiol. Biochem. 2017, 43, 2367–2378. [Google Scholar] [CrossRef] [PubMed]
- Sulaiman, S.A.; Muhsin, N.I.A.; Jamal, R. Regulatory Non-coding RNAs Network in Non-alcoholic Fatty Liver Disease. Front. Physiol. 2019, 10, 279. [Google Scholar] [CrossRef]
- Ji, E.; Kim, C.; Kim, W.; Lee, E.K. Role of long non-coding RNAs in metabolic control. Biochim. Biophys. Acta (BBA) Bioenerg. 2020, 1863, 194348. [Google Scholar] [CrossRef]
- Sookoian, S.; Rohr, C.; Salatino, A.; Dopazo, H.; Gianotti, T.F.; Castaño, G.O.; Pirola, C.J. Genetic variation in long noncoding RNAs and the risk of nonalcoholic fatty liver disease. Oncotarget 2017, 8, 22917–22926. [Google Scholar] [CrossRef]
- Kunej, T.; Skok, D.J.; Zorc, M.; Ogrinc, A.; Michal, J.J.; Kovac, M.; Jiang, Z. Obesity Gene Atlas in Mammals. Int. J. Genom. 2013, 1, 45–55. [Google Scholar] [CrossRef][Green Version]
- Klöting, N.; Berthold, S.; Kovacs, P.; Schön, M.; Fasshauer, M.; Ruschke, K.; Stumvoll, M.; Blüher, M. MicroRNA Expression in Human Omental and Subcutaneous Adipose Tissue. PLoS ONE 2009, 4, e4699. [Google Scholar] [CrossRef]
- Ortega, F.J.; Moreno-Navarrete, J.M.; Pardo, G.; Sabater, M.; Hummel, M.; Ferrer, A.; Rodriguez-Hermosa, J.I.; Ruiz, B.; Ricart, W.; Peral, B.; et al. MiRNA Expression Profile of Human Subcutaneous Adipose and during Adipocyte Differentiation. PLoS ONE 2010, 5, e9022. [Google Scholar] [CrossRef]
- Kim, Y.; Kim, O.-K. Potential Roles of Adipocyte Extracellular Vesicle–Derived miRNAs in Obesity-Mediated Insulin Resistance. Adv. Nutr. 2021, 12, 566–574. [Google Scholar] [CrossRef] [PubMed]
- Heneghan, H.; Miller, N.; McAnena, O.J.; O’Brien, T.; Kerin, M.J. Differential miRNA Expression in Omental Adipose Tissue and in the Circulation of Obese Patients Identifies Novel Metabolic Biomarkers. J. Clin. Endocrinol. Metab. 2011, 96, E846–E850. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.K.; Lee, M.-J.; Abdelmohsen, K.; Kim, W.; Kim, M.M.; Srikantan, S.; Martindale, J.L.; Hutchison, E.R.; Kim, H.H.; Marasa, B.S.; et al. miR-130 Suppresses Adipogenesis by Inhibiting Peroxisome Proliferator-Activated Receptor Expression. Mol. Cell. Biol. 2010, 31, 626–638. [Google Scholar] [CrossRef] [PubMed]
- Prats-Puig, A.; Ortega, F.J.; Mercader, J.M.; Moreno-Navarrete, J.M.; Moreno, M.; Bonet, N.; Ricart, W.; López-Bermejo, A.; Fernández-Real, J.M. Changes in Circulating MicroRNAs Are Associated With Childhood Obesity. J. Clin. Endocrinol. Metab. 2013, 98, E1655–E1660. [Google Scholar] [CrossRef]
- Cui, X.; You, L.; Zhu, L.; Wang, X.; Zhou, Y.; Li, Y.; Wen, J.; Xia, Y.; Wang, X.; Ji, C.; et al. Change in circulating microRNA profile of obese children indicates future risk of adult diabetes. Metabolism 2018, 78, 95–105. [Google Scholar] [CrossRef]
- Poy, M.N.; Eliasson, L.; Krutzfeldt, J.; Kuwajima, S.; Ma, X.; MacDonald, P.; Pfeffer, S.; Tuschl, T.; Rajewsky, N.; Rorsman, P.; et al. A pancreatic islet-specific microRNA regulates insulin secretion. Nat. Cell Biol. 2004, 432, 226–230. [Google Scholar] [CrossRef]
- Nesca, V.; Guay, C.; Jacovetti, C.; Menoud, V.; Peyot, M.-L.; Laybutt, D.R.; Prentki, M.; Regazzi, R. Identification of particular groups of microRNAs that positively or negatively impact on beta cell function in obese models of type 2 diabetes. Diabetologia 2013, 56, 2203–2212. [Google Scholar] [CrossRef]
- Tattikota, S.G.; Rathjen, T.; McAnulty, S.J.; Wessels, H.-H.; Akerman, I.; van de Bunt, M.; Hausser, J.; Esguerra, J.; Musahl, A.; Pandey, A.; et al. Argonaute2 Mediates Compensatory Expansion of the Pancreatic β Cell. Cell Metab. 2014, 19, 122–134. [Google Scholar] [CrossRef]
