FOXE1-Dependent Regulation of Macrophage Chemotaxis by Thyroid Cells In Vitro and In Vivo
Abstract
:1. Introduction
2. Results
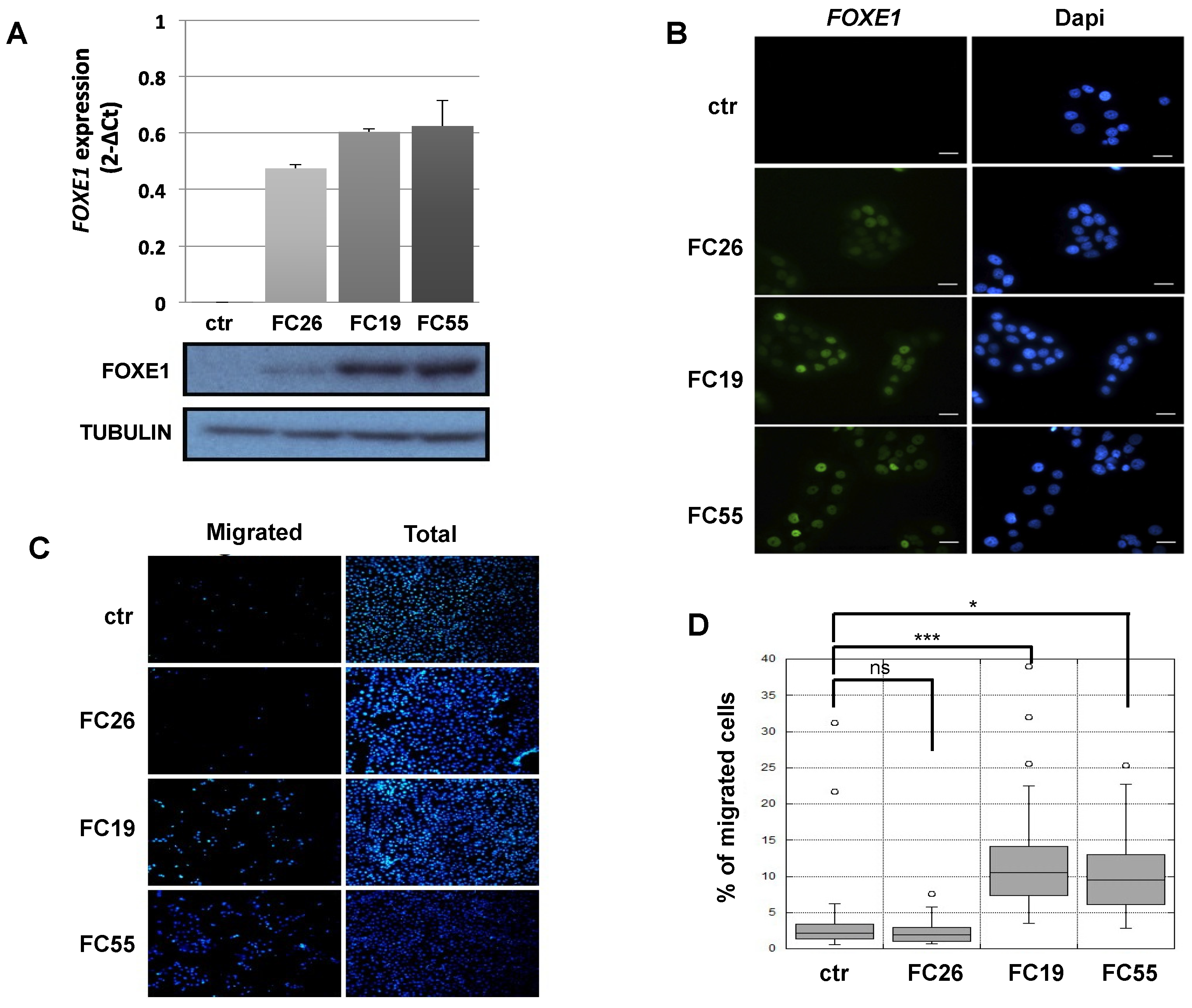
2.1. FOXE1 Expression Induces a Migratory Phenotype in FRT Cells
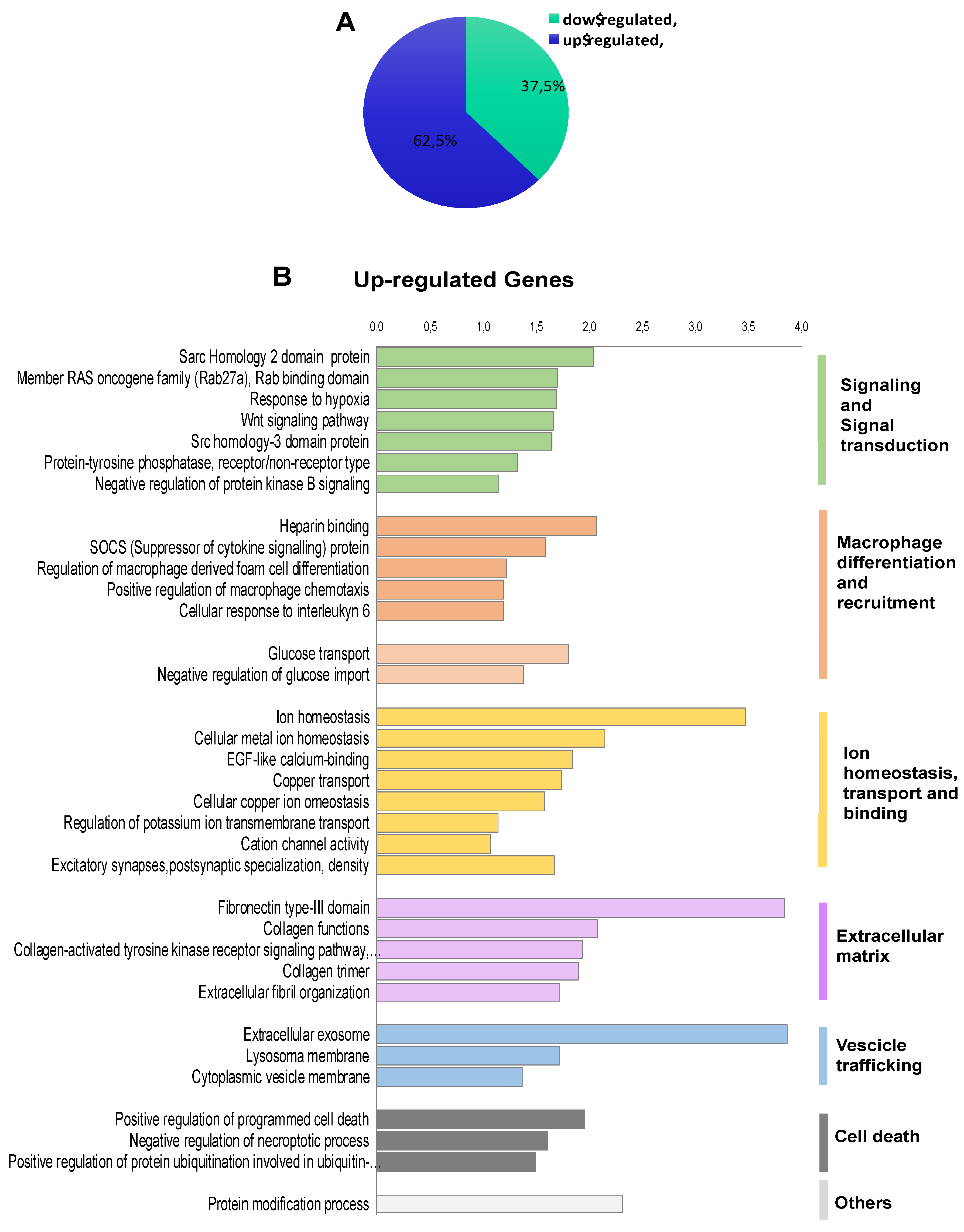
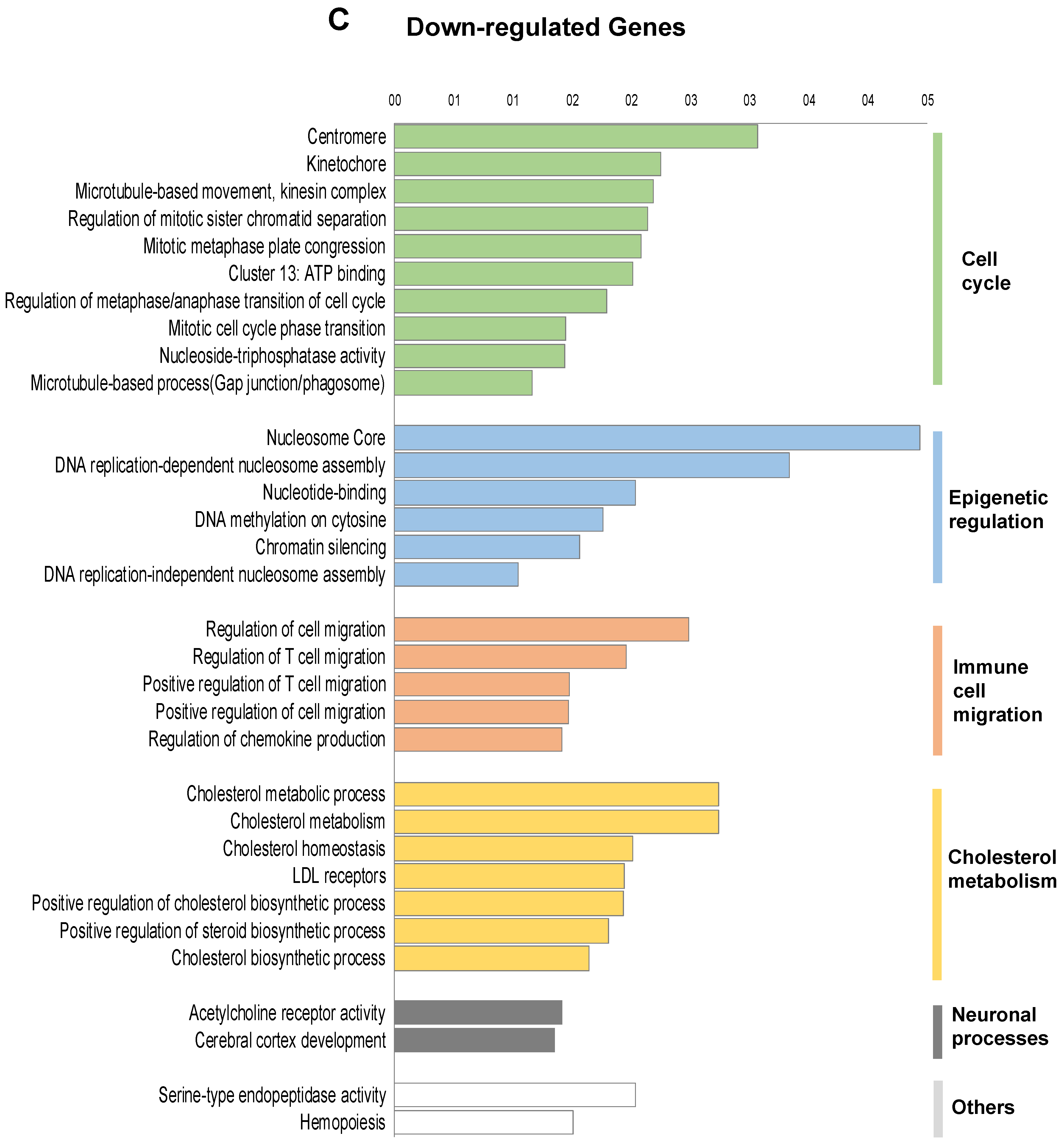
2.2. Transcriptomic Landscape of FOXE1-Expressing FRT Cells
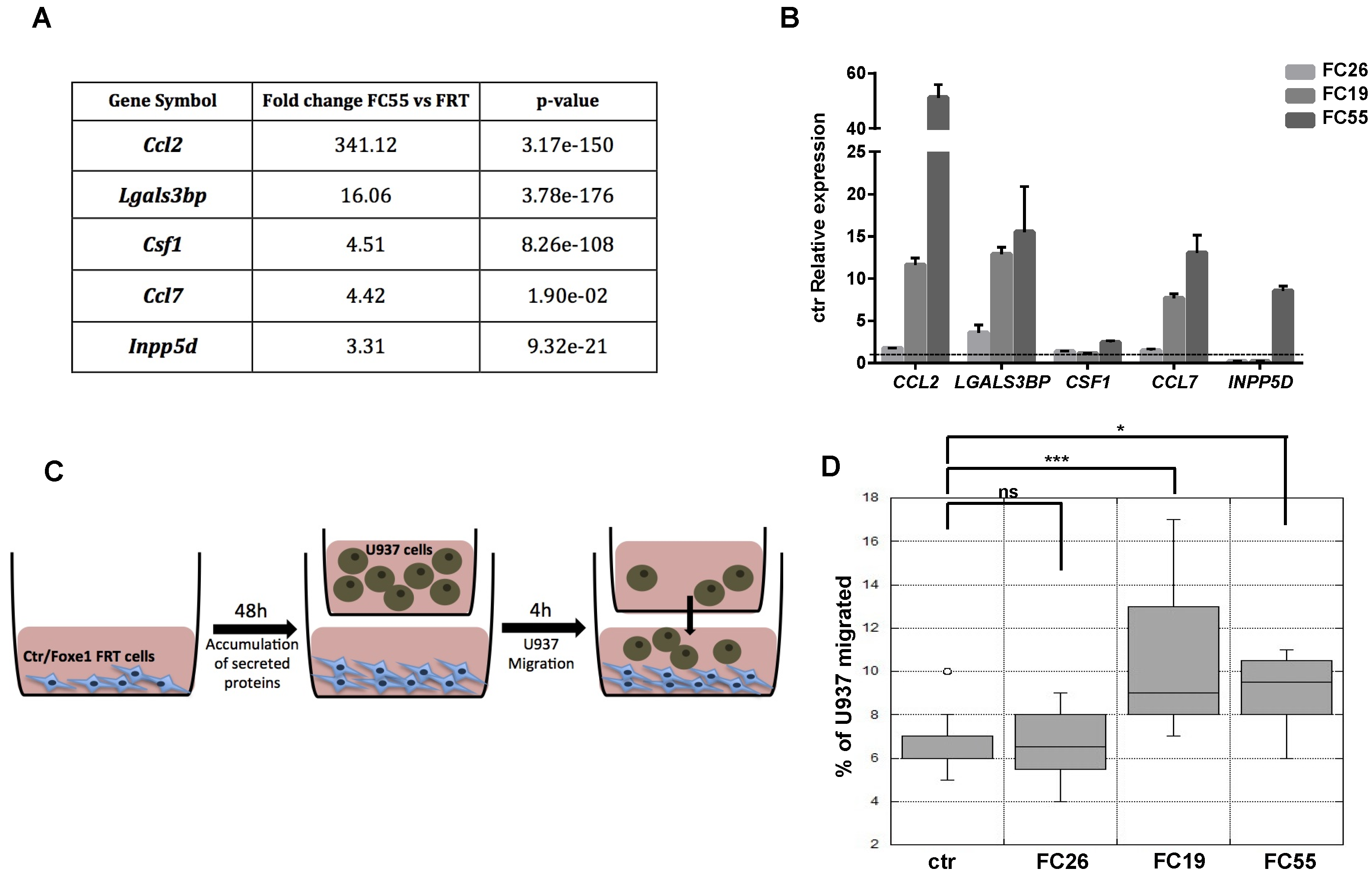
2.3. FOXE1 Enhances Chemoattractant Ability of FRT Cells
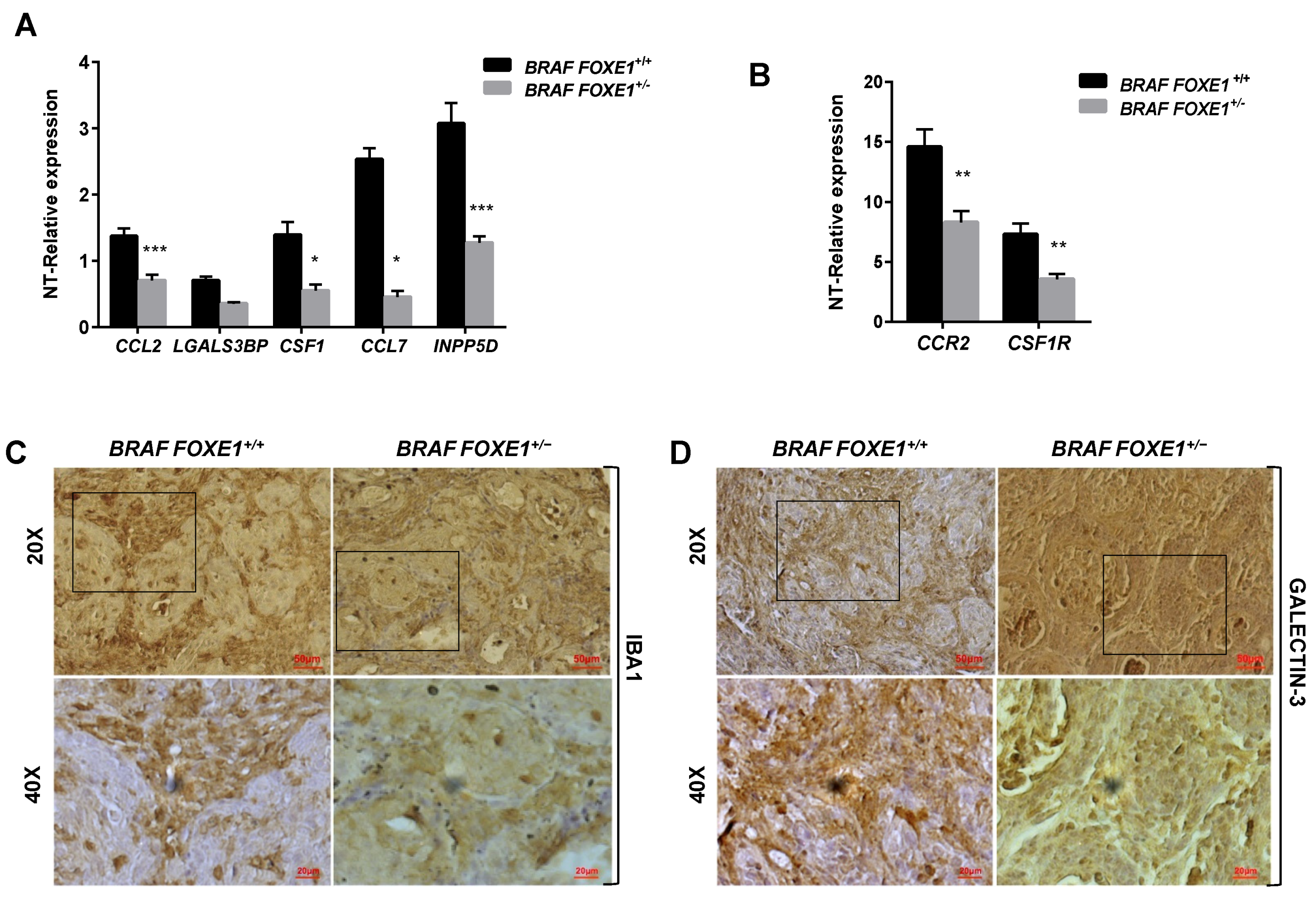
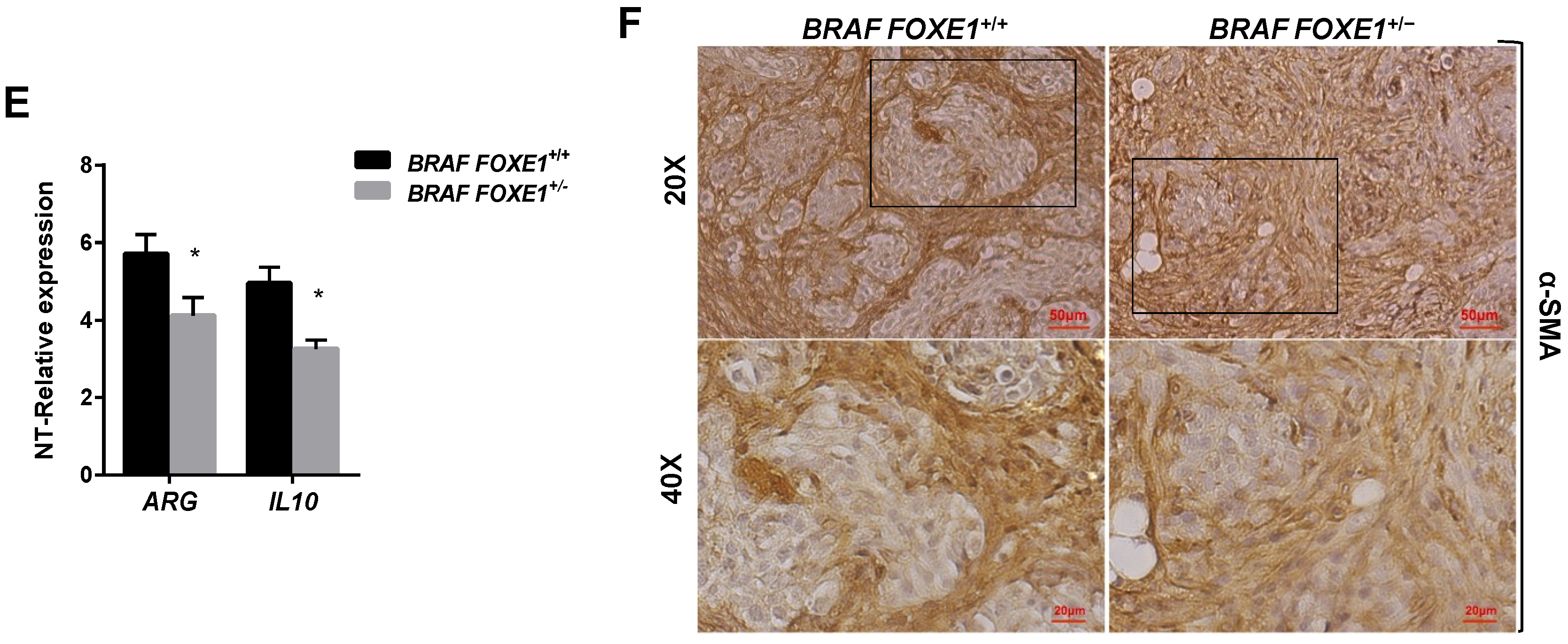
2.4. Chemokine Expression and Immune Cell Recruitment Are FOXE1-Dependent in an In Vivo Thyroid Cancer Model
3. Discussion
4. Materials and Methods
4.1. Plasmid Preparation
4.2. Cell Culture and Transfection
4.3. RNA Sequencing and Bioinformatic Analysis
4.4. Immunoblotting
4.5. Immunofluorescence Assay
4.6. Transwell Assay
4.7. Quantitative Real-Time PCR
4.8. Mice Treatment and Genotyping
4.9. Immunohistochemistry
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Conflicts of Interest
References
- Kitahara, C.M.; Sosa, J.A. The changing incidence of thyroid cancer. Nat. Rev. Endocrinol. 2016, 12, 646–653. [Google Scholar] [CrossRef] [PubMed]
- Douglas, E.H.; Rhoads, A.; Thomas, A.; Aloi, J.; Suhl, J.; Lycan, T., Jr.; Oleson, J.; Conway, M.K.; Klubo-Gwiezdzinska, J.; Lynch, C.F.; et al. Incidence and Survival in Reproductive-Aged Women with Differentiated Thyroid Cancer: United States SEER 18 2000–2016. Thyroid 2020, 30, 1781–1791. [Google Scholar] [CrossRef]
- Rahbari, R.; Zhang, L.; Kebebew, E. Thyroid cancer gender disparity. Future Oncol. 2010, 6. [Google Scholar] [CrossRef] [PubMed]