- Zhu, Y.; Sun, Y.; Zhou, Y.; Zhang, Y.; Zhang, T.; Li, Y.; You, W.; Chang, X.; Yuan, L.; Han, X. MicroRNA-24 promotes pancreatic beta cells toward dedifferentiation to avoid endoplasmic reticulum stress-induced apoptosis. J. Mol. Cell Biol. 2019, 11, 747–760. [Google Scholar] [CrossRef]
- Xu, G.; Chen, J.; Jing, G.; Shalev, A. Thioredoxin-interacting protein regulates insulin transcription through microRNA-204. Nat. Med. 2013, 19, 1141–1146. [Google Scholar] [CrossRef]
- Jo, S.; Chen, J.; Xu, G.; Grayson, T.B.; Thielen, L.A.; Shalev, A. miR-204 Controls Glucagon-Like Peptide 1 Receptor Expression and Agonist Function. Diabetes 2018, 67, 256–264. [Google Scholar] [CrossRef]
- Laudadio, I.; Manfroid, I.; Achouri, Y.; Schmidt, D.; Wilson, M.; Cordi, S.; Thorrez, L.; Knoops, L.; Jacquemin, P.; Schuit, F.; et al. A Feedback Loop between the Liver-Enriched Transcription Factor Network and Mir-122 Controls Hepatocyte Differentiation. Gastroenterology 2012, 142, 119–129. [Google Scholar] [CrossRef]
- Chai, C.; Rivkin, M.; Berkovits, L.; Simerzin, A.; Zorde-Khvalevsky, E.; Rosenberg, N.; Klein, S.; Yaish, D.; Durst, R.; Shpitzen, S.; et al. Metabolic Circuit Involving Free Fatty Acids, microRNA 122, and Triglyceride Synthesis in Liver and Muscle Tissues. Gastroenterology 2017, 153, 1404–1415. [Google Scholar] [CrossRef]
- Mattis, A.N.; Song, G.; Hitchner, K.; Kim, R.; Lee, A.Y.; Sharma, A.D.; Malato, Y.; McManus, M.; Esau, C.C.; Koller, E.; et al. A screen in mice uncovers repression of lipoprotein lipase by microRNA-29a as a mechanism for lipid distribution away from the liver. Hepatology 2015, 61, 141–152. [Google Scholar] [CrossRef]
- Sun, C.; Huang, F.; Liu, X.; Xiao, X.; Yang, M.; Chuanzheng, S.; Liu, H.; Liao, L. miR-21 regulates triglyceride and cholesterol metabolism in non-alcoholic fatty liver disease by targeting HMGCR. Int. J. Mol. Med. 2015, 35, 847–853. [Google Scholar] [CrossRef]
- Wu, H.; Ng, R.; Chen, X.; Steer, C.J.; Song, G. MicroRNA-21 is a potential link between non-alcoholic fatty liver disease and hepatocellular carcinoma via modulation of the HBP1-p53-Srebp1c pathway. Gut 2016, 65, 1850–1860. [Google Scholar] [CrossRef]
- Ahn, J.; Lee, H.; Jung, C.H.; Ha, T. Lycopene inhibits hepatic steatosis via microRNA-21-induced downregulation of fatty acid-binding protein 7 in mice fed a high-fat diet. Mol. Nutr. Food Res. 2012, 56, 1665–1674. [Google Scholar] [CrossRef]
- Meng, F.; Henson, R.; Wehbe–Janek, H.; Ghoshal, K.; Jacob, S.T.; Patel, T. MicroRNA-21 Regulates Expression of the PTEN Tumor Suppressor Gene in Human Hepatocellular Cancer. Gastroenterology 2007, 133, 647–658. [Google Scholar] [CrossRef]
- Vega-Badillo, J.; Gutiérrez-Vidal, R.; Hernández-Pérez, H.A.; Villamil-Ramírez, H.; León, P.; Sánchez-Muñoz, F.; Morán-Ramos, S.; Larrieta-Carrasco, E.; Fernández-Silva, I.; Méndez-Sánchez, N.; et al. Hepatic miR-33a/miR-144 and their target gene ABCA1 are associated with steatohepatitis in morbidly obese subjects. Liver Int. 2016, 36, 1383–1391. [Google Scholar] [CrossRef]
- Rayner, K.; Esau, C.C.; Hussain, F.N.; McDaniel, A.L.; Marshall, S.M.; van Gils, J.; Ray, T.D.; Sheedy, F.; Goedeke, L.; Liu, X.; et al. Inhibition of miR-33a/b in non-human primates raises plasma HDL and lowers VLDL triglycerides. Nat. Cell Biol. 2011, 478, 404–407. [Google Scholar] [CrossRef]
- Davalos, A.; Goedeke, L.; Smibert, P.; Ramírez, C.; Warrier, N.P.; Andreo, U.; Salinas, D.C.; Rayner, K.; Suresh, U.; Pastor-Pareja, J.C.; et al. miR-33a/b contribute to the regulation of fatty acid metabolism and insulin signaling. Proc. Natl. Acad. Sci. USA 2011, 108, 9232–9237. [Google Scholar] [CrossRef] [PubMed]