- Handkiewicz-Junak, D.; Czarniecka, A.; Jarzab, B. Molecular prognostic markers in papillary and follicular thyroid cancer. Current status and future directions. Mol. Cell. Endocrinol. 2010, 322, 8–28. [Google Scholar] [CrossRef]
- Kim, S.; Cho, S.W.; Min, H.S.; Kim, K.M.; Yeom, G.Y.; Kim, E.Y.; Lee, K.E.; Yun, Y.G.; Park, D.J.; Park, Y.J. The Expression of Tumor-Associated Macrophages in Papillary Thyroid Carcinoma. Endocrinol. Metab. 2013, 28, 192–198. [Google Scholar] [CrossRef]
- Qing, W.; Fang, W.Y.; Ye, L.; Shen, L.Y.; Zhang, X.F.; Fei, X.C.; Chen, X.; Wang, W.Q.; Li, X.Y.; Xiao, J.C.; et al. Density of tumor-associated macrophages correlates with lymph node metastasis in papillary thyroid carcinoma. Thyroid 2012, 9, 905–910. [Google Scholar] [CrossRef]
- Jung, K.Y.; Cho, S.W.; Kim, Y.A.; Kim, D.; Oh, B.C.; Park, D.J.; Park, Y.J. Cancers with Higher Density of Tumor-Associated Macrophages Were Associated with Poor Survival Rates. J. Pathol. Transl. Med. 2015, 49, 318–324. [Google Scholar] [CrossRef]
- Ryder, M.; Ghossein, R.A.; Ricarte-Filho, J.C.; Knauf, J.A.; Fagin, J.A. Increased density of tumor-associated macrophages is associated with decreased survival in advanced thyroid cancer. Endocr. Relat. Cancer 2008, 15, 1069–1074. [Google Scholar] [CrossRef]
- Caillou, B.; Talbot, M.; Weyemi, U.; Pioche-Durieu, C.; Al Ghuzlan, A.; Bidart, J.M.; Chouaib, S.; Schlumberger, M.; Dupuy, C. Tumor-associated macrophages (TAMs) form an interconnected cellular supportive network in anaplastic thyroid carcinoma. PLoS ONE 2011, 6, e22567. [Google Scholar] [CrossRef] [PubMed]