- Ramírez, C.; Goedeke, L.; Rotllan, N.; Yoon, J.-H.; Salinas, D.C.; Mattison, J.A.; Suárez, Y.; de Cabo, R.; Gorospe, M.; Fernández-Hernando, C. MicroRNA 33 Regulates Glucose Metabolism. Mol. Cell. Biol. 2013, 33, 2891–2902. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.-J.; Ou-Yang, P.-H.; Han, X.-P. Profibrotic effect of miR-33a with Akt activation in hepatic stellate cells. Cell. Signal. 2014, 26, 141–148. [Google Scholar] [CrossRef]
- Ding, J.; Li, M.; Wan, X.; Jin, X.; Chen, S.; Yu, C.; Li, Y. Effect of miR-34a in regulating steatosis by targeting PPARα expression in nonalcoholic fatty liver disease. Sci. Rep. 2015, 5, 13729. [Google Scholar] [CrossRef] [PubMed]
- Keller, P.; Gburcik, V.; Petrovic, N.; Gallagher, I.J.; Nedergaard, J.; Cannon, B.; Timmons, J.A. Gene-chip studies of adipogenesis-regulated microRNAs in mouse primary adipocytes and human obesity. BMC Endocr. Disord. 2011, 11, 7. [Google Scholar] [CrossRef] [PubMed]
- Ortega, F.J.; Mercader, J.M.; Catalan, V.; Moreno-Navarrete, J.M.; Pueyo, N.; Sabater, M.; Gomez-Ambrosi, J.; Anglada, R.; Formoso, J.A.F.; Ricart, W.; et al. Targeting the Circulating MicroRNA Signature of Obesity. Clin. Chem. 2013, 59, 781–792. [Google Scholar] [CrossRef]
- Pescador, N.; Pérez-Barba, M.; Ibarra, J.M.; Corbatón, A.; Martínez-Larrad, M.T.; Serrano-Ríos, M. Serum Circulating microRNA Profiling for Identification of Potential Type 2 Diabetes and Obesity Biomarkers. PLoS ONE 2013, 8, e77251. [Google Scholar] [CrossRef]
- Wang, R.; Hong, J.; Cao, Y.; Shi, J.; Gu, W.; Ning, G.; Zhang, Y.; Wang, W. Elevated circulating microRNA-122 is associated with obesity and insulin resistance in young adults. Eur. J. Endocrinol. 2015, 172, 291–300. [Google Scholar] [CrossRef]
- Iacomino, G.; Russo, P.; Stillitano, I.; Lauria, F.; Marena, P.; Ahrens, W.; De Luca, P.; Siani, A. Circulating microRNAs are deregulated in overweight/obese children: Preliminary results of the I.Family study. Genes Nutr. 2016, 11, 7. [Google Scholar] [CrossRef]
- Brovkina, O.; Nikitin, A.; Khodyrev, D.; Shestakova, E.; Sklyanik, I.; Panevina, A.; Stafeev, I.; Menshikov, M.; Kobelyatskaya, A.; Yurasov, A.; et al. Role of MicroRNAs in the Regulation of Subcutaneous White Adipose Tissue in Individuals With Obesity and Without Type 2 Diabetes. Front. Endocrinol. 2019, 10, 840. [Google Scholar] [CrossRef]
- Palmer, J.; Soule, B.P.; Simone, B.A.; Zaorsky, N.; Jin, L.; Simone, N. MicroRNA expression altered by diet: Can food be medicinal? Ageing Res. Rev. 2014, 17, 16–24. [Google Scholar] [CrossRef]
- Kristensen, M.M.; Davidsen, P.K.; Vigelsø, A.; Hansen, C.N.; Jensen, L.J.; Jessen, N.; Bruun, J.M.; Dela, F.; Helge, J.W. miRNAs in human subcutaneous adipose tissue: Effects of weight loss induced by hypocaloric diet and exercise. Obesity 2017, 25, 572–580. [Google Scholar] [CrossRef]
- Manning, P.; Munasinghe, P.E.; Papannarao, J.B.; Gray, A.R.; Sutherland, W.; Katare, R. Acute Weight Loss Restores Dysregulated Circulating MicroRNAs in Individuals Who Are Obese. J. Clin. Endocrinol. Metab. 2019, 104, 1239–1248. [Google Scholar] [CrossRef]
- Grieco, G.; Brusco, N.; Licata, G.; Fignani, D.; Formichi, C.; Nigi, L.; Sebastiani, G.; Dotta, F. The Landscape of microRNAs in βCell: Between Phenotype Maintenance and Protection. Int. J. Mol. Sci. 2021, 22, 803. [Google Scholar] [CrossRef]
- Jiménez-Lucena, R.; Camargo, A.; Alcala-Diaz, J.F.; Romero-Baldonado, C.; Luque, R.M.; Van Ommen, B.; Delgado-Lista, J.; Ordovás, J.M.; Pérez-Martínez, P.; Rangel-Zuñiga, O.A.; et al. A plasma circulating miRNAs profile predicts type 2 diabetes mellitus and prediabetes: From the CORDIOPREV study. Exp. Mol. Med. 2018, 50, 1–12. [Google Scholar] [CrossRef]
- Sebastiani, G.; Grieco, G.E.; Brusco, N.; Ventriglia, G.; Formichi, C.; Marselli, L.; Marchetti, P.; Dotta, F. MicroRNA Expression Analysis of In Vitro Dedifferentiated Human Pancreatic Islet Cells Reveals the Activation of the Pluripotency-Related MicroRNA Cluster miR-302s. Int. J. Mol. Sci. 2018, 19, 1170. [Google Scholar] [CrossRef]
- Belgardt, B.-F.; Ahmed, K.; Spranger, M.; Latreille, M.; Denzler, R.; Kondratiuk, N.; Von Meyenn, F.; Villena, F.N.; Herrmanns, K.; Bosco, D.; et al. The microRNA-200 family regulates pancreatic beta cell survival in type 2 diabetes. Nat. Med. 2015, 21, 619–627. [Google Scholar] [CrossRef]
- Grieco, G.E.; Brusco, N.; Nigi, L.; Formichi, C.; Fignani, D.; Licata, G.; Marselli, L.; Marchetti, P.; Salvini, L.; Tinti, L.; et al. Reduced MiR-184-3p Expression Occurring in Type 2 Diabetic Pancreatic Islets Protects β-Cells from Lipotoxic and Proinflammatory Apoptosis via a CRTC1-Dependent Mechanism. Cell Biol. 2021. [Google Scholar] [CrossRef]
- Ciccacci, C.; Di Fusco, D.; Cacciotti, L.; Morganti, R.; D’Amato, C.; Greco, C.; Rufini, S.; Novelli, G.; Sangiuolo, F.; Spallone, V.; et al. MicroRNA genetic variations: Association with type 2 diabetes. Acta Diabetol. 2013, 50, 867–872. [Google Scholar] [CrossRef]
- Gjorgjieva, M.; Sobolewski, C.; Dolicka, D.; De Sousa, M.C.; Foti, M. miRNAs and NAFLD: From pathophysiology to therapy. Gut 2019, 68, 2065–2079. [Google Scholar] [CrossRef]
- Vienberg, S.; Geiger, J.; Madsen, S.; Dalgaard, L.T. MicroRNAs in metabolism. Acta Physiol. 2016, 219, 346–361. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.-Y.; Yang, Y.-L.; Wang, P.-W.; Wang, F.-S.; Huang, Y.-H. The Emerging Role of MicroRNAs in NAFLD: Highlight of MicroRNA-29a in Modulating Oxidative Stress, Inflammation, and Beyond. Cells 2020, 9, 1041. [Google Scholar] [CrossRef] [PubMed]
- Pirola, C.J.; Scian, R.; Gianotti, T.F.; Dopazo, H.; Rohr, C.; Martino, J.S.; Castaño, G.O.; Sookoian, S. Epigenetic Modifications in the Biology of Nonalcoholic Fatty Liver Disease: The Role of DNA Hydroxymethylation and TET Proteins. Medicine 2015, 94, e1480. [Google Scholar] [CrossRef] [PubMed]
- Lakner, A.M.; Bonkovsky, H.L.; Schrum, L.W. microRNAs: Fad or future of liver disease. World J. Gastroenterol. 2011, 17, 2536–2542. [Google Scholar] [CrossRef]
- Cheung, O.; Puri, P.; Eicken, C.; Contos, M.J.; Mirshahi, F.; Maher, J.W.; Kellum, J.M.; Min, H.; Luketic, V.A.; Sanyal, A.J. Nonalcoholic steatohepatitis is associated with altered hepatic MicroRNA expression. Hepatology 2008, 48, 1810–1820. [Google Scholar] [CrossRef]
- Liu, C.-H.; Ampuero, J.; Gil-Gomez, A.; Montero-Vallejo, R.; Rojas, Á.; Muñoz-Hernández, R.; Gallego-Durán, R.; Romero-Gómez, M. miRNAs in patients with non-alcoholic fatty liver disease: A systematic review and meta-analysis. J. Hepatology 2018, 69, 1335–1348. [Google Scholar] [CrossRef]
- Hsu, S.-H.; Wang, B.; Kota, J.; Yu, J.; Costinean, S.; Kutay, H.; Yu, L.; Bai, S.; La Perle, K.; Chivukula, R.R.; et al. Essential metabolic, anti-inflammatory, and anti-tumorigenic functions of miR-122 in liver. J. Clin. Investig. 2012, 122, 2871–2883. [Google Scholar] [CrossRef]
- Tsai, W.-C.; Hsu, S.-D.; Hsu, C.-S.; Lai, T.-C.; Chen, S.-J.; Shen, R.; Huang, Y.; Chen, H.-C.; Lee, C.-H.; Tsai, T.-F.; et al. MicroRNA-122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J. Clin. Investig. 2012, 122, 2884–2897. [Google Scholar] [CrossRef]
- Xing, T.; Jiang, D.; Huang, J.; Xu, Z. Expression and clinical significance of miR-122 and miR-29 in hepatitis B virus-related liver disease. Genet. Mol. Res. 2014, 13, 7912–7918. [Google Scholar] [CrossRef]
- Jampoka, K.; Muangpaisarn, P.; Khongnomnan, K.; Treeprasertsuk, S.; Tangkijvanich, P.; Payungporn, S. Serum miR-29a and miR-122 as Potential Biomarkers for Non-Alcoholic Fatty Liver Disease (NAFLD). MicroRNA 2018, 7, 215–222. [Google Scholar] [CrossRef]