- Komohara, Y.; Jinushi, M.; Takeya, M. Clinical significance of macrophage heterogeneity in human malignant tumors. Cancer Sci. 2014, 105, 1–8. [Google Scholar] [CrossRef]
- Mantovani, A.; Sozzani, S.; Locati, M.; Allavena, P.; Sica, A. Macrophage polarization: Tumor-associated macrophages as a paradigm for polarized M2 mononuclear phagocytes. Trends Immunol. 2002, 23, 549–555. [Google Scholar] [CrossRef]
- Ryder, M.; Gild, M.; Hohl, T.M.; Pamer, E.; Knauf, J.; Ghossein, R.; Joyce, J.A.; Fagin, J.A. Genetic and pharmacological targeting of CSF-1/CSF-1R inhibits tumor-associated macrophages and impairs BRAF-induced thyroid cancer progression. PLoS ONE 2013, 8, e54302. [Google Scholar] [CrossRef]
- Ohno, S.; Suzuki, N.; Ohno, Y.; Inagawa, H.; Soma, G.I.; Inoue, M. Tumor-associated macrophages: Foe or accomplice of tumors? Anticancer Res. 2003, 23, 4395–4409. [Google Scholar] [PubMed]
- He, H.; Li, W.; Liyanarachchi, S.; Jendrzejewski, J.; Srinivas, M.; Davuluri, R.V.; Nagy, R.; de la Chapelle, A. Genetic predisposition to papillary thyroid carcinoma: Involvement of FOXE1, TSHR, and a novel lincRNA gene, PTCSC2. J. Clin. Endocrinol. Metab. 2015, 100, E164–E172. [Google Scholar] [CrossRef] [PubMed]
- Tomaz, R.A.; Sousa, I.; Silva, J.G.; Santos, C.; Teixeira, M.R.; Leite, V.; Cavaco, B.M. FOXE1 polymorphisms are associated with familial and sporadic nonmedullary thyroid cancer susceptibility. Clin. Endocrinol. 2012, 77, 926–933. [Google Scholar] [CrossRef]
- Somuncu, E.; Karatas, A.; Ferahman, S.; Saygili, N.; Yilmaz, E.; Ozturk, O.; Kapan, M. The investigation of foxe1 variations in papillary thyroid carcinoma. Int. J. Clin. Exp. Pathol. 2015, 8, 13458–13464. [Google Scholar] [PubMed]
- Takahashi, M.; Saenko, V.A.; Rogounovitch, T.I.; Kawaguchi, T.; Drozd, V.M.; Takigawa-Imamura, H.; Akulevich, N.M.; Ratanajaraya, C.; Mitsutake, N.; Takamura, N.; et al. The FOXE1 locus is a major genetic determinant for radiation-related thyroid carcinoma in Chernobyl. Hum. Mol. Genet. 2010, 19, 2516–2523. [Google Scholar] [CrossRef]
- Landa, I.; Ruiz-Llorente, S.; Montero-Conde, C.; Inglada-Perez, L.; Schiavi, F.; Leskela, S.; Pita, G.; Milne, R.; Maravall, J.; Ramos, I.; et al. The variant rs1867277 in FOXE1 gene confers thyroid cancer susceptibility through the recruitment of USF1/USF2 transcription factors. PLoS Genet. 2009, 5, e1000637. [Google Scholar] [CrossRef]
- Chakravarty, D.; Santos, E.; Ryder, M.; Knauf, J.A.; Liao, X.H.; West, B.L.; Bollag, G.; Kolesnick, R.; Thin, T.H.; Rosen, N.; et al. Small-molecule MAPK inhibitors restore radioiodine incorporation in mouse thyroid cancers with conditional BRAF activation. J. Clin. Investig. 2011, 121, 4700–4711. [Google Scholar] [CrossRef]
- Credendino, S.C.; Moccia, C.; Amendola, E.; D’Avino, G.; Di Guida, L.; Clery, E.; Greco, A.; Bellevicine, C.; Brunetti, A.; De Felice, M.; et al. Foxe1 Gene Dosage Affects thyroid Cancer Histology and Differentiation in Vivo. Int. J. Mol. Sci. 2020, 22, 25. [Google Scholar] [CrossRef]
- Ambesi-Impiombato, F.S.; Coon, H.G. Thyroid cells in culture. Int. Rev. Cytol. 1979, 10, 163–172. [Google Scholar] [CrossRef]
- De Felice, M.; Ovitt, C.; Biffali, E.; Rodriguez-Mallon, A.; Arra, C.; Anastassiadis, K.; Macchia, P.E.; Mattei, M.G.; Mariano, A.; Scholer, H.; et al. A mouse model for hereditary thyroid dysgenesis and cleft palate. Nat. Genet. 1998, 19, 395–398. [Google Scholar] [CrossRef] [PubMed]