- Becker, P.P.; Rau, M.; Schmitt, J.; Malsch, C.; Hammer, C.; Bantel, H.; Mullhaupt, B.; Geier, A. Performance of Serum microRNAs -122, -192 and -21 as Biomarkers in Patients with Non-Alcoholic Steatohepatitis. PLoS ONE 2015, 10, e0142661. [Google Scholar] [CrossRef]
- Calo, N.; Ramadori, P.; Sobolewski, C.; Romero, Y.; Maeder, C.; Fournier, M.; Rantakari, P.; Zhang, F.-P.; Poutanen, M.; Dufour, J.-F.; et al. Stress-activatedmiR-21/miR-21*in hepatocytes promotes lipid and glucose metabolic disorders associated with high-fat diet consumption. Gut 2016, 65, 1871–1881. [Google Scholar] [CrossRef]
- Auguet, T.; Aragonès, G.; Berlanga, A.; Guiu-Jurado, E.; Martí, A.; Martínez, S.; Sabench, F.; Hernández, M.; Aguilar, C.; Sirvent, J.J.; et al. miR33a/miR33b* and miR122 as Possible Contributors to Hepatic Lipid Metabolism in Obese Women with Nonalcoholic Fatty Liver Disease. Int. J. Mol. Sci. 2016, 17, 1620. [Google Scholar] [CrossRef]
- Salvoza, N.C.; Klinzing, D.C.; Gopez-Cervantes, J.; Baclig, M.O. Association of Circulating Serum miR-34a and miR-122 with Dyslipidemia among Patients with Non-Alcoholic Fatty Liver Disease. PLoS ONE 2016, 11, e0153497. [Google Scholar] [CrossRef]
- Tryndyak, V.; Latendresse, J.R.; Montgomery, B.; Ross, S.A.; Beland, F.A.; Rusyn, I.; Pogribny, I.P. Plasma microRNAs are sensitive indicators of inter-strain differences in the severity of liver injury induced in mice by a choline- and folate-deficient diet. Toxicol. Appl. Pharmacol. 2012, 262, 52–59. [Google Scholar] [CrossRef]
- Cermelli, S.; Ruggieri, A.; Marrero, J.A.; Ioannou, G.N.; Beretta, L. Circulating MicroRNAs in Patients with Chronic Hepatitis C and Non-Alcoholic Fatty Liver Disease. PLoS ONE 2011, 6, e23937. [Google Scholar] [CrossRef]
- Liu, X.-L.; Pan, Q.; Zhang, R.-N.; Shen, F.; Yan, S.-Y.; Sun, C.; Xu, Z.-J.; Chen, Y.-W.; Fan, J.-G. Disease-specific miR-34a as diagnostic marker of non-alcoholic steatohepatitis in a Chinese population. World J. Gastroenterol. 2016, 22, 9844–9852. [Google Scholar] [CrossRef]
- Li, A.; Huang, W.; Zhang, X.; Xie, L.; Miao, X. Identification and Characterization of CircRNAs of Two Pig Breeds as a New Biomarker in Metabolism-Related Diseases. Cell. Physiol. Biochem. 2018, 47, 2458–2470. [Google Scholar] [CrossRef]
- Wang, T.; Li, M.; Guan, J.; Li, P.; Wang, H.; Guo, Y.; Shuai, S.; Li, X. MicroRNAs miR-27a and miR-143 Regulate Porcine Adipocyte Lipid Metabolism. Int. J. Mol. Sci. 2011, 12, 7950–7959. [Google Scholar] [CrossRef]
- Kim, S.Y.; Kim, A.Y.; Lee, H.W.; Son, Y.H.; Lee, G.Y.; Lee, J.-W.; Lee, Y.S.; Kim, J.B. miR-27a is a negative regulator of adipocyte differentiation via suppressing PPARγ expression. Biochem. Biophys. Res. Commun. 2010, 392, 323–328. [Google Scholar] [CrossRef]
- Guo, X.-Y.; Chen, J.-N.; Sun, F.; Wang, Y.-Q.; Pan, Q.; Fan, J.-G. circRNA_0046367 Prevents Hepatoxicity of Lipid Peroxidation: An Inhibitory Role against Hepatic Steatosis. Oxidative Med. Cell. Longev. 2017, 2017, 3960197. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; Jiang, Z.; Yang, X.; Lin, J.; Cai, Q.; Li, X. Circular RNA HIPK3 contributes to hyperglycemia and insulin homeostasis by sponging miR-192-5p and upregulating transcription factor forkhead box O1. Endocr. J. 2020, 67, 397–408. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Wang, Q.; Li, Z.; Yang, Q.; Liu, Y.; Du, Z.; Zhang, G.; Song, Y. Circular RNA CDR1as promotes adipogenic and suppresses osteogenic differentiation of BMSCs in steroid-induced osteonecrosis of the femoral head. Bone 2020, 133, 115258. [Google Scholar] [CrossRef] [PubMed]
- Arcinas, C.; Tan, W.; Fang, W.; Desai, T.P.; Teh, D.C.S.; Degirmenci, U.; Xu, D.; Foo, R.; Sun, L. Adipose circular RNAs exhibit dynamic regulation in obesity and functional role in adipogenesis. Nat. Metab. 2019, 1, 688–703. [Google Scholar] [CrossRef]