- Morillo-Bernal, J.; Fernández, L.P.; Santisteban, P. FOXE1 regulates migration and invasion in thyroid cancer cells and targets ZEB1. Endocr. Relat. Cancer 2020, 27, 137–151. [Google Scholar] [CrossRef]
- Dobranowski, P.; Sly, L.M. SHIP negatively regulates type II immune responses in mast cells and macrophages. J. Leukoc. Biol. 2018, 103, 1053–1064. [Google Scholar] [CrossRef]
- Loimaranta, V.; Hepojoki, J.; Laaksoaho, O.; Pulliainen, A.T. Galectin-3-binding protein: A multitask glycoprotein with innate immunity functions in viral and bacterial infections. J. Leukoc. Biol. 2018, 104, 777–786. [Google Scholar] [CrossRef] [PubMed]
- Pauls, S.D.; Marshall, A.J. Regulation of immune cell signaling by SHIP1: A phosphatase, scaffold protein, and potential therapeutic target. Eur. J. Immunol. 2017, 47, 932–945. [Google Scholar] [CrossRef]
- MacKinnon, A.C.; Farnworth, S.L.; Hodkinson, P.S.; Henderson, N.C.; Atkinson, K.M.; Leffler, H.; Nilsson, U.J.; Haslett, C.; Forbes, S.J.; Sethi, T. Regulation of alternative macrophage activation by galectin-3. J. Immunol. 2008, 180, 2650–2658. [Google Scholar] [CrossRef]
- Ohsawa, K.; Imai, Y.; Kanazawa, H.; Sasaki, Y.; Kohsaka, S. Involvement of Iba1 in membrane ruffling and phagocytosis of macrophages/microglia. J. Cell Sci. 2000, 113 Pt 17, 3073–3084. [Google Scholar] [CrossRef]
- Sano, H.; Hsu, D.K.; Apgar, J.R.; Yu, L.; Sharma, B.B.; Kuwabara, I.; Izui, S.; Liu, F.T. Critical role of galectin-3 in phagocytosis by macrophages. J. Clin. Investig. 2003, 112, 389–397. [Google Scholar] [CrossRef]
- Chung, H.S.; Lee, B.S.; Ma, J.Y. Ethanol Extract of Mylabris phalerata Inhibits M2 Polarization Induced by Recombinant IL-4 and IL-13 in Murine Macrophages. Evid. Based Complementary Altern. Med. Ecam 2017, 2017, 4218468. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Choksi, S.; Chen, K.; Pobezinskaya, Y.; Linnoila, I.; Liu, Z.G. ROS play a critical role in the differentiation of alternatively activated macrophages and the occurrence of tumor-associated macrophages. Cell Res. 2013, 23, 898–914. [Google Scholar] [CrossRef]
- Fernandes, S.; Iyer, S.; Kerr, W.G. Role of SHIP1 in cancer and mucosal inflammation. Ann. N. Y. Acad. Sci. 2013, 1280, 6–10. [Google Scholar] [CrossRef] [PubMed]
- Woodman, N.; Pinder, S.E.; Tajadura, V.; Le Bourhis, X.; Gillett, C.; Delannoy, P.; Burchell, J.M.; Julien, S. Two E-selectin ligands, BST-2 and LGALS3BP, predict metastasis and poor survival of ER-negative breast cancer. Int. J. Oncol. 2016, 49, 265–275. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Ding, H.; Lu, Z.; Ding, L.; Song, Y.; Jing, Y.; Hu, Q.; Dong, Y.; Ni, Y. Increased LGALS3BP promotes proliferation and migration of oral squamous cell carcinoma via PI3K/AKT pathway. Cell. Signal. 2019, 63, 109359. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; He, W.; Huang, J.; Wang, B.; Li, H.; Cai, Q.; Su, F.; Bi, J.; Liu, H.; Zhang, B.; et al. LNMAT1 promotes lymphatic metastasis of bladder cancer via CCL2 dependent macrophage recruitment. Nat. Commun. 2018, 9, 3826. [Google Scholar] [CrossRef] [PubMed]
- Richardsen, E.; Uglehus, R.D.; Johnsen, S.H.; Busund, L.T. Macrophage-colony stimulating factor (CSF1) predicts breast cancer progression and mortality. Anticancer Res. 2015, 35, 865–874. [Google Scholar]