- Stoll, L.; Rodríguez-Trejo, A.; Guay, C.; Brozzi, F.; Bayazit, M.B.; Gattesco, S.; Menoud, V.; Sobel, J.; Marques, A.C.; Venø, M.T.; et al. A circular RNA generated from an intron of the insulin gene controls insulin secretion. Nat. Commun. 2020, 11, 5611. [Google Scholar] [CrossRef]
- Zhao, Z.; Li, X.; Jian, D.; Hao, P.; Rao, L.; Li, M. Hsa_circ_0054633 in peripheral blood can be used as a diagnostic biomarker of pre-diabetes and type 2 diabetes mellitus. Acta Diabetol. 2017, 54, 237–245. [Google Scholar] [CrossRef]
- Fang, Y.; Wang, X.; Li, W.; Han, J.; Jin, J.; Su, F.; Zhang, J.; Huang, W.; Xiao, F.; Pan, Q.; et al. Screening of circular RNAs and validation of circANKRD36 associated with inflammation in patients with type 2 diabetes mellitus. Int. J. Mol. Med. 2018, 42, 1865–1874. [Google Scholar] [CrossRef]
- Guo, X.-Y.; He, C.-X.; Wang, Y.-Q.; Sun, C.; Li, G.-M.; Su, Q.; Pan, Q.; Fan, J.-G. Circular RNA Profiling and Bioinformatic Modeling Identify Its Regulatory Role in Hepatic Steatosis. BioMed Res. Int. 2017, 2017, 5936171. [Google Scholar] [CrossRef]
- Guo, X.-Y.; Sun, F.; Chen, J.-N.; Wang, Y.-Q.; Pan, Q.; Fan, J.-G. circRNA_0046366 inhibits hepatocellular steatosis by normalization of PPAR signaling. World J. Gastroenterol. 2018, 24, 323–337. [Google Scholar] [CrossRef]
- Zaiou, M. circRNAs Signature as Potential Diagnostic and Prognostic Biomarker for Diabetes Mellitus and Related Cardiovascular Complications. Cells 2020, 9, 659. [Google Scholar] [CrossRef]
- Zhu, H.; Leung, S.W. Identification of microRNA biomarkers in type 2 diabetes: A meta-analysis of controlled profiling studies. Diabetologia 2015, 58, 900–911. [Google Scholar] [CrossRef]
- Grieco, G.E.; Sebastiani, G.; Fignani, D.; Brusco, N.; Nigi, L.; Formichi, C.; Licata, G.; Bruttini, M.; D’Aurizio, R.; Mathieu, C.; et al. Protocol to analyze circulating small non-coding RNAs by high-throughput RNA sequencing from human plasma samples. STAR Protoc. 2021, 2, 100606. [Google Scholar] [CrossRef]
- Oses, M.; Sanchez, J.M.; Portillo, M.P.; Aguilera, C.M.; Labayen, I. Circulating miRNAs as Biomarkers of Obesity and Obesity-Associated Comorbidities in Children and Adolescents: A Systematic Review. Nutrients 2019, 11, 2890. [Google Scholar] [CrossRef]
- Zampetaki, A.; Kiechl, S.; Drozdov, I.; Willeit, P.; Mayr, U.; Prokopi, M.; Mayr, A.; Weger, S.; Oberhollenzer, F.; Bonora, E.; et al. Plasma MicroRNA Profiling Reveals Loss of Endothelial MiR-126 and Other MicroRNAs in Type 2 Diabetes. Circ. Res. 2010, 107, 810–817. [Google Scholar] [CrossRef]
- Seyhan, A.; Lopez, Y.N.; Xie, H.; Yi, F.; Mathews, C.; Pasarica, M.; Pratley, R.E. Pancreas-enriched miRNAs are altered in the circulation of subjects with diabetes: A pilot cross-sectional study. Sci. Rep. 2016, 6, 31479. [Google Scholar] [CrossRef]
- Pillai, S.S.; Lakhani, H.V.; Zehra, M.; Wang, J.; Dilip, A.; Puri, N.; O’Hanlon, K.; Sodhi, K. Predicting Nonalcoholic Fatty Liver Disease through a Panel of Plasma Biomarkers and MicroRNAs in Female West Virginia Population. Int. J. Mol. Sci. 2020, 21, 6698. [Google Scholar] [CrossRef]
- Tan, Y.; Ge, G.; Pan, T.; Wen, D.; Gan, J. A Pilot Study of Serum MicroRNAs Panel as Potential Biomarkers for Diagnosis of Nonalcoholic Fatty Liver Disease. PLoS ONE 2014, 9, e105192. [Google Scholar] [CrossRef]
- Piazzolla, V.A.; Mangia, A. Noninvasive Diagnosis of NAFLD and NASH. Cells 2020, 9, 1005. [Google Scholar] [CrossRef]
- Willeit, P.; Skroblin, P.; Kiechl, S.; Fernández-Hernando, C.; Mayr, M. Liver microRNAs: Potential mediators and biomarkers for metabolic and cardiovascular disease? Eur. Hear. J. 2016, 37, 3260–3266. [Google Scholar] [CrossRef]
- Leti, F.; Distefano, J.K. Long Noncoding RNAs as Diagnostic and Therapeutic Targets in Type 2 Diabetes and Related Complications. Genes 2017, 8, 207. [Google Scholar] [CrossRef]
- El-Hefnway, S.; Al-Sheikh, N.; Kasem, H.; El-Esrigy, F. Plasma Circular RNA (0054633) Expression as a Biomarker for Prediabetes and Type 2 diabetes mellitus. Bull. Egypt. Soc. Physiol. Sci. 2018, 38, 77–88. [Google Scholar] [CrossRef]
- Wang, T.; Pan, W.; Hu, J.; Zhang, Z.; Li, G.; Liang, Y. Circular RNAs in Metabolic Diseases. Adv. Exp. Med. Biol. 2018, 1087, 275–285. [Google Scholar] [CrossRef]
- Grieco, G.E.; Brusco, N.; Licata, G.; Nigi, L.; Formichi, C.; Dotta, F.; Sebastiani, G. Targeting microRNAs as a Therapeutic Strategy to Reduce Oxidative Stress in Diabetes. Int. J. Mol. Sci. 2019, 20, 6358. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Trautwein, C.; Luedde, T.; Roderburg, C. A General Overview on Non-coding RNA-Based Diagnostic and Therapeutic Approaches for Liver Diseases. Front. Pharmacol. 2018, 9, 805. [Google Scholar] [CrossRef] [PubMed]
- Zaiou, M.; El Amri, H.; Bakillah, A. The clinical potential of adipogenesis and obesity-related microRNAs. Nutr. Metab. Cardiovasc. Dis. 2018, 28, 91–111. [Google Scholar] [CrossRef] [PubMed]
| Name | Functions | Targets |
|---|---|---|
| SRA | co-activator of PPARγ2 in adipogenesis [102]; promotes steatosis, ↓FFAs oxidation [102] | PPARγ2 [102]; FoxO1 [102] |
| ASMER1, ASMER2 | adipogenesis, lipolysis, adiponectin secretion [103] | - |
| ADNCR | adipocyte differentiation inhibition [104] | miRNA-204/SIRT1 [104] |
| H19 | ↓expression of adipogenic factors [102]; inhibition of BMMSCs adipogenic differentiation [105]; promotes hepatic lipid accumulation [111]; promotes gluconeogenesis [102,112] | HDACs 4,5,6 [102]; miR-675/HDACs [105]; PTBP1/SREBP-1c [111]; FoxO1 [102], HNF4a [112] |
| HOTAIR | promotes pre-adipocyte differentiation, ↑adipogenic genes expression (PPARγ and LPL) [101,106]; activates fibrogenesis and promotes TG accumulation [21] | Genes at HOXD locus [101]; MMP2, MMP9 [21] |
| BlnC1 | ↑UCP1 expression, ↑ mitochondiral genes expression [3]; promotes adipose tissue browning [102] | Ebf2 [3,101,102] |
| MALAT1 | ↑ROS and pro-inflammatory cytokines, promotes angiogenesis [112]; involved in the pathogenesis of diabetic gastroparesis [113]; promotes liver steatosis and fibrosis [21,102,114,115] | SAA3 [112]; α-SMA and SM-MHC [113]; CXCL5 [21,114,115] |
| ANRIL | promotes atherosclerosis, ECs proliferation and adhesion [116]; causes cardiac dysfunction, myocardial apoptosis and fibrosis [117] | INK4B [116] |
| MEG3 | promotes ECs functions, ↑microvascular permeability and inflammation upon silencing [116]; fibrosis progression upon downregulation, possibile interaction with HOTAIR [21] | PI3K/Akt [116] |
| CASC2 | key role in renal tumorigenesis, possible involvement in renal failure [118] | - |
| MIAT | ↑TNFa and VEGF, regulates ECs function [116] | miR-150-5p/VEGF [116] |
| NONRATT021972 | involved in nociception and neuropathic pain ↑TNFa [119] | TNFa [119] |
| lncARSR | ↑hepatic lipid accumulation [120] | PI3K/Akt /SREBP-1c [120] |
| Lnc18q22.2 | associated with fibrosis and lobular inflammation [121,122] | - |
| APTR | involved in collagen accumulation and HSCs activation [21] | - |
| Name | Functions | Targets |
|---|---|---|
| miR-130 | ↓ adipogenesis [135] | PPARγ [135] |
| miR-486 | ↑preadipocyte proliferation and myotube glucose intolerance [137] | preadipocyte [137] |
| miR-146b and miR-15b | ↓glucose-stimulated insulin secretion [137] | - |
| miR-375 | ↓GSIS process [138] | insulin gene transcription [138] |