- Evrard, D.; Szturz, P.; Tijeras-Raballand, A.; Astorgues-Xerri, L.; Abitbol, C.; Paradis, V.; Raymond, E.; Albert, S.; Barry, B.; Faivre, S. Macrophages in the microenvironment of head and neck cancer: Potential targets for cancer therapy. Oral Oncol. 2019, 88, 29–38. [Google Scholar] [CrossRef]
- Li, X.; Yao, W.; Yuan, Y.; Chen, P.; Li, B.; Li, J.; Chu, R.; Song, H.; Xie, D.; Jiang, X.; et al. Targeting of tumour-infiltrating macrophages via CCL2/CCR2 signalling as a therapeutic strategy against hepatocellular carcinoma. Gut 2017, 66, 157–167. [Google Scholar] [CrossRef]
- Pyonteck, S.M.; Akkari, L.; Schuhmacher, A.J.; Bowman, R.L.; Sevenich, L.; Quail, D.F.; Olson, O.C.; Quick, M.L.; Huse, J.T.; Teijeiro, V.; et al. CSF-1R inhibition alters macrophage polarization and blocks glioma progression. Nat. Med. 2013, 19, 1264–1272. [Google Scholar] [CrossRef]
- Varricchi, G.; Loffredo, S.; Marone, G.; Modestino, L.; Fallahi, P.; Ferrari, S.M.; de Paulis, A.; Antonelli, A.; Galdiero, M.R. The Immune Landscape of Thyroid Cancer in the Context of Immune Checkpoint Inhibitin. Int. J. Mol. Sci. 2019, 20, 3934. [Google Scholar] [CrossRef]
- Gascard, P.; Tlsty, T.D. Carcinoma-associated fibroblasts: Orchestrating the composition of malignancy. Genes Dev. 2016, 30, 1002–1019. [Google Scholar] [CrossRef]
- Shiga, K.; Hara, M.; Nagasaki, T.; Sato, T.; Takahashi, H.; Takeyama, H. Cancer-Associated Fibroblasts: Their Characteristics and Their Roles in Tumor Growth. Cancers 2015, 7, 2443–2458. [Google Scholar] [CrossRef]
- Comito, G.; Giannoni, E.; Segura, C.P.; Barcellos-de-Souza, P.; Raspollini, M.R.; Baroni, G.; Lanciotti, M.; Serni, S.; Chiarugi, P. Cancer-associated fibroblasts and M2-polarized macrophages synergize during prostate carcinoma progression. Oncogene 2014, 33, 2423–2431. [Google Scholar] [CrossRef] [PubMed]