| miR-7 | ↓expression of genes involved in the process of fusion of insulin granules with plasma membrane and SNARE proteins [97] | genes involved in insulin granules fusion with plasma membrane and SNARE proteins [97] |
| miR-184 | ↑ β-cell proliferation, ↑insulin secretion in response to glucose challenge and protects β-cells against apoptosis induced by chronic exposure to proinflammatory cytokines or FAs [139,140] | β-cells in pancreatic islets [139,140] |
| miR-24 | implicated in de-differentiation and protection against apoptosis of β-cells [141] | β-cells in pancreatic islets [141] |
| miR-204 | ↓insulin production [142]; ↑ improved glucose tolerance and insulin secretion [143] | TXNIP/miR-204/MafA/insulin pathway [142]; GLP1R [143] |
| miR-122 | ↑Hepatocyte proliferation and maturation [144] ↓TG synthesis [145] | HNF6 [144] |
| miR-29a | ↑Uptake of lipids from the circulation in hepatocytes [146] | Lpl [146] |
| miR-21 | ↓TG, free cholesterol, and total cholesterol levels [147] ↑Lipid oxidation [148] | HMGCR and FABP7 [147,149]; PTEN [150]; PPARα [148] |
| miR-33 | Cholesterol and FAs metabolism [151]; ↓Gluconeogenesis [152,153,154,155] | ABCA1 and ABCG1, CPT1A and AMPKα [151]; PCK1, G6PC [152,153,154,155] |
| miR-34a | ↓FAs catabolism [156] | PPARα, SIRT1 [156] |
| Name | Functions | Targets |
|---|---|---|
| circRNA_1897 | ↑Adipocytes differentiation and lipid metabolism [192] | miR-27a and miR-27b-3p [192] |
| circRNA_26852 | Lipolysis and inhibition of adipocyte differentiation via a PPARγ-dependent mechanism [192] | miR-874 and miR-486 [192] |
| circ_0046367 | Activation of genes involved in lipid metabolism such as Cpt2 and Acbd3 [193] | miR-34a/PPARα [193] |
| circHIPK3 | ↑Lipid droplets accumulation following oleate treatment [194] | miR-192-5p/Foxo1 [194] |
| circ_0001946(circCDR1) | ↑Adipogenic differentiation ↓Osteogenic differentiation of BMSCs [195]; ↓Insulin secretion [197] | miR-7-5p/Wnt5 [195]; miR-7/Snca, Cspa, Cplx1 [197] |
| circRNA_0095570 (circH19) | ↓Lipids droplets formation and adipogenesis in ACDCs cells | Pparγ, Srebp1c and Cebp1- α |
| circ_Arhgap5-2 | ↓Adipogenesis markers [196] | Pparγ, Cebpα, Fabp4 and AdipoQ [196] |
| circRNA_0054633 | Biomarker of prediabetes in peripheral blood [198] | - |
| circ_ANKRD36 | Potential role in inflammatory mechanisms occurring in T2D [199] | - |
| circRNA_021412 | ↓long chain acyl-CoA synthetases and development of hepatic steatosis [200] | miR-1972/LPN1 [200] |
| circRNA_0046366 | ↑Hepatocyte steatosis [193] | miR-34a/PPARα [193] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Formichi, C.; Nigi, L.; Grieco, G.E.; Maccora, C.; Fignani, D.; Brusco, N.; Licata, G.; Sebastiani, G.; Dotta, F. Non-Coding RNAs: Novel Players in Insulin Resistance and Related Diseases. Int. J. Mol. Sci. 2021, 22, 7716. https://doi.org/10.3390/ijms22147716
Formichi C, Nigi L, Grieco GE, Maccora C, Fignani D, Brusco N, Licata G, Sebastiani G, Dotta F. Non-Coding RNAs: Novel Players in Insulin Resistance and Related Diseases. International Journal of Molecular Sciences. 2021; 22(14):7716. https://doi.org/10.3390/ijms22147716
Chicago/Turabian StyleFormichi, Caterina, Laura Nigi, Giuseppina Emanuela Grieco, Carla Maccora, Daniela Fignani, Noemi Brusco, Giada Licata, Guido Sebastiani, and Francesco Dotta. 2021. "Non-Coding RNAs: Novel Players in Insulin Resistance and Related Diseases" International Journal of Molecular Sciences 22, no. 14: 7716. https://doi.org/10.3390/ijms22147716
APA StyleFormichi, C., Nigi, L., Grieco, G. E., Maccora, C., Fignani, D., Brusco, N., Licata, G., Sebastiani, G., & Dotta, F. (2021). Non-Coding RNAs: Novel Players in Insulin Resistance and Related Diseases. International Journal of Molecular Sciences, 22(14), 7716. https://doi.org/10.3390/ijms22147716