- Perrone, L.; Pasca di Magliano, M.; Zannini, M.; Di Lauro, R. The thyroid transcription factor 2 (TTF-2) is a promoter-specific DNA-binding independent transcriptional repressor. Biochem. Biophys. Res. Commun. 2000, 275, 203–208. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Credendino, S.C.; De Menna, M.; Cantone, I.; Moccia, C.; Esposito, M.; Di Guida, L.; De Felice, M.; De Vita, G. FOXE1-Dependent Regulation of Macrophage Chemotaxis by Thyroid Cells In Vitro and In Vivo. Int. J. Mol. Sci. 2021, 22, 7666. https://doi.org/10.3390/ijms22147666
Credendino SC, De Menna M, Cantone I, Moccia C, Esposito M, Di Guida L, De Felice M, De Vita G. FOXE1-Dependent Regulation of Macrophage Chemotaxis by Thyroid Cells In Vitro and In Vivo. International Journal of Molecular Sciences. 2021; 22(14):7666. https://doi.org/10.3390/ijms22147666
Chicago/Turabian StyleCredendino, Sara C., Marta De Menna, Irene Cantone, Carmen Moccia, Matteo Esposito, Luigi Di Guida, Mario De Felice, and Gabriella De Vita. 2021. "FOXE1-Dependent Regulation of Macrophage Chemotaxis by Thyroid Cells In Vitro and In Vivo" International Journal of Molecular Sciences 22, no. 14: 7666. https://doi.org/10.3390/ijms22147666
APA StyleCredendino, S. C., De Menna, M., Cantone, I., Moccia, C., Esposito, M., Di Guida, L., De Felice, M., & De Vita, G. (2021). FOXE1-Dependent Regulation of Macrophage Chemotaxis by Thyroid Cells In Vitro and In Vivo. International Journal of Molecular Sciences, 22(14), 7666. https://doi.org/10.3390/ijms22147666





